Please be patient as the page loads
|
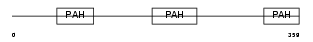
SIN3B_HUMAN
|
||||||
| SwissProt Accessions | O75182, Q68GC2, Q6P4B8, Q8TB34, Q9BSC8 | Gene names | SIN3B, KIAA0700 | |||
|
Domain Architecture |
|
|||||
| Description | Paired amphipathic helix protein Sin3b (Transcriptional corepressor Sin3b) (Histone deacetylase complex subunit Sin3b). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
SIN3A_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.946537 (rank : 4) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q96ST3, Q8N8N4, Q8NC83, Q8WV18, Q96L98, Q9UFQ1 | Gene names | SIN3A | |||
|
Domain Architecture |

|
|||||
| Description | Paired amphipathic helix protein Sin3a (Transcriptional corepressor Sin3a) (Histone deacetylase complex subunit Sin3a). | |||||
|
SIN3A_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.950005 (rank : 3) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q60520, Q60820, Q62139, Q62140, Q7TPU8, Q7TSZ2 | Gene names | Sin3a | |||
|
Domain Architecture |

|
|||||
| Description | Paired amphipathic helix protein Sin3a (Transcriptional corepressor Sin3a) (Histone deacetylase complex subunit Sin3a). | |||||
|
SIN3B_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | O75182, Q68GC2, Q6P4B8, Q8TB34, Q9BSC8 | Gene names | SIN3B, KIAA0700 | |||
|
Domain Architecture |

|
|||||
| Description | Paired amphipathic helix protein Sin3b (Transcriptional corepressor Sin3b) (Histone deacetylase complex subunit Sin3b). | |||||
|
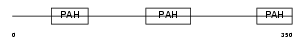
SIN3B_MOUSE
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.988426 (rank : 2) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q62141, O54976, Q6A013, Q8VCB8, Q8VDZ5 | Gene names | Sin3b, Kiaa0700 | |||
|
Domain Architecture |
|
|||||
| Description | Paired amphipathic helix protein Sin3b (Transcriptional corepressor Sin3b) (Histone deacetylase complex subunit Sin3b). | |||||
|
TXLNB_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 5) | NC score | 0.034783 (rank : 16) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 912 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8N3L3, Q76L25, Q86T52, Q8N3S2 | Gene names | TXLNB, C6orf198, MDP77 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-taxilin (Muscle-derived protein 77) (hMDP77). | |||||
|
ACE_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 6) | NC score | 0.049988 (rank : 7) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P09470, Q6GTS2 | Gene names | Ace, Dcp1 | |||
|
Domain Architecture |

|
|||||
| Description | Angiotensin-converting enzyme, somatic isoform precursor (EC 3.4.15.1) (Dipeptidyl carboxypeptidase I) (Kininase II) [Contains: Angiotensin- converting enzyme, somatic isoform, soluble form]. | |||||
|
MAML2_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 7) | NC score | 0.045399 (rank : 9) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8IZL2, Q6AI23, Q6Y3A3, Q8IUL3, Q96JK6 | Gene names | MAML2, KIAA1819 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mastermind-like protein 2 (Mam-2). | |||||
|
NEK11_HUMAN
|
||||||
| θ value | 0.163984 (rank : 8) | NC score | 0.010353 (rank : 63) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1043 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8NG66, Q5JPC0, Q8NG65, Q8TBY1, Q9H5F4 | Gene names | NEK11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase Nek11 (EC 2.7.11.1) (NimA-related protein kinase 11) (Never in mitosis A-related kinase 11). | |||||
|
OGFR_HUMAN
|
||||||
| θ value | 0.163984 (rank : 9) | NC score | 0.023982 (rank : 36) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
CENPE_HUMAN
|
||||||
| θ value | 0.21417 (rank : 10) | NC score | 0.032558 (rank : 19) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 2060 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q02224 | Gene names | CENPE | |||
|
Domain Architecture |

|
|||||
| Description | Centromeric protein E (CENP-E). | |||||
|
SMC1A_MOUSE
|
||||||
| θ value | 0.21417 (rank : 11) | NC score | 0.027252 (rank : 26) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1384 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9CU62, Q3V480, Q9CUX9, Q9D959, Q9WTU1 | Gene names | Smc1l1, Sb1.8, Smc1, Smcb | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 1-like 1 protein (SMC1alpha protein) (Chromosome segregation protein SmcB) (Sb1.8). | |||||
|
NDE1_HUMAN
|
||||||
| θ value | 0.279714 (rank : 12) | NC score | 0.034809 (rank : 15) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 823 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9NXR1, Q49AQ2 | Gene names | NDE1, NUDE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear distribution protein nudE homolog 1 (NudE). | |||||
|
RFTN1_MOUSE
|
||||||
| θ value | 0.279714 (rank : 13) | NC score | 0.036662 (rank : 11) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6A0D4, Q3U654, Q3U8P6, Q80US1 | Gene names | Rftn1, Kiaa0084 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Raftlin (Raft-linking protein). | |||||
|
CCD19_HUMAN
|
||||||
| θ value | 0.813845 (rank : 14) | NC score | 0.046142 (rank : 8) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 639 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UL16 | Gene names | CCDC19, NESG1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 19 (Nasopharyngeal epithelium- specific protein 1). | |||||
|
CGRE1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 15) | NC score | 0.030127 (rank : 21) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q99674 | Gene names | CGREF1, CGR11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell growth regulator with EF hand domain 1 (Cell growth regulatory gene 11 protein). | |||||
|
ITSN2_MOUSE
|
||||||
| θ value | 0.813845 (rank : 16) | NC score | 0.019464 (rank : 44) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1371 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9Z0R6, Q9Z0R5 | Gene names | Itsn2, Ese2, Sh3d1B | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-2 (SH3 domain-containing protein 1B) (EH and SH3 domains protein 2) (EH domain and SH3 domain regulator of endocytosis 2). | |||||
|
NDE1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 17) | NC score | 0.035093 (rank : 14) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 602 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9CZA6, Q3UBS6, Q3UIC1, Q9ERR0 | Gene names | Nde1, Nude | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear distribution protein nudE homolog 1 (NudE) (mNudE). | |||||
|
PCNT_MOUSE
|
||||||
| θ value | 0.813845 (rank : 18) | NC score | 0.022355 (rank : 39) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1207 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P48725 | Gene names | Pcnt, Pcnt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pericentrin. | |||||
|
CN145_HUMAN
|
||||||
| θ value | 1.06291 (rank : 19) | NC score | 0.025822 (rank : 27) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1196 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q6ZU80, Q96ML4 | Gene names | C14orf145 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf145. | |||||
|
GON4L_MOUSE
|
||||||
| θ value | 1.06291 (rank : 20) | NC score | 0.054031 (rank : 6) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9DB00, Q80TB4, Q91YI9 | Gene names | Gon4l, Gon4, KIAA1606 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GON-4-like protein (GON-4 homolog). | |||||
|
PEG3_MOUSE
|
||||||
| θ value | 1.06291 (rank : 21) | NC score | 0.008521 (rank : 68) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1101 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q3URU2, O54978, Q3TQ69, Q5EBP7, Q61138, Q6GQS0, Q80U47, Q8R5N0, Q9QX53 | Gene names | Peg3, Kiaa0287, Pw1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Paternally expressed gene 3 protein (ASF-1). | |||||
|
DOK1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 22) | NC score | 0.020966 (rank : 41) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P97465, Q9R213 | Gene names | Dok1, Dok | |||
|
Domain Architecture |

