Please be patient as the page loads
|
RNF25_MOUSE
|
||||||
| SwissProt Accessions | Q9QZR0, Q3TAL8, Q9DCW7 | Gene names | Rnf25 | |||
|
Domain Architecture |
|
|||||
| Description | RING finger protein 25 (EC 6.3.2.-) (RING finger protein AO7). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
RNF25_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.949209 (rank : 2) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q96BH1, Q9H874 | Gene names | RNF25 | |||
|
Domain Architecture |
|
|||||
| Description | RING finger protein 25 (EC 6.3.2.-). | |||||
|
RNF25_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 106 | |
| SwissProt Accessions | Q9QZR0, Q3TAL8, Q9DCW7 | Gene names | Rnf25 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 25 (EC 6.3.2.-) (RING finger protein AO7). | |||||
|
E2AK4_MOUSE
|
||||||
| θ value | 9.59137e-10 (rank : 3) | NC score | 0.116055 (rank : 37) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 984 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9QZ05, Q6GQT4, Q6ZPT5, Q8C5S0, Q8CIF5, Q9CT30, Q9CUV9, Q9ESB6, Q9ESB7, Q9ESB8 | Gene names | Eif2ak4, Gcn2, Kiaa1338 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 2-alpha kinase 4 (EC 2.7.11.1) (GCN2-like protein) (mGCN2). | |||||
|
E2AK4_HUMAN
|
||||||
| θ value | 1.06045e-08 (rank : 4) | NC score | 0.109564 (rank : 38) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 1012 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9P2K8, Q69YL7, Q6DC97, Q96GN6, Q9H5K1, Q9NSQ3, Q9NSZ5, Q9UJ56 | Gene names | EIF2AK4, GCN2, KIAA1338 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 2-alpha kinase 4 (EC 2.7.11.1) (GCN2-like protein). | |||||
|
RWDD1_HUMAN
|
||||||
| θ value | 4.1701e-05 (rank : 5) | NC score | 0.349776 (rank : 3) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9H446, Q9Y313, Q9Y6B3 | Gene names | RWDD1 | |||
|
Domain Architecture |
|
|||||
| Description | RWD domain-containing protein 1. | |||||
|
RWDD1_MOUSE
|
||||||
| θ value | 4.1701e-05 (rank : 6) | NC score | 0.348046 (rank : 4) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9CQK7 | Gene names | Rwdd1 | |||
|
Domain Architecture |
|
|||||
| Description | RWD domain-containing protein 1 (IH1). | |||||
|
RNF24_MOUSE
|
||||||
| θ value | 0.00665767 (rank : 7) | NC score | 0.252604 (rank : 5) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8BGI1 | Gene names | Rnf24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 24. | |||||
|
RNF24_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 8) | NC score | 0.241979 (rank : 6) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9Y225, Q9UMH1 | Gene names | RNF24 | |||
|
Domain Architecture |
|
|||||
| Description | RING finger protein 24. | |||||
|
CBP_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 9) | NC score | 0.079819 (rank : 44) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 1051 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P45481 | Gene names | Crebbp, Cbp | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
RNF12_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 10) | NC score | 0.221792 (rank : 9) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9WTV7 | Gene names | Rnf12, Rlim | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 12 (LIM domain-interacting RING finger protein) (RING finger LIM domain-binding protein) (R-LIM). | |||||
|
PJA1_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 11) | NC score | 0.209013 (rank : 15) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8NG27, Q8NG28, Q9HAC1 | Gene names | PJA1, RNF70 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin protein ligase Praja1 (EC 6.3.2.-) (RING finger protein 70). | |||||
|
PJA1_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 12) | NC score | 0.213813 (rank : 12) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O55176, Q8CFU2, Q99MJ1, Q99MJ2, Q99MJ3, Q9DB04 | Gene names | Pja1 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin protein ligase Praja1 (EC 6.3.2.-). | |||||
|
CBP_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 13) | NC score | 0.088749 (rank : 39) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 941 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q92793, O00147, Q16376 | Gene names | CREBBP, CBP | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
RNF38_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 14) | NC score | 0.214384 (rank : 11) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9H0F5, Q8N0Y0 | Gene names | RNF38 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 38. | |||||
|
CU006_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 15) | NC score | 0.116622 (rank : 36) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q99M03, Q9JLH4 | Gene names | ORF5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C21orf6 homolog. | |||||
|
RNF38_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 16) | NC score | 0.212701 (rank : 13) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8BI21, Q6P5B1 | Gene names | Rnf38 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 38. | |||||
|
RN122_HUMAN
|
||||||
| θ value | 0.125558 (rank : 17) | NC score | 0.232746 (rank : 8) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9H9V4, Q52LK3 | Gene names | RNF122 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 122. | |||||
|
RN122_MOUSE
|
||||||
| θ value | 0.125558 (rank : 18) | NC score | 0.233368 (rank : 7) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8BP31, Q80VA7, Q8BGD3 | Gene names | Rnf122 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 122. | |||||
|
RNF6_HUMAN
|
||||||
| θ value | 0.125558 (rank : 19) | NC score | 0.215397 (rank : 10) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9Y252, Q9UF41 | Gene names | RNF6 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 6 (RING-H2 protein). | |||||
|
RNF12_HUMAN
|
||||||
| θ value | 0.163984 (rank : 20) | NC score | 0.209346 (rank : 14) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9NVW2, Q9Y598 | Gene names | RNF12, RLIM | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 12 (LIM domain-interacting RING finger protein) (RING finger LIM domain-binding protein) (R-LIM) (NY-REN-43 antigen). | |||||
|
CU006_HUMAN
|
||||||
| θ value | 0.21417 (rank : 21) | NC score | 0.086148 (rank : 41) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P57060 | Gene names | C21orf6 | |||
|
Domain Architecture |
|
|||||
| Description | Protein C21orf6. | |||||
|
NEK4_MOUSE
|
||||||
| θ value | 0.21417 (rank : 22) | NC score | 0.013578 (rank : 128) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 899 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Z1J2, O35673, Q9R1J1 | Gene names | Nek4, Stk2 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase Nek4 (EC 2.7.11.1) (NimA-related protein kinase 4) (Serine/threonine-protein kinase 2). | |||||
|
ZN364_HUMAN
|
||||||
| θ value | 0.21417 (rank : 23) | NC score | 0.195464 (rank : 18) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9Y4L5, Q5T2V9, Q7Z2J2 | Gene names | ZNF364, RNF115 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 364 (Rabring 7) (RING finger protein 115). | |||||
|
ZSWM2_HUMAN
|
||||||
| θ value | 0.21417 (rank : 24) | NC score | 0.167603 (rank : 27) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8NEG5 | Gene names | ZSWIM2 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger SWIM domain-containing protein 2. | |||||
|
ZN364_MOUSE
|
||||||
| θ value | 0.279714 (rank : 25) | NC score | 0.192532 (rank : 20) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9D0C1, Q8R5A1, Q9D885 | Gene names | Znf364, Zfp364 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 364 (Rabring 7). | |||||
|
CO4A1_MOUSE
|
||||||
| θ value | 0.365318 (rank : 26) | NC score | 0.024673 (rank : 111) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 450 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P02463 | Gene names | Col4a1 | |||
|
Domain Architecture |
|
|||||
| Description | Collagen alpha-1(IV) chain precursor. | |||||
|
FHOD1_HUMAN
|
||||||
| θ value | 0.365318 (rank : 27) | NC score | 0.043378 (rank : 91) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Y613, Q59F76, Q6Y1F2, Q76MS8, Q8N521 | Gene names | FHOD1, FHOS | |||
|
Domain Architecture |

|
|||||
| Description | FH1/FH2 domain-containing protein (Formin homolog overexpressed in spleen) (FHOS) (Formin homology 2 domain-containing protein 1). | |||||
|
RNF13_MOUSE
|
||||||
| θ value | 0.365318 (rank : 28) | NC score | 0.189828 (rank : 21) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 164 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O54965, O54966 | Gene names | Rnf13, Rzf | |||
|
Domain Architecture |
|
|||||
| Description | RING finger protein 13. | |||||
|
GOLI_MOUSE
|
||||||
| θ value | 0.47712 (rank : 29) | NC score | 0.189616 (rank : 22) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8VEM1, Q80VL7, Q9QZQ6 | Gene names | Rnf130, G1rp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Goliath homolog precursor (RING finger protein 130) (G1-related zinc finger protein). | |||||
|
RN165_HUMAN
|
||||||
| θ value | 0.47712 (rank : 30) | NC score | 0.194199 (rank : 19) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q6ZSG1 | Gene names | RNF165 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 165. | |||||
|
RNF13_HUMAN
|
||||||
| θ value | 0.47712 (rank : 31) | NC score | 0.186276 (rank : 24) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O43567 | Gene names | RNF13, RZF | |||
|
Domain Architecture |
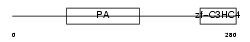
|
|||||
| Description | RING finger protein 13. | |||||
|
TRML2_MOUSE
|
||||||
| θ value | 0.47712 (rank : 32) | NC score | 0.058038 (rank : 74) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q2LA85 | Gene names | Treml2, Tlt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trem-like transcript 2 protein precursor (TLT-2) (Triggering receptor expressed on myeloid cells-like protein 2). | |||||
|
GAK11_HUMAN
|
||||||
| θ value | 0.62314 (rank : 33) | NC score | 0.069617 (rank : 56) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P63145, Q9UKI1 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_22q11.21 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K101 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK12_HUMAN
|
||||||
| θ value | 0.62314 (rank : 34) | NC score | 0.070155 (rank : 54) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P63130, Q9UKI0 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_1q22 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K102 Gag protein) (HERV-K(III) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK4_HUMAN
|
||||||
| θ value | 0.62314 (rank : 35) | NC score | 0.069641 (rank : 55) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 415 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P63126, Q9UKH4 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_6q14.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K109 Gag protein) (HERV-K(C6) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GOLI_HUMAN
|
||||||
| θ value | 0.62314 (rank : 36) | NC score | 0.187060 (rank : 23) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q86XS8, Q641P9, Q6P177, Q7L2U2, Q9P0J9 | Gene names | RNF130 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Goliath homolog precursor (RING finger protein 130). | |||||
|
RN139_HUMAN
|
||||||
| θ value | 0.62314 (rank : 37) | NC score | 0.164270 (rank : 30) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8WU17, O75485, Q7LDL3 | Gene names | RNF139, TRC8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 139 (Translocation in renal carcinoma on chromosome 8). | |||||
|
GAK1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 38) | NC score | 0.069235 (rank : 57) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 268 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P62683 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_12q14.1 provirus ancestral Gag polyprotein (Gag polyprotein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
YLPM1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 39) | NC score | 0.059599 (rank : 73) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P49750, P49752, Q96I64, Q9P1V7 | Gene names | YLPM1, C14orf170, ZAP3 | |||
|
Domain Architecture |

