Please be patient as the page loads
|
PIGQ_HUMAN
|
||||||
| SwissProt Accessions | Q9BRB3, O14927, Q96G00, Q96S22, Q9UJH4 | Gene names | PIGQ, GPI1 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol N-acetylglucosaminyltransferase subunit Q (EC 2.4.1.198) (Phosphatidylinositol-glycan biosynthesis class Q protein) (PIG-Q) (N-acetylglucosamyl transferase component GPI1). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
PIGQ_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 84 | |
| SwissProt Accessions | Q9BRB3, O14927, Q96G00, Q96S22, Q9UJH4 | Gene names | PIGQ, GPI1 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol N-acetylglucosaminyltransferase subunit Q (EC 2.4.1.198) (Phosphatidylinositol-glycan biosynthesis class Q protein) (PIG-Q) (N-acetylglucosamyl transferase component GPI1). | |||||
|
PIGQ_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.965759 (rank : 2) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 5 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9QYT7, O35120, O35456, Q99L11 | Gene names | Pigq, Gpi1h, Mgpi1 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol N-acetylglucosaminyltransferase subunit Q (EC 2.4.1.198) (Phosphatidylinositol-glycan biosynthesis class Q protein) (PIG-Q) (N-acetylglucosamyl transferase component GPI1) (MGpi1p). | |||||
|
SSBP4_HUMAN
|
||||||
| θ value | 0.000461057 (rank : 3) | NC score | 0.083414 (rank : 3) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 753 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9BWG4 | Gene names | SSBP4 | |||
|
Domain Architecture |

|
|||||
| Description | Single-stranded DNA-binding protein 4. | |||||
|
PROP_MOUSE
|
||||||
| θ value | 0.00665767 (rank : 4) | NC score | 0.057920 (rank : 8) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P11680, Q3TB98, Q3U779 | Gene names | Cfp, Pfc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Properdin precursor (Factor P). | |||||
|
SPR1A_MOUSE
|
||||||
| θ value | 0.00869519 (rank : 5) | NC score | 0.074802 (rank : 5) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q62266 | Gene names | Sprr1a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cornifin A (Small proline-rich protein 1A) (SPR1A) (SPR1 A). | |||||
|
STAB1_HUMAN
|
||||||
| θ value | 0.125558 (rank : 6) | NC score | 0.023077 (rank : 37) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 546 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9NY15, Q8IUH0, Q8IUH1, Q93072 | Gene names | STAB1, FEEL1, KIAA0246 | |||
|
Domain Architecture |

|
|||||
| Description | Stabilin-1 precursor (FEEL-1 protein) (MS-1 antigen). | |||||
|
CN032_HUMAN
|
||||||
| θ value | 0.21417 (rank : 7) | NC score | 0.073416 (rank : 6) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8NDC0, Q96BG5 | Gene names | C14orf32 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf32. | |||||
|
TAF4_HUMAN
|
||||||
| θ value | 0.21417 (rank : 8) | NC score | 0.059598 (rank : 7) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 687 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O00268, Q5TBP6, Q99721, Q9BR40, Q9BX42 | Gene names | TAF4, TAF2C, TAF2C1, TAF4A, TAFII130, TAFII135 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 4 (TBP-associated factor 4) (Transcription initiation factor TFIID 135 kDa subunit) (TAF(II)135) (TAFII-135) (TAFII135) (TAFII-130) (TAFII130). | |||||
|
MDFI_HUMAN
|
||||||
| θ value | 0.365318 (rank : 9) | NC score | 0.074821 (rank : 4) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99750 | Gene names | MDFI | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MyoD family inhibitor (Myogenic repressor I-mf). | |||||
|
DMWD_MOUSE
|
||||||
| θ value | 0.47712 (rank : 10) | NC score | 0.037881 (rank : 21) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q08274 | Gene names | Dmwd, Dm9 | |||
|
Domain Architecture |

|
|||||
| Description | Dystrophia myotonica WD repeat-containing protein (Dystrophia myotonica-containing WD repeat motif protein) (DMR-N9 protein). | |||||
|
PCX3_MOUSE
|
||||||
| θ value | 0.47712 (rank : 11) | NC score | 0.052492 (rank : 9) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8VI59 | Gene names | Pcnxl3 | |||
|
Domain Architecture |

