Please be patient as the page loads
|
ANC1_HUMAN
|
||||||
| SwissProt Accessions | Q9H1A4, Q9BSE6, Q9H8D0 | Gene names | ANAPC1, TSG24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Anaphase-promoting complex subunit 1 (APC1) (Cyclosome subunit 1) (Protein Tsg24) (Mitotic checkpoint regulator). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
ANC1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9H1A4, Q9BSE6, Q9H8D0 | Gene names | ANAPC1, TSG24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Anaphase-promoting complex subunit 1 (APC1) (Cyclosome subunit 1) (Protein Tsg24) (Mitotic checkpoint regulator). | |||||
|
ANC1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.993552 (rank : 2) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P53995, Q8BP33, Q8C772 | Gene names | Anapc1, Mcpr, Tsg24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Anaphase-promoting complex subunit 1 (APC1) (Cyclosome subunit 1) (Protein Tsg24) (Mitotic checkpoint regulator). | |||||
|
PHC3_MOUSE
|
||||||
| θ value | 0.0148317 (rank : 3) | NC score | 0.079429 (rank : 3) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8CHP6, Q3TLV5 | Gene names | Phc3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polyhomeotic-like protein 3. | |||||
|
PHC3_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 4) | NC score | 0.079171 (rank : 4) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8NDX5, Q5HYF0, Q6NSG2, Q8NFT7, Q8NFZ1, Q8TBM2, Q9H971, Q9H9I4 | Gene names | PHC3, EDR3, PH3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polyhomeotic-like protein 3 (hPH3) (Homolog of polyhomeotic 3) (Early development regulatory protein 3). | |||||
|
RBP26_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 5) | NC score | 0.045142 (rank : 14) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q68DN6, Q6V1X0 | Gene names | RANBP2L6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ran-binding protein 2-like 6 (RanBP2L6). | |||||
|
COE1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 6) | NC score | 0.060347 (rank : 5) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UH73, Q8IW11 | Gene names | EBF1, COE1, EBF | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor COE1 (OE-1) (O/E-1) (Early B-cell factor). | |||||
|
COE1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 7) | NC score | 0.060347 (rank : 6) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q07802 | Gene names | Ebf1, Coe1, Ebf | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor COE1 (OE-1) (O/E-1) (Early B-cell factor). | |||||
|
TREF1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 8) | NC score | 0.024620 (rank : 22) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 651 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96PN7, Q7Z6T2, Q7Z6T3, Q9NQ72, Q9NQ73, Q9NUN9 | Gene names | TRERF1, RAPA, TREP132 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcriptional-regulating factor 1 (Transcriptional-regulating protein 132) (Zinc finger transcription factor TReP-132) (Zinc finger protein rapa). | |||||
|
FLOT2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 9) | NC score | 0.035586 (rank : 16) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14254 | Gene names | FLOT2, ESA1, M17S1 | |||
|
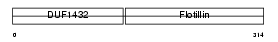
Domain Architecture |
|
|||||
| Description | Flotillin-2 (Epidermal surface antigen) (ESA). | |||||
|
YETS2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 10) | NC score | 0.026479 (rank : 21) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9ULM3, Q641P6, Q9NW96 | Gene names | YEATS2, KIAA1197 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YEATS domain-containing protein 2. | |||||
|
ANK1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 11) | NC score | 0.010740 (rank : 34) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P16157, O43400, Q13768, Q59FP2, Q8N604, Q99407 | Gene names | ANK1, ANK | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-1 (Erythrocyte ankyrin) (Ankyrin-R). | |||||
