Please be patient as the page loads
|
DUS11_HUMAN
|
||||||
| SwissProt Accessions | O75319, Q6AI47, Q9BWE3 | Gene names | DUSP11, PIR1 | |||
|
Domain Architecture |
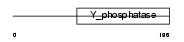
|
|||||
| Description | RNA/RNP complex-1-interacting phosphatase (EC 3.1.3.-) (Phosphatase that interacts with RNA/RNP complex 1) (Dual specificity protein phosphatase 11). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
DUS11_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O75319, Q6AI47, Q9BWE3 | Gene names | DUSP11, PIR1 | |||
|
Domain Architecture |

|
|||||
| Description | RNA/RNP complex-1-interacting phosphatase (EC 3.1.3.-) (Phosphatase that interacts with RNA/RNP complex 1) (Dual specificity protein phosphatase 11). | |||||
|
DUS11_MOUSE
|
||||||
| θ value | 3.89193e-128 (rank : 2) | NC score | 0.977154 (rank : 2) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q6NXK5, Q8BTR4, Q8BYE4 | Gene names | Dusp11, Pir1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA/RNP complex-1-interacting phosphatase (EC 3.1.3.-) (Phosphatase that interacts with RNA/RNP complex 1) (Dual specificity protein phosphatase 11). | |||||
|
MCE1_HUMAN
|
||||||
| θ value | 1.5773e-20 (rank : 3) | NC score | 0.656453 (rank : 3) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O60942, O43483, O60257, O60351 | Gene names | RNGTT, CAP1A | |||
|
Domain Architecture |

|
|||||
| Description | mRNA capping enzyme (HCE) (HCAP1) [Includes: Polynucleotide 5'- triphosphatase (EC 3.1.3.33) (mRNA 5'-triphosphatase) (TPase); mRNA guanylyltransferase (EC 2.7.7.50) (GTP--RNA guanylyltransferase) (GTase)]. | |||||
|
MCE1_MOUSE
|
||||||
| θ value | 1.74391e-19 (rank : 4) | NC score | 0.651511 (rank : 4) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O55236 | Gene names | Rngtt, Cap1a | |||
|
Domain Architecture |

|
|||||
| Description | mRNA capping enzyme (HCE) (MCE1) [Includes: Polynucleotide 5'- triphosphatase (EC 3.1.3.33) (mRNA 5'-triphosphatase) (TPase); mRNA guanylyltransferase (EC 2.7.7.50) (GTP--RNA guanylyltransferase) (GTase)]. | |||||
|
TPTE_HUMAN
|
||||||
| θ value | 0.00228821 (rank : 5) | NC score | 0.207489 (rank : 8) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P56180, Q6XPS4, Q6XPS5, Q71JA8, Q8NCS8 | Gene names | TPTE | |||
|
Domain Architecture |

