Please be patient as the page loads
|
MA1A2_HUMAN
|
||||||
| SwissProt Accessions | O60476, Q9H510 | Gene names | MAN1A2, MAN1B | |||
|
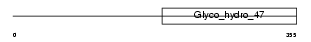
Domain Architecture |
|
|||||
| Description | Mannosyl-oligosaccharide 1,2-alpha-mannosidase IB (EC 3.2.1.113) (Processing alpha-1,2-mannosidase IB) (Alpha-1,2-mannosidase IB) (Mannosidase alpha class 1A member 2). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
MA1A1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.992195 (rank : 3) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P33908, Q9NU44, Q9UJI3 | Gene names | MAN1A1 | |||
|
Domain Architecture |

|
|||||
| Description | Mannosyl-oligosaccharide 1,2-alpha-mannosidase IA (EC 3.2.1.113) (Processing alpha-1,2-mannosidase IA) (Alpha-1,2-mannosidase IA) (Mannosidase alpha class 1A member 1) (Man(9)-alpha-mannosidase) (Man9-mannosidase). | |||||
|
MA1A1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.991064 (rank : 4) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P45700 | Gene names | Man1a1, Man1a | |||
|
Domain Architecture |

|
|||||
| Description | Mannosyl-oligosaccharide 1,2-alpha-mannosidase IA (EC 3.2.1.113) (Processing alpha-1,2-mannosidase IA) (Alpha-1,2-mannosidase IA) (Mannosidase alpha class 1A member 1) (Man(9)-alpha-mannosidase) (Man9-mannosidase). | |||||
|
MA1A2_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O60476, Q9H510 | Gene names | MAN1A2, MAN1B | |||
|
Domain Architecture |

|
|||||
| Description | Mannosyl-oligosaccharide 1,2-alpha-mannosidase IB (EC 3.2.1.113) (Processing alpha-1,2-mannosidase IB) (Alpha-1,2-mannosidase IB) (Mannosidase alpha class 1A member 2). | |||||
|
MA1A2_MOUSE
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.998265 (rank : 2) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P39098, Q5SUY6, Q5SUY7, Q60599, Q9CUY9 | Gene names | Man1a2, Man1b | |||
|
Domain Architecture |

|
|||||
| Description | Mannosyl-oligosaccharide 1,2-alpha-mannosidase IB (EC 3.2.1.113) (Processing alpha-1,2-mannosidase IB) (Alpha-1,2-mannosidase IB) (Mannosidase alpha class 1A member 2). | |||||
|
MA1C1_HUMAN
|
||||||
| θ value | 1.04765e-165 (rank : 5) | NC score | 0.988076 (rank : 5) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NR34, Q9Y545 | Gene names | MAN1C1, MAN1A3, MAN1C | |||
|
Domain Architecture |

|
|||||
| Description | Mannosyl-oligosaccharide 1,2-alpha-mannosidase IC (EC 3.2.1.113) (Processing alpha-1,2-mannosidase IC) (Alpha-1,2-mannosidase IC) (Mannosidase alpha class 1C member 1) (HMIC). | |||||
|
MA1B1_HUMAN
|
||||||
| θ value | 3.93732e-88 (rank : 6) | NC score | 0.930330 (rank : 6) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9UKM7, Q5VSG3, Q9BRS9, Q9Y5K7 | Gene names | MAN1B1 | |||
|
Domain Architecture |

|
|||||
| Description | Endoplasmic reticulum mannosyl-oligosaccharide 1,2-alpha-mannosidase (EC 3.2.1.113) (ER alpha-1,2-mannosidase) (Mannosidase alpha class 1B member 1) (Man9GlcNAc2-specific-processing alpha-mannosidase). | |||||
|
EDEM1_HUMAN
|
||||||
| θ value | 1.01529e-43 (rank : 7) | NC score | 0.838424 (rank : 8) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q92611 | Gene names | EDEM1, EDEM, KIAA0212 | |||
|
Domain Architecture |

|
|||||
| Description | ER degradation-enhancing alpha-mannosidase-like 1. | |||||
|
EDEM1_MOUSE
|
||||||
| θ value | 3.85808e-43 (rank : 8) | NC score | 0.837837 (rank : 9) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q925U4 | Gene names | Edem1, Edem | |||
|
Domain Architecture |

|
|||||
| Description | ER degradation-enhancing alpha-mannosidase-like 1. | |||||
|
EDEM2_HUMAN
|
||||||
| θ value | 2.76489e-41 (rank : 9) | NC score | 0.873601 (rank : 7) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9BV94, Q6IA89, Q6UWZ4, Q9H4U0, Q9H886, Q9NTL9, Q9NVE6 | Gene names | EDEM2, C20orf31, C20orf49 | |||
|
Domain Architecture |