|
ANK1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 12) | NC score | 0.010723 (rank : 35) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 430 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q02357, P70440, P97446, P97941, Q3TZ35, Q3UH42, Q3UHP2, Q3UYY7, Q61302, Q61303, Q78E45 | Gene names | Ank1, Ank-1 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-1 (Erythrocyte ankyrin). | |||||
|
PMS2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 13) | NC score | 0.033121 (rank : 18) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P54279 | Gene names | Pms2 | |||
|
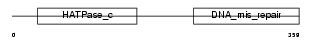
Domain Architecture |
|
|||||
| Description | PMS1 protein homolog 2 (DNA mismatch repair protein PMS2). | |||||
|
PSMD2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 14) | NC score | 0.050647 (rank : 9) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13200, Q12932, Q15321, Q96I12 | Gene names | PSMD2, TRAP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 26S proteasome non-ATPase regulatory subunit 2 (26S proteasome regulatory subunit RPN1) (26S proteasome regulatory subunit S2) (26S proteasome subunit p97) (Tumor necrosis factor type 1 receptor- associated protein 2) (55.11 protein). | |||||
|
PSMD2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 15) | NC score | 0.050362 (rank : 10) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8VDM4 | Gene names | Psmd2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 26S proteasome non-ATPase regulatory subunit 2 (26S proteasome regulatory subunit RPN1) (26S proteasome regulatory subunit S2) (26S proteasome subunit p97). | |||||
|
YAP1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 16) | NC score | 0.046964 (rank : 12) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P46937 | Gene names | YAP1, YAP65 | |||
|
Domain Architecture |

|
|||||
| Description | 65 kDa Yes-associated protein (YAP65). | |||||
|
COE3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 17) | NC score | 0.045135 (rank : 15) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9H4W6, Q5T6H9, Q9H4W5 | Gene names | EBF3, COE3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor COE3 (Early B-cell factor 3) (EBF-3) (Olf-1/EBF- like 2) (OE-2) (O/E-2). | |||||
|
COE3_MOUSE
|
||||||
| θ value | 3.0926 (rank : 18) | NC score | 0.045165 (rank : 13) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O08791, O08793 | Gene names | Ebf3, Coe3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor COE3 (Early B-cell factor 3) (EBF-3) (Olf-1/EBF- like 2) (OE-2) (O/E-2). | |||||
|
DCR1B_MOUSE
|
||||||
| θ value | 3.0926 (rank : 19) | NC score | 0.026982 (rank : 20) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8C7W7, Q3U4P2, Q6NXL4, Q8BN95, Q8BQS8, Q921S0 | Gene names | Dclre1b, Snm1b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA cross-link repair 1B protein. | |||||
|
FLOT2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 20) | NC score | 0.031262 (rank : 19) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q60634, Q6NS75 | Gene names | Flot2, Esa1, M17s1 | |||
|
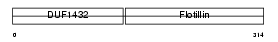
Domain Architecture |
|
|||||
| Description | Flotillin-2 (Epidermal surface antigen) (ESA). | |||||
|
YAP1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 21) | NC score | 0.033980 (rank : 17) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P46938, Q52KJ5 | Gene names | Yap1, Yap, Yap65 | |||
|
Domain Architecture |

|
|||||
| Description | 65 kDa Yes-associated protein (YAP65). | |||||
|
IF2P_HUMAN
|
||||||
| θ value | 4.03905 (rank : 22) | NC score | 0.011556 (rank : 33) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 889 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O60841, O95805, Q9UF81, Q9UMN7 | Gene names | EIF5B, IF2, KIAA0741 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 5B (eIF-5B) (Translation initiation factor IF-2). | |||||