|
|||||
| Description | Pecanex-like protein 3. | |||||
|
WIRE_MOUSE
|
||||||
| θ value | 0.62314 (rank : 12) | NC score | 0.045583 (rank : 15) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q6PEV3 | Gene names | Wire | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WIP-related protein (WASP-interacting protein-related protein). | |||||
|
ZC3H3_MOUSE
|
||||||
| θ value | 0.62314 (rank : 13) | NC score | 0.042949 (rank : 17) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8CHP0, Q80X78 | Gene names | Zc3h3, Kiaa0150, Zc3hdc3, Zfp623 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 3. | |||||
|
TDGF1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 14) | NC score | 0.038518 (rank : 20) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P51865 | Gene names | Tdgf1, Cripto | |||
|
Domain Architecture |
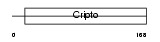
|
|||||
| Description | Teratocarcinoma-derived growth factor precursor (Epidermal growth factor-like Cripto protein) (Cripto growth factor). | |||||
|
DIAP1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 15) | NC score | 0.027718 (rank : 28) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 784 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O60610, Q9UC76 | Gene names | DIAPH1, DIAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein diaphanous homolog 1 (Diaphanous-related formin-1) (DRF1). | |||||
|
PLXB1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 16) | NC score | 0.025972 (rank : 29) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8CJH3, Q6ZQC3, Q80ZZ1 | Gene names | Plxnb1, Kiaa0407 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-B1 precursor. | |||||
|
ADA15_HUMAN
|
||||||
| θ value | 1.38821 (rank : 17) | NC score | 0.012180 (rank : 68) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 288 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13444, Q13493, Q96C78 | Gene names | ADAM15, MDC15 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 15 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 15) (Metalloproteinase-like, disintegrin-like, and cysteine- rich protein 15) (MDC-15) (Metalloprotease RGD disintegrin protein) (Metargidin). | |||||
|
IF35_HUMAN
|
||||||
| θ value | 1.38821 (rank : 18) | NC score | 0.047117 (rank : 12) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O00303, Q8N978 | Gene names | EIF3S5 | |||
|
Domain Architecture |
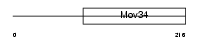
|
|||||
| Description | Eukaryotic translation initiation factor 3 subunit 5 (eIF-3 epsilon) (eIF3 p47 subunit) (eIF3f). | |||||
|
PROP_HUMAN
|
||||||
| θ value | 1.38821 (rank : 19) | NC score | 0.043225 (rank : 16) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P27918, O15134, O15135, O15136, O75826 | Gene names | CFP, PFC | |||
|
Domain Architecture |
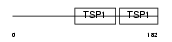
|
|||||
| Description | Properdin precursor (Factor P). | |||||
|
RBM12_MOUSE
|
||||||
| θ value | 1.38821 (rank : 20) | NC score | 0.038639 (rank : 19) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8R4X3, Q8K373, Q8R302, Q9CS80 | Gene names | Rbm12 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 12 (RNA-binding motif protein 12) (SH3/WW domain anchor protein in the nucleus) (SWAN). | |||||
|
PRR12_HUMAN
|
||||||
| θ value | 1.81305 (rank : 21) | NC score | 0.046511 (rank : 14) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 460 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9ULL5, Q8N4J6 | Gene names | PRR12, KIAA1205 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 12. | |||||
|
RXRB_MOUSE
|
||||||
| θ value | 1.81305 (rank : 22) | NC score | 0.009739 (rank : 72) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P28704, P33243 | Gene names | Rxrb, Nr2b2 | |||
|
Domain Architecture |
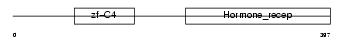
|
|||||
| Description | Retinoic acid receptor RXR-beta (Retinoid X receptor beta) (MHC class I regulatory element-binding protein H-2RIIBP). | |||||
|
ENAH_MOUSE
|
||||||
| θ value | 2.36792 (rank : 23) | NC score | 0.022687 (rank : 39) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 919 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q03173, P70430, P70431, P70432, P70433, Q5D053 | Gene names | Enah, Mena, Ndpp1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein enabled homolog (NPC-derived proline-rich protein 1) (NDPP-1). | |||||
|
IF35_MOUSE
|
||||||
| θ value | 2.36792 (rank : 24) | NC score | 0.042692 (rank : 18) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9DCH4 | Gene names | Eif3s5 | |||
|
Domain Architecture |
|
|||||
| Description | Eukaryotic translation initiation factor 3 subunit 5 (eIF-3 epsilon) (eIF3 p47 subunit) (eIF3f). | |||||
|
NCOR2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 25) | NC score | 0.021268 (rank : 43) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 870 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9Y618, O00613, O15416, Q13354, Q9Y5U0 | Gene names | NCOR2, CTG26 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC) (CTG repeat protein 26) (SMAP270). | |||||
|
PRRT1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 26) | NC score | 0.051231 (rank : 10) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q99946, Q96DW3, Q96NQ8 | Gene names | PRRT1, C6orf31, NG5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich transmembrane protein 1. | |||||
|
PRRT1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 27) | NC score | 0.049686 (rank : 11) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O35449 | Gene names | Prrt1, Ng5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich transmembrane protein 1. | |||||
|
PTN23_MOUSE
|
||||||
| θ value | 2.36792 (rank : 28) | NC score | 0.016827 (rank : 56) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 726 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q6PB44, Q69ZJ0, Q8R1Z5, Q923E6 | Gene names | Ptpn23, Kiaa1471 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 23 (EC 3.1.3.48). | |||||
|
RN167_HUMAN
|
||||||
| θ value | 2.36792 (rank : 29) | NC score | 0.020841 (rank : 44) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H6Y7, Q6XYE0, Q8NDC1, Q9Y3V1 | Gene names | RNF167 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 167 precursor. | |||||
|
ZN414_HUMAN
|
||||||
| θ value | 2.36792 (rank : 30) | NC score | 0.023867 (rank : 35) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 548 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96IQ9 | Gene names | ZNF414 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 414. | |||||
|
ARI1A_HUMAN
|
||||||
| θ value | 3.0926 (rank : 31) | NC score | 0.030347 (rank : 24) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 787 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O14497, Q53FK9, Q5T0W1, Q5T0W2, Q5T0W3, Q8NFD6, Q96T89, Q9BY33, Q9HBJ5, Q9UPZ1 | Gene names | ARID1A, C1orf4, OSA1, SMARCF1 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 1A (ARID domain- containing protein 1A) (SWI/SNF-related, matrix-associated, actin- dependent regulator of chromatin subfamily F member 1) (SWI-SNF complex protein p270) (B120) (SWI-like protein) (Osa homolog 1) (hOSA1) (hELD) (BRG1-associated factor 250) (BAF250) (BRG1-associated factor 250a) (BAF250A). | |||||
|
COE4_MOUSE
|
||||||
| θ value | 3.0926 (rank : 32) | NC score | 0.022644 (rank : 40) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8K4J2, Q8K4J1, Q8K4J3, Q8K4J4, Q8K4J5 | Gene names | Ebf4, Coe4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor COE4 (Early B-cell factor 4) (EBF-4) (Olf-1/EBF- like 4) (OE-4) (O/E-4). | |||||
|
JUN_MOUSE
|
||||||
| θ value | 3.0926 (rank : 33) | NC score | 0.021780 (rank : 42) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P05627 | Gene names | Jun | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor AP-1 (Activator protein 1) (AP1) (Proto-oncogene c-jun) (V-jun avian sarcoma virus 17 oncogene homolog) (Jun A) (AH119). | |||||
|
MBD6_HUMAN
|
||||||
| θ value | 3.0926 (rank : 34) | NC score | 0.036268 (rank : 22) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q96DN6, Q8N3M0, Q8NA81, Q96Q00 | Gene names | MBD6, KIAA1887 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 6 (Methyl-CpG-binding protein MBD6). | |||||
|
SEM5B_HUMAN
|
||||||
| θ value | 3.0926 (rank : 35) | NC score | 0.019897 (rank : 48) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9P283, Q6UY12 | Gene names | SEMA5B, KIAA1445 | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-5B precursor. | |||||
|
ST5_HUMAN
|
||||||
| θ value | 3.0926 (rank : 36) | NC score | 0.015815 (rank : 61) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P78524, P78523, Q16492, Q7KYY2, Q7KZ12, Q8NE12, Q9BQQ6 | Gene names | ST5, DENND2B, HTS1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppression of tumorigenicity 5 (HeLa tumor suppression 1) (DENN domain-containing protein 2B). | |||||
|
COHA1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 37) | NC score | 0.016492 (rank : 57) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 400 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UMD9, Q02802, Q99018, Q9NQK9, Q9UC14 | Gene names | COL17A1, BP180, BPAG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XVII) chain (Bullous pemphigoid antigen 2) (180 kDa bullous pemphigoid antigen 2). | |||||
|
DYR1B_HUMAN
|
||||||
| θ value | 4.03905 (rank : 38) | NC score | 0.004064 (rank : 79) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 831 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Y463, O75258, O75788, O75789 | Gene names | DYRK1B, MIRK | |||
|
Domain Architecture |

|
|||||
| Description | Dual specificity tyrosine-phosphorylation-regulated kinase 1B (EC 2.7.12.1) (Mirk protein kinase) (Minibrain-related kinase). | |||||
|
GP152_HUMAN
|
||||||
| θ value | 4.03905 (rank : 39) | NC score | 0.013955 (rank : 64) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 636 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8TDT2, Q86SM0 | Gene names | GPR152, PGR5 | |||
|
Domain Architecture |

