Please be patient as the page loads
|
SYEP_HUMAN
|
||||||
| SwissProt Accessions | P07814 | Gene names | EPRS, GLNS, PARS, QPRS | |||
|
Domain Architecture |

|
|||||
| Description | Bifunctional aminoacyl-tRNA synthetase [Includes: Glutamyl-tRNA synthetase (EC 6.1.1.17) (Glutamate--tRNA ligase); Prolyl-tRNA synthetase (EC 6.1.1.15) (Proline--tRNA ligase)]. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
SYEP_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | P07814 | Gene names | EPRS, GLNS, PARS, QPRS | |||
|
Domain Architecture |

|
|||||
| Description | Bifunctional aminoacyl-tRNA synthetase [Includes: Glutamyl-tRNA synthetase (EC 6.1.1.17) (Glutamate--tRNA ligase); Prolyl-tRNA synthetase (EC 6.1.1.15) (Proline--tRNA ligase)]. | |||||
|
SYEP_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.965214 (rank : 2) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q8CGC7 | Gene names | Eprs, Qprs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bifunctional aminoacyl-tRNA synthetase [Includes: Glutamyl-tRNA synthetase (EC 6.1.1.17) (Glutamate--tRNA ligase); Prolyl-tRNA synthetase (EC 6.1.1.15) (Proline--tRNA ligase)]. | |||||
|
SYQ_HUMAN
|
||||||
| θ value | 8.53533e-67 (rank : 3) | NC score | 0.799268 (rank : 3) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P47897 | Gene names | QARS | |||
|
Domain Architecture |

|
|||||
| Description | Glutaminyl-tRNA synthetase (EC 6.1.1.18) (Glutamine--tRNA ligase) (GlnRS). | |||||
|
SYW_MOUSE
|
||||||
| θ value | 2.0648e-12 (rank : 4) | NC score | 0.404288 (rank : 4) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P32921, Q80ZY4, Q99J58, Q9DC65 | Gene names | Wars, Wrs | |||
|
Domain Architecture |

|
|||||
| Description | Tryptophanyl-tRNA synthetase (EC 6.1.1.2) (Tryptophan--tRNA ligase) (TrpRS). | |||||
|
SYW_HUMAN
|
||||||
| θ value | 1.25267e-09 (rank : 5) | NC score | 0.370530 (rank : 5) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P23381, P78535, Q9UDL3 | Gene names | WARS, WRS | |||
|
Domain Architecture |

|
|||||
| Description | Tryptophanyl-tRNA synthetase (EC 6.1.1.2) (Tryptophan--tRNA ligase) (TrpRS) (IFP53) (hWRS). | |||||
|
SYM_HUMAN
|
||||||
| θ value | 5.44631e-05 (rank : 6) | NC score | 0.213020 (rank : 7) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P56192, Q14895, Q53H14, Q96A15, Q96BZ0, Q9NSE0 | Gene names | MARS | |||
|
Domain Architecture |

|
|||||
| Description | Methionyl-tRNA synthetase (EC 6.1.1.10) (Methionine--tRNA ligase) (MetRS). | |||||
|
SYG_MOUSE
|
||||||
| θ value | 0.00102713 (rank : 7) | NC score | 0.165709 (rank : 10) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9CZD3, Q8VC67 | Gene names | Gars | |||
|
Domain Architecture |

|
|||||
| Description | Glycyl-tRNA synthetase (EC 6.1.1.14) (Glycine--tRNA ligase) (GlyRS). | |||||
|
SYM_MOUSE
|
||||||
| θ value | 0.00102713 (rank : 8) | NC score | 0.191728 (rank : 9) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q68FL6 | Gene names | Mars | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methionyl-tRNA synthetase (EC 6.1.1.10) (Methionine--tRNA ligase) (MetRS). | |||||
|
MCA3_MOUSE
|
||||||
| θ value | 0.00665767 (rank : 9) | NC score | 0.226003 (rank : 6) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9D1M4 | Gene names | Eef1e1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation elongation factor 1 epsilon-1 (Multisynthetase complex auxiliary component p18) (Elongation factor p18). | |||||
|
PTPRZ_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 10) | NC score | 0.032002 (rank : 33) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P23471 | Gene names | PTPRZ1, PTPRZ, PTPZ | |||
|
Domain Architecture |

|
|||||
| Description | Receptor-type tyrosine-protein phosphatase zeta precursor (EC 3.1.3.48) (R-PTP-zeta). | |||||
|
SYTC_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 11) | NC score | 0.087674 (rank : 15) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P26639, Q96FP5, Q9BWA6 | Gene names | TARS | |||
|
Domain Architecture |

|
|||||
| Description | Threonyl-tRNA synthetase, cytoplasmic (EC 6.1.1.3) (Threonine--tRNA ligase) (ThrRS). | |||||
|
SYTC_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 12) | NC score | 0.087605 (rank : 16) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9D0R2, Q3TL42, Q7TMQ2, Q8BMI6, Q9CX03 | Gene names | Tars | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Threonyl-tRNA synthetase, cytoplasmic (EC 6.1.1.3) (Threonine--tRNA ligase) (ThrRS). | |||||
|
MCA3_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 13) | NC score | 0.206086 (rank : 8) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O43324, Q5THS2 | Gene names | EEF1E1 | |||
|
Domain Architecture |
|
|||||
| Description | Eukaryotic translation elongation factor 1 epsilon-1 (Multisynthetase complex auxiliary component p18) (Elongation factor p18). | |||||
|
RRBP1_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 14) | NC score | 0.040605 (rank : 27) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1306 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9P2E9, O75300, O75301, Q5W165, Q96SB2, Q9BWP1, Q9H476 | Gene names | RRBP1, KIAA1398 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (180 kDa ribosome receptor homolog) (ES/130-related protein). | |||||
|
SYG_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 15) | NC score | 0.147897 (rank : 11) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P41250, Q969Y1 | Gene names | GARS | |||
|
Domain Architecture |