|
LV1A_HUMAN
|
||||||
| θ value | 4.03905 (rank : 23) | NC score | 0.009767 (rank : 37) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P01699 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ig lambda chain V-I region VOR. | |||||
|
MAP1A_HUMAN
|
||||||
| θ value | 4.03905 (rank : 24) | NC score | 0.009885 (rank : 36) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P78559, O95643, Q12973, Q15882, Q9UJT4 | Gene names | MAP1A | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 1A (MAP 1A) (Proliferation-related protein p80) [Contains: MAP1 light chain LC2]. | |||||
|
CAR14_HUMAN
|
||||||
| θ value | 5.27518 (rank : 25) | NC score | 0.009740 (rank : 38) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 449 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9BXL6, Q9BVB5 | Gene names | CARD14, CARMA2 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 14 (CARD-containing MAGUK protein 2) (Carma 2). | |||||
|
HUWE1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 26) | NC score | 0.008230 (rank : 41) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 443 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q7Z6Z7, O15029, Q4G2Z2, Q5H961, Q6P4D0, Q8NG67, Q9BUI0, Q9HCJ4, Q9NSL6, Q9P0A9 | Gene names | HUWE1, KIAA0312, KIAA1578, UREB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT, UBA and WWE domain-containing protein 1 (EC 6.3.2.-) (E3 ubiquitin protein ligase URE-B1) (Mcl-1 ubiquitin ligase E3) (Mule) (ARF-binding protein 1) (ARF-BP1). | |||||
|
KIF6_HUMAN
|
||||||
| θ value | 5.27518 (rank : 27) | NC score | 0.006276 (rank : 44) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6ZMV9, Q2MDE3, Q2MDE4, Q5T8J6, Q6ZWE3, Q86T87, Q8WTV4 | Gene names | KIF6, C6orf102 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin-like protein KIF6. | |||||
|
REPS2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 28) | NC score | 0.023964 (rank : 23) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q80XA6 | Gene names | Reps2, Pob1 | |||
|
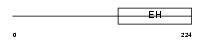
Domain Architecture |
|
|||||
| Description | RalBP1-associated Eps domain-containing protein 2 (RalBP1-interacting protein 2) (Partner of RalBP1). | |||||
|
SMD1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 29) | NC score | 0.053728 (rank : 7) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P62314, P13641 | Gene names | SNRPD1 | |||
|
Domain Architecture |
|
|||||
| Description | Small nuclear ribonucleoprotein Sm D1 (snRNP core protein D1) (Sm-D1) (Sm-D autoantigen). | |||||
|
SMD1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 30) | NC score | 0.053728 (rank : 8) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P62315, P13641, Q3UYM0 | Gene names | Snrpd1 | |||
|
Domain Architecture |
|
|||||
| Description | Small nuclear ribonucleoprotein Sm D1 (snRNP core protein D1) (Sm-D1) (Sm-D autoantigen). | |||||
|
TR102_MOUSE
|
||||||
| θ value | 5.27518 (rank : 31) | NC score | 0.006785 (rank : 43) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7M717, Q71QD0 | Gene names | Tas2r102, T2r51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Taste receptor type 2 member 102 (T2R102) (mT2R51) (STC9-7). | |||||
|
AQR_MOUSE
|
||||||
| θ value | 6.88961 (rank : 32) | NC score | 0.012847 (rank : 31) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8CFQ3, P97871, Q3U9N1, Q3ULE8, Q80TX8 | Gene names | Aqr, Kiaa0560 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intron-binding protein aquarius. | |||||
|
CDKA2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 33) | NC score | 0.019924 (rank : 26) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O75956 | Gene names | CDK2AP2, DOC1R | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-dependent kinase 2-associated protein 2 (CDK2-associated protein 2) (DOC-1-related protein) (DOC-1R). | |||||
|
ECM29_HUMAN
|
||||||
