Please be patient as the page loads
|
LYRIC_MOUSE
|
||||||
| SwissProt Accessions | Q80WJ7, Q3U9F8, Q3UAQ8, Q8BN67, Q8CBT9, Q8CDL0, Q8CGI7, Q9D052 | Gene names | Mtdh, Lyric | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LYRIC (Lysine-rich CEACAM1 co-isolated protein) (3D3/LYRIC) (Metastasis adhesion protein) (Metadherin). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
LYRIC_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.943042 (rank : 2) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q86UE4, Q6PK07, Q8TCX3 | Gene names | MTDH, AEG1, LYRIC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LYRIC (Lysine-rich CEACAM1 co-isolated protein) (3D3/lyric) (Metastasis adhesion protein) (Metadherin) (Astrocyte elevated gene-1 protein) (AEG-1). | |||||
|
LYRIC_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 126 | |
| SwissProt Accessions | Q80WJ7, Q3U9F8, Q3UAQ8, Q8BN67, Q8CBT9, Q8CDL0, Q8CGI7, Q9D052 | Gene names | Mtdh, Lyric | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LYRIC (Lysine-rich CEACAM1 co-isolated protein) (3D3/LYRIC) (Metastasis adhesion protein) (Metadherin). | |||||
|
DSPP_HUMAN
|
||||||
| θ value | 6.43352e-06 (rank : 3) | NC score | 0.179111 (rank : 3) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9NZW4, O95815 | Gene names | DSPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
NIPBL_HUMAN
|
||||||
| θ value | 1.43324e-05 (rank : 4) | NC score | 0.166847 (rank : 6) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
NPBL_MOUSE
|
||||||
| θ value | 0.000461057 (rank : 5) | NC score | 0.163972 (rank : 8) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
AFF4_MOUSE
|
||||||
| θ value | 0.00175202 (rank : 6) | NC score | 0.169258 (rank : 5) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9ESC8, Q8C6K4, Q8C6W3, Q8CCH3 | Gene names | Aff4, Alf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4. | |||||
|
DSPP_MOUSE
|
||||||
| θ value | 0.00175202 (rank : 7) | NC score | 0.171916 (rank : 4) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P97399, O70567 | Gene names | Dspp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor (Dentin matrix protein 3) (DMP-3) [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
LAD1_HUMAN
|
||||||
| θ value | 0.00869519 (rank : 8) | NC score | 0.122070 (rank : 9) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 441 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O00515, O95614 | Gene names | LAD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ladinin 1 (Lad-1) (120 kDa linear IgA bullous dermatosis antigen) (97 kDa linear IgA bullous dermatosis antigen) (Linear IgA disease antigen homolog) (LadA). | |||||
|
AFF4_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 9) | NC score | 0.165554 (rank : 7) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 611 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9UHB7, Q498B2, Q59FB3, Q6P592, Q8TDR1, Q9P0E4 | Gene names | AFF4, AF5Q31, MCEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4 (ALL1-fused gene from chromosome 5q31) (Major CDK9 elongation factor-associated protein). | |||||
|
SELPL_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 10) | NC score | 0.104002 (rank : 13) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 588 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q62170 | Gene names | Selplg, Selp1, Selpl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | P-selectin glycoprotein ligand 1 precursor (PSGL-1) (Selectin P ligand). | |||||
|
SMRC2_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 11) | NC score | 0.098053 (rank : 18) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 487 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q6PDG5, Q6P6P3 | Gene names | Smarcc2, Baf170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 2 (SWI/SNF complex 170 kDa subunit) (BRG1-associated factor 170). | |||||
|
SMRC2_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 12) | NC score | 0.101281 (rank : 16) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 499 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8TAQ2, Q92923, Q96E12, Q96GY4 | Gene names | SMARCC2, BAF170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 2 (SWI/SNF complex 170 kDa subunit) (BRG1-associated factor 170). | |||||
|
ACINU_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 13) | NC score | 0.102428 (rank : 15) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q9UKV3, O75158, Q9UG91, Q9UKV1, Q9UKV2 | Gene names | ACIN1, ACINUS, KIAA0670 | |||
|
Domain Architecture |

|
|||||
| Description | Apoptotic chromatin condensation inducer in the nucleus (Acinus). | |||||
|
DPOLZ_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 14) | NC score | 0.082754 (rank : 34) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O60673, O43214 | Gene names | REV3L, POLZ, REV3 | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase zeta catalytic subunit (EC 2.7.7.7) (hREV3). | |||||
|
BPTF_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 15) | NC score | 0.082957 (rank : 33) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 786 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q12830, Q6NX67, Q7Z7D6, Q9UIG2 | Gene names | FALZ, BPTF, FAC1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleosome remodeling factor subunit BPTF (Bromodomain and PHD finger- containing transcription factor) (Fetal Alzheimer antigen) (Fetal Alz- 50 clone 1 protein). | |||||
|
LAP2_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 16) | NC score | 0.033111 (rank : 140) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 482 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q80TH2, Q8BQ14, Q8CE41, Q8K171, Q99JU3, Q9JI47 | Gene names | Erbb2ip, Erbin, Kiaa1225, Lap2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LAP2 (Erbb2-interacting protein) (Erbin) (Densin-180-like protein). | |||||
|
DMP1_MOUSE
|
||||||
| θ value | 0.125558 (rank : 17) | NC score | 0.118337 (rank : 10) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O55188 | Gene names | Dmp1, Dmp | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1) (AG1). | |||||
|
GP73_MOUSE
|
||||||
| θ value | 0.125558 (rank : 18) | NC score | 0.054683 (rank : 93) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q91XA2 | Gene names | Golph2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi phosphoprotein 2 (Golgi membrane protein GP73). | |||||
|
R51A1_MOUSE
|
||||||
| θ value | 0.125558 (rank : 19) | NC score | 0.109374 (rank : 12) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 164 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8C551, O55219, Q8BP36, Q99L94, Q9D0J0 | Gene names | Rad51ap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RAD51-associated protein 1 (RAD51-interacting protein) (RAB22). | |||||
|
ASXL1_MOUSE
|
||||||
| θ value | 0.21417 (rank : 20) | NC score | 0.068640 (rank : 53) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 182 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P59598 | Gene names | Asxl1, Kiaa0978 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative Polycomb group protein ASXL1 (Additional sex combs-like protein 1). | |||||
|
RPGR_HUMAN
|
||||||
| θ value | 0.21417 (rank : 21) | NC score | 0.063602 (rank : 60) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q92834, O00702, O00737, Q8N5T6, Q93039, Q9HD29, Q9UMR1 | Gene names | RPGR, RP3, XLRP3 | |||
|
Domain Architecture |

|
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator. | |||||
|
SETD8_HUMAN
|
||||||
| θ value | 0.21417 (rank : 22) | NC score | 0.091636 (rank : 22) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9NQR1, Q86W83, Q8TD09 | Gene names | SETD8, PRSET7, SET07, SET8 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H4 lysine-20 specific (EC 2.1.1.43) (Histone H4-K20 methyltransferase) (H4-K20-HMTase) (SET domain-containing lysine methyltransferase 8) (SET domain-containing protein 8) (PR/SET domain-containing protein 07) (PR/SET07) (PR-Set7). | |||||
|
UBP31_HUMAN
|
||||||
| θ value | 0.21417 (rank : 23) | NC score | 0.051068 (rank : 112) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 293 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q70CQ4, Q6AW97, Q6ZTC0, Q6ZTN2, Q9ULL7 | Gene names | USP31, KIAA1203 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 31 (EC 3.1.2.15) (Ubiquitin thioesterase 31) (Ubiquitin-specific-processing protease 31) (Deubiquitinating enzyme 31). | |||||
|
LAP2_HUMAN
|
||||||
| θ value | 0.279714 (rank : 24) | NC score | 0.034349 (rank : 139) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 499 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q96RT1, Q86W38, Q9NR18, Q9NW48, Q9ULJ5 | Gene names | ERBB2IP, ERBIN, KIAA1225, LAP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LAP2 (Erbb2-interacting protein) (Erbin) (Densin-180-like protein). | |||||
|
MDC1_HUMAN
|
||||||
| θ value | 0.279714 (rank : 25) | NC score | 0.085463 (rank : 31) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
SH24A_HUMAN
|
||||||
| θ value | 0.279714 (rank : 26) | NC score | 0.046723 (rank : 121) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9H788, Q5XKC1, Q6NXE9, Q86YM2, Q96C88, Q9H7F7 | Gene names | SH2D4A, SH2A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH2 domain-containing protein 4A (Protein SH(2)A). | |||||
|
CEP57_MOUSE
|
||||||
| θ value | 0.365318 (rank : 27) | NC score | 0.032892 (rank : 141) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 580 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8CEE0, Q6ZQJ3, Q7TN18, Q80X65, Q810F2, Q9D4J4, Q9D5S4, Q9D5W5 | Gene names | Cep57, Kiaa0092, Tsp57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 57 kDa (Cep57 protein) (Testis-specific protein 57) (Translokin). | |||||
|
PALB2_MOUSE
|
||||||
| θ value | 0.365318 (rank : 28) | NC score | 0.053024 (rank : 103) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q3U0P1, Q6NZG9, Q7TMQ4, Q8CEA9 | Gene names | Palb2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Partner and localizer of BRCA2. | |||||
|
ADDB_MOUSE
|
||||||
| θ value | 0.47712 (rank : 29) | NC score | 0.073618 (rank : 42) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 574 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9QYB8, Q3U0E1, Q80VH9, Q8C1C4, Q9CXE3, Q9JLE5, Q9QYB7 | Gene names | Add2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-adducin (Erythrocyte adducin subunit beta) (Add97). | |||||
|
CJ120_HUMAN
|
||||||
| θ value | 0.47712 (rank : 30) | NC score | 0.098340 (rank : 17) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5SQS8 | Gene names | C10orf120 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative uncharacterized protein C10orf120. | |||||
|
EDD1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 31) | NC score | 0.054289 (rank : 95) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O95071, O94970, Q9NPL3 | Gene names | EDD1, EDD, HYD, KIAA0896 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-protein ligase EDD1 (EC 6.3.2.-) (Hyperplastic discs protein homolog) (hHYD) (Progestin-induced protein). | |||||
|
NACAM_MOUSE
|
||||||
| θ value | 0.47712 (rank : 32) | NC score | 0.065808 (rank : 55) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
NFH_HUMAN
|
||||||
| θ value | 0.47712 (rank : 33) | NC score | 0.056826 (rank : 83) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1242 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | P12036, Q9UJS7, Q9UQ14 | Gene names | NEFH, KIAA0845, NFH | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
SETD8_MOUSE
|
||||||
| θ value | 0.47712 (rank : 34) | NC score | 0.092372 (rank : 21) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q2YDW7, Q8C0J9 | Gene names | SETD8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H4 lysine-20 specific (EC 2.1.1.43) (Histone H4-K20 methyltransferase) (H4-K20-HMTase) (SET domain-containing lysine methyltransferase 8) (SET domain-containing protein 8). | |||||
|
TRIPC_HUMAN
|
||||||
| θ value | 0.47712 (rank : 35) | NC score | 0.052313 (rank : 107) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 319 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q14669, Q15644 | Gene names | TRIP12, KIAA0045 | |||
|
Domain Architecture |

|
|||||
| Description | Thyroid receptor-interacting protein 12 (TRIP12). | |||||
|
ARFG1_MOUSE
|
||||||
| θ value | 0.62314 (rank : 36) | NC score | 0.060204 (rank : 71) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9EPJ9, Q3TI52, Q8BMM6, Q8BMQ7 | Gene names | Arfgap1, Arf1gap | |||
|
Domain Architecture |
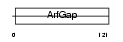
|
|||||
| Description | ADP-ribosylation factor GTPase-activating protein 1 (ADP-ribosylation factor 1 GTPase-activating protein) (ARF1 GAP) (ARF1-directed GTPase- activating protein) (GAP protein). | |||||
|
CI039_HUMAN
|
||||||
| θ value | 0.62314 (rank : 37) | NC score | 0.023328 (rank : 154) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1171 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9NXG0, Q5VYJ0, Q9HAJ5 | Gene names | C9orf39 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C9orf39. | |||||
|
ENAH_MOUSE
|
||||||
| θ value | 0.62314 (rank : 38) | NC score | 0.043347 (rank : 129) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 919 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q03173, P70430, P70431, P70432, P70433, Q5D053 | Gene names | Enah, Mena, Ndpp1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein enabled homolog (NPC-derived proline-rich protein 1) (NDPP-1). | |||||
|
BAZ2A_MOUSE
|
||||||
| θ value | 0.813845 (rank : 39) | NC score | 0.047709 (rank : 119) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q91YE5 | Gene names | Baz2a, Tip5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5). | |||||
|
EDD1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 40) | NC score | 0.052657 (rank : 104) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q80TP3, Q698K9, Q6PEQ8, Q6PFQ9, Q80VL4, Q810V6, Q9CXE9 | Gene names | Edd1, Edd, Kiaa0896 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase EDD1 (EC 6.3.2.-) (Hyperplastic discs protein homolog). | |||||
|
MAP1B_MOUSE
|
||||||
| θ value | 0.813845 (rank : 41) | NC score | 0.103190 (rank : 14) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 977 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | P14873 | Gene names | Map1b, Mtap1b, Mtap5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) (MAP1.2) (MAP1(X)) [Contains: MAP1 light chain LC1]. | |||||
|
MTMR7_HUMAN
|
||||||
| θ value | 0.813845 (rank : 42) | NC score | 0.051575 (rank : 108) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Y216, Q68DX4 | Gene names | MTMR7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myotubularin-related protein 7 (EC 3.1.3.-). | |||||
|
NOLC1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 43) | NC score | 0.114400 (rank : 11) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q14978, Q15030 | Gene names | NOLC1, KIAA0035 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar phosphoprotein p130 (Nucleolar 130 kDa protein) (140 kDa nucleolar phosphoprotein) (Nopp140) (Nucleolar and coiled-body phosphoprotein 1). | |||||
|
SYLC_MOUSE
|
||||||
| θ value | 0.813845 (rank : 44) | NC score | 0.055700 (rank : 87) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BMJ2, Q8BKW9 | Gene names | Lars | |||
|
Domain Architecture |