|
|||||
| Description | ER degradation-enhancing alpha-mannosidase-like 2 precursor. | |||||
|
EDEM3_HUMAN
|
||||||
| θ value | 2.86122e-38 (rank : 10) | NC score | 0.824776 (rank : 10) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9BZQ6, Q5TEZ0, Q9HCW1, Q9UFV7 | Gene names | EDEM3, C1orf22 | |||
|
Domain Architecture |

|
|||||
| Description | ER degradation-enhancing alpha-mannosidase-like 3. | |||||
|
RAB3I_HUMAN
|
||||||
| θ value | 0.365318 (rank : 11) | NC score | 0.029734 (rank : 11) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 270 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96QF0, Q6PCE4, Q96A24, Q96QE6, Q96QE7, Q96QE8, Q96QE9, Q96QF1, Q9H673 | Gene names | RAB3IP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RAB3A-interacting protein (Rabin-3) (SSX2-interacting protein). | |||||
|
C8AP2_MOUSE
|
||||||
| θ value | 0.813845 (rank : 12) | NC score | 0.028465 (rank : 12) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 405 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9WUF3, Q9CSW2 | Gene names | Casp8ap2, Flash | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CASP8-associated protein 2 (FLICE-associated huge protein). | |||||
|
IFT74_MOUSE
|
||||||
| θ value | 1.06291 (rank : 13) | NC score | 0.019832 (rank : 15) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 939 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8BKE9, Q80ZI3, Q9CSY1, Q9CUS0, Q9D9T5 | Gene names | Ift74, Ccdc2, Cmg1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intraflagellar transport 74 homolog (Coiled-coil domain-containing protein 2) (Capillary morphogenesis protein 1) (CMG-1). | |||||
|
GRAP1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 14) | NC score | 0.019927 (rank : 14) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 944 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q4V328, Q4V327, Q4V330, Q5HYG1, Q6N046, Q96DH8, Q9NQ43, Q9ULQ3 | Gene names | GRIPAP1, KIAA1167 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GRIP1-associated protein 1 (GRASP-1). | |||||
|
GRAP1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 15) | NC score | 0.019377 (rank : 16) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1234 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8VD04, O35693, Q3T9C3, Q69ZP9 | Gene names | Gripap1, DXImx47e, Kiaa1167 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GRIP1-associated protein 1 (GRASP-1) (HCMV-interacting protein). | |||||
|
IFT74_HUMAN
|
||||||
| θ value | 3.0926 (rank : 16) | NC score | 0.013745 (rank : 17) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 949 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q96LB3, Q6PGQ8, Q9H643, Q9H8G7 | Gene names | IFT74, CCDC2, CMG1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intraflagellar transport 74 homolog (Coiled-coil domain-containing protein 2) (Capillary morphogenesis protein 1) (CMG-1). | |||||
|
TTC17_HUMAN
|
||||||
| θ value | 3.0926 (rank : 17) | NC score | 0.020737 (rank : 13) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96AE7 | Gene names | TTC17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tetratricopeptide repeat protein 17 (TPR repeat protein 17). | |||||
|
CASP_HUMAN
|
||||||
| θ value | 4.03905 (rank : 18) | NC score | 0.009063 (rank : 21) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1131 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q13948, Q53GU9, Q8TBS3 | Gene names | CUTL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein CASP. | |||||
|
CUTL1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 19) | NC score | 0.008269 (rank : 23) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1574 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P39880, Q6NYH4, Q9UEV5 | Gene names | CUTL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP). | |||||
|
PIBF1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 20) | NC score | 0.008563 (rank : 22) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8WXW3, O95664, Q6U9V2, Q6UG50, Q86V07, Q96SF4 | Gene names | PIBF1, C13orf24, PIBF | |||
|
Domain Architecture |