|
|||||
| Description | Docking protein 1 (Downstream of tyrosine kinase 1) (p62(dok)). | |||||
|
SAPS3_MOUSE
|
||||||
| θ value | 1.38821 (rank : 23) | NC score | 0.017480 (rank : 48) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q922D4, Q6ZPM9, Q8BTW6, Q9D2X9 | Gene names | Saps3, D19Ertd703e, Kiaa1558 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SAPS domain family member 3. | |||||
|
SMC1A_HUMAN
|
||||||
| θ value | 1.38821 (rank : 24) | NC score | 0.024621 (rank : 34) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1346 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q14683, O14995, Q16351 | Gene names | SMC1L1, DXS423E, KIAA0178, SB1.8, SMC1 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 1-like 1 protein (SMC1alpha protein) (Sb1.8). | |||||
|
IF5_HUMAN
|
||||||
| θ value | 1.81305 (rank : 25) | NC score | 0.034461 (rank : 18) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P55010, Q9H5N2, Q9UG48 | Gene names | EIF5 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 5 (eIF-5). | |||||
|
MUG2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 26) | NC score | 0.019617 (rank : 43) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P28666 | Gene names | Mug2, Mug-2 | |||
|
Domain Architecture |

|
|||||
| Description | Murinoglobulin-2 precursor (MuG2). | |||||
|
RAI14_HUMAN
|
||||||
| θ value | 1.81305 (rank : 27) | NC score | 0.012638 (rank : 60) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1367 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9P0K7, Q6V1W9, Q7Z5I4, Q7Z733, Q9P2L2, Q9Y3T5 | Gene names | RAI14, KIAA1334, NORPEG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankycorbin (Ankyrin repeat and coiled-coil structure-containing protein) (Retinoic acid-induced protein 14) (Novel retinal pigment epithelial cell protein). | |||||
|
COX10_MOUSE
|
||||||
| θ value | 2.36792 (rank : 28) | NC score | 0.023571 (rank : 38) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CFY5 | Gene names | Cox10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protoheme IX farnesyltransferase, mitochondrial precursor (EC 2.5.1.-) (Heme O synthase). | |||||
|
HDAC5_HUMAN
|
||||||
| θ value | 2.36792 (rank : 29) | NC score | 0.013430 (rank : 58) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 353 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9UQL6, O60340, O60528, Q96DY4 | Gene names | HDAC5, KIAA0600 | |||
|
Domain Architecture |

|
|||||
| Description | Histone deacetylase 5 (HD5) (Antigen NY-CO-9). | |||||
|
HOOK3_HUMAN
|
||||||
| θ value | 2.36792 (rank : 30) | NC score | 0.024963 (rank : 32) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1070 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q86VS8, Q8NBH0, Q9BY13 | Gene names | HOOK3 | |||
|
Domain Architecture |

|
|||||
| Description | Hook homolog 3 (hHK3). | |||||
|
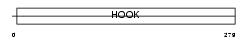
HOOK3_MOUSE
|
||||||
| θ value | 2.36792 (rank : 31) | NC score | 0.025111 (rank : 31) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1007 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8BUK6, Q8BV48, Q8BW57, Q8BY41 | Gene names | Hook3 | |||
|
Domain Architecture |
|
|||||
| Description | Hook homolog 3. | |||||
|
INP4A_MOUSE
|
||||||
| θ value | 2.36792 (rank : 32) | NC score | 0.016340 (rank : 50) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9EPW0, Q8CBJ8 | Gene names | Inpp4a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Type I inositol-3,4-bisphosphate 4-phosphatase (EC 3.1.3.66) (Inositol polyphosphate 4-phosphatase type I). | |||||
|
MTSS1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 33) | NC score | 0.024700 (rank : 33) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O43312, Q8TCA2, Q96RX2 | Gene names | MTSS1, KIAA0429, MIM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metastasis suppressor protein 1 (Missing in metastasis protein) (Metastasis suppressor YGL-1). | |||||
|
PTN21_HUMAN
|
||||||
| θ value | 2.36792 (rank : 34) | NC score | 0.006980 (rank : 75) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q16825 | Gene names | PTPN21, PTPD1 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 21 (EC 3.1.3.48) (Protein-tyrosine phosphatase D1). | |||||
|
SLK_HUMAN
|
||||||
| θ value | 2.36792 (rank : 35) | NC score | 0.007784 (rank : 71) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1775 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9H2G2, O00211, Q6P1Z4, Q86WU7, Q86WW1, Q92603, Q9NQL0, Q9NQL1 | Gene names | SLK, KIAA0204, STK2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | STE20-like serine/threonine-protein kinase (EC 2.7.11.1) (STE20-like kinase) (STE20-related serine/threonine-protein kinase) (STE20-related kinase) (hSLK) (Serine/threonine-protein kinase 2) (CTCL tumor antigen se20-9). | |||||
|
DAB2P_HUMAN
|
||||||
| θ value | 3.0926 (rank : 36) | NC score | 0.025665 (rank : 29) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q5VWQ8, Q8TDL2, Q96SE1, Q9C0C0 | Gene names | DAB2IP, AF9Q34, AIP1, KIAA1743 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Disabled homolog 2-interacting protein (DAB2-interacting protein) (DAB2 interaction protein) (ASK-interacting protein 1). | |||||
|
DAB2P_MOUSE
|
||||||
| θ value | 3.0926 (rank : 37) | NC score | 0.025407 (rank : 30) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 284 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q3UHC7, Q3TPD5, Q3UH44, Q6JTV1, Q80T97 | Gene names | Dab2ip, Kiaa1743 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Disabled homolog 2-interacting protein (DAB2-interacting protein). | |||||
|
SCG3_MOUSE
|
||||||
| θ value | 3.0926 (rank : 38) | NC score | 0.023947 (rank : 37) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P47867 | Gene names | Scg3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Secretogranin-3 precursor (Secretogranin III) (SgIII). | |||||
|
TRIP4_MOUSE
|
||||||
| θ value | 3.0926 (rank : 39) | NC score | 0.035973 (rank : 12) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9QXN3, Q8CAD5 | Gene names | Trip4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Activating signal cointegrator 1 (ASC-1) (Thyroid receptor-interacting protein 4) (Trip-4). | |||||
|
UBXD2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 40) | NC score | 0.029415 (rank : 23) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q92575, Q8IYM5 | Gene names | UBXD2, KIAA0242, UBXDC1 | |||
|
Domain Architecture |