| θ value | 6.88961 (rank : 34) | NC score | 0.013895 (rank : 29) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q5VYK3, O15074, Q8WU82 | Gene names | ECM29, KIAA0368 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteasome-associated protein ECM29 homolog (Ecm29). | |||||
|
MBD5_HUMAN
|
||||||
| θ value | 6.88961 (rank : 35) | NC score | 0.017687 (rank : 27) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9P267, Q9NUV6 | Gene names | MBD5, KIAA1461 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 5 (Methyl-CpG-binding protein MBD5). | |||||
|
PCDGB_HUMAN
|
||||||
| θ value | 6.88961 (rank : 36) | NC score | 0.002213 (rank : 45) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y5H2, Q9Y5D8, Q9Y5D9 | Gene names | PCDHGA11 | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin gamma A11 precursor (PCDH-gamma-A11). | |||||
|
STCH_HUMAN
|
||||||
| θ value | 6.88961 (rank : 37) | NC score | 0.014408 (rank : 28) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P48723, Q8NE40 | Gene names | STCH | |||
|
Domain Architecture |
|
|||||
| Description | Stress 70 protein chaperone microsome-associated 60 kDa protein precursor (Microsomal stress 70 protein ATPase core). | |||||
|
ALEX_MOUSE
|
||||||
| θ value | 8.99809 (rank : 38) | NC score | 0.013486 (rank : 30) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6R0H6, Q8BIR3 | Gene names | Gnas, Gnas1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein ALEX (Alternative gene product encoded by XL-exon). | |||||
|
DCR1B_HUMAN
|
||||||
| θ value | 8.99809 (rank : 39) | NC score | 0.020439 (rank : 25) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H816, Q9H9E5 | Gene names | DCLRE1B, SNM1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA cross-link repair 1B protein (hSNM1B). | |||||
|
GAK_HUMAN
|
||||||
| θ value | 8.99809 (rank : 40) | NC score | 0.001077 (rank : 46) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1028 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O14976, Q9BVY6 | Gene names | GAK | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin G-associated kinase (EC 2.7.11.1). | |||||
|
RAPH1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 41) | NC score | 0.008730 (rank : 40) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 593 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q70E73, Q96Q37, Q9C0I2 | Gene names | RAPH1, ALS2CR9, KIAA1681, LPD, PREL2, RMO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras-associated and pleckstrin homology domains-containing protein 1 (RAPH1) (Lamellipodin) (Proline-rich EVH1 ligand 2) (PREL-2) (Protein RMO1) (Amyotrophic lateral sclerosis 2 chromosomal region candidate 9 gene protein). | |||||
|
STCH_MOUSE
|
||||||
| θ value | 8.99809 (rank : 42) | NC score | 0.012793 (rank : 32) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BM72, Q3TII6, Q9D1X5 | Gene names | Stch | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Stress 70 protein chaperone microsome-associated 60 kDa protein precursor (Microsomal stress 70 protein ATPase core). | |||||
|
TEX15_HUMAN
|
||||||
| θ value | 8.99809 (rank : 43) | NC score | 0.021525 (rank : 24) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9BXT5 | Gene names | TEX15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Testis-expressed sequence 15 protein. | |||||
|
UTX_HUMAN
|
||||||
| θ value | 8.99809 (rank : 44) | NC score | 0.009546 (rank : 39) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O15550 | Gene names | UTX | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitously transcribed X chromosome tetratricopeptide repeat protein (Ubiquitously transcribed TPR protein on the X chromosome). | |||||
|
YIPF5_MOUSE
|
||||||
| θ value | 8.99809 (rank : 45) | NC score | 0.050279 (rank : 11) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9EQQ2 | Gene names | Yipf5, Yip1a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein YIPF5 (YIP1 family member 5) (YPT-interacting protein 1 A). | |||||
|
ZDH19_MOUSE
