Please be patient as the page loads
|
ZCHC6_MOUSE
|
||||||
| SwissProt Accessions | Q5BLK4, Q8CIH3 | Gene names | Zcchc6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 6. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
ZCHC6_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.979667 (rank : 2) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q5VYS8, Q5H9T0, Q5VYS5, Q5VYS7, Q658Z9, Q659A2, Q6MZJ3, Q8N5F0, Q96N57, Q96NE8, Q9C0F2, Q9H8M6 | Gene names | ZCCHC6, KIAA1711 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 6. | |||||
|
ZCHC6_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 90 | |
| SwissProt Accessions | Q5BLK4, Q8CIH3 | Gene names | Zcchc6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 6. | |||||
|
ZCH11_HUMAN
|
||||||
| θ value | 9.45368e-175 (rank : 3) | NC score | 0.870640 (rank : 3) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q5TAX3, Q12764, Q5TAX4, Q86XZ3 | Gene names | ZCCHC11, KIAA0191 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 11. | |||||
|
PAPD1_HUMAN
|
||||||
| θ value | 4.29542e-18 (rank : 4) | NC score | 0.552802 (rank : 5) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9NVV4, Q659E3, Q6P7E5, Q9HA74 | Gene names | PAPD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly(A) RNA polymerase, mitochondrial precursor (EC 2.7.7.19) (PAP) (mtPAP) (Polynucleotide adenylyltransferase) (PAP-associated domain- containing protein 1). | |||||
|
PAPD1_MOUSE
|
||||||
| θ value | 4.29542e-18 (rank : 5) | NC score | 0.564380 (rank : 4) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9D0D3, Q3UXJ1, Q8C651 | Gene names | Papd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly(A) RNA polymerase, mitochondrial precursor (EC 2.7.7.19) (PAP) (Polynucleotide adenylyltransferase) (PAP-associated domain-containing protein 1). | |||||
|
PAD5_MOUSE
|
||||||
| θ value | 2.52405e-10 (rank : 6) | NC score | 0.396870 (rank : 7) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q68ED3, Q8C0K6 | Gene names | Papd5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PAP-associated domain-containing protein 5 (EC 2.7.7.-) (Topoisomerase-related function protein 4-2) (TRF4-2). | |||||
|
PAPD5_HUMAN
|
||||||
| θ value | 2.52405e-10 (rank : 7) | NC score | 0.399735 (rank : 6) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8NDF8, Q9NW67, Q9Y6C0 | Gene names | PAPD5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PAP-associated domain-containing protein 5 (EC 2.7.7.-) (Topoisomerase-related function protein 4-2) (TRF4-2). | |||||
|
POLS_HUMAN
|
||||||
| θ value | 6.21693e-09 (rank : 8) | NC score | 0.386165 (rank : 8) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q5XG87, O43289, Q9Y6C1 | Gene names | POLS, TRF4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA polymerase sigma (EC 2.7.7.7) (Topoisomerase-related function protein 4-1) (TRF4-1) (LAK-1) (DNA polymerase kappa). | |||||
|
POLS_MOUSE
|
||||||
| θ value | 6.21693e-09 (rank : 9) | NC score | 0.379497 (rank : 9) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q6PB75 | Gene names | Pols | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA polymerase sigma (EC 2.7.7.7). | |||||
|
CCD96_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 10) | NC score | 0.043394 (rank : 24) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9CR92 | Gene names | Ccdc96 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 96. | |||||
|
G6PE_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 11) | NC score | 0.063572 (rank : 18) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O95479, Q4TT33, Q66I35, Q68DT3 | Gene names | H6PD, GDH | |||
|
Domain Architecture |

|
|||||
| Description | GDH/6PGL endoplasmic bifunctional protein precursor [Includes: Glucose 1-dehydrogenase (EC 1.1.1.47) (Hexose-6-phosphate dehydrogenase); 6- phosphogluconolactonase (EC 3.1.1.31) (6PGL)]. | |||||
|
TEX14_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 12) | NC score | 0.020439 (rank : 61) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1103 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8IWB6, Q7RTP3, Q8ND97, Q9BXT9 | Gene names | TEX14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Testis-expressed protein 14 (Testis-expressed sequence 14). | |||||
|
ZCHC7_HUMAN
|
||||||
| θ value | 0.125558 (rank : 13) | NC score | 0.123660 (rank : 13) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8N3Z6, Q5T0Q8, Q5T0Q9, Q5T0R0, Q8N2M1, Q8N4J2, Q8TBK8, Q9H648, Q9P0F0 | Gene names | ZCCHC7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 7. | |||||
|
PAPOG_HUMAN
|
||||||
| θ value | 0.163984 (rank : 14) | NC score | 0.078121 (rank : 14) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9BWT3, Q969N1, Q9H8L2, Q9HAD0 | Gene names | PAPOLG, PAP2, PAPG | |||
|
Domain Architecture |

|
|||||
| Description | Poly(A) polymerase gamma (EC 2.7.7.19) (PAP gamma) (Polynucleotide adenylyltransferase gamma) (SRP RNA 3' adenylating enzyme) (Neo- poly(A) polymerase) (Neo-PAP). | |||||
|
PAPOG_MOUSE
|
||||||
| θ value | 0.163984 (rank : 15) | NC score | 0.077374 (rank : 15) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6PCL9, Q8BZC9 | Gene names | Papolg | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly(A) polymerase gamma (EC 2.7.7.19) (PAP gamma) (Polynucleotide adenylyltransferase gamma) (SRP RNA 3' adenylating enzyme). | |||||
|
CENB5_HUMAN
|
||||||
| θ value | 0.279714 (rank : 16) | NC score | 0.022985 (rank : 58) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96P50, Q9BSR9, Q9C0E7 | Gene names | CENTB5, KIAA1716 | |||
|
Domain Architecture |

|
|||||
| Description | Centaurin-beta 5 (Cnt-b5). | |||||
|
K0555_HUMAN
|
||||||
| θ value | 0.279714 (rank : 17) | NC score | 0.033903 (rank : 44) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 880 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q96AA8, O60302 | Gene names | KIAA0555 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0555. | |||||
|
ZCH13_HUMAN
|
||||||
| θ value | 0.279714 (rank : 18) | NC score | 0.138119 (rank : 12) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8WW36 | Gene names | ZCCHC13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 13. | |||||
|
LEO1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 19) | NC score | 0.042477 (rank : 25) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
CNBP_HUMAN
|
||||||
| θ value | 0.62314 (rank : 20) | NC score | 0.151778 (rank : 10) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P62633, P20694, Q5QJR0, Q6PJI7, Q96NV3 | Gene names | CNBP, ZNF9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cellular nucleic acid-binding protein (CNBP) (Zinc finger protein 9). | |||||
|
CNBP_MOUSE
|
||||||
| θ value | 0.62314 (rank : 21) | NC score | 0.142584 (rank : 11) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P53996, Q80Y06, Q8BP23 | Gene names | Cnbp, Cnbp1, Znf9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cellular nucleic acid-binding protein (CNBP) (Zinc finger protein 9). | |||||
|
PLCB3_HUMAN
|
||||||
| θ value | 0.813845 (rank : 22) | NC score | 0.019666 (rank : 62) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q01970 | Gene names | PLCB3 | |||
|
Domain Architecture |