|
|||||
| Description | UBX domain-containing protein 2. | |||||
|
APBA2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 41) | NC score | 0.010418 (rank : 62) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P98084, Q6PAJ2 | Gene names | Apba2, X11l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 precursor protein-binding family A member 2 (Neuron- specific X11L protein) (Neuronal Munc18-1-interacting protein 2) (Mint-2) (Adapter protein X11beta). | |||||
|
G137B_HUMAN
|
||||||
| θ value | 4.03905 (rank : 42) | NC score | 0.025761 (rank : 28) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O60478 | Gene names | GPR137B, TM7SF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Integral membrane protein GPR137B (Transmembrane 7 superfamily member 1). | |||||
|
OFD1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 43) | NC score | 0.035514 (rank : 13) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 753 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O75665, O75666 | Gene names | OFD1, CXorf5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Oral-facial-digital syndrome 1 protein (Protein 71-7A). | |||||
|
RRP5_HUMAN
|
||||||
| θ value | 4.03905 (rank : 44) | NC score | 0.030410 (rank : 20) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 291 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q14690, Q86VQ8 | Gene names | PDCD11, KIAA0185 | |||
|
Domain Architecture |

|
|||||
| Description | RRP5 protein homolog (Programmed cell death protein 11). | |||||
|
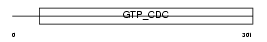
SEPT6_HUMAN
|
||||||
| θ value | 4.03905 (rank : 45) | NC score | 0.019425 (rank : 45) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 220 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q14141, Q5JTK0, Q969W5, Q96A13, Q96GR1, Q96P86, Q96P87 | Gene names | SEPT6, KIAA0128, SEP2 | |||
|
Domain Architecture |
|
|||||
| Description | Septin-6. | |||||
|
UBXD2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 46) | NC score | 0.028168 (rank : 25) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8VCH8, Q8BW17 | Gene names | Ubxd2, Ubxdc1 | |||
|
Domain Architecture |

|
|||||
| Description | UBX domain-containing protein 2. | |||||
|
ACE_HUMAN
|
||||||
| θ value | 5.27518 (rank : 47) | NC score | 0.029879 (rank : 22) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P12821, Q53YX9, Q59GY8 | Gene names | ACE, DCP, DCP1 | |||
|
Domain Architecture |

|
|||||
| Description | Angiotensin-converting enzyme, somatic isoform precursor (EC 3.4.15.1) (Dipeptidyl carboxypeptidase I) (Kininase II) (CD143 antigen) [Contains: Angiotensin-converting enzyme, somatic isoform, soluble form]. | |||||
|
BRAP_MOUSE
|
||||||
| θ value | 5.27518 (rank : 48) | NC score | 0.038118 (rank : 10) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q99MP8, Q8CC00, Q8CHX1, Q99MP7, Q9CXX8 | Gene names | Brap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BRCA1-associated protein (EC 6.3.2.-) (BRAP2) (Impedes mitogenic signal propagation) (IMP). | |||||
|
CTR9_MOUSE
|
||||||
| θ value | 5.27518 (rank : 49) | NC score | 0.018251 (rank : 46) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q62018, Q3UFF5, Q3UY40, Q66JX4, Q7TPS6, Q8BND9, Q8BRD1, Q8C9W7, Q8C9Y3, Q8CHI1 | Gene names | Ctr9, Kiaa0155, Sh2bp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein CTR9 homolog (SH2 domain-binding protein 1) (Tetratricopeptide repeat-containing, SH2-binding phosphoprotein of 150 kDa) (TPR-containing, SH2-binding phosphoprotein of 150 kDa) (p150TSP). | |||||
|
DZIP3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 50) | NC score | 0.020838 (rank : 42) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 414 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q7TPV2, Q80TU4, Q8BYK7 | Gene names | Dzip3, Kiaa0675 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin ligase protein DZIP3 (EC 6.3.2.-) (DAZ-interacting protein 3 homolog). | |||||
|
IF5_MOUSE
|
||||||
| θ value | 5.27518 (rank : 51) | NC score | 0.028761 (rank : 24) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P59325 | Gene names | Eif5 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 5 (eIF-5). | |||||
|
UTRO_HUMAN
|
||||||
| θ value | 5.27518 (rank : 52) | NC score | 0.014752 (rank : 55) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P46939 | Gene names | UTRN, DMDL | |||
|
Domain Architecture |

|
|||||
| Description | Utrophin (Dystrophin-related protein 1) (DRP1) (DRP). | |||||
|
WAPL_MOUSE
|
||||||
| θ value | 5.27518 (rank : 53) | NC score | 0.013656 (rank : 57) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q65Z40, Q6ZQE9 | Gene names | Wapal, Kiaa0261, Wapl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wings apart-like protein homolog (Dioxin-inducible factor 2) (DIF-2). | |||||
|
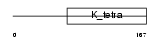
ZBTB3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 54) | NC score | 0.003180 (rank : 80) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 910 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9H5J0 | Gene names | ZBTB3 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger and BTB domain-containing protein 3. | |||||
|
ASCC2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 55) | NC score | 0.012697 (rank : 59) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9H1I8, Q4TT54, Q8TAZ0, Q9H711, Q9H9D6 | Gene names | ASCC2, ASC1P100 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Activating signal cointegrator 1 complex subunit 2 (ASC-1 complex subunit p100) (Trip4 complex subunit p100). | |||||
|
CENPK_HUMAN
|
||||||
| θ value | 6.88961 (rank : 56) | NC score | 0.023983 (rank : 35) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 273 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9BS16, Q9H4L0 | Gene names | CENPK, ICEN37 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein K (CENP-K) (Interphase centromere complex protein 37) (Protein AF-5alpha) (p33). | |||||
|
CI039_HUMAN
|
||||||
| θ value | 6.88961 (rank : 57) | NC score | 0.017684 (rank : 47) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1171 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9NXG0, Q5VYJ0, Q9HAJ5 | Gene names | C9orf39 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C9orf39. | |||||
|
CTR9_HUMAN
|
||||||
| θ value | 6.88961 (rank : 58) | NC score | 0.017453 (rank : 49) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 383 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6PD62, Q15015 | Gene names | CTR9, KIAA0155, SH2BP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein CTR9 homolog (SH2 domain-binding protein 1). | |||||
|
CTRO_HUMAN
|
||||||
| θ value | 6.88961 (rank : 59) | NC score | 0.008567 (rank : 65) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 2092 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O14578, Q6XUH8, Q86UQ9, Q9UPZ7 | Gene names | CIT, KIAA0949, STK21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Citron Rho-interacting kinase (EC 2.7.11.1) (CRIK) (Rho-interacting, serine/threonine-protein kinase 21). | |||||
|
CTRO_MOUSE
|
||||||
| θ value | 6.88961 (rank : 60) | NC score | 0.009220 (rank : 64) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 2344 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P49025, O88528, O88937, O88938, Q3UM99, Q8CIJ1 | Gene names | Cit | |||
|
Domain Architecture |