|
||||||
| θ value | 8.99809 (rank : 46) | NC score | 0.007668 (rank : 42) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q810M5 | Gene names | Zdhhc19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable palmitoyltransferase ZDHHC19 (EC 2.3.1.-) (Zinc finger DHHC domain-containing protein 19) (DHHC-19). | |||||
|
ANC1_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9H1A4, Q9BSE6, Q9H8D0 | Gene names | ANAPC1, TSG24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Anaphase-promoting complex subunit 1 (APC1) (Cyclosome subunit 1) (Protein Tsg24) (Mitotic checkpoint regulator). | |||||
|
ANC1_MOUSE
|
||||||
| NC score | 0.993552 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P53995, Q8BP33, Q8C772 | Gene names | Anapc1, Mcpr, Tsg24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Anaphase-promoting complex subunit 1 (APC1) (Cyclosome subunit 1) (Protein Tsg24) (Mitotic checkpoint regulator). | |||||
|
PHC3_MOUSE
|
||||||
| NC score | 0.079429 (rank : 3) | θ value | 0.0148317 (rank : 3) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 194 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8CHP6, Q3TLV5 | Gene names | Phc3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polyhomeotic-like protein 3. | |||||
|
PHC3_HUMAN
|
||||||
| NC score | 0.079171 (rank : 4) | θ value | 0.0736092 (rank : 4) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8NDX5, Q5HYF0, Q6NSG2, Q8NFT7, Q8NFZ1, Q8TBM2, Q9H971, Q9H9I4 | Gene names | PHC3, EDR3, PH3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polyhomeotic-like protein 3 (hPH3) (Homolog of polyhomeotic 3) (Early development regulatory protein 3). | |||||
|
COE1_HUMAN
|
||||||
| NC score | 0.060347 (rank : 5) | θ value | 0.813845 (rank : 6) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UH73, Q8IW11 | Gene names | EBF1, COE1, EBF | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor COE1 (OE-1) (O/E-1) (Early B-cell factor). | |||||
|
COE1_MOUSE
|
||||||
| NC score | 0.060347 (rank : 6) | θ value | 0.813845 (rank : 7) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q07802 | Gene names | Ebf1, Coe1, Ebf | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor COE1 (OE-1) (O/E-1) (Early B-cell factor). | |||||
|
SMD1_HUMAN
|
||||||
| NC score | 0.053728 (rank : 7) | θ value | 5.27518 (rank : 29) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P62314, P13641 | Gene names | SNRPD1 | |||
|
Domain Architecture |

|
|||||
| Description | Small nuclear ribonucleoprotein Sm D1 (snRNP core protein D1) (Sm-D1) (Sm-D autoantigen). | |||||
|
SMD1_MOUSE
|
||||||
| NC score | 0.053728 (rank : 8) | θ value | 5.27518 (rank : 30) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P62315, P13641, Q3UYM0 | Gene names | Snrpd1 | |||
|
Domain Architecture |

|
|||||
| Description | Small nuclear ribonucleoprotein Sm D1 (snRNP core protein D1) (Sm-D1) (Sm-D autoantigen). | |||||
|
PSMD2_HUMAN
|
||||||
| NC score | 0.050647 (rank : 9) | θ value | 2.36792 (rank : 14) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13200, Q12932, Q15321, Q96I12 | Gene names | PSMD2, TRAP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 26S proteasome non-ATPase regulatory subunit 2 (26S proteasome regulatory subunit RPN1) (26S proteasome regulatory subunit S2) (26S proteasome subunit p97) (Tumor necrosis factor type 1 receptor- associated protein 2) (55.11 protein). | |||||
|
PSMD2_MOUSE
|
||||||
| NC score | 0.050362 (rank : 10) | θ value | 2.36792 (rank : 15) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8VDM4 | Gene names | Psmd2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 26S proteasome non-ATPase regulatory subunit 2 (26S proteasome regulatory subunit RPN1) (26S proteasome regulatory subunit S2) (26S proteasome subunit p97). | |||||
|
YIPF5_MOUSE
|
||||||
| NC score | 0.050279 (rank : 11) | θ value | 8.99809 (rank : 45) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9EQQ2 | Gene names | Yipf5, Yip1a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein YIPF5 (YIP1 family member 5) (YPT-interacting protein 1 A). | |||||