|
|||||
| Description | Progesterone-induced-blocking factor 1. | |||||
|
KI21B_HUMAN
|
||||||
| θ value | 5.27518 (rank : 21) | NC score | 0.007616 (rank : 24) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1017 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O75037, Q5T4J3 | Gene names | KIF21B, KIAA0449 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin family member 21B. | |||||
|
KI21B_MOUSE
|
||||||
| θ value | 5.27518 (rank : 22) | NC score | 0.006698 (rank : 25) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 997 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9QXL1, P97424 | Gene names | Kif21b, Kif6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin family member 21B (Kinesin-like protein KIF6). | |||||
|
MYH11_MOUSE
|
||||||
| θ value | 5.27518 (rank : 23) | NC score | 0.005571 (rank : 30) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1777 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O08638, O08639, Q62462, Q64195 | Gene names | Myh11 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
LEO1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 24) | NC score | 0.010267 (rank : 18) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
BTBD7_MOUSE
|
||||||
| θ value | 8.99809 (rank : 25) | NC score | 0.009989 (rank : 19) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8CFE5, Q80TC3, Q8BZY9 | Gene names | Btbd7, Kiaa1525 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein 7. | |||||
|
CCD41_HUMAN
|
||||||
| θ value | 8.99809 (rank : 26) | NC score | 0.006313 (rank : 28) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1139 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Y592 | Gene names | CCDC41 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 41 (NY-REN-58 antigen). | |||||
|
FLIP1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 27) | NC score | 0.006192 (rank : 29) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1176 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9CS72 | Gene names | Filip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filamin-A-interacting protein 1 (FILIP). | |||||
|
LAMB1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 28) | NC score | 0.004376 (rank : 31) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1018 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P07942 | Gene names | LAMB1 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-1 chain precursor (Laminin B1 chain). | |||||
|
PCNT_HUMAN
|
||||||
| θ value | 8.99809 (rank : 29) | NC score | 0.009667 (rank : 20) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1551 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O95613, O43152, Q7Z7C9 | Gene names | PCNT, KIAA0402, PCNT2 | |||
|
Domain Architecture |

|
|||||
| Description | Pericentrin (Pericentrin B) (Kendrin). | |||||
|
STIM1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 30) | NC score | 0.006571 (rank : 26) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 665 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q13586 | Gene names | STIM1, GOK | |||
|
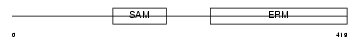
Domain Architecture |
|
|||||
| Description | Stromal interaction molecule 1 precursor. | |||||
|
SYCP1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 31) | NC score | 0.006425 (rank : 27) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1313 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q15431, O14963 | Gene names | SYCP1, SCP1 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptonemal complex protein 1 (SCP-1). | |||||
|
MA1A2_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O60476, Q9H510 | Gene names | MAN1A2, MAN1B | |||
|
Domain Architecture |

|
|||||
| Description | Mannosyl-oligosaccharide 1,2-alpha-mannosidase IB (EC 3.2.1.113) (Processing alpha-1,2-mannosidase IB) (Alpha-1,2-mannosidase IB) (Mannosidase alpha class 1A member 2). | |||||
|
MA1A2_MOUSE
|
||||||
| NC score | 0.998265 (rank : 2) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P39098, Q5SUY6, Q5SUY7, Q60599, Q9CUY9 | Gene names | Man1a2, Man1b | |||
|
Domain Architecture |

|
|||||
| Description | Mannosyl-oligosaccharide 1,2-alpha-mannosidase IB (EC 3.2.1.113) (Processing alpha-1,2-mannosidase IB) (Alpha-1,2-mannosidase IB) (Mannosidase alpha class 1A member 2). | |||||
|
MA1A1_HUMAN
|
||||||
| NC score | 0.992195 (rank : 3) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P33908, Q9NU44, Q9UJI3 | Gene names | MAN1A1 | |||
|
Domain Architecture |

|
|||||
| Description | Mannosyl-oligosaccharide 1,2-alpha-mannosidase IA (EC 3.2.1.113) (Processing alpha-1,2-mannosidase IA) (Alpha-1,2-mannosidase IA) (Mannosidase alpha class 1A member 1) (Man(9)-alpha-mannosidase) (Man9-mannosidase). | |||||
|
MA1A1_MOUSE
|
||||||
| NC score | 0.991064 (rank : 4) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P45700 | Gene names | Man1a1, Man1a | |||
|
Domain Architecture |

|
|||||
| Description | Mannosyl-oligosaccharide 1,2-alpha-mannosidase IA (EC 3.2.1.113) (Processing alpha-1,2-mannosidase IA) (Alpha-1,2-mannosidase IA) (Mannosidase alpha class 1A member 1) (Man(9)-alpha-mannosidase) (Man9-mannosidase). | |||||
|
MA1C1_HUMAN
|
||||||
| NC score | 0.988076 (rank : 5) | θ value | 1.04765e-165 (rank : 5) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NR34, Q9Y545 | Gene names | MAN1C1, MAN1A3, MAN1C | |||
|
Domain Architecture |

