Please be patient as the page loads
|
HM20B_HUMAN
|
||||||
| SwissProt Accessions | Q9P0W2, Q6IBP8, Q8NBD5, Q9HD21, Q9Y491, Q9Y4A2 | Gene names | HMG20B, BRAF35, HMGX2, SMARCE1R | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily E member 1-related (SMARCE1-related protein) (HMG domain protein HMG20B) (Structural DNA-binding protein BRAF35) (BRCA2- associated factor 35) (Sox-like transcriptional factor). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
HM20B_HUMAN
|
||||||
| θ value | 5.27232e-117 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 126 | |
| SwissProt Accessions | Q9P0W2, Q6IBP8, Q8NBD5, Q9HD21, Q9Y491, Q9Y4A2 | Gene names | HMG20B, BRAF35, HMGX2, SMARCE1R | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily E member 1-related (SMARCE1-related protein) (HMG domain protein HMG20B) (Structural DNA-binding protein BRAF35) (BRCA2- associated factor 35) (Sox-like transcriptional factor). | |||||
|
HM20B_MOUSE
|
||||||
| θ value | 1.17455e-116 (rank : 2) | NC score | 0.994391 (rank : 2) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 111 | |
| SwissProt Accessions | Q9Z104, Q9NSF7, Q9NSF8 | Gene names | Hmg20b, Braf35, Hmgx2, Smarce1r | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily E member 1-related (SMARCE1-related protein) (HMG domain protein HMG20B) (Structural DNA-binding protein BRAF35) (BRCA2- associated factor 35). | |||||
|
HM20A_HUMAN
|
||||||
| θ value | 4.67263e-73 (rank : 3) | NC score | 0.888577 (rank : 3) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 91 | |
| SwissProt Accessions | Q9NP66, Q53G31, Q9NSF6 | Gene names | HMG20A, HMGX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | High mobility group protein 20A (HMG box-containing protein 20A) (HMG domain-containing protein HMGX1). | |||||
|
HM20A_MOUSE
|
||||||
| θ value | 3.02872e-72 (rank : 4) | NC score | 0.885809 (rank : 4) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 348 | Shared Neighborhood Hits | 91 | |
| SwissProt Accessions | Q9DC33, Q3LSF9, Q6NV87, Q8BSK1, Q8C3C1, Q8CAA0, Q9CYG2 | Gene names | Hmg20a, Ibraf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | High mobility group protein 20A (HMG box-containing protein 20A) (iBRAF) (Inhibitor of BRAF35) (HMG domain protein HMGX1). | |||||
|
SMCE1_HUMAN
|
||||||
| θ value | 3.29651e-10 (rank : 5) | NC score | 0.588235 (rank : 25) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 272 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | Q969G3, O43539 | Gene names | SMARCE1, BAF57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily E member 1 (BRG1-associated factor 57). | |||||
|
SMCE1_MOUSE
|
||||||
| θ value | 3.29651e-10 (rank : 6) | NC score | 0.555227 (rank : 48) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | O54941, Q8BPD9 | Gene names | Smarce1, Baf57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator chromatin subfamily E member 1 (BRG1-associated factor 57). | |||||
|
HMGB2_HUMAN
|
||||||
| θ value | 6.21693e-09 (rank : 7) | NC score | 0.683952 (rank : 7) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | P26583 | Gene names | HMGB2, HMG2 | |||
|
Domain Architecture |
|
|||||
| Description | High mobility group protein B2 (High mobility group protein 2) (HMG- 2). | |||||
|
HMGB3_HUMAN
|
||||||
| θ value | 8.11959e-09 (rank : 8) | NC score | 0.699913 (rank : 6) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | O15347, O95556, Q6NS40 | Gene names | HMGB3, HMG2A, HMG4 | |||
|
Domain Architecture |
|
|||||
| Description | High mobility group protein B3 (High mobility group protein 4) (HMG-4) (High mobility group protein 2a) (HMG-2a). | |||||
|
HMGB2_MOUSE
|
||||||
| θ value | 1.06045e-08 (rank : 9) | NC score | 0.678617 (rank : 8) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 77 | |
| SwissProt Accessions | P30681, Q3UXT1, Q9EQD5 | Gene names | Hmgb2, Hmg2 | |||
|
Domain Architecture |
|
|||||
| Description | High mobility group protein B2 (High mobility group protein 2) (HMG- 2). | |||||
|
TOX_MOUSE
|
||||||
| θ value | 1.06045e-08 (rank : 10) | NC score | 0.564673 (rank : 39) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q66JW3, Q8BKH9, Q8BYQ5, Q8R4H0 | Gene names | Tox | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thymus high mobility group box protein TOX. | |||||
|
HMGB3_MOUSE
|
||||||
| θ value | 1.38499e-08 (rank : 11) | NC score | 0.699929 (rank : 5) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | O54879 | Gene names | Hmgb3, Hmg2a, Hmg4 | |||
|
Domain Architecture |
|
|||||
| Description | High mobility group protein B3 (High mobility group protein 4) (HMG-4) (High mobility group protein 2a) (HMG-2a). | |||||
|
TOX_HUMAN
|
||||||
| θ value | 1.80886e-08 (rank : 12) | NC score | 0.562286 (rank : 41) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | O94900, Q96AV5 | Gene names | TOX, KIAA0808 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thymus high mobility group box protein TOX. | |||||
|
CN092_HUMAN
|
||||||
| θ value | 3.08544e-08 (rank : 13) | NC score | 0.569718 (rank : 34) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | O94842 | Gene names | C14orf92, KIAA0737 | |||
|
Domain Architecture |

|
|||||
| Description | Epidermal Langerhans cell protein LCP1. | |||||
|
CN092_MOUSE
|
||||||
| θ value | 3.08544e-08 (rank : 14) | NC score | 0.559636 (rank : 42) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q8BU11, Q80UI2, Q99PN9, Q9CS16 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | Epidermal Langerhans cell protein LCP1. | |||||
|
HMGB1_HUMAN
|
||||||
| θ value | 5.26297e-08 (rank : 15) | NC score | 0.671960 (rank : 10) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | P09429, Q6IBE1 | Gene names | HMGB1, HMG1 | |||
|
Domain Architecture |
|
|||||
| Description | High mobility group protein B1 (High mobility group protein 1) (HMG- 1). | |||||
|
HMGB1_MOUSE
|
||||||
| θ value | 5.26297e-08 (rank : 16) | NC score | 0.672519 (rank : 9) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | P63158, P07155, P27109, P27428 | Gene names | Hmgb1, Hmg-1, Hmg1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | High mobility group protein B1 (High mobility group protein 1) (HMG- 1). | |||||
|
UBF1_HUMAN
|
||||||
| θ value | 5.26297e-08 (rank : 17) | NC score | 0.567110 (rank : 37) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 415 | Shared Neighborhood Hits | 86 | |
| SwissProt Accessions | P17480 | Gene names | UBTF, UBF, UBF1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar transcription factor 1 (Upstream-binding factor 1) (UBF-1) (Autoantigen NOR-90). | |||||
|
UBF1_MOUSE
|
||||||
| θ value | 5.26297e-08 (rank : 18) | NC score | 0.558850 (rank : 43) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 86 | |
| SwissProt Accessions | P25976 | Gene names | Ubtf, Tcfubf, Ubf-1, Ubf1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar transcription factor 1 (Upstream-binding factor 1) (UBF-1). | |||||
|
SSRP1_MOUSE
|
||||||
| θ value | 8.97725e-08 (rank : 19) | NC score | 0.644507 (rank : 11) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | Q08943, Q3U9Z2, Q3UJ75, Q4V9U4, Q8CGA6 | Gene names | Ssrp1 | |||
|
Domain Architecture |

|
|||||
| Description | FACT complex subunit SSRP1 (Facilitates chromatin transcription complex subunit SSRP1) (Structure-specific recognition protein 1) (Recombination signal sequence recognition protein 1) (T160). | |||||
|
PB1_HUMAN
|
||||||
| θ value | 1.53129e-07 (rank : 20) | NC score | 0.394359 (rank : 73) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 261 | Shared Neighborhood Hits | 80 | |
| SwissProt Accessions | Q86U86, Q9H2T3, Q9H2T4, Q9H2T5, Q9H301, Q9H314 | Gene names | PB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein polybromo-1 (hPB1) (Polybromo-1D) (BRG1-associated factor 180) (BAF180). | |||||
|
SSRP1_HUMAN
|
||||||
| θ value | 1.53129e-07 (rank : 21) | NC score | 0.643644 (rank : 12) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | Q08945, Q5BJG8 | Gene names | SSRP1, FACT80 | |||
|
Domain Architecture |