|
|||||
| Description | 1-phosphatidylinositol-4,5-bisphosphate phosphodiesterase beta 3 (EC 3.1.4.11) (Phosphoinositide phospholipase C) (Phospholipase C- beta-3) (PLC-beta-3). | |||||
|
RBM6_HUMAN
|
||||||
| θ value | 1.06291 (rank : 23) | NC score | 0.032120 (rank : 46) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P78332, O60549, O75524 | Gene names | RBM6, DEF3 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 6 (RNA-binding motif protein 6) (RNA-binding protein DEF-3) (Lung cancer antigen NY-LU-12) (Protein G16). | |||||
|
BAZ1B_MOUSE
|
||||||
| θ value | 1.38821 (rank : 24) | NC score | 0.028206 (rank : 55) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9Z277, Q9CU68 | Gene names | Baz1b, Wbscr9 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1B (Williams-Beuren syndrome chromosome region 9 protein homolog) (WBRS9). | |||||
|
BRPF1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 25) | NC score | 0.018089 (rank : 65) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P55201, Q9UHI0 | Gene names | BRPF1, BR140 | |||
|
Domain Architecture |

|
|||||
| Description | Peregrin (Bromodomain and PHD finger-containing protein 1) (BR140 protein). | |||||
|
SODC_MOUSE
|
||||||
| θ value | 1.38821 (rank : 26) | NC score | 0.029669 (rank : 52) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P08228 | Gene names | Sod1 | |||
|
Domain Architecture |
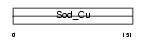
|
|||||
| Description | Superoxide dismutase [Cu-Zn] (EC 1.15.1.1). | |||||
|
BAZ1B_HUMAN
|
||||||
| θ value | 1.81305 (rank : 27) | NC score | 0.026083 (rank : 56) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 701 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9UIG0, O95039, O95247, O95277 | Gene names | BAZ1B, WBSC10, WBSCR9, WSTF | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1B (Williams-Beuren syndrome chromosome region 9 protein) (Williams syndrome transcription factor) (hWALP2). | |||||
|
HIP1R_MOUSE
|
||||||
| θ value | 1.81305 (rank : 28) | NC score | 0.012491 (rank : 72) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 959 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9JKY5 | Gene names | Hip1r | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin-interacting protein 1-related protein (Hip1-related). | |||||
|
RBM4_HUMAN
|
||||||
| θ value | 1.81305 (rank : 29) | NC score | 0.036016 (rank : 39) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9BWF3, O02916, Q6P1P2, Q8WU85 | Gene names | RBM4, RBM4A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein 4 (RNA-binding motif protein 4) (RNA-binding motif protein 4a) (Lark homolog) (hLark). | |||||
|
RBM4_MOUSE
|
||||||
| θ value | 1.81305 (rank : 30) | NC score | 0.036005 (rank : 40) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8C7Q4, O08752, Q8BN66 | Gene names | Rbm4, Rbm4a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein 4 (RNA-binding motif protein 4) (RNA-binding motif protein 4a) (Lark homolog) (mLark). | |||||
|
TRAT1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 31) | NC score | 0.051249 (rank : 20) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6PIZ9, Q9NZX5 | Gene names | TRAT1, TCRIM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | T-cell receptor-associated transmembrane adapter 1 (T-cell receptor- interacting molecule) (TRIM) (pp29/30). | |||||
|
CCD96_HUMAN
|
||||||
| θ value | 2.36792 (rank : 32) | NC score | 0.029843 (rank : 50) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 595 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q2M329, Q8N2I7 | Gene names | CCDC96 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 96. | |||||
|
FGD6_MOUSE
|
||||||
| θ value | 2.36792 (rank : 33) | NC score | 0.010046 (rank : 75) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q69ZL1, Q8C8W5, Q8K3B0, Q9D3Y7 | Gene names | Fgd6, Kiaa1362 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 6. | |||||
|
GAK2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 34) | NC score | 0.045261 (rank : 21) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q7LDI9, Q9UKH5, Q9Y6I1, Q9YNA6, Q9YNB0 | Gene names | ERVK6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_7p22.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K(HML-2.HOM) Gag protein) (HERV-K108 Gag protein) (HERV-K(C7) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
MDN1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 35) | NC score | 0.032245 (rank : 45) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 662 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9NU22, O15019 | Gene names | MDN1, KIAA0301 | |||
|
Domain Architecture |

|
|||||
| Description | Midasin (MIDAS-containing protein). | |||||
|
NUFP2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 36) | NC score | 0.024583 (rank : 57) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7Z417, Q9P2M5 | Gene names | NUFIP2, KIAA1321, PIG1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear fragile X mental retardation-interacting protein 2 (FMRP- interacting protein 2) (82 kDa FMRP-interacting protein) (82-FIP) (Proliferation-inducing gene 1 protein). | |||||
|
BSN_MOUSE
|
||||||
| θ value | 3.0926 (rank : 37) | NC score | 0.021087 (rank : 60) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
GPDM_MOUSE
|
||||||
| θ value | 3.0926 (rank : 38) | NC score | 0.035730 (rank : 42) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q64521, Q61507 | Gene names | Gpd2, Gdm1 | |||
|
Domain Architecture |

