Please be patient as the page loads
|
MA1B1_HUMAN
|
||||||
| SwissProt Accessions | Q9UKM7, Q5VSG3, Q9BRS9, Q9Y5K7 | Gene names | MAN1B1 | |||
|
Domain Architecture |

|
|||||
| Description | Endoplasmic reticulum mannosyl-oligosaccharide 1,2-alpha-mannosidase (EC 3.2.1.113) (ER alpha-1,2-mannosidase) (Mannosidase alpha class 1B member 1) (Man9GlcNAc2-specific-processing alpha-mannosidase). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
MA1B1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 125 | |
| SwissProt Accessions | Q9UKM7, Q5VSG3, Q9BRS9, Q9Y5K7 | Gene names | MAN1B1 | |||
|
Domain Architecture |

|
|||||
| Description | Endoplasmic reticulum mannosyl-oligosaccharide 1,2-alpha-mannosidase (EC 3.2.1.113) (ER alpha-1,2-mannosidase) (Mannosidase alpha class 1B member 1) (Man9GlcNAc2-specific-processing alpha-mannosidase). | |||||
|
MA1A1_MOUSE
|
||||||
| θ value | 1.39714e-93 (rank : 2) | NC score | 0.931012 (rank : 4) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P45700 | Gene names | Man1a1, Man1a | |||
|
Domain Architecture |

|
|||||
| Description | Mannosyl-oligosaccharide 1,2-alpha-mannosidase IA (EC 3.2.1.113) (Processing alpha-1,2-mannosidase IA) (Alpha-1,2-mannosidase IA) (Mannosidase alpha class 1A member 1) (Man(9)-alpha-mannosidase) (Man9-mannosidase). | |||||
|
MA1A1_HUMAN
|
||||||
| θ value | 6.07444e-89 (rank : 3) | NC score | 0.931227 (rank : 3) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P33908, Q9NU44, Q9UJI3 | Gene names | MAN1A1 | |||
|
Domain Architecture |

|
|||||
| Description | Mannosyl-oligosaccharide 1,2-alpha-mannosidase IA (EC 3.2.1.113) (Processing alpha-1,2-mannosidase IA) (Alpha-1,2-mannosidase IA) (Mannosidase alpha class 1A member 1) (Man(9)-alpha-mannosidase) (Man9-mannosidase). | |||||
|
MA1A2_MOUSE
|
||||||
| θ value | 2.30828e-88 (rank : 4) | NC score | 0.930026 (rank : 6) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P39098, Q5SUY6, Q5SUY7, Q60599, Q9CUY9 | Gene names | Man1a2, Man1b | |||
|
Domain Architecture |

|
|||||
| Description | Mannosyl-oligosaccharide 1,2-alpha-mannosidase IB (EC 3.2.1.113) (Processing alpha-1,2-mannosidase IB) (Alpha-1,2-mannosidase IB) (Mannosidase alpha class 1A member 2). | |||||
|
MA1A2_HUMAN
|
||||||
| θ value | 3.93732e-88 (rank : 5) | NC score | 0.930330 (rank : 5) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O60476, Q9H510 | Gene names | MAN1A2, MAN1B | |||
|
Domain Architecture |
|
|||||
| Description | Mannosyl-oligosaccharide 1,2-alpha-mannosidase IB (EC 3.2.1.113) (Processing alpha-1,2-mannosidase IB) (Alpha-1,2-mannosidase IB) (Mannosidase alpha class 1A member 2). | |||||
|
MA1C1_HUMAN
|
||||||
| θ value | 1.40038e-85 (rank : 6) | NC score | 0.934773 (rank : 2) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9NR34, Q9Y545 | Gene names | MAN1C1, MAN1A3, MAN1C | |||
|
Domain Architecture |

|
|||||
| Description | Mannosyl-oligosaccharide 1,2-alpha-mannosidase IC (EC 3.2.1.113) (Processing alpha-1,2-mannosidase IC) (Alpha-1,2-mannosidase IC) (Mannosidase alpha class 1C member 1) (HMIC). | |||||
|
EDEM3_HUMAN
|
||||||
| θ value | 2.25659e-51 (rank : 7) | NC score | 0.850558 (rank : 10) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9BZQ6, Q5TEZ0, Q9HCW1, Q9UFV7 | Gene names | EDEM3, C1orf22 | |||
|
Domain Architecture |

|
|||||
| Description | ER degradation-enhancing alpha-mannosidase-like 3. | |||||
|
EDEM1_MOUSE
|
||||||
| θ value | 5.20228e-48 (rank : 8) | NC score | 0.860645 (rank : 9) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q925U4 | Gene names | Edem1, Edem | |||
|
Domain Architecture |

|
|||||
| Description | ER degradation-enhancing alpha-mannosidase-like 1. | |||||
|
EDEM1_HUMAN
|
||||||
| θ value | 1.51363e-47 (rank : 9) | NC score | 0.861070 (rank : 8) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q92611 | Gene names | EDEM1, EDEM, KIAA0212 | |||
|
Domain Architecture |

|
|||||
| Description | ER degradation-enhancing alpha-mannosidase-like 1. | |||||
|
EDEM2_HUMAN
|
||||||
| θ value | 5.03882e-43 (rank : 10) | NC score | 0.884530 (rank : 7) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9BV94, Q6IA89, Q6UWZ4, Q9H4U0, Q9H886, Q9NTL9, Q9NVE6 | Gene names | EDEM2, C20orf31, C20orf49 | |||
|
Domain Architecture |

|
|||||
| Description | ER degradation-enhancing alpha-mannosidase-like 2 precursor. | |||||
|
SFPQ_MOUSE
|
||||||
| θ value | 0.00228821 (rank : 11) | NC score | 0.046285 (rank : 29) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8VIJ6, Q9ERW2 | Gene names | Sfpq | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, proline- and glutamine-rich (Polypyrimidine tract- binding protein-associated-splicing factor) (PTB-associated-splicing factor) (PSF) (DNA-binding p52/p100 complex, 100 kDa subunit). | |||||
|
NCOA6_HUMAN
|
||||||
| θ value | 0.00509761 (rank : 12) | NC score | 0.087132 (rank : 11) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1202 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q14686, Q9NTZ9, Q9UH74, Q9UK86 | Gene names | NCOA6, AIB3, KIAA0181, RAP250, TRBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (PRIP) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC RAP250) (Thyroid hormone receptor-binding protein). | |||||
|
MNT_MOUSE
|
||||||
| θ value | 0.00869519 (rank : 13) | NC score | 0.076802 (rank : 13) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 432 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O08789, P97349 | Gene names | Mnt, Rox | |||
|
Domain Architecture |

|
|||||
| Description | Max-binding protein MNT (Protein ROX) (Myc antagonist MNT). | |||||
|
PRP1_HUMAN
|
||||||
| θ value | 0.00869519 (rank : 14) | NC score | 0.059001 (rank : 17) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P04280, Q08805, Q15186, Q15187, Q15214, Q15215, Q16038 | Gene names | PRB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 1 precursor (Salivary proline-rich protein) [Contains: Basic peptide IB-6; Peptide P-H]. | |||||
|
ANDR_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 15) | NC score | 0.022192 (rank : 78) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P19091 | Gene names | Ar, Nr3c4 | |||
|
Domain Architecture |

|
|||||
| Description | Androgen receptor (Dihydrotestosterone receptor). | |||||
|
PRB3_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 16) | NC score | 0.053287 (rank : 22) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q04118, Q15188, Q7M4M9 | Gene names | PRB3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 3 precursor (Parotid salivary glycoprotein G1) (Proline-rich protein G1). | |||||
|
CCNK_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 17) | NC score | 0.047301 (rank : 27) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O88874, Q8R068 | Gene names | Ccnk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-K. | |||||
|
MILK1_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 18) | NC score | 0.051284 (rank : 24) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 788 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q8BGT6, Q8BJ60 | Gene names | Mirab13, D15Mit260, Kiaa1668 | |||
|
Domain Architecture |

|
|||||
| Description | Molecule interacting with Rab13 (MIRab13) (MICAL-like protein 1). | |||||
|
PRB4S_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 19) | NC score | 0.051763 (rank : 23) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P10163, P02813 | Gene names | PRB4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 4 allele S precursor (Salivary proline-rich protein Po) (Parotid o protein) [Contains: Protein N1; Glycosylated protein A]. | |||||
|
ADDA_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 20) | NC score | 0.038988 (rank : 40) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9QYC0, Q9JLE3 | Gene names | Add1 | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-adducin (Erythrocyte adducin subunit alpha). | |||||
|
MADCA_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 21) | NC score | 0.064186 (rank : 15) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q13477, O60222, O75867 | Gene names | MADCAM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucosal addressin cell adhesion molecule 1 precursor (MAdCAM-1) (hMAdCAM-1). | |||||
|
MNT_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 22) | NC score | 0.070932 (rank : 14) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 496 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q99583 | Gene names | MNT, ROX | |||
|
Domain Architecture |

|
|||||
| Description | Max-binding protein MNT (Protein ROX) (Myc antagonist MNT). | |||||
|
PRP2_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 23) | NC score | 0.048883 (rank : 26) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P05142 | Gene names | Prh1, Prp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein MP-2 precursor. | |||||
|
TITIN_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 24) | NC score | 0.009854 (rank : 108) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 2923 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8WZ42, Q10465, Q10466, Q15598, Q2XUS3, Q32Q60, Q4U1Z6, Q6NSG0, Q6PDB1, Q6PJP0, Q7KYM2, Q7KYN4, Q7KYN5, Q7LDM3, Q7Z2X3, Q8TCG8, Q8WZ51, Q8WZ52, Q8WZ53, Q8WZB3, Q92761, Q92762, Q9UD97, Q9UP84, Q9Y6L9 | Gene names | TTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Titin (EC 2.7.11.1) (Connectin) (Rhabdomyosarcoma antigen MU-RMS- 40.14). | |||||
|
B3A3_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 25) | NC score | 0.030534 (rank : 54) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P48751 | Gene names | SLC4A3, AE3 | |||
|
Domain Architecture |

|
|||||
| Description | Anion exchange protein 3 (Neuronal band 3-like protein) (Solute carrier family 4 member 3) (Cardiac/brain band 3-like protein) (CAE3/BAE3). | |||||
|
SPTB2_HUMAN
|
||||||
| θ value | 0.163984 (rank : 26) | NC score | 0.012544 (rank : 99) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 748 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q01082, O60837, Q16057 | Gene names | SPTBN1, SPTB2 | |||
|
Domain Architecture |