|
|||||
| Description | Citron Rho-interacting kinase (EC 2.7.11.1) (CRIK) (Rho-interacting, serine/threonine-protein kinase 21). | |||||
|
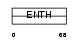
EPN2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 61) | NC score | 0.008561 (rank : 66) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8CHU3, O70447, Q8BZ85 | Gene names | Epn2 | |||
|
Domain Architecture |
|
|||||
| Description | Epsin-2 (EPS-15-interacting protein 2) (Intersectin-EH-binding protein 2) (Ibp2). | |||||
|
GON4L_HUMAN
|
||||||
| θ value | 6.88961 (rank : 62) | NC score | 0.034513 (rank : 17) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 555 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q3T8J9, Q3T8J8, Q5VYZ5, Q5W0D5, Q6AWA6, Q6P1Q6, Q7Z3L3, Q8IY79, Q9BQI1, Q9HCG6 | Gene names | GON4L, GON4, KIAA1606 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GON-4-like protein (GON-4 homolog). | |||||
|
MYO6_MOUSE
|
||||||
| θ value | 6.88961 (rank : 63) | NC score | 0.006985 (rank : 74) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 586 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q64331 | Gene names | Myo6, Sv | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-6 (Myosin VI). | |||||
|
PIAS1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 64) | NC score | 0.010975 (rank : 61) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O75925, Q99751, Q9UN02 | Gene names | PIAS1, DDXBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein inhibitor of activated STAT protein 1 (Gu-binding protein) (GBP) (RNA helicase II-binding protein) (DEAD/H box-binding protein 1). | |||||
|
SYCP1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 65) | NC score | 0.015601 (rank : 52) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1456 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q62209, O09205, P70192, Q62329 | Gene names | Sycp1, Scp1 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptonemal complex protein 1 (SCP-1). | |||||
|
TAU_HUMAN
|
||||||
| θ value | 6.88961 (rank : 66) | NC score | 0.008503 (rank : 69) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 448 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P10636, P18518, Q14799, Q15549, Q15550, Q15551, Q9UDJ3, Q9UMH0, Q9UQ96 | Gene names | MAPT, MTBT1, TAU | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein tau (Neurofibrillary tangle protein) (Paired helical filament-tau) (PHF-tau). | |||||
|
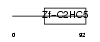
TRIP4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 67) | NC score | 0.022174 (rank : 40) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q15650, Q96ED7, Q9UKH0 | Gene names | TRIP4 | |||
|
Domain Architecture |
|
|||||
| Description | Activating signal cointegrator 1 (ASC-1) (Thyroid receptor-interacting protein 4) (TRIP-4). | |||||
|
UT14C_HUMAN
|
||||||
| θ value | 6.88961 (rank : 68) | NC score | 0.016148 (rank : 51) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q5TAP6, Q5FWG3, Q92555 | Gene names | UTP14C, KIAA0266 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U3 small nucleolar RNA-associated protein 14 homolog C. | |||||
|
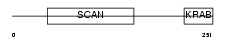
ZFP95_MOUSE
|
||||||
| θ value | 6.88961 (rank : 69) | NC score | 0.001066 (rank : 83) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 787 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Z1D8 | Gene names | Zfp95 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger protein 95 (Zfp-95). | |||||
|
ZN287_MOUSE
|
||||||
| θ value | 6.88961 (rank : 70) | NC score | 0.002632 (rank : 81) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 771 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9EQB9 | Gene names | Znf287, Skat2, Zfp287 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger protein 287 (Zfp-287) (Zinc finger protein SKAT-2). | |||||
|
AFF2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 71) | NC score | 0.007264 (rank : 72) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O55112 | Gene names | Aff2, Fmr2, Ox19 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 2 (Fragile X mental retardation protein 2 homolog) (Protein FMR-2) (FMR2P) (Protein Ox19). | |||||
|
CAN3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 72) | NC score | 0.003643 (rank : 79) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P20807, Q9BTU4, Q9Y5S6, Q9Y5S7 | Gene names | CAPN3, CANP3, CANPL3, NCL1 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-3 (EC 3.4.22.-) (Calpain L3) (Calpain p94) (Calcium-activated neutral proteinase 3) (CANP 3) (Muscle-specific calcium-activated neutral protease 3) (nCL-1). | |||||
|
CENPJ_HUMAN
|
||||||
| θ value | 8.99809 (rank : 73) | NC score | 0.014906 (rank : 54) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 721 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9HC77, Q5T6R5, Q96KS5, Q9C067 | Gene names | CENPJ, CPAP, LAP, LIP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein J (CENP-J) (Centrosomal P4.1-associated protein) (LAG-3-associated protein) (LYST-interacting protein 1). | |||||
|
CLMN_MOUSE
|
||||||
| θ value | 8.99809 (rank : 74) | NC score | 0.006458 (rank : 76) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8C5W0, Q91V71, Q91XT7, Q91XT8, Q91XU9 | Gene names | Clmn | |||
|
Domain Architecture |
|
|||||
| Description | Calmin. | |||||
|
CX022_HUMAN
|
||||||
| θ value | 8.99809 (rank : 75) | NC score | 0.013968 (rank : 56) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6ZTR5, Q5JRM8, Q8N6X8 | Gene names | CXorf22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein CXorf22. | |||||
|
E41L2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 76) | NC score | 0.006018 (rank : 77) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 287 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O43491 | Gene names | EPB41L2 | |||
|
Domain Architecture |

|
|||||
| Description | Band 4.1-like protein 2 (Generally expressed protein 4.1) (4.1G). | |||||
|
JPH1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 77) | NC score | 0.005996 (rank : 78) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9ET80, Q9EQZ3 | Gene names | Jph1, Jp1 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-1 (Junctophilin type 1) (JP-1). | |||||
|
OXRP_HUMAN
|
||||||
| θ value | 8.99809 (rank : 78) | NC score | 0.008524 (rank : 67) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 544 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Y4L1 | Gene names | HYOU1, ORP150 | |||
|
Domain Architecture |

|
|||||
| Description | 150 kDa oxygen-regulated protein precursor (Orp150) (Hypoxia up- regulated 1). | |||||
|
PUM2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 79) | NC score | 0.007104 (rank : 73) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8TB72, O00234, Q53TV7, Q8WY43, Q9HAN2 | Gene names | PUM2, KIAA0235, PUMH2 | |||
|
Domain Architecture |

|
|||||
| Description | Pumilio homolog 2 (Pumilio-2). | |||||
|
RGS3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 80) | NC score | 0.014910 (rank : 53) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
SHAN2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 81) | NC score | 0.008248 (rank : 70) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 346 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UPX8, Q3Y8G9, Q52LK2, Q9UKP1 | Gene names | SHANK2, KIAA1022 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 and multiple ankyrin repeat domains protein 2 (Shank2). | |||||
|
ZN287_HUMAN
|
||||||
| θ value | 8.99809 (rank : 82) | NC score | 0.002214 (rank : 82) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 777 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9HBT7 | Gene names | ZNF287 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger protein 287. | |||||
|
ZN645_HUMAN
|
||||||
| θ value | θ > 10 (rank : 83) | NC score | 0.064277 (rank : 5) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 265 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8N7E2, Q6DJY9 | Gene names | ZNF645 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 645. | |||||
|
SIN3B_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | O75182, Q68GC2, Q6P4B8, Q8TB34, Q9BSC8 | Gene names | SIN3B, KIAA0700 | |||
|
Domain Architecture |

|
|||||
| Description | Paired amphipathic helix protein Sin3b (Transcriptional corepressor Sin3b) (Histone deacetylase complex subunit Sin3b). | |||||
|
SIN3B_MOUSE
|
||||||
| NC score | 0.988426 (rank : 2) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q62141, O54976, Q6A013, Q8VCB8, Q8VDZ5 | Gene names | Sin3b, Kiaa0700 | |||
|
Domain Architecture |