|
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3) (ZAP113). | |||||
|
EP300_HUMAN
|
||||||
| θ value | 1.06291 (rank : 40) | NC score | 0.087362 (rank : 40) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q09472 | Gene names | EP300, P300 | |||
|
Domain Architecture |

|
|||||
| Description | E1A-associated protein p300 (EC 2.3.1.48). | |||||
|
RN139_MOUSE
|
||||||
| θ value | 1.06291 (rank : 41) | NC score | 0.157628 (rank : 31) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q7TMV1, Q8BZU9 | Gene names | Rnf139 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 139. | |||||
|
RNF11_HUMAN
|
||||||
| θ value | 1.06291 (rank : 42) | NC score | 0.199784 (rank : 16) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9Y3C5 | Gene names | RNF11 | |||
|
Domain Architecture |
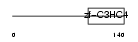
|
|||||
| Description | RING finger protein 11 (Sid 1669). | |||||
|
RNF11_MOUSE
|
||||||
| θ value | 1.06291 (rank : 43) | NC score | 0.199204 (rank : 17) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9QYK7, Q9EQI1 | Gene names | Rnf11, N4wbp2, Sid1669 | |||
|
Domain Architecture |
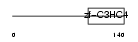
|
|||||
| Description | RING finger protein 11 (NEDD4 WW domain-binding protein 2) (Sid 1669). | |||||
|
SYN2_MOUSE
|
||||||
| θ value | 1.06291 (rank : 44) | NC score | 0.050809 (rank : 87) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q64332, Q6NZR0, Q9QWV7 | Gene names | Syn2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synapsin-2 (Synapsin II). | |||||
|
BSN_MOUSE
|
||||||
| θ value | 1.38821 (rank : 45) | NC score | 0.042326 (rank : 94) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
HNRL1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 46) | NC score | 0.033351 (rank : 102) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8VDM6, Q3U201, Q3UPB0, Q6AZA7, Q8BY45, Q8K365 | Gene names | Hnrpul1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heterogeneous nuclear ribonucleoprotein U-like protein 1. | |||||
|
NCOA6_MOUSE
|
||||||
| θ value | 1.38821 (rank : 47) | NC score | 0.047705 (rank : 88) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 1076 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9JL19, Q9JLT9 | Gene names | Ncoa6, Aib3, Prip, Rap250, Trbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC) (Thyroid hormone receptor-binding protein). | |||||
|
PERQ1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 48) | NC score | 0.043486 (rank : 90) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O75420 | Gene names | PERQ1 | |||
|
Domain Architecture |

|
|||||
| Description | PERQ amino acid rich with GYF domain protein 1. | |||||
|
RN103_HUMAN
|
||||||
| θ value | 1.38821 (rank : 49) | NC score | 0.148278 (rank : 35) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O00237, Q8IVB9 | Gene names | RNF103 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 103 (Zinc finger protein 103 homolog) (Zfp-103) (KF-1) (hKF-1). | |||||
|
RN103_MOUSE
|
||||||
| θ value | 1.38821 (rank : 50) | NC score | 0.149324 (rank : 34) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9R1W3, O08670, O08883 | Gene names | Rnf103, Zfp103 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 103 (Zinc finger protein 103) (Zfp-103) (KF-1) (mKF-1). | |||||
|
TXLNB_MOUSE
|
||||||
| θ value | 1.38821 (rank : 51) | NC score | 0.017265 (rank : 125) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 640 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8VBT1, Q3UVB8, Q8BUK2 | Gene names | Txlnb, Mdp77 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-taxilin (Muscle-derived protein 77). | |||||
|
GAK2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 52) | NC score | 0.067443 (rank : 60) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q7LDI9, Q9UKH5, Q9Y6I1, Q9YNA6, Q9YNB0 | Gene names | ERVK6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_7p22.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K(HML-2.HOM) Gag protein) (HERV-K108 Gag protein) (HERV-K(C7) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK3_HUMAN
|
||||||
| θ value | 1.81305 (rank : 53) | NC score | 0.067607 (rank : 59) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 182 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9YNA8 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_19q12 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K(C19) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK6_HUMAN
|
||||||
| θ value | 1.81305 (rank : 54) | NC score | 0.067994 (rank : 58) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P62685 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_8p23.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K115 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
PRG4_HUMAN
|
||||||
| θ value | 1.81305 (rank : 55) | NC score | 0.029818 (rank : 103) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q92954, Q6DNC4, Q6DNC5, Q6ZMZ5, Q9BX49 | Gene names | PRG4, MSF, SZP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
SYN3_HUMAN
|
||||||
| θ value | 1.81305 (rank : 56) | NC score | 0.051999 (rank : 84) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O14994 | Gene names | SYN3 | |||
|
Domain Architecture |
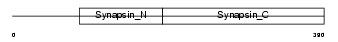
|
|||||
| Description | Synapsin-3 (Synapsin III). | |||||
|
SYQ_HUMAN
|
||||||
| θ value | 1.81305 (rank : 57) | NC score | 0.037118 (rank : 100) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P47897 | Gene names | QARS | |||
|
Domain Architecture |

|
|||||
| Description | Glutaminyl-tRNA synthetase (EC 6.1.1.18) (Glutamine--tRNA ligase) (GlnRS). | |||||
|
CN004_HUMAN
|
||||||
| θ value | 2.36792 (rank : 58) | NC score | 0.041630 (rank : 96) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9H1B7, Q8NDQ2, Q96JG2, Q9H3I7 | Gene names | C14orf4, KIAA1865 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf4. | |||||
|
CRF_HUMAN
|
||||||
| θ value | 2.36792 (rank : 59) | NC score | 0.042776 (rank : 92) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P06850 | Gene names | CRH | |||
|
Domain Architecture |

|
|||||
| Description | Corticoliberin precursor (Corticotropin-releasing factor) (CRF) (Corticotropin-releasing hormone). | |||||
|
EVPL_MOUSE
|
||||||
| θ value | 2.36792 (rank : 60) | NC score | 0.013131 (rank : 130) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 952 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9D952 | Gene names | Evpl | |||
|
Domain Architecture |

|
|||||
| Description | Envoplakin (p210) (210 kDa cornified envelope precursor protein). | |||||
|
IF4G3_HUMAN
|
||||||
| θ value | 2.36792 (rank : 61) | NC score | 0.019728 (rank : 120) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O43432, Q15597, Q8NEN1 | Gene names | EIF4G3 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 3 (eIF-4-gamma 3) (eIF-4G 3) (eIF4G 3) (eIF-4-gamma II) (eIF4GII). | |||||
|
NADAP_HUMAN
|
||||||
| θ value | 2.36792 (rank : 62) | NC score | 0.038946 (rank : 98) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9BWU0, Q4KMT1, Q4KMX0, Q7Z5Q9, Q9NVN2 | Gene names | SLC4A1AP, HLC3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kanadaptin (Kidney anion exchanger adapter protein) (Solute carrier family 4 anion exchanger member 1 adapter protein) (Lung cancer oncogene 3 protein). | |||||
|
PRG4_MOUSE
|
||||||
| θ value | 2.36792 (rank : 63) | NC score | 0.025890 (rank : 108) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 624 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9JM99, Q3UEL1, Q3V198 | Gene names | Prg4, Msf, Szp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
RFC1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 64) | NC score | 0.024884 (rank : 110) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P35601 | Gene names | Rfc1, Ibf-1, Recc1 | |||
|
Domain Architecture |

|
|||||
| Description | Replication factor C subunit 1 (Replication factor C large subunit) (RF-C 140 kDa subunit) (Activator 1 140 kDa subunit) (Activator 1 large subunit) (A1 140 kDa subunit) (A1-P145) (Differentiation- specific element-binding protein) (ISRE-binding protein). | |||||
|
TFAP4_HUMAN
|
||||||
| θ value | 2.36792 (rank : 65) | NC score | 0.028156 (rank : 105) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q01664, O60409 | Gene names | TFAP4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor AP-4 (Activating enhancer-binding protein 4). | |||||
|
ZSWM2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 66) | NC score | 0.153074 (rank : 32) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9D9X6 | Gene names | Zswim2 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger SWIM domain-containing protein 2. | |||||
|
DYN3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 67) | NC score | 0.026505 (rank : 106) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UQ16, O14982, O95555, Q5W129, Q9H0P3, Q9H548, Q9NQ68, Q9NQN6 | Gene names | DNM3, KIAA0820 | |||
|
Domain Architecture |