|
|||||
| Description | Glycerol-3-phosphate dehydrogenase, mitochondrial precursor (EC 1.1.99.5) (GPD-M) (GPDH-M). | |||||
|
NCOA3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 39) | NC score | 0.012367 (rank : 73) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y6Q9, Q5JYD9, Q5JYE0, Q9BR49, Q9UPC9, Q9UPG4, Q9UPG7 | Gene names | NCOA3, AIB1, RAC3, TRAM1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP-interacting protein) (pCIP). | |||||
|
TAF1L_HUMAN
|
||||||
| θ value | 3.0926 (rank : 40) | NC score | 0.039305 (rank : 30) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8IZX4 | Gene names | TAF1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID 210 kDa subunit (TBP-associated factor 210 kDa) (TAF(II)210) (TBP-associated factor 1-like). | |||||
|
TAF1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 41) | NC score | 0.042466 (rank : 26) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P21675, Q6IUZ1 | Gene names | TAF1, BA2R, CCG1, TAF2A | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 1 (EC 2.7.11.1) (Transcription initiation factor TFIID 250 kDa subunit) (TAF(II)250) (TAFII-250) (TAFII250) (TBP-associated factor 250 kDa) (p250) (Cell cycle gene 1 protein). | |||||
|
CF152_MOUSE
|
||||||
| θ value | 4.03905 (rank : 42) | NC score | 0.015156 (rank : 66) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 597 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q80ST9, Q9CYM9, Q9D5J9 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf152 homolog. | |||||
|
GAK18_HUMAN
|
||||||
| θ value | 4.03905 (rank : 43) | NC score | 0.040383 (rank : 29) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P62690 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_22q11.23 provirus ancestral Gag polyprotein (Gag polyprotein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK8_HUMAN
|
||||||
| θ value | 4.03905 (rank : 44) | NC score | 0.043599 (rank : 23) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9HDB9 | Gene names | ERVK5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_3q12.3 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K(II) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GSUB_HUMAN
|
||||||
| θ value | 4.03905 (rank : 45) | NC score | 0.055913 (rank : 19) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 3 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O96001, Q9UDQ0 | Gene names | GSBS, C7orf16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G-substrate. | |||||
|
RBCC1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 46) | NC score | 0.008694 (rank : 76) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1048 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9ESK9, Q3TXX2, Q61384, Q8BRY9, Q9JK14 | Gene names | Rb1cc1, Cc1, Kiaa0203 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RB1-inducible coiled-coil protein 1 (Coiled-coil-forming protein 1) (LaXp180 protein). | |||||
|
RBM4B_HUMAN
|
||||||
| θ value | 4.03905 (rank : 47) | NC score | 0.031505 (rank : 47) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9BQ04 | Gene names | RBM4B, RBM30 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein 4B (RNA-binding motif protein 4B) (RNA-binding protein 30) (RNA-binding motif protein 30). | |||||
|
RBM4B_MOUSE
|
||||||
| θ value | 4.03905 (rank : 48) | NC score | 0.031481 (rank : 48) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8VE92 | Gene names | Rbm4b, Rbm30 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein 4B (RNA-binding motif protein 4B) (RNA-binding protein 30) (RNA-binding motif protein 30). | |||||
|
ZCHC8_HUMAN
|
||||||
| θ value | 4.03905 (rank : 49) | NC score | 0.041311 (rank : 28) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q6NZY4, Q7L2P6, Q8N2K5, Q96SK7, Q9NSS2, Q9NSS3 | Gene names | ZCCHC8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 8. | |||||
|
BC11B_HUMAN
|
||||||
| θ value | 5.27518 (rank : 50) | NC score | 0.003096 (rank : 85) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 761 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9C0K0, Q9H162 | Gene names | BCL11B, CTIP2, RIT1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma/leukemia 11B (B-cell CLL/lymphoma 11B) (Radiation- induced tumor suppressor gene 1 protein) (hRit1) (COUP-TF-interacting protein 2). | |||||
|
CD2L1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 51) | NC score | 0.001433 (rank : 86) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1463 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P24788, Q3UI03, Q61399, Q7TST4, Q8BP53 | Gene names | Cdc2l1 | |||
|
Domain Architecture |

|
|||||
| Description | PITSLRE serine/threonine-protein kinase CDC2L1 (EC 2.7.11.22) (Galactosyltransferase-associated protein kinase p58/GTA) (Cell division cycle 2-like protein kinase 1). | |||||
|
GPDM_HUMAN
|
||||||
| θ value | 5.27518 (rank : 52) | NC score | 0.030412 (rank : 49) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P43304, Q59FR1, Q9HAP9 | Gene names | GPD2 | |||
|
Domain Architecture |

|
|||||
| Description | Glycerol-3-phosphate dehydrogenase, mitochondrial precursor (EC 1.1.99.5) (GPD-M) (GPDH-M) (mtGPD). | |||||
|
IWS1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 53) | NC score | 0.013865 (rank : 68) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
MYLK_HUMAN
|
||||||
| θ value | 5.27518 (rank : 54) | NC score | -0.000361 (rank : 88) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1328 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q15746, O95796, O95797, O95798, O95799, Q14844, Q16794, Q5MY99, Q5MYA0, Q7Z4J0, Q9C0L5, Q9UBG5, Q9UIT9 | Gene names | MYLK, MLCK | |||
|
Domain Architecture |

|
|||||
| Description | Myosin light chain kinase, smooth muscle (EC 2.7.11.18) (MLCK) (Telokin) (Kinase-related protein) (KRP). | |||||
|
NIN_HUMAN
|
||||||
| θ value | 5.27518 (rank : 55) | NC score | 0.007398 (rank : 79) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1416 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8N4C6, Q6P0P6, Q9BWU6, Q9C012, Q9C013, Q9C014, Q9H5I6, Q9HAT7, Q9HBY5, Q9HCK7, Q9UH61 | Gene names | NIN, KIAA1565 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ninein (hNinein) (Glycogen synthase kinase 3 beta-interacting protein) (GSK3B-interacting protein). | |||||
|
NMDE1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 56) | NC score | 0.008652 (rank : 77) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q12879, O00669 | Gene names | GRIN2A | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate [NMDA] receptor subunit epsilon 1 precursor (N-methyl D- aspartate receptor subtype 2A) (NR2A) (NMDAR2A) (hNR2A). | |||||
|
ZCHC9_HUMAN
|
||||||
| θ value | 5.27518 (rank : 57) | NC score | 0.074066 (rank : 16) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8N567, Q9H027 | Gene names | ZCCHC9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 9. | |||||
|
ZCHC9_MOUSE
|
||||||
| θ value | 5.27518 (rank : 58) | NC score | 0.069376 (rank : 17) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8R1J3, Q921T6 | Gene names | Zcchc9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 9. | |||||
|
ZN291_HUMAN
|
||||||
| θ value | 5.27518 (rank : 59) | NC score | 0.028775 (rank : 54) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 647 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9BY12 | Gene names | ZNF291 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 291. | |||||
|
ZN403_MOUSE
|
||||||
| θ value | 5.27518 (rank : 60) | NC score | 0.029009 (rank : 53) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5SV77, Q5SV75, Q5SV76, Q5SV78, Q6GVH6, Q6P9J4, Q920N4 | Gene names | Znf403, Ggnbp2, Zfp403 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein ZNF403 (Gametogenetin-binding protein 2) (Dioxin-inducible factor 3) (DIF-3). | |||||
|
GAK10_HUMAN
|
||||||
| θ value | 6.88961 (rank : 61) | NC score | 0.038123 (rank : 32) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P87889, P10263, P10264, Q69385, Q9UKH6 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_5q33.3 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K10 Gag protein) (HERV-K107 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK11_HUMAN
|
||||||
| θ value | 6.88961 (rank : 62) | NC score | 0.037613 (rank : 35) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P63145, Q9UKI1 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_22q11.21 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K101 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK12_HUMAN
|
||||||
| θ value | 6.88961 (rank : 63) | NC score | 0.041715 (rank : 27) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P63130, Q9UKI0 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_1q22 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K102 Gag protein) (HERV-K(III) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 64) | NC score | 0.038306 (rank : 31) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 268 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P62683 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_12q14.1 provirus ancestral Gag polyprotein (Gag polyprotein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 65) | NC score | 0.044817 (rank : 22) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 182 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9YNA8 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_19q12 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K(C19) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 66) | NC score | 0.037648 (rank : 33) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 415 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P63126, Q9UKH4 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_6q14.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K109 Gag protein) (HERV-K(C6) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK5_HUMAN
|
||||||
| θ value | 6.88961 (rank : 67) | NC score | 0.037504 (rank : 36) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 429 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P62684 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_19p13.11 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K113 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK6_HUMAN
|
||||||
| θ value | 6.88961 (rank : 68) | NC score | 0.037642 (rank : 34) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P62685 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_8p23.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K115 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
LEO1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 69) | NC score | 0.035770 (rank : 41) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
PAPOA_HUMAN
|
||||||
| θ value | 6.88961 (rank : 70) | NC score | 0.037266 (rank : 37) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P51003, Q86SX4, Q86TV0, Q8IYF5, Q9BVU2 | Gene names | PAPOLA, PAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly(A) polymerase alpha (EC 2.7.7.19) (PAP) (Polynucleotide adenylyltransferase alpha). | |||||
|
ZCHC5_HUMAN
|
||||||
| θ value | 6.88961 (rank : 71) | NC score | 0.029771 (rank : 51) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8N8U3 | Gene names | ZCCHC5 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger CCHC domain-containing protein 5. | |||||
|
ZN403_HUMAN
|
||||||
| θ value | 6.88961 (rank : 72) | NC score | 0.021686 (rank : 59) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H3C7, Q96T90, Q9GZR8, Q9H767 | Gene names | ZNF403, LCRG1, LZK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein ZNF403 (Laryngeal carcinoma-related protein 1). | |||||
|
ZN694_HUMAN
|
||||||
| θ value | 6.88961 (rank : 73) | NC score | -0.001089 (rank : 90) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q63HK3, Q6ZN77 | Gene names | ZNF694 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 694. | |||||
|
BANK1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 74) | NC score | 0.012222 (rank : 74) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80VH0, Q3U178, Q8BRV6, Q8BRY8 | Gene names | Bank1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell scaffold protein with ankyrin repeats (Protein AVIEF). | |||||
|
BRD8_MOUSE
|
||||||
| θ value | 8.99809 (rank : 75) | NC score | 0.034855 (rank : 43) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 422 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8R3B7, Q8C049, Q8R583, Q8VDP0 | Gene names | Brd8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 8. | |||||
|
COIA1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 76) | NC score | 0.003691 (rank : 83) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 481 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P39061, Q60672, Q61437, Q62001, Q62002, Q9JK63 | Gene names | Col18a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XVIII) chain precursor [Contains: Endostatin]. | |||||
|
DOCK2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 77) | NC score | 0.005700 (rank : 81) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8C3J5, Q99M79 | Gene names | Dock2 | |||
|
Domain Architecture |