|
|||||
| Description | Paired amphipathic helix protein Sin3b (Transcriptional corepressor Sin3b) (Histone deacetylase complex subunit Sin3b). | |||||
|
SIN3A_MOUSE
|
||||||
| NC score | 0.950005 (rank : 3) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q60520, Q60820, Q62139, Q62140, Q7TPU8, Q7TSZ2 | Gene names | Sin3a | |||
|
Domain Architecture |

|
|||||
| Description | Paired amphipathic helix protein Sin3a (Transcriptional corepressor Sin3a) (Histone deacetylase complex subunit Sin3a). | |||||
|
SIN3A_HUMAN
|
||||||
| NC score | 0.946537 (rank : 4) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q96ST3, Q8N8N4, Q8NC83, Q8WV18, Q96L98, Q9UFQ1 | Gene names | SIN3A | |||
|
Domain Architecture |

|
|||||
| Description | Paired amphipathic helix protein Sin3a (Transcriptional corepressor Sin3a) (Histone deacetylase complex subunit Sin3a). | |||||
|
ZN645_HUMAN
|
||||||
| NC score | 0.064277 (rank : 5) | θ value | θ > 10 (rank : 83) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 265 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8N7E2, Q6DJY9 | Gene names | ZNF645 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 645. | |||||
|
GON4L_MOUSE
|
||||||
| NC score | 0.054031 (rank : 6) | θ value | 1.06291 (rank : 20) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9DB00, Q80TB4, Q91YI9 | Gene names | Gon4l, Gon4, KIAA1606 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GON-4-like protein (GON-4 homolog). | |||||
|
ACE_MOUSE
|
||||||
| NC score | 0.049988 (rank : 7) | θ value | 0.0961366 (rank : 6) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P09470, Q6GTS2 | Gene names | Ace, Dcp1 | |||
|
Domain Architecture |

|
|||||
| Description | Angiotensin-converting enzyme, somatic isoform precursor (EC 3.4.15.1) (Dipeptidyl carboxypeptidase I) (Kininase II) [Contains: Angiotensin- converting enzyme, somatic isoform, soluble form]. | |||||
|
CCD19_HUMAN
|
||||||
| NC score | 0.046142 (rank : 8) | θ value | 0.813845 (rank : 14) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 639 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UL16 | Gene names | CCDC19, NESG1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 19 (Nasopharyngeal epithelium- specific protein 1). | |||||
|
MAML2_HUMAN
|
||||||
| NC score | 0.045399 (rank : 9) | θ value | 0.0961366 (rank : 7) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8IZL2, Q6AI23, Q6Y3A3, Q8IUL3, Q96JK6 | Gene names | MAML2, KIAA1819 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mastermind-like protein 2 (Mam-2). | |||||
|
BRAP_MOUSE
|
||||||
| NC score | 0.038118 (rank : 10) | θ value | 5.27518 (rank : 48) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q99MP8, Q8CC00, Q8CHX1, Q99MP7, Q9CXX8 | Gene names | Brap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BRCA1-associated protein (EC 6.3.2.-) (BRAP2) (Impedes mitogenic signal propagation) (IMP). | |||||
|
RFTN1_MOUSE
|
||||||
| NC score | 0.036662 (rank : 11) | θ value | 0.279714 (rank : 13) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6A0D4, Q3U654, Q3U8P6, Q80US1 | Gene names | Rftn1, Kiaa0084 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Raftlin (Raft-linking protein). | |||||
|
TRIP4_MOUSE
|
||||||
| NC score | 0.035973 (rank : 12) | θ value | 3.0926 (rank : 39) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9QXN3, Q8CAD5 | Gene names | Trip4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Activating signal cointegrator 1 (ASC-1) (Thyroid receptor-interacting protein 4) (Trip-4). | |||||
|
OFD1_HUMAN
|
||||||
| NC score | 0.035514 (rank : 13) | θ value | 4.03905 (rank : 43) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 753 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O75665, O75666 | Gene names | OFD1, CXorf5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Oral-facial-digital syndrome 1 protein (Protein 71-7A). | |||||
|
NDE1_MOUSE
|
||||||
| NC score | 0.035093 (rank : 14) | θ value | 0.813845 (rank : 17) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 602 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9CZA6, Q3UBS6, Q3UIC1, Q9ERR0 | Gene names | Nde1, Nude | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear distribution protein nudE homolog 1 (NudE) (mNudE). | |||||
|
NDE1_HUMAN
|
||||||
| NC score | 0.034809 (rank : 15) | θ value | 0.279714 (rank : 12) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 823 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9NXR1, Q49AQ2 | Gene names | NDE1, NUDE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear distribution protein nudE homolog 1 (NudE). | |||||
|
TXLNB_HUMAN
|
||||||
| NC score | 0.034783 (rank : 16) | θ value | 0.0563607 (rank : 5) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 912 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8N3L3, Q76L25, Q86T52, Q8N3S2 | Gene names | TXLNB, C6orf198, MDP77 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-taxilin (Muscle-derived protein 77) (hMDP77). | |||||
|
GON4L_HUMAN
|
||||||
| NC score | 0.034513 (rank : 17) | θ value | 6.88961 (rank : 62) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 555 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q3T8J9, Q3T8J8, Q5VYZ5, Q5W0D5, Q6AWA6, Q6P1Q6, Q7Z3L3, Q8IY79, Q9BQI1, Q9HCG6 | Gene names | GON4L, GON4, KIAA1606 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GON-4-like protein (GON-4 homolog). | |||||
|
IF5_HUMAN
|
||||||
| NC score | 0.034461 (rank : 18) | θ value | 1.81305 (rank : 25) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P55010, Q9H5N2, Q9UG48 | Gene names | EIF5 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 5 (eIF-5). | |||||
|
CENPE_HUMAN
|
||||||
| NC score | 0.032558 (rank : 19) | θ value | 0.21417 (rank : 10) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 2060 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q02224 | Gene names | CENPE | |||
|
Domain Architecture |

|
|||||
| Description | Centromeric protein E (CENP-E). | |||||
|
RRP5_HUMAN
|
||||||
| NC score | 0.030410 (rank : 20) | θ value | 4.03905 (rank : 44) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 291 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q14690, Q86VQ8 | Gene names | PDCD11, KIAA0185 | |||
|
Domain Architecture |

|
|||||
| Description | RRP5 protein homolog (Programmed cell death protein 11). | |||||
|
CGRE1_HUMAN
|
||||||
| NC score | 0.030127 (rank : 21) | θ value | 0.813845 (rank : 15) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 354 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q99674 | Gene names | CGREF1, CGR11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell growth regulator with EF hand domain 1 (Cell growth regulatory gene 11 protein). | |||||
|
ACE_HUMAN
|
||||||
| NC score | 0.029879 (rank : 22) | θ value | 5.27518 (rank : 47) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P12821, Q53YX9, Q59GY8 | Gene names | ACE, DCP, DCP1 | |||
|
Domain Architecture |