|
|||||
| Description | Dynamin-3 (EC 3.6.5.5) (Dynamin, testicular) (T-dynamin). | |||||
|
HCN4_HUMAN
|
||||||
| θ value | 3.0926 (rank : 68) | NC score | 0.020149 (rank : 118) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9Y3Q4, Q9UMQ7 | Gene names | HCN4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 4. | |||||
|
LR37B_HUMAN
|
||||||
| θ value | 3.0926 (rank : 69) | NC score | 0.018210 (rank : 123) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q96QE4, Q5YKG6 | Gene names | LRRC37B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 37B precursor (C66 SLIT-like testicular protein). | |||||
|
MAT1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 70) | NC score | 0.025951 (rank : 107) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P51949, Q3TZP0, Q9D8A0, Q9D8D2 | Gene names | Mnat1, Mat1 | |||
|
Domain Architecture |
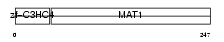
|
|||||
| Description | CDK-activating kinase assembly factor MAT1 (RING finger protein MAT1) (Menage a trois) (CDK7/cyclin H assembly factor) (p36) (p35). | |||||
|
RN126_MOUSE
|
||||||
| θ value | 3.0926 (rank : 71) | NC score | 0.164856 (rank : 29) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q91YL2 | Gene names | Rnf126 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 126. | |||||
|
CCD1A_MOUSE
|
||||||
| θ value | 4.03905 (rank : 72) | NC score | 0.024651 (rank : 112) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8K1A6, Q2MJB5, Q8R3Z4 | Gene names | Cc2d1a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil and C2 domain-containing protein 1A (Five repressor element under dual repression-binding protein 1) (FRE under dual repression-binding protein 1) (Freud-1). | |||||
|
CCDC4_HUMAN
|
||||||
| θ value | 4.03905 (rank : 73) | NC score | 0.037989 (rank : 99) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6ZU67, Q58A26, Q58A27 | Gene names | CCDC4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 4. | |||||
|
CO4A_HUMAN
|
||||||
| θ value | 4.03905 (rank : 74) | NC score | 0.021172 (rank : 114) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P0C0L4, P01028, P78445, Q13160, Q13906, Q14033, Q14835, Q5JQM8, Q9NPK5, Q9UIP5 | Gene names | C4A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Complement C4-A precursor (Acidic complement C4) [Contains: Complement C4 beta chain; Complement C4-A alpha chain; C4a anaphylatoxin; C4b-A; C4d-A; Complement C4 gamma chain]. | |||||
|
CO4B_HUMAN
|
||||||
| θ value | 4.03905 (rank : 75) | NC score | 0.021161 (rank : 115) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P0C0L5, P01028, P78445, Q13160, Q13906, Q14033, Q14835, Q9NPK5, Q9UIP5 | Gene names | C4B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Complement C4-B precursor (Basic complement C4) [Contains: Complement C4 beta chain; Complement C4-B alpha chain; C4a anaphylatoxin; C4b-B; C4d-B; Complement C4 gamma chain]. | |||||
|
INVS_HUMAN
|
||||||
| θ value | 4.03905 (rank : 76) | NC score | 0.011523 (rank : 131) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9Y283, Q5W0T6, Q8IVX8, Q9BRB9, Q9Y488, Q9Y498 | Gene names | INVS, INV, NPHP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inversin (Inversion of embryo turning homolog) (Nephrocystin-2). | |||||
|
RN167_HUMAN
|
||||||
| θ value | 4.03905 (rank : 77) | NC score | 0.172362 (rank : 25) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9H6Y7, Q6XYE0, Q8NDC1, Q9Y3V1 | Gene names | RNF167 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 167 precursor. | |||||
|
RN167_MOUSE
|
||||||
| θ value | 4.03905 (rank : 78) | NC score | 0.167099 (rank : 28) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q91XF4, Q3U4S5 | Gene names | Rnf167 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 167 precursor. | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 79) | NC score | 0.029508 (rank : 104) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
BAZ2A_HUMAN
|
||||||
| θ value | 5.27518 (rank : 80) | NC score | 0.020060 (rank : 119) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 780 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9UIF9, O00536, O15030, Q96H26 | Gene names | BAZ2A, KIAA0314, TIP5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5) (hWALp3). | |||||
|
DJBP_MOUSE
|
||||||
| θ value | 5.27518 (rank : 81) | NC score | 0.017986 (rank : 124) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6P1E8, Q5DTV8, Q6PGK8, Q9D2D8 | Gene names | Djbp, Kiaa1672 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DJ-1-binding protein. | |||||
|
MLL2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 82) | NC score | 0.033652 (rank : 101) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
RN133_HUMAN
|
||||||
| θ value | 5.27518 (rank : 83) | NC score | 0.169777 (rank : 26) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q8WVZ7, Q8N7G7 | Gene names | RNF133 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 133. | |||||
|
SIRT1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 84) | NC score | 0.016137 (rank : 127) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q923E4, Q9QXG8 | Gene names | Sirt1, Sir2l1 | |||
|
Domain Architecture |

|
|||||
| Description | NAD-dependent deacetylase sirtuin-1 (EC 3.5.1.-) (SIR2alpha) (mSIR2a) (Sir2) (SIR2-like protein 1). | |||||
|
SYN2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 85) | NC score | 0.042415 (rank : 93) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q92777 | Gene names | SYN2 | |||
|
Domain Architecture |
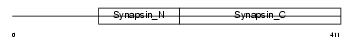
|
|||||
| Description | Synapsin-2 (Synapsin II). | |||||
|
ANDR_HUMAN
|
||||||
| θ value | 6.88961 (rank : 86) | NC score | 0.005670 (rank : 136) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P10275 | Gene names | AR, DHTR, NR3C4 | |||
|
Domain Architecture |

|
|||||
| Description | Androgen receptor (Dihydrotestosterone receptor). | |||||
|
AZI1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 87) | NC score | 0.013431 (rank : 129) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 990 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q62036 | Gene names | Azi1, Az1, Azi | |||
|
Domain Architecture |

|
|||||
| Description | 5-azacytidine-induced protein 1 (Pre-acrosome localization protein 1). | |||||
|
EVPL_HUMAN
|
||||||
| θ value | 6.88961 (rank : 88) | NC score | 0.011511 (rank : 132) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 895 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q92817 | Gene names | EVPL | |||
|
Domain Architecture |

|
|||||
| Description | Envoplakin (210 kDa paraneoplastic pemphigus antigen) (p210) (210 kDa cornified envelope precursor protein). | |||||
|
FOXP4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 89) | NC score | 0.010135 (rank : 133) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8IVH2, Q96E19 | Gene names | FOXP4, FKHLA | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein P4 (Fork head-related protein-like A). | |||||
|
MAML2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 90) | NC score | 0.060315 (rank : 70) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8IZL2, Q6AI23, Q6Y3A3, Q8IUL3, Q96JK6 | Gene names | MAML2, KIAA1819 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mastermind-like protein 2 (Mam-2). | |||||
|
NFAT5_HUMAN
|
||||||
| θ value | 6.88961 (rank : 91) | NC score | 0.020165 (rank : 117) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O94916, O95693, Q9UN18 | Gene names | NFAT5, KIAA0827, TONEBP | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor of activated T-cells 5 (T-cell transcription factor NFAT5) (NF-AT5) (Tonicity-responsive enhancer-binding protein) (TonE- binding protein) (TonEBP). | |||||
|
RP1L1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 92) | NC score | 0.020596 (rank : 116) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 1072 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8IWN7, Q86SQ1, Q8IWN8, Q8IWN9, Q8IWP0, Q8IWP1, Q8IWP2 | Gene names | RP1L1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinitis pigmentosa 1-like 1 protein. | |||||
|
RWDD3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 93) | NC score | 0.042258 (rank : 95) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Y3V2, Q6FID3, Q9BX35 | Gene names | RWDD3 | |||
|
Domain Architecture |
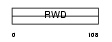
|
|||||
| Description | RWD domain-containing protein 3. | |||||
|
ATBF1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 94) | NC score | 0.004233 (rank : 138) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 1547 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q15911, O15101, Q13719 | Gene names | ATBF1 | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-fetoprotein enhancer-binding protein (AT motif-binding factor) (AT-binding transcription factor 1). | |||||
|
AZI1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 95) | NC score | 0.009271 (rank : 134) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 1000 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UPN4, Q96F50 | Gene names | AZI1, CEP131, KIAA1118 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 5-azacytidine-induced protein 1 (Pre-acrosome localization protein 1) (Centrosomal protein of 131 kDa) (Cep131 protein). | |||||
|
BAT2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 96) | NC score | 0.025819 (rank : 109) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P48634, O95875, Q5SQ29, Q5SQ30, Q5ST84, Q5STX6, Q5STX7, Q68DW9, Q6P9P7, Q6PIN1, Q96QC6 | Gene names | BAT2, G2 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
FOXP4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 97) | NC score | 0.009161 (rank : 135) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9DBY0, Q80V92, Q8CG10, Q8CIS1 | Gene names | Foxp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Forkhead box protein P4 (Fork head-related protein like A) (mFKHLA). | |||||
|
GAK5_HUMAN
|
||||||
| θ value | 8.99809 (rank : 98) | NC score | 0.063620 (rank : 62) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 429 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P62684 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_19p13.11 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K113 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
IF140_HUMAN
|
||||||
| θ value | 8.99809 (rank : 99) | NC score | 0.016569 (rank : 126) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96RY7, O60332 | Gene names | IFT140, KIAA0590, WDTC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intraflagellar transport 140 homolog (WD and tetratricopeptide repeats protein 2). | |||||
|
MLL3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 100) | NC score | 0.022587 (rank : 113) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
NCOA6_HUMAN
|
||||||
| θ value | 8.99809 (rank : 101) | NC score | 0.040702 (rank : 97) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 1202 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q14686, Q9NTZ9, Q9UH74, Q9UK86 | Gene names | NCOA6, AIB3, KIAA0181, RAP250, TRBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (PRIP) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC RAP250) (Thyroid hormone receptor-binding protein). | |||||
|
RN126_HUMAN
|
||||||
| θ value | 8.99809 (rank : 102) | NC score | 0.150170 (rank : 33) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9BV68, Q9NWX1 | Gene names | RNF126 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 126. | |||||
|
SYN3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 103) | NC score | 0.043539 (rank : 89) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8JZP2 | Gene names | Syn3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synapsin-3 (Synapsin III). | |||||
|
TMF1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 104) | NC score | 0.004256 (rank : 137) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 987 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P82094 | Gene names | TMF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TATA element modulatory factor (TMF). | |||||
|
UBR2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 105) | NC score | 0.019606 (rank : 122) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8IWV8, O15057, Q4VXK2, Q5TFH6, Q6P2I2, Q6ZUD0 | Gene names | UBR2, C6orf133, KIAA0349 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase E3 component N-recognin-2 (EC 6.-.-.-) (Ubiquitin-protein ligase E3-alpha-2) (Ubiquitin-protein ligase E3- alpha-II). | |||||
|
UBR2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 106) | NC score | 0.019667 (rank : 121) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6WKZ8, Q6DIB9, Q80U31, Q8BUL9, Q8CGW0, Q8K2I6, Q8R0V7, Q8R130 | Gene names | Ubr2, Kiaa0349 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase E3 component N-recognin-2 (EC 6.-.-.-) (Ubiquitin-protein ligase E3-alpha-2) (Ubiquitin-protein ligase E3- alpha-II). | |||||
|
AMFR2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 107) | NC score | 0.082702 (rank : 43) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9UKV5, Q8IZ70 | Gene names | AMFR | |||
|
Domain Architecture |