|
|||||
| Description | Putative tyrosine-protein phosphatase TPTE (EC 3.1.3.48) (Transmembrane phosphatase with tensin homology) (Protein BJ-HCC-5). | |||||
|
PTPM1_MOUSE
|
||||||
| θ value | 0.0193708 (rank : 6) | NC score | 0.234877 (rank : 7) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q66GT5, Q9CSJ8, Q9D622 | Gene names | Ptpmt1, Plip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein-tyrosine phosphatase mitochondrial 1, mitochondrial precursor (EC 3.1.3.48) (EC 3.1.3.16) (PTEN-like phosphatase) (Phosphoinositide lipid phosphatase). | |||||
|
TPTE2_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 7) | NC score | 0.183673 (rank : 9) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q6XPS3, Q5VUH2, Q8WWL4, Q8WWL5 | Gene names | TPTE2, TPIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol-3,4,5-trisphosphate 3-phosphatase TPTE2 (EC 3.1.3.67) (TPTE and PTEN homologous inositol lipid phosphatase) (Lipid phosphatase TPIP). | |||||
|
DUS23_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 8) | NC score | 0.246008 (rank : 5) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9BVJ7, Q9NX48 | Gene names | DUSP23, LDP3, VHZ | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dual specificity protein phosphatase 23 (EC 3.1.3.48) (EC 3.1.3.16) (Low molecular mass dual specificity phosphatase 3) (LDP-3) (VH1-like phosphatase Z). | |||||
|
DUS23_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 9) | NC score | 0.241904 (rank : 6) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q6NT99, Q9CW48 | Gene names | Dusp23, Ldp3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dual specificity protein phosphatase 23 (EC 3.1.3.48) (EC 3.1.3.16) (Low molecular mass dual specificity phosphatase 3) (LDP-3). | |||||
|
PTEN_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 10) | NC score | 0.164622 (rank : 11) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P60484, O00633, O02679, Q6ICT7 | Gene names | PTEN, MMAC1, TEP1 | |||
|
Domain Architecture |
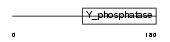
|
|||||
| Description | Phosphatidylinositol-3,4,5-trisphosphate 3-phosphatase and dual- specificity protein phosphatase PTEN (EC 3.1.3.67) (EC 3.1.3.16) (EC 3.1.3.48) (Phosphatase and tensin homolog) (Mutated in multiple advanced cancers 1). | |||||
|
PTEN_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 11) | NC score | 0.164621 (rank : 12) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O08586, Q542G1 | Gene names | Pten, Mmac1 | |||
|
Domain Architecture |
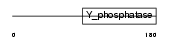
|
|||||
| Description | Phosphatidylinositol-3,4,5-trisphosphate 3-phosphatase PTEN (EC 3.1.3.67) (Mutated in multiple advanced cancers 1). | |||||
|
DUS2_MOUSE
|
||||||
| θ value | 0.365318 (rank : 12) | NC score | 0.068815 (rank : 17) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q05922, Q60640 | Gene names | Dusp2, Pac-1, Pac1 | |||
|
Domain Architecture |
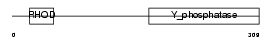
|
|||||
| Description | Dual specificity protein phosphatase 2 (EC 3.1.3.48) (EC 3.1.3.16) (Dual specificity protein phosphatase PAC-1). | |||||
|
DUS2_HUMAN
|
||||||
| θ value | 0.47712 (rank : 13) | NC score | 0.063267 (rank : 19) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q05923 | Gene names | DUSP2, PAC1 | |||
|
Domain Architecture |
|
|||||
| Description | Dual specificity protein phosphatase 2 (EC 3.1.3.48) (EC 3.1.3.16) (Dual specificity protein phosphatase PAC-1). | |||||
|
PTPRJ_HUMAN
|
||||||
| θ value | 0.62314 (rank : 14) | NC score | 0.051221 (rank : 21) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q12913, Q15255, Q8NHM2 | Gene names | PTPRJ, DEP1 | |||
|
Domain Architecture |

|
|||||
| Description | Receptor-type tyrosine-protein phosphatase eta precursor (EC 3.1.3.48) (Protein-tyrosine phosphatase eta) (R-PTP-eta) (HPTP eta) (Protein- tyrosine phosphatase receptor type J) (Density-enhanced phosphatase 1) (DEP-1) (CD148 antigen). | |||||
|
CC14A_HUMAN
|
||||||
| θ value | 1.38821 (rank : 15) | NC score | 0.120060 (rank : 14) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9UNH5, O43171, O60727, O60728, Q8IXX0 | Gene names | CDC14A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dual specificity protein phosphatase CDC14A (EC 3.1.3.48) (EC 3.1.3.16) (CDC14 cell division cycle 14 homolog A). | |||||
|
PTPRJ_MOUSE
|
||||||
| θ value | 1.81305 (rank : 16) | NC score | 0.045319 (rank : 25) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q64455 | Gene names | Ptprj, Byp, Scc1 | |||
|
Domain Architecture |