|
|||||
| Description | Dedicator of cytokinesis protein 2 (Hch protein). | |||||
|
EPN4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 78) | NC score | 0.013889 (rank : 67) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q14677, Q8NAF1, Q96E05 | Gene names | CLINT1, ENTH, EPN4, EPNR, KIAA0171 | |||
|
Domain Architecture |
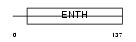
|
|||||
| Description | Clathrin interactor 1 (Epsin-4) (Epsin-related protein) (EpsinR) (Enthoprotin) (Clathrin-interacting protein localized in the trans- Golgi region) (Clint). | |||||
|
EPN4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 79) | NC score | 0.013632 (rank : 70) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q99KN9, Q8CFH4 | Gene names | Clint1, Enth, Epn4, Epnr, Kiaa0171 | |||
|
Domain Architecture |

|
|||||
| Description | Clathrin interactor 1 (Epsin-4) (Epsin-related protein) (EpsinR) (Enthoprotin). | |||||
|
NMDE1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 80) | NC score | 0.007552 (rank : 78) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P35436 | Gene names | Grin2a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate [NMDA] receptor subunit epsilon 1 precursor (N-methyl D- aspartate receptor subtype 2A) (NR2A) (NMDAR2A). | |||||
|
PAPOA_MOUSE
|
||||||
| θ value | 8.99809 (rank : 81) | NC score | 0.036187 (rank : 38) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q61183, Q61208, Q61209, Q8K4X2 | Gene names | Papola, Pap, Plap | |||
|
Domain Architecture |

|
|||||
| Description | Poly(A) polymerase alpha (EC 2.7.7.19) (PAP) (Polynucleotide adenylyltransferase). | |||||
|
PKP4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 82) | NC score | 0.005370 (rank : 82) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q68FH0, Q640N0, Q68G56, Q8BK47, Q8BVH1, Q9CRE3 | Gene names | Pkp4, Armrp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plakophilin-4 (Armadillo-related protein). | |||||
|
PLK4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 83) | NC score | -0.000759 (rank : 89) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 879 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q64702, Q6PEP6, Q78EG6, Q8R0I5, Q9CVR6, Q9CVU6 | Gene names | Plk4, Sak, Stk18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase PLK4 (EC 2.7.11.21) (Polo-like kinase 4) (PLK-4) (Serine/threonine-protein kinase Sak) (Serine/threonine- protein kinase 18). | |||||
|
PTPR2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 84) | NC score | 0.003355 (rank : 84) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q92932, Q8N4I5, Q92662 | Gene names | PTPRN2 | |||
|
Domain Architecture |