|
|||||
| Description | FACT complex subunit SSRP1 (Facilitates chromatin transcription complex subunit SSRP1) (FACT 80 kDa subunit) (FACTp80) (Chromatin- specific transcription elongation factor 80 kDa subunit) (Structure- specific recognition protein 1) (hSSRP1) (Recombination signal sequence recognition protein 1) (T160). | |||||
|
SP100_HUMAN
|
||||||
| θ value | 1.99992e-07 (rank : 22) | NC score | 0.493442 (rank : 64) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | P23497, O75450, Q13343, Q96F70, Q96T24, Q96T95, Q9NP33, Q9UE32 | Gene names | SP100 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear autoantigen Sp-100 (Speckled 100 kDa) (Nuclear dot-associated Sp100 protein) (Lysp100b). | |||||
|
SOX7_HUMAN
|
||||||
| θ value | 5.81887e-07 (rank : 23) | NC score | 0.601826 (rank : 18) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 78 | |
| SwissProt Accessions | Q9BT81 | Gene names | SOX7 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor SOX-7. | |||||
|
SOX7_MOUSE
|
||||||
| θ value | 7.59969e-07 (rank : 24) | NC score | 0.605177 (rank : 17) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 78 | |
| SwissProt Accessions | P40646, Q9R1T6 | Gene names | Sox7, Sox-7 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-7 (mSOX7). | |||||
|
SOX14_HUMAN
|
||||||
| θ value | 1.29631e-06 (rank : 25) | NC score | 0.605928 (rank : 16) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | O95416, Q3KPH7 | Gene names | SOX14, SOX28 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-14. | |||||
|
SOX14_MOUSE
|
||||||
| θ value | 1.29631e-06 (rank : 26) | NC score | 0.605998 (rank : 15) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | Q04892, Q9JLC9, Q9JLD0, Q9QXU4 | Gene names | Sox14, Sox-14 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-14. | |||||
|
SOX17_MOUSE
|
||||||
| θ value | 1.29631e-06 (rank : 27) | NC score | 0.584013 (rank : 27) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | Q61473, Q61472, Q62248 | Gene names | Sox17, Sox-17 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-17. | |||||
|
SOX2_HUMAN
|
||||||
| θ value | 1.29631e-06 (rank : 28) | NC score | 0.592516 (rank : 20) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | P48431, Q14537 | Gene names | SOX2 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor SOX-2. | |||||
|
SOX2_MOUSE
|
||||||
| θ value | 1.29631e-06 (rank : 29) | NC score | 0.592602 (rank : 19) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | P48432 | Gene names | Sox2, Sox-2 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor SOX-2. | |||||
|
HMG1X_HUMAN
|
||||||
| θ value | 2.21117e-06 (rank : 30) | NC score | 0.642862 (rank : 13) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q9UGV6 | Gene names | HMG1L10 | |||
|
Domain Architecture |
|
|||||
| Description | High mobility group protein 1-like 10 (HMG-1L10). | |||||
|
TFAM_HUMAN
|
||||||
| θ value | 3.77169e-06 (rank : 31) | NC score | 0.568002 (rank : 36) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q00059 | Gene names | TFAM, TCF6L2 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor A, mitochondrial precursor (mtTFA) (Mitochondrial transcription factor 1) (MtTF1) (Transcription factor 6-like 2). | |||||
|
SOX17_HUMAN
|
||||||
| θ value | 1.09739e-05 (rank : 32) | NC score | 0.582435 (rank : 28) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | Q9H6I2 | Gene names | SOX17 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor SOX-17. | |||||
|
SOX21_HUMAN
|
||||||
| θ value | 1.09739e-05 (rank : 33) | NC score | 0.591673 (rank : 21) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 78 | |
| SwissProt Accessions | Q9Y651, P35715, Q15504 | Gene names | SOX21, SOX25, SOXA | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-21 (SOX-A). | |||||
|
SOX21_MOUSE
|
||||||
| θ value | 1.09739e-05 (rank : 34) | NC score | 0.591671 (rank : 22) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 78 | |
| SwissProt Accessions | Q811W0, Q8VH35 | Gene names | Sox21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor SOX-21. | |||||
|
SOX8_MOUSE
|
||||||
| θ value | 1.43324e-05 (rank : 35) | NC score | 0.548552 (rank : 50) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q04886 | Gene names | Sox8, Sox-8 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-8. | |||||
|
TFAM_MOUSE
|
||||||
| θ value | 1.87187e-05 (rank : 36) | NC score | 0.612231 (rank : 14) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | P40630, P97894, P97906, Q543I8, Q9DBM9 | Gene names | Tfam, Hmgts | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor A, mitochondrial precursor (mtTFA) (Testis- specific high mobility group protein) (TS-HMG). | |||||
|
BBX_HUMAN
|
||||||
| θ value | 2.44474e-05 (rank : 37) | NC score | 0.528037 (rank : 53) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q8WY36, Q2TAJ1, Q7L3J8, Q7LBY8, Q8NDB0, Q8WY35, Q9H0J6 | Gene names | BBX, HBP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HMG box transcription factor BBX (Bobby sox homolog) (HMG box- containing protein 2). | |||||
|
BBX_MOUSE
|
||||||
| θ value | 2.44474e-05 (rank : 38) | NC score | 0.536765 (rank : 52) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | Q8VBW5, Q3TZK1, Q6NXY8, Q6PEU3, Q8BQJ7, Q8C7E0, Q8CDQ0, Q8CDV1, Q8VI48, Q8VI49, Q8VI50, Q9CS94 | Gene names | Bbx, Hbp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HMG box transcription factor BBX (Bobby sox homolog) (HMG box- containing protein 2). | |||||
|
SOX18_HUMAN
|
||||||
| θ value | 4.1701e-05 (rank : 39) | NC score | 0.586701 (rank : 26) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | P35713, Q9NPH8 | Gene names | SOX18 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-18. | |||||
|
SOX18_MOUSE
|
||||||
| θ value | 4.1701e-05 (rank : 40) | NC score | 0.582126 (rank : 29) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | P43680, Q9EQ73 | Gene names | Sox18, Sox-18 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-18. | |||||
|
SOX1_HUMAN
|
||||||
| θ value | 4.1701e-05 (rank : 41) | NC score | 0.579709 (rank : 31) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | O00570 | Gene names | SOX1 | |||
|
Domain Architecture |
|
|||||
| Description | SOX-1 protein. | |||||
|
SOX1_MOUSE
|
||||||
| θ value | 4.1701e-05 (rank : 42) | NC score | 0.579348 (rank : 32) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | P53783 | Gene names | Sox1, Sox-1 | |||
|
Domain Architecture |
|
|||||
| Description | SOX-1 protein. | |||||
|
SOX3_HUMAN
|
||||||
| θ value | 4.1701e-05 (rank : 43) | NC score | 0.576648 (rank : 33) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 78 | |
| SwissProt Accessions | P41225, P35714, Q9NP49 | Gene names | SOX3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-3. | |||||
|
SOX3_MOUSE
|
||||||
| θ value | 4.1701e-05 (rank : 44) | NC score | 0.579738 (rank : 30) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | P53784 | Gene names | Sox3, Sox-3 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor SOX-3. | |||||
|
SOX8_HUMAN
|
||||||
| θ value | 9.29e-05 (rank : 45) | NC score | 0.548427 (rank : 51) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | P57073, Q9NZW2 | Gene names | SOX8 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-8. | |||||
|
SOX15_HUMAN
|
||||||
| θ value | 0.000121331 (rank : 46) | NC score | 0.589441 (rank : 23) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | O60248, P35717, Q9Y6W7 | Gene names | SOX15, SOX12, SOX20, SOX26, SOX27 | |||
|
Domain Architecture |

|
|||||
| Description | SOX-15 protein (SOX-20 protein) (SOX-12 protein). | |||||
|
CIC_HUMAN
|
||||||
| θ value | 0.00020696 (rank : 47) | NC score | 0.453363 (rank : 69) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 992 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | Q96RK0, Q7LGI1, Q9UEG5, Q9Y6T1 | Gene names | CIC, KIAA0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
CIC_MOUSE
|
||||||
| θ value | 0.00020696 (rank : 48) | NC score | 0.482185 (rank : 65) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 742 | Shared Neighborhood Hits | 78 | |
| SwissProt Accessions | Q924A2, Q6PDJ8, Q8CGE4, Q8CHH0, Q9CW61 | Gene names | Cic, Kiaa0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
SOX12_HUMAN
|
||||||
| θ value | 0.000270298 (rank : 49) | NC score | 0.564390 (rank : 40) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | O15370, Q9NUD4 | Gene names | SOX12, SOX22 | |||
|
Domain Architecture |
|
|||||
| Description | SOX-12 protein (SOX-22 protein). | |||||
|
SOX12_MOUSE
|
||||||
| θ value | 0.000270298 (rank : 50) | NC score | 0.565213 (rank : 38) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q04890, P70417, Q6NXL2 | Gene names | Sox12, Sox-12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SOX-12 protein. | |||||
|
SOX15_MOUSE
|
||||||
| θ value | 0.000270298 (rank : 51) | NC score | 0.589049 (rank : 24) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | P43267, O70204, P70418, Q62246, Q91V00, Q91V43, Q920T1, Q9JLG2 | Gene names | Sox15, Sox-15 | |||
|
Domain Architecture |

|
|||||
| Description | SOX-15 protein. | |||||
|
GCX1_HUMAN
|
||||||
| θ value | 0.000461057 (rank : 52) | NC score | 0.471189 (rank : 66) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q96NM4, Q5TE33, Q5TE34, Q5TE35, Q96IC9, Q9BQN5 | Gene names | GCX1, C20orf100 | |||
|
Domain Architecture |

|
|||||
| Description | Granulosa cell HMG box protein 1 (GCX-1). | |||||
|
SOX4_HUMAN
|
||||||
| θ value | 0.000461057 (rank : 53) | NC score | 0.555869 (rank : 47) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 77 | |
| SwissProt Accessions | Q06945 | Gene names | SOX4 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor SOX-4. | |||||
|
SOX13_HUMAN
|
||||||
| θ value | 0.000602161 (rank : 54) | NC score | 0.504556 (rank : 61) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 464 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | Q9UN79, O95275, O95826, Q9UHW7 | Gene names | SOX13 | |||
|
Domain Architecture |

|
|||||
| Description | SOX-13 protein (Type 1 diabetes autoantigen ICA12) (Islet cell antigen 12). | |||||
|
SOX13_MOUSE
|
||||||
| θ value | 0.000602161 (rank : 55) | NC score | 0.507390 (rank : 60) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 495 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | Q04891, Q922L3 | Gene names | Sox13, Sox-13 | |||
|
Domain Architecture |

|
|||||
| Description | SOX-13 protein. | |||||
|
SOX9_HUMAN
|
||||||
| θ value | 0.000602161 (rank : 56) | NC score | 0.522775 (rank : 55) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | P48436 | Gene names | SOX9 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor SOX-9. | |||||
|
SOX9_MOUSE
|
||||||
| θ value | 0.000602161 (rank : 57) | NC score | 0.523716 (rank : 54) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q04887, Q8C7L2, Q91ZK2, Q99KQ0 | Gene names | Sox9, Sox-9 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-9. | |||||
|
SOX11_HUMAN
|
||||||
| θ value | 0.000786445 (rank : 58) | NC score | 0.558420 (rank : 44) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | P35716 | Gene names | SOX11 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-11. | |||||
|
SOX11_MOUSE
|
||||||
| θ value | 0.000786445 (rank : 59) | NC score | 0.557644 (rank : 45) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 77 | |
| SwissProt Accessions | Q7M6Y2, O35178, O89036, Q04889, Q80XF0 | Gene names | Sox11, Sox-11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor SOX-11. | |||||
|
SOX5_HUMAN
|
||||||
| θ value | 0.000786445 (rank : 60) | NC score | 0.511260 (rank : 59) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 85 | |
| SwissProt Accessions | P35711, Q86UK8, Q8J017, Q8J018, Q8J019, Q8J020, Q8N1D9, Q8N7E0, Q8TEA4 | Gene names | SOX5 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-5. | |||||
|
SOX5_MOUSE
|
||||||
| θ value | 0.000786445 (rank : 61) | NC score | 0.512428 (rank : 58) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 86 | |
| SwissProt Accessions | P35710, O88184, O89018 | Gene names | Sox5, Sox-5 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-5. | |||||
|
SOX4_MOUSE
|
||||||
| θ value | 0.00134147 (rank : 62) | NC score | 0.555221 (rank : 49) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 77 | |
| SwissProt Accessions | Q06831 | Gene names | Sox4, Sox-4 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor SOX-4. | |||||
|
SOX30_HUMAN
|
||||||
| θ value | 0.00298849 (rank : 63) | NC score | 0.515104 (rank : 57) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 77 | |
| SwissProt Accessions | O94993, O94995, Q8IYX6 | Gene names | SOX30 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-30. | |||||
|
SOX30_MOUSE
|
||||||
| θ value | 0.00298849 (rank : 64) | NC score | 0.518149 (rank : 56) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 77 | |
| SwissProt Accessions | Q8CGW4 | Gene names | Sox30 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-30. | |||||
|
SOX6_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 65) | NC score | 0.500000 (rank : 62) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 96 | |
| SwissProt Accessions | P35712, Q9BXQ3, Q9BXQ4, Q9BXQ5, Q9H0I8 | Gene names | SOX6 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-6. | |||||
|
SOX6_MOUSE
|
||||||
| θ value | 0.00390308 (rank : 66) | NC score | 0.498831 (rank : 63) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 96 | |
| SwissProt Accessions | P40645, Q62250, Q9QWS5 | Gene names | Sox6, Sox-6 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-6 (SOX-LZ). | |||||
|
SRY_MOUSE
|
||||||
| θ value | 0.00869519 (rank : 67) | NC score | 0.557479 (rank : 46) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | Q05738 | Gene names | Sry, Tdf, Tdy | |||
|
Domain Architecture |