|
YAP1_HUMAN
|
||||||
| NC score | 0.046964 (rank : 12) | θ value | 2.36792 (rank : 16) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P46937 | Gene names | YAP1, YAP65 | |||
|
Domain Architecture |

|
|||||
| Description | 65 kDa Yes-associated protein (YAP65). | |||||
|
COE3_MOUSE
|
||||||
| NC score | 0.045165 (rank : 13) | θ value | 3.0926 (rank : 18) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O08791, O08793 | Gene names | Ebf3, Coe3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor COE3 (Early B-cell factor 3) (EBF-3) (Olf-1/EBF- like 2) (OE-2) (O/E-2). | |||||
|
RBP26_HUMAN
|
||||||
| NC score | 0.045142 (rank : 14) | θ value | 0.0736092 (rank : 5) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q68DN6, Q6V1X0 | Gene names | RANBP2L6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ran-binding protein 2-like 6 (RanBP2L6). | |||||
|
COE3_HUMAN
|
||||||
| NC score | 0.045135 (rank : 15) | θ value | 3.0926 (rank : 17) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9H4W6, Q5T6H9, Q9H4W5 | Gene names | EBF3, COE3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor COE3 (Early B-cell factor 3) (EBF-3) (Olf-1/EBF- like 2) (OE-2) (O/E-2). | |||||
|
FLOT2_HUMAN
|
||||||
| NC score | 0.035586 (rank : 16) | θ value | 1.38821 (rank : 9) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14254 | Gene names | FLOT2, ESA1, M17S1 | |||
|
Domain Architecture |

|
|||||
| Description | Flotillin-2 (Epidermal surface antigen) (ESA). | |||||
|
YAP1_MOUSE
|
||||||
| NC score | 0.033980 (rank : 17) | θ value | 3.0926 (rank : 21) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P46938, Q52KJ5 | Gene names | Yap1, Yap, Yap65 | |||
|
Domain Architecture |

|
|||||
| Description | 65 kDa Yes-associated protein (YAP65). | |||||
|
PMS2_MOUSE
|
||||||
| NC score | 0.033121 (rank : 18) | θ value | 2.36792 (rank : 13) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P54279 | Gene names | Pms2 | |||
|
Domain Architecture |

|
|||||
| Description | PMS1 protein homolog 2 (DNA mismatch repair protein PMS2). | |||||
|
FLOT2_MOUSE
|
||||||
| NC score | 0.031262 (rank : 19) | θ value | 3.0926 (rank : 20) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q60634, Q6NS75 | Gene names | Flot2, Esa1, M17s1 | |||
|
Domain Architecture |

|
|||||
| Description | Flotillin-2 (Epidermal surface antigen) (ESA). | |||||
|
DCR1B_MOUSE
|
||||||
| NC score | 0.026982 (rank : 20) | θ value | 3.0926 (rank : 19) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8C7W7, Q3U4P2, Q6NXL4, Q8BN95, Q8BQS8, Q921S0 | Gene names | Dclre1b, Snm1b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA cross-link repair 1B protein. | |||||
|
YETS2_HUMAN
|
||||||
| NC score | 0.026479 (rank : 21) | θ value | 1.81305 (rank : 10) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9ULM3, Q641P6, Q9NW96 | Gene names | YEATS2, KIAA1197 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YEATS domain-containing protein 2. | |||||
|
TREF1_HUMAN
|
||||||
| NC score | 0.024620 (rank : 22) | θ value | 0.813845 (rank : 8) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 651 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96PN7, Q7Z6T2, Q7Z6T3, Q9NQ72, Q9NQ73, Q9NUN9 | Gene names | TRERF1, RAPA, TREP132 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcriptional-regulating factor 1 (Transcriptional-regulating protein 132) (Zinc finger transcription factor TReP-132) (Zinc finger protein rapa). | |||||
|
REPS2_MOUSE
|
||||||
| NC score | 0.023964 (rank : 23) | θ value | 5.27518 (rank : 28) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q80XA6 | Gene names | Reps2, Pob1 | |||
|
Domain Architecture |

|
|||||
| Description | RalBP1-associated Eps domain-containing protein 2 (RalBP1-interacting protein 2) (Partner of RalBP1). | |||||
|
TEX15_HUMAN
|