|
|||||
| Description | Receptor-type tyrosine-protein phosphatase N2 precursor (EC 3.1.3.48) (R-PTP-N2) (Islet cell autoantigen-related protein) (ICAAR) (IAR) (Phogrin). | |||||
|
SNX1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 85) | NC score | 0.006859 (rank : 80) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q13596, O60750, O60751 | Gene names | SNX1 | |||
|
Domain Architecture |
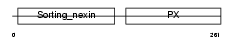
|
|||||
| Description | Sorting nexin-1. | |||||
|
XRCC1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 86) | NC score | 0.013693 (rank : 69) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P18887, Q9HCB1 | Gene names | XRCC1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-repair protein XRCC1 (X-ray repair cross-complementing protein 1). | |||||
|
ZMAT2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 87) | NC score | 0.019322 (rank : 63) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q96NC0 | Gene names | ZMAT2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger matrin type 2. | |||||
|
ZMAT2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 88) | NC score | 0.019322 (rank : 64) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9CPW7, Q5EBK0 | Gene names | Zmat2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger matrin type 2. | |||||
|
ZN318_MOUSE
|
||||||
| θ value | 8.99809 (rank : 89) | NC score | 0.012631 (rank : 71) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 642 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q99PP2, Q3TZL5, Q9JJ01 | Gene names | Znf318, Tzf, Zfp318 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 318 (Testicular zinc finger protein). | |||||
|
ZN516_MOUSE
|
||||||
| θ value | 8.99809 (rank : 90) | NC score | -0.000139 (rank : 87) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1197 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q7TSH3 | Gene names | Znf516, Kiaa0222, Zfp516 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 516. | |||||
|
ZCHC6_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 90 | |
| SwissProt Accessions | Q5BLK4, Q8CIH3 | Gene names | Zcchc6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 6. | |||||
|
ZCHC6_HUMAN
|
||||||
| NC score | 0.979667 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q5VYS8, Q5H9T0, Q5VYS5, Q5VYS7, Q658Z9, Q659A2, Q6MZJ3, Q8N5F0, Q96N57, Q96NE8, Q9C0F2, Q9H8M6 | Gene names | ZCCHC6, KIAA1711 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 6. | |||||
|
ZCH11_HUMAN
|
||||||
| NC score | 0.870640 (rank : 3) | θ value | 9.45368e-175 (rank : 3) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 278 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q5TAX3, Q12764, Q5TAX4, Q86XZ3 | Gene names | ZCCHC11, KIAA0191 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 11. | |||||
|
PAPD1_MOUSE
|
||||||
| NC score | 0.564380 (rank : 4) | θ value | 4.29542e-18 (rank : 5) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9D0D3, Q3UXJ1, Q8C651 | Gene names | Papd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly(A) RNA polymerase, mitochondrial precursor (EC 2.7.7.19) (PAP) (Polynucleotide adenylyltransferase) (PAP-associated domain-containing protein 1). | |||||
|
PAPD1_HUMAN
|
||||||
| NC score | 0.552802 (rank : 5) | θ value | 4.29542e-18 (rank : 4) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9NVV4, Q659E3, Q6P7E5, Q9HA74 | Gene names | PAPD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly(A) RNA polymerase, mitochondrial precursor (EC 2.7.7.19) (PAP) (mtPAP) (Polynucleotide adenylyltransferase) (PAP-associated domain- containing protein 1). | |||||
|
PAPD5_HUMAN
|
||||||
| NC score | 0.399735 (rank : 6) | θ value | 2.52405e-10 (rank : 7) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8NDF8, Q9NW67, Q9Y6C0 | Gene names | PAPD5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PAP-associated domain-containing protein 5 (EC 2.7.7.-) (Topoisomerase-related function protein 4-2) (TRF4-2). | |||||
|
PAD5_MOUSE
|
||||||
| NC score | 0.396870 (rank : 7) | θ value | 2.52405e-10 (rank : 6) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q68ED3, Q8C0K6 | Gene names | Papd5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PAP-associated domain-containing protein 5 (EC 2.7.7.-) (Topoisomerase-related function protein 4-2) (TRF4-2). | |||||
|
POLS_HUMAN
|
||||||
| NC score | 0.386165 (rank : 8) | θ value | 6.21693e-09 (rank : 8) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q5XG87, O43289, Q9Y6C1 | Gene names | POLS, TRF4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA polymerase sigma (EC 2.7.7.7) (Topoisomerase-related function protein 4-1) (TRF4-1) (LAK-1) (DNA polymerase kappa). | |||||
|
POLS_MOUSE
|
||||||
| NC score | 0.379497 (rank : 9) | θ value | 6.21693e-09 (rank : 9) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q6PB75 | Gene names | Pols | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA polymerase sigma (EC 2.7.7.7). | |||||
|
CNBP_HUMAN
|
||||||
| NC score | 0.151778 (rank : 10) | θ value | 0.62314 (rank : 20) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P62633, P20694, Q5QJR0, Q6PJI7, Q96NV3 | Gene names | CNBP, ZNF9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cellular nucleic acid-binding protein (CNBP) (Zinc finger protein 9). | |||||
|
CNBP_MOUSE
|
||||||
| NC score | 0.142584 (rank : 11) | θ value | 0.62314 (rank : 21) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 215 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P53996, Q80Y06, Q8BP23 | Gene names | Cnbp, Cnbp1, Znf9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cellular nucleic acid-binding protein (CNBP) (Zinc finger protein 9). | |||||
|
ZCH13_HUMAN
|
||||||
| NC score | 0.138119 (rank : 12) | θ value | 0.279714 (rank : 18) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8WW36 | Gene names | ZCCHC13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 13. | |||||
|
ZCHC7_HUMAN
|
||||||
| NC score | 0.123660 (rank : 13) | θ value | 0.125558 (rank : 13) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8N3Z6, Q5T0Q8, Q5T0Q9, Q5T0R0, Q8N2M1, Q8N4J2, Q8TBK8, Q9H648, Q9P0F0 | Gene names | ZCCHC7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 7. | |||||
|
PAPOG_HUMAN
|
||||||
| NC score | 0.078121 (rank : 14) | θ value | 0.163984 (rank : 14) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9BWT3, Q969N1, Q9H8L2, Q9HAD0 | Gene names | PAPOLG, PAP2, PAPG | |||
|
Domain Architecture |

|
|||||
| Description | Poly(A) polymerase gamma (EC 2.7.7.19) (PAP gamma) (Polynucleotide adenylyltransferase gamma) (SRP RNA 3' adenylating enzyme) (Neo- poly(A) polymerase) (Neo-PAP). | |||||
|
PAPOG_MOUSE
|
||||||
| NC score | 0.077374 (rank : 15) | θ value | 0.163984 (rank : 15) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6PCL9, Q8BZC9 | Gene names | Papolg | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly(A) polymerase gamma (EC 2.7.7.19) (PAP gamma) (Polynucleotide adenylyltransferase gamma) (SRP RNA 3' adenylating enzyme). | |||||
|
ZCHC9_HUMAN
|
||||||
| NC score | 0.074066 (rank : 16) | θ value | 5.27518 (rank : 57) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8N567, Q9H027 | Gene names | ZCCHC9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 9. | |||||
|
ZCHC9_MOUSE
|
||||||
| NC score | 0.069376 (rank : 17) | θ value | 5.27518 (rank : 58) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8R1J3, Q921T6 | Gene names | Zcchc9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 9. | |||||
|
G6PE_HUMAN
|
||||||
| NC score | 0.063572 (rank : 18) | θ value | 0.0961366 (rank : 11) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O95479, Q4TT33, Q66I35, Q68DT3 | Gene names | H6PD, GDH | |||
|
Domain Architecture |