|
|||||
| Description | Spectrin beta chain, brain 1 (Spectrin, non-erythroid beta chain 1) (Beta-II spectrin) (Fodrin beta chain). | |||||
|
BSN_HUMAN
|
||||||
| θ value | 0.21417 (rank : 27) | NC score | 0.045568 (rank : 30) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
CBP_HUMAN
|
||||||
| θ value | 0.279714 (rank : 28) | NC score | 0.042854 (rank : 37) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 941 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q92793, O00147, Q16376 | Gene names | CREBBP, CBP | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
ANK2_HUMAN
|
||||||
| θ value | 0.365318 (rank : 29) | NC score | 0.024279 (rank : 72) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
BAT2_HUMAN
|
||||||
| θ value | 0.365318 (rank : 30) | NC score | 0.053305 (rank : 21) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P48634, O95875, Q5SQ29, Q5SQ30, Q5ST84, Q5STX6, Q5STX7, Q68DW9, Q6P9P7, Q6PIN1, Q96QC6 | Gene names | BAT2, G2 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
SATB1_MOUSE
|
||||||
| θ value | 0.365318 (rank : 31) | NC score | 0.037711 (rank : 43) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q60611 | Gene names | Satb1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-binding protein SATB1 (Special AT-rich sequence-binding protein 1). | |||||
|
SGIP1_HUMAN
|
||||||
| θ value | 0.365318 (rank : 32) | NC score | 0.045304 (rank : 31) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9BQI5, Q4LE32, Q5VYE2, Q5VYE3, Q5VYE4, Q68D76, Q6MZY6, Q8IWC2 | Gene names | SGIP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH3-containing GRB2-like protein 3-interacting protein 1 (Endophilin- 3-interacting protein). | |||||
|
BC11B_HUMAN
|
||||||
| θ value | 0.47712 (rank : 33) | NC score | 0.011292 (rank : 103) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 761 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9C0K0, Q9H162 | Gene names | BCL11B, CTIP2, RIT1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma/leukemia 11B (B-cell CLL/lymphoma 11B) (Radiation- induced tumor suppressor gene 1 protein) (hRit1) (COUP-TF-interacting protein 2). | |||||
|
ENL_HUMAN
|
||||||
| θ value | 0.47712 (rank : 34) | NC score | 0.043306 (rank : 35) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q03111, Q14768 | Gene names | MLLT1, ENL, LTG19, YEATS1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein ENL (YEATS domain-containing protein 1). | |||||
|
EVL_MOUSE
|
||||||
| θ value | 0.47712 (rank : 35) | NC score | 0.039917 (rank : 39) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P70429, Q9ERU8 | Gene names | Evl | |||
|
Domain Architecture |
|
|||||
| Description | Ena/VASP-like protein (Ena/vasodilator-stimulated phosphoprotein- like). | |||||
|
NCOA6_MOUSE
|
||||||
| θ value | 0.47712 (rank : 36) | NC score | 0.077598 (rank : 12) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1076 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9JL19, Q9JLT9 | Gene names | Ncoa6, Aib3, Prip, Rap250, Trbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC) (Thyroid hormone receptor-binding protein). | |||||
|
PRPC_HUMAN
|
||||||
| θ value | 0.47712 (rank : 37) | NC score | 0.059034 (rank : 16) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P02810, Q4VBP2, Q53XA2, Q6P2F6 | Gene names | PRH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Salivary acidic proline-rich phosphoprotein 1/2 precursor (PRP-1/PRP- 2) (Parotid proline-rich protein 1/2) (Pr1/Pr2) (Protein C) (Parotid acidic protein) (Pa) (Parotid isoelectric focusing variant protein) (PIF-S) (Parotid double-band protein) (Db-s) [Contains: Salivary acidic proline-rich phosphoprotein 1/2; Salivary acidic proline-rich phosphoprotein 3/4 (PRP-3/PRP-4) (Protein A) (PIF-F) (Db-F); Peptide P-C]. | |||||
|
SRRM1_MOUSE
|
||||||
| θ value | 0.47712 (rank : 38) | NC score | 0.058525 (rank : 18) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
ZFHX2_HUMAN
|
||||||
| θ value | 0.47712 (rank : 39) | NC score | 0.020019 (rank : 84) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1178 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9C0A1 | Gene names | ZFHX2, KIAA1762 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger homeobox protein 2. | |||||
|
B3A3_MOUSE
|
||||||
| θ value | 0.62314 (rank : 40) | NC score | 0.027844 (rank : 63) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P16283 | Gene names | Slc4a3, Ae3 | |||
|
Domain Architecture |

|
|||||
| Description | Anion exchange protein 3 (Neuronal band 3-like protein) (Solute carrier family 4 member 3). | |||||
|
MAGI2_MOUSE
|
||||||
| θ value | 0.62314 (rank : 41) | NC score | 0.022609 (rank : 77) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 370 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9WVQ1, Q3UH81, Q6GT88, Q8BYT1, Q8CA85 | Gene names | Magi2, Acvrinp1, Aip1, Arip1 | |||
|
Domain Architecture |

|
|||||
| Description | Membrane-associated guanylate kinase, WW and PDZ domain-containing protein 2 (Membrane-associated guanylate kinase inverted 2) (MAGI-2) (Atrophin-1-interacting protein 1) (Activin receptor-interacting protein 1) (Acvrip1). | |||||
|
BC11A_HUMAN
|
||||||
| θ value | 0.813845 (rank : 42) | NC score | 0.010898 (rank : 105) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 745 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9H165, Q86W14, Q8WU92, Q96JL6, Q9H163, Q9H164, Q9H3G9, Q9NWA7 | Gene names | BCL11A, CTIP1, EVI9, KIAA1809 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma/leukemia 11A (B-cell CLL/lymphoma 11A) (COUP-TF- interacting protein 1) (Ecotropic viral integration site 9 protein) (EVI-9). | |||||
|
BC11A_MOUSE
|
||||||
| θ value | 0.813845 (rank : 43) | NC score | 0.013768 (rank : 96) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 732 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9QYE3, Q80T89, Q8BLC7, Q8BLR4, Q8BWX3, Q921V4, Q9D0V2, Q9JIT4, Q9JLK8, Q9JLK9 | Gene names | Bcl11a, Ctip1, Evi9, Kiaa1809 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma/leukemia 11A (B-cell CLL/lymphoma 11A) (COUP-TF- interacting protein 1) (Ecotropic viral integration site 9 protein) (EVI-9). | |||||
|
DNL1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 44) | NC score | 0.027909 (rank : 62) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P18858 | Gene names | LIG1 | |||
|
Domain Architecture |
|
|||||
| Description | DNA ligase 1 (EC 6.5.1.1) (DNA ligase I) (Polydeoxyribonucleotide synthase [ATP] 1). | |||||
|
MAP1A_HUMAN
|
||||||
| θ value | 0.813845 (rank : 45) | NC score | 0.028026 (rank : 60) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | P78559, O95643, Q12973, Q15882, Q9UJT4 | Gene names | MAP1A | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 1A (MAP 1A) (Proliferation-related protein p80) [Contains: MAP1 light chain LC2]. | |||||
|
SPEG_MOUSE
|
||||||
| θ value | 0.813845 (rank : 46) | NC score | 0.011659 (rank : 101) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1959 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q62407, Q3TPH8, Q6P5V1, Q80TF7, Q80ZN0, Q8BZF4, Q9EQJ5 | Gene names | Speg, Apeg1, Kiaa1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle-specific serine/threonine protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
SRRM1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 47) | NC score | 0.044785 (rank : 33) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
BSN_MOUSE
|
||||||
| θ value | 1.06291 (rank : 48) | NC score | 0.053346 (rank : 20) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
EVL_HUMAN
|
||||||
| θ value | 1.06291 (rank : 49) | NC score | 0.041227 (rank : 38) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9UI08, O95884, Q7Z522, Q8TBV1, Q9UF25, Q9UIC2 | Gene names | EVL, RNB6 | |||
|
Domain Architecture |

|
|||||
| Description | Ena/VASP-like protein (Ena/vasodilator-stimulated phosphoprotein- like). | |||||
|
GP73_MOUSE
|
||||||
| θ value | 1.06291 (rank : 50) | NC score | 0.022640 (rank : 76) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q91XA2 | Gene names | Golph2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi phosphoprotein 2 (Golgi membrane protein GP73). | |||||
|
MLL2_HUMAN
|
||||||
| θ value | 1.06291 (rank : 51) | NC score | 0.028305 (rank : 57) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
PRG4_HUMAN
|
||||||
| θ value | 1.06291 (rank : 52) | NC score | 0.031274 (rank : 50) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q92954, Q6DNC4, Q6DNC5, Q6ZMZ5, Q9BX49 | Gene names | PRG4, MSF, SZP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
SON_HUMAN
|
||||||
| θ value | 1.06291 (rank : 53) | NC score | 0.046940 (rank : 28) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
SYP2L_HUMAN
|
||||||
| θ value | 1.06291 (rank : 54) | NC score | 0.031276 (rank : 49) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 400 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9H987 | Gene names | SYNPO2L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptopodin 2-like protein. | |||||
|
WASF3_HUMAN
|
||||||
| θ value | 1.06291 (rank : 55) | NC score | 0.026866 (rank : 64) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9UPY6, O94974 | Gene names | WASF3, KIAA0900, SCAR3, WAVE3 | |||
|
Domain Architecture |

|
|||||
| Description | Wiskott-Aldrich syndrome protein family member 3 (WASP-family protein member 3) (Protein WAVE-3) (Verprolin homology domain-containing protein 3). | |||||
|
ZAR1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 56) | NC score | 0.043002 (rank : 36) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q86SH2 | Gene names | ZAR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zygote arrest 1 (Oocyte-specific maternal effect factor). | |||||
|
EP400_MOUSE
|
||||||
| θ value | 1.38821 (rank : 57) | NC score | 0.025964 (rank : 66) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 592 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q8CHI8, Q80TC8, Q8BXI5, Q8BYW3, Q8C0P6, Q8CHI7, Q8VDF4, Q9DA54 | Gene names | Ep400, Kiaa1498 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E1A-binding protein p400 (EC 3.6.1.-) (p400 kDa SWI2/SNF2-related protein) (Domino homolog) (mDomino). | |||||
|
MAGI2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 58) | NC score | 0.020975 (rank : 80) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 373 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q86UL8, O60434, O60510, Q86UI7, Q9NP44, Q9UDQ5, Q9UDU1 | Gene names | MAGI2, ACVRINP1, AIP1, KIAA0705 | |||
|
Domain Architecture |