|
|||||
| Description | Glycyl-tRNA synthetase (EC 6.1.1.14) (Glycine--tRNA ligase) (GlyRS). | |||||
|
SYPM_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 16) | NC score | 0.115314 (rank : 13) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8CFI5, Q8R0C4 | Gene names | Pars2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable prolyl-tRNA synthetase, mitochondrial precursor (EC 6.1.1.15) (Proline--tRNA ligase) (ProRS). | |||||
|
ZN217_HUMAN
|
||||||
| θ value | 0.125558 (rank : 17) | NC score | 0.009921 (rank : 67) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 905 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O75362 | Gene names | ZNF217, ZABC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 217. | |||||
|
RRBP1_MOUSE
|
||||||
| θ value | 0.163984 (rank : 18) | NC score | 0.033558 (rank : 32) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1650 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q99PL5, Q99PK5, Q99PK6, Q99PK7, Q99PK8, Q99PK9, Q99PL0, Q99PL1, Q99PL2, Q99PL3, Q99PL4, Q9CS20 | Gene names | Rrbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (mRRp). | |||||
|
SYPM_HUMAN
|
||||||
| θ value | 0.163984 (rank : 19) | NC score | 0.089815 (rank : 14) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q7L3T8, Q9H6S5, Q9UFT1 | Gene names | PARS2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable prolyl-tRNA synthetase, mitochondrial precursor (EC 6.1.1.15) (Proline--tRNA ligase) (ProRS). | |||||
|
MCA2_HUMAN
|
||||||
| θ value | 0.21417 (rank : 20) | NC score | 0.135605 (rank : 12) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q13155, Q96CZ5, Q9P1L2 | Gene names | JTV1 | |||
|
Domain Architecture |
|
|||||
| Description | Multisynthetase complex auxiliary component p38 (Protein JTV-1). | |||||
|
GP155_HUMAN
|
||||||
| θ value | 0.279714 (rank : 21) | NC score | 0.071812 (rank : 20) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7Z3F1, Q53SJ3, Q53TA8, Q69YG8, Q86SP9, Q8N261, Q8N639, Q8N8K3, Q96MV6 | Gene names | GPR155, PGR22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Integral membrane protein GPR155 (G-protein coupled receptor PGR22). | |||||
|
LAMA2_MOUSE
|
||||||
| θ value | 0.279714 (rank : 22) | NC score | 0.018554 (rank : 44) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1198 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q60675, Q05003, Q64061 | Gene names | Lama2 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-2 chain precursor (Laminin M chain) (Merosin heavy chain). | |||||
|
SYH_MOUSE
|
||||||
| θ value | 0.279714 (rank : 23) | NC score | 0.083006 (rank : 17) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q61035 | Gene names | Hars | |||
|
Domain Architecture |

|
|||||
| Description | Histidyl-tRNA synthetase (EC 6.1.1.21) (Histidine--tRNA ligase) (HisRS). | |||||
|
NCF2_HUMAN
|
||||||
| θ value | 0.365318 (rank : 24) | NC score | 0.053010 (rank : 21) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 229 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P19878, Q2PP06, Q8NFC7, Q9BV51 | Gene names | NCF2 | |||
|
Domain Architecture |

|
|||||
| Description | Neutrophil cytosol factor 2 (NCF-2) (Neutrophil NADPH oxidase factor 2) (67 kDa neutrophil oxidase factor) (p67-phox) (NOXA2). | |||||
|
CN145_HUMAN
|
||||||
| θ value | 0.813845 (rank : 25) | NC score | 0.018772 (rank : 41) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1196 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q6ZU80, Q96ML4 | Gene names | C14orf145 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf145. | |||||
|
SYH_HUMAN
|
||||||
| θ value | 0.813845 (rank : 26) | NC score | 0.080436 (rank : 18) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P12081 | Gene names | HARS, HRS | |||
|
Domain Architecture |

|
|||||
| Description | Histidyl-tRNA synthetase (EC 6.1.1.21) (Histidine--tRNA ligase) (HisRS). | |||||
|
CD2AP_MOUSE
|
||||||
| θ value | 1.38821 (rank : 27) | NC score | 0.044815 (rank : 26) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 336 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9JLQ0, O88903, Q8K4Z1, Q8VCI9 | Gene names | Cd2ap, Mets1 | |||
|
Domain Architecture |
|
|||||
| Description | CD2-associated protein (Mesenchyme-to-epithelium transition protein with SH3 domains 1) (METS-1). | |||||
|
ESCO1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 28) | NC score | 0.035430 (rank : 30) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 361 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q5FWF5, Q69YG4, Q69YS3, Q6IMD7, Q8N3Z5, Q8NBG2, Q96PX7 | Gene names | ESCO1, EFO1, KIAA1911 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyltransferase ESCO1 (EC 2.3.1.-) (Establishment of cohesion 1 homolog 1) (ECO1 homolog 1) (ESO1 homolog 1) (Establishment factor- like protein 1) (EFO1p) (hEFO1) (CTF7 homolog 1). | |||||
|
SRBS1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 29) | NC score | 0.023995 (rank : 39) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q62417, Q80TF8, Q8BZI3, Q8K3Y2, Q921F8, Q9Z0Z8, Q9Z0Z9 | Gene names | Sorbs1, Kiaa1296, Sh3d5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sorbin and SH3 domain-containing protein 1 (Ponsin) (c-Cbl-associated protein) (CAP) (SH3 domain protein 5) (SH3P12). | |||||
|
SYCC_MOUSE
|
||||||
| θ value | 1.81305 (rank : 30) | NC score | 0.046042 (rank : 25) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9ER72, O88303, Q8BP81, Q8CAY7, Q8CCE3, Q8K0S4, Q9ER68 | Gene names | Cars | |||
|
Domain Architecture |