|
|||||
| Description | Mannosyl-oligosaccharide 1,2-alpha-mannosidase IC (EC 3.2.1.113) (Processing alpha-1,2-mannosidase IC) (Alpha-1,2-mannosidase IC) (Mannosidase alpha class 1C member 1) (HMIC). | |||||
|
MA1B1_HUMAN
|
||||||
| NC score | 0.930330 (rank : 6) | θ value | 3.93732e-88 (rank : 6) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9UKM7, Q5VSG3, Q9BRS9, Q9Y5K7 | Gene names | MAN1B1 | |||
|
Domain Architecture |

|
|||||
| Description | Endoplasmic reticulum mannosyl-oligosaccharide 1,2-alpha-mannosidase (EC 3.2.1.113) (ER alpha-1,2-mannosidase) (Mannosidase alpha class 1B member 1) (Man9GlcNAc2-specific-processing alpha-mannosidase). | |||||
|
EDEM2_HUMAN
|
||||||
| NC score | 0.873601 (rank : 7) | θ value | 2.76489e-41 (rank : 9) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9BV94, Q6IA89, Q6UWZ4, Q9H4U0, Q9H886, Q9NTL9, Q9NVE6 | Gene names | EDEM2, C20orf31, C20orf49 | |||
|
Domain Architecture |

|
|||||
| Description | ER degradation-enhancing alpha-mannosidase-like 2 precursor. | |||||
|
EDEM1_HUMAN
|
||||||
| NC score | 0.838424 (rank : 8) | θ value | 1.01529e-43 (rank : 7) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q92611 | Gene names | EDEM1, EDEM, KIAA0212 | |||
|
Domain Architecture |

|
|||||
| Description | ER degradation-enhancing alpha-mannosidase-like 1. | |||||
|
EDEM1_MOUSE
|
||||||
| NC score | 0.837837 (rank : 9) | θ value | 3.85808e-43 (rank : 8) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q925U4 | Gene names | Edem1, Edem | |||
|
Domain Architecture |

|
|||||
| Description | ER degradation-enhancing alpha-mannosidase-like 1. | |||||
|
EDEM3_HUMAN
|
||||||
| NC score | 0.824776 (rank : 10) | θ value | 2.86122e-38 (rank : 10) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9BZQ6, Q5TEZ0, Q9HCW1, Q9UFV7 | Gene names | EDEM3, C1orf22 | |||
|
Domain Architecture |

|
|||||
| Description | ER degradation-enhancing alpha-mannosidase-like 3. | |||||
|
RAB3I_HUMAN
|
||||||
| NC score | 0.029734 (rank : 11) | θ value | 0.365318 (rank : 11) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 270 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96QF0, Q6PCE4, Q96A24, Q96QE6, Q96QE7, Q96QE8, Q96QE9, Q96QF1, Q9H673 | Gene names | RAB3IP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RAB3A-interacting protein (Rabin-3) (SSX2-interacting protein). | |||||
|
C8AP2_MOUSE
|
||||||
| NC score | 0.028465 (rank : 12) | θ value | 0.813845 (rank : 12) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 405 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9WUF3, Q9CSW2 | Gene names | Casp8ap2, Flash | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CASP8-associated protein 2 (FLICE-associated huge protein). | |||||
|
TTC17_HUMAN
|
||||||
| NC score | 0.020737 (rank : 13) | θ value | 3.0926 (rank : 17) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96AE7 | Gene names | TTC17 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tetratricopeptide repeat protein 17 (TPR repeat protein 17). | |||||
|
GRAP1_HUMAN
|
||||||
| NC score | 0.019927 (rank : 14) | θ value | 1.81305 (rank : 14) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 944 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q4V328, Q4V327, Q4V330, Q5HYG1, Q6N046, Q96DH8, Q9NQ43, Q9ULQ3 | Gene names | GRIPAP1, KIAA1167 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GRIP1-associated protein 1 (GRASP-1). | |||||
|
IFT74_MOUSE
|
||||||
| NC score | 0.019832 (rank : 15) | θ value | 1.06291 (rank : 13) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 939 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8BKE9, Q80ZI3, Q9CSY1, Q9CUS0, Q9D9T5 | Gene names | Ift74, Ccdc2, Cmg1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intraflagellar transport 74 homolog (Coiled-coil domain-containing protein 2) (Capillary morphogenesis protein 1) (CMG-1). | |||||
|
GRAP1_MOUSE
|
||||||
| NC score | 0.019377 (rank : 16) | θ value | 1.81305 (rank : 15) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1234 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8VD04, O35693, Q3T9C3, Q69ZP9 | Gene names | Gripap1, DXImx47e, Kiaa1167 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GRIP1-associated protein 1 (GRASP-1) (HCMV-interacting protein). | |||||
|
IFT74_HUMAN
|
||||||
| NC score | 0.013745 (rank : 17) | θ value | 3.0926 (rank : 16) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 949 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q96LB3, Q6PGQ8, Q9H643, Q9H8G7 | Gene names | IFT74, CCDC2, CMG1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intraflagellar transport 74 homolog (Coiled-coil domain-containing protein 2) (Capillary morphogenesis protein 1) (CMG-1). | |||||
|
LEO1_MOUSE
|
||||||
| NC score | 0.010267 (rank : 18) | θ value | 6.88961 (rank : 24) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
BTBD7_MOUSE
|
||||||
| NC score | 0.009989 (rank : 19) | θ value | 8.99809 (rank : 25) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8CFE5, Q80TC3, Q8BZY9 | Gene names | Btbd7, Kiaa1525 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein 7. | |||||
|
PCNT_HUMAN
|
||||||
| NC score | 0.009667 (rank : 20) | θ value | 8.99809 (rank : 29) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1551 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O95613, O43152, Q7Z7C9 | Gene names | PCNT, KIAA0402, PCNT2 | |||
|
Domain Architecture |