|
|||||
| Description | Leucyl-tRNA synthetase, cytoplasmic (EC 6.1.1.4) (Leucine--tRNA ligase) (LeuRS). | |||||
|
AFF3_HUMAN
|
||||||
| θ value | 1.06291 (rank : 45) | NC score | 0.096959 (rank : 19) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P51826 | Gene names | AFF3, LAF4 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 3 (Protein LAF-4) (Lymphoid nuclear protein related to AF4). | |||||
|
BRD8_HUMAN
|
||||||
| θ value | 1.06291 (rank : 46) | NC score | 0.059948 (rank : 74) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9H0E9, O43178, Q15355, Q59GN0, Q969M9 | Gene names | BRD8, SMAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 8 (p120) (Skeletal muscle abundant protein) (Thyroid hormone receptor coactivating protein 120kDa) (TrCP120). | |||||
|
BRD8_MOUSE
|
||||||
| θ value | 1.06291 (rank : 47) | NC score | 0.070079 (rank : 48) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 422 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8R3B7, Q8C049, Q8R583, Q8VDP0 | Gene names | Brd8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 8. | |||||
|
PRG4_HUMAN
|
||||||
| θ value | 1.06291 (rank : 48) | NC score | 0.076259 (rank : 41) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q92954, Q6DNC4, Q6DNC5, Q6ZMZ5, Q9BX49 | Gene names | PRG4, MSF, SZP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
AP3B2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 49) | NC score | 0.046616 (rank : 122) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q13367, O14808 | Gene names | AP3B2 | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit beta-2 (Adapter-related protein complex 3 beta-2 subunit) (Beta3B-adaptin) (Adaptor protein complex AP-3 beta-2 subunit) (AP-3 complex beta-2 subunit) (Clathrin assembly protein complex 3 beta-2 large chain) (Neuron-specific vesicle coat protein beta-NAP). | |||||
|
CSPG2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 50) | NC score | 0.019370 (rank : 159) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 711 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q62059, Q62058, Q9CUU0 | Gene names | Cspg2 | |||
|
Domain Architecture |

|
|||||
| Description | Versican core protein precursor (Large fibroblast proteoglycan) (Chondroitin sulfate proteoglycan core protein 2) (PG-M). | |||||
|
NOP14_HUMAN
|
||||||
| θ value | 1.38821 (rank : 51) | NC score | 0.051180 (rank : 111) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 573 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P78316, Q7LGI5, Q7Z6K0, Q8TBR6 | Gene names | NOP14, C4orf9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable nucleolar complex protein 14. | |||||
|
PCLO_HUMAN
|
||||||
| θ value | 1.38821 (rank : 52) | NC score | 0.059203 (rank : 75) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
ZMYM5_HUMAN
|
||||||
| θ value | 1.38821 (rank : 53) | NC score | 0.040364 (rank : 133) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UJ78, Q5T6E2, Q96IY6, Q9NZY5, Q9UBW0, Q9UJ77 | Gene names | ZMYM5, ZNF198L1, ZNF237 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger MYM-type protein 5 (Zinc finger protein 237) (Zinc finger protein 198-like 1). | |||||
|
ARI5B_HUMAN
|
||||||
| θ value | 1.81305 (rank : 54) | NC score | 0.060535 (rank : 68) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q14865, Q7Z3M4, Q8N421, Q9H786 | Gene names | ARID5B, DESRT, MRF2 | |||
|
Domain Architecture |
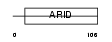
|
|||||
| Description | AT-rich interactive domain-containing protein 5B (ARID domain- containing protein 5B) (Mrf1-like) (Modulator recognition factor 2) (MRF-2). | |||||
|
CDC6_MOUSE
|
||||||
| θ value | 1.81305 (rank : 55) | NC score | 0.048244 (rank : 118) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O89033, Q3TMD0, Q8C3S5, Q9CZT0 | Gene names | Cdc6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division control protein 6 homolog (CDC6-related protein) (p62(cdc6)). | |||||
|
MADCA_HUMAN
|
||||||
| θ value | 1.81305 (rank : 56) | NC score | 0.057529 (rank : 81) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q13477, O60222, O75867 | Gene names | MADCAM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucosal addressin cell adhesion molecule 1 precursor (MAdCAM-1) (hMAdCAM-1). | |||||
|
MYLK_MOUSE
|
||||||
| θ value | 1.81305 (rank : 57) | NC score | 0.011291 (rank : 169) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1489 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q6PDN3, Q3TSJ7, Q80UX0, Q80YN7, Q80YN8, Q8K026, Q924D2, Q9ERD3 | Gene names | Mylk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin light chain kinase, smooth muscle (EC 2.7.11.18) (MLCK) (Telokin) (Kinase-related protein) (KRP). | |||||
|
PHF14_MOUSE
|
||||||
| θ value | 1.81305 (rank : 58) | NC score | 0.060053 (rank : 72) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9D4H9, Q810Z6, Q8C284, Q8CGF8, Q9D1Q8 | Gene names | Phf14 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 14. | |||||
|
SEMG1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 59) | NC score | 0.044009 (rank : 126) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P04279, Q6X4I9, Q6Y809, Q6Y822, Q6Y823, Q86U64, Q96QM3 | Gene names | SEMG1, SEMG | |||
|
Domain Architecture |

|
|||||
| Description | Semenogelin-1 precursor (Semenogelin I) (SGI) [Contains: Alpha- inhibin-92; Alpha-inhibin-31; Seminal basic protein]. | |||||
|
YTHD3_HUMAN
|
||||||
| θ value | 1.81305 (rank : 60) | NC score | 0.026719 (rank : 148) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q7Z739, Q63Z37, Q659A3 | Gene names | YTHDF3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YTH domain family protein 3. | |||||
|
ARX_MOUSE
|
||||||
| θ value | 2.36792 (rank : 61) | NC score | 0.007476 (rank : 178) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 398 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O35085, Q9QYT4 | Gene names | Arx | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein ARX (Aristaless-related homeobox). | |||||
|
CDC6_HUMAN
|
||||||
| θ value | 2.36792 (rank : 62) | NC score | 0.043701 (rank : 127) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99741, Q8TB30 | Gene names | CDC6, CDC18L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division control protein 6 homolog (CDC6-related protein) (p62(cdc6)) (HsCDC6) (HsCDC18). | |||||
|
FA44A_HUMAN
|
||||||
| θ value | 2.36792 (rank : 63) | NC score | 0.060019 (rank : 73) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8NFC6, Q6P0M8 | Gene names | FAM44A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM44A. | |||||
|
MTMR7_MOUSE
|
||||||
| θ value | 2.36792 (rank : 64) | NC score | 0.046770 (rank : 120) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9Z2C9, Q8C4J6 | Gene names | Mtmr7 | |||
|
Domain Architecture |
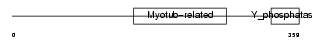
|
|||||
| Description | Myotubularin-related protein 7 (EC 3.1.3.-). | |||||
|
NEB1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 65) | NC score | 0.030260 (rank : 143) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 829 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9ULJ8, O76059, Q9NXT2 | Gene names | PPP1R9A, KIAA1222 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neurabin-1 (Neurabin-I) (Neural tissue-specific F-actin-binding protein I) (Protein phosphatase 1 regulatory subunit 9A). | |||||
|
PHF2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 66) | NC score | 0.026967 (rank : 147) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O75151, Q8N3K2, Q9Y6N4 | Gene names | PHF2, KIAA0662 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 2 (GRC5). | |||||
|
PK1IP_MOUSE
|
||||||
| θ value | 2.36792 (rank : 67) | NC score | 0.023803 (rank : 153) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9DCE5, Q3UBK7, Q80UT4, Q8C5N6, Q923K2, Q9D3E9 | Gene names | Pak1ip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | p21-activated protein kinase-interacting protein 1 (PAK1-interacting protein 1) (Putative PAK inhibitor Skb15). | |||||
|
REV1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 68) | NC score | 0.026528 (rank : 149) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q920Q2, Q9QXV2 | Gene names | Rev1l, Rev1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein REV1 (EC 2.7.7.-) (Rev1-like terminal deoxycytidyl transferase). | |||||
|
SCN5A_HUMAN
|
||||||
| θ value | 2.36792 (rank : 69) | NC score | 0.010768 (rank : 170) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q14524 | Gene names | SCN5A | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 5 subunit alpha (Sodium channel protein type V subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.5) (Sodium channel protein, cardiac muscle alpha-subunit) (HH1). | |||||
|
SETX_HUMAN
|
||||||
| θ value | 2.36792 (rank : 70) | NC score | 0.046530 (rank : 123) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q7Z333, O75120, Q3KQX4, Q68DW5, Q6AZD7, Q7Z3J6, Q8WX33, Q9H9D1, Q9NVP9 | Gene names | SETX, ALS4, KIAA0625, SCAR1 | |||
|
Domain Architecture |

|
|||||
| Description | Probable helicase senataxin (EC 3.6.1.-) (SEN1 homolog). | |||||
|
SMRCD_HUMAN
|
||||||
| θ value | 2.36792 (rank : 71) | NC score | 0.016859 (rank : 160) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9H4L7, Q96SX1, Q9H017, Q9H860, Q9NPU9, Q9ULU7 | Gene names | SMARCAD1, KIAA1122 | |||
|
Domain Architecture |

|
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A containing DEAD/H box 1 (EC 3.6.1.-) (ATP- dependent helicase 1) (hHEL1). | |||||
|
TNC6B_HUMAN
|
||||||
| θ value | 2.36792 (rank : 72) | NC score | 0.091592 (rank : 23) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 256 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9UPQ9, Q5TH52, Q8TBX2 | Gene names | TNRC6B, KIAA1093 | |||
|
Domain Architecture |

|
|||||
| Description | Trinucleotide repeat-containing 6B protein. | |||||
|
ANKR6_HUMAN
|
||||||
| θ value | 3.0926 (rank : 73) | NC score | 0.008544 (rank : 174) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 439 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Y2G4, Q9NU24, Q9UFQ9 | Gene names | ANKRD6, KIAA0957 | |||
|
Domain Architecture |
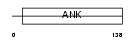
|
|||||
| Description | Ankyrin repeat domain-containing protein 6. | |||||
|
ARHG7_MOUSE
|
||||||
| θ value | 3.0926 (rank : 74) | NC score | 0.012216 (rank : 168) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 396 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9ES28, O08757, Q9ES27 | Gene names | Arhgef7, Pak3bp | |||
|
Domain Architecture |
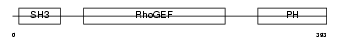
|
|||||
| Description | Rho guanine nucleotide exchange factor 7 (PAK-interacting exchange factor beta) (Beta-Pix) (p85SPR). | |||||
|
MINT_MOUSE
|
||||||
| θ value | 3.0926 (rank : 75) | NC score | 0.063261 (rank : 61) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
MLL3_MOUSE
|
||||||
| θ value | 3.0926 (rank : 76) | NC score | 0.050251 (rank : 116) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
NASP_HUMAN
|
||||||
| θ value | 3.0926 (rank : 77) | NC score | 0.080460 (rank : 37) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P49321, Q96A69, Q9BTW2 | Gene names | NASP | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear autoantigenic sperm protein (NASP). | |||||
|
PRG4_MOUSE
|
||||||
| θ value | 3.0926 (rank : 78) | NC score | 0.078016 (rank : 38) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 624 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9JM99, Q3UEL1, Q3V198 | Gene names | Prg4, Msf, Szp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
WDHD1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 79) | NC score | 0.042374 (rank : 130) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 414 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P59328, Q6P408 | Gene names | Wdhd1, And1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat and HMG-box DNA-binding protein 1 (Acidic nucleoplasmic DNA- binding protein 1) (And-1). | |||||
|
PHF14_HUMAN
|
||||||
| θ value | 4.03905 (rank : 80) | NC score | 0.063019 (rank : 63) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O94880 | Gene names | PHF14, KIAA0783 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 14. | |||||
|
PKHA6_MOUSE
|
||||||
| θ value | 4.03905 (rank : 81) | NC score | 0.029805 (rank : 144) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 591 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q7TQG1, Q8K0J5 | Gene names | Plekha6, Pepp3 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 6 (Phosphoinositol 3-phosphate-binding protein 3) (PEPP-3). | |||||
|
RAD18_MOUSE
|
||||||
| θ value | 4.03905 (rank : 82) | NC score | 0.037503 (rank : 136) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9QXK2, Q9CZB8 | Gene names | Rad18, Rad18sc | |||
|
Domain Architecture |

|
|||||
| Description | Postreplication repair protein RAD18 (mRAD18Sc). | |||||
|
RFC1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 83) | NC score | 0.046350 (rank : 124) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P35251, Q5XKF5, Q6PKU0, Q86V41, Q86V46 | Gene names | RFC1, RFC140 | |||
|
Domain Architecture |