|
|||||
| Description | Probable G-protein coupled receptor 152 (G-protein coupled receptor PGR5). | |||||
|
LBXCO_HUMAN
|
||||||
| θ value | 4.03905 (rank : 40) | NC score | 0.022880 (rank : 38) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P84550 | Gene names | LBXCOR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ladybird homeobox corepressor 1. | |||||
|
PLXB1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 41) | NC score | 0.025810 (rank : 30) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O43157, Q6NY20, Q9UIV7, Q9UJ92, Q9UJ93 | Gene names | PLXNB1, KIAA0407, SEP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-B1 precursor (Semaphorin receptor SEP). | |||||
|
COE4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 42) | NC score | 0.020555 (rank : 46) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9BQW3, Q5JY53, Q9NUB6, Q9P2A6 | Gene names | EBF4, COE4, KIAA1442 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor COE4 (Early B-cell factor 4) (EBF-4) (Olf-1/EBF- like 4) (OE-4) (O/E-4). | |||||
|
F113A_HUMAN
|
||||||
| θ value | 5.27518 (rank : 43) | NC score | 0.024368 (rank : 34) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H1Q7, Q5JUA5, Q5JUA6, Q6PK19, Q86WF5, Q96CG7, Q9H1Q6, Q9H6D1 | Gene names | FAM113A, C20orf81 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM113A (Sarcoma antigen NY-SAR-23). | |||||
|
FOG1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 44) | NC score | 0.006850 (rank : 75) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 981 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8IX07 | Gene names | ZFPM1, FOG1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein ZFPM1 (Zinc finger protein multitype 1) (Friend of GATA protein 1) (Friend of GATA-1) (FOG-1). | |||||
|
FUBP1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 45) | NC score | 0.024437 (rank : 31) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96AE4, Q12828 | Gene names | FUBP1 | |||
|
Domain Architecture |
|
|||||
| Description | Far upstream element-binding protein 1 (FUSE-binding protein 1) (FBP) (DNA helicase V) (HDH V). | |||||
|
FUBP1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 46) | NC score | 0.024420 (rank : 33) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q91WJ8, Q8C0Y8 | Gene names | Fubp1, D3Ertd330e | |||
|
Domain Architecture |
|
|||||
| Description | Far upstream element-binding protein 1 (FUSE-binding protein 1) (FBP). | |||||
|
GRIP1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 47) | NC score | 0.010709 (rank : 70) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q925T6, Q8BLQ3, Q8C0T3, Q925T5, Q925T7 | Gene names | Grip1 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor-interacting protein 1 (GRIP1 protein). | |||||
|
IRAK1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 48) | NC score | 0.004109 (rank : 78) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 820 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P51617, Q7Z5V4, Q96RL2 | Gene names | IRAK1, IRAK | |||
|
Domain Architecture |
|
|||||
| Description | Interleukin-1 receptor-associated kinase 1 (EC 2.7.11.1) (IRAK-1). | |||||
|
IRS2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 49) | NC score | 0.019903 (rank : 47) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9Y4H2, Q96RR2, Q9BZG0, Q9Y6I5 | Gene names | IRS2 | |||
|
Domain Architecture |
|
|||||
| Description | Insulin receptor substrate 2 (IRS-2). | |||||
|
MUC5B_HUMAN
|
||||||
| θ value | 5.27518 (rank : 50) | NC score | 0.013480 (rank : 65) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1020 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9HC84, O00447, O00573, O14985, O15494, O95291, O95451, Q14881, Q99552, Q9UE28 | Gene names | MUC5B, MUC5 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-5B precursor (Mucin-5 subtype B, tracheobronchial) (High molecular weight salivary mucin MG1) (Sublingual gland mucin). | |||||
|
PRGC2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 51) | NC score | 0.019705 (rank : 49) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 272 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q86YN6, Q86YN3, Q86YN4, Q86YN5, Q8N1N9, Q8TDE4, Q8TDE5 | Gene names | PPARGC1B, PERC, PGC1, PGC1B, PPARGC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor gamma coactivator 1-beta (PPAR gamma coactivator-1beta) (PGC-1-related estrogen receptor alpha coactivator) (PPARGC-1-beta) (PGC-1-beta). | |||||
|
RFX1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 52) | NC score | 0.018548 (rank : 52) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 256 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P22670 | Gene names | RFX1 | |||
|
Domain Architecture |

|
|||||
| Description | MHC class II regulatory factor RFX1 (RFX) (Enhancer factor C) (EF-C). | |||||
|
SIAH2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 53) | NC score | 0.015987 (rank : 58) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q06986 | Gene names | Siah2 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin ligase SIAH2 (EC 6.3.2.-) (Seven in absentia homolog 2) (Siah-2) (mSiah2). | |||||
|
SMCA2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 54) | NC score | 0.010533 (rank : 71) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 503 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P51531 | Gene names | SMARCA2, BRM, SNF2A, SNF2L2 | |||
|
Domain Architecture |

|
|||||
| Description | Probable global transcription activator SNF2L2 (EC 3.6.1.-) (ATP- dependent helicase SMARCA2) (SNF2-alpha) (SWI/SNF-related matrix- associated actin-dependent regulator of chromatin subfamily A member 2) (hBRM). | |||||
|
ZXDA_HUMAN
|
||||||
| θ value | 5.27518 (rank : 55) | NC score | 0.001595 (rank : 82) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 750 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P98168, Q9UJP7 | Gene names | ZXDA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger X-linked protein ZXDA. | |||||
|
3BP1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 56) | NC score | 0.012235 (rank : 67) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 376 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9Y3L3, Q6IBZ2, Q6ZVL9, Q96HQ5, Q9NSQ9 | Gene names | SH3BP1 | |||
|
Domain Architecture |
|
|||||
| Description | SH3 domain-binding protein 1 (3BP-1). | |||||
|
CAF1B_MOUSE
|
||||||
| θ value | 6.88961 (rank : 57) | NC score | 0.009205 (rank : 73) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9D0N7 | Gene names | Chaf1b | |||
|
Domain Architecture |
|
|||||
| Description | Chromatin assembly factor 1 subunit B (CAF-1 subunit B) (Chromatin assembly factor I p60 subunit) (CAF-I 60 kDa subunit) (CAF-Ip60). | |||||
|
CD5_HUMAN
|
||||||
| θ value | 6.88961 (rank : 58) | NC score | 0.017515 (rank : 55) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P06127 | Gene names | CD5, LEU1 | |||
|
Domain Architecture |
|
|||||
| Description | T-cell surface glycoprotein CD5 precursor (Lymphocyte antigen T1/Leu- 1). | |||||
|
DCC_MOUSE
|
||||||
| θ value | 6.88961 (rank : 59) | NC score | 0.007972 (rank : 74) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P70211 | Gene names | Dcc | |||
|
Domain Architecture |

|
|||||
| Description | Netrin receptor DCC precursor (Tumor suppressor protein DCC). | |||||
|
DMRT2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 60) | NC score | 0.019492 (rank : 51) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y5R5 | Gene names | DMRT2, DSXL2 | |||
|
Domain Architecture |

|
|||||
| Description | Doublesex- and mab-3-related transcription factor 2 (Doublesex-like 2 protein) (DSXL-2). | |||||
|
FMN1A_MOUSE
|
||||||
| θ value | 6.88961 (rank : 61) | NC score | 0.028856 (rank : 25) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 423 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q05860 | Gene names | Fmn, Ld | |||
|
Domain Architecture |

|
|||||
| Description | Formin-1 isoforms I/II/III (Limb deformity protein). | |||||
|
FMN1B_MOUSE
|
||||||
| θ value | 6.88961 (rank : 62) | NC score | 0.027871 (rank : 27) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q05859 | Gene names | Fmn, Ld | |||
|
Domain Architecture |