|
|||||
| Description | Sex-determining region Y protein (Testis-determining factor). | |||||
|
ST18_MOUSE
|
||||||
| θ value | 0.00869519 (rank : 68) | NC score | 0.076559 (rank : 83) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q80TY4, Q3UH00, Q3URH9, Q3UVB9, Q3UZN9, Q811B4, Q8K098 | Gene names | St18, Kiaa0535 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppression of tumorigenicity protein 18. | |||||
|
ST18_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 69) | NC score | 0.075575 (rank : 84) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 252 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O60284 | Gene names | ST18, KIAA0535, ZNF387 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppression of tumorigenicity protein 18 (Zinc finger protein 387). | |||||
|
SRY_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 70) | NC score | 0.568453 (rank : 35) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | Q05066 | Gene names | SRY, TDF | |||
|
Domain Architecture |
|
|||||
| Description | Sex-determining region Y protein (Testis-determining factor). | |||||
|
LEF1_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 71) | NC score | 0.364272 (rank : 75) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q9UJU2, Q9HAZ0 | Gene names | LEF1 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphoid enhancer-binding factor 1 (LEF-1) (T cell-specific transcription factor 1-alpha) (TCF1-alpha). | |||||
|
LEF1_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 72) | NC score | 0.363913 (rank : 76) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | P27782 | Gene names | Lef1, Lef-1 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphoid enhancer-binding factor 1 (LEF-1). | |||||
|
TCF7_HUMAN
|
||||||
| θ value | 0.21417 (rank : 73) | NC score | 0.386693 (rank : 74) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 186 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | P36402, Q9UKI4 | Gene names | TCF7, TCF1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 7 (T-cell-specific transcription factor 1) (TCF- 1) (T-cell factor 1). | |||||
|
CP250_HUMAN
|
||||||
| θ value | 0.279714 (rank : 74) | NC score | 0.036115 (rank : 92) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1805 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q9BV73, O14812, O60588, Q9H450 | Gene names | CEP250, CEP2, CNAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Centrosome-associated protein CEP250 (Centrosomal protein 2) (Centrosomal Nek2-associated protein 1) (C-Nap1). | |||||
|
CAN3_MOUSE
|
||||||
| θ value | 0.47712 (rank : 75) | NC score | 0.025885 (rank : 107) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q64691, Q9WUC5 | Gene names | Capn3 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-3 (EC 3.4.22.-) (Calpain L3) (Calpain p94) (Calcium-activated neutral proteinase 3) (CANP 3) (Muscle-specific calcium-activated neutral protease 3). | |||||
|
CF060_HUMAN
|
||||||
| θ value | 0.47712 (rank : 76) | NC score | 0.039439 (rank : 89) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1270 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q8NB25, Q96GY8, Q9H851 | Gene names | C6orf60 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C6orf60. | |||||
|
LIPA1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 77) | NC score | 0.035276 (rank : 94) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1294 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q13136, Q13135, Q14567, Q8N4I2 | Gene names | PPFIA1, LIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-1 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-1) (PTPRF-interacting protein alpha-1) (LAR-interacting protein 1) (LIP.1). | |||||
|
REST_HUMAN
|
||||||
| θ value | 0.47712 (rank : 78) | NC score | 0.035182 (rank : 95) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1605 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P30622 | Gene names | RSN, CYLN1 | |||
|
Domain Architecture |

|
|||||
| Description | Restin (Cytoplasmic linker protein 170 alpha-2) (CLIP-170) (Reed- Sternberg intermediate filament-associated protein) (Cytoplasmic linker protein 1). | |||||
|
CROCC_MOUSE
|
||||||
| θ value | 0.62314 (rank : 79) | NC score | 0.034050 (rank : 96) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1460 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8CJ40, Q7TQL2, Q80U01, Q8R0B9 | Gene names | Crocc, Kiaa0445 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rootletin (Ciliary rootlet coiled-coil protein). | |||||
|
GOGA1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 80) | NC score | 0.036560 (rank : 91) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1222 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q92805, Q8IYZ9 | Gene names | GOLGA1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 1 (Golgin-97). | |||||
|
HM2L1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 81) | NC score | 0.445396 (rank : 70) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9UGU5, O75672, O75673, Q9UMT5 | Gene names | HMG2L1, HMGBCG | |||
|
Domain Architecture |

|
|||||
| Description | High mobility group protein 2-like 1 (Protein HMGBCG). | |||||
|
TCF7_MOUSE
|
||||||
| θ value | 0.62314 (rank : 82) | NC score | 0.359894 (rank : 77) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q00417 | Gene names | Tcf7, Tcf-1, Tcf1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 7 (T-cell-specific transcription factor 1) (TCF- 1) (T-cell factor 1). | |||||
|
TF7L2_HUMAN
|
||||||
| θ value | 0.62314 (rank : 83) | NC score | 0.346487 (rank : 79) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9NQB0, O00185, Q9NQB1, Q9NQB2, Q9NQB3, Q9NQB4, Q9NQB5, Q9NQB6, Q9NQB7, Q9ULC2 | Gene names | TCF7L2, TCF4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 7-like 2 (HMG box transcription factor 4) (T- cell-specific transcription factor 4) (TCF-4) (hTCF-4). | |||||
|
TF7L2_MOUSE
|
||||||
| θ value | 0.62314 (rank : 84) | NC score | 0.347678 (rank : 78) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q924A0, O70574, Q91XP2, Q91XP3, Q91XP4, Q924A1, Q9Z0V3, Q9Z0V4 | Gene names | Tcf7l2, Tcf4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 7-like 2 (HMG box transcription factor 4) (T- cell-specific transcription factor 4) (TCF-4) (mTCF-4). | |||||
|
GABP2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 85) | NC score | 0.014564 (rank : 117) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 454 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q06547, Q06545, Q12940, Q12941, Q12942, Q8IYD0 | Gene names | GABPB2, E4TF1B, GABPB, GABPB1 | |||
|
Domain Architecture |
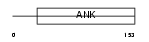
|
|||||
| Description | GA-binding protein beta chain (GABP subunit beta-2) (GABP-2) (GABP subunit beta-1) (GABPB-1) (Transcription factor E4TF1-53) (Transcription factor E4TF1-47) (Nuclear respiratory factor 2). | |||||
|
CK5P2_HUMAN
|
||||||
| θ value | 1.06291 (rank : 86) | NC score | 0.029231 (rank : 101) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1433 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q96SN8, Q7Z3L4, Q7Z3U1, Q7Z7I6, Q9BSW0, Q9H6J6, Q9HCD9, Q9NV90, Q9UIW9 | Gene names | CDK5RAP2, CEP215, KIAA1633 | |||
|
Domain Architecture |

|
|||||
| Description | CDK5 regulatory subunit-associated protein 2 (CDK5 activator-binding protein C48) (Centrosome-associated protein 215). | |||||
|
GABP2_MOUSE
|
||||||
| θ value | 1.06291 (rank : 87) | NC score | 0.014395 (rank : 118) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 429 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q00420, Q00421, Q8BS31, Q91YZ0, Q9QVV2 | Gene names | Gabpb2, Gabpb, Gabpb1 | |||
|
Domain Architecture |
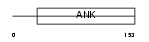
|
|||||
| Description | GA-binding protein beta chain (GABP subunit beta-2) (GABP-2) (GABP subunit beta-1) (GABPB-1). | |||||
|
GOGA1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 88) | NC score | 0.033348 (rank : 97) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 994 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9CW79, Q80YB0 | Gene names | Golga1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 1 (Golgin-97). | |||||
|
HBP1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 89) | NC score | 0.409471 (rank : 71) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | O60381, Q8TBM1, Q8TE93, Q96AJ2 | Gene names | HBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HMG box-containing protein 1 (HMG box transcription factor 1) (High mobility group box transcription factor 1). | |||||
|
HBP1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 90) | NC score | 0.409116 (rank : 72) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q8R316, Q3V0I4, Q8BUS3, Q8C199 | Gene names | Hbp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HMG box-containing protein 1 (HMG box transcription factor 1) (High mobility group box transcription factor 1). | |||||
|
NEMO_HUMAN
|
||||||
| θ value | 1.06291 (rank : 91) | NC score | 0.032393 (rank : 98) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 820 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9Y6K9 | Gene names | IKBKG, FIP3, NEMO | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NF-kappa-B essential modulator (NEMO) (NF-kappa-B essential modifier) (Inhibitor of nuclear factor kappa-B kinase subunit gamma) (IkB kinase subunit gamma) (I-kappa-B kinase gamma) (IKK-gamma) (IKKG) (IkB kinase-associated protein 1) (IKKAP1) (FIP-3). | |||||
|
KIF14_HUMAN
|
||||||
| θ value | 1.38821 (rank : 92) | NC score | 0.016983 (rank : 116) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 732 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q15058 | Gene names | KIF14, KIAA0042 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin-like protein KIF14. | |||||
|
RC3H1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 93) | NC score | 0.040513 (rank : 88) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 244 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q4VGL6, Q69Z31 | Gene names | Rc3h1, Gm551, Kiaa2025 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Roquin (RING finger and C3H zinc finger protein 1) (Sanroque protein). | |||||
|
LAMB3_HUMAN
|
||||||
| θ value | 1.81305 (rank : 94) | NC score | 0.010379 (rank : 126) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 461 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q13751, O14947, Q14733, Q9UJK4, Q9UJL1 | Gene names | LAMB3 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-3 chain precursor (Laminin 5 beta 3) (Laminin B1k chain) (Kalinin B1 chain). | |||||
|
TF7L1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 95) | NC score | 0.327175 (rank : 81) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q9HCS4, Q53R97, Q6PD70, Q9NP00 | Gene names | TCF7L1, TCF3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 7-like 1 (HMG box transcription factor 3) (TCF- 3). | |||||
|
TF7L1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 96) | NC score | 0.327654 (rank : 80) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q9Z1J1, O70450, O70573 | Gene names | Tcf7l1, Tcf3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 7-like 1 (HMG box transcription factor 3) (TCF-3) (mTCF-3). | |||||
|
K0555_HUMAN
|
||||||
| θ value | 2.36792 (rank : 97) | NC score | 0.035931 (rank : 93) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 880 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q96AA8, O60302 | Gene names | KIAA0555 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0555. | |||||
|
KIF23_HUMAN
|
||||||
| θ value | 2.36792 (rank : 98) | NC score | 0.017511 (rank : 114) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 618 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q02241 | Gene names | KIF23, KNSL5, MKLP1 | |||
|
Domain Architecture |
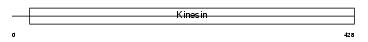
|
|||||
| Description | Kinesin-like protein KIF23 (Mitotic kinesin-like protein 1) (Kinesin- like protein 5). | |||||
|
REST_MOUSE
|
||||||
| θ value | 2.36792 (rank : 99) | NC score | 0.037456 (rank : 90) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1506 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q922J3, Q571L7 | Gene names | Rsn, Kiaa4046 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Restin. | |||||
|
TRI38_HUMAN
|
||||||
| θ value | 2.36792 (rank : 100) | NC score | 0.009593 (rank : 130) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 373 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O00635 | Gene names | TRIM38, RNF15, RORET | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 38 (RING finger protein 15) (Zinc finger protein RoRet). | |||||
|
NIN_MOUSE
|
||||||
| θ value | 3.0926 (rank : 101) | NC score | 0.025455 (rank : 108) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1376 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q61043, Q6ZPM7 | Gene names | Nin, Kiaa1565 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ninein. | |||||
|
TRI41_HUMAN
|
||||||
| θ value | 3.0926 (rank : 102) | NC score | 0.009915 (rank : 129) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8WV44, Q5BKT0, Q7L484, Q96Q10, Q9BSL8 | Gene names | TRIM41 | |||
|
Domain Architecture |
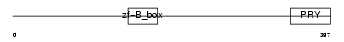
|
|||||
| Description | Tripartite motif-containing protein 41. | |||||
|
CLC4F_MOUSE
|
||||||
| θ value | 4.03905 (rank : 103) | NC score | 0.017362 (rank : 115) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 572 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P70194, Q8BLZ8 | Gene names | Clec4f, Clecsf13, Kclr | |||
|
Domain Architecture |