|
|||||
| Description | Replication factor C subunit 1 (Replication factor C large subunit) (RF-C 140 kDa subunit) (Activator 1 140 kDa subunit) (Activator 1 large subunit) (A1 140 kDa subunit) (DNA-binding protein PO-GA). | |||||
|
SMC2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 84) | NC score | 0.015160 (rank : 164) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1131 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8CG48, Q61076, Q9CS17, Q9CSD8 | Gene names | Smc2l1, Cape, Fin16, Smc2 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 2-like 1 protein (Chromosome- associated protein E) (XCAP-E homolog) (FGF-inducible protein 16). | |||||
|
TTC16_MOUSE
|
||||||
| θ value | 4.03905 (rank : 85) | NC score | 0.024040 (rank : 152) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8C1F5 | Gene names | Ttc16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tetratricopeptide repeat protein 16 (TPR repeat protein 16). | |||||
|
DEK_HUMAN
|
||||||
| θ value | 5.27518 (rank : 86) | NC score | 0.061936 (rank : 66) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P35659, Q5TGV4, Q5TGV5 | Gene names | DEK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein DEK. | |||||
|
ELOA1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 87) | NC score | 0.049370 (rank : 117) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q14241, Q8IXH1 | Gene names | TCEB3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription elongation factor B polypeptide 3 (RNA polymerase II transcription factor SIII subunit A1) (SIII p110) (Elongin-A) (EloA) (Elongin 110 kDa subunit). | |||||
|
EMIL2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 88) | NC score | 0.021105 (rank : 156) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 485 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8K482 | Gene names | Emilin2 | |||
|
Domain Architecture |

|
|||||
| Description | EMILIN-2 precursor (Elastin microfibril interface-located protein 2) (Elastin microfibril interfacer 2) (Basilin). | |||||
|
FBLN2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 89) | NC score | 0.007573 (rank : 177) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 505 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P37889, Q9WUI2 | Gene names | Fbln2 | |||
|
Domain Architecture |

|
|||||
| Description | Fibulin-2 precursor. | |||||
|
GASP2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 90) | NC score | 0.030421 (rank : 142) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8BUY8 | Gene names | Gprasp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G-protein coupled receptor-associated sorting protein 2 (GASP-2). | |||||
|
K0586_HUMAN
|
||||||
| θ value | 5.27518 (rank : 91) | NC score | 0.029131 (rank : 145) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9BVV6, O60328 | Gene names | KIAA0586 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0586. | |||||
|
LAD1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 92) | NC score | 0.091039 (rank : 25) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 391 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P57016 | Gene names | Lad1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ladinin 1 (Lad-1) (Linear IgA disease autoantigen). | |||||
|
LEO1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 93) | NC score | 0.092774 (rank : 20) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
MBB1A_HUMAN
|
||||||
| θ value | 5.27518 (rank : 94) | NC score | 0.061115 (rank : 67) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 665 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9BQG0, Q86VM3, Q9BW49, Q9P0V5, Q9UF99 | Gene names | MYBBP1A, P160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myb-binding protein 1A. | |||||
|
NFH_MOUSE
|
||||||
| θ value | 5.27518 (rank : 95) | NC score | 0.053134 (rank : 102) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
PARP1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 96) | NC score | 0.016595 (rank : 161) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P11103, Q9JLX4, Q9QVQ3 | Gene names | Parp1, Adprp, Adprt, Adprt1 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 1 (EC 2.4.2.30) (PARP-1) (ADPRT) (NAD(+) ADP-ribosyltransferase 1) (Poly[ADP-ribose] synthetase 1) (msPARP). | |||||
|
PCLO_MOUSE
|
||||||
| θ value | 5.27518 (rank : 97) | NC score | 0.056212 (rank : 84) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
TSH2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 98) | NC score | 0.026343 (rank : 150) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9NRE2, Q4VXM4, Q6N003, Q8N260 | Gene names | TSHZ2, C20orf17, TSH2, ZNF218 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Teashirt homolog 2 (Zinc finger protein 218) (Ovarian cancer-related protein 10-2) (OVC10-2). | |||||
|
TSH2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 99) | NC score | 0.036619 (rank : 137) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q68FE9, Q5DTF1, Q6AXH5, Q9JL71 | Gene names | Tshz2, Kiaa4248, Sdccag33l, Tsh2, Zfp218, Znf218 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Teashirt homolog 2 (Zinc finger protein 218) (SDCCAG33-like protein). | |||||
|
YSK4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 100) | NC score | 0.000753 (rank : 181) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 921 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q56UN5, Q56UN1, Q56UN2, Q56UN3, Q56UN4, Q8N4E9, Q9H5T2 | Gene names | YSK4, RCK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SPS1/STE20-related protein kinase YSK4 (EC 2.7.11.1) (Regulated in COPD, protein kinase). | |||||
|
AFF2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 101) | NC score | 0.084385 (rank : 32) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O55112 | Gene names | Aff2, Fmr2, Ox19 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 2 (Fragile X mental retardation protein 2 homolog) (Protein FMR-2) (FMR2P) (Protein Ox19). | |||||
|
AP3B1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 102) | NC score | 0.040993 (rank : 132) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O00203, O00580, Q7Z393, Q9HD66 | Gene names | AP3B1, ADTB3A | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit beta-1 (Adapter-related protein complex 3 beta-1 subunit) (Beta3A-adaptin) (Adaptor protein complex AP-3 beta-1 subunit) (Clathrin assembly protein complex 3 beta-1 large chain). | |||||
|
BNC1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 103) | NC score | 0.010103 (rank : 171) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 582 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O35914, O54886 | Gene names | Bnc1, Bnc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein basonuclin-1. | |||||
|
F120C_HUMAN
|
||||||
| θ value | 6.88961 (rank : 104) | NC score | 0.014110 (rank : 165) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NX05 | Gene names | FAM120C, CXorf17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UPF0318 protein FAM120C (Tumor antigen BJ-HCC-21). | |||||
|
JPH2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 105) | NC score | 0.019736 (rank : 157) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9ET78, Q9ET79 | Gene names | Jph2, Jp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Junctophilin-2 (Junctophilin type 2) (JP-2). | |||||
|
MLL2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 106) | NC score | 0.043358 (rank : 128) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
SNX9_MOUSE
|
||||||
| θ value | 6.88961 (rank : 107) | NC score | 0.041346 (rank : 131) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q91VH2 | Gene names | Snx9 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-9. | |||||
|
ANLN_HUMAN
|
||||||
| θ value | 8.99809 (rank : 108) | NC score | 0.025137 (rank : 151) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9NQW6, Q5CZ78, Q6NSK5, Q9H8Y4, Q9NVN9, Q9NVP0 | Gene names | ANLN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Actin-binding protein anillin. | |||||
|
ASB1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 109) | NC score | 0.004849 (rank : 179) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 291 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y576, Q9ULS4 | Gene names | ASB1, KIAA1146 | |||
|
Domain Architecture |
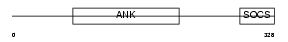
|
|||||
| Description | Ankyrin repeat and SOCS box protein 1 (ASB-1). | |||||
|
CDYL_MOUSE
|
||||||
| θ value | 8.99809 (rank : 110) | NC score | 0.009054 (rank : 172) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9WTK2 | Gene names | Cdyl | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain Y-like protein (CDY-like). | |||||
|
CUTL2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 111) | NC score | 0.013079 (rank : 167) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O14529 | Gene names | CUTL2, KIAA0293 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 2 (Homeobox protein Cux-2) (Cut-like 2). | |||||
|
CXXC6_HUMAN
|
||||||
| θ value | 8.99809 (rank : 112) | NC score | 0.019398 (rank : 158) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8NFU7, Q5VUP7, Q7Z6B6, Q8TCR1, Q9C0I7 | Gene names | CXXC6, KIAA1676, LCX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CXXC-type zinc finger protein 6 (Leukemia-associated protein with a CXXC domain). | |||||
|
ERIC1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 113) | NC score | 0.045104 (rank : 125) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q86X53, Q9P063 | Gene names | ERICH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate-rich protein 1. | |||||
|
FNBP2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 114) | NC score | 0.008086 (rank : 176) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 432 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O75044 | Gene names | SRGAP2, FNBP2, KIAA0456 | |||
|
Domain Architecture |

|
|||||
| Description | SLIT-ROBO Rho GTPase-activating protein 2 (srGAP2) (Formin-binding protein 2). | |||||
|
HRB2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 115) | NC score | 0.015776 (rank : 163) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BGA5, Q8C029 | Gene names | Hrb2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HIV-1 Rev-binding protein 2 homolog. | |||||
|
HUCE1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 116) | NC score | 0.016402 (rank : 162) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9DB85, Q8BHW3 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cerebral protein 1 homolog. | |||||
|
LIPB2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 117) | NC score | 0.008817 (rank : 173) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 599 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8ND30, O75337, Q8WW26 | Gene names | PPFIBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-beta-2 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein-binding protein 2) (PTPRF-interacting protein-binding protein 2). | |||||
|
LYAR_MOUSE
|
||||||
| θ value | 8.99809 (rank : 118) | NC score | 0.038651 (rank : 135) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q08288, Q9D9X2 | Gene names | Lyar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell growth-regulating nucleolar protein. | |||||
|
MAP1B_HUMAN
|
||||||
| θ value | 8.99809 (rank : 119) | NC score | 0.091154 (rank : 24) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 903 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | P46821 | Gene names | MAP1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) [Contains: MAP1 light chain LC1]. | |||||
|
PA24B_HUMAN
|
||||||
| θ value | 8.99809 (rank : 120) | NC score | 0.008273 (rank : 175) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O95712, Q59GF9, Q8TB10, Q9UKV7 | Gene names | PLA2G4B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytosolic phospholipase A2 beta (EC 3.1.1.4) (cPLA2-beta) (Phospholipase A2 group IVB). | |||||
|
RBMX2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 121) | NC score | 0.039592 (rank : 134) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9Y388, Q5JY82, Q9Y3I8 | Gene names | RBMX2 | |||
|
Domain Architecture |
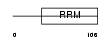
|
|||||
| Description | RNA-binding motif protein, X-linked 2. | |||||
|
SRPR_HUMAN
|
||||||
| θ value | 8.99809 (rank : 122) | NC score | 0.014062 (rank : 166) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P08240, Q9BVJ4 | Gene names | SRPR | |||
|
Domain Architecture |

|
|||||
| Description | Signal recognition particle receptor subunit alpha (SR-alpha) (Docking protein alpha) (DP-alpha). | |||||
|
TENX_HUMAN
|
||||||
| θ value | 8.99809 (rank : 123) | NC score | 0.001971 (rank : 180) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 789 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P22105, P78530, P78531, Q08424, Q9UMG7 | Gene names | TNXB, HXBL, TNX, XB | |||
|
Domain Architecture |

|
|||||
| Description | Tenascin-X precursor (TN-X) (Hexabrachion-like protein). | |||||
|
TERF1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 124) | NC score | 0.027149 (rank : 146) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P54274, Q15553, Q93029 | Gene names | TERF1, PIN2, TRF, TRF1 | |||
|
Domain Architecture |
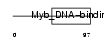
|
|||||
| Description | Telomeric repeat-binding factor 1 (TTAGGG repeat-binding factor 1) (NIMA-interacting protein 2) (Telomeric protein Pin2/TRF1). | |||||
|
ZC11A_MOUSE
|
||||||
| θ value | 8.99809 (rank : 125) | NC score | 0.021879 (rank : 155) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q6NZF1, Q6NXW9, Q6PDR6, Q80TU7, Q8C5L5, Q99JN6 | Gene names | Zc3h11a, Kiaa0663 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 11A. | |||||
|
ZCH10_MOUSE
|
||||||
| θ value | 8.99809 (rank : 126) | NC score | 0.036164 (rank : 138) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9CX48 | Gene names | Zcchc10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 10. | |||||
|
ACINU_MOUSE
|
||||||
| θ value | θ > 10 (rank : 127) | NC score | 0.058274 (rank : 78) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 712 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9JIX8, Q9CSN7, Q9CSR9, Q9CSX7, Q9R046, Q9R047 | Gene names | Acin1, Acinus | |||
|
Domain Architecture |

|
|||||
| Description | Apoptotic chromatin condensation inducer in the nucleus (Acinus). | |||||
|
AFF1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 128) | NC score | 0.090430 (rank : 26) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P51825 | Gene names | AFF1, AF4, FEL, MLLT2 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 1 (Protein AF-4) (Proto-oncogene AF4) (Protein FEL). | |||||
|
AFF1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 129) | NC score | 0.071565 (rank : 44) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O88573 | Gene names | Aff1, Mllt2, Mllt2h | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 1 (Protein AF-4) (Proto-oncogene AF4). | |||||
|
AFF2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 130) | NC score | 0.068946 (rank : 52) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P51816, O43786, O60215, P78407, Q13521, Q14323, Q9UNA5 | Gene names | AFF2, FMR2, OX19 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 2 (Fragile X mental retardation 2 protein) (Protein FMR-2) (FMR2P) (Protein Ox19) (Fragile X E mental retardation syndrome protein). | |||||
|
AKA12_HUMAN
|
||||||
| θ value | θ > 10 (rank : 131) | NC score | 0.064394 (rank : 57) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q02952, O00310, O00498, Q99970 | Gene names | AKAP12, AKAP250 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 12 (A-kinase anchor protein 250 kDa) (AKAP 250) (Myasthenia gravis autoantigen gravin). | |||||
|
ATRX_MOUSE
|
||||||
| θ value | θ > 10 (rank : 132) | NC score | 0.053593 (rank : 100) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 682 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q61687 | Gene names | Atrx, Hp1bp2, Xnp | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ATRX (EC 3.6.1.-) (ATP-dependent helicase ATRX) (X-linked nuclear protein) (Heterochromatin protein 2) (HP1 alpha-interacting protein) (HP1-BP38 protein). | |||||
|
BASP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 133) | NC score | 0.055154 (rank : 91) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P80723, O43596, Q5U0S0 | Gene names | BASP1, NAP22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
BASP_MOUSE
|
||||||
| θ value | θ > 10 (rank : 134) | NC score | 0.052478 (rank : 106) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q91XV3 | Gene names | Basp1, Nap22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
CJ012_HUMAN
|
||||||
| θ value | θ > 10 (rank : 135) | NC score | 0.070253 (rank : 47) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8N655, Q9H945, Q9Y457 | Gene names | C10orf12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf12. | |||||
|
CYLC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 136) | NC score | 0.064472 (rank : 56) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P35663, Q5JQQ9 | Gene names | CYLC1, CYL, CYL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-1 (Cylicin I) (Multiple-band polypeptide I). | |||||
|
CYLC2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 137) | NC score | 0.070271 (rank : 45) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q14093 | Gene names | CYLC2, CYL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-2 (Cylicin II) (Multiple-band polypeptide II). | |||||
|
DMP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 138) | NC score | 0.085712 (rank : 30) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q13316, O43265 | Gene names | DMP1 | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1). | |||||
|
ENL_HUMAN
|
||||||
| θ value | θ > 10 (rank : 139) | NC score | 0.057920 (rank : 80) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q03111, Q14768 | Gene names | MLLT1, ENL, LTG19, YEATS1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein ENL (YEATS domain-containing protein 1). | |||||
|
FILA_HUMAN
|
||||||
| θ value | θ > 10 (rank : 140) | NC score | 0.054274 (rank : 96) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 685 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P20930, Q01720, Q5T583 | Gene names | FLG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filaggrin. | |||||
|
GPTC8_HUMAN
|
||||||
| θ value | θ > 10 (rank : 141) | NC score | 0.051382 (rank : 110) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 366 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9UKJ3, O60300 | Gene names | GPATC8, KIAA0553 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G patch domain-containing protein 8. | |||||
|
HIRP3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 142) | NC score | 0.063819 (rank : 59) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9BW71, O75707, O75708 | Gene names | HIRIP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HIRA-interacting protein 3. | |||||
|
IWS1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 143) | NC score | 0.086665 (rank : 28) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
IWS1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 144) | NC score | 0.077146 (rank : 39) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
KI67_HUMAN
|
||||||
| θ value | θ > 10 (rank : 145) | NC score | 0.058584 (rank : 76) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P46013 | Gene names | MKI67 | |||
|
Domain Architecture |