|
|||||
| Description | Formin-1 isoform IV (Limb deformity protein). | |||||
|
GPNMB_MOUSE
|
||||||
| θ value | 6.88961 (rank : 63) | NC score | 0.017552 (rank : 54) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99P91, Q8BVV9, Q8BXL4, Q9QXA0 | Gene names | Gpnmb, Dchil, Nmb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane glycoprotein NMB precursor (Dendritic cell-associated transmembrane protein) (DC-HIL). | |||||
|
K0310_HUMAN
|
||||||
| θ value | 6.88961 (rank : 64) | NC score | 0.031987 (rank : 23) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 420 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O15027, Q5SXP0, Q5SXP1, Q8N347, Q96HP1 | Gene names | KIAA0310 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0310. | |||||
|
KCNC3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 65) | NC score | 0.006610 (rank : 76) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 256 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q14003 | Gene names | KCNC3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily C member 3 (Voltage-gated potassium channel subunit Kv3.3) (KSHIIID). | |||||
|
MLL2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 66) | NC score | 0.019609 (rank : 50) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
SN1L2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 67) | NC score | 0.000604 (rank : 84) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 944 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CFH6 | Gene names | Snf1lk2, Sik2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SNF1-like kinase 2 (EC 2.7.11.1) (Salt-inducible kinase 2). | |||||
|
SPR2B_MOUSE
|
||||||
| θ value | 6.88961 (rank : 68) | NC score | 0.046932 (rank : 13) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O70554, Q7TSB2 | Gene names | Sprr2b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Small proline-rich protein 2B. | |||||
|
CBP_MOUSE
|
||||||
| θ value | 8.99809 (rank : 69) | NC score | 0.024421 (rank : 32) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1051 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P45481 | Gene names | Crebbp, Cbp | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
CDX2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 70) | NC score | 0.003326 (rank : 80) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99626, O00503, Q969L8 | Gene names | CDX2, CDX3 | |||
|
Domain Architecture |
|
|||||
| Description | Homeobox protein CDX-2 (Caudal-type homeobox protein 2) (CDX-3). | |||||
|
CS007_HUMAN
|
||||||
| θ value | 8.99809 (rank : 71) | NC score | 0.020616 (rank : 45) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UPT8, Q9Y420 | Gene names | C19orf7, KIAA1064 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein C19orf7. | |||||
|
DIAP2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 72) | NC score | 0.022285 (rank : 41) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O70566, Q8C2G8 | Gene names | Diaph2, Diap2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein diaphanous homolog 2 (Diaphanous-related formin-2) (DRF2) (mDia3). | |||||
|
FGRL1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 73) | NC score | 0.005664 (rank : 77) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8N441, Q6PJN1, Q9BXN7, Q9H4D7 | Gene names | FGFRL1, FGFR5, FHFR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fibroblast growth factor receptor-like 1 precursor (FGF receptor-like protein 1) (Fibroblast growth factor receptor 5) (FGFR-like protein) (FGF homologous factor receptor). | |||||
|
FRAS1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 74) | NC score | 0.010873 (rank : 69) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 759 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q86XX4, Q86UZ4, Q8N3U9, Q8NAU7, Q96JW7, Q9H6N9, Q9P228 | Gene names | FRAS1, KIAA1500 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Extracellular matrix protein FRAS1 precursor. | |||||
|
GR65_HUMAN
|
||||||
| θ value | 8.99809 (rank : 75) | NC score | 0.015230 (rank : 62) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9BQQ3, Q8N272, Q96H42 | Gene names | GORASP1, GRASP65 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi reassembly-stacking protein 1 (Golgi reassembly-stacking protein of 65 kDa) (GRASP65) (Golgi peripheral membrane protein p65) (Golgi phosphoprotein 5) (GOLPH5). | |||||
|
HD_HUMAN
|
||||||
| θ value | 8.99809 (rank : 76) | NC score | 0.015837 (rank : 60) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P42858, Q9UQB7 | Gene names | HD, IT15 | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin (Huntington disease protein) (HD protein). | |||||
|
KLF2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 77) | NC score | 0.001543 (rank : 83) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 738 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q60843 | Gene names | Klf2, Lklf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Krueppel-like factor 2 (Lung krueppel-like factor). | |||||
|
M3K14_HUMAN
|
||||||
| θ value | 8.99809 (rank : 78) | NC score | 0.002116 (rank : 81) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 887 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q99558, Q8IYN1 | Gene names | MAP3K14, NIK | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase kinase kinase 14 (EC 2.7.11.25) (NF- kappa beta-inducing kinase) (Serine/threonine-protein kinase NIK) (HsNIK). | |||||
|
MLL4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 79) | NC score | 0.015925 (rank : 59) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 722 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9UMN6, O15022, O95836, Q96GP2, Q96IP3, Q9UK25, Q9Y668, Q9Y669 | Gene names | MLL4, HRX2, KIAA0304, MLL2, TRX2 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 4 (Trithorax homolog 2). | |||||
|
MTCH1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 80) | NC score | 0.012675 (rank : 66) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NZJ7, Q6PK60, Q6UX45, Q7L465, Q9BW23, Q9NZR6, Q9UJZ5 | Gene names | MTCH1, PSAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitochondrial carrier homolog 1 (Presenilin-associated protein). | |||||
|
PRB3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 81) | NC score | 0.023259 (rank : 36) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q04118, Q15188, Q7M4M9 | Gene names | PRB3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 3 precursor (Parotid salivary glycoprotein G1) (Proline-rich protein G1). | |||||
|
PRCC_HUMAN
|
||||||
| θ value | 8.99809 (rank : 82) | NC score | 0.028202 (rank : 26) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q92733, O00665, O00724 | Gene names | PRCC, TPRC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein PRCC (Papillary renal cell carcinoma translocation-associated gene protein). | |||||
|
SASH1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 83) | NC score | 0.014073 (rank : 63) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O94885, Q8TAI0, Q9H7R7 | Gene names | SASH1, KIAA0790, PEPE1 | |||
|
Domain Architecture |

|
|||||
| Description | SAM and SH3 domain-containing protein 1 (Proline-glutamate repeat- containing protein). | |||||
|
SSXT_MOUSE
|
||||||
| θ value | 8.99809 (rank : 84) | NC score | 0.017758 (rank : 53) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q62280 | Gene names | Ss18, Ssxt, Syt | |||
|
Domain Architecture |
|
|||||
| Description | SSXT protein (SYT protein) (Synovial sarcoma-associated Ss18-alpha). | |||||
|
PIGQ_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 84 | |
| SwissProt Accessions | Q9BRB3, O14927, Q96G00, Q96S22, Q9UJH4 | Gene names | PIGQ, GPI1 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol N-acetylglucosaminyltransferase subunit Q (EC 2.4.1.198) (Phosphatidylinositol-glycan biosynthesis class Q protein) (PIG-Q) (N-acetylglucosamyl transferase component GPI1). | |||||
|
PIGQ_MOUSE
|
||||||
| NC score | 0.965759 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 5 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9QYT7, O35120, O35456, Q99L11 | Gene names | Pigq, Gpi1h, Mgpi1 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol N-acetylglucosaminyltransferase subunit Q (EC 2.4.1.198) (Phosphatidylinositol-glycan biosynthesis class Q protein) (PIG-Q) (N-acetylglucosamyl transferase component GPI1) (MGpi1p). | |||||
|
SSBP4_HUMAN
|
||||||
| NC score | 0.083414 (rank : 3) | θ value | 0.000461057 (rank : 3) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 753 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q9BWG4 | Gene names | SSBP4 | |||
|
Domain Architecture |