|
|||||
| Description | Membrane-associated guanylate kinase, WW and PDZ domain-containing protein 2 (Membrane-associated guanylate kinase inverted 2) (MAGI-2) (Atrophin-1-interacting protein 1) (Atrophin-1-interacting protein A). | |||||
|
NCOR2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 59) | NC score | 0.031107 (rank : 51) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 870 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9Y618, O00613, O15416, Q13354, Q9Y5U0 | Gene names | NCOR2, CTG26 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC) (CTG repeat protein 26) (SMAP270). | |||||
|
REPS2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 60) | NC score | 0.029696 (rank : 56) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q80XA6 | Gene names | Reps2, Pob1 | |||
|
Domain Architecture |
|
|||||
| Description | RalBP1-associated Eps domain-containing protein 2 (RalBP1-interacting protein 2) (Partner of RalBP1). | |||||
|
SATB1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 61) | NC score | 0.033810 (rank : 48) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q01826 | Gene names | SATB1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-binding protein SATB1 (Special AT-rich sequence-binding protein 1). | |||||
|
SYNJ2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 62) | NC score | 0.018172 (rank : 89) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O15056, Q5TA13, Q5TA16, Q5TA19, Q86XK0, Q8IZA8, Q9H226 | Gene names | SYNJ2, KIAA0348 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptojanin-2 (EC 3.1.3.36) (Synaptic inositol-1,4,5-trisphosphate 5- phosphatase 2). | |||||
|
AAK1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 63) | NC score | 0.009425 (rank : 113) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1077 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q2M2I8, Q4ZFZ3, Q53RX6, Q9UPV4 | Gene names | AAK1, KIAA1048 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AP2-associated protein kinase 1 (EC 2.7.11.1) (Adaptor-associated kinase 1). | |||||
|
CG020_HUMAN
|
||||||
| θ value | 1.81305 (rank : 64) | NC score | 0.026516 (rank : 65) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 4 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7L5D6, Q9UFC9, Q9Y309 | Gene names | C7orf20 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UPF0363 protein C7orf20. | |||||
|
IWS1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 65) | NC score | 0.035531 (rank : 45) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
K1949_MOUSE
|
||||||
| θ value | 1.81305 (rank : 66) | NC score | 0.028014 (rank : 61) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8BQ30, Q3UDS6, Q8CEY9, Q8R3D8 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1949 homolog. | |||||
|
PYGO2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 67) | NC score | 0.038234 (rank : 41) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9BRQ0, Q8WYZ4, Q96CY2 | Gene names | PYGO2 | |||
|
Domain Architecture |

|
|||||
| Description | Pygopus homolog 2. | |||||
|
RAPH1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 68) | NC score | 0.030428 (rank : 55) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 593 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q70E73, Q96Q37, Q9C0I2 | Gene names | RAPH1, ALS2CR9, KIAA1681, LPD, PREL2, RMO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras-associated and pleckstrin homology domains-containing protein 1 (RAPH1) (Lamellipodin) (Proline-rich EVH1 ligand 2) (PREL-2) (Protein RMO1) (Amyotrophic lateral sclerosis 2 chromosomal region candidate 9 gene protein). | |||||
|
TARA_MOUSE
|
||||||
| θ value | 1.81305 (rank : 69) | NC score | 0.028088 (rank : 59) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1133 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q99KW3, Q2PZW9, Q2Q402, Q2Q403, Q2Q404, Q6ZPK4, Q8C6T3, Q8CG90 | Gene names | Triobp, Kiaa1662, Tara | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
TROAP_HUMAN
|
||||||
| θ value | 1.81305 (rank : 70) | NC score | 0.044868 (rank : 32) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q12815 | Gene names | TROAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trophinin-associated protein (Tastin) (Trophinin-assisting protein). | |||||
|
ZCHC5_HUMAN
|
||||||
| θ value | 1.81305 (rank : 71) | NC score | 0.034796 (rank : 46) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8N8U3 | Gene names | ZCCHC5 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger CCHC domain-containing protein 5. | |||||
|
CD5_MOUSE
|
||||||
| θ value | 2.36792 (rank : 72) | NC score | 0.019141 (rank : 86) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P13379 | Gene names | Cd5, Ly-1 | |||
|
Domain Architecture |
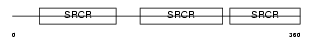
|
|||||
| Description | T-cell surface glycoprotein CD5 precursor (Lymphocyte antigen Ly-1) (Lyt-1). | |||||
|
FYB_MOUSE
|
||||||
| θ value | 2.36792 (rank : 73) | NC score | 0.024418 (rank : 70) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O35601, Q9Z2H3 | Gene names | Fyb | |||
|
Domain Architecture |

|
|||||
| Description | FYN-binding protein (FYN-T-binding protein) (FYB-120/130) (p120/p130) (SLP-76-associated phosphoprotein) (SLAP-130). | |||||
|
STAT6_MOUSE
|
||||||
| θ value | 2.36792 (rank : 74) | NC score | 0.013759 (rank : 97) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P52633 | Gene names | Stat6 | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and transcription activator 6. | |||||
|
WASIP_MOUSE
|
||||||
| θ value | 2.36792 (rank : 75) | NC score | 0.037793 (rank : 42) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8K1I7, Q3U0U8 | Gene names | Waspip, Wip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein). | |||||
|
ADA19_MOUSE
|
||||||
| θ value | 3.0926 (rank : 76) | NC score | 0.005384 (rank : 121) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O35674 | Gene names | Adam19, Mltnb | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 19 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 19) (Meltrin beta). | |||||
|
CHD3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 77) | NC score | 0.010786 (rank : 107) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q12873, Q9Y4I0 | Gene names | CHD3 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain helicase-DNA-binding protein 3 (EC 3.6.1.-) (ATP- dependent helicase CHD3) (CHD-3) (Mi-2 autoantigen 240 kDa protein) (Mi2-alpha) (Zinc-finger helicase) (hZFH). | |||||
|
CJ026_MOUSE
|
||||||
| θ value | 3.0926 (rank : 78) | NC score | 0.021895 (rank : 79) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8BGW2, Q3TF48, Q3U3M2, Q6PD43, Q8R0W8 | Gene names | D19Wsu162e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C10orf26 homolog. | |||||
|
CSKI1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 79) | NC score | 0.018148 (rank : 90) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 997 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8WXD9, Q9P2P0 | Gene names | CASKIN1, KIAA1306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Caskin-1 (CASK-interacting protein 1). | |||||
|
DGKK_HUMAN
|
||||||
| θ value | 3.0926 (rank : 80) | NC score | 0.020655 (rank : 82) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 378 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q5KSL6 | Gene names | DGKK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Diacylglycerol kinase kappa (EC 2.7.1.107) (Diglyceride kinase kappa) (DGK-kappa) (DAG kinase kappa) (142 kDa diacylglycerol kinase). | |||||
|
GSCR1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 81) | NC score | 0.033814 (rank : 47) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 674 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9NZM4 | Gene names | GLTSCR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glioma tumor suppressor candidate region gene 1 protein. | |||||
|
MOR2A_MOUSE
|
||||||
| θ value | 3.0926 (rank : 82) | NC score | 0.016730 (rank : 93) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 220 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q69ZX6, Q5QNQ7, Q6P547 | Gene names | Morc2a, Kiaa0852, Zcwcc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORC family CW-type zinc finger protein 2A (Zinc finger CW-type coiled-coil domain protein 1). | |||||
|
OXR1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 83) | NC score | 0.017469 (rank : 92) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 516 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q4KMM3, Q5FWW1, Q99L06, Q99MK1, Q99MP4 | Gene names | Oxr1, C7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Oxidation resistance protein 1 (Protein C7). | |||||
|
BRD4_MOUSE
|
||||||
| θ value | 4.03905 (rank : 84) | NC score | 0.019856 (rank : 85) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 845 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9ESU6, Q8VHF7, Q8VHF8 | Gene names | Brd4, Mcap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 4 (Mitotic chromosome-associated protein) (MCAP). | |||||
|
CSKI1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 85) | NC score | 0.020899 (rank : 81) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1031 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q6P9K8, Q6ZPU2, Q8BWU2, Q8BX99, Q9CXH0 | Gene names | Caskin1, Kiaa1306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Caskin-1 (CASK-interacting protein 1). | |||||
|
ENAH_MOUSE
|
||||||
| θ value | 4.03905 (rank : 86) | NC score | 0.030782 (rank : 52) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 919 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q03173, P70430, P70431, P70432, P70433, Q5D053 | Gene names | Enah, Mena, Ndpp1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein enabled homolog (NPC-derived proline-rich protein 1) (NDPP-1). | |||||
|
ES8L3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 87) | NC score | 0.011604 (rank : 102) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8TE67, Q5T8Q6, Q5T8Q7, Q5T8Q8, Q96E47, Q9H719 | Gene names | EPS8L3, EPS8R3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Epidermal growth factor receptor kinase substrate 8-like protein 3 (Epidermal growth factor receptor pathway substrate 8-related protein 3) (EPS8-like protein 3). | |||||
|
GPC2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 88) | NC score | 0.009541 (rank : 110) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8N158 | Gene names | GPC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glypican-2 precursor. | |||||
|
MDC1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 89) | NC score | 0.057241 (rank : 19) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
PO6F1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 90) | NC score | 0.006716 (rank : 120) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 370 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q14863 | Gene names | POU6F1, BRN5, MPOU, TCFB1 | |||
|
Domain Architecture |
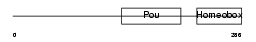
|
|||||
| Description | POU domain, class 6, transcription factor 1 (mPOU homeobox protein) (Brain-specific homeobox/POU domain protein 5) (Brain-5) (Brn-5 protein). | |||||
|
REPS2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 91) | NC score | 0.024681 (rank : 69) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8NFH8, O43428, Q8NFI5 | Gene names | REPS2, POB1 | |||
|
Domain Architecture |

|
|||||
| Description | RalBP1-associated Eps domain-containing protein 2 (RalBP1-interacting protein 2) (Partner of RalBP1). | |||||
|
SETBP_MOUSE
|
||||||
| θ value | 4.03905 (rank : 92) | NC score | 0.028127 (rank : 58) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9Z180, Q66JL8 | Gene names | Setbp1, Kiaa0437 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SET-binding protein (SEB). | |||||
|
ZN646_HUMAN
|
||||||
| θ value | 4.03905 (rank : 93) | NC score | 0.000404 (rank : 124) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 813 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O15015 | Gene names | ZNF646, KIAA0296 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 646. | |||||
|
BC11B_MOUSE
|
||||||
| θ value | 5.27518 (rank : 94) | NC score | 0.009530 (rank : 111) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 767 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q99PV8, Q8C2I1, Q99PV6, Q99PV7, Q9JLF8 | Gene names | Bcl11b, Ctip2, Rit1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma/leukemia 11B (B-cell CLL/lymphoma 11B) (Radiation- induced tumor suppressor gene 1 protein) (mRit1) (COUP-TF-interacting protein 2). | |||||
|
CEBPA_MOUSE
|
||||||
| θ value | 5.27518 (rank : 95) | NC score | 0.011194 (rank : 104) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P53566, Q91XB6 | Gene names | Cebpa | |||
|
Domain Architecture |

|
|||||
| Description | CCAAT/enhancer-binding protein alpha (C/EBP alpha). | |||||
|
PHC1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 96) | NC score | 0.018955 (rank : 87) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P78364, Q8WVM3, Q9BU63 | Gene names | PHC1, EDR1, PH1 | |||
|
Domain Architecture |