|
|||||
| Description | Antigen KI-67. | |||||
|
LEO1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 146) | NC score | 0.081569 (rank : 36) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
LRRF1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 147) | NC score | 0.051384 (rank : 109) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 440 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q3UZ39, O70323, Q3TS94 | Gene names | Lrrfip1, Flap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat flightless-interacting protein 1 (LRR FLII- interacting protein 1) (FLI-LRR-associated protein 1) (Flap-1) (H186 FLAP). | |||||
|
MBB1A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 148) | NC score | 0.064011 (rank : 58) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q7TPV4, O35851, Q80Y66, Q8R4X2, Q99KP0 | Gene names | Mybbp1a, P160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myb-binding protein 1A (Myb-binding protein of 160 kDa). | |||||
|
MDC1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 149) | NC score | 0.058401 (rank : 77) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
MDN1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 150) | NC score | 0.053859 (rank : 99) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 662 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9NU22, O15019 | Gene names | MDN1, KIAA0301 | |||
|
Domain Architecture |

|
|||||
| Description | Midasin (MIDAS-containing protein). | |||||
|
NSBP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 151) | NC score | 0.072085 (rank : 43) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P82970 | Gene names | NSBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1. | |||||
|
NSBP1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 152) | NC score | 0.052505 (rank : 105) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9JL35, O88832, Q8VC71, Q9CUW1 | Gene names | Nsbp1, Garp45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1 (Nucleosome-binding protein 45) (NBP-45) (GARP45 protein). | |||||
|
NUCKS_HUMAN
|
||||||
| θ value | θ > 10 (rank : 153) | NC score | 0.055362 (rank : 89) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9H1E3, Q9H1D6, Q9H723 | Gene names | NUCKS1, NUCKS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (P1). | |||||
|
OXR1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 154) | NC score | 0.050464 (rank : 114) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 516 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q4KMM3, Q5FWW1, Q99L06, Q99MK1, Q99MP4 | Gene names | Oxr1, C7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Oxidation resistance protein 1 (Protein C7). | |||||
|
RBBP6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 155) | NC score | 0.069565 (rank : 50) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q7Z6E9, Q15290, Q6DKH4, Q6P4C2, Q6YNC9, Q7Z6E8, Q8N0V2, Q96PH3, Q9H3I8, Q9H5M5, Q9NPX4 | Gene names | RBBP6, P2PR, PACT, RBQ1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R) (Retinoblastoma-binding Q protein 1) (Protein RBQ-1). | |||||
|
RBBP6_MOUSE
|
||||||
| θ value | θ > 10 (rank : 156) | NC score | 0.069123 (rank : 51) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1014 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P97868, P70287, Q3TTR9, Q3TUM7, Q3UMP7, Q4U217, Q7TT06, Q8BNY8, Q8R399 | Gene names | Rbbp6, P2pr, Pact | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R). | |||||
|
RL1D1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 157) | NC score | 0.054017 (rank : 98) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O76021, Q6PL22, Q8IWS7, Q8WUZ1, Q9HDA9, Q9Y3Z9 | Gene names | RSL1D1, CATX11, CSIG, PBK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ribosomal L1 domain-containing protein 1 (Cellular senescence- inhibited gene protein) (Protein PBK1) (CATX-11). | |||||
|
RSF1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 158) | NC score | 0.054155 (rank : 97) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 539 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q96T23, Q86X86, Q9H3L8, Q9NVZ8, Q9NYU0 | Gene names | RSF1, HBXAP, XAP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Remodeling and spacing factor 1 (Rsf-1) (Hepatitis B virus X- associated protein) (HBV pX-associated protein 8) (p325 subunit of RSF chromatin remodelling complex). | |||||
|
SET1A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 159) | NC score | 0.055752 (rank : 86) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | O15047, Q6PIF3, Q8TAJ6 | Gene names | SETD1A, KIAA0339, SET1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET1 (EC 2.1.1.43) (Set1/Ash2 histone methyltransferase complex subunit SET1) (SET-domain-containing protein 1A). | |||||
|
SFR11_HUMAN
|
||||||
| θ value | θ > 10 (rank : 160) | NC score | 0.062521 (rank : 65) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q05519 | Gene names | SFRS11 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor arginine/serine-rich 11 (Arginine-rich 54 kDa nuclear protein) (p54). | |||||
|
SFR12_HUMAN
|
||||||
| θ value | θ > 10 (rank : 161) | NC score | 0.054895 (rank : 92) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 553 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8WXA9, Q86X37 | Gene names | SFRS12, SRRP86 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 12 (Serine-arginine-rich- splicing regulatory protein 86) (SRrp86) (Splicing regulatory protein 508) (SRrp508). | |||||
|
SFR12_MOUSE
|
||||||
| θ value | θ > 10 (rank : 162) | NC score | 0.054632 (rank : 94) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 486 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8BZX4, Q8BLM8, Q8BWK7, Q8BYK0, Q8BYZ4 | Gene names | Sfrs12, Srrp86 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 12 (Serine-arginine-rich- splicing regulatory protein 86) (SRrp86). | |||||
|
SFR17_HUMAN
|
||||||
| θ value | θ > 10 (rank : 163) | NC score | 0.050632 (rank : 113) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8TF01, Q5T064, Q6P2B4, Q6P2N4, Q6PJ93, Q6PK36, Q7Z640, Q8TF00, Q96K10, Q96SM5, Q9P076, Q9P0C0, Q9Y4N3 | Gene names | SRRP130, C6orf111 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, arginine/serine-rich 130 (Serine-arginine-rich- splicing regulatory protein 130) (SRrp130) (SR-rich protein) (SR- related protein). | |||||
|
SFRIP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 164) | NC score | 0.070263 (rank : 46) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 530 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q99590, Q8IW59 | Gene names | SFRS2IP, CASP11, SIP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SFRS2-interacting protein (Splicing factor, arginine/serine-rich 2- interacting protein) (SC35-interacting protein 1) (CTD-associated SR protein 11) (Splicing regulatory protein 129) (SRrp129) (NY-REN-40 antigen). | |||||
|
SMRC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 165) | NC score | 0.057050 (rank : 82) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 375 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q92922, Q6P172, Q8IWH2 | Gene names | SMARCC1, BAF155 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 1 (SWI/SNF complex 155 kDa subunit) (BRG1-associated factor 155). | |||||
|
SMRC1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 166) | NC score | 0.056190 (rank : 85) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 459 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P97496, Q7TS80, Q7TT29 | Gene names | Smarcc1, Baf155, Srg3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 1 (SWI/SNF complex 155 kDa subunit) (BRG1-associated factor 155) (SWI3-related protein). | |||||
|
SRRM1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 167) | NC score | 0.060278 (rank : 70) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
SRRM1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 168) | NC score | 0.060459 (rank : 69) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
SRRM2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 169) | NC score | 0.069939 (rank : 49) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 170) | NC score | 0.063200 (rank : 62) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
T2FA_HUMAN
|
||||||
| θ value | θ > 10 (rank : 171) | NC score | 0.089652 (rank : 27) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P35269, Q9BWN0 | Gene names | GTF2F1, RAP74 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor IIF alpha subunit (EC 2.7.11.1) (TFIIF-alpha) (Transcription initiation factor RAP74) (General transcription factor IIF polypeptide 1 74 kDa subunit protein). | |||||
|
TARA_HUMAN
|
||||||
| θ value | θ > 10 (rank : 172) | NC score | 0.062954 (rank : 64) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9H2D6, O94797, Q2PZW8, Q2Q3Z9, Q2Q400, Q5R3M6, Q96DW1, Q9BT77, Q9BTL7, Q9BY98, Q9Y3L4 | Gene names | TRIOBP, KIAA1662, TARA | |||
|
Domain Architecture |

|
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
TGON2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 173) | NC score | 0.086178 (rank : 29) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1037 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | O43493, O15282, O43492, O43499, O43500, O43501, Q92760 | Gene names | TGOLN2, TGN46, TGN51 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (Trans-Golgi network protein TGN51) (TGN46) (TGN48) (TGN38 homolog). | |||||
|
TGON2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 174) | NC score | 0.058189 (rank : 79) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q62314 | Gene names | Tgoln2, Ttgn2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (TGN38B). | |||||
|
THOC2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 175) | NC score | 0.055327 (rank : 90) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8NI27, Q9H8I6 | Gene names | THOC2, CXorf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | THO complex subunit 2 (Tho2). | |||||
|
TNR6A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 176) | NC score | 0.050422 (rank : 115) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8NDV7, O15408, Q658L5, Q6NVB5, Q8NEZ0, Q8TBT8, Q8TCR0, Q9NV59, Q9P268 | Gene names | TNRC6A, CAGH26, KIAA1460, TNRC6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trinucleotide repeat-containing gene 6A protein (CAG repeat protein 26) (Glycine-tryptophan protein of 182 kDa) (GW182 autoantigen) (Protein GW1) (EMSY interactor protein). | |||||
|
TRDN_HUMAN
|
||||||
| θ value | θ > 10 (rank : 177) | NC score | 0.068422 (rank : 54) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 542 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q13061 | Gene names | TRDN | |||
|
Domain Architecture |

|
|||||
| Description | Triadin. | |||||
|
TTC16_HUMAN
|
||||||
| θ value | θ > 10 (rank : 178) | NC score | 0.055488 (rank : 88) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8NEE8, Q5JU66, Q96M72 | Gene names | TTC16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tetratricopeptide repeat protein 16 (TPR repeat protein 16). | |||||
|
TXND2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 179) | NC score | 0.076427 (rank : 40) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 793 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q86VQ3, Q8N7U4, Q96RX3, Q9H0L8 | Gene names | TXNDC2, SPTRX, SPTRX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1). | |||||
|
TXND2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 180) | NC score | 0.082502 (rank : 35) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
YTDC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 181) | NC score | 0.053436 (rank : 101) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q96MU7, Q7Z622, Q8TF35 | Gene names | YTHDC1, KIAA1966, YT521 | |||
|
Domain Architecture |

|
|||||
| Description | YTH domain-containing protein 1 (Putative splicing factor YT521). | |||||
|
LYRIC_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 126 | |
| SwissProt Accessions | Q80WJ7, Q3U9F8, Q3UAQ8, Q8BN67, Q8CBT9, Q8CDL0, Q8CGI7, Q9D052 | Gene names | Mtdh, Lyric | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LYRIC (Lysine-rich CEACAM1 co-isolated protein) (3D3/LYRIC) (Metastasis adhesion protein) (Metadherin). | |||||
|
LYRIC_HUMAN
|
||||||
| NC score | 0.943042 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q86UE4, Q6PK07, Q8TCX3 | Gene names | MTDH, AEG1, LYRIC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LYRIC (Lysine-rich CEACAM1 co-isolated protein) (3D3/lyric) (Metastasis adhesion protein) (Metadherin) (Astrocyte elevated gene-1 protein) (AEG-1). | |||||
|
DSPP_HUMAN
|
||||||
| NC score | 0.179111 (rank : 3) | θ value | 6.43352e-06 (rank : 3) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q9NZW4, O95815 | Gene names | DSPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
DSPP_MOUSE
|
||||||
| NC score | 0.171916 (rank : 4) | θ value | 0.00175202 (rank : 7) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P97399, O70567 | Gene names | Dspp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor (Dentin matrix protein 3) (DMP-3) [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
AFF4_MOUSE
|
||||||
| NC score | 0.169258 (rank : 5) | θ value | 0.00175202 (rank : 6) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9ESC8, Q8C6K4, Q8C6W3, Q8CCH3 | Gene names | Aff4, Alf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4. | |||||
|
NIPBL_HUMAN
|
||||||
| NC score | 0.166847 (rank : 6) | θ value | 1.43324e-05 (rank : 4) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
AFF4_HUMAN
|
||||||
| NC score | 0.165554 (rank : 7) | θ value | 0.0148317 (rank : 9) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 611 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9UHB7, Q498B2, Q59FB3, Q6P592, Q8TDR1, Q9P0E4 | Gene names | AFF4, AF5Q31, MCEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4 (ALL1-fused gene from chromosome 5q31) (Major CDK9 elongation factor-associated protein). | |||||
|
NPBL_MOUSE
|
||||||
| NC score | 0.163972 (rank : 8) | θ value | 0.000461057 (rank : 5) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
LAD1_HUMAN
|
||||||
| NC score | 0.122070 (rank : 9) | θ value | 0.00869519 (rank : 8) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 441 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O00515, O95614 | Gene names | LAD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ladinin 1 (Lad-1) (120 kDa linear IgA bullous dermatosis antigen) (97 kDa linear IgA bullous dermatosis antigen) (Linear IgA disease antigen homolog) (LadA). | |||||
|
DMP1_MOUSE
|
||||||
| NC score | 0.118337 (rank : 10) | θ value | 0.125558 (rank : 17) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O55188 | Gene names | Dmp1, Dmp | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1) (AG1). | |||||
|
NOLC1_HUMAN
|
||||||
| NC score | 0.114400 (rank : 11) | θ value | 0.813845 (rank : 43) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q14978, Q15030 | Gene names | NOLC1, KIAA0035 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar phosphoprotein p130 (Nucleolar 130 kDa protein) (140 kDa nucleolar phosphoprotein) (Nopp140) (Nucleolar and coiled-body phosphoprotein 1). | |||||
|
R51A1_MOUSE
|
||||||
| NC score | 0.109374 (rank : 12) | θ value | 0.125558 (rank : 19) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 164 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8C551, O55219, Q8BP36, Q99L94, Q9D0J0 | Gene names | Rad51ap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RAD51-associated protein 1 (RAD51-interacting protein) (RAB22). | |||||
|
SELPL_MOUSE
|
||||||
| NC score | 0.104002 (rank : 13) | θ value | 0.0252991 (rank : 10) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 588 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q62170 | Gene names | Selplg, Selp1, Selpl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | P-selectin glycoprotein ligand 1 precursor (PSGL-1) (Selectin P ligand). | |||||
|
MAP1B_MOUSE
|
||||||
| NC score | 0.103190 (rank : 14) | θ value | 0.813845 (rank : 41) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 977 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | P14873 | Gene names | Map1b, Mtap1b, Mtap5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) (MAP1.2) (MAP1(X)) [Contains: MAP1 light chain LC1]. | |||||
|
ACINU_HUMAN
|
||||||
| NC score | 0.102428 (rank : 15) | θ value | 0.0563607 (rank : 13) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q9UKV3, O75158, Q9UG91, Q9UKV1, Q9UKV2 | Gene names | ACIN1, ACINUS, KIAA0670 | |||
|
Domain Architecture |