|
|||||
| Description | Receptor-type tyrosine-protein phosphatase eta precursor (EC 3.1.3.48) (Protein-tyrosine phosphatase eta) (R-PTP-eta) (HPTP beta-like tyrosine phosphatase) (Protein-tyrosine phosphatase receptor type J) (Susceptibility to colon cancer 1). | |||||
|
DUS16_HUMAN
|
||||||
| θ value | 2.36792 (rank : 17) | NC score | 0.066894 (rank : 18) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9BY84, Q9C0G3 | Gene names | DUSP16, KIAA1700, MKP7 | |||
|
Domain Architecture |
|
|||||
| Description | Dual specificity protein phosphatase 16 (EC 3.1.3.48) (EC 3.1.3.16) (Mitogen-activated protein kinase phosphatase 7) (MAP kinase phosphatase 7) (MKP-7). | |||||
|
DUS1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 18) | NC score | 0.049647 (rank : 23) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P28563 | Gene names | Dusp1, 3ch134, Mkp1, Ptpn10, Ptpn16 | |||
|
Domain Architecture |
|
|||||
| Description | Dual specificity protein phosphatase 1 (EC 3.1.3.48) (EC 3.1.3.16) (MAP kinase phosphatase 1) (MKP-1) (Protein-tyrosine phosphatase 3CH134) (Protein-tyrosine phosphatase ERP). | |||||
|
CC14A_MOUSE
|
||||||
| θ value | 3.0926 (rank : 19) | NC score | 0.115125 (rank : 15) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q6GQT0, Q8BZ66 | Gene names | Cdc14a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dual specificity protein phosphatase CDC14A (EC 3.1.3.48) (EC 3.1.3.16) (CDC14 cell division cycle 14 homolog A). | |||||
|
PTPM1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 20) | NC score | 0.182446 (rank : 10) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8WUK0, Q7Z557, Q96CR2, Q9BXV8 | Gene names | PTPMT1, MOSP, PLIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein-tyrosine phosphatase mitochondrial 1, mitochondrial precursor (EC 3.1.3.48) (EC 3.1.3.16) (PTEN-like phosphatase) (Phosphoinositide lipid phosphatase). | |||||
|
ZBT24_HUMAN
|
||||||
| θ value | 3.0926 (rank : 21) | NC score | 0.000785 (rank : 40) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 870 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O43167, Q5TED5, Q8N455 | Gene names | ZBTB24, KIAA0441, ZNF450 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger and BTB domain-containing protein 24 (Zinc finger protein 450). | |||||
|
BRD4_HUMAN
|
||||||
| θ value | 4.03905 (rank : 22) | NC score | 0.011241 (rank : 36) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 851 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O60885, O60433, Q96PD3 | Gene names | BRD4, HUNK1 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 4 (HUNK1 protein). | |||||
|
CASR_HUMAN
|
||||||
| θ value | 4.03905 (rank : 23) | NC score | 0.010070 (rank : 37) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P41180, Q13912, Q16108, Q16109, Q16110, Q16379, Q4PJ19 | Gene names | CASR, GPRC2A, PCAR1 | |||
|
Domain Architecture |

|
|||||
| Description | Extracellular calcium-sensing receptor precursor (CaSR) (Parathyroid Cell calcium-sensing receptor). | |||||
|
PTN14_HUMAN
|
||||||
| θ value | 4.03905 (rank : 24) | NC score | 0.024702 (rank : 28) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q15678 | Gene names | PTPN14, PEZ | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 14 (EC 3.1.3.48) (Protein-tyrosine phosphatase pez). | |||||
|
AMOT_HUMAN
|
||||||
| θ value | 5.27518 (rank : 25) | NC score | 0.011262 (rank : 35) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 712 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q4VCS5, Q504X5, Q9HD27, Q9UPT1 | Gene names | AMOT, KIAA1071 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin. | |||||
|
CC14B_HUMAN
|
||||||
| θ value | 5.27518 (rank : 26) | NC score | 0.127533 (rank : 13) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O60729, O43183, O60730 | Gene names | CDC14B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dual specificity protein phosphatase CDC14B (EC 3.1.3.48) (EC 3.1.3.16) (CDC14 cell division cycle 14 homolog B). | |||||
|
PTPRA_HUMAN
|
||||||
| θ value | 6.88961 (rank : 27) | NC score | 0.034481 (rank : 27) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P18433, Q14513, Q7Z2I2, Q96TD9 | Gene names | PTPRA, PTPA | |||
|
Domain Architecture |

|
|||||
| Description | Receptor-type tyrosine-protein phosphatase alpha precursor (EC 3.1.3.48) (Protein-tyrosine phosphatase alpha) (R-PTP-alpha). | |||||
|
PTPRA_MOUSE
|
||||||
| θ value | 6.88961 (rank : 28) | NC score | 0.034876 (rank : 26) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P18052, Q61808 | Gene names | Ptpra, Lrp, Ptpa | |||
|
Domain Architecture |

|
|||||
| Description | Receptor-type tyrosine-protein phosphatase alpha precursor (EC 3.1.3.48) (Protein-tyrosine phosphatase alpha) (R-PTP-alpha) (LCA- related phosphatase) (PTPTY-28). | |||||
|
PTPRD_HUMAN
|
||||||
| θ value | 6.88961 (rank : 29) | NC score | 0.024332 (rank : 30) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 485 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P23468 | Gene names | PTPRD | |||
|
Domain Architecture |