|
|||||
| Description | Autocrine motility factor receptor, isoform 2 (EC 6.3.2.-) (AMF receptor) (gp78). | |||||
|
AMFR_MOUSE
|
||||||
| θ value | θ > 10 (rank : 108) | NC score | 0.083274 (rank : 42) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9R049, Q8K008, Q99LH5 | Gene names | Amfr | |||
|
Domain Architecture |

|
|||||
| Description | Autocrine motility factor receptor (EC 6.3.2.19) (AMF receptor). | |||||
|
APC11_HUMAN
|
||||||
| θ value | θ > 10 (rank : 109) | NC score | 0.051957 (rank : 85) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9NYG5, Q502X9, Q9BW64, Q9P0R2 | Gene names | ANAPC11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Anaphase-promoting complex subunit 11 (APC11) (Cyclosome subunit 11) (Hepatocellular carcinoma-associated RING finger protein). | |||||
|
APC11_MOUSE
|
||||||
| θ value | θ > 10 (rank : 110) | NC score | 0.053834 (rank : 79) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9CPX9, Q3UWH1, Q9CTG0 | Gene names | Anapc11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Anaphase-promoting complex subunit 11 (APC11) (Cyclosome subunit 11). | |||||
|
DZIP3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 111) | NC score | 0.076293 (rank : 48) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 399 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q86Y13, O75162, Q6P3R9, Q6PH82, Q86Y14, Q86Y15, Q86Y16, Q8IWI0, Q96RS9 | Gene names | DZIP3, KIAA0675 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin ligase protein DZIP3 (EC 6.3.2.-) (DAZ-interacting protein 3) (RNA-binding ubiquitin ligase of 138 kDa) (hRUL138). | |||||
|
DZIP3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 112) | NC score | 0.078305 (rank : 46) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 414 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q7TPV2, Q80TU4, Q8BYK7 | Gene names | Dzip3, Kiaa0675 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin ligase protein DZIP3 (EC 6.3.2.-) (DAZ-interacting protein 3 homolog). | |||||
|
GAK10_HUMAN
|
||||||
| θ value | θ > 10 (rank : 113) | NC score | 0.061103 (rank : 68) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P87889, P10263, P10264, Q69385, Q9UKH6 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_5q33.3 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K10 Gag protein) (HERV-K107 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK14_HUMAN
|
||||||
| θ value | θ > 10 (rank : 114) | NC score | 0.051131 (rank : 86) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P62687 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_16p3.3 provirus ancestral Gag polyprotein (Gag polyprotein). | |||||
|
GAK15_HUMAN
|
||||||
| θ value | θ > 10 (rank : 115) | NC score | 0.054929 (rank : 78) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P62688 | Gene names | ERVK4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_3q21.2 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K(I) Gag protein). | |||||
|
GAK17_HUMAN
|
||||||
| θ value | θ > 10 (rank : 116) | NC score | 0.055259 (rank : 76) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P62689 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_11q22.1 provirus ancestral Gag polyprotein (Gag polyprotein) [Contains: Matrix protein]. | |||||
|
GAK18_HUMAN
|
||||||
| θ value | θ > 10 (rank : 117) | NC score | 0.060195 (rank : 71) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P62690 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_22q11.23 provirus ancestral Gag polyprotein (Gag polyprotein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK19_HUMAN
|
||||||
| θ value | θ > 10 (rank : 118) | NC score | 0.052910 (rank : 81) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P62691 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_4p16.1 provirus ancestral Gag polyprotein (Gag polyprotein). | |||||
|
GAK7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 119) | NC score | 0.057234 (rank : 75) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q96PI4, Q9QC08 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_1q23.3 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K18 Gag protein) (HERV-K110 Gag protein) (HERV-K(C1a) Gag protein) [Contains: Matrix protein]. | |||||
|
GAK8_HUMAN
|
||||||
| θ value | θ > 10 (rank : 120) | NC score | 0.060068 (rank : 72) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9HDB9 | Gene names | ERVK5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_3q12.3 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K(II) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK9_HUMAN
|
||||||
| θ value | θ > 10 (rank : 121) | NC score | 0.055178 (rank : 77) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UKH8 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_5q13.3 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K104 Gag protein) [Contains: Matrix protein]. | |||||
|
PEX10_HUMAN
|
||||||
| θ value | θ > 10 (rank : 122) | NC score | 0.053011 (rank : 80) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O60683, Q5T095, Q9BW90 | Gene names | PEX10, RNF69 | |||
|
Domain Architecture |
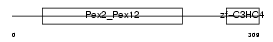
|
|||||
| Description | Peroxisome assembly protein 10 (Peroxin-10) (Peroxisome biogenesis factor 10) (RING finger protein 69). | |||||
|
PRB3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 123) | NC score | 0.060341 (rank : 69) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q04118, Q15188, Q7M4M9 | Gene names | PRB3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 3 precursor (Parotid salivary glycoprotein G1) (Proline-rich protein G1). | |||||
|
PRB4S_HUMAN
|
||||||
| θ value | θ > 10 (rank : 124) | NC score | 0.062639 (rank : 64) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P10163, P02813 | Gene names | PRB4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 4 allele S precursor (Salivary proline-rich protein Po) (Parotid o protein) [Contains: Protein N1; Glycosylated protein A]. | |||||
|
RBX1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 125) | NC score | 0.052455 (rank : 82) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P62877, Q8N6Z8, Q9D1S2, Q9WUK9, Q9Y254 | Gene names | RBX1, RNF75, ROC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING-box protein 1 (Rbx1) (Regulator of cullins 1) (RING finger protein 75) (Protein ZYP). | |||||
|
RBX1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 126) | NC score | 0.052455 (rank : 83) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P62878, Q8N6Z8, Q9D1S2, Q9WUK9, Q9Y254 | Gene names | Rbx1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING-box protein 1 (Rbx1). | |||||
|
RBX2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 127) | NC score | 0.062181 (rank : 67) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9UBF6, Q9BXN8, Q9Y5M7 | Gene names | RNF7, RBX2, ROC2, SAG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING-box protein 2 (Rbx2) (RING finger protein 7) (Regulator of cullins 2) (CKII beta-binding protein 1) (CKBBP1) (Sensitive to apoptosis gene protein). | |||||
|
RBX2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 128) | NC score | 0.062332 (rank : 66) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9WTZ1, Q3UKF8 | Gene names | Rnf7, Rbx2, Sag | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING-box protein 2 (Rbx2) (RING finger protein 7) (Sensitive to apoptosis gene protein). | |||||
|
RN141_HUMAN
|
||||||
| θ value | θ > 10 (rank : 129) | NC score | 0.078127 (rank : 47) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8WVD5, Q9NZB4 | Gene names | RNF141, ZNF230 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 141 (Zinc finger protein 230). | |||||
|
RN141_MOUSE
|
||||||
| θ value | θ > 10 (rank : 130) | NC score | 0.078981 (rank : 45) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q99MB7, Q9D040, Q9D5C4, Q9D9L0 | Gene names | Rnf141, Znf230 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 141 (Zinc finger protein 230). | |||||
|
RN175_HUMAN
|
||||||
| θ value | θ > 10 (rank : 131) | NC score | 0.062424 (rank : 65) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8N4F7 | Gene names | RNF175 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 175. | |||||
|
RNF32_HUMAN
|
||||||
| θ value | θ > 10 (rank : 132) | NC score | 0.065859 (rank : 61) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9H0A6, Q6FIB3, Q6X7T4, Q8N6V8, Q8TDG0, Q96BM5, Q9Y6U1 | Gene names | RNF32 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 32. | |||||
|
RNF32_MOUSE
|
||||||
| θ value | θ > 10 (rank : 133) | NC score | 0.063499 (rank : 63) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9JIT1, Q9D9X0, Q9DAJ3 | Gene names | Rnf32, Lmbr2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 32 (Limb region protein 2). | |||||
|
SYHH_HUMAN
|
||||||
| θ value | θ > 10 (rank : 134) | NC score | 0.071267 (rank : 53) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P49590 | Gene names | HARSL, HARSR, HO3 | |||
|
Domain Architecture |

|
|||||
| Description | Histidyl-tRNA synthetase homolog (EC 6.1.1.21) (Histidine--tRNA ligase homolog) (HisRS). | |||||
|
SYH_HUMAN
|
||||||
| θ value | θ > 10 (rank : 135) | NC score | 0.075292 (rank : 50) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P12081 | Gene names | HARS, HRS | |||
|
Domain Architecture |

|
|||||
| Description | Histidyl-tRNA synthetase (EC 6.1.1.21) (Histidine--tRNA ligase) (HisRS). | |||||
|
SYH_MOUSE
|
||||||
| θ value | θ > 10 (rank : 136) | NC score | 0.076228 (rank : 49) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q61035 | Gene names | Hars | |||
|
Domain Architecture |