|
|||||
| Description | C-type lectin domain family 4 member F (C-type lectin superfamily member 13) (C-type lectin 13) (Kupffer cell receptor). | |||||
|
GOGA3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 104) | NC score | 0.026989 (rank : 104) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1317 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q08378, O43241, Q6P9C7, Q86XW3, Q8TDA9, Q8WZA3 | Gene names | GOLGA3 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 3 (Golgin-160) (Golgi complex-associated protein of 170 kDa) (GCP170). | |||||
|
LRSM1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 105) | NC score | 0.013649 (rank : 119) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1078 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q6UWE0, Q5VVV0, Q8NB40, Q96GT5, Q96MX5, Q96MZ7 | Gene names | LRSAM1, TAL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin ligase protein LRSAM1 (EC 6.3.2.-) (Leucine-rich repeat and sterile alpha motif-containing protein 1) (Tsg101-associated ligase) (hTAL). | |||||
|
TEX14_MOUSE
|
||||||
| θ value | 4.03905 (rank : 106) | NC score | 0.003835 (rank : 131) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1333 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q7M6U3, Q3UT36, Q5NC10, Q5NC11, Q8CGK1, Q8CGK2, Q99MV8 | Gene names | Tex14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Testis-expressed protein 14 (Testis-expressed sequence 14). | |||||
|
UBN1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 107) | NC score | 0.064338 (rank : 87) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NPG3, Q13079, Q9P1P7 | Gene names | UBN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubinuclein (Ubiquitously expressed nuclear protein) (VT4). | |||||
|
CROCC_HUMAN
|
||||||
| θ value | 5.27518 (rank : 108) | NC score | 0.026790 (rank : 105) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1409 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q5TZA2, Q2VHY3, Q66GT7, Q7Z2L4, Q7Z5D7 | Gene names | CROCC, KIAA0445 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rootletin (Ciliary rootlet coiled-coil protein). | |||||
|
IQGA2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 109) | NC score | 0.010890 (rank : 125) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q13576 | Gene names | IQGAP2 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP2. | |||||
|
MORF4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 110) | NC score | 0.010006 (rank : 128) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y690 | Gene names | MORF4 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor-like protein MORF4 (Mortality factor 4) (Cellular senescence-related protein 1) (SEN1). | |||||
|
POT15_HUMAN
|
||||||
| θ value | 5.27518 (rank : 111) | NC score | 0.011478 (rank : 123) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 893 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q6S5H4, Q6NXN7, Q6S5H7 | Gene names | POTE15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prostate, ovary, testis-expressed protein on chromosome 15. | |||||
|
SPT16_HUMAN
|
||||||
| θ value | 5.27518 (rank : 112) | NC score | 0.021014 (rank : 111) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Y5B9, Q6GMT8, Q6P2F1, Q6PJM1, Q9NRX0 | Gene names | SUPT16H, FACT140, FACTP140 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FACT complex subunit SPT16 (Facilitates chromatin transcription complex subunit SPT16) (hSPT16) (FACT 140 kDa subunit) (FACTp140) (Chromatin-specific transcription elongation factor 140 kDa subunit). | |||||
|
SPT16_MOUSE
|
||||||
| θ value | 5.27518 (rank : 113) | NC score | 0.021362 (rank : 110) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q920B9, Q3TZ48, Q3UFH0, Q921H4 | Gene names | Supt16h, Fact140, Factp140 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FACT complex subunit SPT16 (Facilitates chromatin transcription complex subunit SPT16) (FACT 140 kDa subunit) (FACTp140) (Chromatin- specific transcription elongation factor 140 kDa subunit). | |||||
|
ANR21_HUMAN
|
||||||
| θ value | 6.88961 (rank : 114) | NC score | 0.011109 (rank : 124) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 864 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q86YR6 | Gene names | ANKRD21, POTE | |||
|
Domain Architecture |
|
|||||
| Description | Ankyrin repeat domain-containing protein 21 (Protein POTE) (Prostate, ovary, testis-expressed protein). | |||||
|
NFH_MOUSE
|
||||||
| θ value | 6.88961 (rank : 115) | NC score | 0.010160 (rank : 127) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
RC3H1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 116) | NC score | 0.030652 (rank : 99) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 433 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q5TC82, Q5W180, Q5W181, Q8IVE6, Q8N9V1 | Gene names | RC3H1, KIAA2025 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Roquin (RING finger and C3H zinc finger protein 1). | |||||
|
TEX9_HUMAN
|
||||||
| θ value | 6.88961 (rank : 117) | NC score | 0.020577 (rank : 112) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 541 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8N6V9 | Gene names | TEX9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Testis-expressed sequence 9 protein. | |||||
|
TRI32_MOUSE
|
||||||
| θ value | 6.88961 (rank : 118) | NC score | 0.011906 (rank : 122) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8CH72, Q8K055 | Gene names | Trim32 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 32 (EC 6.3.2.-). | |||||
|
BPAEA_HUMAN
|
||||||
| θ value | 8.99809 (rank : 119) | NC score | 0.026485 (rank : 106) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1369 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | O94833, Q5TBT0, Q8N1T8, Q8N8J3, Q8WXK9, Q96AK9, Q96DQ5, Q9H555 | Gene names | DST, BPAG1, DMH, DT, KIAA0728 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoforms 6/9/10 (Trabeculin-beta) (Bullous pemphigoid antigen) (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
CCD41_HUMAN
|
||||||
| θ value | 8.99809 (rank : 120) | NC score | 0.022455 (rank : 109) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1139 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9Y592 | Gene names | CCDC41 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 41 (NY-REN-58 antigen). | |||||
|
CCD41_MOUSE
|
||||||
| θ value | 8.99809 (rank : 121) | NC score | 0.019587 (rank : 113) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9D5R3, Q3U7X7, Q3UX57, Q80VF0 | Gene names | Ccdc41 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 41. | |||||
|
GOPC_HUMAN
|
||||||
| θ value | 8.99809 (rank : 122) | NC score | 0.012876 (rank : 120) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 377 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9HD26, Q59FS4, Q969U8 | Gene names | GOPC, CAL, FIG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi-associated PDZ and coiled-coil motif-containing protein (PDZ protein interacting specifically with TC10) (PIST) (CFTR-associated ligand) (Fused in glioblastoma). | |||||
|
GOPC_MOUSE
|
||||||
| θ value | 8.99809 (rank : 123) | NC score | 0.012394 (rank : 121) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8BH60, Q8BSV4, Q920R1, Q9ET11 | Gene names | Gopc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi-associated PDZ and coiled-coil motif-containing protein (PDZ protein interacting specifically with TC10) (PIST). | |||||
|
LIPA2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 124) | NC score | 0.029090 (rank : 102) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1044 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O75334 | Gene names | PPFIA2 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-2 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-2) (PTPRF-interacting protein alpha-2). | |||||
|
LIPA2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 125) | NC score | 0.029606 (rank : 100) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1282 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8BSS9, Q8BN73 | Gene names | Ppfia2 | |||
|
Domain Architecture |
|
|||||
| Description | Liprin-alpha-2 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-2) (PTPRF-interacting protein alpha-2). | |||||
|
LIPA3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 126) | NC score | 0.028077 (rank : 103) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O75145, Q9H8B5, Q9UEW4 | Gene names | PPFIA3, KIAA0654 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-3 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-3) (PTPRF-interacting protein alpha-3). | |||||
|
MYT1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 127) | NC score | 0.069037 (rank : 85) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 475 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q01538, O94922, Q9UPV2 | Gene names | MYT1, KIAA0835, KIAA1050, MTF1, MYTI, PLPB1 | |||
|
Domain Architecture |

|
|||||
| Description | Myelin transcription factor 1 (MyT1) (MyTI) (Proteolipid protein- binding protein) (PLPB1). | |||||
|
MYT1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 128) | NC score | 0.065405 (rank : 86) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8CFC2, O08995, Q8CFH1 | Gene names | Myt1, Kiaa0835, Nzf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myelin transcription factor 1 (MyT1) (Neural zinc finger factor 2) (NZF-2). | |||||
|
PMS1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 129) | NC score | 0.263624 (rank : 82) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | P54277 | Gene names | PMS1, PMSL1 | |||
|
Domain Architecture |