|
|||||
| Description | Cysteinyl-tRNA synthetase, cytoplasmic (EC 6.1.1.16) (Cysteine--tRNA ligase) (CysRS). | |||||
|
LZTS1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 31) | NC score | 0.027772 (rank : 36) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 540 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P60853 | Gene names | Lzts1, Fez1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine zipper putative tumor suppressor 1 (F37/Esophageal cancer- related gene-coding leucine-zipper motif) (Fez1). | |||||
|
MCES_MOUSE
|
||||||
| θ value | 2.36792 (rank : 32) | NC score | 0.040151 (rank : 28) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9D0L8, Q3V3U9, Q6ZQC6, Q9D5F1 | Gene names | Rnmt, Kiaa0398 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | mRNA cap guanine-N7 methyltransferase (EC 2.1.1.56) (mRNA (guanine- N(7)-)-methyltransferase) (RG7MT1) (mRNA cap methyltransferase). | |||||
|
MYO6_HUMAN
|
||||||
| θ value | 2.36792 (rank : 33) | NC score | 0.017412 (rank : 46) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 727 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9UM54, Q9BZZ7, Q9UEG2 | Gene names | MYO6, KIAA0389 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-6 (Myosin VI). | |||||
|
NRBF2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 34) | NC score | 0.031671 (rank : 34) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96F24, Q86UR2, Q96NP6, Q9H0S9, Q9H2I2 | Gene names | NRBF2, COPR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor-binding factor 2 (NRBF-2) (Comodulator of PPAR and RXR). | |||||
|
CD2L5_MOUSE
|
||||||
| θ value | 3.0926 (rank : 35) | NC score | 0.002177 (rank : 70) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1011 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q69ZA1, Q80V11, Q8BZG1, Q8K0A4 | Gene names | Cdc2l5, Kiaa1791 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle 2-like protein kinase 5 (EC 2.7.11.22) (CDC2- related protein kinase 5). | |||||
|
IWS1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 36) | NC score | 0.017754 (rank : 45) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
KMHN1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 37) | NC score | 0.014208 (rank : 56) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 949 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q3V125, Q3TTQ8 | Gene names | Gm172 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cancer/testis antigen KM-HN-1 homolog. | |||||
|
SYV_MOUSE
|
||||||
| θ value | 3.0926 (rank : 38) | NC score | 0.049778 (rank : 23) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Z1Q9, Q9QUN2 | Gene names | Vars, Bat6, G7a, Vars2 | |||
|
Domain Architecture |
|
|||||
| Description | Valyl-tRNA synthetase (EC 6.1.1.9) (Valine--tRNA ligase) (ValRS) (Protein G7a). | |||||
|
XYLT1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 39) | NC score | 0.016814 (rank : 49) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q86Y38, Q9H1B6 | Gene names | XYLT1, XT1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Xylosyltransferase 1 (EC 2.4.2.26) (Xylosyltransferase I) (XylT-I) (XT-I) (Peptide O-xylosyltransferase 1). | |||||
|
DOCK4_HUMAN
|
||||||
| θ value | 4.03905 (rank : 40) | NC score | 0.016356 (rank : 51) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 237 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8N1I0, O14584, O94824, Q8NB45 | Gene names | DOCK4, KIAA0716 | |||
|
Domain Architecture |

|
|||||
| Description | Dedicator of cytokinesis protein 4. | |||||
|
DOCK4_MOUSE
|
||||||
| θ value | 4.03905 (rank : 41) | NC score | 0.016289 (rank : 52) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P59764 | Gene names | Dock4, Kiaa0716 | |||
|
Domain Architecture |

|
|||||
| Description | Dedicator of cytokinesis protein 4. | |||||
|
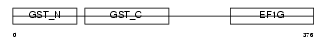
EF1G_HUMAN
|
||||||
| θ value | 4.03905 (rank : 42) | NC score | 0.048473 (rank : 24) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P26641, Q6PJ62, Q6PK31, Q96CU2, Q9P196 | Gene names | EEF1G, EF1G | |||
|
Domain Architecture |
|
|||||
| Description | Elongation factor 1-gamma (EF-1-gamma) (eEF-1B gamma). | |||||
|
MECR_MOUSE
|
||||||
| θ value | 4.03905 (rank : 43) | NC score | 0.017221 (rank : 47) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9DCS3, Q99L39 | Gene names | Mecr, Nrbf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trans-2-enoyl-CoA reductase, mitochondrial precursor (EC 1.3.1.38). | |||||
|
MYH13_HUMAN
|
||||||
| θ value | 4.03905 (rank : 44) | NC score | 0.013815 (rank : 57) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1504 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9UKX3, O95252 | Gene names | MYH13 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-13 (Myosin heavy chain, skeletal muscle, extraocular) (MyHC- eo). | |||||
|
RBM28_MOUSE
|
||||||
| θ value | 4.03905 (rank : 45) | NC score | 0.011946 (rank : 61) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8CGC6, Q8BG25, Q8VEJ8, Q9CS22, Q9CSE6 | Gene names | Rbm28 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein 28 (RNA-binding motif protein 28). | |||||
|
REST_MOUSE
|
||||||
| θ value | 4.03905 (rank : 46) | NC score | 0.012659 (rank : 60) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1506 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q922J3, Q571L7 | Gene names | Rsn, Kiaa4046 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Restin. | |||||
|
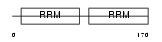
SF3B4_HUMAN
|
||||||
| θ value | 4.03905 (rank : 47) | NC score | 0.009769 (rank : 68) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q15427 | Gene names | SF3B4, SAP49 | |||
|
Domain Architecture |
|
|||||
| Description | Splicing factor 3B subunit 4 (Spliceosome-associated protein 49) (SAP 49) (SF3b50) (Pre-mRNA-splicing factor SF3b 49 kDa subunit). | |||||
|
AFF2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 48) | NC score | 0.014578 (rank : 55) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P51816, O43786, O60215, P78407, Q13521, Q14323, Q9UNA5 | Gene names | AFF2, FMR2, OX19 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 2 (Fragile X mental retardation 2 protein) (Protein FMR-2) (FMR2P) (Protein Ox19) (Fragile X E mental retardation syndrome protein). | |||||
|
ATRX_MOUSE
|
||||||
| θ value | 5.27518 (rank : 49) | NC score | 0.015365 (rank : 53) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 682 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q61687 | Gene names | Atrx, Hp1bp2, Xnp | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ATRX (EC 3.6.1.-) (ATP-dependent helicase ATRX) (X-linked nuclear protein) (Heterochromatin protein 2) (HP1 alpha-interacting protein) (HP1-BP38 protein). | |||||
|
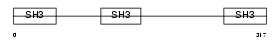
CD2AP_HUMAN
|
||||||
| θ value | 5.27518 (rank : 50) | NC score | 0.034412 (rank : 31) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y5K6, Q9UG97 | Gene names | CD2AP | |||
|
Domain Architecture |
|
|||||
| Description | CD2-associated protein (Cas ligand with multiple SH3 domains) (Adapter protein CMS). | |||||
|
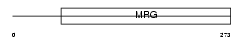
MO4L2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 51) | NC score | 0.028307 (rank : 35) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q15014, Q567V0, Q8J026 | Gene names | MORF4L2, KIAA0026, MRGX | |||
|
Domain Architecture |
|
|||||
| Description | Mortality factor 4-like protein 2 (MORF-related gene X protein) (Transcription factor-like protein MRGX) (MSL3-2 protein). | |||||
|
MYH7_HUMAN
|
||||||
| θ value | 5.27518 (rank : 52) | NC score | 0.011777 (rank : 62) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P12883, Q14836, Q14837, Q14904, Q16579, Q92679, Q9H1D5, Q9UMM8 | Gene names | MYH7, MYHCB | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle beta isoform (MyHC-beta). | |||||
|
SCG2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 53) | NC score | 0.036679 (rank : 29) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q03517, Q80Y79, Q9CW80 | Gene names | Scg2, Chgc, Scg-2 | |||
|
Domain Architecture |