|
|||||
| Description | Single-stranded DNA-binding protein 4. | |||||
|
MDFI_HUMAN
|
||||||
| NC score | 0.074821 (rank : 4) | θ value | 0.365318 (rank : 9) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99750 | Gene names | MDFI | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MyoD family inhibitor (Myogenic repressor I-mf). | |||||
|
SPR1A_MOUSE
|
||||||
| NC score | 0.074802 (rank : 5) | θ value | 0.00869519 (rank : 5) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q62266 | Gene names | Sprr1a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cornifin A (Small proline-rich protein 1A) (SPR1A) (SPR1 A). | |||||
|
CN032_HUMAN
|
||||||
| NC score | 0.073416 (rank : 6) | θ value | 0.21417 (rank : 7) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8NDC0, Q96BG5 | Gene names | C14orf32 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf32. | |||||
|
TAF4_HUMAN
|
||||||
| NC score | 0.059598 (rank : 7) | θ value | 0.21417 (rank : 8) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 687 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O00268, Q5TBP6, Q99721, Q9BR40, Q9BX42 | Gene names | TAF4, TAF2C, TAF2C1, TAF4A, TAFII130, TAFII135 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 4 (TBP-associated factor 4) (Transcription initiation factor TFIID 135 kDa subunit) (TAF(II)135) (TAFII-135) (TAFII135) (TAFII-130) (TAFII130). | |||||
|
PROP_MOUSE
|
||||||
| NC score | 0.057920 (rank : 8) | θ value | 0.00665767 (rank : 4) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P11680, Q3TB98, Q3U779 | Gene names | Cfp, Pfc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Properdin precursor (Factor P). | |||||
|
PCX3_MOUSE
|
||||||
| NC score | 0.052492 (rank : 9) | θ value | 0.47712 (rank : 11) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8VI59 | Gene names | Pcnxl3 | |||
|
Domain Architecture |

|
|||||
| Description | Pecanex-like protein 3. | |||||
|
PRRT1_HUMAN
|
||||||
| NC score | 0.051231 (rank : 10) | θ value | 2.36792 (rank : 26) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q99946, Q96DW3, Q96NQ8 | Gene names | PRRT1, C6orf31, NG5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich transmembrane protein 1. | |||||
|
PRRT1_MOUSE
|
||||||
| NC score | 0.049686 (rank : 11) | θ value | 2.36792 (rank : 27) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O35449 | Gene names | Prrt1, Ng5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich transmembrane protein 1. | |||||
|
IF35_HUMAN
|
||||||
| NC score | 0.047117 (rank : 12) | θ value | 1.38821 (rank : 18) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O00303, Q8N978 | Gene names | EIF3S5 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 3 subunit 5 (eIF-3 epsilon) (eIF3 p47 subunit) (eIF3f). | |||||
|
SPR2B_MOUSE
|
||||||
| NC score | 0.046932 (rank : 13) | θ value | 6.88961 (rank : 68) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O70554, Q7TSB2 | Gene names | Sprr2b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Small proline-rich protein 2B. | |||||
|
PRR12_HUMAN
|
||||||
| NC score | 0.046511 (rank : 14) | θ value | 1.81305 (rank : 21) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 460 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9ULL5, Q8N4J6 | Gene names | PRR12, KIAA1205 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 12. | |||||
|
WIRE_MOUSE
|
||||||
| NC score | 0.045583 (rank : 15) | θ value | 0.62314 (rank : 12) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q6PEV3 | Gene names | Wire | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WIP-related protein (WASP-interacting protein-related protein). | |||||
|
PROP_HUMAN
|
||||||
| NC score | 0.043225 (rank : 16) | θ value | 1.38821 (rank : 19) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P27918, O15134, O15135, O15136, O75826 | Gene names | CFP, PFC | |||
|
Domain Architecture |

|
|||||
| Description | Properdin precursor (Factor P). | |||||
|
ZC3H3_MOUSE
|
||||||
| NC score | 0.042949 (rank : 17) | θ value | 0.62314 (rank : 13) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8CHP0, Q80X78 | Gene names | Zc3h3, Kiaa0150, Zc3hdc3, Zfp623 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 3. | |||||
|
IF35_MOUSE
|
||||||
| NC score | 0.042692 (rank : 18) | θ value | 2.36792 (rank : 24) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9DCH4 | Gene names | Eif3s5 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 3 subunit 5 (eIF-3 epsilon) (eIF3 p47 subunit) (eIF3f). | |||||
|
RBM12_MOUSE
|
||||||
| NC score | 0.038639 (rank : 19) | θ value | 1.38821 (rank : 20) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 313 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8R4X3, Q8K373, Q8R302, Q9CS80 | Gene names | Rbm12 | |||
|
Domain Architecture |

|
|||||
| Description | RNA-binding protein 12 (RNA-binding motif protein 12) (SH3/WW domain anchor protein in the nucleus) (SWAN). | |||||
|
TDGF1_MOUSE
|
||||||
| NC score | 0.038518 (rank : 20) | θ value | 0.813845 (rank : 14) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P51865 | Gene names | Tdgf1, Cripto | |||
|
Domain Architecture |

|
|||||
| Description | Teratocarcinoma-derived growth factor precursor (Epidermal growth factor-like Cripto protein) (Cripto growth factor). | |||||
|
DMWD_MOUSE
|
||||||
| NC score | 0.037881 (rank : 21) | θ value | 0.47712 (rank : 10) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q08274 | Gene names | Dmwd, Dm9 | |||
|
Domain Architecture |

|
|||||
| Description | Dystrophia myotonica WD repeat-containing protein (Dystrophia myotonica-containing WD repeat motif protein) (DMR-N9 protein). | |||||
|
MBD6_HUMAN
|
||||||
| NC score | 0.036268 (rank : 22) | θ value | 3.0926 (rank : 34) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q96DN6, Q8N3M0, Q8NA81, Q96Q00 | Gene names | MBD6, KIAA1887 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 6 (Methyl-CpG-binding protein MBD6). | |||||
|
K0310_HUMAN
|
||||||
| NC score | 0.031987 (rank : 23) | θ value | 6.88961 (rank : 64) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 420 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O15027, Q5SXP0, Q5SXP1, Q8N347, Q96HP1 | Gene names | KIAA0310 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0310. | |||||
|
ARI1A_HUMAN
|
||||||
| NC score | 0.030347 (rank : 24) | θ value | 3.0926 (rank : 31) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 787 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O14497, Q53FK9, Q5T0W1, Q5T0W2, Q5T0W3, Q8NFD6, Q96T89, Q9BY33, Q9HBJ5, Q9UPZ1 | Gene names | ARID1A, C1orf4, OSA1, SMARCF1 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 1A (ARID domain- containing protein 1A) (SWI/SNF-related, matrix-associated, actin- dependent regulator of chromatin subfamily F member 1) (SWI-SNF complex protein p270) (B120) (SWI-like protein) (Osa homolog 1) (hOSA1) (hELD) (BRG1-associated factor 250) (BAF250) (BRG1-associated factor 250a) (BAF250A). | |||||
|
FMN1A_MOUSE
|
||||||
| NC score | 0.028856 (rank : 25) | θ value | 6.88961 (rank : 61) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 423 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q05860 | Gene names | Fmn, Ld | |||
|
Domain Architecture |