|
|||||
| Description | PMS1 protein homolog 1 (DNA mismatch repair protein PMS1). | |||||
|
SOX10_HUMAN
|
||||||
| θ value | θ > 10 (rank : 130) | NC score | 0.471061 (rank : 67) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | P56693 | Gene names | SOX10 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor SOX-10. | |||||
|
SOX10_MOUSE
|
||||||
| θ value | θ > 10 (rank : 131) | NC score | 0.470910 (rank : 68) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q04888, O08518, O09141, O54856, P70416 | Gene names | Sox10, Sox-10, Sox21 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-10 (SOX-21) (Transcription factor SOX-M). | |||||
|
HM20B_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 5.27232e-117 (rank : 1) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 126 | |
| SwissProt Accessions | Q9P0W2, Q6IBP8, Q8NBD5, Q9HD21, Q9Y491, Q9Y4A2 | Gene names | HMG20B, BRAF35, HMGX2, SMARCE1R | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily E member 1-related (SMARCE1-related protein) (HMG domain protein HMG20B) (Structural DNA-binding protein BRAF35) (BRCA2- associated factor 35) (Sox-like transcriptional factor). | |||||
|
HM20B_MOUSE
|
||||||
| NC score | 0.994391 (rank : 2) | θ value | 1.17455e-116 (rank : 2) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 111 | |
| SwissProt Accessions | Q9Z104, Q9NSF7, Q9NSF8 | Gene names | Hmg20b, Braf35, Hmgx2, Smarce1r | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily E member 1-related (SMARCE1-related protein) (HMG domain protein HMG20B) (Structural DNA-binding protein BRAF35) (BRCA2- associated factor 35). | |||||
|
HM20A_HUMAN
|
||||||
| NC score | 0.888577 (rank : 3) | θ value | 4.67263e-73 (rank : 3) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 91 | |
| SwissProt Accessions | Q9NP66, Q53G31, Q9NSF6 | Gene names | HMG20A, HMGX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | High mobility group protein 20A (HMG box-containing protein 20A) (HMG domain-containing protein HMGX1). | |||||
|
HM20A_MOUSE
|
||||||
| NC score | 0.885809 (rank : 4) | θ value | 3.02872e-72 (rank : 4) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 348 | Shared Neighborhood Hits | 91 | |
| SwissProt Accessions | Q9DC33, Q3LSF9, Q6NV87, Q8BSK1, Q8C3C1, Q8CAA0, Q9CYG2 | Gene names | Hmg20a, Ibraf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | High mobility group protein 20A (HMG box-containing protein 20A) (iBRAF) (Inhibitor of BRAF35) (HMG domain protein HMGX1). | |||||
|
HMGB3_MOUSE
|
||||||
| NC score | 0.699929 (rank : 5) | θ value | 1.38499e-08 (rank : 11) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | O54879 | Gene names | Hmgb3, Hmg2a, Hmg4 | |||
|
Domain Architecture |

|
|||||
| Description | High mobility group protein B3 (High mobility group protein 4) (HMG-4) (High mobility group protein 2a) (HMG-2a). | |||||
|
HMGB3_HUMAN
|
||||||
| NC score | 0.699913 (rank : 6) | θ value | 8.11959e-09 (rank : 8) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | O15347, O95556, Q6NS40 | Gene names | HMGB3, HMG2A, HMG4 | |||
|
Domain Architecture |

|
|||||
| Description | High mobility group protein B3 (High mobility group protein 4) (HMG-4) (High mobility group protein 2a) (HMG-2a). | |||||
|
HMGB2_HUMAN
|
||||||
| NC score | 0.683952 (rank : 7) | θ value | 6.21693e-09 (rank : 7) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | P26583 | Gene names | HMGB2, HMG2 | |||
|
Domain Architecture |

|
|||||
| Description | High mobility group protein B2 (High mobility group protein 2) (HMG- 2). | |||||
|
HMGB2_MOUSE
|
||||||
| NC score | 0.678617 (rank : 8) | θ value | 1.06045e-08 (rank : 9) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 77 | |
| SwissProt Accessions | P30681, Q3UXT1, Q9EQD5 | Gene names | Hmgb2, Hmg2 | |||
|
Domain Architecture |

|
|||||
| Description | High mobility group protein B2 (High mobility group protein 2) (HMG- 2). | |||||
|
HMGB1_MOUSE
|
||||||
| NC score | 0.672519 (rank : 9) | θ value | 5.26297e-08 (rank : 16) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | P63158, P07155, P27109, P27428 | Gene names | Hmgb1, Hmg-1, Hmg1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | High mobility group protein B1 (High mobility group protein 1) (HMG- 1). | |||||
|
HMGB1_HUMAN
|
||||||
| NC score | 0.671960 (rank : 10) | θ value | 5.26297e-08 (rank : 15) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | P09429, Q6IBE1 | Gene names | HMGB1, HMG1 | |||
|
Domain Architecture |

|
|||||
| Description | High mobility group protein B1 (High mobility group protein 1) (HMG- 1). | |||||
|
SSRP1_MOUSE
|
||||||
| NC score | 0.644507 (rank : 11) | θ value | 8.97725e-08 (rank : 19) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | Q08943, Q3U9Z2, Q3UJ75, Q4V9U4, Q8CGA6 | Gene names | Ssrp1 | |||
|
Domain Architecture |

|
|||||
| Description | FACT complex subunit SSRP1 (Facilitates chromatin transcription complex subunit SSRP1) (Structure-specific recognition protein 1) (Recombination signal sequence recognition protein 1) (T160). | |||||
|
SSRP1_HUMAN
|
||||||
| NC score | 0.643644 (rank : 12) | θ value | 1.53129e-07 (rank : 21) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | Q08945, Q5BJG8 | Gene names | SSRP1, FACT80 | |||
|
Domain Architecture |

|
|||||
| Description | FACT complex subunit SSRP1 (Facilitates chromatin transcription complex subunit SSRP1) (FACT 80 kDa subunit) (FACTp80) (Chromatin- specific transcription elongation factor 80 kDa subunit) (Structure- specific recognition protein 1) (hSSRP1) (Recombination signal sequence recognition protein 1) (T160). | |||||
|
HMG1X_HUMAN
|
||||||
| NC score | 0.642862 (rank : 13) | θ value | 2.21117e-06 (rank : 30) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q9UGV6 | Gene names | HMG1L10 | |||
|
Domain Architecture |

|
|||||
| Description | High mobility group protein 1-like 10 (HMG-1L10). | |||||
|
TFAM_MOUSE
|
||||||
| NC score | 0.612231 (rank : 14) | θ value | 1.87187e-05 (rank : 36) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | P40630, P97894, P97906, Q543I8, Q9DBM9 | Gene names | Tfam, Hmgts | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor A, mitochondrial precursor (mtTFA) (Testis- specific high mobility group protein) (TS-HMG). | |||||
|
SOX14_MOUSE
|
||||||
| NC score | 0.605998 (rank : 15) | θ value | 1.29631e-06 (rank : 26) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | Q04892, Q9JLC9, Q9JLD0, Q9QXU4 | Gene names | Sox14, Sox-14 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-14. | |||||
|
SOX14_HUMAN
|
||||||
| NC score | 0.605928 (rank : 16) | θ value | 1.29631e-06 (rank : 25) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | O95416, Q3KPH7 | Gene names | SOX14, SOX28 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-14. | |||||
|
SOX7_MOUSE
|
||||||
| NC score | 0.605177 (rank : 17) | θ value | 7.59969e-07 (rank : 24) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 78 | |
| SwissProt Accessions | P40646, Q9R1T6 | Gene names | Sox7, Sox-7 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-7 (mSOX7). | |||||
|
SOX7_HUMAN
|
||||||
| NC score | 0.601826 (rank : 18) | θ value | 5.81887e-07 (rank : 23) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 78 | |
| SwissProt Accessions | Q9BT81 | Gene names | SOX7 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-7. | |||||
|
SOX2_MOUSE
|
||||||
| NC score | 0.592602 (rank : 19) | θ value | 1.29631e-06 (rank : 29) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | P48432 | Gene names | Sox2, Sox-2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-2. | |||||
|
SOX2_HUMAN
|
||||||
| NC score | 0.592516 (rank : 20) | θ value | 1.29631e-06 (rank : 28) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | P48431, Q14537 | Gene names | SOX2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-2. | |||||
|
SOX21_HUMAN
|
||||||
| NC score | 0.591673 (rank : 21) | θ value | 1.09739e-05 (rank : 33) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 78 | |
| SwissProt Accessions | Q9Y651, P35715, Q15504 | Gene names | SOX21, SOX25, SOXA | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-21 (SOX-A). | |||||
|
SOX21_MOUSE
|
||||||
| NC score | 0.591671 (rank : 22) | θ value | 1.09739e-05 (rank : 34) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 78 | |
| SwissProt Accessions | Q811W0, Q8VH35 | Gene names | Sox21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor SOX-21. | |||||
|
SOX15_HUMAN
|
||||||
| NC score | 0.589441 (rank : 23) | θ value | 0.000121331 (rank : 46) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | O60248, P35717, Q9Y6W7 | Gene names | SOX15, SOX12, SOX20, SOX26, SOX27 | |||
|
Domain Architecture |

|
|||||
| Description | SOX-15 protein (SOX-20 protein) (SOX-12 protein). | |||||
|
SOX15_MOUSE
|
||||||
| NC score | 0.589049 (rank : 24) | θ value | 0.000270298 (rank : 51) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | P43267, O70204, P70418, Q62246, Q91V00, Q91V43, Q920T1, Q9JLG2 | Gene names | Sox15, Sox-15 | |||
|
Domain Architecture |

|
|||||
| Description | SOX-15 protein. | |||||
|
SMCE1_HUMAN
|
||||||
| NC score | 0.588235 (rank : 25) | θ value | 3.29651e-10 (rank : 5) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 272 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | Q969G3, O43539 | Gene names | SMARCE1, BAF57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily E member 1 (BRG1-associated factor 57). | |||||
|
SOX18_HUMAN
|
||||||
| NC score | 0.586701 (rank : 26) | θ value | 4.1701e-05 (rank : 39) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | P35713, Q9NPH8 | Gene names | SOX18 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-18. | |||||
|
SOX17_MOUSE
|
||||||
| NC score | 0.584013 (rank : 27) | θ value | 1.29631e-06 (rank : 27) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | Q61473, Q61472, Q62248 | Gene names | Sox17, Sox-17 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-17. | |||||
|
SOX17_HUMAN
|
||||||
| NC score | 0.582435 (rank : 28) | θ value | 1.09739e-05 (rank : 32) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | Q9H6I2 | Gene names | SOX17 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-17. | |||||
|
SOX18_MOUSE
|
||||||
| NC score | 0.582126 (rank : 29) | θ value | 4.1701e-05 (rank : 40) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | P43680, Q9EQ73 | Gene names | Sox18, Sox-18 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-18. | |||||
|
SOX3_MOUSE
|
||||||
| NC score | 0.579738 (rank : 30) | θ value | 4.1701e-05 (rank : 44) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | P53784 | Gene names | Sox3, Sox-3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-3. | |||||
|
SOX1_HUMAN
|
||||||
| NC score | 0.579709 (rank : 31) | θ value | 4.1701e-05 (rank : 41) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | O00570 | Gene names | SOX1 | |||
|
Domain Architecture |

|
|||||
| Description | SOX-1 protein. | |||||
|
SOX1_MOUSE
|
||||||
| NC score | 0.579348 (rank : 32) | θ value | 4.1701e-05 (rank : 42) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | P53783 | Gene names | Sox1, Sox-1 | |||
|
Domain Architecture |

|
|||||
| Description | SOX-1 protein. | |||||
|
SOX3_HUMAN
|
||||||
| NC score | 0.576648 (rank : 33) | θ value | 4.1701e-05 (rank : 43) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 78 | |
| SwissProt Accessions | P41225, P35714, Q9NP49 | Gene names | SOX3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-3. | |||||
|
CN092_HUMAN
|
||||||
| NC score | 0.569718 (rank : 34) | θ value | 3.08544e-08 (rank : 13) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | O94842 | Gene names | C14orf92, KIAA0737 | |||
|
Domain Architecture |