|
|||||
| Description | Secretogranin-2 precursor (Secretogranin II) (SgII) (Chromogranin C) [Contains: Secretoneurin (SN)]. | |||||
|
TBC14_HUMAN
|
||||||
| θ value | 5.27518 (rank : 54) | NC score | 0.013404 (rank : 59) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9P2M4, Q8IW15 | Gene names | TBC1D14, KIAA1322 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 14. | |||||
|
CKAP2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 55) | NC score | 0.021200 (rank : 40) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q3V1H1, Q66LN2, Q8BSF0 | Gene names | Ckap2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoskeleton-associated protein 2. | |||||
|
IWS1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 56) | NC score | 0.013691 (rank : 58) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
MATR3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 57) | NC score | 0.016773 (rank : 50) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P43243, Q9UHW0, Q9UQ27 | Gene names | MATR3, KIAA0723 | |||
|
Domain Architecture |

|
|||||
| Description | Matrin-3. | |||||
|
NIPBL_HUMAN
|
||||||
| θ value | 6.88961 (rank : 58) | NC score | 0.025296 (rank : 37) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
PRG4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 59) | NC score | 0.024422 (rank : 38) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q92954, Q6DNC4, Q6DNC5, Q6ZMZ5, Q9BX49 | Gene names | PRG4, MSF, SZP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
RRP5_HUMAN
|
||||||
| θ value | 6.88961 (rank : 60) | NC score | 0.018681 (rank : 42) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 291 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q14690, Q86VQ8 | Gene names | PDCD11, KIAA0185 | |||
|
Domain Architecture |

|
|||||
| Description | RRP5 protein homolog (Programmed cell death protein 11). | |||||
|
SYNE1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 61) | NC score | 0.011305 (rank : 64) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1484 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8NF91, O94890, Q5JV19, Q5JV22, Q8N9P7, Q8TCP1, Q8WWW6, Q8WWW7, Q8WXF6, Q96N17, Q9C0A7, Q9H525, Q9H526, Q9NS36, Q9NU50, Q9UJ06, Q9UJ07, Q9ULF8 | Gene names | SYNE1, KIAA0796, KIAA1262, KIAA1756, MYNE1 | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-1 (Nuclear envelope spectrin repeat protein 1) (Synaptic nuclear envelope protein 1) (Syne-1) (Myocyte nuclear envelope protein 1) (Myne-1) (Enaptin). | |||||
|
TITIN_HUMAN
|
||||||
| θ value | 6.88961 (rank : 62) | NC score | 0.001035 (rank : 71) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 2923 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8WZ42, Q10465, Q10466, Q15598, Q2XUS3, Q32Q60, Q4U1Z6, Q6NSG0, Q6PDB1, Q6PJP0, Q7KYM2, Q7KYN4, Q7KYN5, Q7LDM3, Q7Z2X3, Q8TCG8, Q8WZ51, Q8WZ52, Q8WZ53, Q8WZB3, Q92761, Q92762, Q9UD97, Q9UP84, Q9Y6L9 | Gene names | TTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Titin (EC 2.7.11.1) (Connectin) (Rhabdomyosarcoma antigen MU-RMS- 40.14). | |||||
|
CDC7_HUMAN
|
||||||
| θ value | 8.99809 (rank : 63) | NC score | 0.004339 (rank : 69) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 652 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O00311, O00558 | Gene names | CDC7, CDC7L1 | |||
|
Domain Architecture |

|
|||||
| Description | Cell division cycle 7-related protein kinase (EC 2.7.11.1) (CDC7- related kinase) (HsCdc7) (huCdc7). | |||||
|
CF182_HUMAN
|
||||||
| θ value | 8.99809 (rank : 64) | NC score | 0.014581 (rank : 54) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 474 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8IYX8 | Gene names | C6orf182 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf182. | |||||
|
CKAP2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 65) | NC score | 0.018660 (rank : 43) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8WWK9, Q3KRA5, Q5VXB4, Q8IWV5, Q8IWV6, Q96FH9, Q9H012, Q9H0D0, Q9H988, Q9HC49, Q9NVG4 | Gene names | CKAP2, LB1, TMAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoskeleton-associated protein 2 (Tumor-associated microtubule- associated protein) (CTCL tumor antigen se20-10). | |||||
|
FEZ2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 66) | NC score | 0.011663 (rank : 63) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UHY8, Q76LN0, Q99690 | Gene names | FEZ2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fasciculation and elongation protein zeta 2 (Zygin-2) (Zygin II). | |||||
|
GOGA4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 67) | NC score | 0.010691 (rank : 65) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 2034 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q13439, Q13270, Q13654, Q14436 | Gene names | GOLGA4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (Trans-Golgi p230) (256 kDa golgin) (Golgin-245) (Protein 72.1). | |||||
|
MAP1A_HUMAN
|
||||||
| θ value | 8.99809 (rank : 68) | NC score | 0.017046 (rank : 48) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P78559, O95643, Q12973, Q15882, Q9UJT4 | Gene names | MAP1A | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 1A (MAP 1A) (Proliferation-related protein p80) [Contains: MAP1 light chain LC2]. | |||||
|
PTN6_HUMAN
|
||||||
| θ value | 8.99809 (rank : 69) | NC score | 0.010068 (rank : 66) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 234 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P29350, Q969V8 | Gene names | PTPN6, HCP, PTP1C | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 6 (EC 3.1.3.48) (Protein-tyrosine phosphatase 1C) (PTP-1C) (Hematopoietic cell protein-tyrosine phosphatase) (SH-PTP1) (Protein-tyrosine phosphatase SHP-1). | |||||
|
ZN583_HUMAN
|
||||||
| θ value | 8.99809 (rank : 70) | NC score | -0.000366 (rank : 72) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 774 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96ND8, O14850, Q2NKK3 | Gene names | ZNF583 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 583 (Zinc finger protein L3-5). | |||||
|
BASP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 71) | NC score | 0.076421 (rank : 19) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P80723, O43596, Q5U0S0 | Gene names | BASP1, NAP22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
TRDN_HUMAN
|
||||||
| θ value | θ > 10 (rank : 72) | NC score | 0.050923 (rank : 22) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 542 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q13061 | Gene names | TRDN | |||
|
Domain Architecture |

|
|||||
| Description | Triadin. | |||||
|
SYEP_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | P07814 | Gene names | EPRS, GLNS, PARS, QPRS | |||
|
Domain Architecture |