|
|||||
| Description | Angiotensin-converting enzyme, somatic isoform precursor (EC 3.4.15.1) (Dipeptidyl carboxypeptidase I) (Kininase II) (CD143 antigen) [Contains: Angiotensin-converting enzyme, somatic isoform, soluble form]. | |||||
|
UBXD2_HUMAN
|
||||||
| NC score | 0.029415 (rank : 23) | θ value | 3.0926 (rank : 40) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q92575, Q8IYM5 | Gene names | UBXD2, KIAA0242, UBXDC1 | |||
|
Domain Architecture |

|
|||||
| Description | UBX domain-containing protein 2. | |||||
|
IF5_MOUSE
|
||||||
| NC score | 0.028761 (rank : 24) | θ value | 5.27518 (rank : 51) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P59325 | Gene names | Eif5 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 5 (eIF-5). | |||||
|
UBXD2_MOUSE
|
||||||
| NC score | 0.028168 (rank : 25) | θ value | 4.03905 (rank : 46) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8VCH8, Q8BW17 | Gene names | Ubxd2, Ubxdc1 | |||
|
Domain Architecture |

|
|||||
| Description | UBX domain-containing protein 2. | |||||
|
SMC1A_MOUSE
|
||||||
| NC score | 0.027252 (rank : 26) | θ value | 0.21417 (rank : 11) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1384 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9CU62, Q3V480, Q9CUX9, Q9D959, Q9WTU1 | Gene names | Smc1l1, Sb1.8, Smc1, Smcb | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 1-like 1 protein (SMC1alpha protein) (Chromosome segregation protein SmcB) (Sb1.8). | |||||
|
CN145_HUMAN
|
||||||
| NC score | 0.025822 (rank : 27) | θ value | 1.06291 (rank : 19) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1196 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q6ZU80, Q96ML4 | Gene names | C14orf145 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf145. | |||||
|
G137B_HUMAN
|
||||||
| NC score | 0.025761 (rank : 28) | θ value | 4.03905 (rank : 42) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O60478 | Gene names | GPR137B, TM7SF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Integral membrane protein GPR137B (Transmembrane 7 superfamily member 1). | |||||
|
DAB2P_HUMAN
|
||||||
| NC score | 0.025665 (rank : 29) | θ value | 3.0926 (rank : 36) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q5VWQ8, Q8TDL2, Q96SE1, Q9C0C0 | Gene names | DAB2IP, AF9Q34, AIP1, KIAA1743 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Disabled homolog 2-interacting protein (DAB2-interacting protein) (DAB2 interaction protein) (ASK-interacting protein 1). | |||||
|
DAB2P_MOUSE
|
||||||
| NC score | 0.025407 (rank : 30) | θ value | 3.0926 (rank : 37) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 284 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q3UHC7, Q3TPD5, Q3UH44, Q6JTV1, Q80T97 | Gene names | Dab2ip, Kiaa1743 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Disabled homolog 2-interacting protein (DAB2-interacting protein). | |||||
|
HOOK3_MOUSE
|
||||||
| NC score | 0.025111 (rank : 31) | θ value | 2.36792 (rank : 31) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1007 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8BUK6, Q8BV48, Q8BW57, Q8BY41 | Gene names | Hook3 | |||
|
Domain Architecture |

|
|||||
| Description | Hook homolog 3. | |||||
|
HOOK3_HUMAN
|
||||||
| NC score | 0.024963 (rank : 32) | θ value | 2.36792 (rank : 30) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1070 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q86VS8, Q8NBH0, Q9BY13 | Gene names | HOOK3 | |||
|
Domain Architecture |

|
|||||
| Description | Hook homolog 3 (hHK3). | |||||
|
MTSS1_HUMAN
|
||||||
| NC score | 0.024700 (rank : 33) | θ value | 2.36792 (rank : 33) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O43312, Q8TCA2, Q96RX2 | Gene names | MTSS1, KIAA0429, MIM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metastasis suppressor protein 1 (Missing in metastasis protein) (Metastasis suppressor YGL-1). | |||||
|
SMC1A_HUMAN
|
||||||
| NC score | 0.024621 (rank : 34) | θ value | 1.38821 (rank : 24) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1346 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q14683, O14995, Q16351 | Gene names | SMC1L1, DXS423E, KIAA0178, SB1.8, SMC1 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 1-like 1 protein (SMC1alpha protein) (Sb1.8). | |||||
|
CENPK_HUMAN
|
||||||
| NC score | 0.023983 (rank : 35) | θ value | 6.88961 (rank : 56) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 273 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9BS16, Q9H4L0 | Gene names | CENPK, ICEN37 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein K (CENP-K) (Interphase centromere complex protein 37) (Protein AF-5alpha) (p33). | |||||
|
OGFR_HUMAN
|
||||||
| NC score | 0.023982 (rank : 36) | θ value | 0.163984 (rank : 9) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
SCG3_MOUSE
|
||||||
| NC score | 0.023947 (rank : 37) | θ value | 3.0926 (rank : 38) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P47867 | Gene names | Scg3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Secretogranin-3 precursor (Secretogranin III) (SgIII). | |||||
|
COX10_MOUSE
|
||||||
| NC score | 0.023571 (rank : 38) | θ value | 2.36792 (rank : 28) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CFY5 | Gene names | Cox10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protoheme IX farnesyltransferase, mitochondrial precursor (EC 2.5.1.-) (Heme O synthase). | |||||
|
PCNT_MOUSE
|
||||||
| NC score | 0.022355 (rank : 39) | θ value | 0.813845 (rank : 18) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1207 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P48725 | Gene names | Pcnt, Pcnt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pericentrin. | |||||
|
TRIP4_HUMAN
|
||||||
| NC score | 0.022174 (rank : 40) | θ value | 6.88961 (rank : 67) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q15650, Q96ED7, Q9UKH0 | Gene names | TRIP4 | |||
|
Domain Architecture |

|
|||||
| Description | Activating signal cointegrator 1 (ASC-1) (Thyroid receptor-interacting protein 4) (TRIP-4). | |||||
|
DOK1_MOUSE
|
||||||
| NC score | 0.020966 (rank : 41) | θ value | 1.38821 (rank : 22) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P97465, Q9R213 | Gene names | Dok1, Dok | |||
|
Domain Architecture |

|
|||||
| Description | Docking protein 1 (Downstream of tyrosine kinase 1) (p62(dok)). | |||||
|
DZIP3_MOUSE
|
||||||
| NC score | 0.020838 (rank : 42) | θ value | 5.27518 (rank : 50) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 414 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q7TPV2, Q80TU4, Q8BYK7 | Gene names | Dzip3, Kiaa0675 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin ligase protein DZIP3 (EC 6.3.2.-) (DAZ-interacting protein 3 homolog). | |||||
|
MUG2_MOUSE
|
||||||
| NC score | 0.019617 (rank : 43) | θ value | 1.81305 (rank : 26) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P28666 | Gene names | Mug2, Mug-2 | |||
|
Domain Architecture |

|
|||||
| Description | Murinoglobulin-2 precursor (MuG2). | |||||
|
ITSN2_MOUSE
|
||||||
| NC score | 0.019464 (rank : 44) | θ value | 0.813845 (rank : 16) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1371 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9Z0R6, Q9Z0R5 | Gene names | Itsn2, Ese2, Sh3d1B | |||
|
Domain Architecture |