|
|||||
| Description | Formin-1 isoforms I/II/III (Limb deformity protein). | |||||
|
PRCC_HUMAN
|
||||||
| NC score | 0.028202 (rank : 26) | θ value | 8.99809 (rank : 82) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q92733, O00665, O00724 | Gene names | PRCC, TPRC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein PRCC (Papillary renal cell carcinoma translocation-associated gene protein). | |||||
|
FMN1B_MOUSE
|
||||||
| NC score | 0.027871 (rank : 27) | θ value | 6.88961 (rank : 62) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q05859 | Gene names | Fmn, Ld | |||
|
Domain Architecture |

|
|||||
| Description | Formin-1 isoform IV (Limb deformity protein). | |||||
|
DIAP1_HUMAN
|
||||||
| NC score | 0.027718 (rank : 28) | θ value | 1.06291 (rank : 15) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 784 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O60610, Q9UC76 | Gene names | DIAPH1, DIAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein diaphanous homolog 1 (Diaphanous-related formin-1) (DRF1). | |||||
|
PLXB1_MOUSE
|
||||||
| NC score | 0.025972 (rank : 29) | θ value | 1.06291 (rank : 16) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8CJH3, Q6ZQC3, Q80ZZ1 | Gene names | Plxnb1, Kiaa0407 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-B1 precursor. | |||||
|
PLXB1_HUMAN
|
||||||
| NC score | 0.025810 (rank : 30) | θ value | 4.03905 (rank : 41) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O43157, Q6NY20, Q9UIV7, Q9UJ92, Q9UJ93 | Gene names | PLXNB1, KIAA0407, SEP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plexin-B1 precursor (Semaphorin receptor SEP). | |||||
|
FUBP1_HUMAN
|
||||||
| NC score | 0.024437 (rank : 31) | θ value | 5.27518 (rank : 45) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96AE4, Q12828 | Gene names | FUBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Far upstream element-binding protein 1 (FUSE-binding protein 1) (FBP) (DNA helicase V) (HDH V). | |||||
|
CBP_MOUSE
|
||||||
| NC score | 0.024421 (rank : 32) | θ value | 8.99809 (rank : 69) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1051 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P45481 | Gene names | Crebbp, Cbp | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
FUBP1_MOUSE
|
||||||
| NC score | 0.024420 (rank : 33) | θ value | 5.27518 (rank : 46) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q91WJ8, Q8C0Y8 | Gene names | Fubp1, D3Ertd330e | |||
|
Domain Architecture |

|
|||||
| Description | Far upstream element-binding protein 1 (FUSE-binding protein 1) (FBP). | |||||
|
F113A_HUMAN
|
||||||
| NC score | 0.024368 (rank : 34) | θ value | 5.27518 (rank : 43) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H1Q7, Q5JUA5, Q5JUA6, Q6PK19, Q86WF5, Q96CG7, Q9H1Q6, Q9H6D1 | Gene names | FAM113A, C20orf81 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM113A (Sarcoma antigen NY-SAR-23). | |||||
|
ZN414_HUMAN
|
||||||
| NC score | 0.023867 (rank : 35) | θ value | 2.36792 (rank : 30) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 548 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96IQ9 | Gene names | ZNF414 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 414. | |||||
|
PRB3_HUMAN
|
||||||
| NC score | 0.023259 (rank : 36) | θ value | 8.99809 (rank : 81) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q04118, Q15188, Q7M4M9 | Gene names | PRB3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 3 precursor (Parotid salivary glycoprotein G1) (Proline-rich protein G1). | |||||
|
STAB1_HUMAN
|
||||||
| NC score | 0.023077 (rank : 37) | θ value | 0.125558 (rank : 6) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 546 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9NY15, Q8IUH0, Q8IUH1, Q93072 | Gene names | STAB1, FEEL1, KIAA0246 | |||
|
Domain Architecture |

|
|||||
| Description | Stabilin-1 precursor (FEEL-1 protein) (MS-1 antigen). | |||||
|
LBXCO_HUMAN
|
||||||
| NC score | 0.022880 (rank : 38) | θ value | 4.03905 (rank : 40) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P84550 | Gene names | LBXCOR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ladybird homeobox corepressor 1. | |||||
|
ENAH_MOUSE
|
||||||
| NC score | 0.022687 (rank : 39) | θ value | 2.36792 (rank : 23) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 919 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q03173, P70430, P70431, P70432, P70433, Q5D053 | Gene names | Enah, Mena, Ndpp1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein enabled homolog (NPC-derived proline-rich protein 1) (NDPP-1). | |||||
|
COE4_MOUSE
|
||||||
| NC score | 0.022644 (rank : 40) | θ value | 3.0926 (rank : 32) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8K4J2, Q8K4J1, Q8K4J3, Q8K4J4, Q8K4J5 | Gene names | Ebf4, Coe4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor COE4 (Early B-cell factor 4) (EBF-4) (Olf-1/EBF- like 4) (OE-4) (O/E-4). | |||||
|
DIAP2_MOUSE
|
||||||
| NC score | 0.022285 (rank : 41) | θ value | 8.99809 (rank : 72) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O70566, Q8C2G8 | Gene names | Diaph2, Diap2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein diaphanous homolog 2 (Diaphanous-related formin-2) (DRF2) (mDia3). | |||||
|
JUN_MOUSE
|
||||||
| NC score | 0.021780 (rank : 42) | θ value | 3.0926 (rank : 33) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P05627 | Gene names | Jun | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor AP-1 (Activator protein 1) (AP1) (Proto-oncogene c-jun) (V-jun avian sarcoma virus 17 oncogene homolog) (Jun A) (AH119). | |||||
|
NCOR2_HUMAN
|
||||||
| NC score | 0.021268 (rank : 43) | θ value | 2.36792 (rank : 25) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 870 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9Y618, O00613, O15416, Q13354, Q9Y5U0 | Gene names | NCOR2, CTG26 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC) (CTG repeat protein 26) (SMAP270). | |||||
|
RN167_HUMAN
|
||||||
| NC score | 0.020841 (rank : 44) | θ value | 2.36792 (rank : 29) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H6Y7, Q6XYE0, Q8NDC1, Q9Y3V1 | Gene names | RNF167 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 167 precursor. | |||||
|
CS007_HUMAN
|
||||||
| NC score | 0.020616 (rank : 45) | θ value | 8.99809 (rank : 71) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UPT8, Q9Y420 | Gene names | C19orf7, KIAA1064 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein C19orf7. | |||||
|
COE4_HUMAN
|
||||||
| NC score | 0.020555 (rank : 46) | θ value | 5.27518 (rank : 42) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9BQW3, Q5JY53, Q9NUB6, Q9P2A6 | Gene names | EBF4, COE4, KIAA1442 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor COE4 (Early B-cell factor 4) (EBF-4) (Olf-1/EBF- like 4) (OE-4) (O/E-4). | |||||
|
IRS2_HUMAN
|
||||||
| NC score | 0.019903 (rank : 47) | θ value | 5.27518 (rank : 49) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9Y4H2, Q96RR2, Q9BZG0, Q9Y6I5 | Gene names | IRS2 | |||
|
Domain Architecture |