|
|||||
| Description | Receptor-type tyrosine-protein phosphatase delta precursor (EC 3.1.3.48) (Protein-tyrosine phosphatase delta) (R-PTP-delta). | |||||
|
PTPRD_MOUSE
|
||||||
| θ value | 6.88961 (rank : 30) | NC score | 0.024532 (rank : 29) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q64487, Q64486, Q64488, Q64495 | Gene names | Ptprd | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Receptor-type tyrosine-protein phosphatase delta precursor (EC 3.1.3.48) (Protein-tyrosine phosphatase delta) (R-PTP-delta). | |||||
|
PTPRS_HUMAN
|
||||||
| θ value | 6.88961 (rank : 31) | NC score | 0.024224 (rank : 31) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 472 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q13332, O75255, O75870, Q15718, Q16341 | Gene names | PTPRS | |||
|
Domain Architecture |

|
|||||
| Description | Receptor-type tyrosine-protein phosphatase S precursor (EC 3.1.3.48) (R-PTP-S) (Protein-tyrosine phosphatase sigma) (R-PTP-sigma). | |||||
|
CBX7_HUMAN
|
||||||
| θ value | 8.99809 (rank : 32) | NC score | 0.011766 (rank : 34) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O95931, Q86T17 | Gene names | CBX7 | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 7. | |||||
|
DUS1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 33) | NC score | 0.049452 (rank : 24) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P28562, Q2V508 | Gene names | DUSP1, CL100, MKP1, PTPN10, VH1 | |||
|
Domain Architecture |
|
|||||
| Description | Dual specificity protein phosphatase 1 (EC 3.1.3.48) (EC 3.1.3.16) (MAP kinase phosphatase 1) (MKP-1) (Protein-tyrosine phosphatase CL100) (Dual specificity protein phosphatase hVH1). | |||||
|
K0408_HUMAN
|
||||||
| θ value | 8.99809 (rank : 34) | NC score | 0.020152 (rank : 33) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6ZU52, O43158, Q5TF20, Q7L2M2 | Gene names | KIAA0408 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0408. | |||||
|
PTN3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 35) | NC score | 0.021956 (rank : 32) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P26045 | Gene names | PTPN3, PTPH1 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 3 (EC 3.1.3.48) (Protein-tyrosine phosphatase H1) (PTP-H1). | |||||
|
SLIT2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 36) | NC score | 0.003741 (rank : 38) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 708 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O94813, O95710, Q9Y5Q7 | Gene names | SLIT2, SLIL3 | |||
|
Domain Architecture |

|
|||||
| Description | Slit homolog 2 protein precursor (Slit-2) [Contains: Slit homolog 2 protein N-product; Slit homolog 2 protein C-product]. | |||||
|
SLIT2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 37) | NC score | 0.003735 (rank : 39) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 684 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9R1B9, Q9Z166 | Gene names | Slit2 | |||
|
Domain Architecture |

|
|||||
| Description | Slit homolog 2 protein precursor (Slit-2) [Contains: Slit homolog 2 protein N-product; Slit homolog 2 protein C-product]. | |||||
|
CC14B_MOUSE
|
||||||
| θ value | θ > 10 (rank : 38) | NC score | 0.106316 (rank : 16) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q6PFY9, Q80WC4, Q8BLV5 | Gene names | Cdc14b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dual specificity protein phosphatase CDC14B (EC 3.1.3.48) (EC 3.1.3.16) (CDC14 cell division cycle 14 homolog B). | |||||
|
DUS8_MOUSE
|
||||||
| θ value | θ > 10 (rank : 39) | NC score | 0.054183 (rank : 20) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O09112 | Gene names | Dusp8, Nttp1 | |||
|
Domain Architecture |
|
|||||
| Description | Dual specificity protein phosphatase 8 (EC 3.1.3.48) (EC 3.1.3.16) (Neuronal tyrosine threonine phosphatase 1). | |||||
|
TP4A3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 40) | NC score | 0.050798 (rank : 22) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9D658, O70275, Q3T9Z5, Q9CTC8 | Gene names | Ptp4a3, Prl3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein tyrosine phosphatase type IVA protein 3 (EC 3.1.3.48) (Protein-tyrosine phosphatase 4a3) (Protein-tyrosine phosphatase of regenerating liver 3) (PRL-3). | |||||
|
DUS11_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O75319, Q6AI47, Q9BWE3 | Gene names | DUSP11, PIR1 | |||
|
Domain Architecture |