||||||
| NC score | 0.021525 (rank : 24) | θ value | 8.99809 (rank : 43) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9BXT5 | Gene names | TEX15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Testis-expressed sequence 15 protein. | |||||
|
DCR1B_HUMAN
|
||||||
| NC score | 0.020439 (rank : 25) | θ value | 8.99809 (rank : 39) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H816, Q9H9E5 | Gene names | DCLRE1B, SNM1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA cross-link repair 1B protein (hSNM1B). | |||||
|
CDKA2_HUMAN
|
||||||
| NC score | 0.019924 (rank : 26) | θ value | 6.88961 (rank : 33) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O75956 | Gene names | CDK2AP2, DOC1R | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-dependent kinase 2-associated protein 2 (CDK2-associated protein 2) (DOC-1-related protein) (DOC-1R). | |||||
|
MBD5_HUMAN
|
||||||
| NC score | 0.017687 (rank : 27) | θ value | 6.88961 (rank : 35) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9P267, Q9NUV6 | Gene names | MBD5, KIAA1461 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 5 (Methyl-CpG-binding protein MBD5). | |||||
|
STCH_HUMAN
|
||||||
| NC score | 0.014408 (rank : 28) | θ value | 6.88961 (rank : 37) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P48723, Q8NE40 | Gene names | STCH | |||
|
Domain Architecture |

|
|||||
| Description | Stress 70 protein chaperone microsome-associated 60 kDa protein precursor (Microsomal stress 70 protein ATPase core). | |||||
|
ECM29_HUMAN
|
||||||
| NC score | 0.013895 (rank : 29) | θ value | 6.88961 (rank : 34) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q5VYK3, O15074, Q8WU82 | Gene names | ECM29, KIAA0368 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteasome-associated protein ECM29 homolog (Ecm29). | |||||
|
ALEX_MOUSE
|
||||||
| NC score | 0.013486 (rank : 30) | θ value | 8.99809 (rank : 38) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6R0H6, Q8BIR3 | Gene names | Gnas, Gnas1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein ALEX (Alternative gene product encoded by XL-exon). | |||||
|
AQR_MOUSE
|
||||||
| NC score | 0.012847 (rank : 31) | θ value | 6.88961 (rank : 32) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8CFQ3, P97871, Q3U9N1, Q3ULE8, Q80TX8 | Gene names | Aqr, Kiaa0560 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intron-binding protein aquarius. | |||||
|
STCH_MOUSE
|
||||||
| NC score | 0.012793 (rank : 32) | θ value | 8.99809 (rank : 42) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BM72, Q3TII6, Q9D1X5 | Gene names | Stch | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Stress 70 protein chaperone microsome-associated 60 kDa protein precursor (Microsomal stress 70 protein ATPase core). | |||||
|
IF2P_HUMAN
|
||||||
| NC score | 0.011556 (rank : 33) | θ value | 4.03905 (rank : 22) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 889 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O60841, O95805, Q9UF81, Q9UMN7 | Gene names | EIF5B, IF2, KIAA0741 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 5B (eIF-5B) (Translation initiation factor IF-2). | |||||
|
ANK1_HUMAN
|
||||||
| NC score | 0.010740 (rank : 34) | θ value | 2.36792 (rank : 11) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P16157, O43400, Q13768, Q59FP2, Q8N604, Q99407 | Gene names | ANK1, ANK | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-1 (Erythrocyte ankyrin) (Ankyrin-R). | |||||
|
ANK1_MOUSE
|
||||||
| NC score | 0.010723 (rank : 35) | θ value | 2.36792 (rank : 12) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 430 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q02357, P70440, P97446, P97941, Q3TZ35, Q3UH42, Q3UHP2, Q3UYY7, Q61302, Q61303, Q78E45 | Gene names | Ank1, Ank-1 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-1 (Erythrocyte ankyrin). | |||||
|
MAP1A_HUMAN
|
||||||