|
|||||
| Description | Polyhomeotic-like protein 1 (hPH1) (Early development regulatory protein 1). | |||||
|
PPRB_MOUSE
|
||||||
| θ value | 5.27518 (rank : 97) | NC score | 0.018887 (rank : 88) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q925J9, O88323, Q3UHV0, Q6AXD5, Q8BW37, Q8BX19, Q8VDQ7, Q925K0 | Gene names | Pparbp, Trip2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor-binding protein (PBP) (PPAR-binding protein) (Thyroid hormone receptor-associated protein complex 220 kDa component) (Trap220) (Thyroid receptor-interacting protein 2) (TRIP-2). | |||||
|
RERE_HUMAN
|
||||||
| θ value | 5.27518 (rank : 98) | NC score | 0.023620 (rank : 74) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 779 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9P2R6, O43393, O75046, O75359, Q5VXL9, Q6P6B9, Q9Y2W4 | Gene names | RERE, ARG, ARP, ATN1L, KIAA0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-1-like protein) (Atrophin-1-related protein). | |||||
|
SFPQ_HUMAN
|
||||||
| θ value | 5.27518 (rank : 99) | NC score | 0.030598 (rank : 53) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 986 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P23246, P30808 | Gene names | SFPQ, PSF | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, proline- and glutamine-rich (Polypyrimidine tract- binding protein-associated-splicing factor) (PTB-associated-splicing factor) (PSF) (DNA-binding p52/p100 complex, 100 kDa subunit) (100 kDa DNA-pairing protein) (hPOMp100). | |||||
|
SYP2L_MOUSE
|
||||||
| θ value | 5.27518 (rank : 100) | NC score | 0.025682 (rank : 67) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8BWB1 | Gene names | Synpo2l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptopodin 2-like protein. | |||||
|
ZN579_HUMAN
|
||||||
| θ value | 5.27518 (rank : 101) | NC score | 0.000322 (rank : 125) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 726 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8NAF0 | Gene names | ZNF579 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 579. | |||||
|
ADDA_HUMAN
|
||||||
| θ value | 6.88961 (rank : 102) | NC score | 0.025537 (rank : 68) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P35611, Q13734, Q14729, Q16156, Q9UJB6 | Gene names | ADD1, ADDA | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-adducin (Erythrocyte adducin subunit alpha). | |||||
|
ALEX_MOUSE
|
||||||
| θ value | 6.88961 (rank : 103) | NC score | 0.023191 (rank : 75) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q6R0H6, Q8BIR3 | Gene names | Gnas, Gnas1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein ALEX (Alternative gene product encoded by XL-exon). | |||||
|
GLU2B_MOUSE
|
||||||
| θ value | 6.88961 (rank : 104) | NC score | 0.013713 (rank : 98) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O08795, Q921X2 | Gene names | Prkcsh | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glucosidase 2 subunit beta precursor (Glucosidase II subunit beta) (Protein kinase C substrate, 60.1 kDa protein, heavy chain) (PKCSH) (80K-H protein). | |||||
|
IWS1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 105) | NC score | 0.036239 (rank : 44) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
KCNN3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 106) | NC score | 0.006938 (rank : 119) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P58391 | Gene names | Kcnn3, Sk3 | |||
|
Domain Architecture |
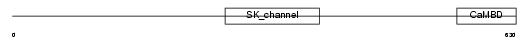
|
|||||
| Description | Small conductance calcium-activated potassium channel protein 3 (SK3). | |||||
|
KI21B_HUMAN
|
||||||
| θ value | 6.88961 (rank : 107) | NC score | 0.001578 (rank : 122) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1017 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O75037, Q5T4J3 | Gene names | KIF21B, KIAA0449 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin family member 21B. | |||||
|
PHF2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 108) | NC score | 0.008093 (rank : 117) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9WTU0, Q80WA8 | Gene names | Phf2 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 2 (GRC5). | |||||
|
PRP4B_HUMAN
|
||||||
| θ value | 6.88961 (rank : 109) | NC score | 0.001482 (rank : 123) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1030 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q13523, Q8TDP2, Q96QT7, Q9UEE6 | Gene names | PRPF4B, KIAA0536, PRP4, PRP4H, PRP4K | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase PRP4 homolog (EC 2.7.11.1) (PRP4 pre- mRNA-processing factor 4 homolog) (PRP4 kinase). | |||||
|
RUSC1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 110) | NC score | 0.009502 (rank : 112) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 226 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9BVN2, Q9UPY4, Q9Y4T5 | Gene names | RUSC1, NESCA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RUN and SH3 domain-containing protein 1 (New molecule containing SH3 at the carboxy-terminus) (Nesca). | |||||
|
RYR1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 111) | NC score | 0.008529 (rank : 114) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P21817, Q16314, Q16368, Q9NPK1, Q9P1U4 | Gene names | RYR1, RYDR | |||
|
Domain Architecture |

|
|||||
| Description | Ryanodine receptor 1 (Skeletal muscle-type ryanodine receptor) (RyR1) (RYR-1) (Skeletal muscle calcium release channel). | |||||
|
SYN2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 112) | NC score | 0.015111 (rank : 95) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q92777 | Gene names | SYN2 | |||
|
Domain Architecture |
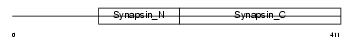
|
|||||
| Description | Synapsin-2 (Synapsin II). | |||||
|
TNNT1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 113) | NC score | 0.009556 (rank : 109) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P13805, O95472, Q16061 | Gene names | TNNT1, TNT | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Troponin T, slow skeletal muscle (TnTs) (Slow skeletal muscle troponin T) (sTnT). | |||||
|
EP400_HUMAN
|
||||||
| θ value | 8.99809 (rank : 114) | NC score | 0.020575 (rank : 83) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 556 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q96L91, O15411, Q6P2F5, Q8N8Q7, Q8NE05, Q96JK7, Q9P230 | Gene names | EP400, CAGH32, KIAA1498, KIAA1818, TNRC12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E1A-binding protein p400 (EC 3.6.1.-) (p400 kDa SWI2/SNF2-related protein) (Domino homolog) (hDomino) (CAG repeat protein 32) (Trinucleotide repeat-containing gene 12 protein). | |||||
|
FBX46_MOUSE
|
||||||
| θ value | 8.99809 (rank : 115) | NC score | 0.015142 (rank : 94) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8BG80 | Gene names | Fbxo46, Fbx46, Fbxo34l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 46 (F-box only protein 34-like). | |||||
|
K0174_MOUSE
|
||||||
| θ value | 8.99809 (rank : 116) | NC score | 0.011918 (rank : 100) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9CX00, Q80U68 | Gene names | Kiaa0174 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0174. | |||||
|
LBXCO_HUMAN
|
||||||
| θ value | 8.99809 (rank : 117) | NC score | 0.007847 (rank : 118) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P84550 | Gene names | LBXCOR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ladybird homeobox corepressor 1. | |||||
|
LR37B_HUMAN
|
||||||
| θ value | 8.99809 (rank : 118) | NC score | 0.008415 (rank : 115) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q96QE4, Q5YKG6 | Gene names | LRRC37B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 37B precursor (C66 SLIT-like testicular protein). | |||||
|
RGS3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 119) | NC score | 0.044293 (rank : 34) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
S4A7_HUMAN
|
||||||
| θ value | 8.99809 (rank : 120) | NC score | 0.017623 (rank : 91) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Y6M7, O60350, Q6AHZ9, Q9HC88, Q9UIB9 | Gene names | SLC4A7, BT, NBC2, NBC2B, NBC3, SBC2, SLC4A6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium bicarbonate cotransporter 3 (Sodium bicarbonate cotransporter 2) (Sodium bicarbonate cotransporter 2b) (Bicarbonate transporter) (Solute carrier family 4 member 7). | |||||
|
SBK1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 121) | NC score | -0.000537 (rank : 126) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 824 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8QZX0 | Gene names | Sbk1, Sbk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SBK1 (EC 2.7.11.1) (SH3-binding kinase 1). | |||||
|
SEPT9_HUMAN
|
||||||
| θ value | 8.99809 (rank : 122) | NC score | 0.024349 (rank : 71) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 706 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q9UHD8, Q96QF3, Q96QF4, Q96QF5, Q9HA04, Q9UG40, Q9Y5W4 | Gene names | SEPT9, KIAA0991, MSF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Septin-9 (MLL septin-like fusion protein) (MLL septin-like fusion protein MSF-A) (Ovarian/Breast septin) (Ov/Br septin) (Septin D1). | |||||
|
USH1C_MOUSE
|
||||||
| θ value | 8.99809 (rank : 123) | NC score | 0.010873 (rank : 106) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 557 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9ES64, Q91XD1, Q9CVG7, Q9ES65 | Gene names | Ush1c | |||
|
Domain Architecture |

|
|||||
| Description | Harmonin (Usher syndrome type-1C protein homolog) (PDZ domain- containing protein). | |||||
|
YBOX2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 124) | NC score | 0.008227 (rank : 116) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Z2C8, Q5NCW8, Q5NCW9, Q9Z2C7 | Gene names | Ybx2, Msy2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Y-box-binding protein 2 (Germ cell-specific Y-box-binding protein) (FRGY2 homolog). | |||||
|
ZN207_HUMAN
|
||||||
| θ value | 8.99809 (rank : 125) | NC score | 0.023983 (rank : 73) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 326 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O43670, Q96HW5, Q9BUQ7 | Gene names | ZNF207 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 207. | |||||
|
PRRT3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 126) | NC score | 0.050922 (rank : 25) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q5FWE3, Q49AD0, Q6UXY6, Q8NBC9 | Gene names | PRRT3, UNQ5823/PRO19642 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich transmembrane protein 3. | |||||
|
MA1B1_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 125 | |
| SwissProt Accessions | Q9UKM7, Q5VSG3, Q9BRS9, Q9Y5K7 | Gene names | MAN1B1 | |||
|
Domain Architecture |

|
|||||
| Description | Endoplasmic reticulum mannosyl-oligosaccharide 1,2-alpha-mannosidase (EC 3.2.1.113) (ER alpha-1,2-mannosidase) (Mannosidase alpha class 1B member 1) (Man9GlcNAc2-specific-processing alpha-mannosidase). | |||||
|
MA1C1_HUMAN
|
||||||
| NC score | 0.934773 (rank : 2) | θ value | 1.40038e-85 (rank : 6) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9NR34, Q9Y545 | Gene names | MAN1C1, MAN1A3, MAN1C | |||
|
Domain Architecture |

|
|||||
| Description | Mannosyl-oligosaccharide 1,2-alpha-mannosidase IC (EC 3.2.1.113) (Processing alpha-1,2-mannosidase IC) (Alpha-1,2-mannosidase IC) (Mannosidase alpha class 1C member 1) (HMIC). | |||||
|
MA1A1_HUMAN
|
||||||
| NC score | 0.931227 (rank : 3) | θ value | 6.07444e-89 (rank : 3) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P33908, Q9NU44, Q9UJI3 | Gene names | MAN1A1 | |||
|
Domain Architecture |