|
|||||
| Description | GDH/6PGL endoplasmic bifunctional protein precursor [Includes: Glucose 1-dehydrogenase (EC 1.1.1.47) (Hexose-6-phosphate dehydrogenase); 6- phosphogluconolactonase (EC 3.1.1.31) (6PGL)]. | |||||
|
GSUB_HUMAN
|
||||||
| NC score | 0.055913 (rank : 19) | θ value | 4.03905 (rank : 45) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 3 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O96001, Q9UDQ0 | Gene names | GSBS, C7orf16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G-substrate. | |||||
|
TRAT1_HUMAN
|
||||||
| NC score | 0.051249 (rank : 20) | θ value | 1.81305 (rank : 31) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6PIZ9, Q9NZX5 | Gene names | TRAT1, TCRIM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | T-cell receptor-associated transmembrane adapter 1 (T-cell receptor- interacting molecule) (TRIM) (pp29/30). | |||||
|
GAK2_HUMAN
|
||||||
| NC score | 0.045261 (rank : 21) | θ value | 2.36792 (rank : 34) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q7LDI9, Q9UKH5, Q9Y6I1, Q9YNA6, Q9YNB0 | Gene names | ERVK6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_7p22.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K(HML-2.HOM) Gag protein) (HERV-K108 Gag protein) (HERV-K(C7) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK3_HUMAN
|
||||||
| NC score | 0.044817 (rank : 22) | θ value | 6.88961 (rank : 65) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 182 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9YNA8 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_19q12 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K(C19) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK8_HUMAN
|
||||||
| NC score | 0.043599 (rank : 23) | θ value | 4.03905 (rank : 44) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9HDB9 | Gene names | ERVK5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_3q12.3 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K(II) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
CCD96_MOUSE
|
||||||
| NC score | 0.043394 (rank : 24) | θ value | 0.0563607 (rank : 10) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9CR92 | Gene names | Ccdc96 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 96. | |||||
|
LEO1_HUMAN
|
||||||
| NC score | 0.042477 (rank : 25) | θ value | 0.47712 (rank : 19) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
TAF1_HUMAN
|
||||||
| NC score | 0.042466 (rank : 26) | θ value | 3.0926 (rank : 41) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P21675, Q6IUZ1 | Gene names | TAF1, BA2R, CCG1, TAF2A | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 1 (EC 2.7.11.1) (Transcription initiation factor TFIID 250 kDa subunit) (TAF(II)250) (TAFII-250) (TAFII250) (TBP-associated factor 250 kDa) (p250) (Cell cycle gene 1 protein). | |||||
|
GAK12_HUMAN
|
||||||
| NC score | 0.041715 (rank : 27) | θ value | 6.88961 (rank : 63) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P63130, Q9UKI0 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_1q22 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K102 Gag protein) (HERV-K(III) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
ZCHC8_HUMAN
|
||||||
| NC score | 0.041311 (rank : 28) | θ value | 4.03905 (rank : 49) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q6NZY4, Q7L2P6, Q8N2K5, Q96SK7, Q9NSS2, Q9NSS3 | Gene names | ZCCHC8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 8. | |||||
|
GAK18_HUMAN
|
||||||
| NC score | 0.040383 (rank : 29) | θ value | 4.03905 (rank : 43) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P62690 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_22q11.23 provirus ancestral Gag polyprotein (Gag polyprotein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
TAF1L_HUMAN
|
||||||
| NC score | 0.039305 (rank : 30) | θ value | 3.0926 (rank : 40) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8IZX4 | Gene names | TAF1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID 210 kDa subunit (TBP-associated factor 210 kDa) (TAF(II)210) (TBP-associated factor 1-like). | |||||
|
GAK1_HUMAN
|
||||||
| NC score | 0.038306 (rank : 31) | θ value | 6.88961 (rank : 64) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 268 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P62683 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_12q14.1 provirus ancestral Gag polyprotein (Gag polyprotein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK10_HUMAN
|
||||||
| NC score | 0.038123 (rank : 32) | θ value | 6.88961 (rank : 61) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P87889, P10263, P10264, Q69385, Q9UKH6 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_5q33.3 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K10 Gag protein) (HERV-K107 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK4_HUMAN
|
||||||
| NC score | 0.037648 (rank : 33) | θ value | 6.88961 (rank : 66) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 415 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P63126, Q9UKH4 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_6q14.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K109 Gag protein) (HERV-K(C6) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK6_HUMAN
|
||||||
| NC score | 0.037642 (rank : 34) | θ value | 6.88961 (rank : 68) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P62685 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_8p23.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K115 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK11_HUMAN
|
||||||
| NC score | 0.037613 (rank : 35) | θ value | 6.88961 (rank : 62) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P63145, Q9UKI1 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_22q11.21 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K101 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK5_HUMAN
|
||||||
| NC score | 0.037504 (rank : 36) | θ value | 6.88961 (rank : 67) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 429 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P62684 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_19p13.11 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K113 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
PAPOA_HUMAN
|
||||||
| NC score | 0.037266 (rank : 37) | θ value | 6.88961 (rank : 70) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P51003, Q86SX4, Q86TV0, Q8IYF5, Q9BVU2 | Gene names | PAPOLA, PAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly(A) polymerase alpha (EC 2.7.7.19) (PAP) (Polynucleotide adenylyltransferase alpha). | |||||
|
PAPOA_MOUSE
|
||||||
| NC score | 0.036187 (rank : 38) | θ value | 8.99809 (rank : 81) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q61183, Q61208, Q61209, Q8K4X2 | Gene names | Papola, Pap, Plap | |||
|
Domain Architecture |

|
|||||
| Description | Poly(A) polymerase alpha (EC 2.7.7.19) (PAP) (Polynucleotide adenylyltransferase). | |||||
|
RBM4_HUMAN
|
||||||
| NC score | 0.036016 (rank : 39) | θ value | 1.81305 (rank : 29) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9BWF3, O02916, Q6P1P2, Q8WU85 | Gene names | RBM4, RBM4A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein 4 (RNA-binding motif protein 4) (RNA-binding motif protein 4a) (Lark homolog) (hLark). | |||||
|
RBM4_MOUSE
|
||||||
| NC score | 0.036005 (rank : 40) | θ value | 1.81305 (rank : 30) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8C7Q4, O08752, Q8BN66 | Gene names | Rbm4, Rbm4a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein 4 (RNA-binding motif protein 4) (RNA-binding motif protein 4a) (Lark homolog) (mLark). | |||||
|
LEO1_MOUSE
|
||||||
| NC score | 0.035770 (rank : 41) | θ value | 6.88961 (rank : 69) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
GPDM_MOUSE
|
||||||
| NC score | 0.035730 (rank : 42) | θ value | 3.0926 (rank : 38) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q64521, Q61507 | Gene names | Gpd2, Gdm1 | |||
|
Domain Architecture |

|
|||||
| Description | Glycerol-3-phosphate dehydrogenase, mitochondrial precursor (EC 1.1.99.5) (GPD-M) (GPDH-M). | |||||
|
BRD8_MOUSE
|
||||||
| NC score | 0.034855 (rank : 43) | θ value | 8.99809 (rank : 75) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 422 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8R3B7, Q8C049, Q8R583, Q8VDP0 | Gene names | Brd8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 8. | |||||
|
K0555_HUMAN
|
||||||
| NC score | 0.033903 (rank : 44) | θ value | 0.279714 (rank : 17) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 880 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q96AA8, O60302 | Gene names | KIAA0555 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0555. | |||||
|
MDN1_HUMAN
|
||||||
| NC score | 0.032245 (rank : 45) | θ value | 2.36792 (rank : 35) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 662 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9NU22, O15019 | Gene names | MDN1, KIAA0301 | |||
|
Domain Architecture |

|
|||||
| Description | Midasin (MIDAS-containing protein). | |||||
|
RBM6_HUMAN
|
||||||
| NC score | 0.032120 (rank : 46) | θ value | 1.06291 (rank : 23) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P78332, O60549, O75524 | Gene names | RBM6, DEF3 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 6 (RNA-binding motif protein 6) (RNA-binding protein DEF-3) (Lung cancer antigen NY-LU-12) (Protein G16). | |||||
|
RBM4B_HUMAN
|
||||||
| NC score | 0.031505 (rank : 47) | θ value | 4.03905 (rank : 47) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9BQ04 | Gene names | RBM4B, RBM30 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein 4B (RNA-binding motif protein 4B) (RNA-binding protein 30) (RNA-binding motif protein 30). | |||||
|
RBM4B_MOUSE
|
||||||
| NC score | 0.031481 (rank : 48) | θ value | 4.03905 (rank : 48) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8VE92 | Gene names | Rbm4b, Rbm30 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein 4B (RNA-binding motif protein 4B) (RNA-binding protein 30) (RNA-binding motif protein 30). | |||||
|
GPDM_HUMAN
|
||||||
| NC score | 0.030412 (rank : 49) | θ value | 5.27518 (rank : 52) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P43304, Q59FR1, Q9HAP9 | Gene names | GPD2 | |||
|
Domain Architecture |