|
|||||
| Description | Histidyl-tRNA synthetase (EC 6.1.1.21) (Histidine--tRNA ligase) (HisRS). | |||||
|
ZN363_HUMAN
|
||||||
| θ value | θ > 10 (rank : 137) | NC score | 0.072486 (rank : 52) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q96PM5, Q96PR5 | Gene names | RCHY1, ARNIP, CHIMP, PIRH2, ZNF363 | |||
|
Domain Architecture |
|
|||||
| Description | RING finger and CHY zinc finger domain-containing protein 1 (Zinc finger protein 363) (CH-rich-interacting match with PLAG1) (Androgen receptor N-terminal-interacting protein) (p53 induced RING-H2 protein) (hPirh2). | |||||
|
ZN363_MOUSE
|
||||||
| θ value | θ > 10 (rank : 138) | NC score | 0.075146 (rank : 51) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9CR50, Q920H0 | Gene names | Rchy1, Arnip, Chimp, Zfp363, Znf363 | |||
|
Domain Architecture |
|
|||||
| Description | RING finger and CHY zinc finger domain-containing protein 1 (Zinc finger protein 363) (CH-rich-interacting match with PLAG1) (Androgen receptor N-terminal-interacting protein). | |||||
|
RNF25_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 106 | |
| SwissProt Accessions | Q9QZR0, Q3TAL8, Q9DCW7 | Gene names | Rnf25 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 25 (EC 6.3.2.-) (RING finger protein AO7). | |||||
|
RNF25_HUMAN
|
||||||
| NC score | 0.949209 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q96BH1, Q9H874 | Gene names | RNF25 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 25 (EC 6.3.2.-). | |||||
|
RWDD1_HUMAN
|
||||||
| NC score | 0.349776 (rank : 3) | θ value | 4.1701e-05 (rank : 5) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9H446, Q9Y313, Q9Y6B3 | Gene names | RWDD1 | |||
|
Domain Architecture |

|
|||||
| Description | RWD domain-containing protein 1. | |||||
|
RWDD1_MOUSE
|
||||||
| NC score | 0.348046 (rank : 4) | θ value | 4.1701e-05 (rank : 6) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9CQK7 | Gene names | Rwdd1 | |||
|
Domain Architecture |

|
|||||
| Description | RWD domain-containing protein 1 (IH1). | |||||
|
RNF24_MOUSE
|
||||||
| NC score | 0.252604 (rank : 5) | θ value | 0.00665767 (rank : 7) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8BGI1 | Gene names | Rnf24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 24. | |||||
|
RNF24_HUMAN
|
||||||
| NC score | 0.241979 (rank : 6) | θ value | 0.0252991 (rank : 8) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9Y225, Q9UMH1 | Gene names | RNF24 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 24. | |||||
|
RN122_MOUSE
|
||||||
| NC score | 0.233368 (rank : 7) | θ value | 0.125558 (rank : 18) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8BP31, Q80VA7, Q8BGD3 | Gene names | Rnf122 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 122. | |||||
|
RN122_HUMAN
|
||||||
| NC score | 0.232746 (rank : 8) | θ value | 0.125558 (rank : 17) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9H9V4, Q52LK3 | Gene names | RNF122 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 122. | |||||
|
RNF12_MOUSE
|
||||||
| NC score | 0.221792 (rank : 9) | θ value | 0.0431538 (rank : 10) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9WTV7 | Gene names | Rnf12, Rlim | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 12 (LIM domain-interacting RING finger protein) (RING finger LIM domain-binding protein) (R-LIM). | |||||
|
RNF6_HUMAN
|
||||||
| NC score | 0.215397 (rank : 10) | θ value | 0.125558 (rank : 19) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9Y252, Q9UF41 | Gene names | RNF6 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 6 (RING-H2 protein). | |||||
|
RNF38_HUMAN
|
||||||
| NC score | 0.214384 (rank : 11) | θ value | 0.0736092 (rank : 14) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9H0F5, Q8N0Y0 | Gene names | RNF38 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 38. | |||||
|
PJA1_MOUSE
|
||||||
| NC score | 0.213813 (rank : 12) | θ value | 0.0563607 (rank : 12) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O55176, Q8CFU2, Q99MJ1, Q99MJ2, Q99MJ3, Q9DB04 | Gene names | Pja1 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin protein ligase Praja1 (EC 6.3.2.-). | |||||
|
RNF38_MOUSE
|
||||||
| NC score | 0.212701 (rank : 13) | θ value | 0.0961366 (rank : 16) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8BI21, Q6P5B1 | Gene names | Rnf38 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 38. | |||||
|
RNF12_HUMAN
|
||||||
| NC score | 0.209346 (rank : 14) | θ value | 0.163984 (rank : 20) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9NVW2, Q9Y598 | Gene names | RNF12, RLIM | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 12 (LIM domain-interacting RING finger protein) (RING finger LIM domain-binding protein) (R-LIM) (NY-REN-43 antigen). | |||||
|
PJA1_HUMAN
|
||||||
| NC score | 0.209013 (rank : 15) | θ value | 0.0563607 (rank : 11) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8NG27, Q8NG28, Q9HAC1 | Gene names | PJA1, RNF70 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin protein ligase Praja1 (EC 6.3.2.-) (RING finger protein 70). | |||||
|
RNF11_HUMAN
|
||||||
| NC score | 0.199784 (rank : 16) | θ value | 1.06291 (rank : 42) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9Y3C5 | Gene names | RNF11 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 11 (Sid 1669). | |||||
|
RNF11_MOUSE
|
||||||
| NC score | 0.199204 (rank : 17) | θ value | 1.06291 (rank : 43) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9QYK7, Q9EQI1 | Gene names | Rnf11, N4wbp2, Sid1669 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 11 (NEDD4 WW domain-binding protein 2) (Sid 1669). | |||||
|
ZN364_HUMAN
|
||||||
| NC score | 0.195464 (rank : 18) | θ value | 0.21417 (rank : 23) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9Y4L5, Q5T2V9, Q7Z2J2 | Gene names | ZNF364, RNF115 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 364 (Rabring 7) (RING finger protein 115). | |||||
|
RN165_HUMAN
|
||||||
| NC score | 0.194199 (rank : 19) | θ value | 0.47712 (rank : 30) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q6ZSG1 | Gene names | RNF165 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 165. | |||||
|
ZN364_MOUSE
|
||||||
| NC score | 0.192532 (rank : 20) | θ value | 0.279714 (rank : 25) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9D0C1, Q8R5A1, Q9D885 | Gene names | Znf364, Zfp364 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 364 (Rabring 7). | |||||
|
RNF13_MOUSE
|
||||||
| NC score | 0.189828 (rank : 21) | θ value | 0.365318 (rank : 28) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 164 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O54965, O54966 | Gene names | Rnf13, Rzf | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 13. | |||||
|
GOLI_MOUSE
|
||||||
| NC score | 0.189616 (rank : 22) | θ value | 0.47712 (rank : 29) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8VEM1, Q80VL7, Q9QZQ6 | Gene names | Rnf130, G1rp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Goliath homolog precursor (RING finger protein 130) (G1-related zinc finger protein). | |||||
|
GOLI_HUMAN
|
||||||
| NC score | 0.187060 (rank : 23) | θ value | 0.62314 (rank : 36) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q86XS8, Q641P9, Q6P177, Q7L2U2, Q9P0J9 | Gene names | RNF130 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Goliath homolog precursor (RING finger protein 130). | |||||
|
RNF13_HUMAN
|
||||||
| NC score | 0.186276 (rank : 24) | θ value | 0.47712 (rank : 31) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O43567 | Gene names | RNF13, RZF | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 13. | |||||
|
RN167_HUMAN
|
||||||
| NC score | 0.172362 (rank : 25) | θ value | 4.03905 (rank : 77) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9H6Y7, Q6XYE0, Q8NDC1, Q9Y3V1 | Gene names | RNF167 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 167 precursor. | |||||
|
RN133_HUMAN
|
||||||
| NC score | 0.169777 (rank : 26) | θ value | 5.27518 (rank : 83) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q8WVZ7, Q8N7G7 | Gene names | RNF133 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 133. | |||||
|
ZSWM2_HUMAN
|
||||||
| NC score | 0.167603 (rank : 27) | θ value | 0.21417 (rank : 24) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8NEG5 | Gene names | ZSWIM2 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger SWIM domain-containing protein 2. | |||||
|
RN167_MOUSE
|
||||||
| NC score | 0.167099 (rank : 28) | θ value | 4.03905 (rank : 78) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q91XF4, Q3U4S5 | Gene names | Rnf167 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 167 precursor. | |||||
|
RN126_MOUSE
|
||||||
| NC score | 0.164856 (rank : 29) | θ value | 3.0926 (rank : 71) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q91YL2 | Gene names | Rnf126 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 126. | |||||
|
RN139_HUMAN
|
||||||
| NC score | 0.164270 (rank : 30) | θ value | 0.62314 (rank : 37) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8WU17, O75485, Q7LDL3 | Gene names | RNF139, TRC8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 139 (Translocation in renal carcinoma on chromosome 8). | |||||
|
RN139_MOUSE
|
||||||
| NC score | 0.157628 (rank : 31) | θ value | 1.06291 (rank : 41) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q7TMV1, Q8BZU9 | Gene names | Rnf139 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 139. | |||||
|
ZSWM2_MOUSE
|
||||||
| NC score | 0.153074 (rank : 32) | θ value | 2.36792 (rank : 66) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9D9X6 | Gene names | Zswim2 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger SWIM domain-containing protein 2. | |||||
|
RN126_HUMAN
|
||||||
| NC score | 0.150170 (rank : 33) | θ value | 8.99809 (rank : 102) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9BV68, Q9NWX1 | Gene names | RNF126 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 126. | |||||
|
RN103_MOUSE
|
||||||
| NC score | 0.149324 (rank : 34) | θ value | 1.38821 (rank : 50) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9R1W3, O08670, O08883 | Gene names | Rnf103, Zfp103 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 103 (Zinc finger protein 103) (Zfp-103) (KF-1) (mKF-1). | |||||
|
RN103_HUMAN
|
||||||
| NC score | 0.148278 (rank : 35) | θ value | 1.38821 (rank : 49) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O00237, Q8IVB9 | Gene names | RNF103 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 103 (Zinc finger protein 103 homolog) (Zfp-103) (KF-1) (hKF-1). | |||||
|
CU006_MOUSE
|
||||||
| NC score | 0.116622 (rank : 36) | θ value | 0.0961366 (rank : 15) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q99M03, Q9JLH4 | Gene names | ORF5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C21orf6 homolog. | |||||
|
E2AK4_MOUSE
|
||||||
| NC score | 0.116055 (rank : 37) | θ value | 9.59137e-10 (rank : 3) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 984 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9QZ05, Q6GQT4, Q6ZPT5, Q8C5S0, Q8CIF5, Q9CT30, Q9CUV9, Q9ESB6, Q9ESB7, Q9ESB8 | Gene names | Eif2ak4, Gcn2, Kiaa1338 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 2-alpha kinase 4 (EC 2.7.11.1) (GCN2-like protein) (mGCN2). | |||||
|
E2AK4_HUMAN
|
||||||
| NC score | 0.109564 (rank : 38) | θ value | 1.06045e-08 (rank : 4) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 1012 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9P2K8, Q69YL7, Q6DC97, Q96GN6, Q9H5K1, Q9NSQ3, Q9NSZ5, Q9UJ56 | Gene names | EIF2AK4, GCN2, KIAA1338 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 2-alpha kinase 4 (EC 2.7.11.1) (GCN2-like protein). | |||||
|
CBP_HUMAN
|
||||||
| NC score | 0.088749 (rank : 39) | θ value | 0.0736092 (rank : 13) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 941 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q92793, O00147, Q16376 | Gene names | CREBBP, CBP | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
EP300_HUMAN
|
||||||
| NC score | 0.087362 (rank : 40) | θ value | 1.06291 (rank : 40) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q09472 | Gene names | EP300, P300 | |||
|
Domain Architecture |