|
|||||
| Description | Mannosyl-oligosaccharide 1,2-alpha-mannosidase IA (EC 3.2.1.113) (Processing alpha-1,2-mannosidase IA) (Alpha-1,2-mannosidase IA) (Mannosidase alpha class 1A member 1) (Man(9)-alpha-mannosidase) (Man9-mannosidase). | |||||
|
MA1A1_MOUSE
|
||||||
| NC score | 0.931012 (rank : 4) | θ value | 1.39714e-93 (rank : 2) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P45700 | Gene names | Man1a1, Man1a | |||
|
Domain Architecture |

|
|||||
| Description | Mannosyl-oligosaccharide 1,2-alpha-mannosidase IA (EC 3.2.1.113) (Processing alpha-1,2-mannosidase IA) (Alpha-1,2-mannosidase IA) (Mannosidase alpha class 1A member 1) (Man(9)-alpha-mannosidase) (Man9-mannosidase). | |||||
|
MA1A2_HUMAN
|
||||||
| NC score | 0.930330 (rank : 5) | θ value | 3.93732e-88 (rank : 5) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O60476, Q9H510 | Gene names | MAN1A2, MAN1B | |||
|
Domain Architecture |

|
|||||
| Description | Mannosyl-oligosaccharide 1,2-alpha-mannosidase IB (EC 3.2.1.113) (Processing alpha-1,2-mannosidase IB) (Alpha-1,2-mannosidase IB) (Mannosidase alpha class 1A member 2). | |||||
|
MA1A2_MOUSE
|
||||||
| NC score | 0.930026 (rank : 6) | θ value | 2.30828e-88 (rank : 4) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P39098, Q5SUY6, Q5SUY7, Q60599, Q9CUY9 | Gene names | Man1a2, Man1b | |||
|
Domain Architecture |

|
|||||
| Description | Mannosyl-oligosaccharide 1,2-alpha-mannosidase IB (EC 3.2.1.113) (Processing alpha-1,2-mannosidase IB) (Alpha-1,2-mannosidase IB) (Mannosidase alpha class 1A member 2). | |||||
|
EDEM2_HUMAN
|
||||||
| NC score | 0.884530 (rank : 7) | θ value | 5.03882e-43 (rank : 10) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9BV94, Q6IA89, Q6UWZ4, Q9H4U0, Q9H886, Q9NTL9, Q9NVE6 | Gene names | EDEM2, C20orf31, C20orf49 | |||
|
Domain Architecture |

|
|||||
| Description | ER degradation-enhancing alpha-mannosidase-like 2 precursor. | |||||
|
EDEM1_HUMAN
|
||||||
| NC score | 0.861070 (rank : 8) | θ value | 1.51363e-47 (rank : 9) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q92611 | Gene names | EDEM1, EDEM, KIAA0212 | |||
|
Domain Architecture |

|
|||||
| Description | ER degradation-enhancing alpha-mannosidase-like 1. | |||||
|
EDEM1_MOUSE
|
||||||
| NC score | 0.860645 (rank : 9) | θ value | 5.20228e-48 (rank : 8) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q925U4 | Gene names | Edem1, Edem | |||
|
Domain Architecture |

|
|||||
| Description | ER degradation-enhancing alpha-mannosidase-like 1. | |||||
|
EDEM3_HUMAN
|
||||||
| NC score | 0.850558 (rank : 10) | θ value | 2.25659e-51 (rank : 7) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9BZQ6, Q5TEZ0, Q9HCW1, Q9UFV7 | Gene names | EDEM3, C1orf22 | |||
|
Domain Architecture |

|
|||||
| Description | ER degradation-enhancing alpha-mannosidase-like 3. | |||||
|
NCOA6_HUMAN
|
||||||
| NC score | 0.087132 (rank : 11) | θ value | 0.00509761 (rank : 12) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1202 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q14686, Q9NTZ9, Q9UH74, Q9UK86 | Gene names | NCOA6, AIB3, KIAA0181, RAP250, TRBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (PRIP) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC RAP250) (Thyroid hormone receptor-binding protein). | |||||
|
NCOA6_MOUSE
|
||||||
| NC score | 0.077598 (rank : 12) | θ value | 0.47712 (rank : 36) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1076 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9JL19, Q9JLT9 | Gene names | Ncoa6, Aib3, Prip, Rap250, Trbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC) (Thyroid hormone receptor-binding protein). | |||||
|
MNT_MOUSE
|
||||||
| NC score | 0.076802 (rank : 13) | θ value | 0.00869519 (rank : 13) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 432 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O08789, P97349 | Gene names | Mnt, Rox | |||
|
Domain Architecture |

|
|||||
| Description | Max-binding protein MNT (Protein ROX) (Myc antagonist MNT). | |||||
|
MNT_HUMAN
|
||||||
| NC score | 0.070932 (rank : 14) | θ value | 0.0736092 (rank : 22) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 496 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q99583 | Gene names | MNT, ROX | |||
|
Domain Architecture |

|
|||||
| Description | Max-binding protein MNT (Protein ROX) (Myc antagonist MNT). | |||||
|
MADCA_HUMAN
|
||||||
| NC score | 0.064186 (rank : 15) | θ value | 0.0563607 (rank : 21) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q13477, O60222, O75867 | Gene names | MADCAM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucosal addressin cell adhesion molecule 1 precursor (MAdCAM-1) (hMAdCAM-1). | |||||
|
PRPC_HUMAN
|
||||||
| NC score | 0.059034 (rank : 16) | θ value | 0.47712 (rank : 37) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P02810, Q4VBP2, Q53XA2, Q6P2F6 | Gene names | PRH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Salivary acidic proline-rich phosphoprotein 1/2 precursor (PRP-1/PRP- 2) (Parotid proline-rich protein 1/2) (Pr1/Pr2) (Protein C) (Parotid acidic protein) (Pa) (Parotid isoelectric focusing variant protein) (PIF-S) (Parotid double-band protein) (Db-s) [Contains: Salivary acidic proline-rich phosphoprotein 1/2; Salivary acidic proline-rich phosphoprotein 3/4 (PRP-3/PRP-4) (Protein A) (PIF-F) (Db-F); Peptide P-C]. | |||||
|
PRP1_HUMAN
|
||||||
| NC score | 0.059001 (rank : 17) | θ value | 0.00869519 (rank : 14) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P04280, Q08805, Q15186, Q15187, Q15214, Q15215, Q16038 | Gene names | PRB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 1 precursor (Salivary proline-rich protein) [Contains: Basic peptide IB-6; Peptide P-H]. | |||||
|
SRRM1_MOUSE
|
||||||
| NC score | 0.058525 (rank : 18) | θ value | 0.47712 (rank : 38) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
MDC1_MOUSE
|
||||||
| NC score | 0.057241 (rank : 19) | θ value | 4.03905 (rank : 89) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
BSN_MOUSE
|
||||||
| NC score | 0.053346 (rank : 20) | θ value | 1.06291 (rank : 48) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
BAT2_HUMAN
|
||||||
| NC score | 0.053305 (rank : 21) | θ value | 0.365318 (rank : 30) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | P48634, O95875, Q5SQ29, Q5SQ30, Q5ST84, Q5STX6, Q5STX7, Q68DW9, Q6P9P7, Q6PIN1, Q96QC6 | Gene names | BAT2, G2 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
PRB3_HUMAN
|
||||||
| NC score | 0.053287 (rank : 22) | θ value | 0.0148317 (rank : 16) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q04118, Q15188, Q7M4M9 | Gene names | PRB3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 3 precursor (Parotid salivary glycoprotein G1) (Proline-rich protein G1). | |||||
|
PRB4S_HUMAN
|
||||||
| NC score | 0.051763 (rank : 23) | θ value | 0.0330416 (rank : 19) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P10163, P02813 | Gene names | PRB4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 4 allele S precursor (Salivary proline-rich protein Po) (Parotid o protein) [Contains: Protein N1; Glycosylated protein A]. | |||||
|
MILK1_MOUSE
|
||||||
| NC score | 0.051284 (rank : 24) | θ value | 0.0330416 (rank : 18) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 788 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q8BGT6, Q8BJ60 | Gene names | Mirab13, D15Mit260, Kiaa1668 | |||
|
Domain Architecture |

|
|||||
| Description | Molecule interacting with Rab13 (MIRab13) (MICAL-like protein 1). | |||||
|
PRRT3_HUMAN
|
||||||
| NC score | 0.050922 (rank : 25) | θ value | θ > 10 (rank : 126) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q5FWE3, Q49AD0, Q6UXY6, Q8NBC9 | Gene names | PRRT3, UNQ5823/PRO19642 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich transmembrane protein 3. | |||||
|
PRP2_MOUSE
|
||||||
| NC score | 0.048883 (rank : 26) | θ value | 0.0736092 (rank : 23) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P05142 | Gene names | Prh1, Prp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein MP-2 precursor. | |||||
|
CCNK_MOUSE
|
||||||
| NC score | 0.047301 (rank : 27) | θ value | 0.0330416 (rank : 17) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O88874, Q8R068 | Gene names | Ccnk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-K. | |||||
|
SON_HUMAN
|
||||||
| NC score | 0.046940 (rank : 28) | θ value | 1.06291 (rank : 53) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
SFPQ_MOUSE
|
||||||
| NC score | 0.046285 (rank : 29) | θ value | 0.00228821 (rank : 11) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8VIJ6, Q9ERW2 | Gene names | Sfpq | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, proline- and glutamine-rich (Polypyrimidine tract- binding protein-associated-splicing factor) (PTB-associated-splicing factor) (PSF) (DNA-binding p52/p100 complex, 100 kDa subunit). | |||||
|
BSN_HUMAN
|
||||||
| NC score | 0.045568 (rank : 30) | θ value | 0.21417 (rank : 27) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
SGIP1_HUMAN
|
||||||
| NC score | 0.045304 (rank : 31) | θ value | 0.365318 (rank : 32) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9BQI5, Q4LE32, Q5VYE2, Q5VYE3, Q5VYE4, Q68D76, Q6MZY6, Q8IWC2 | Gene names | SGIP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH3-containing GRB2-like protein 3-interacting protein 1 (Endophilin- 3-interacting protein). | |||||
|
TROAP_HUMAN
|
||||||
| NC score | 0.044868 (rank : 32) | θ value | 1.81305 (rank : 70) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q12815 | Gene names | TROAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trophinin-associated protein (Tastin) (Trophinin-assisting protein). | |||||
|
SRRM1_HUMAN
|
||||||
| NC score | 0.044785 (rank : 33) | θ value | 0.813845 (rank : 47) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
RGS3_MOUSE
|
||||||
| NC score | 0.044293 (rank : 34) | θ value | 8.99809 (rank : 119) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
ENL_HUMAN
|
||||||
| NC score | 0.043306 (rank : 35) | θ value | 0.47712 (rank : 34) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q03111, Q14768 | Gene names | MLLT1, ENL, LTG19, YEATS1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein ENL (YEATS domain-containing protein 1). | |||||
|
ZAR1_HUMAN
|
||||||
| NC score | 0.043002 (rank : 36) | θ value | 1.06291 (rank : 56) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q86SH2 | Gene names | ZAR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zygote arrest 1 (Oocyte-specific maternal effect factor). | |||||
|
CBP_HUMAN
|
||||||
| NC score | 0.042854 (rank : 37) | θ value | 0.279714 (rank : 28) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 941 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q92793, O00147, Q16376 | Gene names | CREBBP, CBP | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
EVL_HUMAN
|
||||||
| NC score | 0.041227 (rank : 38) | θ value | 1.06291 (rank : 49) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9UI08, O95884, Q7Z522, Q8TBV1, Q9UF25, Q9UIC2 | Gene names | EVL, RNB6 | |||
|
Domain Architecture |