|
|||||
| Description | Intersectin-2 (SH3 domain-containing protein 1B) (EH and SH3 domains protein 2) (EH domain and SH3 domain regulator of endocytosis 2). | |||||
|
SEPT6_HUMAN
|
||||||
| NC score | 0.019425 (rank : 45) | θ value | 4.03905 (rank : 45) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 220 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q14141, Q5JTK0, Q969W5, Q96A13, Q96GR1, Q96P86, Q96P87 | Gene names | SEPT6, KIAA0128, SEP2 | |||
|
Domain Architecture |

|
|||||
| Description | Septin-6. | |||||
|
CTR9_MOUSE
|
||||||
| NC score | 0.018251 (rank : 46) | θ value | 5.27518 (rank : 49) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q62018, Q3UFF5, Q3UY40, Q66JX4, Q7TPS6, Q8BND9, Q8BRD1, Q8C9W7, Q8C9Y3, Q8CHI1 | Gene names | Ctr9, Kiaa0155, Sh2bp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein CTR9 homolog (SH2 domain-binding protein 1) (Tetratricopeptide repeat-containing, SH2-binding phosphoprotein of 150 kDa) (TPR-containing, SH2-binding phosphoprotein of 150 kDa) (p150TSP). | |||||
|
CI039_HUMAN
|
||||||
| NC score | 0.017684 (rank : 47) | θ value | 6.88961 (rank : 57) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1171 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9NXG0, Q5VYJ0, Q9HAJ5 | Gene names | C9orf39 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C9orf39. | |||||
|
SAPS3_MOUSE
|
||||||
| NC score | 0.017480 (rank : 48) | θ value | 1.38821 (rank : 23) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q922D4, Q6ZPM9, Q8BTW6, Q9D2X9 | Gene names | Saps3, D19Ertd703e, Kiaa1558 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SAPS domain family member 3. | |||||
|
CTR9_HUMAN
|
||||||
| NC score | 0.017453 (rank : 49) | θ value | 6.88961 (rank : 58) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 383 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6PD62, Q15015 | Gene names | CTR9, KIAA0155, SH2BP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein CTR9 homolog (SH2 domain-binding protein 1). | |||||
|
INP4A_MOUSE
|
||||||
| NC score | 0.016340 (rank : 50) | θ value | 2.36792 (rank : 32) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9EPW0, Q8CBJ8 | Gene names | Inpp4a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Type I inositol-3,4-bisphosphate 4-phosphatase (EC 3.1.3.66) (Inositol polyphosphate 4-phosphatase type I). | |||||
|
UT14C_HUMAN
|
||||||
| NC score | 0.016148 (rank : 51) | θ value | 6.88961 (rank : 68) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q5TAP6, Q5FWG3, Q92555 | Gene names | UTP14C, KIAA0266 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | U3 small nucleolar RNA-associated protein 14 homolog C. | |||||
|
SYCP1_MOUSE
|
||||||
| NC score | 0.015601 (rank : 52) | θ value | 6.88961 (rank : 65) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1456 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q62209, O09205, P70192, Q62329 | Gene names | Sycp1, Scp1 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptonemal complex protein 1 (SCP-1). | |||||
|
RGS3_HUMAN
|
||||||
| NC score | 0.014910 (rank : 53) | θ value | 8.99809 (rank : 80) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
CENPJ_HUMAN
|
||||||
| NC score | 0.014906 (rank : 54) | θ value | 8.99809 (rank : 73) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 721 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9HC77, Q5T6R5, Q96KS5, Q9C067 | Gene names | CENPJ, CPAP, LAP, LIP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein J (CENP-J) (Centrosomal P4.1-associated protein) (LAG-3-associated protein) (LYST-interacting protein 1). | |||||
|
UTRO_HUMAN
|
||||||
| NC score | 0.014752 (rank : 55) | θ value | 5.27518 (rank : 52) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P46939 | Gene names | UTRN, DMDL | |||
|
Domain Architecture |

|
|||||
| Description | Utrophin (Dystrophin-related protein 1) (DRP1) (DRP). | |||||
|
CX022_HUMAN
|
||||||
| NC score | 0.013968 (rank : 56) | θ value | 8.99809 (rank : 75) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6ZTR5, Q5JRM8, Q8N6X8 | Gene names | CXorf22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein CXorf22. | |||||
|
WAPL_MOUSE
|
||||||
| NC score | 0.013656 (rank : 57) | θ value | 5.27518 (rank : 53) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q65Z40, Q6ZQE9 | Gene names | Wapal, Kiaa0261, Wapl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wings apart-like protein homolog (Dioxin-inducible factor 2) (DIF-2). | |||||
|
HDAC5_HUMAN
|
||||||
| NC score | 0.013430 (rank : 58) | θ value | 2.36792 (rank : 29) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 353 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9UQL6, O60340, O60528, Q96DY4 | Gene names | HDAC5, KIAA0600 | |||
|
Domain Architecture |

|
|||||
| Description | Histone deacetylase 5 (HD5) (Antigen NY-CO-9). | |||||
|
ASCC2_HUMAN
|
||||||
| NC score | 0.012697 (rank : 59) | θ value | 6.88961 (rank : 55) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9H1I8, Q4TT54, Q8TAZ0, Q9H711, Q9H9D6 | Gene names | ASCC2, ASC1P100 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Activating signal cointegrator 1 complex subunit 2 (ASC-1 complex subunit p100) (Trip4 complex subunit p100). | |||||
|
RAI14_HUMAN
|
||||||
| NC score | 0.012638 (rank : 60) | θ value | 1.81305 (rank : 27) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1367 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9P0K7, Q6V1W9, Q7Z5I4, Q7Z733, Q9P2L2, Q9Y3T5 | Gene names | RAI14, KIAA1334, NORPEG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankycorbin (Ankyrin repeat and coiled-coil structure-containing protein) (Retinoic acid-induced protein 14) (Novel retinal pigment epithelial cell protein). | |||||
|
PIAS1_HUMAN
|
||||||
| NC score | 0.010975 (rank : 61) | θ value | 6.88961 (rank : 64) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O75925, Q99751, Q9UN02 | Gene names | PIAS1, DDXBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein inhibitor of activated STAT protein 1 (Gu-binding protein) (GBP) (RNA helicase II-binding protein) (DEAD/H box-binding protein 1). | |||||
|
APBA2_MOUSE
|
||||||
| NC score | 0.010418 (rank : 62) | θ value | 4.03905 (rank : 41) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P98084, Q6PAJ2 | Gene names | Apba2, X11l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 precursor protein-binding family A member 2 (Neuron- specific X11L protein) (Neuronal Munc18-1-interacting protein 2) (Mint-2) (Adapter protein X11beta). | |||||
|
NEK11_HUMAN
|
||||||
| NC score | 0.010353 (rank : 63) | θ value | 0.163984 (rank : 8) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1043 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8NG66, Q5JPC0, Q8NG65, Q8TBY1, Q9H5F4 | Gene names | NEK11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase Nek11 (EC 2.7.11.1) (NimA-related protein kinase 11) (Never in mitosis A-related kinase 11). | |||||
|
CTRO_MOUSE
|
||||||
| NC score | 0.009220 (rank : 64) | θ value | 6.88961 (rank : 60) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 2344 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P49025, O88528, O88937, O88938, Q3UM99, Q8CIJ1 | Gene names | Cit | |||
|
Domain Architecture |