|
|||||
| Description | Bifunctional aminoacyl-tRNA synthetase [Includes: Glutamyl-tRNA synthetase (EC 6.1.1.17) (Glutamate--tRNA ligase); Prolyl-tRNA synthetase (EC 6.1.1.15) (Proline--tRNA ligase)]. | |||||
|
SYEP_MOUSE
|
||||||
| NC score | 0.965214 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q8CGC7 | Gene names | Eprs, Qprs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bifunctional aminoacyl-tRNA synthetase [Includes: Glutamyl-tRNA synthetase (EC 6.1.1.17) (Glutamate--tRNA ligase); Prolyl-tRNA synthetase (EC 6.1.1.15) (Proline--tRNA ligase)]. | |||||
|
SYQ_HUMAN
|
||||||
| NC score | 0.799268 (rank : 3) | θ value | 8.53533e-67 (rank : 3) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P47897 | Gene names | QARS | |||
|
Domain Architecture |

|
|||||
| Description | Glutaminyl-tRNA synthetase (EC 6.1.1.18) (Glutamine--tRNA ligase) (GlnRS). | |||||
|
SYW_MOUSE
|
||||||
| NC score | 0.404288 (rank : 4) | θ value | 2.0648e-12 (rank : 4) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P32921, Q80ZY4, Q99J58, Q9DC65 | Gene names | Wars, Wrs | |||
|
Domain Architecture |

|
|||||
| Description | Tryptophanyl-tRNA synthetase (EC 6.1.1.2) (Tryptophan--tRNA ligase) (TrpRS). | |||||
|
SYW_HUMAN
|
||||||
| NC score | 0.370530 (rank : 5) | θ value | 1.25267e-09 (rank : 5) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P23381, P78535, Q9UDL3 | Gene names | WARS, WRS | |||
|
Domain Architecture |

|
|||||
| Description | Tryptophanyl-tRNA synthetase (EC 6.1.1.2) (Tryptophan--tRNA ligase) (TrpRS) (IFP53) (hWRS). | |||||
|
MCA3_MOUSE
|
||||||
| NC score | 0.226003 (rank : 6) | θ value | 0.00665767 (rank : 9) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9D1M4 | Gene names | Eef1e1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation elongation factor 1 epsilon-1 (Multisynthetase complex auxiliary component p18) (Elongation factor p18). | |||||
|
SYM_HUMAN
|
||||||
| NC score | 0.213020 (rank : 7) | θ value | 5.44631e-05 (rank : 6) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P56192, Q14895, Q53H14, Q96A15, Q96BZ0, Q9NSE0 | Gene names | MARS | |||
|
Domain Architecture |

|
|||||
| Description | Methionyl-tRNA synthetase (EC 6.1.1.10) (Methionine--tRNA ligase) (MetRS). | |||||
|
MCA3_HUMAN
|
||||||
| NC score | 0.206086 (rank : 8) | θ value | 0.0961366 (rank : 13) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O43324, Q5THS2 | Gene names | EEF1E1 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation elongation factor 1 epsilon-1 (Multisynthetase complex auxiliary component p18) (Elongation factor p18). | |||||
|
SYM_MOUSE
|
||||||
| NC score | 0.191728 (rank : 9) | θ value | 0.00102713 (rank : 8) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q68FL6 | Gene names | Mars | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methionyl-tRNA synthetase (EC 6.1.1.10) (Methionine--tRNA ligase) (MetRS). | |||||
|
SYG_MOUSE
|
||||||
| NC score | 0.165709 (rank : 10) | θ value | 0.00102713 (rank : 7) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9CZD3, Q8VC67 | Gene names | Gars | |||
|
Domain Architecture |

|
|||||
| Description | Glycyl-tRNA synthetase (EC 6.1.1.14) (Glycine--tRNA ligase) (GlyRS). | |||||
|
SYG_HUMAN
|
||||||
| NC score | 0.147897 (rank : 11) | θ value | 0.0961366 (rank : 15) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P41250, Q969Y1 | Gene names | GARS | |||
|
Domain Architecture |

|
|||||
| Description | Glycyl-tRNA synthetase (EC 6.1.1.14) (Glycine--tRNA ligase) (GlyRS). | |||||
|
MCA2_HUMAN
|
||||||
| NC score | 0.135605 (rank : 12) | θ value | 0.21417 (rank : 20) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q13155, Q96CZ5, Q9P1L2 | Gene names | JTV1 | |||
|
Domain Architecture |

|
|||||
| Description | Multisynthetase complex auxiliary component p38 (Protein JTV-1). | |||||
|
SYPM_MOUSE
|
||||||
| NC score | 0.115314 (rank : 13) | θ value | 0.0961366 (rank : 16) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8CFI5, Q8R0C4 | Gene names | Pars2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable prolyl-tRNA synthetase, mitochondrial precursor (EC 6.1.1.15) (Proline--tRNA ligase) (ProRS). | |||||
|
SYPM_HUMAN
|
||||||
| NC score | 0.089815 (rank : 14) | θ value | 0.163984 (rank : 19) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q7L3T8, Q9H6S5, Q9UFT1 | Gene names | PARS2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable prolyl-tRNA synthetase, mitochondrial precursor (EC 6.1.1.15) (Proline--tRNA ligase) (ProRS). | |||||
|
SYTC_HUMAN
|
||||||
| NC score | 0.087674 (rank : 15) | θ value | 0.0563607 (rank : 11) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P26639, Q96FP5, Q9BWA6 | Gene names | TARS | |||
|
Domain Architecture |

|
|||||
| Description | Threonyl-tRNA synthetase, cytoplasmic (EC 6.1.1.3) (Threonine--tRNA ligase) (ThrRS). | |||||
|
SYTC_MOUSE
|
||||||
| NC score | 0.087605 (rank : 16) | θ value | 0.0563607 (rank : 12) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9D0R2, Q3TL42, Q7TMQ2, Q8BMI6, Q9CX03 | Gene names | Tars | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Threonyl-tRNA synthetase, cytoplasmic (EC 6.1.1.3) (Threonine--tRNA ligase) (ThrRS). | |||||
|
SYH_MOUSE
|
||||||
| NC score | 0.083006 (rank : 17) | θ value | 0.279714 (rank : 23) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q61035 | Gene names | Hars | |||
|
Domain Architecture |

|
|||||
| Description | Histidyl-tRNA synthetase (EC 6.1.1.21) (Histidine--tRNA ligase) (HisRS). | |||||
|
SYH_HUMAN
|
||||||
| NC score | 0.080436 (rank : 18) | θ value | 0.813845 (rank : 26) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P12081 | Gene names | HARS, HRS | |||
|
Domain Architecture |