|
|||||
| Description | Apoptotic chromatin condensation inducer in the nucleus (Acinus). | |||||
|
SMRC2_HUMAN
|
||||||
| NC score | 0.101281 (rank : 16) | θ value | 0.0330416 (rank : 12) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 499 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8TAQ2, Q92923, Q96E12, Q96GY4 | Gene names | SMARCC2, BAF170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 2 (SWI/SNF complex 170 kDa subunit) (BRG1-associated factor 170). | |||||
|
CJ120_HUMAN
|
||||||
| NC score | 0.098340 (rank : 17) | θ value | 0.47712 (rank : 30) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q5SQS8 | Gene names | C10orf120 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative uncharacterized protein C10orf120. | |||||
|
SMRC2_MOUSE
|
||||||
| NC score | 0.098053 (rank : 18) | θ value | 0.0252991 (rank : 11) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 487 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q6PDG5, Q6P6P3 | Gene names | Smarcc2, Baf170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 2 (SWI/SNF complex 170 kDa subunit) (BRG1-associated factor 170). | |||||
|
AFF3_HUMAN
|
||||||
| NC score | 0.096959 (rank : 19) | θ value | 1.06291 (rank : 45) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 190 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P51826 | Gene names | AFF3, LAF4 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 3 (Protein LAF-4) (Lymphoid nuclear protein related to AF4). | |||||
|
LEO1_MOUSE
|
||||||
| NC score | 0.092774 (rank : 20) | θ value | 5.27518 (rank : 93) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
SETD8_MOUSE
|
||||||
| NC score | 0.092372 (rank : 21) | θ value | 0.47712 (rank : 34) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q2YDW7, Q8C0J9 | Gene names | SETD8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H4 lysine-20 specific (EC 2.1.1.43) (Histone H4-K20 methyltransferase) (H4-K20-HMTase) (SET domain-containing lysine methyltransferase 8) (SET domain-containing protein 8). | |||||
|
SETD8_HUMAN
|
||||||
| NC score | 0.091636 (rank : 22) | θ value | 0.21417 (rank : 22) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9NQR1, Q86W83, Q8TD09 | Gene names | SETD8, PRSET7, SET07, SET8 | |||
|
Domain Architecture |

|
|||||
| Description | Histone-lysine N-methyltransferase, H4 lysine-20 specific (EC 2.1.1.43) (Histone H4-K20 methyltransferase) (H4-K20-HMTase) (SET domain-containing lysine methyltransferase 8) (SET domain-containing protein 8) (PR/SET domain-containing protein 07) (PR/SET07) (PR-Set7). | |||||
|
TNC6B_HUMAN
|
||||||
| NC score | 0.091592 (rank : 23) | θ value | 2.36792 (rank : 72) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 256 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9UPQ9, Q5TH52, Q8TBX2 | Gene names | TNRC6B, KIAA1093 | |||
|
Domain Architecture |

|
|||||
| Description | Trinucleotide repeat-containing 6B protein. | |||||
|
MAP1B_HUMAN
|
||||||
| NC score | 0.091154 (rank : 24) | θ value | 8.99809 (rank : 119) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 903 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | P46821 | Gene names | MAP1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) [Contains: MAP1 light chain LC1]. | |||||
|
LAD1_MOUSE
|
||||||
| NC score | 0.091039 (rank : 25) | θ value | 5.27518 (rank : 92) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 391 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P57016 | Gene names | Lad1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ladinin 1 (Lad-1) (Linear IgA disease autoantigen). | |||||
|
AFF1_HUMAN
|
||||||
| NC score | 0.090430 (rank : 26) | θ value | θ > 10 (rank : 128) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P51825 | Gene names | AFF1, AF4, FEL, MLLT2 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 1 (Protein AF-4) (Proto-oncogene AF4) (Protein FEL). | |||||
|
T2FA_HUMAN
|
||||||
| NC score | 0.089652 (rank : 27) | θ value | θ > 10 (rank : 171) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P35269, Q9BWN0 | Gene names | GTF2F1, RAP74 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor IIF alpha subunit (EC 2.7.11.1) (TFIIF-alpha) (Transcription initiation factor RAP74) (General transcription factor IIF polypeptide 1 74 kDa subunit protein). | |||||
|
IWS1_HUMAN
|
||||||
| NC score | 0.086665 (rank : 28) | θ value | θ > 10 (rank : 143) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
TGON2_HUMAN
|
||||||
| NC score | 0.086178 (rank : 29) | θ value | θ > 10 (rank : 173) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1037 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | O43493, O15282, O43492, O43499, O43500, O43501, Q92760 | Gene names | TGOLN2, TGN46, TGN51 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (Trans-Golgi network protein TGN51) (TGN46) (TGN48) (TGN38 homolog). | |||||
|
DMP1_HUMAN
|
||||||
| NC score | 0.085712 (rank : 30) | θ value | θ > 10 (rank : 138) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q13316, O43265 | Gene names | DMP1 | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1). | |||||
|
MDC1_HUMAN
|
||||||
| NC score | 0.085463 (rank : 31) | θ value | 0.279714 (rank : 25) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
AFF2_MOUSE
|
||||||
| NC score | 0.084385 (rank : 32) | θ value | 6.88961 (rank : 101) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O55112 | Gene names | Aff2, Fmr2, Ox19 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 2 (Fragile X mental retardation protein 2 homolog) (Protein FMR-2) (FMR2P) (Protein Ox19). | |||||
|
BPTF_HUMAN
|
||||||
| NC score | 0.082957 (rank : 33) | θ value | 0.0736092 (rank : 15) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 786 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q12830, Q6NX67, Q7Z7D6, Q9UIG2 | Gene names | FALZ, BPTF, FAC1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleosome remodeling factor subunit BPTF (Bromodomain and PHD finger- containing transcription factor) (Fetal Alzheimer antigen) (Fetal Alz- 50 clone 1 protein). | |||||
|
DPOLZ_HUMAN
|
||||||
| NC score | 0.082754 (rank : 34) | θ value | 0.0563607 (rank : 14) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O60673, O43214 | Gene names | REV3L, POLZ, REV3 | |||
|
Domain Architecture |

|
|||||
| Description | DNA polymerase zeta catalytic subunit (EC 2.7.7.7) (hREV3). | |||||
|
TXND2_MOUSE
|
||||||
| NC score | 0.082502 (rank : 35) | θ value | θ > 10 (rank : 180) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
LEO1_HUMAN
|
||||||
| NC score | 0.081569 (rank : 36) | θ value | θ > 10 (rank : 146) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
NASP_HUMAN
|
||||||
| NC score | 0.080460 (rank : 37) | θ value | 3.0926 (rank : 77) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P49321, Q96A69, Q9BTW2 | Gene names | NASP | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear autoantigenic sperm protein (NASP). | |||||
|
PRG4_MOUSE
|
||||||
| NC score | 0.078016 (rank : 38) | θ value | 3.0926 (rank : 78) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 624 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q9JM99, Q3UEL1, Q3V198 | Gene names | Prg4, Msf, Szp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
IWS1_MOUSE
|
||||||
| NC score | 0.077146 (rank : 39) | θ value | θ > 10 (rank : 144) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
TXND2_HUMAN
|
||||||
| NC score | 0.076427 (rank : 40) | θ value | θ > 10 (rank : 179) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 793 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q86VQ3, Q8N7U4, Q96RX3, Q9H0L8 | Gene names | TXNDC2, SPTRX, SPTRX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1). | |||||
|
PRG4_HUMAN
|
||||||
| NC score | 0.076259 (rank : 41) | θ value | 1.06291 (rank : 48) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q92954, Q6DNC4, Q6DNC5, Q6ZMZ5, Q9BX49 | Gene names | PRG4, MSF, SZP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
ADDB_MOUSE
|
||||||
| NC score | 0.073618 (rank : 42) | θ value | 0.47712 (rank : 29) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 574 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9QYB8, Q3U0E1, Q80VH9, Q8C1C4, Q9CXE3, Q9JLE5, Q9QYB7 | Gene names | Add2 | |||
|
Domain Architecture |

|
|||||
| Description | Beta-adducin (Erythrocyte adducin subunit beta) (Add97). | |||||
|
NSBP1_HUMAN
|
||||||
| NC score | 0.072085 (rank : 43) | θ value | θ > 10 (rank : 151) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P82970 | Gene names | NSBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1. | |||||
|
AFF1_MOUSE
|
||||||
| NC score | 0.071565 (rank : 44) | θ value | θ > 10 (rank : 129) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O88573 | Gene names | Aff1, Mllt2, Mllt2h | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 1 (Protein AF-4) (Proto-oncogene AF4). | |||||
|
CYLC2_HUMAN
|
||||||
| NC score | 0.070271 (rank : 45) | θ value | θ > 10 (rank : 137) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q14093 | Gene names | CYLC2, CYL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-2 (Cylicin II) (Multiple-band polypeptide II). | |||||
|
SFRIP_HUMAN
|
||||||
| NC score | 0.070263 (rank : 46) | θ value | θ > 10 (rank : 164) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 530 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q99590, Q8IW59 | Gene names | SFRS2IP, CASP11, SIP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SFRS2-interacting protein (Splicing factor, arginine/serine-rich 2- interacting protein) (SC35-interacting protein 1) (CTD-associated SR protein 11) (Splicing regulatory protein 129) (SRrp129) (NY-REN-40 antigen). | |||||
|
CJ012_HUMAN
|
||||||
| NC score | 0.070253 (rank : 47) | θ value | θ > 10 (rank : 135) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q8N655, Q9H945, Q9Y457 | Gene names | C10orf12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf12. | |||||
|
BRD8_MOUSE
|
||||||
| NC score | 0.070079 (rank : 48) | θ value | 1.06291 (rank : 47) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 422 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8R3B7, Q8C049, Q8R583, Q8VDP0 | Gene names | Brd8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 8. | |||||
|
SRRM2_HUMAN
|
||||||
| NC score | 0.069939 (rank : 49) | θ value | θ > 10 (rank : 169) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
RBBP6_HUMAN
|
||||||
| NC score | 0.069565 (rank : 50) | θ value | θ > 10 (rank : 155) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q7Z6E9, Q15290, Q6DKH4, Q6P4C2, Q6YNC9, Q7Z6E8, Q8N0V2, Q96PH3, Q9H3I8, Q9H5M5, Q9NPX4 | Gene names | RBBP6, P2PR, PACT, RBQ1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R) (Retinoblastoma-binding Q protein 1) (Protein RBQ-1). | |||||
|
RBBP6_MOUSE
|
||||||
| NC score | 0.069123 (rank : 51) | θ value | θ > 10 (rank : 156) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1014 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P97868, P70287, Q3TTR9, Q3TUM7, Q3UMP7, Q4U217, Q7TT06, Q8BNY8, Q8R399 | Gene names | Rbbp6, P2pr, Pact | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R). | |||||
|
AFF2_HUMAN
|
||||||
| NC score | 0.068946 (rank : 52) | θ value | θ > 10 (rank : 130) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P51816, O43786, O60215, P78407, Q13521, Q14323, Q9UNA5 | Gene names | AFF2, FMR2, OX19 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 2 (Fragile X mental retardation 2 protein) (Protein FMR-2) (FMR2P) (Protein Ox19) (Fragile X E mental retardation syndrome protein). | |||||
|
ASXL1_MOUSE
|
||||||
| NC score | 0.068640 (rank : 53) | θ value | 0.21417 (rank : 20) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 182 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P59598 | Gene names | Asxl1, Kiaa0978 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative Polycomb group protein ASXL1 (Additional sex combs-like protein 1). | |||||
|
TRDN_HUMAN
|
||||||
| NC score | 0.068422 (rank : 54) | θ value | θ > 10 (rank : 177) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 542 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q13061 | Gene names | TRDN | |||
|
Domain Architecture |