|
|||||
| Description | Citron Rho-interacting kinase (EC 2.7.11.1) (CRIK) (Rho-interacting, serine/threonine-protein kinase 21). | |||||
|
CTRO_HUMAN
|
||||||
| NC score | 0.008567 (rank : 65) | θ value | 6.88961 (rank : 59) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 2092 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O14578, Q6XUH8, Q86UQ9, Q9UPZ7 | Gene names | CIT, KIAA0949, STK21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Citron Rho-interacting kinase (EC 2.7.11.1) (CRIK) (Rho-interacting, serine/threonine-protein kinase 21). | |||||
|
EPN2_MOUSE
|
||||||
| NC score | 0.008561 (rank : 66) | θ value | 6.88961 (rank : 61) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8CHU3, O70447, Q8BZ85 | Gene names | Epn2 | |||
|
Domain Architecture |

|
|||||
| Description | Epsin-2 (EPS-15-interacting protein 2) (Intersectin-EH-binding protein 2) (Ibp2). | |||||
|
OXRP_HUMAN
|
||||||
| NC score | 0.008524 (rank : 67) | θ value | 8.99809 (rank : 78) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 544 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Y4L1 | Gene names | HYOU1, ORP150 | |||
|
Domain Architecture |

|
|||||
| Description | 150 kDa oxygen-regulated protein precursor (Orp150) (Hypoxia up- regulated 1). | |||||
|
PEG3_MOUSE
|
||||||
| NC score | 0.008521 (rank : 68) | θ value | 1.06291 (rank : 21) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1101 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q3URU2, O54978, Q3TQ69, Q5EBP7, Q61138, Q6GQS0, Q80U47, Q8R5N0, Q9QX53 | Gene names | Peg3, Kiaa0287, Pw1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Paternally expressed gene 3 protein (ASF-1). | |||||
|
TAU_HUMAN
|
||||||
| NC score | 0.008503 (rank : 69) | θ value | 6.88961 (rank : 66) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 448 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P10636, P18518, Q14799, Q15549, Q15550, Q15551, Q9UDJ3, Q9UMH0, Q9UQ96 | Gene names | MAPT, MTBT1, TAU | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein tau (Neurofibrillary tangle protein) (Paired helical filament-tau) (PHF-tau). | |||||
|
SHAN2_HUMAN
|
||||||
| NC score | 0.008248 (rank : 70) | θ value | 8.99809 (rank : 81) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 346 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UPX8, Q3Y8G9, Q52LK2, Q9UKP1 | Gene names | SHANK2, KIAA1022 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 and multiple ankyrin repeat domains protein 2 (Shank2). | |||||
|
SLK_HUMAN
|
||||||
| NC score | 0.007784 (rank : 71) | θ value | 2.36792 (rank : 35) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1775 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9H2G2, O00211, Q6P1Z4, Q86WU7, Q86WW1, Q92603, Q9NQL0, Q9NQL1 | Gene names | SLK, KIAA0204, STK2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | STE20-like serine/threonine-protein kinase (EC 2.7.11.1) (STE20-like kinase) (STE20-related serine/threonine-protein kinase) (STE20-related kinase) (hSLK) (Serine/threonine-protein kinase 2) (CTCL tumor antigen se20-9). | |||||
|
AFF2_MOUSE
|
||||||
| NC score | 0.007264 (rank : 72) | θ value | 8.99809 (rank : 71) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O55112 | Gene names | Aff2, Fmr2, Ox19 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 2 (Fragile X mental retardation protein 2 homolog) (Protein FMR-2) (FMR2P) (Protein Ox19). | |||||
|
PUM2_HUMAN
|
||||||
| NC score | 0.007104 (rank : 73) | θ value | 8.99809 (rank : 79) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8TB72, O00234, Q53TV7, Q8WY43, Q9HAN2 | Gene names | PUM2, KIAA0235, PUMH2 | |||
|
Domain Architecture |

|
|||||
| Description | Pumilio homolog 2 (Pumilio-2). | |||||
|
MYO6_MOUSE
|
||||||
| NC score | 0.006985 (rank : 74) | θ value | 6.88961 (rank : 63) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 586 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q64331 | Gene names | Myo6, Sv | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-6 (Myosin VI). | |||||
|
PTN21_HUMAN
|
||||||
| NC score | 0.006980 (rank : 75) | θ value | 2.36792 (rank : 34) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q16825 | Gene names | PTPN21, PTPD1 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 21 (EC 3.1.3.48) (Protein-tyrosine phosphatase D1). | |||||
|
CLMN_MOUSE
|
||||||
| NC score | 0.006458 (rank : 76) | θ value | 8.99809 (rank : 74) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8C5W0, Q91V71, Q91XT7, Q91XT8, Q91XU9 | Gene names | Clmn | |||
|
Domain Architecture |

|
|||||
| Description | Calmin. | |||||
|
E41L2_HUMAN
|
||||||
| NC score | 0.006018 (rank : 77) | θ value | 8.99809 (rank : 76) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 287 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O43491 | Gene names | EPB41L2 | |||
|
Domain Architecture |

|
|||||
| Description | Band 4.1-like protein 2 (Generally expressed protein 4.1) (4.1G). | |||||
|
JPH1_MOUSE
|
||||||
| NC score | 0.005996 (rank : 78) | θ value | 8.99809 (rank : 77) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9ET80, Q9EQZ3 | Gene names | Jph1, Jp1 | |||
|
Domain Architecture |

|
|||||
| Description | Junctophilin-1 (Junctophilin type 1) (JP-1). | |||||
|
CAN3_HUMAN
|
||||||
| NC score | 0.003643 (rank : 79) | θ value | 8.99809 (rank : 72) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P20807, Q9BTU4, Q9Y5S6, Q9Y5S7 | Gene names | CAPN3, CANP3, CANPL3, NCL1 | |||
|
Domain Architecture |

|
|||||
| Description | Calpain-3 (EC 3.4.22.-) (Calpain L3) (Calpain p94) (Calcium-activated neutral proteinase 3) (CANP 3) (Muscle-specific calcium-activated neutral protease 3) (nCL-1). | |||||
|
ZBTB3_HUMAN
|
||||||
| NC score | 0.003180 (rank : 80) | θ value | 5.27518 (rank : 54) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 910 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9H5J0 | Gene names | ZBTB3 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and BTB domain-containing protein 3. | |||||
|
ZN287_MOUSE
|
||||||
| NC score | 0.002632 (rank : 81) | θ value | 6.88961 (rank : 70) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 771 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9EQB9 | Gene names | Znf287, Skat2, Zfp287 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 287 (Zfp-287) (Zinc finger protein SKAT-2). | |||||
|
ZN287_HUMAN
|
||||||
| NC score | 0.002214 (rank : 82) | θ value | 8.99809 (rank : 82) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 777 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9HBT7 | Gene names | ZNF287 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 287. | |||||
|
ZFP95_MOUSE
|
||||||
| NC score | 0.001066 (rank : 83) | θ value | 6.88961 (rank : 69) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 787 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Z1D8 | Gene names | Zfp95 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 95 (Zfp-95). | |||||