|
|||||
| Description | Insulin receptor substrate 2 (IRS-2). | |||||
|
SEM5B_HUMAN
|
||||||
| NC score | 0.019897 (rank : 48) | θ value | 3.0926 (rank : 35) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9P283, Q6UY12 | Gene names | SEMA5B, KIAA1445 | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-5B precursor. | |||||
|
PRGC2_HUMAN
|
||||||
| NC score | 0.019705 (rank : 49) | θ value | 5.27518 (rank : 51) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 272 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q86YN6, Q86YN3, Q86YN4, Q86YN5, Q8N1N9, Q8TDE4, Q8TDE5 | Gene names | PPARGC1B, PERC, PGC1, PGC1B, PPARGC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor gamma coactivator 1-beta (PPAR gamma coactivator-1beta) (PGC-1-related estrogen receptor alpha coactivator) (PPARGC-1-beta) (PGC-1-beta). | |||||
|
MLL2_HUMAN
|
||||||
| NC score | 0.019609 (rank : 50) | θ value | 6.88961 (rank : 66) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
DMRT2_HUMAN
|
||||||
| NC score | 0.019492 (rank : 51) | θ value | 6.88961 (rank : 60) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y5R5 | Gene names | DMRT2, DSXL2 | |||
|
Domain Architecture |

|
|||||
| Description | Doublesex- and mab-3-related transcription factor 2 (Doublesex-like 2 protein) (DSXL-2). | |||||
|
RFX1_HUMAN
|
||||||
| NC score | 0.018548 (rank : 52) | θ value | 5.27518 (rank : 52) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 256 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P22670 | Gene names | RFX1 | |||
|
Domain Architecture |

|
|||||
| Description | MHC class II regulatory factor RFX1 (RFX) (Enhancer factor C) (EF-C). | |||||
|
SSXT_MOUSE
|
||||||
| NC score | 0.017758 (rank : 53) | θ value | 8.99809 (rank : 84) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q62280 | Gene names | Ss18, Ssxt, Syt | |||
|
Domain Architecture |

|
|||||
| Description | SSXT protein (SYT protein) (Synovial sarcoma-associated Ss18-alpha). | |||||
|
GPNMB_MOUSE
|
||||||
| NC score | 0.017552 (rank : 54) | θ value | 6.88961 (rank : 63) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99P91, Q8BVV9, Q8BXL4, Q9QXA0 | Gene names | Gpnmb, Dchil, Nmb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane glycoprotein NMB precursor (Dendritic cell-associated transmembrane protein) (DC-HIL). | |||||
|
CD5_HUMAN
|
||||||
| NC score | 0.017515 (rank : 55) | θ value | 6.88961 (rank : 58) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P06127 | Gene names | CD5, LEU1 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell surface glycoprotein CD5 precursor (Lymphocyte antigen T1/Leu- 1). | |||||
|
PTN23_MOUSE
|
||||||
| NC score | 0.016827 (rank : 56) | θ value | 2.36792 (rank : 28) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 726 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q6PB44, Q69ZJ0, Q8R1Z5, Q923E6 | Gene names | Ptpn23, Kiaa1471 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 23 (EC 3.1.3.48). | |||||
|
COHA1_HUMAN
|
||||||
| NC score | 0.016492 (rank : 57) | θ value | 4.03905 (rank : 37) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 400 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UMD9, Q02802, Q99018, Q9NQK9, Q9UC14 | Gene names | COL17A1, BP180, BPAG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XVII) chain (Bullous pemphigoid antigen 2) (180 kDa bullous pemphigoid antigen 2). | |||||
|
SIAH2_MOUSE
|
||||||
| NC score | 0.015987 (rank : 58) | θ value | 5.27518 (rank : 53) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q06986 | Gene names | Siah2 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin ligase SIAH2 (EC 6.3.2.-) (Seven in absentia homolog 2) (Siah-2) (mSiah2). | |||||
|
MLL4_HUMAN
|
||||||
| NC score | 0.015925 (rank : 59) | θ value | 8.99809 (rank : 79) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 722 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9UMN6, O15022, O95836, Q96GP2, Q96IP3, Q9UK25, Q9Y668, Q9Y669 | Gene names | MLL4, HRX2, KIAA0304, MLL2, TRX2 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 4 (Trithorax homolog 2). | |||||
|
HD_HUMAN
|
||||||
| NC score | 0.015837 (rank : 60) | θ value | 8.99809 (rank : 76) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P42858, Q9UQB7 | Gene names | HD, IT15 | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin (Huntington disease protein) (HD protein). | |||||
|
ST5_HUMAN
|
||||||
| NC score | 0.015815 (rank : 61) | θ value | 3.0926 (rank : 36) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P78524, P78523, Q16492, Q7KYY2, Q7KZ12, Q8NE12, Q9BQQ6 | Gene names | ST5, DENND2B, HTS1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppression of tumorigenicity 5 (HeLa tumor suppression 1) (DENN domain-containing protein 2B). | |||||
|
GR65_HUMAN
|
||||||
| NC score | 0.015230 (rank : 62) | θ value | 8.99809 (rank : 75) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9BQQ3, Q8N272, Q96H42 | Gene names | GORASP1, GRASP65 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi reassembly-stacking protein 1 (Golgi reassembly-stacking protein of 65 kDa) (GRASP65) (Golgi peripheral membrane protein p65) (Golgi phosphoprotein 5) (GOLPH5). | |||||
|
SASH1_HUMAN
|
||||||
| NC score | 0.014073 (rank : 63) | θ value | 8.99809 (rank : 83) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O94885, Q8TAI0, Q9H7R7 | Gene names | SASH1, KIAA0790, PEPE1 | |||
|
Domain Architecture |

|
|||||
| Description | SAM and SH3 domain-containing protein 1 (Proline-glutamate repeat- containing protein). | |||||
|
GP152_HUMAN
|
||||||
| NC score | 0.013955 (rank : 64) | θ value | 4.03905 (rank : 39) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 636 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8TDT2, Q86SM0 | Gene names | GPR152, PGR5 | |||
|
Domain Architecture |