|
|||||
| Description | Epidermal Langerhans cell protein LCP1. | |||||
|
SRY_HUMAN
|
||||||
| NC score | 0.568453 (rank : 35) | θ value | 0.0148317 (rank : 70) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | Q05066 | Gene names | SRY, TDF | |||
|
Domain Architecture |

|
|||||
| Description | Sex-determining region Y protein (Testis-determining factor). | |||||
|
TFAM_HUMAN
|
||||||
| NC score | 0.568002 (rank : 36) | θ value | 3.77169e-06 (rank : 31) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q00059 | Gene names | TFAM, TCF6L2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor A, mitochondrial precursor (mtTFA) (Mitochondrial transcription factor 1) (MtTF1) (Transcription factor 6-like 2). | |||||
|
UBF1_HUMAN
|
||||||
| NC score | 0.567110 (rank : 37) | θ value | 5.26297e-08 (rank : 17) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 415 | Shared Neighborhood Hits | 86 | |
| SwissProt Accessions | P17480 | Gene names | UBTF, UBF, UBF1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar transcription factor 1 (Upstream-binding factor 1) (UBF-1) (Autoantigen NOR-90). | |||||
|
SOX12_MOUSE
|
||||||
| NC score | 0.565213 (rank : 38) | θ value | 0.000270298 (rank : 50) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q04890, P70417, Q6NXL2 | Gene names | Sox12, Sox-12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SOX-12 protein. | |||||
|
TOX_MOUSE
|
||||||
| NC score | 0.564673 (rank : 39) | θ value | 1.06045e-08 (rank : 10) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q66JW3, Q8BKH9, Q8BYQ5, Q8R4H0 | Gene names | Tox | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thymus high mobility group box protein TOX. | |||||
|
SOX12_HUMAN
|
||||||
| NC score | 0.564390 (rank : 40) | θ value | 0.000270298 (rank : 49) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | O15370, Q9NUD4 | Gene names | SOX12, SOX22 | |||
|
Domain Architecture |

|
|||||
| Description | SOX-12 protein (SOX-22 protein). | |||||
|
TOX_HUMAN
|
||||||
| NC score | 0.562286 (rank : 41) | θ value | 1.80886e-08 (rank : 12) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | O94900, Q96AV5 | Gene names | TOX, KIAA0808 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thymus high mobility group box protein TOX. | |||||
|
CN092_MOUSE
|
||||||
| NC score | 0.559636 (rank : 42) | θ value | 3.08544e-08 (rank : 14) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q8BU11, Q80UI2, Q99PN9, Q9CS16 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | Epidermal Langerhans cell protein LCP1. | |||||
|
UBF1_MOUSE
|
||||||
| NC score | 0.558850 (rank : 43) | θ value | 5.26297e-08 (rank : 18) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 86 | |
| SwissProt Accessions | P25976 | Gene names | Ubtf, Tcfubf, Ubf-1, Ubf1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar transcription factor 1 (Upstream-binding factor 1) (UBF-1). | |||||
|
SOX11_HUMAN
|
||||||
| NC score | 0.558420 (rank : 44) | θ value | 0.000786445 (rank : 58) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | P35716 | Gene names | SOX11 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-11. | |||||
|
SOX11_MOUSE
|
||||||
| NC score | 0.557644 (rank : 45) | θ value | 0.000786445 (rank : 59) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 77 | |
| SwissProt Accessions | Q7M6Y2, O35178, O89036, Q04889, Q80XF0 | Gene names | Sox11, Sox-11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor SOX-11. | |||||
|
SRY_MOUSE
|
||||||
| NC score | 0.557479 (rank : 46) | θ value | 0.00869519 (rank : 67) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | Q05738 | Gene names | Sry, Tdf, Tdy | |||
|
Domain Architecture |

|
|||||
| Description | Sex-determining region Y protein (Testis-determining factor). | |||||
|
SOX4_HUMAN
|
||||||
| NC score | 0.555869 (rank : 47) | θ value | 0.000461057 (rank : 53) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 77 | |
| SwissProt Accessions | Q06945 | Gene names | SOX4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-4. | |||||
|
SMCE1_MOUSE
|
||||||
| NC score | 0.555227 (rank : 48) | θ value | 3.29651e-10 (rank : 6) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | O54941, Q8BPD9 | Gene names | Smarce1, Baf57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator chromatin subfamily E member 1 (BRG1-associated factor 57). | |||||
|
SOX4_MOUSE
|
||||||
| NC score | 0.555221 (rank : 49) | θ value | 0.00134147 (rank : 62) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 77 | |
| SwissProt Accessions | Q06831 | Gene names | Sox4, Sox-4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-4. | |||||
|
SOX8_MOUSE
|
||||||
| NC score | 0.548552 (rank : 50) | θ value | 1.43324e-05 (rank : 35) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q04886 | Gene names | Sox8, Sox-8 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-8. | |||||
|
SOX8_HUMAN
|
||||||
| NC score | 0.548427 (rank : 51) | θ value | 9.29e-05 (rank : 45) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | P57073, Q9NZW2 | Gene names | SOX8 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-8. | |||||
|
BBX_MOUSE
|
||||||
| NC score | 0.536765 (rank : 52) | θ value | 2.44474e-05 (rank : 38) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | Q8VBW5, Q3TZK1, Q6NXY8, Q6PEU3, Q8BQJ7, Q8C7E0, Q8CDQ0, Q8CDV1, Q8VI48, Q8VI49, Q8VI50, Q9CS94 | Gene names | Bbx, Hbp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HMG box transcription factor BBX (Bobby sox homolog) (HMG box- containing protein 2). | |||||
|
BBX_HUMAN
|
||||||
| NC score | 0.528037 (rank : 53) | θ value | 2.44474e-05 (rank : 37) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q8WY36, Q2TAJ1, Q7L3J8, Q7LBY8, Q8NDB0, Q8WY35, Q9H0J6 | Gene names | BBX, HBP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HMG box transcription factor BBX (Bobby sox homolog) (HMG box- containing protein 2). | |||||
|
SOX9_MOUSE
|
||||||
| NC score | 0.523716 (rank : 54) | θ value | 0.000602161 (rank : 57) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q04887, Q8C7L2, Q91ZK2, Q99KQ0 | Gene names | Sox9, Sox-9 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-9. | |||||
|
SOX9_HUMAN
|
||||||
| NC score | 0.522775 (rank : 55) | θ value | 0.000602161 (rank : 56) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | P48436 | Gene names | SOX9 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-9. | |||||
|
SOX30_MOUSE
|
||||||
| NC score | 0.518149 (rank : 56) | θ value | 0.00298849 (rank : 64) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 77 | |
| SwissProt Accessions | Q8CGW4 | Gene names | Sox30 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-30. | |||||
|
SOX30_HUMAN
|
||||||
| NC score | 0.515104 (rank : 57) | θ value | 0.00298849 (rank : 63) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 77 | |
| SwissProt Accessions | O94993, O94995, Q8IYX6 | Gene names | SOX30 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-30. | |||||
|
SOX5_MOUSE
|
||||||
| NC score | 0.512428 (rank : 58) | θ value | 0.000786445 (rank : 61) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 86 | |
| SwissProt Accessions | P35710, O88184, O89018 | Gene names | Sox5, Sox-5 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-5. | |||||
|
SOX5_HUMAN
|
||||||
| NC score | 0.511260 (rank : 59) | θ value | 0.000786445 (rank : 60) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 85 | |
| SwissProt Accessions | P35711, Q86UK8, Q8J017, Q8J018, Q8J019, Q8J020, Q8N1D9, Q8N7E0, Q8TEA4 | Gene names | SOX5 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-5. | |||||
|
SOX13_MOUSE
|
||||||
| NC score | 0.507390 (rank : 60) | θ value | 0.000602161 (rank : 55) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 495 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | Q04891, Q922L3 | Gene names | Sox13, Sox-13 | |||
|
Domain Architecture |

|
|||||
| Description | SOX-13 protein. | |||||
|
SOX13_HUMAN
|
||||||
| NC score | 0.504556 (rank : 61) | θ value | 0.000602161 (rank : 54) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 464 | Shared Neighborhood Hits | 83 | |
| SwissProt Accessions | Q9UN79, O95275, O95826, Q9UHW7 | Gene names | SOX13 | |||
|
Domain Architecture |

|
|||||
| Description | SOX-13 protein (Type 1 diabetes autoantigen ICA12) (Islet cell antigen 12). | |||||
|
SOX6_HUMAN
|
||||||
| NC score | 0.500000 (rank : 62) | θ value | 0.00390308 (rank : 65) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 96 | |
| SwissProt Accessions | P35712, Q9BXQ3, Q9BXQ4, Q9BXQ5, Q9H0I8 | Gene names | SOX6 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-6. | |||||
|
SOX6_MOUSE
|
||||||
| NC score | 0.498831 (rank : 63) | θ value | 0.00390308 (rank : 66) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 96 | |
| SwissProt Accessions | P40645, Q62250, Q9QWS5 | Gene names | Sox6, Sox-6 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-6 (SOX-LZ). | |||||
|
SP100_HUMAN
|
||||||
| NC score | 0.493442 (rank : 64) | θ value | 1.99992e-07 (rank : 22) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | P23497, O75450, Q13343, Q96F70, Q96T24, Q96T95, Q9NP33, Q9UE32 | Gene names | SP100 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear autoantigen Sp-100 (Speckled 100 kDa) (Nuclear dot-associated Sp100 protein) (Lysp100b). | |||||
|
CIC_MOUSE
|
||||||
| NC score | 0.482185 (rank : 65) | θ value | 0.00020696 (rank : 48) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 742 | Shared Neighborhood Hits | 78 | |
| SwissProt Accessions | Q924A2, Q6PDJ8, Q8CGE4, Q8CHH0, Q9CW61 | Gene names | Cic, Kiaa0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
GCX1_HUMAN
|
||||||
| NC score | 0.471189 (rank : 66) | θ value | 0.000461057 (rank : 52) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q96NM4, Q5TE33, Q5TE34, Q5TE35, Q96IC9, Q9BQN5 | Gene names | GCX1, C20orf100 | |||
|
Domain Architecture |

|
|||||
| Description | Granulosa cell HMG box protein 1 (GCX-1). | |||||
|
SOX10_HUMAN
|
||||||
| NC score | 0.471061 (rank : 67) | θ value | θ > 10 (rank : 130) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | P56693 | Gene names | SOX10 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-10. | |||||
|
SOX10_MOUSE
|
||||||
| NC score | 0.470910 (rank : 68) | θ value | θ > 10 (rank : 131) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q04888, O08518, O09141, O54856, P70416 | Gene names | Sox10, Sox-10, Sox21 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-10 (SOX-21) (Transcription factor SOX-M). | |||||
|
CIC_HUMAN
|
||||||
| NC score | 0.453363 (rank : 69) | θ value | 0.00020696 (rank : 47) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 992 | Shared Neighborhood Hits | 79 | |
| SwissProt Accessions | Q96RK0, Q7LGI1, Q9UEG5, Q9Y6T1 | Gene names | CIC, KIAA0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
HM2L1_HUMAN
|
||||||
| NC score | 0.445396 (rank : 70) | θ value | 0.62314 (rank : 81) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9UGU5, O75672, O75673, Q9UMT5 | Gene names | HMG2L1, HMGBCG | |||
|
Domain Architecture |