|
|||||
| Description | RNA/RNP complex-1-interacting phosphatase (EC 3.1.3.-) (Phosphatase that interacts with RNA/RNP complex 1) (Dual specificity protein phosphatase 11). | |||||
|
DUS11_MOUSE
|
||||||
| NC score | 0.977154 (rank : 2) | θ value | 3.89193e-128 (rank : 2) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q6NXK5, Q8BTR4, Q8BYE4 | Gene names | Dusp11, Pir1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA/RNP complex-1-interacting phosphatase (EC 3.1.3.-) (Phosphatase that interacts with RNA/RNP complex 1) (Dual specificity protein phosphatase 11). | |||||
|
MCE1_HUMAN
|
||||||
| NC score | 0.656453 (rank : 3) | θ value | 1.5773e-20 (rank : 3) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O60942, O43483, O60257, O60351 | Gene names | RNGTT, CAP1A | |||
|
Domain Architecture |

|
|||||
| Description | mRNA capping enzyme (HCE) (HCAP1) [Includes: Polynucleotide 5'- triphosphatase (EC 3.1.3.33) (mRNA 5'-triphosphatase) (TPase); mRNA guanylyltransferase (EC 2.7.7.50) (GTP--RNA guanylyltransferase) (GTase)]. | |||||
|
MCE1_MOUSE
|
||||||
| NC score | 0.651511 (rank : 4) | θ value | 1.74391e-19 (rank : 4) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O55236 | Gene names | Rngtt, Cap1a | |||
|
Domain Architecture |

|
|||||
| Description | mRNA capping enzyme (HCE) (MCE1) [Includes: Polynucleotide 5'- triphosphatase (EC 3.1.3.33) (mRNA 5'-triphosphatase) (TPase); mRNA guanylyltransferase (EC 2.7.7.50) (GTP--RNA guanylyltransferase) (GTase)]. | |||||
|
DUS23_HUMAN
|
||||||
| NC score | 0.246008 (rank : 5) | θ value | 0.0431538 (rank : 8) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9BVJ7, Q9NX48 | Gene names | DUSP23, LDP3, VHZ | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dual specificity protein phosphatase 23 (EC 3.1.3.48) (EC 3.1.3.16) (Low molecular mass dual specificity phosphatase 3) (LDP-3) (VH1-like phosphatase Z). | |||||
|
DUS23_MOUSE
|
||||||
| NC score | 0.241904 (rank : 6) | θ value | 0.0961366 (rank : 9) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q6NT99, Q9CW48 | Gene names | Dusp23, Ldp3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dual specificity protein phosphatase 23 (EC 3.1.3.48) (EC 3.1.3.16) (Low molecular mass dual specificity phosphatase 3) (LDP-3). | |||||
|
PTPM1_MOUSE
|
||||||
| NC score | 0.234877 (rank : 7) | θ value | 0.0193708 (rank : 6) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q66GT5, Q9CSJ8, Q9D622 | Gene names | Ptpmt1, Plip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein-tyrosine phosphatase mitochondrial 1, mitochondrial precursor (EC 3.1.3.48) (EC 3.1.3.16) (PTEN-like phosphatase) (Phosphoinositide lipid phosphatase). | |||||
|
TPTE_HUMAN
|
||||||
| NC score | 0.207489 (rank : 8) | θ value | 0.00228821 (rank : 5) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P56180, Q6XPS4, Q6XPS5, Q71JA8, Q8NCS8 | Gene names | TPTE | |||
|
Domain Architecture |