|
|||||
| Description | Glycerol-3-phosphate dehydrogenase, mitochondrial precursor (EC 1.1.99.5) (GPD-M) (GPDH-M) (mtGPD). | |||||
|
CCD96_HUMAN
|
||||||
| NC score | 0.029843 (rank : 50) | θ value | 2.36792 (rank : 32) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 595 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q2M329, Q8N2I7 | Gene names | CCDC96 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 96. | |||||
|
ZCHC5_HUMAN
|
||||||
| NC score | 0.029771 (rank : 51) | θ value | 6.88961 (rank : 71) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8N8U3 | Gene names | ZCCHC5 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger CCHC domain-containing protein 5. | |||||
|
SODC_MOUSE
|
||||||
| NC score | 0.029669 (rank : 52) | θ value | 1.38821 (rank : 26) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P08228 | Gene names | Sod1 | |||
|
Domain Architecture |

|
|||||
| Description | Superoxide dismutase [Cu-Zn] (EC 1.15.1.1). | |||||
|
ZN403_MOUSE
|
||||||
| NC score | 0.029009 (rank : 53) | θ value | 5.27518 (rank : 60) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5SV77, Q5SV75, Q5SV76, Q5SV78, Q6GVH6, Q6P9J4, Q920N4 | Gene names | Znf403, Ggnbp2, Zfp403 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein ZNF403 (Gametogenetin-binding protein 2) (Dioxin-inducible factor 3) (DIF-3). | |||||
|
ZN291_HUMAN
|
||||||
| NC score | 0.028775 (rank : 54) | θ value | 5.27518 (rank : 59) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 647 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9BY12 | Gene names | ZNF291 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 291. | |||||
|
BAZ1B_MOUSE
|
||||||
| NC score | 0.028206 (rank : 55) | θ value | 1.38821 (rank : 24) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 394 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9Z277, Q9CU68 | Gene names | Baz1b, Wbscr9 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1B (Williams-Beuren syndrome chromosome region 9 protein homolog) (WBRS9). | |||||
|
BAZ1B_HUMAN
|
||||||
| NC score | 0.026083 (rank : 56) | θ value | 1.81305 (rank : 27) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 701 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9UIG0, O95039, O95247, O95277 | Gene names | BAZ1B, WBSC10, WBSCR9, WSTF | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain protein 1B (Williams-Beuren syndrome chromosome region 9 protein) (Williams syndrome transcription factor) (hWALP2). | |||||
|
NUFP2_HUMAN
|
||||||
| NC score | 0.024583 (rank : 57) | θ value | 2.36792 (rank : 36) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7Z417, Q9P2M5 | Gene names | NUFIP2, KIAA1321, PIG1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear fragile X mental retardation-interacting protein 2 (FMRP- interacting protein 2) (82 kDa FMRP-interacting protein) (82-FIP) (Proliferation-inducing gene 1 protein). | |||||
|
CENB5_HUMAN
|
||||||
| NC score | 0.022985 (rank : 58) | θ value | 0.279714 (rank : 16) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 395 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96P50, Q9BSR9, Q9C0E7 | Gene names | CENTB5, KIAA1716 | |||
|
Domain Architecture |

|
|||||
| Description | Centaurin-beta 5 (Cnt-b5). | |||||
|
ZN403_HUMAN
|
||||||
| NC score | 0.021686 (rank : 59) | θ value | 6.88961 (rank : 72) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H3C7, Q96T90, Q9GZR8, Q9H767 | Gene names | ZNF403, LCRG1, LZK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein ZNF403 (Laryngeal carcinoma-related protein 1). | |||||
|
BSN_MOUSE
|
||||||
| NC score | 0.021087 (rank : 60) | θ value | 3.0926 (rank : 37) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
TEX14_HUMAN
|
||||||
| NC score | 0.020439 (rank : 61) | θ value | 0.0961366 (rank : 12) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1103 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8IWB6, Q7RTP3, Q8ND97, Q9BXT9 | Gene names | TEX14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Testis-expressed protein 14 (Testis-expressed sequence 14). | |||||
|
PLCB3_HUMAN
|
||||||
| NC score | 0.019666 (rank : 62) | θ value | 0.813845 (rank : 22) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q01970 | Gene names | PLCB3 | |||
|
Domain Architecture |

|
|||||
| Description | 1-phosphatidylinositol-4,5-bisphosphate phosphodiesterase beta 3 (EC 3.1.4.11) (Phosphoinositide phospholipase C) (Phospholipase C- beta-3) (PLC-beta-3). | |||||
|
ZMAT2_HUMAN
|
||||||
| NC score | 0.019322 (rank : 63) | θ value | 8.99809 (rank : 87) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q96NC0 | Gene names | ZMAT2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger matrin type 2. | |||||
|
ZMAT2_MOUSE
|
||||||
| NC score | 0.019322 (rank : 64) | θ value | 8.99809 (rank : 88) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9CPW7, Q5EBK0 | Gene names | Zmat2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger matrin type 2. | |||||
|
BRPF1_HUMAN
|
||||||
| NC score | 0.018089 (rank : 65) | θ value | 1.38821 (rank : 25) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P55201, Q9UHI0 | Gene names | BRPF1, BR140 | |||
|
Domain Architecture |

|
|||||
| Description | Peregrin (Bromodomain and PHD finger-containing protein 1) (BR140 protein). | |||||
|
CF152_MOUSE
|
||||||
| NC score | 0.015156 (rank : 66) | θ value | 4.03905 (rank : 42) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 597 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q80ST9, Q9CYM9, Q9D5J9 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf152 homolog. | |||||
|
EPN4_HUMAN
|
||||||
| NC score | 0.013889 (rank : 67) | θ value | 8.99809 (rank : 78) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q14677, Q8NAF1, Q96E05 | Gene names | CLINT1, ENTH, EPN4, EPNR, KIAA0171 | |||
|
Domain Architecture |

|
|||||
| Description | Clathrin interactor 1 (Epsin-4) (Epsin-related protein) (EpsinR) (Enthoprotin) (Clathrin-interacting protein localized in the trans- Golgi region) (Clint). | |||||
|
IWS1_HUMAN
|
||||||
| NC score | 0.013865 (rank : 68) | θ value | 5.27518 (rank : 53) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
XRCC1_HUMAN
|
||||||
| NC score | 0.013693 (rank : 69) | θ value | 8.99809 (rank : 86) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P18887, Q9HCB1 | Gene names | XRCC1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-repair protein XRCC1 (X-ray repair cross-complementing protein 1). | |||||
|
EPN4_MOUSE
|
||||||
| NC score | 0.013632 (rank : 70) | θ value | 8.99809 (rank : 79) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q99KN9, Q8CFH4 | Gene names | Clint1, Enth, Epn4, Epnr, Kiaa0171 | |||
|
Domain Architecture |