|
|||||
| Description | E1A-associated protein p300 (EC 2.3.1.48). | |||||
|
CU006_HUMAN
|
||||||
| NC score | 0.086148 (rank : 41) | θ value | 0.21417 (rank : 21) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P57060 | Gene names | C21orf6 | |||
|
Domain Architecture |

|
|||||
| Description | Protein C21orf6. | |||||
|
AMFR_MOUSE
|
||||||
| NC score | 0.083274 (rank : 42) | θ value | θ > 10 (rank : 108) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9R049, Q8K008, Q99LH5 | Gene names | Amfr | |||
|
Domain Architecture |

|
|||||
| Description | Autocrine motility factor receptor (EC 6.3.2.19) (AMF receptor). | |||||
|
AMFR2_HUMAN
|
||||||
| NC score | 0.082702 (rank : 43) | θ value | θ > 10 (rank : 107) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9UKV5, Q8IZ70 | Gene names | AMFR | |||
|
Domain Architecture |

|
|||||
| Description | Autocrine motility factor receptor, isoform 2 (EC 6.3.2.-) (AMF receptor) (gp78). | |||||
|
CBP_MOUSE
|
||||||
| NC score | 0.079819 (rank : 44) | θ value | 0.0431538 (rank : 9) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 1051 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P45481 | Gene names | Crebbp, Cbp | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
RN141_MOUSE
|
||||||
| NC score | 0.078981 (rank : 45) | θ value | θ > 10 (rank : 130) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q99MB7, Q9D040, Q9D5C4, Q9D9L0 | Gene names | Rnf141, Znf230 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 141 (Zinc finger protein 230). | |||||
|
DZIP3_MOUSE
|
||||||
| NC score | 0.078305 (rank : 46) | θ value | θ > 10 (rank : 112) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 414 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q7TPV2, Q80TU4, Q8BYK7 | Gene names | Dzip3, Kiaa0675 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin ligase protein DZIP3 (EC 6.3.2.-) (DAZ-interacting protein 3 homolog). | |||||
|
RN141_HUMAN
|
||||||
| NC score | 0.078127 (rank : 47) | θ value | θ > 10 (rank : 129) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8WVD5, Q9NZB4 | Gene names | RNF141, ZNF230 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 141 (Zinc finger protein 230). | |||||
|
DZIP3_HUMAN
|
||||||
| NC score | 0.076293 (rank : 48) | θ value | θ > 10 (rank : 111) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 399 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q86Y13, O75162, Q6P3R9, Q6PH82, Q86Y14, Q86Y15, Q86Y16, Q8IWI0, Q96RS9 | Gene names | DZIP3, KIAA0675 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin ligase protein DZIP3 (EC 6.3.2.-) (DAZ-interacting protein 3) (RNA-binding ubiquitin ligase of 138 kDa) (hRUL138). | |||||
|
SYH_MOUSE
|
||||||
| NC score | 0.076228 (rank : 49) | θ value | θ > 10 (rank : 136) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q61035 | Gene names | Hars | |||
|
Domain Architecture |

|
|||||
| Description | Histidyl-tRNA synthetase (EC 6.1.1.21) (Histidine--tRNA ligase) (HisRS). | |||||
|
SYH_HUMAN
|
||||||
| NC score | 0.075292 (rank : 50) | θ value | θ > 10 (rank : 135) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P12081 | Gene names | HARS, HRS | |||
|
Domain Architecture |

|
|||||
| Description | Histidyl-tRNA synthetase (EC 6.1.1.21) (Histidine--tRNA ligase) (HisRS). | |||||
|
ZN363_MOUSE
|
||||||
| NC score | 0.075146 (rank : 51) | θ value | θ > 10 (rank : 138) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9CR50, Q920H0 | Gene names | Rchy1, Arnip, Chimp, Zfp363, Znf363 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger and CHY zinc finger domain-containing protein 1 (Zinc finger protein 363) (CH-rich-interacting match with PLAG1) (Androgen receptor N-terminal-interacting protein). | |||||
|
ZN363_HUMAN
|
||||||
| NC score | 0.072486 (rank : 52) | θ value | θ > 10 (rank : 137) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q96PM5, Q96PR5 | Gene names | RCHY1, ARNIP, CHIMP, PIRH2, ZNF363 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger and CHY zinc finger domain-containing protein 1 (Zinc finger protein 363) (CH-rich-interacting match with PLAG1) (Androgen receptor N-terminal-interacting protein) (p53 induced RING-H2 protein) (hPirh2). | |||||
|
SYHH_HUMAN
|
||||||
| NC score | 0.071267 (rank : 53) | θ value | θ > 10 (rank : 134) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P49590 | Gene names | HARSL, HARSR, HO3 | |||
|
Domain Architecture |

|
|||||
| Description | Histidyl-tRNA synthetase homolog (EC 6.1.1.21) (Histidine--tRNA ligase homolog) (HisRS). | |||||
|
GAK12_HUMAN
|
||||||
| NC score | 0.070155 (rank : 54) | θ value | 0.62314 (rank : 34) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P63130, Q9UKI0 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_1q22 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K102 Gag protein) (HERV-K(III) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK4_HUMAN
|
||||||
| NC score | 0.069641 (rank : 55) | θ value | 0.62314 (rank : 35) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 415 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P63126, Q9UKH4 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_6q14.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K109 Gag protein) (HERV-K(C6) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK11_HUMAN
|
||||||
| NC score | 0.069617 (rank : 56) | θ value | 0.62314 (rank : 33) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P63145, Q9UKI1 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_22q11.21 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K101 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK1_HUMAN
|
||||||
| NC score | 0.069235 (rank : 57) | θ value | 0.813845 (rank : 38) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 268 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P62683 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_12q14.1 provirus ancestral Gag polyprotein (Gag polyprotein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK6_HUMAN
|
||||||
| NC score | 0.067994 (rank : 58) | θ value | 1.81305 (rank : 54) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P62685 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_8p23.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K115 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK3_HUMAN
|
||||||
| NC score | 0.067607 (rank : 59) | θ value | 1.81305 (rank : 53) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 182 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9YNA8 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_19q12 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K(C19) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK2_HUMAN
|
||||||
| NC score | 0.067443 (rank : 60) | θ value | 1.81305 (rank : 52) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q7LDI9, Q9UKH5, Q9Y6I1, Q9YNA6, Q9YNB0 | Gene names | ERVK6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_7p22.1 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K(HML-2.HOM) Gag protein) (HERV-K108 Gag protein) (HERV-K(C7) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
RNF32_HUMAN
|
||||||
| NC score | 0.065859 (rank : 61) | θ value | θ > 10 (rank : 132) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9H0A6, Q6FIB3, Q6X7T4, Q8N6V8, Q8TDG0, Q96BM5, Q9Y6U1 | Gene names | RNF32 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 32. | |||||
|
GAK5_HUMAN
|
||||||
| NC score | 0.063620 (rank : 62) | θ value | 8.99809 (rank : 98) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 429 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P62684 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_19p13.11 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K113 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
RNF32_MOUSE
|
||||||
| NC score | 0.063499 (rank : 63) | θ value | θ > 10 (rank : 133) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9JIT1, Q9D9X0, Q9DAJ3 | Gene names | Rnf32, Lmbr2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 32 (Limb region protein 2). | |||||
|
PRB4S_HUMAN
|
||||||
| NC score | 0.062639 (rank : 64) | θ value | θ > 10 (rank : 124) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P10163, P02813 | Gene names | PRB4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 4 allele S precursor (Salivary proline-rich protein Po) (Parotid o protein) [Contains: Protein N1; Glycosylated protein A]. | |||||
|
RN175_HUMAN
|
||||||
| NC score | 0.062424 (rank : 65) | θ value | θ > 10 (rank : 131) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8N4F7 | Gene names | RNF175 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 175. | |||||
|
RBX2_MOUSE
|
||||||
| NC score | 0.062332 (rank : 66) | θ value | θ > 10 (rank : 128) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9WTZ1, Q3UKF8 | Gene names | Rnf7, Rbx2, Sag | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING-box protein 2 (Rbx2) (RING finger protein 7) (Sensitive to apoptosis gene protein). | |||||
|
RBX2_HUMAN
|
||||||
| NC score | 0.062181 (rank : 67) | θ value | θ > 10 (rank : 127) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9UBF6, Q9BXN8, Q9Y5M7 | Gene names | RNF7, RBX2, ROC2, SAG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING-box protein 2 (Rbx2) (RING finger protein 7) (Regulator of cullins 2) (CKII beta-binding protein 1) (CKBBP1) (Sensitive to apoptosis gene protein). | |||||
|
GAK10_HUMAN
|
||||||
| NC score | 0.061103 (rank : 68) | θ value | θ > 10 (rank : 113) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P87889, P10263, P10264, Q69385, Q9UKH6 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_5q33.3 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K10 Gag protein) (HERV-K107 Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
PRB3_HUMAN
|
||||||
| NC score | 0.060341 (rank : 69) | θ value | θ > 10 (rank : 123) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q04118, Q15188, Q7M4M9 | Gene names | PRB3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 3 precursor (Parotid salivary glycoprotein G1) (Proline-rich protein G1). | |||||
|
MAML2_HUMAN
|
||||||
| NC score | 0.060315 (rank : 70) | θ value | 6.88961 (rank : 90) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8IZL2, Q6AI23, Q6Y3A3, Q8IUL3, Q96JK6 | Gene names | MAML2, KIAA1819 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mastermind-like protein 2 (Mam-2). | |||||
|
GAK18_HUMAN
|
||||||
| NC score | 0.060195 (rank : 71) | θ value | θ > 10 (rank : 117) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P62690 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_22q11.23 provirus ancestral Gag polyprotein (Gag polyprotein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
GAK8_HUMAN
|
||||||
| NC score | 0.060068 (rank : 72) | θ value | θ > 10 (rank : 120) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9HDB9 | Gene names | ERVK5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_3q12.3 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K(II) Gag protein) [Contains: Matrix protein; Capsid protein; Nucleocapsid protein]. | |||||
|
YLPM1_HUMAN
|
||||||
| NC score | 0.059599 (rank : 73) | θ value | 0.813845 (rank : 39) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P49750, P49752, Q96I64, Q9P1V7 | Gene names | YLPM1, C14orf170, ZAP3 | |||
|
Domain Architecture |