|
|||||
| Description | Putative tyrosine-protein phosphatase TPTE (EC 3.1.3.48) (Transmembrane phosphatase with tensin homology) (Protein BJ-HCC-5). | |||||
|
TPTE2_HUMAN
|
||||||
| NC score | 0.183673 (rank : 9) | θ value | 0.0193708 (rank : 7) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q6XPS3, Q5VUH2, Q8WWL4, Q8WWL5 | Gene names | TPTE2, TPIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphatidylinositol-3,4,5-trisphosphate 3-phosphatase TPTE2 (EC 3.1.3.67) (TPTE and PTEN homologous inositol lipid phosphatase) (Lipid phosphatase TPIP). | |||||
|
PTPM1_HUMAN
|
||||||
| NC score | 0.182446 (rank : 10) | θ value | 3.0926 (rank : 20) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8WUK0, Q7Z557, Q96CR2, Q9BXV8 | Gene names | PTPMT1, MOSP, PLIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein-tyrosine phosphatase mitochondrial 1, mitochondrial precursor (EC 3.1.3.48) (EC 3.1.3.16) (PTEN-like phosphatase) (Phosphoinositide lipid phosphatase). | |||||
|
PTEN_HUMAN
|
||||||
| NC score | 0.164622 (rank : 11) | θ value | 0.0961366 (rank : 10) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P60484, O00633, O02679, Q6ICT7 | Gene names | PTEN, MMAC1, TEP1 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-3,4,5-trisphosphate 3-phosphatase and dual- specificity protein phosphatase PTEN (EC 3.1.3.67) (EC 3.1.3.16) (EC 3.1.3.48) (Phosphatase and tensin homolog) (Mutated in multiple advanced cancers 1). | |||||
|
PTEN_MOUSE
|
||||||
| NC score | 0.164621 (rank : 12) | θ value | 0.0961366 (rank : 11) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O08586, Q542G1 | Gene names | Pten, Mmac1 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-3,4,5-trisphosphate 3-phosphatase PTEN (EC 3.1.3.67) (Mutated in multiple advanced cancers 1). | |||||
|
CC14B_HUMAN
|
||||||
| NC score | 0.127533 (rank : 13) | θ value | 5.27518 (rank : 26) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O60729, O43183, O60730 | Gene names | CDC14B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dual specificity protein phosphatase CDC14B (EC 3.1.3.48) (EC 3.1.3.16) (CDC14 cell division cycle 14 homolog B). | |||||
|
CC14A_HUMAN
|
||||||
| NC score | 0.120060 (rank : 14) | θ value | 1.38821 (rank : 15) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9UNH5, O43171, O60727, O60728, Q8IXX0 | Gene names | CDC14A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dual specificity protein phosphatase CDC14A (EC 3.1.3.48) (EC 3.1.3.16) (CDC14 cell division cycle 14 homolog A). | |||||
|
CC14A_MOUSE
|
||||||
| NC score | 0.115125 (rank : 15) | θ value | 3.0926 (rank : 19) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q6GQT0, Q8BZ66 | Gene names | Cdc14a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dual specificity protein phosphatase CDC14A (EC 3.1.3.48) (EC 3.1.3.16) (CDC14 cell division cycle 14 homolog A). | |||||
|
CC14B_MOUSE
|
||||||
| NC score | 0.106316 (rank : 16) | θ value | θ > 10 (rank : 38) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q6PFY9, Q80WC4, Q8BLV5 | Gene names | Cdc14b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dual specificity protein phosphatase CDC14B (EC 3.1.3.48) (EC 3.1.3.16) (CDC14 cell division cycle 14 homolog B). | |||||
|
DUS2_MOUSE
|
||||||
| NC score | 0.068815 (rank : 17) | θ value | 0.365318 (rank : 12) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q05922, Q60640 | Gene names | Dusp2, Pac-1, Pac1 | |||
|
Domain Architecture |

|
|||||
| Description | Dual specificity protein phosphatase 2 (EC 3.1.3.48) (EC 3.1.3.16) (Dual specificity protein phosphatase PAC-1). | |||||
|
DUS16_HUMAN
|
||||||
| NC score | 0.066894 (rank : 18) | θ value | 2.36792 (rank : 17) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9BY84, Q9C0G3 | Gene names | DUSP16, KIAA1700, MKP7 | |||
|
Domain Architecture |

|
|||||
| Description | Dual specificity protein phosphatase 16 (EC 3.1.3.48) (EC 3.1.3.16) (Mitogen-activated protein kinase phosphatase 7) (MAP kinase phosphatase 7) (MKP-7). | |||||
|
DUS2_HUMAN
|
||||||
| NC score | 0.063267 (rank : 19) | θ value | 0.47712 (rank : 13) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q05923 | Gene names | DUSP2, PAC1 | |||
|
Domain Architecture |

|
|||||
| Description | Dual specificity protein phosphatase 2 (EC 3.1.3.48) (EC 3.1.3.16) (Dual specificity protein phosphatase PAC-1). | |||||
|
DUS8_MOUSE
|
||||||
| NC score | 0.054183 (rank : 20) | θ value | θ > 10 (rank : 39) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O09112 | Gene names | Dusp8, Nttp1 | |||
|
Domain Architecture |