|
|||||
| Description | Histidyl-tRNA synthetase (EC 6.1.1.21) (Histidine--tRNA ligase) (HisRS). | |||||
|
BASP_HUMAN
|
||||||
| NC score | 0.076421 (rank : 19) | θ value | θ > 10 (rank : 71) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P80723, O43596, Q5U0S0 | Gene names | BASP1, NAP22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
GP155_HUMAN
|
||||||
| NC score | 0.071812 (rank : 20) | θ value | 0.279714 (rank : 21) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7Z3F1, Q53SJ3, Q53TA8, Q69YG8, Q86SP9, Q8N261, Q8N639, Q8N8K3, Q96MV6 | Gene names | GPR155, PGR22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Integral membrane protein GPR155 (G-protein coupled receptor PGR22). | |||||
|
NCF2_HUMAN
|
||||||
| NC score | 0.053010 (rank : 21) | θ value | 0.365318 (rank : 24) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 229 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P19878, Q2PP06, Q8NFC7, Q9BV51 | Gene names | NCF2 | |||
|
Domain Architecture |

|
|||||
| Description | Neutrophil cytosol factor 2 (NCF-2) (Neutrophil NADPH oxidase factor 2) (67 kDa neutrophil oxidase factor) (p67-phox) (NOXA2). | |||||
|
TRDN_HUMAN
|
||||||
| NC score | 0.050923 (rank : 22) | θ value | θ > 10 (rank : 72) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 542 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q13061 | Gene names | TRDN | |||
|
Domain Architecture |

|
|||||
| Description | Triadin. | |||||
|
SYV_MOUSE
|
||||||
| NC score | 0.049778 (rank : 23) | θ value | 3.0926 (rank : 38) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Z1Q9, Q9QUN2 | Gene names | Vars, Bat6, G7a, Vars2 | |||
|
Domain Architecture |

|
|||||
| Description | Valyl-tRNA synthetase (EC 6.1.1.9) (Valine--tRNA ligase) (ValRS) (Protein G7a). | |||||
|
EF1G_HUMAN
|
||||||
| NC score | 0.048473 (rank : 24) | θ value | 4.03905 (rank : 42) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P26641, Q6PJ62, Q6PK31, Q96CU2, Q9P196 | Gene names | EEF1G, EF1G | |||
|
Domain Architecture |

|
|||||
| Description | Elongation factor 1-gamma (EF-1-gamma) (eEF-1B gamma). | |||||
|
SYCC_MOUSE
|
||||||
| NC score | 0.046042 (rank : 25) | θ value | 1.81305 (rank : 30) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9ER72, O88303, Q8BP81, Q8CAY7, Q8CCE3, Q8K0S4, Q9ER68 | Gene names | Cars | |||
|
Domain Architecture |

|
|||||
| Description | Cysteinyl-tRNA synthetase, cytoplasmic (EC 6.1.1.16) (Cysteine--tRNA ligase) (CysRS). | |||||
|
CD2AP_MOUSE
|
||||||
| NC score | 0.044815 (rank : 26) | θ value | 1.38821 (rank : 27) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 336 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9JLQ0, O88903, Q8K4Z1, Q8VCI9 | Gene names | Cd2ap, Mets1 | |||
|
Domain Architecture |

|
|||||
| Description | CD2-associated protein (Mesenchyme-to-epithelium transition protein with SH3 domains 1) (METS-1). | |||||
|
RRBP1_HUMAN
|
||||||
| NC score | 0.040605 (rank : 27) | θ value | 0.0961366 (rank : 14) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1306 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9P2E9, O75300, O75301, Q5W165, Q96SB2, Q9BWP1, Q9H476 | Gene names | RRBP1, KIAA1398 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (180 kDa ribosome receptor homolog) (ES/130-related protein). | |||||
|
MCES_MOUSE
|
||||||
| NC score | 0.040151 (rank : 28) | θ value | 2.36792 (rank : 32) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9D0L8, Q3V3U9, Q6ZQC6, Q9D5F1 | Gene names | Rnmt, Kiaa0398 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | mRNA cap guanine-N7 methyltransferase (EC 2.1.1.56) (mRNA (guanine- N(7)-)-methyltransferase) (RG7MT1) (mRNA cap methyltransferase). | |||||
|
SCG2_MOUSE
|
||||||
| NC score | 0.036679 (rank : 29) | θ value | 5.27518 (rank : 53) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q03517, Q80Y79, Q9CW80 | Gene names | Scg2, Chgc, Scg-2 | |||
|
Domain Architecture |

|
|||||
| Description | Secretogranin-2 precursor (Secretogranin II) (SgII) (Chromogranin C) [Contains: Secretoneurin (SN)]. | |||||
|
ESCO1_HUMAN
|
||||||
| NC score | 0.035430 (rank : 30) | θ value | 1.38821 (rank : 28) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 361 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q5FWF5, Q69YG4, Q69YS3, Q6IMD7, Q8N3Z5, Q8NBG2, Q96PX7 | Gene names | ESCO1, EFO1, KIAA1911 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyltransferase ESCO1 (EC 2.3.1.-) (Establishment of cohesion 1 homolog 1) (ECO1 homolog 1) (ESO1 homolog 1) (Establishment factor- like protein 1) (EFO1p) (hEFO1) (CTF7 homolog 1). | |||||
|
CD2AP_HUMAN
|
||||||
| NC score | 0.034412 (rank : 31) | θ value | 5.27518 (rank : 50) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y5K6, Q9UG97 | Gene names | CD2AP | |||
|
Domain Architecture |

|
|||||
| Description | CD2-associated protein (Cas ligand with multiple SH3 domains) (Adapter protein CMS). | |||||
|
RRBP1_MOUSE
|
||||||
| NC score | 0.033558 (rank : 32) | θ value | 0.163984 (rank : 18) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1650 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q99PL5, Q99PK5, Q99PK6, Q99PK7, Q99PK8, Q99PK9, Q99PL0, Q99PL1, Q99PL2, Q99PL3, Q99PL4, Q9CS20 | Gene names | Rrbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (mRRp). | |||||
|
PTPRZ_HUMAN
|
||||||
| NC score | 0.032002 (rank : 33) | θ value | 0.0252991 (rank : 10) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P23471 | Gene names | PTPRZ1, PTPRZ, PTPZ | |||
|
Domain Architecture |

|
|||||
| Description | Receptor-type tyrosine-protein phosphatase zeta precursor (EC 3.1.3.48) (R-PTP-zeta). | |||||
|
NRBF2_HUMAN
|
||||||
| NC score | 0.031671 (rank : 34) | θ value | 2.36792 (rank : 34) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96F24, Q86UR2, Q96NP6, Q9H0S9, Q9H2I2 | Gene names | NRBF2, COPR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor-binding factor 2 (NRBF-2) (Comodulator of PPAR and RXR). | |||||
|
MO4L2_HUMAN
|
||||||
| NC score | 0.028307 (rank : 35) | θ value | 5.27518 (rank : 51) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q15014, Q567V0, Q8J026 | Gene names | MORF4L2, KIAA0026, MRGX | |||
|
Domain Architecture |