| NC score | 0.009885 (rank : 36) | θ value | 4.03905 (rank : 24) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P78559, O95643, Q12973, Q15882, Q9UJT4 | Gene names | MAP1A | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 1A (MAP 1A) (Proliferation-related protein p80) [Contains: MAP1 light chain LC2]. | |||||
|
LV1A_HUMAN
|
||||||
| NC score | 0.009767 (rank : 37) | θ value | 4.03905 (rank : 23) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P01699 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ig lambda chain V-I region VOR. | |||||
|
CAR14_HUMAN
|
||||||
| NC score | 0.009740 (rank : 38) | θ value | 5.27518 (rank : 25) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 449 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9BXL6, Q9BVB5 | Gene names | CARD14, CARMA2 | |||
|
Domain Architecture |

|
|||||
| Description | Caspase recruitment domain-containing protein 14 (CARD-containing MAGUK protein 2) (Carma 2). | |||||
|
UTX_HUMAN
|
||||||
| NC score | 0.009546 (rank : 39) | θ value | 8.99809 (rank : 44) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O15550 | Gene names | UTX | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitously transcribed X chromosome tetratricopeptide repeat protein (Ubiquitously transcribed TPR protein on the X chromosome). | |||||
|
RAPH1_HUMAN
|
||||||
| NC score | 0.008730 (rank : 40) | θ value | 8.99809 (rank : 41) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 593 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q70E73, Q96Q37, Q9C0I2 | Gene names | RAPH1, ALS2CR9, KIAA1681, LPD, PREL2, RMO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras-associated and pleckstrin homology domains-containing protein 1 (RAPH1) (Lamellipodin) (Proline-rich EVH1 ligand 2) (PREL-2) (Protein RMO1) (Amyotrophic lateral sclerosis 2 chromosomal region candidate 9 gene protein). | |||||
|
HUWE1_HUMAN
|
||||||
| NC score | 0.008230 (rank : 41) | θ value | 5.27518 (rank : 26) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 443 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q7Z6Z7, O15029, Q4G2Z2, Q5H961, Q6P4D0, Q8NG67, Q9BUI0, Q9HCJ4, Q9NSL6, Q9P0A9 | Gene names | HUWE1, KIAA0312, KIAA1578, UREB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT, UBA and WWE domain-containing protein 1 (EC 6.3.2.-) (E3 ubiquitin protein ligase URE-B1) (Mcl-1 ubiquitin ligase E3) (Mule) (ARF-binding protein 1) (ARF-BP1). | |||||
|
ZDH19_MOUSE
|
||||||
| NC score | 0.007668 (rank : 42) | θ value | 8.99809 (rank : 46) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q810M5 | Gene names | Zdhhc19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable palmitoyltransferase ZDHHC19 (EC 2.3.1.-) (Zinc finger DHHC domain-containing protein 19) (DHHC-19). | |||||
|
TR102_MOUSE
|
||||||
| NC score | 0.006785 (rank : 43) | θ value | 5.27518 (rank : 31) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7M717, Q71QD0 | Gene names | Tas2r102, T2r51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Taste receptor type 2 member 102 (T2R102) (mT2R51) (STC9-7). | |||||
|
KIF6_HUMAN
|
||||||
| NC score | 0.006276 (rank : 44) | θ value | 5.27518 (rank : 27) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6ZMV9, Q2MDE3, Q2MDE4, Q5T8J6, Q6ZWE3, Q86T87, Q8WTV4 | Gene names | KIF6, C6orf102 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin-like protein KIF6. | |||||
|
PCDGB_HUMAN
|
||||||
| NC score | 0.002213 (rank : 45) | θ value | 6.88961 (rank : 36) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y5H2, Q9Y5D8, Q9Y5D9 | Gene names | PCDHGA11 | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin gamma A11 precursor (PCDH-gamma-A11). | |||||
|
GAK_HUMAN
|
||||||
| NC score | 0.001077 (rank : 46) | θ value | 8.99809 (rank : 40) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1028 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O14976, Q9BVY6 | Gene names | GAK | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin G-associated kinase (EC 2.7.11.1). | |||||