|
|||||
| Description | Clathrin interactor 1 (Epsin-4) (Epsin-related protein) (EpsinR) (Enthoprotin). | |||||
|
ZN318_MOUSE
|
||||||
| NC score | 0.012631 (rank : 71) | θ value | 8.99809 (rank : 89) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 642 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q99PP2, Q3TZL5, Q9JJ01 | Gene names | Znf318, Tzf, Zfp318 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 318 (Testicular zinc finger protein). | |||||
|
HIP1R_MOUSE
|
||||||
| NC score | 0.012491 (rank : 72) | θ value | 1.81305 (rank : 28) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 959 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9JKY5 | Gene names | Hip1r | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin-interacting protein 1-related protein (Hip1-related). | |||||
|
NCOA3_HUMAN
|
||||||
| NC score | 0.012367 (rank : 73) | θ value | 3.0926 (rank : 39) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y6Q9, Q5JYD9, Q5JYE0, Q9BR49, Q9UPC9, Q9UPG4, Q9UPG7 | Gene names | NCOA3, AIB1, RAC3, TRAM1 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 3 (EC 2.3.1.48) (NCoA-3) (Thyroid hormone receptor activator molecule 1) (TRAM-1) (ACTR) (Receptor-associated coactivator 3) (RAC-3) (Amplified in breast cancer-1 protein) (AIB-1) (Steroid receptor coactivator protein 3) (SRC-3) (CBP-interacting protein) (pCIP). | |||||
|
BANK1_MOUSE
|
||||||
| NC score | 0.012222 (rank : 74) | θ value | 8.99809 (rank : 74) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80VH0, Q3U178, Q8BRV6, Q8BRY8 | Gene names | Bank1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell scaffold protein with ankyrin repeats (Protein AVIEF). | |||||
|
FGD6_MOUSE
|
||||||
| NC score | 0.010046 (rank : 75) | θ value | 2.36792 (rank : 33) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q69ZL1, Q8C8W5, Q8K3B0, Q9D3Y7 | Gene names | Fgd6, Kiaa1362 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 6. | |||||
|
RBCC1_MOUSE
|
||||||
| NC score | 0.008694 (rank : 76) | θ value | 4.03905 (rank : 46) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1048 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9ESK9, Q3TXX2, Q61384, Q8BRY9, Q9JK14 | Gene names | Rb1cc1, Cc1, Kiaa0203 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RB1-inducible coiled-coil protein 1 (Coiled-coil-forming protein 1) (LaXp180 protein). | |||||
|
NMDE1_HUMAN
|
||||||
| NC score | 0.008652 (rank : 77) | θ value | 5.27518 (rank : 56) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q12879, O00669 | Gene names | GRIN2A | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate [NMDA] receptor subunit epsilon 1 precursor (N-methyl D- aspartate receptor subtype 2A) (NR2A) (NMDAR2A) (hNR2A). | |||||
|
NMDE1_MOUSE
|
||||||
| NC score | 0.007552 (rank : 78) | θ value | 8.99809 (rank : 80) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P35436 | Gene names | Grin2a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate [NMDA] receptor subunit epsilon 1 precursor (N-methyl D- aspartate receptor subtype 2A) (NR2A) (NMDAR2A). | |||||
|
NIN_HUMAN
|
||||||
| NC score | 0.007398 (rank : 79) | θ value | 5.27518 (rank : 55) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1416 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8N4C6, Q6P0P6, Q9BWU6, Q9C012, Q9C013, Q9C014, Q9H5I6, Q9HAT7, Q9HBY5, Q9HCK7, Q9UH61 | Gene names | NIN, KIAA1565 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ninein (hNinein) (Glycogen synthase kinase 3 beta-interacting protein) (GSK3B-interacting protein). | |||||
|
SNX1_HUMAN
|
||||||
| NC score | 0.006859 (rank : 80) | θ value | 8.99809 (rank : 85) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q13596, O60750, O60751 | Gene names | SNX1 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-1. | |||||
|
DOCK2_MOUSE
|
||||||
| NC score | 0.005700 (rank : 81) | θ value | 8.99809 (rank : 77) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8C3J5, Q99M79 | Gene names | Dock2 | |||
|
Domain Architecture |

|
|||||
| Description | Dedicator of cytokinesis protein 2 (Hch protein). | |||||
|
PKP4_MOUSE
|
||||||
| NC score | 0.005370 (rank : 82) | θ value | 8.99809 (rank : 82) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q68FH0, Q640N0, Q68G56, Q8BK47, Q8BVH1, Q9CRE3 | Gene names | Pkp4, Armrp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plakophilin-4 (Armadillo-related protein). | |||||
|
COIA1_MOUSE
|
||||||
| NC score | 0.003691 (rank : 83) | θ value | 8.99809 (rank : 76) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 481 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P39061, Q60672, Q61437, Q62001, Q62002, Q9JK63 | Gene names | Col18a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XVIII) chain precursor [Contains: Endostatin]. | |||||
|
PTPR2_HUMAN
|
||||||
| NC score | 0.003355 (rank : 84) | θ value | 8.99809 (rank : 84) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q92932, Q8N4I5, Q92662 | Gene names | PTPRN2 | |||
|
Domain Architecture |

|
|||||
| Description | Receptor-type tyrosine-protein phosphatase N2 precursor (EC 3.1.3.48) (R-PTP-N2) (Islet cell autoantigen-related protein) (ICAAR) (IAR) (Phogrin). | |||||
|
BC11B_HUMAN
|
||||||
| NC score | 0.003096 (rank : 85) | θ value | 5.27518 (rank : 50) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 761 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9C0K0, Q9H162 | Gene names | BCL11B, CTIP2, RIT1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma/leukemia 11B (B-cell CLL/lymphoma 11B) (Radiation- induced tumor suppressor gene 1 protein) (hRit1) (COUP-TF-interacting protein 2). | |||||
|
CD2L1_MOUSE
|
||||||
| NC score | 0.001433 (rank : 86) | θ value | 5.27518 (rank : 51) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1463 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P24788, Q3UI03, Q61399, Q7TST4, Q8BP53 | Gene names | Cdc2l1 | |||
|
Domain Architecture |

|
|||||
| Description | PITSLRE serine/threonine-protein kinase CDC2L1 (EC 2.7.11.22) (Galactosyltransferase-associated protein kinase p58/GTA) (Cell division cycle 2-like protein kinase 1). | |||||
|
ZN516_MOUSE
|
||||||
| NC score | -0.000139 (rank : 87) | θ value | 8.99809 (rank : 90) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1197 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q7TSH3 | Gene names | Znf516, Kiaa0222, Zfp516 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 516. | |||||
|
MYLK_HUMAN
|
||||||
| NC score | -0.000361 (rank : 88) | θ value | 5.27518 (rank : 54) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 1328 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q15746, O95796, O95797, O95798, O95799, Q14844, Q16794, Q5MY99, Q5MYA0, Q7Z4J0, Q9C0L5, Q9UBG5, Q9UIT9 | Gene names | MYLK, MLCK | |||
|
Domain Architecture |

|
|||||
| Description | Myosin light chain kinase, smooth muscle (EC 2.7.11.18) (MLCK) (Telokin) (Kinase-related protein) (KRP). | |||||
|
PLK4_MOUSE
|
||||||
| NC score | -0.000759 (rank : 89) | θ value | 8.99809 (rank : 83) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 879 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q64702, Q6PEP6, Q78EG6, Q8R0I5, Q9CVR6, Q9CVU6 | Gene names | Plk4, Sak, Stk18 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase PLK4 (EC 2.7.11.21) (Polo-like kinase 4) (PLK-4) (Serine/threonine-protein kinase Sak) (Serine/threonine- protein kinase 18). | |||||
|
ZN694_HUMAN
|
||||||
| NC score | -0.001089 (rank : 90) | θ value | 6.88961 (rank : 73) | |||
| Query Neighborhood Hits | 90 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q63HK3, Q6ZN77 | Gene names | ZNF694 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 694. | |||||