|
|||||
| Description | Ena/VASP-like protein (Ena/vasodilator-stimulated phosphoprotein- like). | |||||
|
EVL_MOUSE
|
||||||
| NC score | 0.039917 (rank : 39) | θ value | 0.47712 (rank : 35) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P70429, Q9ERU8 | Gene names | Evl | |||
|
Domain Architecture |

|
|||||
| Description | Ena/VASP-like protein (Ena/vasodilator-stimulated phosphoprotein- like). | |||||
|
ADDA_MOUSE
|
||||||
| NC score | 0.038988 (rank : 40) | θ value | 0.0563607 (rank : 20) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9QYC0, Q9JLE3 | Gene names | Add1 | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-adducin (Erythrocyte adducin subunit alpha). | |||||
|
PYGO2_HUMAN
|
||||||
| NC score | 0.038234 (rank : 41) | θ value | 1.81305 (rank : 67) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 189 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9BRQ0, Q8WYZ4, Q96CY2 | Gene names | PYGO2 | |||
|
Domain Architecture |

|
|||||
| Description | Pygopus homolog 2. | |||||
|
WASIP_MOUSE
|
||||||
| NC score | 0.037793 (rank : 42) | θ value | 2.36792 (rank : 75) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q8K1I7, Q3U0U8 | Gene names | Waspip, Wip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein). | |||||
|
SATB1_MOUSE
|
||||||
| NC score | 0.037711 (rank : 43) | θ value | 0.365318 (rank : 31) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q60611 | Gene names | Satb1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-binding protein SATB1 (Special AT-rich sequence-binding protein 1). | |||||
|
IWS1_HUMAN
|
||||||
| NC score | 0.036239 (rank : 44) | θ value | 6.88961 (rank : 105) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
IWS1_MOUSE
|
||||||
| NC score | 0.035531 (rank : 45) | θ value | 1.81305 (rank : 65) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
ZCHC5_HUMAN
|
||||||
| NC score | 0.034796 (rank : 46) | θ value | 1.81305 (rank : 71) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8N8U3 | Gene names | ZCCHC5 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger CCHC domain-containing protein 5. | |||||
|
GSCR1_HUMAN
|
||||||
| NC score | 0.033814 (rank : 47) | θ value | 3.0926 (rank : 81) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 674 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9NZM4 | Gene names | GLTSCR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glioma tumor suppressor candidate region gene 1 protein. | |||||
|
SATB1_HUMAN
|
||||||
| NC score | 0.033810 (rank : 48) | θ value | 1.38821 (rank : 61) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q01826 | Gene names | SATB1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-binding protein SATB1 (Special AT-rich sequence-binding protein 1). | |||||
|
SYP2L_HUMAN
|
||||||
| NC score | 0.031276 (rank : 49) | θ value | 1.06291 (rank : 54) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 400 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9H987 | Gene names | SYNPO2L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptopodin 2-like protein. | |||||
|
PRG4_HUMAN
|
||||||
| NC score | 0.031274 (rank : 50) | θ value | 1.06291 (rank : 52) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q92954, Q6DNC4, Q6DNC5, Q6ZMZ5, Q9BX49 | Gene names | PRG4, MSF, SZP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
NCOR2_HUMAN
|
||||||
| NC score | 0.031107 (rank : 51) | θ value | 1.38821 (rank : 59) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 870 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9Y618, O00613, O15416, Q13354, Q9Y5U0 | Gene names | NCOR2, CTG26 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC) (CTG repeat protein 26) (SMAP270). | |||||
|
ENAH_MOUSE
|
||||||
| NC score | 0.030782 (rank : 52) | θ value | 4.03905 (rank : 86) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 919 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q03173, P70430, P70431, P70432, P70433, Q5D053 | Gene names | Enah, Mena, Ndpp1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein enabled homolog (NPC-derived proline-rich protein 1) (NDPP-1). | |||||
|
SFPQ_HUMAN
|
||||||
| NC score | 0.030598 (rank : 53) | θ value | 5.27518 (rank : 99) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 986 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P23246, P30808 | Gene names | SFPQ, PSF | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, proline- and glutamine-rich (Polypyrimidine tract- binding protein-associated-splicing factor) (PTB-associated-splicing factor) (PSF) (DNA-binding p52/p100 complex, 100 kDa subunit) (100 kDa DNA-pairing protein) (hPOMp100). | |||||
|
B3A3_HUMAN
|
||||||
| NC score | 0.030534 (rank : 54) | θ value | 0.0961366 (rank : 25) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P48751 | Gene names | SLC4A3, AE3 | |||
|
Domain Architecture |

|
|||||
| Description | Anion exchange protein 3 (Neuronal band 3-like protein) (Solute carrier family 4 member 3) (Cardiac/brain band 3-like protein) (CAE3/BAE3). | |||||
|
RAPH1_HUMAN
|
||||||
| NC score | 0.030428 (rank : 55) | θ value | 1.81305 (rank : 68) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 593 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q70E73, Q96Q37, Q9C0I2 | Gene names | RAPH1, ALS2CR9, KIAA1681, LPD, PREL2, RMO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras-associated and pleckstrin homology domains-containing protein 1 (RAPH1) (Lamellipodin) (Proline-rich EVH1 ligand 2) (PREL-2) (Protein RMO1) (Amyotrophic lateral sclerosis 2 chromosomal region candidate 9 gene protein). | |||||
|
REPS2_MOUSE
|
||||||
| NC score | 0.029696 (rank : 56) | θ value | 1.38821 (rank : 60) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q80XA6 | Gene names | Reps2, Pob1 | |||
|
Domain Architecture |

|
|||||
| Description | RalBP1-associated Eps domain-containing protein 2 (RalBP1-interacting protein 2) (Partner of RalBP1). | |||||
|
MLL2_HUMAN
|
||||||
| NC score | 0.028305 (rank : 57) | θ value | 1.06291 (rank : 51) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
SETBP_MOUSE
|
||||||
| NC score | 0.028127 (rank : 58) | θ value | 4.03905 (rank : 92) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9Z180, Q66JL8 | Gene names | Setbp1, Kiaa0437 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SET-binding protein (SEB). | |||||
|
TARA_MOUSE
|
||||||
| NC score | 0.028088 (rank : 59) | θ value | 1.81305 (rank : 69) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1133 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q99KW3, Q2PZW9, Q2Q402, Q2Q403, Q2Q404, Q6ZPK4, Q8C6T3, Q8CG90 | Gene names | Triobp, Kiaa1662, Tara | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
MAP1A_HUMAN
|
||||||
| NC score | 0.028026 (rank : 60) | θ value | 0.813845 (rank : 45) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | P78559, O95643, Q12973, Q15882, Q9UJT4 | Gene names | MAP1A | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 1A (MAP 1A) (Proliferation-related protein p80) [Contains: MAP1 light chain LC2]. | |||||
|
K1949_MOUSE
|
||||||
| NC score | 0.028014 (rank : 61) | θ value | 1.81305 (rank : 66) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8BQ30, Q3UDS6, Q8CEY9, Q8R3D8 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1949 homolog. | |||||
|
DNL1_HUMAN
|
||||||
| NC score | 0.027909 (rank : 62) | θ value | 0.813845 (rank : 44) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P18858 | Gene names | LIG1 | |||
|
Domain Architecture |
|
|||||
| Description | DNA ligase 1 (EC 6.5.1.1) (DNA ligase I) (Polydeoxyribonucleotide synthase [ATP] 1). | |||||
|
B3A3_MOUSE
|
||||||
| NC score | 0.027844 (rank : 63) | θ value | 0.62314 (rank : 40) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P16283 | Gene names | Slc4a3, Ae3 | |||
|
Domain Architecture |

|
|||||
| Description | Anion exchange protein 3 (Neuronal band 3-like protein) (Solute carrier family 4 member 3). | |||||
|
WASF3_HUMAN
|
||||||
| NC score | 0.026866 (rank : 64) | θ value | 1.06291 (rank : 55) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9UPY6, O94974 | Gene names | WASF3, KIAA0900, SCAR3, WAVE3 | |||
|
Domain Architecture |

|
|||||
| Description | Wiskott-Aldrich syndrome protein family member 3 (WASP-family protein member 3) (Protein WAVE-3) (Verprolin homology domain-containing protein 3). | |||||
|
CG020_HUMAN
|
||||||
| NC score | 0.026516 (rank : 65) | θ value | 1.81305 (rank : 64) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 4 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7L5D6, Q9UFC9, Q9Y309 | Gene names | C7orf20 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UPF0363 protein C7orf20. | |||||
|
EP400_MOUSE
|
||||||
| NC score | 0.025964 (rank : 66) | θ value | 1.38821 (rank : 57) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 592 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q8CHI8, Q80TC8, Q8BXI5, Q8BYW3, Q8C0P6, Q8CHI7, Q8VDF4, Q9DA54 | Gene names | Ep400, Kiaa1498 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E1A-binding protein p400 (EC 3.6.1.-) (p400 kDa SWI2/SNF2-related protein) (Domino homolog) (mDomino). | |||||
|
SYP2L_MOUSE
|
||||||
| NC score | 0.025682 (rank : 67) | θ value | 5.27518 (rank : 100) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8BWB1 | Gene names | Synpo2l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptopodin 2-like protein. | |||||
|
ADDA_HUMAN
|
||||||
| NC score | 0.025537 (rank : 68) | θ value | 6.88961 (rank : 102) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P35611, Q13734, Q14729, Q16156, Q9UJB6 | Gene names | ADD1, ADDA | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-adducin (Erythrocyte adducin subunit alpha). | |||||
|
REPS2_HUMAN
|
||||||
| NC score | 0.024681 (rank : 69) | θ value | 4.03905 (rank : 91) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8NFH8, O43428, Q8NFI5 | Gene names | REPS2, POB1 | |||
|
Domain Architecture |