|
|||||
| Description | High mobility group protein 2-like 1 (Protein HMGBCG). | |||||
|
HBP1_HUMAN
|
||||||
| NC score | 0.409471 (rank : 71) | θ value | 1.06291 (rank : 89) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | O60381, Q8TBM1, Q8TE93, Q96AJ2 | Gene names | HBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HMG box-containing protein 1 (HMG box transcription factor 1) (High mobility group box transcription factor 1). | |||||
|
HBP1_MOUSE
|
||||||
| NC score | 0.409116 (rank : 72) | θ value | 1.06291 (rank : 90) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q8R316, Q3V0I4, Q8BUS3, Q8C199 | Gene names | Hbp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HMG box-containing protein 1 (HMG box transcription factor 1) (High mobility group box transcription factor 1). | |||||
|
PB1_HUMAN
|
||||||
| NC score | 0.394359 (rank : 73) | θ value | 1.53129e-07 (rank : 20) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 261 | Shared Neighborhood Hits | 80 | |
| SwissProt Accessions | Q86U86, Q9H2T3, Q9H2T4, Q9H2T5, Q9H301, Q9H314 | Gene names | PB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein polybromo-1 (hPB1) (Polybromo-1D) (BRG1-associated factor 180) (BAF180). | |||||
|
TCF7_HUMAN
|
||||||
| NC score | 0.386693 (rank : 74) | θ value | 0.21417 (rank : 73) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 186 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | P36402, Q9UKI4 | Gene names | TCF7, TCF1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 7 (T-cell-specific transcription factor 1) (TCF- 1) (T-cell factor 1). | |||||
|
LEF1_HUMAN
|
||||||
| NC score | 0.364272 (rank : 75) | θ value | 0.0330416 (rank : 71) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q9UJU2, Q9HAZ0 | Gene names | LEF1 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphoid enhancer-binding factor 1 (LEF-1) (T cell-specific transcription factor 1-alpha) (TCF1-alpha). | |||||
|
LEF1_MOUSE
|
||||||
| NC score | 0.363913 (rank : 76) | θ value | 0.0330416 (rank : 72) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | P27782 | Gene names | Lef1, Lef-1 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphoid enhancer-binding factor 1 (LEF-1). | |||||
|
TCF7_MOUSE
|
||||||
| NC score | 0.359894 (rank : 77) | θ value | 0.62314 (rank : 82) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 277 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q00417 | Gene names | Tcf7, Tcf-1, Tcf1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 7 (T-cell-specific transcription factor 1) (TCF- 1) (T-cell factor 1). | |||||
|
TF7L2_MOUSE
|
||||||
| NC score | 0.347678 (rank : 78) | θ value | 0.62314 (rank : 84) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q924A0, O70574, Q91XP2, Q91XP3, Q91XP4, Q924A1, Q9Z0V3, Q9Z0V4 | Gene names | Tcf7l2, Tcf4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 7-like 2 (HMG box transcription factor 4) (T- cell-specific transcription factor 4) (TCF-4) (mTCF-4). | |||||
|
TF7L2_HUMAN
|
||||||
| NC score | 0.346487 (rank : 79) | θ value | 0.62314 (rank : 83) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9NQB0, O00185, Q9NQB1, Q9NQB2, Q9NQB3, Q9NQB4, Q9NQB5, Q9NQB6, Q9NQB7, Q9ULC2 | Gene names | TCF7L2, TCF4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 7-like 2 (HMG box transcription factor 4) (T- cell-specific transcription factor 4) (TCF-4) (hTCF-4). | |||||
|
TF7L1_MOUSE
|
||||||
| NC score | 0.327654 (rank : 80) | θ value | 1.81305 (rank : 96) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q9Z1J1, O70450, O70573 | Gene names | Tcf7l1, Tcf3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 7-like 1 (HMG box transcription factor 3) (TCF-3) (mTCF-3). | |||||
|
TF7L1_HUMAN
|
||||||
| NC score | 0.327175 (rank : 81) | θ value | 1.81305 (rank : 95) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q9HCS4, Q53R97, Q6PD70, Q9NP00 | Gene names | TCF7L1, TCF3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 7-like 1 (HMG box transcription factor 3) (TCF- 3). | |||||
|
PMS1_HUMAN
|
||||||
| NC score | 0.263624 (rank : 82) | θ value | θ > 10 (rank : 129) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | P54277 | Gene names | PMS1, PMSL1 | |||
|
Domain Architecture |

|
|||||
| Description | PMS1 protein homolog 1 (DNA mismatch repair protein PMS1). | |||||
|
ST18_MOUSE
|
||||||
| NC score | 0.076559 (rank : 83) | θ value | 0.00869519 (rank : 68) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 248 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q80TY4, Q3UH00, Q3URH9, Q3UVB9, Q3UZN9, Q811B4, Q8K098 | Gene names | St18, Kiaa0535 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppression of tumorigenicity protein 18. | |||||
|
ST18_HUMAN
|
||||||
| NC score | 0.075575 (rank : 84) | θ value | 0.0113563 (rank : 69) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 252 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O60284 | Gene names | ST18, KIAA0535, ZNF387 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppression of tumorigenicity protein 18 (Zinc finger protein 387). | |||||
|
MYT1_HUMAN
|
||||||
| NC score | 0.069037 (rank : 85) | θ value | θ > 10 (rank : 127) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 475 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q01538, O94922, Q9UPV2 | Gene names | MYT1, KIAA0835, KIAA1050, MTF1, MYTI, PLPB1 | |||
|
Domain Architecture |

|
|||||
| Description | Myelin transcription factor 1 (MyT1) (MyTI) (Proteolipid protein- binding protein) (PLPB1). | |||||
|
MYT1_MOUSE
|
||||||
| NC score | 0.065405 (rank : 86) | θ value | θ > 10 (rank : 128) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8CFC2, O08995, Q8CFH1 | Gene names | Myt1, Kiaa0835, Nzf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myelin transcription factor 1 (MyT1) (Neural zinc finger factor 2) (NZF-2). | |||||
|
UBN1_HUMAN
|
||||||
| NC score | 0.064338 (rank : 87) | θ value | 4.03905 (rank : 107) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9NPG3, Q13079, Q9P1P7 | Gene names | UBN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubinuclein (Ubiquitously expressed nuclear protein) (VT4). | |||||
|
RC3H1_MOUSE
|
||||||
| NC score | 0.040513 (rank : 88) | θ value | 1.38821 (rank : 93) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 244 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q4VGL6, Q69Z31 | Gene names | Rc3h1, Gm551, Kiaa2025 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Roquin (RING finger and C3H zinc finger protein 1) (Sanroque protein). | |||||
|
CF060_HUMAN
|
||||||
| NC score | 0.039439 (rank : 89) | θ value | 0.47712 (rank : 76) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1270 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q8NB25, Q96GY8, Q9H851 | Gene names | C6orf60 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C6orf60. | |||||
|
REST_MOUSE
|
||||||
| NC score | 0.037456 (rank : 90) | θ value | 2.36792 (rank : 99) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1506 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q922J3, Q571L7 | Gene names | Rsn, Kiaa4046 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Restin. | |||||
|
GOGA1_HUMAN
|
||||||
| NC score | 0.036560 (rank : 91) | θ value | 0.62314 (rank : 80) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1222 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q92805, Q8IYZ9 | Gene names | GOLGA1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 1 (Golgin-97). | |||||
|
CP250_HUMAN
|
||||||
| NC score | 0.036115 (rank : 92) | θ value | 0.279714 (rank : 74) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1805 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q9BV73, O14812, O60588, Q9H450 | Gene names | CEP250, CEP2, CNAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Centrosome-associated protein CEP250 (Centrosomal protein 2) (Centrosomal Nek2-associated protein 1) (C-Nap1). | |||||
|
K0555_HUMAN
|
||||||
| NC score | 0.035931 (rank : 93) | θ value | 2.36792 (rank : 97) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 880 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q96AA8, O60302 | Gene names | KIAA0555 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0555. | |||||
|
LIPA1_HUMAN
|
||||||
| NC score | 0.035276 (rank : 94) | θ value | 0.47712 (rank : 77) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1294 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q13136, Q13135, Q14567, Q8N4I2 | Gene names | PPFIA1, LIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-1 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-1) (PTPRF-interacting protein alpha-1) (LAR-interacting protein 1) (LIP.1). | |||||
|
REST_HUMAN
|
||||||
| NC score | 0.035182 (rank : 95) | θ value | 0.47712 (rank : 78) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1605 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P30622 | Gene names | RSN, CYLN1 | |||
|
Domain Architecture |

|
|||||
| Description | Restin (Cytoplasmic linker protein 170 alpha-2) (CLIP-170) (Reed- Sternberg intermediate filament-associated protein) (Cytoplasmic linker protein 1). | |||||
|
CROCC_MOUSE
|
||||||
| NC score | 0.034050 (rank : 96) | θ value | 0.62314 (rank : 79) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1460 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8CJ40, Q7TQL2, Q80U01, Q8R0B9 | Gene names | Crocc, Kiaa0445 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rootletin (Ciliary rootlet coiled-coil protein). | |||||
|
GOGA1_MOUSE
|
||||||
| NC score | 0.033348 (rank : 97) | θ value | 1.06291 (rank : 88) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 994 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q9CW79, Q80YB0 | Gene names | Golga1 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 1 (Golgin-97). | |||||
|
NEMO_HUMAN
|
||||||
| NC score | 0.032393 (rank : 98) | θ value | 1.06291 (rank : 91) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 820 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9Y6K9 | Gene names | IKBKG, FIP3, NEMO | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NF-kappa-B essential modulator (NEMO) (NF-kappa-B essential modifier) (Inhibitor of nuclear factor kappa-B kinase subunit gamma) (IkB kinase subunit gamma) (I-kappa-B kinase gamma) (IKK-gamma) (IKKG) (IkB kinase-associated protein 1) (IKKAP1) (FIP-3). | |||||
|
RC3H1_HUMAN
|
||||||
| NC score | 0.030652 (rank : 99) | θ value | 6.88961 (rank : 116) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 433 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q5TC82, Q5W180, Q5W181, Q8IVE6, Q8N9V1 | Gene names | RC3H1, KIAA2025 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Roquin (RING finger and C3H zinc finger protein 1). | |||||
|
LIPA2_MOUSE
|
||||||
| NC score | 0.029606 (rank : 100) | θ value | 8.99809 (rank : 125) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1282 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q8BSS9, Q8BN73 | Gene names | Ppfia2 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-2 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-2) (PTPRF-interacting protein alpha-2). | |||||
|
CK5P2_HUMAN
|
||||||
| NC score | 0.029231 (rank : 101) | θ value | 1.06291 (rank : 86) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1433 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q96SN8, Q7Z3L4, Q7Z3U1, Q7Z7I6, Q9BSW0, Q9H6J6, Q9HCD9, Q9NV90, Q9UIW9 | Gene names | CDK5RAP2, CEP215, KIAA1633 | |||
|
Domain Architecture |