|
|||||
| Description | Probable G-protein coupled receptor 152 (G-protein coupled receptor PGR5). | |||||
|
MUC5B_HUMAN
|
||||||
| NC score | 0.013480 (rank : 65) | θ value | 5.27518 (rank : 50) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 1020 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9HC84, O00447, O00573, O14985, O15494, O95291, O95451, Q14881, Q99552, Q9UE28 | Gene names | MUC5B, MUC5 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-5B precursor (Mucin-5 subtype B, tracheobronchial) (High molecular weight salivary mucin MG1) (Sublingual gland mucin). | |||||
|
MTCH1_HUMAN
|
||||||
| NC score | 0.012675 (rank : 66) | θ value | 8.99809 (rank : 80) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NZJ7, Q6PK60, Q6UX45, Q7L465, Q9BW23, Q9NZR6, Q9UJZ5 | Gene names | MTCH1, PSAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitochondrial carrier homolog 1 (Presenilin-associated protein). | |||||
|
3BP1_HUMAN
|
||||||
| NC score | 0.012235 (rank : 67) | θ value | 6.88961 (rank : 56) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 376 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9Y3L3, Q6IBZ2, Q6ZVL9, Q96HQ5, Q9NSQ9 | Gene names | SH3BP1 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 domain-binding protein 1 (3BP-1). | |||||
|
ADA15_HUMAN
|
||||||
| NC score | 0.012180 (rank : 68) | θ value | 1.38821 (rank : 17) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 288 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13444, Q13493, Q96C78 | Gene names | ADAM15, MDC15 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 15 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 15) (Metalloproteinase-like, disintegrin-like, and cysteine- rich protein 15) (MDC-15) (Metalloprotease RGD disintegrin protein) (Metargidin). | |||||
|
FRAS1_HUMAN
|
||||||
| NC score | 0.010873 (rank : 69) | θ value | 8.99809 (rank : 74) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 759 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q86XX4, Q86UZ4, Q8N3U9, Q8NAU7, Q96JW7, Q9H6N9, Q9P228 | Gene names | FRAS1, KIAA1500 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Extracellular matrix protein FRAS1 precursor. | |||||
|
GRIP1_MOUSE
|
||||||
| NC score | 0.010709 (rank : 70) | θ value | 5.27518 (rank : 47) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q925T6, Q8BLQ3, Q8C0T3, Q925T5, Q925T7 | Gene names | Grip1 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate receptor-interacting protein 1 (GRIP1 protein). | |||||
|
SMCA2_HUMAN
|
||||||
| NC score | 0.010533 (rank : 71) | θ value | 5.27518 (rank : 54) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 503 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P51531 | Gene names | SMARCA2, BRM, SNF2A, SNF2L2 | |||
|
Domain Architecture |

|
|||||
| Description | Probable global transcription activator SNF2L2 (EC 3.6.1.-) (ATP- dependent helicase SMARCA2) (SNF2-alpha) (SWI/SNF-related matrix- associated actin-dependent regulator of chromatin subfamily A member 2) (hBRM). | |||||
|
RXRB_MOUSE
|
||||||
| NC score | 0.009739 (rank : 72) | θ value | 1.81305 (rank : 22) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 230 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P28704, P33243 | Gene names | Rxrb, Nr2b2 | |||
|
Domain Architecture |

|
|||||
| Description | Retinoic acid receptor RXR-beta (Retinoid X receptor beta) (MHC class I regulatory element-binding protein H-2RIIBP). | |||||
|
CAF1B_MOUSE
|
||||||
| NC score | 0.009205 (rank : 73) | θ value | 6.88961 (rank : 57) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9D0N7 | Gene names | Chaf1b | |||
|
Domain Architecture |

|
|||||
| Description | Chromatin assembly factor 1 subunit B (CAF-1 subunit B) (Chromatin assembly factor I p60 subunit) (CAF-I 60 kDa subunit) (CAF-Ip60). | |||||
|
DCC_MOUSE
|
||||||
| NC score | 0.007972 (rank : 74) | θ value | 6.88961 (rank : 59) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P70211 | Gene names | Dcc | |||
|
Domain Architecture |

|
|||||
| Description | Netrin receptor DCC precursor (Tumor suppressor protein DCC). | |||||
|
FOG1_HUMAN
|
||||||
| NC score | 0.006850 (rank : 75) | θ value | 5.27518 (rank : 44) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 981 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8IX07 | Gene names | ZFPM1, FOG1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein ZFPM1 (Zinc finger protein multitype 1) (Friend of GATA protein 1) (Friend of GATA-1) (FOG-1). | |||||
|
KCNC3_HUMAN
|
||||||
| NC score | 0.006610 (rank : 76) | θ value | 6.88961 (rank : 65) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 256 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q14003 | Gene names | KCNC3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily C member 3 (Voltage-gated potassium channel subunit Kv3.3) (KSHIIID). | |||||
|
FGRL1_HUMAN
|
||||||
| NC score | 0.005664 (rank : 77) | θ value | 8.99809 (rank : 73) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 335 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8N441, Q6PJN1, Q9BXN7, Q9H4D7 | Gene names | FGFRL1, FGFR5, FHFR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fibroblast growth factor receptor-like 1 precursor (FGF receptor-like protein 1) (Fibroblast growth factor receptor 5) (FGFR-like protein) (FGF homologous factor receptor). | |||||
|
IRAK1_HUMAN
|
||||||
| NC score | 0.004109 (rank : 78) | θ value | 5.27518 (rank : 48) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 820 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P51617, Q7Z5V4, Q96RL2 | Gene names | IRAK1, IRAK | |||
|
Domain Architecture |

|
|||||
| Description | Interleukin-1 receptor-associated kinase 1 (EC 2.7.11.1) (IRAK-1). | |||||
|
DYR1B_HUMAN
|
||||||
| NC score | 0.004064 (rank : 79) | θ value | 4.03905 (rank : 38) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 831 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Y463, O75258, O75788, O75789 | Gene names | DYRK1B, MIRK | |||
|
Domain Architecture |

|
|||||
| Description | Dual specificity tyrosine-phosphorylation-regulated kinase 1B (EC 2.7.12.1) (Mirk protein kinase) (Minibrain-related kinase). | |||||
|
CDX2_HUMAN
|
||||||
| NC score | 0.003326 (rank : 80) | θ value | 8.99809 (rank : 70) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99626, O00503, Q969L8 | Gene names | CDX2, CDX3 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein CDX-2 (Caudal-type homeobox protein 2) (CDX-3). | |||||
|
M3K14_HUMAN
|
||||||
| NC score | 0.002116 (rank : 81) | θ value | 8.99809 (rank : 78) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 887 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q99558, Q8IYN1 | Gene names | MAP3K14, NIK | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase kinase kinase 14 (EC 2.7.11.25) (NF- kappa beta-inducing kinase) (Serine/threonine-protein kinase NIK) (HsNIK). | |||||
|
ZXDA_HUMAN
|
||||||
| NC score | 0.001595 (rank : 82) | θ value | 5.27518 (rank : 55) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 750 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P98168, Q9UJP7 | Gene names | ZXDA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger X-linked protein ZXDA. | |||||
|
KLF2_MOUSE
|
||||||
| NC score | 0.001543 (rank : 83) | θ value | 8.99809 (rank : 77) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 738 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q60843 | Gene names | Klf2, Lklf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Krueppel-like factor 2 (Lung krueppel-like factor). | |||||
|
SN1L2_MOUSE
|
||||||
| NC score | 0.000604 (rank : 84) | θ value | 6.88961 (rank : 67) | |||
| Query Neighborhood Hits | 84 | Target Neighborhood Hits | 944 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CFH6 | Gene names | Snf1lk2, Sik2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SNF1-like kinase 2 (EC 2.7.11.1) (Salt-inducible kinase 2). | |||||