|
|||||
| Description | Triadin. | |||||
|
NACAM_MOUSE
|
||||||
| NC score | 0.065808 (rank : 55) | θ value | 0.47712 (rank : 32) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
CYLC1_HUMAN
|
||||||
| NC score | 0.064472 (rank : 56) | θ value | θ > 10 (rank : 136) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P35663, Q5JQQ9 | Gene names | CYLC1, CYL, CYL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-1 (Cylicin I) (Multiple-band polypeptide I). | |||||
|
AKA12_HUMAN
|
||||||
| NC score | 0.064394 (rank : 57) | θ value | θ > 10 (rank : 131) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q02952, O00310, O00498, Q99970 | Gene names | AKAP12, AKAP250 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 12 (A-kinase anchor protein 250 kDa) (AKAP 250) (Myasthenia gravis autoantigen gravin). | |||||
|
MBB1A_MOUSE
|
||||||
| NC score | 0.064011 (rank : 58) | θ value | θ > 10 (rank : 148) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q7TPV4, O35851, Q80Y66, Q8R4X2, Q99KP0 | Gene names | Mybbp1a, P160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myb-binding protein 1A (Myb-binding protein of 160 kDa). | |||||
|
HIRP3_HUMAN
|
||||||
| NC score | 0.063819 (rank : 59) | θ value | θ > 10 (rank : 142) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9BW71, O75707, O75708 | Gene names | HIRIP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HIRA-interacting protein 3. | |||||
|
RPGR_HUMAN
|
||||||
| NC score | 0.063602 (rank : 60) | θ value | 0.21417 (rank : 21) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q92834, O00702, O00737, Q8N5T6, Q93039, Q9HD29, Q9UMR1 | Gene names | RPGR, RP3, XLRP3 | |||
|
Domain Architecture |

|
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator. | |||||
|
MINT_MOUSE
|
||||||
| NC score | 0.063261 (rank : 61) | θ value | 3.0926 (rank : 75) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.063200 (rank : 62) | θ value | θ > 10 (rank : 170) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
PHF14_HUMAN
|
||||||
| NC score | 0.063019 (rank : 63) | θ value | 4.03905 (rank : 80) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O94880 | Gene names | PHF14, KIAA0783 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 14. | |||||
|
TARA_HUMAN
|
||||||
| NC score | 0.062954 (rank : 64) | θ value | θ > 10 (rank : 172) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9H2D6, O94797, Q2PZW8, Q2Q3Z9, Q2Q400, Q5R3M6, Q96DW1, Q9BT77, Q9BTL7, Q9BY98, Q9Y3L4 | Gene names | TRIOBP, KIAA1662, TARA | |||
|
Domain Architecture |

|
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
SFR11_HUMAN
|
||||||
| NC score | 0.062521 (rank : 65) | θ value | θ > 10 (rank : 160) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q05519 | Gene names | SFRS11 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor arginine/serine-rich 11 (Arginine-rich 54 kDa nuclear protein) (p54). | |||||
|
DEK_HUMAN
|
||||||
| NC score | 0.061936 (rank : 66) | θ value | 5.27518 (rank : 86) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P35659, Q5TGV4, Q5TGV5 | Gene names | DEK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein DEK. | |||||
|
MBB1A_HUMAN
|
||||||
| NC score | 0.061115 (rank : 67) | θ value | 5.27518 (rank : 94) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 665 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9BQG0, Q86VM3, Q9BW49, Q9P0V5, Q9UF99 | Gene names | MYBBP1A, P160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myb-binding protein 1A. | |||||
|
ARI5B_HUMAN
|
||||||
| NC score | 0.060535 (rank : 68) | θ value | 1.81305 (rank : 54) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q14865, Q7Z3M4, Q8N421, Q9H786 | Gene names | ARID5B, DESRT, MRF2 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 5B (ARID domain- containing protein 5B) (Mrf1-like) (Modulator recognition factor 2) (MRF-2). | |||||
|
SRRM1_MOUSE
|
||||||
| NC score | 0.060459 (rank : 69) | θ value | θ > 10 (rank : 168) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
SRRM1_HUMAN
|
||||||
| NC score | 0.060278 (rank : 70) | θ value | θ > 10 (rank : 167) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
ARFG1_MOUSE
|
||||||
| NC score | 0.060204 (rank : 71) | θ value | 0.62314 (rank : 36) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9EPJ9, Q3TI52, Q8BMM6, Q8BMQ7 | Gene names | Arfgap1, Arf1gap | |||
|
Domain Architecture |

|
|||||
| Description | ADP-ribosylation factor GTPase-activating protein 1 (ADP-ribosylation factor 1 GTPase-activating protein) (ARF1 GAP) (ARF1-directed GTPase- activating protein) (GAP protein). | |||||
|
PHF14_MOUSE
|
||||||
| NC score | 0.060053 (rank : 72) | θ value | 1.81305 (rank : 58) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9D4H9, Q810Z6, Q8C284, Q8CGF8, Q9D1Q8 | Gene names | Phf14 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 14. | |||||
|
FA44A_HUMAN
|
||||||
| NC score | 0.060019 (rank : 73) | θ value | 2.36792 (rank : 63) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8NFC6, Q6P0M8 | Gene names | FAM44A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM44A. | |||||
|
BRD8_HUMAN
|
||||||
| NC score | 0.059948 (rank : 74) | θ value | 1.06291 (rank : 46) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9H0E9, O43178, Q15355, Q59GN0, Q969M9 | Gene names | BRD8, SMAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 8 (p120) (Skeletal muscle abundant protein) (Thyroid hormone receptor coactivating protein 120kDa) (TrCP120). | |||||
|
PCLO_HUMAN
|
||||||
| NC score | 0.059203 (rank : 75) | θ value | 1.38821 (rank : 52) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
KI67_HUMAN
|
||||||
| NC score | 0.058584 (rank : 76) | θ value | θ > 10 (rank : 145) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P46013 | Gene names | MKI67 | |||
|
Domain Architecture |

|
|||||
| Description | Antigen KI-67. | |||||
|
MDC1_MOUSE
|
||||||
| NC score | 0.058401 (rank : 77) | θ value | θ > 10 (rank : 149) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
ACINU_MOUSE
|
||||||
| NC score | 0.058274 (rank : 78) | θ value | θ > 10 (rank : 127) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 712 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9JIX8, Q9CSN7, Q9CSR9, Q9CSX7, Q9R046, Q9R047 | Gene names | Acin1, Acinus | |||
|
Domain Architecture |

|
|||||
| Description | Apoptotic chromatin condensation inducer in the nucleus (Acinus). | |||||
|
TGON2_MOUSE
|
||||||
| NC score | 0.058189 (rank : 79) | θ value | θ > 10 (rank : 174) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q62314 | Gene names | Tgoln2, Ttgn2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (TGN38B). | |||||
|
ENL_HUMAN
|
||||||
| NC score | 0.057920 (rank : 80) | θ value | θ > 10 (rank : 139) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q03111, Q14768 | Gene names | MLLT1, ENL, LTG19, YEATS1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein ENL (YEATS domain-containing protein 1). | |||||
|
MADCA_HUMAN
|
||||||
| NC score | 0.057529 (rank : 81) | θ value | 1.81305 (rank : 56) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q13477, O60222, O75867 | Gene names | MADCAM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucosal addressin cell adhesion molecule 1 precursor (MAdCAM-1) (hMAdCAM-1). | |||||
|
SMRC1_HUMAN
|
||||||
| NC score | 0.057050 (rank : 82) | θ value | θ > 10 (rank : 165) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 375 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q92922, Q6P172, Q8IWH2 | Gene names | SMARCC1, BAF155 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 1 (SWI/SNF complex 155 kDa subunit) (BRG1-associated factor 155). | |||||
|
NFH_HUMAN
|
||||||
| NC score | 0.056826 (rank : 83) | θ value | 0.47712 (rank : 33) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1242 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | P12036, Q9UJS7, Q9UQ14 | Gene names | NEFH, KIAA0845, NFH | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
PCLO_MOUSE
|
||||||
| NC score | 0.056212 (rank : 84) | θ value | 5.27518 (rank : 97) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
SMRC1_MOUSE
|
||||||
| NC score | 0.056190 (rank : 85) | θ value | θ > 10 (rank : 166) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 459 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P97496, Q7TS80, Q7TT29 | Gene names | Smarcc1, Baf155, Srg3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 1 (SWI/SNF complex 155 kDa subunit) (BRG1-associated factor 155) (SWI3-related protein). | |||||
|
SET1A_HUMAN
|
||||||
| NC score | 0.055752 (rank : 86) | θ value | θ > 10 (rank : 159) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | O15047, Q6PIF3, Q8TAJ6 | Gene names | SETD1A, KIAA0339, SET1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-4 specific SET1 (EC 2.1.1.43) (Set1/Ash2 histone methyltransferase complex subunit SET1) (SET-domain-containing protein 1A). | |||||
|
SYLC_MOUSE
|
||||||
| NC score | 0.055700 (rank : 87) | θ value | 0.813845 (rank : 44) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BMJ2, Q8BKW9 | Gene names | Lars | |||
|
Domain Architecture |

|
|||||
| Description | Leucyl-tRNA synthetase, cytoplasmic (EC 6.1.1.4) (Leucine--tRNA ligase) (LeuRS). | |||||
|
TTC16_HUMAN
|
||||||
| NC score | 0.055488 (rank : 88) | θ value | θ > 10 (rank : 178) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8NEE8, Q5JU66, Q96M72 | Gene names | TTC16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tetratricopeptide repeat protein 16 (TPR repeat protein 16). | |||||
|
NUCKS_HUMAN
|
||||||
| NC score | 0.055362 (rank : 89) | θ value | θ > 10 (rank : 153) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9H1E3, Q9H1D6, Q9H723 | Gene names | NUCKS1, NUCKS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear ubiquitous casein and cyclin-dependent kinases substrate (P1). | |||||
|
THOC2_HUMAN
|
||||||
| NC score | 0.055327 (rank : 90) | θ value | θ > 10 (rank : 175) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8NI27, Q9H8I6 | Gene names | THOC2, CXorf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | THO complex subunit 2 (Tho2). | |||||
|
BASP_HUMAN
|
||||||
| NC score | 0.055154 (rank : 91) | θ value | θ > 10 (rank : 133) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P80723, O43596, Q5U0S0 | Gene names | BASP1, NAP22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
SFR12_HUMAN
|
||||||
| NC score | 0.054895 (rank : 92) | θ value | θ > 10 (rank : 161) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 553 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8WXA9, Q86X37 | Gene names | SFRS12, SRRP86 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 12 (Serine-arginine-rich- splicing regulatory protein 86) (SRrp86) (Splicing regulatory protein 508) (SRrp508). | |||||
|
GP73_MOUSE
|
||||||
| NC score | 0.054683 (rank : 93) | θ value | 0.125558 (rank : 18) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q91XA2 | Gene names | Golph2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi phosphoprotein 2 (Golgi membrane protein GP73). | |||||
|
SFR12_MOUSE
|
||||||
| NC score | 0.054632 (rank : 94) | θ value | θ > 10 (rank : 162) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 486 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8BZX4, Q8BLM8, Q8BWK7, Q8BYK0, Q8BYZ4 | Gene names | Sfrs12, Srrp86 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 12 (Serine-arginine-rich- splicing regulatory protein 86) (SRrp86). | |||||
|
EDD1_HUMAN
|
||||||
| NC score | 0.054289 (rank : 95) | θ value | 0.47712 (rank : 31) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O95071, O94970, Q9NPL3 | Gene names | EDD1, EDD, HYD, KIAA0896 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-protein ligase EDD1 (EC 6.3.2.-) (Hyperplastic discs protein homolog) (hHYD) (Progestin-induced protein). | |||||
|
FILA_HUMAN
|
||||||
| NC score | 0.054274 (rank : 96) | θ value | θ > 10 (rank : 140) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 685 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P20930, Q01720, Q5T583 | Gene names | FLG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filaggrin. | |||||
|
RSF1_HUMAN
|
||||||
| NC score | 0.054155 (rank : 97) | θ value | θ > 10 (rank : 158) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 539 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q96T23, Q86X86, Q9H3L8, Q9NVZ8, Q9NYU0 | Gene names | RSF1, HBXAP, XAP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Remodeling and spacing factor 1 (Rsf-1) (Hepatitis B virus X- associated protein) (HBV pX-associated protein 8) (p325 subunit of RSF chromatin remodelling complex). | |||||
|
RL1D1_HUMAN
|
||||||
| NC score | 0.054017 (rank : 98) | θ value | θ > 10 (rank : 157) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O76021, Q6PL22, Q8IWS7, Q8WUZ1, Q9HDA9, Q9Y3Z9 | Gene names | RSL1D1, CATX11, CSIG, PBK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ribosomal L1 domain-containing protein 1 (Cellular senescence- inhibited gene protein) (Protein PBK1) (CATX-11). | |||||
|
MDN1_HUMAN
|
||||||
| NC score | 0.053859 (rank : 99) | θ value | θ > 10 (rank : 150) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 662 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9NU22, O15019 | Gene names | MDN1, KIAA0301 | |||
|
Domain Architecture |

|
|||||
| Description | Midasin (MIDAS-containing protein). | |||||
|
ATRX_MOUSE
|
||||||
| NC score | 0.053593 (rank : 100) | θ value | θ > 10 (rank : 132) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 682 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q61687 | Gene names | Atrx, Hp1bp2, Xnp | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ATRX (EC 3.6.1.-) (ATP-dependent helicase ATRX) (X-linked nuclear protein) (Heterochromatin protein 2) (HP1 alpha-interacting protein) (HP1-BP38 protein). | |||||
|
YTDC1_HUMAN
|
||||||
| NC score | 0.053436 (rank : 101) | θ value | θ > 10 (rank : 181) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q96MU7, Q7Z622, Q8TF35 | Gene names | YTHDC1, KIAA1966, YT521 | |||
|
Domain Architecture |