|
|||||
| Description | Pericentrin (Pericentrin B) (Kendrin). | |||||
|
CASP_HUMAN
|
||||||
| NC score | 0.009063 (rank : 21) | θ value | 4.03905 (rank : 18) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1131 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q13948, Q53GU9, Q8TBS3 | Gene names | CUTL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein CASP. | |||||
|
PIBF1_HUMAN
|
||||||
| NC score | 0.008563 (rank : 22) | θ value | 4.03905 (rank : 20) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8WXW3, O95664, Q6U9V2, Q6UG50, Q86V07, Q96SF4 | Gene names | PIBF1, C13orf24, PIBF | |||
|
Domain Architecture |

|
|||||
| Description | Progesterone-induced-blocking factor 1. | |||||
|
CUTL1_HUMAN
|
||||||
| NC score | 0.008269 (rank : 23) | θ value | 4.03905 (rank : 19) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1574 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P39880, Q6NYH4, Q9UEV5 | Gene names | CUTL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein cut-like 1 (CCAAT displacement protein) (CDP). | |||||
|
KI21B_HUMAN
|
||||||
| NC score | 0.007616 (rank : 24) | θ value | 5.27518 (rank : 21) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1017 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O75037, Q5T4J3 | Gene names | KIF21B, KIAA0449 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin family member 21B. | |||||
|
KI21B_MOUSE
|
||||||
| NC score | 0.006698 (rank : 25) | θ value | 5.27518 (rank : 22) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 997 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9QXL1, P97424 | Gene names | Kif21b, Kif6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin family member 21B (Kinesin-like protein KIF6). | |||||
|
STIM1_HUMAN
|
||||||
| NC score | 0.006571 (rank : 26) | θ value | 8.99809 (rank : 30) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 665 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q13586 | Gene names | STIM1, GOK | |||
|
Domain Architecture |

|
|||||
| Description | Stromal interaction molecule 1 precursor. | |||||
|
SYCP1_HUMAN
|
||||||
| NC score | 0.006425 (rank : 27) | θ value | 8.99809 (rank : 31) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1313 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q15431, O14963 | Gene names | SYCP1, SCP1 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptonemal complex protein 1 (SCP-1). | |||||
|
CCD41_HUMAN
|
||||||
| NC score | 0.006313 (rank : 28) | θ value | 8.99809 (rank : 26) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1139 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9Y592 | Gene names | CCDC41 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 41 (NY-REN-58 antigen). | |||||
|
FLIP1_MOUSE
|
||||||
| NC score | 0.006192 (rank : 29) | θ value | 8.99809 (rank : 27) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1176 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9CS72 | Gene names | Filip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filamin-A-interacting protein 1 (FILIP). | |||||
|
MYH11_MOUSE
|
||||||
| NC score | 0.005571 (rank : 30) | θ value | 5.27518 (rank : 23) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1777 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O08638, O08639, Q62462, Q64195 | Gene names | Myh11 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-11 (Myosin heavy chain, smooth muscle isoform) (SMMHC). | |||||
|
LAMB1_HUMAN
|
||||||
| NC score | 0.004376 (rank : 31) | θ value | 8.99809 (rank : 28) | |||
| Query Neighborhood Hits | 31 | Target Neighborhood Hits | 1018 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P07942 | Gene names | LAMB1 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-1 chain precursor (Laminin B1 chain). | |||||