|
|||||
| Description | Mortality factor 4-like protein 2 (MORF-related gene X protein) (Transcription factor-like protein MRGX) (MSL3-2 protein). | |||||
|
LZTS1_MOUSE
|
||||||
| NC score | 0.027772 (rank : 36) | θ value | 2.36792 (rank : 31) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 540 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P60853 | Gene names | Lzts1, Fez1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine zipper putative tumor suppressor 1 (F37/Esophageal cancer- related gene-coding leucine-zipper motif) (Fez1). | |||||
|
NIPBL_HUMAN
|
||||||
| NC score | 0.025296 (rank : 37) | θ value | 6.88961 (rank : 58) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
PRG4_HUMAN
|
||||||
| NC score | 0.024422 (rank : 38) | θ value | 6.88961 (rank : 59) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q92954, Q6DNC4, Q6DNC5, Q6ZMZ5, Q9BX49 | Gene names | PRG4, MSF, SZP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
SRBS1_MOUSE
|
||||||
| NC score | 0.023995 (rank : 39) | θ value | 1.38821 (rank : 29) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q62417, Q80TF8, Q8BZI3, Q8K3Y2, Q921F8, Q9Z0Z8, Q9Z0Z9 | Gene names | Sorbs1, Kiaa1296, Sh3d5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sorbin and SH3 domain-containing protein 1 (Ponsin) (c-Cbl-associated protein) (CAP) (SH3 domain protein 5) (SH3P12). | |||||
|
CKAP2_MOUSE
|
||||||
| NC score | 0.021200 (rank : 40) | θ value | 6.88961 (rank : 55) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q3V1H1, Q66LN2, Q8BSF0 | Gene names | Ckap2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoskeleton-associated protein 2. | |||||
|
CN145_HUMAN
|
||||||
| NC score | 0.018772 (rank : 41) | θ value | 0.813845 (rank : 25) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1196 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q6ZU80, Q96ML4 | Gene names | C14orf145 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf145. | |||||
|
RRP5_HUMAN
|
||||||
| NC score | 0.018681 (rank : 42) | θ value | 6.88961 (rank : 60) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 291 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q14690, Q86VQ8 | Gene names | PDCD11, KIAA0185 | |||
|
Domain Architecture |

|
|||||
| Description | RRP5 protein homolog (Programmed cell death protein 11). | |||||
|
CKAP2_HUMAN
|
||||||
| NC score | 0.018660 (rank : 43) | θ value | 8.99809 (rank : 65) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8WWK9, Q3KRA5, Q5VXB4, Q8IWV5, Q8IWV6, Q96FH9, Q9H012, Q9H0D0, Q9H988, Q9HC49, Q9NVG4 | Gene names | CKAP2, LB1, TMAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoskeleton-associated protein 2 (Tumor-associated microtubule- associated protein) (CTCL tumor antigen se20-10). | |||||
|
LAMA2_MOUSE
|
||||||
| NC score | 0.018554 (rank : 44) | θ value | 0.279714 (rank : 22) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1198 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q60675, Q05003, Q64061 | Gene names | Lama2 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-2 chain precursor (Laminin M chain) (Merosin heavy chain). | |||||
|
IWS1_HUMAN
|
||||||
| NC score | 0.017754 (rank : 45) | θ value | 3.0926 (rank : 36) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
MYO6_HUMAN
|
||||||
| NC score | 0.017412 (rank : 46) | θ value | 2.36792 (rank : 33) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 727 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9UM54, Q9BZZ7, Q9UEG2 | Gene names | MYO6, KIAA0389 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-6 (Myosin VI). | |||||
|
MECR_MOUSE
|
||||||
| NC score | 0.017221 (rank : 47) | θ value | 4.03905 (rank : 43) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9DCS3, Q99L39 | Gene names | Mecr, Nrbf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trans-2-enoyl-CoA reductase, mitochondrial precursor (EC 1.3.1.38). | |||||
|
MAP1A_HUMAN
|
||||||
| NC score | 0.017046 (rank : 48) | θ value | 8.99809 (rank : 68) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1626 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P78559, O95643, Q12973, Q15882, Q9UJT4 | Gene names | MAP1A | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 1A (MAP 1A) (Proliferation-related protein p80) [Contains: MAP1 light chain LC2]. | |||||
|
XYLT1_HUMAN
|
||||||
| NC score | 0.016814 (rank : 49) | θ value | 3.0926 (rank : 39) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q86Y38, Q9H1B6 | Gene names | XYLT1, XT1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Xylosyltransferase 1 (EC 2.4.2.26) (Xylosyltransferase I) (XylT-I) (XT-I) (Peptide O-xylosyltransferase 1). | |||||
|
MATR3_HUMAN
|
||||||
| NC score | 0.016773 (rank : 50) | θ value | 6.88961 (rank : 57) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P43243, Q9UHW0, Q9UQ27 | Gene names | MATR3, KIAA0723 | |||
|
Domain Architecture |

|
|||||
| Description | Matrin-3. | |||||
|
DOCK4_HUMAN
|
||||||
| NC score | 0.016356 (rank : 51) | θ value | 4.03905 (rank : 40) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 237 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8N1I0, O14584, O94824, Q8NB45 | Gene names | DOCK4, KIAA0716 | |||
|
Domain Architecture |

|
|||||
| Description | Dedicator of cytokinesis protein 4. | |||||
|
DOCK4_MOUSE
|
||||||
| NC score | 0.016289 (rank : 52) | θ value | 4.03905 (rank : 41) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P59764 | Gene names | Dock4, Kiaa0716 | |||
|
Domain Architecture |