|
|||||
| Description | Dual specificity protein phosphatase 8 (EC 3.1.3.48) (EC 3.1.3.16) (Neuronal tyrosine threonine phosphatase 1). | |||||
|
PTPRJ_HUMAN
|
||||||
| NC score | 0.051221 (rank : 21) | θ value | 0.62314 (rank : 14) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q12913, Q15255, Q8NHM2 | Gene names | PTPRJ, DEP1 | |||
|
Domain Architecture |

|
|||||
| Description | Receptor-type tyrosine-protein phosphatase eta precursor (EC 3.1.3.48) (Protein-tyrosine phosphatase eta) (R-PTP-eta) (HPTP eta) (Protein- tyrosine phosphatase receptor type J) (Density-enhanced phosphatase 1) (DEP-1) (CD148 antigen). | |||||
|
TP4A3_MOUSE
|
||||||
| NC score | 0.050798 (rank : 22) | θ value | θ > 10 (rank : 40) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9D658, O70275, Q3T9Z5, Q9CTC8 | Gene names | Ptp4a3, Prl3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein tyrosine phosphatase type IVA protein 3 (EC 3.1.3.48) (Protein-tyrosine phosphatase 4a3) (Protein-tyrosine phosphatase of regenerating liver 3) (PRL-3). | |||||
|
DUS1_MOUSE
|
||||||
| NC score | 0.049647 (rank : 23) | θ value | 2.36792 (rank : 18) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P28563 | Gene names | Dusp1, 3ch134, Mkp1, Ptpn10, Ptpn16 | |||
|
Domain Architecture |

|
|||||
| Description | Dual specificity protein phosphatase 1 (EC 3.1.3.48) (EC 3.1.3.16) (MAP kinase phosphatase 1) (MKP-1) (Protein-tyrosine phosphatase 3CH134) (Protein-tyrosine phosphatase ERP). | |||||
|
DUS1_HUMAN
|
||||||
| NC score | 0.049452 (rank : 24) | θ value | 8.99809 (rank : 33) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P28562, Q2V508 | Gene names | DUSP1, CL100, MKP1, PTPN10, VH1 | |||
|
Domain Architecture |

|
|||||
| Description | Dual specificity protein phosphatase 1 (EC 3.1.3.48) (EC 3.1.3.16) (MAP kinase phosphatase 1) (MKP-1) (Protein-tyrosine phosphatase CL100) (Dual specificity protein phosphatase hVH1). | |||||
|
PTPRJ_MOUSE
|
||||||
| NC score | 0.045319 (rank : 25) | θ value | 1.81305 (rank : 16) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q64455 | Gene names | Ptprj, Byp, Scc1 | |||
|
Domain Architecture |

|
|||||
| Description | Receptor-type tyrosine-protein phosphatase eta precursor (EC 3.1.3.48) (Protein-tyrosine phosphatase eta) (R-PTP-eta) (HPTP beta-like tyrosine phosphatase) (Protein-tyrosine phosphatase receptor type J) (Susceptibility to colon cancer 1). | |||||
|
PTPRA_MOUSE
|
||||||
| NC score | 0.034876 (rank : 26) | θ value | 6.88961 (rank : 28) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P18052, Q61808 | Gene names | Ptpra, Lrp, Ptpa | |||
|
Domain Architecture |

|
|||||
| Description | Receptor-type tyrosine-protein phosphatase alpha precursor (EC 3.1.3.48) (Protein-tyrosine phosphatase alpha) (R-PTP-alpha) (LCA- related phosphatase) (PTPTY-28). | |||||
|
PTPRA_HUMAN
|
||||||
| NC score | 0.034481 (rank : 27) | θ value | 6.88961 (rank : 27) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P18433, Q14513, Q7Z2I2, Q96TD9 | Gene names | PTPRA, PTPA | |||
|
Domain Architecture |

|
|||||
| Description | Receptor-type tyrosine-protein phosphatase alpha precursor (EC 3.1.3.48) (Protein-tyrosine phosphatase alpha) (R-PTP-alpha). | |||||
|
PTN14_HUMAN
|
||||||
| NC score | 0.024702 (rank : 28) | θ value | 4.03905 (rank : 24) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q15678 | Gene names | PTPN14, PEZ | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 14 (EC 3.1.3.48) (Protein-tyrosine phosphatase pez). | |||||
|
PTPRD_MOUSE
|
||||||
| NC score | 0.024532 (rank : 29) | θ value | 6.88961 (rank : 30) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q64487, Q64486, Q64488, Q64495 | Gene names | Ptprd | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Receptor-type tyrosine-protein phosphatase delta precursor (EC 3.1.3.48) (Protein-tyrosine phosphatase delta) (R-PTP-delta). | |||||
|
PTPRD_HUMAN
|
||||||
| NC score | 0.024332 (rank : 30) | θ value | 6.88961 (rank : 29) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 485 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P23468 | Gene names | PTPRD | |||
|
Domain Architecture |