|
|||||
| Description | RalBP1-associated Eps domain-containing protein 2 (RalBP1-interacting protein 2) (Partner of RalBP1). | |||||
|
FYB_MOUSE
|
||||||
| NC score | 0.024418 (rank : 70) | θ value | 2.36792 (rank : 73) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O35601, Q9Z2H3 | Gene names | Fyb | |||
|
Domain Architecture |

|
|||||
| Description | FYN-binding protein (FYN-T-binding protein) (FYB-120/130) (p120/p130) (SLP-76-associated phosphoprotein) (SLAP-130). | |||||
|
SEPT9_HUMAN
|
||||||
| NC score | 0.024349 (rank : 71) | θ value | 8.99809 (rank : 122) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 706 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q9UHD8, Q96QF3, Q96QF4, Q96QF5, Q9HA04, Q9UG40, Q9Y5W4 | Gene names | SEPT9, KIAA0991, MSF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Septin-9 (MLL septin-like fusion protein) (MLL septin-like fusion protein MSF-A) (Ovarian/Breast septin) (Ov/Br septin) (Septin D1). | |||||
|
ANK2_HUMAN
|
||||||
| NC score | 0.024279 (rank : 72) | θ value | 0.365318 (rank : 29) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
ZN207_HUMAN
|
||||||
| NC score | 0.023983 (rank : 73) | θ value | 8.99809 (rank : 125) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 326 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O43670, Q96HW5, Q9BUQ7 | Gene names | ZNF207 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 207. | |||||
|
RERE_HUMAN
|
||||||
| NC score | 0.023620 (rank : 74) | θ value | 5.27518 (rank : 98) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 779 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9P2R6, O43393, O75046, O75359, Q5VXL9, Q6P6B9, Q9Y2W4 | Gene names | RERE, ARG, ARP, ATN1L, KIAA0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-1-like protein) (Atrophin-1-related protein). | |||||
|
ALEX_MOUSE
|
||||||
| NC score | 0.023191 (rank : 75) | θ value | 6.88961 (rank : 103) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q6R0H6, Q8BIR3 | Gene names | Gnas, Gnas1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein ALEX (Alternative gene product encoded by XL-exon). | |||||
|
GP73_MOUSE
|
||||||
| NC score | 0.022640 (rank : 76) | θ value | 1.06291 (rank : 50) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q91XA2 | Gene names | Golph2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi phosphoprotein 2 (Golgi membrane protein GP73). | |||||
|
MAGI2_MOUSE
|
||||||
| NC score | 0.022609 (rank : 77) | θ value | 0.62314 (rank : 41) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 370 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9WVQ1, Q3UH81, Q6GT88, Q8BYT1, Q8CA85 | Gene names | Magi2, Acvrinp1, Aip1, Arip1 | |||
|
Domain Architecture |

|
|||||
| Description | Membrane-associated guanylate kinase, WW and PDZ domain-containing protein 2 (Membrane-associated guanylate kinase inverted 2) (MAGI-2) (Atrophin-1-interacting protein 1) (Activin receptor-interacting protein 1) (Acvrip1). | |||||
|
ANDR_MOUSE
|
||||||
| NC score | 0.022192 (rank : 78) | θ value | 0.0113563 (rank : 15) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P19091 | Gene names | Ar, Nr3c4 | |||
|
Domain Architecture |

|
|||||
| Description | Androgen receptor (Dihydrotestosterone receptor). | |||||
|
CJ026_MOUSE
|
||||||
| NC score | 0.021895 (rank : 79) | θ value | 3.0926 (rank : 78) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8BGW2, Q3TF48, Q3U3M2, Q6PD43, Q8R0W8 | Gene names | D19Wsu162e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C10orf26 homolog. | |||||
|
MAGI2_HUMAN
|
||||||
| NC score | 0.020975 (rank : 80) | θ value | 1.38821 (rank : 58) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 373 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q86UL8, O60434, O60510, Q86UI7, Q9NP44, Q9UDQ5, Q9UDU1 | Gene names | MAGI2, ACVRINP1, AIP1, KIAA0705 | |||
|
Domain Architecture |

|
|||||
| Description | Membrane-associated guanylate kinase, WW and PDZ domain-containing protein 2 (Membrane-associated guanylate kinase inverted 2) (MAGI-2) (Atrophin-1-interacting protein 1) (Atrophin-1-interacting protein A). | |||||
|
CSKI1_MOUSE
|
||||||
| NC score | 0.020899 (rank : 81) | θ value | 4.03905 (rank : 85) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1031 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q6P9K8, Q6ZPU2, Q8BWU2, Q8BX99, Q9CXH0 | Gene names | Caskin1, Kiaa1306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Caskin-1 (CASK-interacting protein 1). | |||||
|
DGKK_HUMAN
|
||||||
| NC score | 0.020655 (rank : 82) | θ value | 3.0926 (rank : 80) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 378 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q5KSL6 | Gene names | DGKK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Diacylglycerol kinase kappa (EC 2.7.1.107) (Diglyceride kinase kappa) (DGK-kappa) (DAG kinase kappa) (142 kDa diacylglycerol kinase). | |||||
|
EP400_HUMAN
|
||||||
| NC score | 0.020575 (rank : 83) | θ value | 8.99809 (rank : 114) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 556 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q96L91, O15411, Q6P2F5, Q8N8Q7, Q8NE05, Q96JK7, Q9P230 | Gene names | EP400, CAGH32, KIAA1498, KIAA1818, TNRC12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E1A-binding protein p400 (EC 3.6.1.-) (p400 kDa SWI2/SNF2-related protein) (Domino homolog) (hDomino) (CAG repeat protein 32) (Trinucleotide repeat-containing gene 12 protein). | |||||
|
ZFHX2_HUMAN
|
||||||
| NC score | 0.020019 (rank : 84) | θ value | 0.47712 (rank : 39) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1178 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9C0A1 | Gene names | ZFHX2, KIAA1762 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger homeobox protein 2. | |||||
|
BRD4_MOUSE
|
||||||
| NC score | 0.019856 (rank : 85) | θ value | 4.03905 (rank : 84) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 845 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9ESU6, Q8VHF7, Q8VHF8 | Gene names | Brd4, Mcap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain-containing protein 4 (Mitotic chromosome-associated protein) (MCAP). | |||||
|
CD5_MOUSE
|
||||||
| NC score | 0.019141 (rank : 86) | θ value | 2.36792 (rank : 72) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P13379 | Gene names | Cd5, Ly-1 | |||
|
Domain Architecture |

|
|||||
| Description | T-cell surface glycoprotein CD5 precursor (Lymphocyte antigen Ly-1) (Lyt-1). | |||||
|
PHC1_HUMAN
|
||||||
| NC score | 0.018955 (rank : 87) | θ value | 5.27518 (rank : 96) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P78364, Q8WVM3, Q9BU63 | Gene names | PHC1, EDR1, PH1 | |||
|
Domain Architecture |

|
|||||
| Description | Polyhomeotic-like protein 1 (hPH1) (Early development regulatory protein 1). | |||||
|
PPRB_MOUSE
|
||||||
| NC score | 0.018887 (rank : 88) | θ value | 5.27518 (rank : 97) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q925J9, O88323, Q3UHV0, Q6AXD5, Q8BW37, Q8BX19, Q8VDQ7, Q925K0 | Gene names | Pparbp, Trip2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor-binding protein (PBP) (PPAR-binding protein) (Thyroid hormone receptor-associated protein complex 220 kDa component) (Trap220) (Thyroid receptor-interacting protein 2) (TRIP-2). | |||||
|
SYNJ2_HUMAN
|
||||||
| NC score | 0.018172 (rank : 89) | θ value | 1.38821 (rank : 62) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O15056, Q5TA13, Q5TA16, Q5TA19, Q86XK0, Q8IZA8, Q9H226 | Gene names | SYNJ2, KIAA0348 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptojanin-2 (EC 3.1.3.36) (Synaptic inositol-1,4,5-trisphosphate 5- phosphatase 2). | |||||
|
CSKI1_HUMAN
|
||||||
| NC score | 0.018148 (rank : 90) | θ value | 3.0926 (rank : 79) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 997 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8WXD9, Q9P2P0 | Gene names | CASKIN1, KIAA1306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Caskin-1 (CASK-interacting protein 1). | |||||
|
S4A7_HUMAN
|
||||||
| NC score | 0.017623 (rank : 91) | θ value | 8.99809 (rank : 120) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Y6M7, O60350, Q6AHZ9, Q9HC88, Q9UIB9 | Gene names | SLC4A7, BT, NBC2, NBC2B, NBC3, SBC2, SLC4A6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium bicarbonate cotransporter 3 (Sodium bicarbonate cotransporter 2) (Sodium bicarbonate cotransporter 2b) (Bicarbonate transporter) (Solute carrier family 4 member 7). | |||||
|
OXR1_MOUSE
|
||||||
| NC score | 0.017469 (rank : 92) | θ value | 3.0926 (rank : 83) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 516 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q4KMM3, Q5FWW1, Q99L06, Q99MK1, Q99MP4 | Gene names | Oxr1, C7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Oxidation resistance protein 1 (Protein C7). | |||||
|
MOR2A_MOUSE
|
||||||
| NC score | 0.016730 (rank : 93) | θ value | 3.0926 (rank : 82) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 220 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q69ZX6, Q5QNQ7, Q6P547 | Gene names | Morc2a, Kiaa0852, Zcwcc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MORC family CW-type zinc finger protein 2A (Zinc finger CW-type coiled-coil domain protein 1). | |||||
|
FBX46_MOUSE
|
||||||
| NC score | 0.015142 (rank : 94) | θ value | 8.99809 (rank : 115) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8BG80 | Gene names | Fbxo46, Fbx46, Fbxo34l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 46 (F-box only protein 34-like). | |||||
|
SYN2_HUMAN
|
||||||
| NC score | 0.015111 (rank : 95) | θ value | 6.88961 (rank : 112) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q92777 | Gene names | SYN2 | |||
|
Domain Architecture |

|
|||||
| Description | Synapsin-2 (Synapsin II). | |||||
|
BC11A_MOUSE
|
||||||
| NC score | 0.013768 (rank : 96) | θ value | 0.813845 (rank : 43) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 732 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9QYE3, Q80T89, Q8BLC7, Q8BLR4, Q8BWX3, Q921V4, Q9D0V2, Q9JIT4, Q9JLK8, Q9JLK9 | Gene names | Bcl11a, Ctip1, Evi9, Kiaa1809 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma/leukemia 11A (B-cell CLL/lymphoma 11A) (COUP-TF- interacting protein 1) (Ecotropic viral integration site 9 protein) (EVI-9). | |||||
|
STAT6_MOUSE
|
||||||
| NC score | 0.013759 (rank : 97) | θ value | 2.36792 (rank : 74) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P52633 | Gene names | Stat6 | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and transcription activator 6. | |||||
|
GLU2B_MOUSE
|
||||||
| NC score | 0.013713 (rank : 98) | θ value | 6.88961 (rank : 104) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O08795, Q921X2 | Gene names | Prkcsh | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glucosidase 2 subunit beta precursor (Glucosidase II subunit beta) (Protein kinase C substrate, 60.1 kDa protein, heavy chain) (PKCSH) (80K-H protein). | |||||
|
SPTB2_HUMAN
|
||||||
| NC score | 0.012544 (rank : 99) | θ value | 0.163984 (rank : 26) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 748 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q01082, O60837, Q16057 | Gene names | SPTBN1, SPTB2 | |||
|
Domain Architecture |