|
|||||
| Description | Dedicator of cytokinesis protein 4. | |||||
|
ATRX_MOUSE
|
||||||
| NC score | 0.015365 (rank : 53) | θ value | 5.27518 (rank : 49) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 682 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q61687 | Gene names | Atrx, Hp1bp2, Xnp | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ATRX (EC 3.6.1.-) (ATP-dependent helicase ATRX) (X-linked nuclear protein) (Heterochromatin protein 2) (HP1 alpha-interacting protein) (HP1-BP38 protein). | |||||
|
CF182_HUMAN
|
||||||
| NC score | 0.014581 (rank : 54) | θ value | 8.99809 (rank : 64) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 474 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8IYX8 | Gene names | C6orf182 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C6orf182. | |||||
|
AFF2_HUMAN
|
||||||
| NC score | 0.014578 (rank : 55) | θ value | 5.27518 (rank : 48) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P51816, O43786, O60215, P78407, Q13521, Q14323, Q9UNA5 | Gene names | AFF2, FMR2, OX19 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 2 (Fragile X mental retardation 2 protein) (Protein FMR-2) (FMR2P) (Protein Ox19) (Fragile X E mental retardation syndrome protein). | |||||
|
KMHN1_MOUSE
|
||||||
| NC score | 0.014208 (rank : 56) | θ value | 3.0926 (rank : 37) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 949 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q3V125, Q3TTQ8 | Gene names | Gm172 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cancer/testis antigen KM-HN-1 homolog. | |||||
|
MYH13_HUMAN
|
||||||
| NC score | 0.013815 (rank : 57) | θ value | 4.03905 (rank : 44) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1504 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9UKX3, O95252 | Gene names | MYH13 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-13 (Myosin heavy chain, skeletal muscle, extraocular) (MyHC- eo). | |||||
|
IWS1_MOUSE
|
||||||
| NC score | 0.013691 (rank : 58) | θ value | 6.88961 (rank : 56) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
TBC14_HUMAN
|
||||||
| NC score | 0.013404 (rank : 59) | θ value | 5.27518 (rank : 54) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9P2M4, Q8IW15 | Gene names | TBC1D14, KIAA1322 | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 14. | |||||
|
REST_MOUSE
|
||||||
| NC score | 0.012659 (rank : 60) | θ value | 4.03905 (rank : 46) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1506 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q922J3, Q571L7 | Gene names | Rsn, Kiaa4046 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Restin. | |||||
|
RBM28_MOUSE
|
||||||
| NC score | 0.011946 (rank : 61) | θ value | 4.03905 (rank : 45) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8CGC6, Q8BG25, Q8VEJ8, Q9CS22, Q9CSE6 | Gene names | Rbm28 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein 28 (RNA-binding motif protein 28). | |||||
|
MYH7_HUMAN
|
||||||
| NC score | 0.011777 (rank : 62) | θ value | 5.27518 (rank : 52) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P12883, Q14836, Q14837, Q14904, Q16579, Q92679, Q9H1D5, Q9UMM8 | Gene names | MYH7, MYHCB | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle beta isoform (MyHC-beta). | |||||
|
FEZ2_HUMAN
|
||||||
| NC score | 0.011663 (rank : 63) | θ value | 8.99809 (rank : 66) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UHY8, Q76LN0, Q99690 | Gene names | FEZ2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fasciculation and elongation protein zeta 2 (Zygin-2) (Zygin II). | |||||
|
SYNE1_HUMAN
|
||||||
| NC score | 0.011305 (rank : 64) | θ value | 6.88961 (rank : 61) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1484 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8NF91, O94890, Q5JV19, Q5JV22, Q8N9P7, Q8TCP1, Q8WWW6, Q8WWW7, Q8WXF6, Q96N17, Q9C0A7, Q9H525, Q9H526, Q9NS36, Q9NU50, Q9UJ06, Q9UJ07, Q9ULF8 | Gene names | SYNE1, KIAA0796, KIAA1262, KIAA1756, MYNE1 | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-1 (Nuclear envelope spectrin repeat protein 1) (Synaptic nuclear envelope protein 1) (Syne-1) (Myocyte nuclear envelope protein 1) (Myne-1) (Enaptin). | |||||
|
GOGA4_HUMAN
|
||||||
| NC score | 0.010691 (rank : 65) | θ value | 8.99809 (rank : 67) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 2034 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q13439, Q13270, Q13654, Q14436 | Gene names | GOLGA4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (Trans-Golgi p230) (256 kDa golgin) (Golgin-245) (Protein 72.1). | |||||
|
PTN6_HUMAN
|
||||||
| NC score | 0.010068 (rank : 66) | θ value | 8.99809 (rank : 69) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 234 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P29350, Q969V8 | Gene names | PTPN6, HCP, PTP1C | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 6 (EC 3.1.3.48) (Protein-tyrosine phosphatase 1C) (PTP-1C) (Hematopoietic cell protein-tyrosine phosphatase) (SH-PTP1) (Protein-tyrosine phosphatase SHP-1). | |||||
|
ZN217_HUMAN
|
||||||
| NC score | 0.009921 (rank : 67) | θ value | 0.125558 (rank : 17) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 905 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O75362 | Gene names | ZNF217, ZABC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 217. | |||||
|
SF3B4_HUMAN
|
||||||
| NC score | 0.009769 (rank : 68) | θ value | 4.03905 (rank : 47) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q15427 | Gene names | SF3B4, SAP49 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 3B subunit 4 (Spliceosome-associated protein 49) (SAP 49) (SF3b50) (Pre-mRNA-splicing factor SF3b 49 kDa subunit). | |||||
|
CDC7_HUMAN
|
||||||
| NC score | 0.004339 (rank : 69) | θ value | 8.99809 (rank : 63) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 652 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O00311, O00558 | Gene names | CDC7, CDC7L1 | |||
|
Domain Architecture |

|
|||||
| Description | Cell division cycle 7-related protein kinase (EC 2.7.11.1) (CDC7- related kinase) (HsCdc7) (huCdc7). | |||||
|
CD2L5_MOUSE
|
||||||
| NC score | 0.002177 (rank : 70) | θ value | 3.0926 (rank : 35) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 1011 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q69ZA1, Q80V11, Q8BZG1, Q8K0A4 | Gene names | Cdc2l5, Kiaa1791 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle 2-like protein kinase 5 (EC 2.7.11.22) (CDC2- related protein kinase 5). | |||||
|
TITIN_HUMAN
|
||||||
| NC score | 0.001035 (rank : 71) | θ value | 6.88961 (rank : 62) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 2923 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8WZ42, Q10465, Q10466, Q15598, Q2XUS3, Q32Q60, Q4U1Z6, Q6NSG0, Q6PDB1, Q6PJP0, Q7KYM2, Q7KYN4, Q7KYN5, Q7LDM3, Q7Z2X3, Q8TCG8, Q8WZ51, Q8WZ52, Q8WZ53, Q8WZB3, Q92761, Q92762, Q9UD97, Q9UP84, Q9Y6L9 | Gene names | TTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Titin (EC 2.7.11.1) (Connectin) (Rhabdomyosarcoma antigen MU-RMS- 40.14). | |||||
|
ZN583_HUMAN
|
||||||
| NC score | -0.000366 (rank : 72) | θ value | 8.99809 (rank : 70) | |||
| Query Neighborhood Hits | 70 | Target Neighborhood Hits | 774 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96ND8, O14850, Q2NKK3 | Gene names | ZNF583 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 583 (Zinc finger protein L3-5). | |||||