|
|||||
| Description | YTH domain-containing protein 1 (Putative splicing factor YT521). | |||||
|
NFH_MOUSE
|
||||||
| NC score | 0.053134 (rank : 102) | θ value | 5.27518 (rank : 95) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
PALB2_MOUSE
|
||||||
| NC score | 0.053024 (rank : 103) | θ value | 0.365318 (rank : 28) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q3U0P1, Q6NZG9, Q7TMQ4, Q8CEA9 | Gene names | Palb2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Partner and localizer of BRCA2. | |||||
|
EDD1_MOUSE
|
||||||
| NC score | 0.052657 (rank : 104) | θ value | 0.813845 (rank : 40) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q80TP3, Q698K9, Q6PEQ8, Q6PFQ9, Q80VL4, Q810V6, Q9CXE9 | Gene names | Edd1, Edd, Kiaa0896 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase EDD1 (EC 6.3.2.-) (Hyperplastic discs protein homolog). | |||||
|
NSBP1_MOUSE
|
||||||
| NC score | 0.052505 (rank : 105) | θ value | θ > 10 (rank : 152) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9JL35, O88832, Q8VC71, Q9CUW1 | Gene names | Nsbp1, Garp45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1 (Nucleosome-binding protein 45) (NBP-45) (GARP45 protein). | |||||
|
BASP_MOUSE
|
||||||
| NC score | 0.052478 (rank : 106) | θ value | θ > 10 (rank : 134) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q91XV3 | Gene names | Basp1, Nap22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
TRIPC_HUMAN
|
||||||
| NC score | 0.052313 (rank : 107) | θ value | 0.47712 (rank : 35) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 319 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q14669, Q15644 | Gene names | TRIP12, KIAA0045 | |||
|
Domain Architecture |

|
|||||
| Description | Thyroid receptor-interacting protein 12 (TRIP12). | |||||
|
MTMR7_HUMAN
|
||||||
| NC score | 0.051575 (rank : 108) | θ value | 0.813845 (rank : 42) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Y216, Q68DX4 | Gene names | MTMR7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myotubularin-related protein 7 (EC 3.1.3.-). | |||||
|
LRRF1_MOUSE
|
||||||
| NC score | 0.051384 (rank : 109) | θ value | θ > 10 (rank : 147) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 440 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q3UZ39, O70323, Q3TS94 | Gene names | Lrrfip1, Flap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat flightless-interacting protein 1 (LRR FLII- interacting protein 1) (FLI-LRR-associated protein 1) (Flap-1) (H186 FLAP). | |||||
|
GPTC8_HUMAN
|
||||||
| NC score | 0.051382 (rank : 110) | θ value | θ > 10 (rank : 141) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 366 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9UKJ3, O60300 | Gene names | GPATC8, KIAA0553 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G patch domain-containing protein 8. | |||||
|
NOP14_HUMAN
|
||||||
| NC score | 0.051180 (rank : 111) | θ value | 1.38821 (rank : 51) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 573 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P78316, Q7LGI5, Q7Z6K0, Q8TBR6 | Gene names | NOP14, C4orf9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable nucleolar complex protein 14. | |||||
|
UBP31_HUMAN
|
||||||
| NC score | 0.051068 (rank : 112) | θ value | 0.21417 (rank : 23) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 293 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q70CQ4, Q6AW97, Q6ZTC0, Q6ZTN2, Q9ULL7 | Gene names | USP31, KIAA1203 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 31 (EC 3.1.2.15) (Ubiquitin thioesterase 31) (Ubiquitin-specific-processing protease 31) (Deubiquitinating enzyme 31). | |||||
|
SFR17_HUMAN
|
||||||
| NC score | 0.050632 (rank : 113) | θ value | θ > 10 (rank : 163) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8TF01, Q5T064, Q6P2B4, Q6P2N4, Q6PJ93, Q6PK36, Q7Z640, Q8TF00, Q96K10, Q96SM5, Q9P076, Q9P0C0, Q9Y4N3 | Gene names | SRRP130, C6orf111 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, arginine/serine-rich 130 (Serine-arginine-rich- splicing regulatory protein 130) (SRrp130) (SR-rich protein) (SR- related protein). | |||||
|
OXR1_MOUSE
|
||||||
| NC score | 0.050464 (rank : 114) | θ value | θ > 10 (rank : 154) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 516 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q4KMM3, Q5FWW1, Q99L06, Q99MK1, Q99MP4 | Gene names | Oxr1, C7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Oxidation resistance protein 1 (Protein C7). | |||||
|
TNR6A_HUMAN
|
||||||
| NC score | 0.050422 (rank : 115) | θ value | θ > 10 (rank : 176) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8NDV7, O15408, Q658L5, Q6NVB5, Q8NEZ0, Q8TBT8, Q8TCR0, Q9NV59, Q9P268 | Gene names | TNRC6A, CAGH26, KIAA1460, TNRC6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trinucleotide repeat-containing gene 6A protein (CAG repeat protein 26) (Glycine-tryptophan protein of 182 kDa) (GW182 autoantigen) (Protein GW1) (EMSY interactor protein). | |||||
|
MLL3_MOUSE
|
||||||
| NC score | 0.050251 (rank : 116) | θ value | 3.0926 (rank : 76) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
ELOA1_HUMAN
|
||||||
| NC score | 0.049370 (rank : 117) | θ value | 5.27518 (rank : 87) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q14241, Q8IXH1 | Gene names | TCEB3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription elongation factor B polypeptide 3 (RNA polymerase II transcription factor SIII subunit A1) (SIII p110) (Elongin-A) (EloA) (Elongin 110 kDa subunit). | |||||
|
CDC6_MOUSE
|
||||||
| NC score | 0.048244 (rank : 118) | θ value | 1.81305 (rank : 55) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O89033, Q3TMD0, Q8C3S5, Q9CZT0 | Gene names | Cdc6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division control protein 6 homolog (CDC6-related protein) (p62(cdc6)). | |||||
|
BAZ2A_MOUSE
|
||||||
| NC score | 0.047709 (rank : 119) | θ value | 0.813845 (rank : 39) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q91YE5 | Gene names | Baz2a, Tip5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5). | |||||
|
MTMR7_MOUSE
|
||||||
| NC score | 0.046770 (rank : 120) | θ value | 2.36792 (rank : 64) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9Z2C9, Q8C4J6 | Gene names | Mtmr7 | |||
|
Domain Architecture |

|
|||||
| Description | Myotubularin-related protein 7 (EC 3.1.3.-). | |||||
|
SH24A_HUMAN
|
||||||
| NC score | 0.046723 (rank : 121) | θ value | 0.279714 (rank : 26) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9H788, Q5XKC1, Q6NXE9, Q86YM2, Q96C88, Q9H7F7 | Gene names | SH2D4A, SH2A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH2 domain-containing protein 4A (Protein SH(2)A). | |||||
|
AP3B2_HUMAN
|
||||||
| NC score | 0.046616 (rank : 122) | θ value | 1.38821 (rank : 49) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q13367, O14808 | Gene names | AP3B2 | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit beta-2 (Adapter-related protein complex 3 beta-2 subunit) (Beta3B-adaptin) (Adaptor protein complex AP-3 beta-2 subunit) (AP-3 complex beta-2 subunit) (Clathrin assembly protein complex 3 beta-2 large chain) (Neuron-specific vesicle coat protein beta-NAP). | |||||
|
SETX_HUMAN
|
||||||
| NC score | 0.046530 (rank : 123) | θ value | 2.36792 (rank : 70) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q7Z333, O75120, Q3KQX4, Q68DW5, Q6AZD7, Q7Z3J6, Q8WX33, Q9H9D1, Q9NVP9 | Gene names | SETX, ALS4, KIAA0625, SCAR1 | |||
|
Domain Architecture |

|
|||||
| Description | Probable helicase senataxin (EC 3.6.1.-) (SEN1 homolog). | |||||
|
RFC1_HUMAN
|
||||||
| NC score | 0.046350 (rank : 124) | θ value | 4.03905 (rank : 83) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P35251, Q5XKF5, Q6PKU0, Q86V41, Q86V46 | Gene names | RFC1, RFC140 | |||
|
Domain Architecture |

|
|||||
| Description | Replication factor C subunit 1 (Replication factor C large subunit) (RF-C 140 kDa subunit) (Activator 1 140 kDa subunit) (Activator 1 large subunit) (A1 140 kDa subunit) (DNA-binding protein PO-GA). | |||||
|
ERIC1_HUMAN
|
||||||
| NC score | 0.045104 (rank : 125) | θ value | 8.99809 (rank : 113) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q86X53, Q9P063 | Gene names | ERICH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate-rich protein 1. | |||||
|
SEMG1_HUMAN
|
||||||
| NC score | 0.044009 (rank : 126) | θ value | 1.81305 (rank : 59) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P04279, Q6X4I9, Q6Y809, Q6Y822, Q6Y823, Q86U64, Q96QM3 | Gene names | SEMG1, SEMG | |||
|
Domain Architecture |

|
|||||
| Description | Semenogelin-1 precursor (Semenogelin I) (SGI) [Contains: Alpha- inhibin-92; Alpha-inhibin-31; Seminal basic protein]. | |||||
|
CDC6_HUMAN
|
||||||
| NC score | 0.043701 (rank : 127) | θ value | 2.36792 (rank : 62) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99741, Q8TB30 | Gene names | CDC6, CDC18L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division control protein 6 homolog (CDC6-related protein) (p62(cdc6)) (HsCDC6) (HsCDC18). | |||||
|
MLL2_HUMAN
|
||||||
| NC score | 0.043358 (rank : 128) | θ value | 6.88961 (rank : 106) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
ENAH_MOUSE
|
||||||
| NC score | 0.043347 (rank : 129) | θ value | 0.62314 (rank : 38) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 919 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q03173, P70430, P70431, P70432, P70433, Q5D053 | Gene names | Enah, Mena, Ndpp1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein enabled homolog (NPC-derived proline-rich protein 1) (NDPP-1). | |||||
|
WDHD1_MOUSE
|
||||||
| NC score | 0.042374 (rank : 130) | θ value | 3.0926 (rank : 79) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 414 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P59328, Q6P408 | Gene names | Wdhd1, And1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WD repeat and HMG-box DNA-binding protein 1 (Acidic nucleoplasmic DNA- binding protein 1) (And-1). | |||||
|
SNX9_MOUSE
|
||||||
| NC score | 0.041346 (rank : 131) | θ value | 6.88961 (rank : 107) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q91VH2 | Gene names | Snx9 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-9. | |||||
|
AP3B1_HUMAN
|
||||||
| NC score | 0.040993 (rank : 132) | θ value | 6.88961 (rank : 102) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O00203, O00580, Q7Z393, Q9HD66 | Gene names | AP3B1, ADTB3A | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit beta-1 (Adapter-related protein complex 3 beta-1 subunit) (Beta3A-adaptin) (Adaptor protein complex AP-3 beta-1 subunit) (Clathrin assembly protein complex 3 beta-1 large chain). | |||||
|
ZMYM5_HUMAN
|
||||||
| NC score | 0.040364 (rank : 133) | θ value | 1.38821 (rank : 53) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UJ78, Q5T6E2, Q96IY6, Q9NZY5, Q9UBW0, Q9UJ77 | Gene names | ZMYM5, ZNF198L1, ZNF237 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger MYM-type protein 5 (Zinc finger protein 237) (Zinc finger protein 198-like 1). | |||||
|
RBMX2_HUMAN
|
||||||
| NC score | 0.039592 (rank : 134) | θ value | 8.99809 (rank : 121) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9Y388, Q5JY82, Q9Y3I8 | Gene names | RBMX2 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding motif protein, X-linked 2. | |||||
|
LYAR_MOUSE
|
||||||
| NC score | 0.038651 (rank : 135) | θ value | 8.99809 (rank : 118) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q08288, Q9D9X2 | Gene names | Lyar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell growth-regulating nucleolar protein. | |||||
|
RAD18_MOUSE
|
||||||
| NC score | 0.037503 (rank : 136) | θ value | 4.03905 (rank : 82) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9QXK2, Q9CZB8 | Gene names | Rad18, Rad18sc | |||
|
Domain Architecture |

|
|||||
| Description | Postreplication repair protein RAD18 (mRAD18Sc). | |||||
|
TSH2_MOUSE
|
||||||
| NC score | 0.036619 (rank : 137) | θ value | 5.27518 (rank : 99) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q68FE9, Q5DTF1, Q6AXH5, Q9JL71 | Gene names | Tshz2, Kiaa4248, Sdccag33l, Tsh2, Zfp218, Znf218 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Teashirt homolog 2 (Zinc finger protein 218) (SDCCAG33-like protein). | |||||
|
ZCH10_MOUSE
|
||||||
| NC score | 0.036164 (rank : 138) | θ value | 8.99809 (rank : 126) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9CX48 | Gene names | Zcchc10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 10. | |||||
|
LAP2_HUMAN
|
||||||
| NC score | 0.034349 (rank : 139) | θ value | 0.279714 (rank : 24) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 499 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q96RT1, Q86W38, Q9NR18, Q9NW48, Q9ULJ5 | Gene names | ERBB2IP, ERBIN, KIAA1225, LAP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LAP2 (Erbb2-interacting protein) (Erbin) (Densin-180-like protein). | |||||
|
LAP2_MOUSE
|
||||||
| NC score | 0.033111 (rank : 140) | θ value | 0.0736092 (rank : 16) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 482 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q80TH2, Q8BQ14, Q8CE41, Q8K171, Q99JU3, Q9JI47 | Gene names | Erbb2ip, Erbin, Kiaa1225, Lap2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein LAP2 (Erbb2-interacting protein) (Erbin) (Densin-180-like protein). | |||||
|
CEP57_MOUSE
|
||||||
| NC score | 0.032892 (rank : 141) | θ value | 0.365318 (rank : 27) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 580 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8CEE0, Q6ZQJ3, Q7TN18, Q80X65, Q810F2, Q9D4J4, Q9D5S4, Q9D5W5 | Gene names | Cep57, Kiaa0092, Tsp57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 57 kDa (Cep57 protein) (Testis-specific protein 57) (Translokin). | |||||
|
GASP2_MOUSE
|
||||||
| NC score | 0.030421 (rank : 142) | θ value | 5.27518 (rank : 90) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8BUY8 | Gene names | Gprasp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G-protein coupled receptor-associated sorting protein 2 (GASP-2). | |||||
|
NEB1_HUMAN
|
||||||
| NC score | 0.030260 (rank : 143) | θ value | 2.36792 (rank : 65) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 829 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9ULJ8, O76059, Q9NXT2 | Gene names | PPP1R9A, KIAA1222 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neurabin-1 (Neurabin-I) (Neural tissue-specific F-actin-binding protein I) (Protein phosphatase 1 regulatory subunit 9A). | |||||
|
PKHA6_MOUSE
|
||||||
| NC score | 0.029805 (rank : 144) | θ value | 4.03905 (rank : 81) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 591 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q7TQG1, Q8K0J5 | Gene names | Plekha6, Pepp3 | |||
|
Domain Architecture |