|
|||||
| Description | Spectrin beta chain, brain 1 (Spectrin, non-erythroid beta chain 1) (Beta-II spectrin) (Fodrin beta chain). | |||||
|
K0174_MOUSE
|
||||||
| NC score | 0.011918 (rank : 100) | θ value | 8.99809 (rank : 116) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9CX00, Q80U68 | Gene names | Kiaa0174 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0174. | |||||
|
SPEG_MOUSE
|
||||||
| NC score | 0.011659 (rank : 101) | θ value | 0.813845 (rank : 46) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1959 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q62407, Q3TPH8, Q6P5V1, Q80TF7, Q80ZN0, Q8BZF4, Q9EQJ5 | Gene names | Speg, Apeg1, Kiaa1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle-specific serine/threonine protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
ES8L3_HUMAN
|
||||||
| NC score | 0.011604 (rank : 102) | θ value | 4.03905 (rank : 87) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8TE67, Q5T8Q6, Q5T8Q7, Q5T8Q8, Q96E47, Q9H719 | Gene names | EPS8L3, EPS8R3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Epidermal growth factor receptor kinase substrate 8-like protein 3 (Epidermal growth factor receptor pathway substrate 8-related protein 3) (EPS8-like protein 3). | |||||
|
BC11B_HUMAN
|
||||||
| NC score | 0.011292 (rank : 103) | θ value | 0.47712 (rank : 33) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 761 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9C0K0, Q9H162 | Gene names | BCL11B, CTIP2, RIT1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma/leukemia 11B (B-cell CLL/lymphoma 11B) (Radiation- induced tumor suppressor gene 1 protein) (hRit1) (COUP-TF-interacting protein 2). | |||||
|
CEBPA_MOUSE
|
||||||
| NC score | 0.011194 (rank : 104) | θ value | 5.27518 (rank : 95) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P53566, Q91XB6 | Gene names | Cebpa | |||
|
Domain Architecture |

|
|||||
| Description | CCAAT/enhancer-binding protein alpha (C/EBP alpha). | |||||
|
BC11A_HUMAN
|
||||||
| NC score | 0.010898 (rank : 105) | θ value | 0.813845 (rank : 42) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 745 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9H165, Q86W14, Q8WU92, Q96JL6, Q9H163, Q9H164, Q9H3G9, Q9NWA7 | Gene names | BCL11A, CTIP1, EVI9, KIAA1809 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma/leukemia 11A (B-cell CLL/lymphoma 11A) (COUP-TF- interacting protein 1) (Ecotropic viral integration site 9 protein) (EVI-9). | |||||
|
USH1C_MOUSE
|
||||||
| NC score | 0.010873 (rank : 106) | θ value | 8.99809 (rank : 123) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 557 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9ES64, Q91XD1, Q9CVG7, Q9ES65 | Gene names | Ush1c | |||
|
Domain Architecture |

|
|||||
| Description | Harmonin (Usher syndrome type-1C protein homolog) (PDZ domain- containing protein). | |||||
|
CHD3_HUMAN
|
||||||
| NC score | 0.010786 (rank : 107) | θ value | 3.0926 (rank : 77) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q12873, Q9Y4I0 | Gene names | CHD3 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain helicase-DNA-binding protein 3 (EC 3.6.1.-) (ATP- dependent helicase CHD3) (CHD-3) (Mi-2 autoantigen 240 kDa protein) (Mi2-alpha) (Zinc-finger helicase) (hZFH). | |||||
|
TITIN_HUMAN
|
||||||
| NC score | 0.009854 (rank : 108) | θ value | 0.0736092 (rank : 24) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 2923 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q8WZ42, Q10465, Q10466, Q15598, Q2XUS3, Q32Q60, Q4U1Z6, Q6NSG0, Q6PDB1, Q6PJP0, Q7KYM2, Q7KYN4, Q7KYN5, Q7LDM3, Q7Z2X3, Q8TCG8, Q8WZ51, Q8WZ52, Q8WZ53, Q8WZB3, Q92761, Q92762, Q9UD97, Q9UP84, Q9Y6L9 | Gene names | TTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Titin (EC 2.7.11.1) (Connectin) (Rhabdomyosarcoma antigen MU-RMS- 40.14). | |||||
|
TNNT1_HUMAN
|
||||||
| NC score | 0.009556 (rank : 109) | θ value | 6.88961 (rank : 113) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P13805, O95472, Q16061 | Gene names | TNNT1, TNT | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Troponin T, slow skeletal muscle (TnTs) (Slow skeletal muscle troponin T) (sTnT). | |||||
|
GPC2_HUMAN
|
||||||
| NC score | 0.009541 (rank : 110) | θ value | 4.03905 (rank : 88) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8N158 | Gene names | GPC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glypican-2 precursor. | |||||
|
BC11B_MOUSE
|
||||||
| NC score | 0.009530 (rank : 111) | θ value | 5.27518 (rank : 94) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 767 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q99PV8, Q8C2I1, Q99PV6, Q99PV7, Q9JLF8 | Gene names | Bcl11b, Ctip2, Rit1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | B-cell lymphoma/leukemia 11B (B-cell CLL/lymphoma 11B) (Radiation- induced tumor suppressor gene 1 protein) (mRit1) (COUP-TF-interacting protein 2). | |||||
|
RUSC1_HUMAN
|
||||||
| NC score | 0.009502 (rank : 112) | θ value | 6.88961 (rank : 110) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 226 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9BVN2, Q9UPY4, Q9Y4T5 | Gene names | RUSC1, NESCA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RUN and SH3 domain-containing protein 1 (New molecule containing SH3 at the carboxy-terminus) (Nesca). | |||||
|
AAK1_HUMAN
|
||||||
| NC score | 0.009425 (rank : 113) | θ value | 1.81305 (rank : 63) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1077 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q2M2I8, Q4ZFZ3, Q53RX6, Q9UPV4 | Gene names | AAK1, KIAA1048 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AP2-associated protein kinase 1 (EC 2.7.11.1) (Adaptor-associated kinase 1). | |||||
|
RYR1_HUMAN
|
||||||
| NC score | 0.008529 (rank : 114) | θ value | 6.88961 (rank : 111) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P21817, Q16314, Q16368, Q9NPK1, Q9P1U4 | Gene names | RYR1, RYDR | |||
|
Domain Architecture |

|
|||||
| Description | Ryanodine receptor 1 (Skeletal muscle-type ryanodine receptor) (RyR1) (RYR-1) (Skeletal muscle calcium release channel). | |||||
|
LR37B_HUMAN
|
||||||
| NC score | 0.008415 (rank : 115) | θ value | 8.99809 (rank : 118) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q96QE4, Q5YKG6 | Gene names | LRRC37B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 37B precursor (C66 SLIT-like testicular protein). | |||||
|
YBOX2_MOUSE
|
||||||
| NC score | 0.008227 (rank : 116) | θ value | 8.99809 (rank : 124) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Z2C8, Q5NCW8, Q5NCW9, Q9Z2C7 | Gene names | Ybx2, Msy2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Y-box-binding protein 2 (Germ cell-specific Y-box-binding protein) (FRGY2 homolog). | |||||
|
PHF2_MOUSE
|
||||||
| NC score | 0.008093 (rank : 117) | θ value | 6.88961 (rank : 108) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9WTU0, Q80WA8 | Gene names | Phf2 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 2 (GRC5). | |||||
|
LBXCO_HUMAN
|
||||||
| NC score | 0.007847 (rank : 118) | θ value | 8.99809 (rank : 117) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P84550 | Gene names | LBXCOR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ladybird homeobox corepressor 1. | |||||
|
KCNN3_MOUSE
|
||||||
| NC score | 0.006938 (rank : 119) | θ value | 6.88961 (rank : 106) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P58391 | Gene names | Kcnn3, Sk3 | |||
|
Domain Architecture |
|
|||||
| Description | Small conductance calcium-activated potassium channel protein 3 (SK3). | |||||
|
PO6F1_HUMAN
|
||||||
| NC score | 0.006716 (rank : 120) | θ value | 4.03905 (rank : 90) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 370 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q14863 | Gene names | POU6F1, BRN5, MPOU, TCFB1 | |||
|
Domain Architecture |

|
|||||
| Description | POU domain, class 6, transcription factor 1 (mPOU homeobox protein) (Brain-specific homeobox/POU domain protein 5) (Brain-5) (Brn-5 protein). | |||||
|
ADA19_MOUSE
|
||||||
| NC score | 0.005384 (rank : 121) | θ value | 3.0926 (rank : 76) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O35674 | Gene names | Adam19, Mltnb | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 19 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 19) (Meltrin beta). | |||||
|
KI21B_HUMAN
|
||||||
| NC score | 0.001578 (rank : 122) | θ value | 6.88961 (rank : 107) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1017 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O75037, Q5T4J3 | Gene names | KIF21B, KIAA0449 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin family member 21B. | |||||
|
PRP4B_HUMAN
|
||||||
| NC score | 0.001482 (rank : 123) | θ value | 6.88961 (rank : 109) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 1030 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q13523, Q8TDP2, Q96QT7, Q9UEE6 | Gene names | PRPF4B, KIAA0536, PRP4, PRP4H, PRP4K | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase PRP4 homolog (EC 2.7.11.1) (PRP4 pre- mRNA-processing factor 4 homolog) (PRP4 kinase). | |||||
|
ZN646_HUMAN
|
||||||
| NC score | 0.000404 (rank : 124) | θ value | 4.03905 (rank : 93) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 813 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O15015 | Gene names | ZNF646, KIAA0296 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 646. | |||||
|
ZN579_HUMAN
|
||||||
| NC score | 0.000322 (rank : 125) | θ value | 5.27518 (rank : 101) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 726 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8NAF0 | Gene names | ZNF579 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 579. | |||||
|
SBK1_MOUSE
|
||||||
| NC score | -0.000537 (rank : 126) | θ value | 8.99809 (rank : 121) | |||
| Query Neighborhood Hits | 125 | Target Neighborhood Hits | 824 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8QZX0 | Gene names | Sbk1, Sbk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SBK1 (EC 2.7.11.1) (SH3-binding kinase 1). | |||||