|
|||||
| Description | Receptor-type tyrosine-protein phosphatase delta precursor (EC 3.1.3.48) (Protein-tyrosine phosphatase delta) (R-PTP-delta). | |||||
|
PTPRS_HUMAN
|
||||||
| NC score | 0.024224 (rank : 31) | θ value | 6.88961 (rank : 31) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 472 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q13332, O75255, O75870, Q15718, Q16341 | Gene names | PTPRS | |||
|
Domain Architecture |

|
|||||
| Description | Receptor-type tyrosine-protein phosphatase S precursor (EC 3.1.3.48) (R-PTP-S) (Protein-tyrosine phosphatase sigma) (R-PTP-sigma). | |||||
|
PTN3_HUMAN
|
||||||
| NC score | 0.021956 (rank : 32) | θ value | 8.99809 (rank : 35) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P26045 | Gene names | PTPN3, PTPH1 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 3 (EC 3.1.3.48) (Protein-tyrosine phosphatase H1) (PTP-H1). | |||||
|
K0408_HUMAN
|
||||||
| NC score | 0.020152 (rank : 33) | θ value | 8.99809 (rank : 34) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6ZU52, O43158, Q5TF20, Q7L2M2 | Gene names | KIAA0408 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0408. | |||||
|
CBX7_HUMAN
|
||||||
| NC score | 0.011766 (rank : 34) | θ value | 8.99809 (rank : 32) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O95931, Q86T17 | Gene names | CBX7 | |||
|
Domain Architecture |

|
|||||
| Description | Chromobox protein homolog 7. | |||||
|
AMOT_HUMAN
|
||||||
| NC score | 0.011262 (rank : 35) | θ value | 5.27518 (rank : 25) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 712 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q4VCS5, Q504X5, Q9HD27, Q9UPT1 | Gene names | AMOT, KIAA1071 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin. | |||||
|
BRD4_HUMAN
|
||||||
| NC score | 0.011241 (rank : 36) | θ value | 4.03905 (rank : 22) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 851 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O60885, O60433, Q96PD3 | Gene names | BRD4, HUNK1 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 4 (HUNK1 protein). | |||||
|
CASR_HUMAN
|
||||||
| NC score | 0.010070 (rank : 37) | θ value | 4.03905 (rank : 23) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P41180, Q13912, Q16108, Q16109, Q16110, Q16379, Q4PJ19 | Gene names | CASR, GPRC2A, PCAR1 | |||
|
Domain Architecture |

|
|||||
| Description | Extracellular calcium-sensing receptor precursor (CaSR) (Parathyroid Cell calcium-sensing receptor). | |||||
|
SLIT2_HUMAN
|
||||||
| NC score | 0.003741 (rank : 38) | θ value | 8.99809 (rank : 36) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 708 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O94813, O95710, Q9Y5Q7 | Gene names | SLIT2, SLIL3 | |||
|
Domain Architecture |

|
|||||
| Description | Slit homolog 2 protein precursor (Slit-2) [Contains: Slit homolog 2 protein N-product; Slit homolog 2 protein C-product]. | |||||
|
SLIT2_MOUSE
|
||||||
| NC score | 0.003735 (rank : 39) | θ value | 8.99809 (rank : 37) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 684 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9R1B9, Q9Z166 | Gene names | Slit2 | |||
|
Domain Architecture |

|
|||||
| Description | Slit homolog 2 protein precursor (Slit-2) [Contains: Slit homolog 2 protein N-product; Slit homolog 2 protein C-product]. | |||||
|
ZBT24_HUMAN
|
||||||
| NC score | 0.000785 (rank : 40) | θ value | 3.0926 (rank : 21) | |||
| Query Neighborhood Hits | 37 | Target Neighborhood Hits | 870 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O43167, Q5TED5, Q8N455 | Gene names | ZBTB24, KIAA0441, ZNF450 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger and BTB domain-containing protein 24 (Zinc finger protein 450). | |||||