|
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3) (ZAP113). | |||||
|
TRML2_MOUSE
|
||||||
| NC score | 0.058038 (rank : 74) | θ value | 0.47712 (rank : 32) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q2LA85 | Gene names | Treml2, Tlt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trem-like transcript 2 protein precursor (TLT-2) (Triggering receptor expressed on myeloid cells-like protein 2). | |||||
|
GAK7_HUMAN
|
||||||
| NC score | 0.057234 (rank : 75) | θ value | θ > 10 (rank : 119) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q96PI4, Q9QC08 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_1q23.3 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K18 Gag protein) (HERV-K110 Gag protein) (HERV-K(C1a) Gag protein) [Contains: Matrix protein]. | |||||
|
GAK17_HUMAN
|
||||||
| NC score | 0.055259 (rank : 76) | θ value | θ > 10 (rank : 116) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P62689 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_11q22.1 provirus ancestral Gag polyprotein (Gag polyprotein) [Contains: Matrix protein]. | |||||
|
GAK9_HUMAN
|
||||||
| NC score | 0.055178 (rank : 77) | θ value | θ > 10 (rank : 121) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UKH8 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_5q13.3 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K104 Gag protein) [Contains: Matrix protein]. | |||||
|
GAK15_HUMAN
|
||||||
| NC score | 0.054929 (rank : 78) | θ value | θ > 10 (rank : 115) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P62688 | Gene names | ERVK4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_3q21.2 provirus ancestral Gag polyprotein (Gag polyprotein) (HERV-K(I) Gag protein). | |||||
|
APC11_MOUSE
|
||||||
| NC score | 0.053834 (rank : 79) | θ value | θ > 10 (rank : 110) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9CPX9, Q3UWH1, Q9CTG0 | Gene names | Anapc11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Anaphase-promoting complex subunit 11 (APC11) (Cyclosome subunit 11). | |||||
|
PEX10_HUMAN
|
||||||
| NC score | 0.053011 (rank : 80) | θ value | θ > 10 (rank : 122) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O60683, Q5T095, Q9BW90 | Gene names | PEX10, RNF69 | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisome assembly protein 10 (Peroxin-10) (Peroxisome biogenesis factor 10) (RING finger protein 69). | |||||
|
GAK19_HUMAN
|
||||||
| NC score | 0.052910 (rank : 81) | θ value | θ > 10 (rank : 118) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P62691 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_4p16.1 provirus ancestral Gag polyprotein (Gag polyprotein). | |||||
|
RBX1_HUMAN
|
||||||
| NC score | 0.052455 (rank : 82) | θ value | θ > 10 (rank : 125) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P62877, Q8N6Z8, Q9D1S2, Q9WUK9, Q9Y254 | Gene names | RBX1, RNF75, ROC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING-box protein 1 (Rbx1) (Regulator of cullins 1) (RING finger protein 75) (Protein ZYP). | |||||
|
RBX1_MOUSE
|
||||||
| NC score | 0.052455 (rank : 83) | θ value | θ > 10 (rank : 126) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P62878, Q8N6Z8, Q9D1S2, Q9WUK9, Q9Y254 | Gene names | Rbx1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING-box protein 1 (Rbx1). | |||||
|
SYN3_HUMAN
|
||||||
| NC score | 0.051999 (rank : 84) | θ value | 1.81305 (rank : 56) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 203 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O14994 | Gene names | SYN3 | |||
|
Domain Architecture |

|
|||||
| Description | Synapsin-3 (Synapsin III). | |||||
|
APC11_HUMAN
|
||||||
| NC score | 0.051957 (rank : 85) | θ value | θ > 10 (rank : 109) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9NYG5, Q502X9, Q9BW64, Q9P0R2 | Gene names | ANAPC11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Anaphase-promoting complex subunit 11 (APC11) (Cyclosome subunit 11) (Hepatocellular carcinoma-associated RING finger protein). | |||||
|
GAK14_HUMAN
|
||||||
| NC score | 0.051131 (rank : 86) | θ value | θ > 10 (rank : 114) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P62687 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HERV-K_16p3.3 provirus ancestral Gag polyprotein (Gag polyprotein). | |||||
|
SYN2_MOUSE
|
||||||
| NC score | 0.050809 (rank : 87) | θ value | 1.06291 (rank : 44) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q64332, Q6NZR0, Q9QWV7 | Gene names | Syn2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synapsin-2 (Synapsin II). | |||||
|
NCOA6_MOUSE
|
||||||
| NC score | 0.047705 (rank : 88) | θ value | 1.38821 (rank : 47) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 1076 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9JL19, Q9JLT9 | Gene names | Ncoa6, Aib3, Prip, Rap250, Trbp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC) (Thyroid hormone receptor-binding protein). | |||||
|
SYN3_MOUSE
|
||||||
| NC score | 0.043539 (rank : 89) | θ value | 8.99809 (rank : 103) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8JZP2 | Gene names | Syn3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synapsin-3 (Synapsin III). | |||||
|
PERQ1_HUMAN
|
||||||
| NC score | 0.043486 (rank : 90) | θ value | 1.38821 (rank : 48) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O75420 | Gene names | PERQ1 | |||
|
Domain Architecture |

|
|||||
| Description | PERQ amino acid rich with GYF domain protein 1. | |||||
|
FHOD1_HUMAN
|
||||||
| NC score | 0.043378 (rank : 91) | θ value | 0.365318 (rank : 27) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Y613, Q59F76, Q6Y1F2, Q76MS8, Q8N521 | Gene names | FHOD1, FHOS | |||
|
Domain Architecture |

|
|||||
| Description | FH1/FH2 domain-containing protein (Formin homolog overexpressed in spleen) (FHOS) (Formin homology 2 domain-containing protein 1). | |||||
|
CRF_HUMAN
|
||||||
| NC score | 0.042776 (rank : 92) | θ value | 2.36792 (rank : 59) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P06850 | Gene names | CRH | |||
|
Domain Architecture |

|
|||||
| Description | Corticoliberin precursor (Corticotropin-releasing factor) (CRF) (Corticotropin-releasing hormone). | |||||
|
SYN2_HUMAN
|
||||||
| NC score | 0.042415 (rank : 93) | θ value | 5.27518 (rank : 85) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q92777 | Gene names | SYN2 | |||
|
Domain Architecture |

|
|||||
| Description | Synapsin-2 (Synapsin II). | |||||
|
BSN_MOUSE
|
||||||
| NC score | 0.042326 (rank : 94) | θ value | 1.38821 (rank : 45) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
RWDD3_HUMAN
|
||||||
| NC score | 0.042258 (rank : 95) | θ value | 6.88961 (rank : 93) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Y3V2, Q6FID3, Q9BX35 | Gene names | RWDD3 | |||
|
Domain Architecture |

|
|||||
| Description | RWD domain-containing protein 3. | |||||
|
CN004_HUMAN
|
||||||
| NC score | 0.041630 (rank : 96) | θ value | 2.36792 (rank : 58) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9H1B7, Q8NDQ2, Q96JG2, Q9H3I7 | Gene names | C14orf4, KIAA1865 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf4. | |||||
|
NCOA6_HUMAN
|
||||||
| NC score | 0.040702 (rank : 97) | θ value | 8.99809 (rank : 101) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 1202 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q14686, Q9NTZ9, Q9UH74, Q9UK86 | Gene names | NCOA6, AIB3, KIAA0181, RAP250, TRBP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor coactivator 6 (Amplified in breast cancer protein 3) (Cancer-amplified transcriptional coactivator ASC-2) (Activating signal cointegrator 2) (ASC-2) (Peroxisome proliferator-activated receptor-interacting protein) (PPAR-interacting protein) (PRIP) (Nuclear receptor-activating protein, 250 kDa) (Nuclear receptor coactivator RAP250) (NRC RAP250) (Thyroid hormone receptor-binding protein). | |||||
|
NADAP_HUMAN
|
||||||
| NC score | 0.038946 (rank : 98) | θ value | 2.36792 (rank : 62) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9BWU0, Q4KMT1, Q4KMX0, Q7Z5Q9, Q9NVN2 | Gene names | SLC4A1AP, HLC3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kanadaptin (Kidney anion exchanger adapter protein) (Solute carrier family 4 anion exchanger member 1 adapter protein) (Lung cancer oncogene 3 protein). | |||||
|
CCDC4_HUMAN
|
||||||
| NC score | 0.037989 (rank : 99) | θ value | 4.03905 (rank : 73) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6ZU67, Q58A26, Q58A27 | Gene names | CCDC4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 4. | |||||
|
SYQ_HUMAN
|
||||||
| NC score | 0.037118 (rank : 100) | θ value | 1.81305 (rank : 57) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P47897 | Gene names | QARS | |||
|
Domain Architecture |