|
|||||
| Description | Pleckstrin homology domain-containing family A member 6 (Phosphoinositol 3-phosphate-binding protein 3) (PEPP-3). | |||||
|
K0586_HUMAN
|
||||||
| NC score | 0.029131 (rank : 145) | θ value | 5.27518 (rank : 91) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9BVV6, O60328 | Gene names | KIAA0586 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0586. | |||||
|
TERF1_HUMAN
|
||||||
| NC score | 0.027149 (rank : 146) | θ value | 8.99809 (rank : 124) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P54274, Q15553, Q93029 | Gene names | TERF1, PIN2, TRF, TRF1 | |||
|
Domain Architecture |

|
|||||
| Description | Telomeric repeat-binding factor 1 (TTAGGG repeat-binding factor 1) (NIMA-interacting protein 2) (Telomeric protein Pin2/TRF1). | |||||
|
PHF2_HUMAN
|
||||||
| NC score | 0.026967 (rank : 147) | θ value | 2.36792 (rank : 66) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O75151, Q8N3K2, Q9Y6N4 | Gene names | PHF2, KIAA0662 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 2 (GRC5). | |||||
|
YTHD3_HUMAN
|
||||||
| NC score | 0.026719 (rank : 148) | θ value | 1.81305 (rank : 60) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q7Z739, Q63Z37, Q659A3 | Gene names | YTHDF3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YTH domain family protein 3. | |||||
|
REV1_MOUSE
|
||||||
| NC score | 0.026528 (rank : 149) | θ value | 2.36792 (rank : 68) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q920Q2, Q9QXV2 | Gene names | Rev1l, Rev1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein REV1 (EC 2.7.7.-) (Rev1-like terminal deoxycytidyl transferase). | |||||
|
TSH2_HUMAN
|
||||||
| NC score | 0.026343 (rank : 150) | θ value | 5.27518 (rank : 98) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9NRE2, Q4VXM4, Q6N003, Q8N260 | Gene names | TSHZ2, C20orf17, TSH2, ZNF218 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Teashirt homolog 2 (Zinc finger protein 218) (Ovarian cancer-related protein 10-2) (OVC10-2). | |||||
|
ANLN_HUMAN
|
||||||
| NC score | 0.025137 (rank : 151) | θ value | 8.99809 (rank : 108) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9NQW6, Q5CZ78, Q6NSK5, Q9H8Y4, Q9NVN9, Q9NVP0 | Gene names | ANLN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Actin-binding protein anillin. | |||||
|
TTC16_MOUSE
|
||||||
| NC score | 0.024040 (rank : 152) | θ value | 4.03905 (rank : 85) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8C1F5 | Gene names | Ttc16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tetratricopeptide repeat protein 16 (TPR repeat protein 16). | |||||
|
PK1IP_MOUSE
|
||||||
| NC score | 0.023803 (rank : 153) | θ value | 2.36792 (rank : 67) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9DCE5, Q3UBK7, Q80UT4, Q8C5N6, Q923K2, Q9D3E9 | Gene names | Pak1ip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | p21-activated protein kinase-interacting protein 1 (PAK1-interacting protein 1) (Putative PAK inhibitor Skb15). | |||||
|
CI039_HUMAN
|
||||||
| NC score | 0.023328 (rank : 154) | θ value | 0.62314 (rank : 37) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1171 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9NXG0, Q5VYJ0, Q9HAJ5 | Gene names | C9orf39 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C9orf39. | |||||
|
ZC11A_MOUSE
|
||||||
| NC score | 0.021879 (rank : 155) | θ value | 8.99809 (rank : 125) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q6NZF1, Q6NXW9, Q6PDR6, Q80TU7, Q8C5L5, Q99JN6 | Gene names | Zc3h11a, Kiaa0663 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 11A. | |||||
|
EMIL2_MOUSE
|
||||||
| NC score | 0.021105 (rank : 156) | θ value | 5.27518 (rank : 88) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 485 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8K482 | Gene names | Emilin2 | |||
|
Domain Architecture |

|
|||||
| Description | EMILIN-2 precursor (Elastin microfibril interface-located protein 2) (Elastin microfibril interfacer 2) (Basilin). | |||||
|
JPH2_MOUSE
|
||||||
| NC score | 0.019736 (rank : 157) | θ value | 6.88961 (rank : 105) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9ET78, Q9ET79 | Gene names | Jph2, Jp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Junctophilin-2 (Junctophilin type 2) (JP-2). | |||||
|
CXXC6_HUMAN
|
||||||
| NC score | 0.019398 (rank : 158) | θ value | 8.99809 (rank : 112) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8NFU7, Q5VUP7, Q7Z6B6, Q8TCR1, Q9C0I7 | Gene names | CXXC6, KIAA1676, LCX | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CXXC-type zinc finger protein 6 (Leukemia-associated protein with a CXXC domain). | |||||
|
CSPG2_MOUSE
|
||||||
| NC score | 0.019370 (rank : 159) | θ value | 1.38821 (rank : 50) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 711 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q62059, Q62058, Q9CUU0 | Gene names | Cspg2 | |||
|
Domain Architecture |

|
|||||
| Description | Versican core protein precursor (Large fibroblast proteoglycan) (Chondroitin sulfate proteoglycan core protein 2) (PG-M). | |||||
|
SMRCD_HUMAN
|
||||||
| NC score | 0.016859 (rank : 160) | θ value | 2.36792 (rank : 71) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9H4L7, Q96SX1, Q9H017, Q9H860, Q9NPU9, Q9ULU7 | Gene names | SMARCAD1, KIAA1122 | |||
|
Domain Architecture |

|
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A containing DEAD/H box 1 (EC 3.6.1.-) (ATP- dependent helicase 1) (hHEL1). | |||||
|
PARP1_MOUSE
|
||||||
| NC score | 0.016595 (rank : 161) | θ value | 5.27518 (rank : 96) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P11103, Q9JLX4, Q9QVQ3 | Gene names | Parp1, Adprp, Adprt, Adprt1 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 1 (EC 2.4.2.30) (PARP-1) (ADPRT) (NAD(+) ADP-ribosyltransferase 1) (Poly[ADP-ribose] synthetase 1) (msPARP). | |||||
|
HUCE1_MOUSE
|
||||||
| NC score | 0.016402 (rank : 162) | θ value | 8.99809 (rank : 116) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9DB85, Q8BHW3 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cerebral protein 1 homolog. | |||||
|
HRB2_MOUSE
|
||||||
| NC score | 0.015776 (rank : 163) | θ value | 8.99809 (rank : 115) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BGA5, Q8C029 | Gene names | Hrb2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HIV-1 Rev-binding protein 2 homolog. | |||||
|
SMC2_MOUSE
|
||||||
| NC score | 0.015160 (rank : 164) | θ value | 4.03905 (rank : 84) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1131 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8CG48, Q61076, Q9CS17, Q9CSD8 | Gene names | Smc2l1, Cape, Fin16, Smc2 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 2-like 1 protein (Chromosome- associated protein E) (XCAP-E homolog) (FGF-inducible protein 16). | |||||
|
F120C_HUMAN
|
||||||
| NC score | 0.014110 (rank : 165) | θ value | 6.88961 (rank : 104) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NX05 | Gene names | FAM120C, CXorf17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UPF0318 protein FAM120C (Tumor antigen BJ-HCC-21). | |||||
|
SRPR_HUMAN
|
||||||
| NC score | 0.014062 (rank : 166) | θ value | 8.99809 (rank : 122) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P08240, Q9BVJ4 | Gene names | SRPR | |||
|
Domain Architecture |

|
|||||
| Description | Signal recognition particle receptor subunit alpha (SR-alpha) (Docking protein alpha) (DP-alpha). | |||||
|
CUTL2_HUMAN
|
||||||
| NC score | 0.013079 (rank : 167) | θ value | 8.99809 (rank : 111) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O14529 | Gene names | CUTL2, KIAA0293 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 2 (Homeobox protein Cux-2) (Cut-like 2). | |||||
|
ARHG7_MOUSE
|
||||||
| NC score | 0.012216 (rank : 168) | θ value | 3.0926 (rank : 74) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 396 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9ES28, O08757, Q9ES27 | Gene names | Arhgef7, Pak3bp | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 7 (PAK-interacting exchange factor beta) (Beta-Pix) (p85SPR). | |||||
|
MYLK_MOUSE
|
||||||
| NC score | 0.011291 (rank : 169) | θ value | 1.81305 (rank : 57) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1489 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q6PDN3, Q3TSJ7, Q80UX0, Q80YN7, Q80YN8, Q8K026, Q924D2, Q9ERD3 | Gene names | Mylk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin light chain kinase, smooth muscle (EC 2.7.11.18) (MLCK) (Telokin) (Kinase-related protein) (KRP). | |||||
|
SCN5A_HUMAN
|
||||||
| NC score | 0.010768 (rank : 170) | θ value | 2.36792 (rank : 69) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q14524 | Gene names | SCN5A | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 5 subunit alpha (Sodium channel protein type V subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.5) (Sodium channel protein, cardiac muscle alpha-subunit) (HH1). | |||||
|
BNC1_MOUSE
|
||||||
| NC score | 0.010103 (rank : 171) | θ value | 6.88961 (rank : 103) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 582 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O35914, O54886 | Gene names | Bnc1, Bnc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein basonuclin-1. | |||||
|
CDYL_MOUSE
|
||||||
| NC score | 0.009054 (rank : 172) | θ value | 8.99809 (rank : 110) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9WTK2 | Gene names | Cdyl | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain Y-like protein (CDY-like). | |||||
|
LIPB2_HUMAN
|
||||||
| NC score | 0.008817 (rank : 173) | θ value | 8.99809 (rank : 117) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 599 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8ND30, O75337, Q8WW26 | Gene names | PPFIBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-beta-2 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein-binding protein 2) (PTPRF-interacting protein-binding protein 2). | |||||
|
ANKR6_HUMAN
|
||||||
| NC score | 0.008544 (rank : 174) | θ value | 3.0926 (rank : 73) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 439 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Y2G4, Q9NU24, Q9UFQ9 | Gene names | ANKRD6, KIAA0957 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin repeat domain-containing protein 6. | |||||
|
PA24B_HUMAN
|
||||||
| NC score | 0.008273 (rank : 175) | θ value | 8.99809 (rank : 120) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O95712, Q59GF9, Q8TB10, Q9UKV7 | Gene names | PLA2G4B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytosolic phospholipase A2 beta (EC 3.1.1.4) (cPLA2-beta) (Phospholipase A2 group IVB). | |||||
|
FNBP2_HUMAN
|
||||||
| NC score | 0.008086 (rank : 176) | θ value | 8.99809 (rank : 114) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 432 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O75044 | Gene names | SRGAP2, FNBP2, KIAA0456 | |||
|
Domain Architecture |

|
|||||
| Description | SLIT-ROBO Rho GTPase-activating protein 2 (srGAP2) (Formin-binding protein 2). | |||||
|
FBLN2_MOUSE
|
||||||
| NC score | 0.007573 (rank : 177) | θ value | 5.27518 (rank : 89) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 505 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P37889, Q9WUI2 | Gene names | Fbln2 | |||
|
Domain Architecture |

|
|||||
| Description | Fibulin-2 precursor. | |||||
|
ARX_MOUSE
|
||||||
| NC score | 0.007476 (rank : 178) | θ value | 2.36792 (rank : 61) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 398 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O35085, Q9QYT4 | Gene names | Arx | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein ARX (Aristaless-related homeobox). | |||||
|
ASB1_HUMAN
|
||||||
| NC score | 0.004849 (rank : 179) | θ value | 8.99809 (rank : 109) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 291 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y576, Q9ULS4 | Gene names | ASB1, KIAA1146 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin repeat and SOCS box protein 1 (ASB-1). | |||||
|
TENX_HUMAN
|
||||||
| NC score | 0.001971 (rank : 180) | θ value | 8.99809 (rank : 123) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 789 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P22105, P78530, P78531, Q08424, Q9UMG7 | Gene names | TNXB, HXBL, TNX, XB | |||
|
Domain Architecture |

|
|||||
| Description | Tenascin-X precursor (TN-X) (Hexabrachion-like protein). | |||||
|
YSK4_HUMAN
|
||||||
| NC score | 0.000753 (rank : 181) | θ value | 5.27518 (rank : 100) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 921 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q56UN5, Q56UN1, Q56UN2, Q56UN3, Q56UN4, Q8N4E9, Q9H5T2 | Gene names | YSK4, RCK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SPS1/STE20-related protein kinase YSK4 (EC 2.7.11.1) (Regulated in COPD, protein kinase). | |||||