|
|||||
| Description | CDK5 regulatory subunit-associated protein 2 (CDK5 activator-binding protein C48) (Centrosome-associated protein 215). | |||||
|
LIPA2_HUMAN
|
||||||
| NC score | 0.029090 (rank : 102) | θ value | 8.99809 (rank : 124) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1044 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O75334 | Gene names | PPFIA2 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-2 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-2) (PTPRF-interacting protein alpha-2). | |||||
|
LIPA3_HUMAN
|
||||||
| NC score | 0.028077 (rank : 103) | θ value | 8.99809 (rank : 126) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O75145, Q9H8B5, Q9UEW4 | Gene names | PPFIA3, KIAA0654 | |||
|
Domain Architecture |

|
|||||
| Description | Liprin-alpha-3 (Protein tyrosine phosphatase receptor type f polypeptide-interacting protein alpha-3) (PTPRF-interacting protein alpha-3). | |||||
|
GOGA3_HUMAN
|
||||||
| NC score | 0.026989 (rank : 104) | θ value | 4.03905 (rank : 104) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1317 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q08378, O43241, Q6P9C7, Q86XW3, Q8TDA9, Q8WZA3 | Gene names | GOLGA3 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 3 (Golgin-160) (Golgi complex-associated protein of 170 kDa) (GCP170). | |||||
|
CROCC_HUMAN
|
||||||
| NC score | 0.026790 (rank : 105) | θ value | 5.27518 (rank : 108) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1409 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q5TZA2, Q2VHY3, Q66GT7, Q7Z2L4, Q7Z5D7 | Gene names | CROCC, KIAA0445 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rootletin (Ciliary rootlet coiled-coil protein). | |||||
|
BPAEA_HUMAN
|
||||||
| NC score | 0.026485 (rank : 106) | θ value | 8.99809 (rank : 119) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1369 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | O94833, Q5TBT0, Q8N1T8, Q8N8J3, Q8WXK9, Q96AK9, Q96DQ5, Q9H555 | Gene names | DST, BPAG1, DMH, DT, KIAA0728 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoforms 6/9/10 (Trabeculin-beta) (Bullous pemphigoid antigen) (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
CAN3_MOUSE
|
||||||
| NC score | 0.025885 (rank : 107) | θ value | 0.47712 (rank : 75) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 115 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q64691, Q9WUC5 | Gene names | Capn3 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-3 (EC 3.4.22.-) (Calpain L3) (Calpain p94) (Calcium-activated neutral proteinase 3) (CANP 3) (Muscle-specific calcium-activated neutral protease 3). | |||||
|
NIN_MOUSE
|
||||||
| NC score | 0.025455 (rank : 108) | θ value | 3.0926 (rank : 101) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1376 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q61043, Q6ZPM7 | Gene names | Nin, Kiaa1565 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ninein. | |||||
|
CCD41_HUMAN
|
||||||
| NC score | 0.022455 (rank : 109) | θ value | 8.99809 (rank : 120) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1139 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9Y592 | Gene names | CCDC41 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 41 (NY-REN-58 antigen). | |||||
|
SPT16_MOUSE
|
||||||
| NC score | 0.021362 (rank : 110) | θ value | 5.27518 (rank : 113) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q920B9, Q3TZ48, Q3UFH0, Q921H4 | Gene names | Supt16h, Fact140, Factp140 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FACT complex subunit SPT16 (Facilitates chromatin transcription complex subunit SPT16) (FACT 140 kDa subunit) (FACTp140) (Chromatin- specific transcription elongation factor 140 kDa subunit). | |||||
|
SPT16_HUMAN
|
||||||
| NC score | 0.021014 (rank : 111) | θ value | 5.27518 (rank : 112) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Y5B9, Q6GMT8, Q6P2F1, Q6PJM1, Q9NRX0 | Gene names | SUPT16H, FACT140, FACTP140 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FACT complex subunit SPT16 (Facilitates chromatin transcription complex subunit SPT16) (hSPT16) (FACT 140 kDa subunit) (FACTp140) (Chromatin-specific transcription elongation factor 140 kDa subunit). | |||||
|
TEX9_HUMAN
|
||||||
| NC score | 0.020577 (rank : 112) | θ value | 6.88961 (rank : 117) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 541 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8N6V9 | Gene names | TEX9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Testis-expressed sequence 9 protein. | |||||
|
CCD41_MOUSE
|
||||||
| NC score | 0.019587 (rank : 113) | θ value | 8.99809 (rank : 121) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9D5R3, Q3U7X7, Q3UX57, Q80VF0 | Gene names | Ccdc41 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 41. | |||||
|
KIF23_HUMAN
|
||||||
| NC score | 0.017511 (rank : 114) | θ value | 2.36792 (rank : 98) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 618 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q02241 | Gene names | KIF23, KNSL5, MKLP1 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF23 (Mitotic kinesin-like protein 1) (Kinesin- like protein 5). | |||||
|
CLC4F_MOUSE
|
||||||
| NC score | 0.017362 (rank : 115) | θ value | 4.03905 (rank : 103) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 572 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P70194, Q8BLZ8 | Gene names | Clec4f, Clecsf13, Kclr | |||
|
Domain Architecture |

|
|||||
| Description | C-type lectin domain family 4 member F (C-type lectin superfamily member 13) (C-type lectin 13) (Kupffer cell receptor). | |||||
|
KIF14_HUMAN
|
||||||
| NC score | 0.016983 (rank : 116) | θ value | 1.38821 (rank : 92) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 732 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q15058 | Gene names | KIF14, KIAA0042 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin-like protein KIF14. | |||||
|
GABP2_HUMAN
|
||||||
| NC score | 0.014564 (rank : 117) | θ value | 0.813845 (rank : 85) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 454 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q06547, Q06545, Q12940, Q12941, Q12942, Q8IYD0 | Gene names | GABPB2, E4TF1B, GABPB, GABPB1 | |||
|
Domain Architecture |

|
|||||
| Description | GA-binding protein beta chain (GABP subunit beta-2) (GABP-2) (GABP subunit beta-1) (GABPB-1) (Transcription factor E4TF1-53) (Transcription factor E4TF1-47) (Nuclear respiratory factor 2). | |||||
|
GABP2_MOUSE
|
||||||
| NC score | 0.014395 (rank : 118) | θ value | 1.06291 (rank : 87) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 429 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q00420, Q00421, Q8BS31, Q91YZ0, Q9QVV2 | Gene names | Gabpb2, Gabpb, Gabpb1 | |||
|
Domain Architecture |

|
|||||
| Description | GA-binding protein beta chain (GABP subunit beta-2) (GABP-2) (GABP subunit beta-1) (GABPB-1). | |||||
|
LRSM1_HUMAN
|
||||||
| NC score | 0.013649 (rank : 119) | θ value | 4.03905 (rank : 105) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1078 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q6UWE0, Q5VVV0, Q8NB40, Q96GT5, Q96MX5, Q96MZ7 | Gene names | LRSAM1, TAL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin ligase protein LRSAM1 (EC 6.3.2.-) (Leucine-rich repeat and sterile alpha motif-containing protein 1) (Tsg101-associated ligase) (hTAL). | |||||
|
GOPC_HUMAN
|
||||||
| NC score | 0.012876 (rank : 120) | θ value | 8.99809 (rank : 122) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 377 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9HD26, Q59FS4, Q969U8 | Gene names | GOPC, CAL, FIG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi-associated PDZ and coiled-coil motif-containing protein (PDZ protein interacting specifically with TC10) (PIST) (CFTR-associated ligand) (Fused in glioblastoma). | |||||
|
GOPC_MOUSE
|
||||||
| NC score | 0.012394 (rank : 121) | θ value | 8.99809 (rank : 123) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8BH60, Q8BSV4, Q920R1, Q9ET11 | Gene names | Gopc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi-associated PDZ and coiled-coil motif-containing protein (PDZ protein interacting specifically with TC10) (PIST). | |||||
|
TRI32_MOUSE
|
||||||
| NC score | 0.011906 (rank : 122) | θ value | 6.88961 (rank : 118) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8CH72, Q8K055 | Gene names | Trim32 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 32 (EC 6.3.2.-). | |||||
|
POT15_HUMAN
|
||||||
| NC score | 0.011478 (rank : 123) | θ value | 5.27518 (rank : 111) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 893 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q6S5H4, Q6NXN7, Q6S5H7 | Gene names | POTE15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prostate, ovary, testis-expressed protein on chromosome 15. | |||||
|
ANR21_HUMAN
|
||||||
| NC score | 0.011109 (rank : 124) | θ value | 6.88961 (rank : 114) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 864 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q86YR6 | Gene names | ANKRD21, POTE | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin repeat domain-containing protein 21 (Protein POTE) (Prostate, ovary, testis-expressed protein). | |||||
|
IQGA2_HUMAN
|
||||||
| NC score | 0.010890 (rank : 125) | θ value | 5.27518 (rank : 109) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q13576 | Gene names | IQGAP2 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP2. | |||||
|
LAMB3_HUMAN
|
||||||
| NC score | 0.010379 (rank : 126) | θ value | 1.81305 (rank : 94) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 461 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q13751, O14947, Q14733, Q9UJK4, Q9UJL1 | Gene names | LAMB3 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-3 chain precursor (Laminin 5 beta 3) (Laminin B1k chain) (Kalinin B1 chain). | |||||
|
NFH_MOUSE
|
||||||
| NC score | 0.010160 (rank : 127) | θ value | 6.88961 (rank : 115) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
MORF4_HUMAN
|
||||||
| NC score | 0.010006 (rank : 128) | θ value | 5.27518 (rank : 110) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y690 | Gene names | MORF4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor-like protein MORF4 (Mortality factor 4) (Cellular senescence-related protein 1) (SEN1). | |||||
|
TRI41_HUMAN
|
||||||
| NC score | 0.009915 (rank : 129) | θ value | 3.0926 (rank : 102) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8WV44, Q5BKT0, Q7L484, Q96Q10, Q9BSL8 | Gene names | TRIM41 | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 41. | |||||
|
TRI38_HUMAN
|
||||||
| NC score | 0.009593 (rank : 130) | θ value | 2.36792 (rank : 100) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 373 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O00635 | Gene names | TRIM38, RNF15, RORET | |||
|
Domain Architecture |

|
|||||
| Description | Tripartite motif-containing protein 38 (RING finger protein 15) (Zinc finger protein RoRet). | |||||
|
TEX14_MOUSE
|
||||||
| NC score | 0.003835 (rank : 131) | θ value | 4.03905 (rank : 106) | |||
| Query Neighborhood Hits | 126 | Target Neighborhood Hits | 1333 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q7M6U3, Q3UT36, Q5NC10, Q5NC11, Q8CGK1, Q8CGK2, Q99MV8 | Gene names | Tex14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Testis-expressed protein 14 (Testis-expressed sequence 14). | |||||