|
|||||
| Description | Glutaminyl-tRNA synthetase (EC 6.1.1.18) (Glutamine--tRNA ligase) (GlnRS). | |||||
|
MLL2_HUMAN
|
||||||
| NC score | 0.033652 (rank : 101) | θ value | 5.27518 (rank : 82) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
HNRL1_MOUSE
|
||||||
| NC score | 0.033351 (rank : 102) | θ value | 1.38821 (rank : 46) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 200 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8VDM6, Q3U201, Q3UPB0, Q6AZA7, Q8BY45, Q8K365 | Gene names | Hnrpul1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heterogeneous nuclear ribonucleoprotein U-like protein 1. | |||||
|
PRG4_HUMAN
|
||||||
| NC score | 0.029818 (rank : 103) | θ value | 1.81305 (rank : 55) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q92954, Q6DNC4, Q6DNC5, Q6ZMZ5, Q9BX49 | Gene names | PRG4, MSF, SZP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.029508 (rank : 104) | θ value | 4.03905 (rank : 79) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
TFAP4_HUMAN
|
||||||
| NC score | 0.028156 (rank : 105) | θ value | 2.36792 (rank : 65) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q01664, O60409 | Gene names | TFAP4 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor AP-4 (Activating enhancer-binding protein 4). | |||||
|
DYN3_HUMAN
|
||||||
| NC score | 0.026505 (rank : 106) | θ value | 3.0926 (rank : 67) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UQ16, O14982, O95555, Q5W129, Q9H0P3, Q9H548, Q9NQ68, Q9NQN6 | Gene names | DNM3, KIAA0820 | |||
|
Domain Architecture |

|
|||||
| Description | Dynamin-3 (EC 3.6.5.5) (Dynamin, testicular) (T-dynamin). | |||||
|
MAT1_MOUSE
|
||||||
| NC score | 0.025951 (rank : 107) | θ value | 3.0926 (rank : 70) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P51949, Q3TZP0, Q9D8A0, Q9D8D2 | Gene names | Mnat1, Mat1 | |||
|
Domain Architecture |

|
|||||
| Description | CDK-activating kinase assembly factor MAT1 (RING finger protein MAT1) (Menage a trois) (CDK7/cyclin H assembly factor) (p36) (p35). | |||||
|
PRG4_MOUSE
|
||||||
| NC score | 0.025890 (rank : 108) | θ value | 2.36792 (rank : 63) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 624 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9JM99, Q3UEL1, Q3V198 | Gene names | Prg4, Msf, Szp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
BAT2_HUMAN
|
||||||
| NC score | 0.025819 (rank : 109) | θ value | 8.99809 (rank : 96) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P48634, O95875, Q5SQ29, Q5SQ30, Q5ST84, Q5STX6, Q5STX7, Q68DW9, Q6P9P7, Q6PIN1, Q96QC6 | Gene names | BAT2, G2 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
RFC1_MOUSE
|
||||||
| NC score | 0.024884 (rank : 110) | θ value | 2.36792 (rank : 64) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P35601 | Gene names | Rfc1, Ibf-1, Recc1 | |||
|
Domain Architecture |

|
|||||
| Description | Replication factor C subunit 1 (Replication factor C large subunit) (RF-C 140 kDa subunit) (Activator 1 140 kDa subunit) (Activator 1 large subunit) (A1 140 kDa subunit) (A1-P145) (Differentiation- specific element-binding protein) (ISRE-binding protein). | |||||
|
CO4A1_MOUSE
|
||||||
| NC score | 0.024673 (rank : 111) | θ value | 0.365318 (rank : 26) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 450 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P02463 | Gene names | Col4a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(IV) chain precursor. | |||||
|
CCD1A_MOUSE
|
||||||
| NC score | 0.024651 (rank : 112) | θ value | 4.03905 (rank : 72) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8K1A6, Q2MJB5, Q8R3Z4 | Gene names | Cc2d1a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil and C2 domain-containing protein 1A (Five repressor element under dual repression-binding protein 1) (FRE under dual repression-binding protein 1) (Freud-1). | |||||
|
MLL3_MOUSE
|
||||||
| NC score | 0.022587 (rank : 113) | θ value | 8.99809 (rank : 100) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
CO4A_HUMAN
|
||||||
| NC score | 0.021172 (rank : 114) | θ value | 4.03905 (rank : 74) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P0C0L4, P01028, P78445, Q13160, Q13906, Q14033, Q14835, Q5JQM8, Q9NPK5, Q9UIP5 | Gene names | C4A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Complement C4-A precursor (Acidic complement C4) [Contains: Complement C4 beta chain; Complement C4-A alpha chain; C4a anaphylatoxin; C4b-A; C4d-A; Complement C4 gamma chain]. | |||||
|
CO4B_HUMAN
|
||||||
| NC score | 0.021161 (rank : 115) | θ value | 4.03905 (rank : 75) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P0C0L5, P01028, P78445, Q13160, Q13906, Q14033, Q14835, Q9NPK5, Q9UIP5 | Gene names | C4B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Complement C4-B precursor (Basic complement C4) [Contains: Complement C4 beta chain; Complement C4-B alpha chain; C4a anaphylatoxin; C4b-B; C4d-B; Complement C4 gamma chain]. | |||||
|
RP1L1_HUMAN
|
||||||
| NC score | 0.020596 (rank : 116) | θ value | 6.88961 (rank : 92) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 1072 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8IWN7, Q86SQ1, Q8IWN8, Q8IWN9, Q8IWP0, Q8IWP1, Q8IWP2 | Gene names | RP1L1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinitis pigmentosa 1-like 1 protein. | |||||
|
NFAT5_HUMAN
|
||||||
| NC score | 0.020165 (rank : 117) | θ value | 6.88961 (rank : 91) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O94916, O95693, Q9UN18 | Gene names | NFAT5, KIAA0827, TONEBP | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor of activated T-cells 5 (T-cell transcription factor NFAT5) (NF-AT5) (Tonicity-responsive enhancer-binding protein) (TonE- binding protein) (TonEBP). | |||||
|
HCN4_HUMAN
|
||||||
| NC score | 0.020149 (rank : 118) | θ value | 3.0926 (rank : 68) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9Y3Q4, Q9UMQ7 | Gene names | HCN4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 4. | |||||
|
BAZ2A_HUMAN
|
||||||
| NC score | 0.020060 (rank : 119) | θ value | 5.27518 (rank : 80) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 780 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9UIF9, O00536, O15030, Q96H26 | Gene names | BAZ2A, KIAA0314, TIP5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5) (hWALp3). | |||||
|
IF4G3_HUMAN
|
||||||
| NC score | 0.019728 (rank : 120) | θ value | 2.36792 (rank : 61) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O43432, Q15597, Q8NEN1 | Gene names | EIF4G3 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4 gamma 3 (eIF-4-gamma 3) (eIF-4G 3) (eIF4G 3) (eIF-4-gamma II) (eIF4GII). | |||||
|
UBR2_MOUSE
|
||||||
| NC score | 0.019667 (rank : 121) | θ value | 8.99809 (rank : 106) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6WKZ8, Q6DIB9, Q80U31, Q8BUL9, Q8CGW0, Q8K2I6, Q8R0V7, Q8R130 | Gene names | Ubr2, Kiaa0349 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase E3 component N-recognin-2 (EC 6.-.-.-) (Ubiquitin-protein ligase E3-alpha-2) (Ubiquitin-protein ligase E3- alpha-II). | |||||
|
UBR2_HUMAN
|
||||||
| NC score | 0.019606 (rank : 122) | θ value | 8.99809 (rank : 105) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8IWV8, O15057, Q4VXK2, Q5TFH6, Q6P2I2, Q6ZUD0 | Gene names | UBR2, C6orf133, KIAA0349 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase E3 component N-recognin-2 (EC 6.-.-.-) (Ubiquitin-protein ligase E3-alpha-2) (Ubiquitin-protein ligase E3- alpha-II). | |||||
|
LR37B_HUMAN
|
||||||
| NC score | 0.018210 (rank : 123) | θ value | 3.0926 (rank : 69) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q96QE4, Q5YKG6 | Gene names | LRRC37B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 37B precursor (C66 SLIT-like testicular protein). | |||||
|
DJBP_MOUSE
|
||||||
| NC score | 0.017986 (rank : 124) | θ value | 5.27518 (rank : 81) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6P1E8, Q5DTV8, Q6PGK8, Q9D2D8 | Gene names | Djbp, Kiaa1672 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DJ-1-binding protein. | |||||
|
TXLNB_MOUSE
|
||||||
| NC score | 0.017265 (rank : 125) | θ value | 1.38821 (rank : 51) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 640 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8VBT1, Q3UVB8, Q8BUK2 | Gene names | Txlnb, Mdp77 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-taxilin (Muscle-derived protein 77). | |||||
|
IF140_HUMAN
|
||||||
| NC score | 0.016569 (rank : 126) | θ value | 8.99809 (rank : 99) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96RY7, O60332 | Gene names | IFT140, KIAA0590, WDTC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Intraflagellar transport 140 homolog (WD and tetratricopeptide repeats protein 2). | |||||
|
SIRT1_MOUSE
|
||||||
| NC score | 0.016137 (rank : 127) | θ value | 5.27518 (rank : 84) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q923E4, Q9QXG8 | Gene names | Sirt1, Sir2l1 | |||
|
Domain Architecture |

|
|||||
| Description | NAD-dependent deacetylase sirtuin-1 (EC 3.5.1.-) (SIR2alpha) (mSIR2a) (Sir2) (SIR2-like protein 1). | |||||
|
NEK4_MOUSE
|
||||||
| NC score | 0.013578 (rank : 128) | θ value | 0.21417 (rank : 22) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 899 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Z1J2, O35673, Q9R1J1 | Gene names | Nek4, Stk2 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase Nek4 (EC 2.7.11.1) (NimA-related protein kinase 4) (Serine/threonine-protein kinase 2). | |||||
|
AZI1_MOUSE
|
||||||
| NC score | 0.013431 (rank : 129) | θ value | 6.88961 (rank : 87) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 990 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q62036 | Gene names | Azi1, Az1, Azi | |||
|
Domain Architecture |

|
|||||
| Description | 5-azacytidine-induced protein 1 (Pre-acrosome localization protein 1). | |||||
|
EVPL_MOUSE
|
||||||
| NC score | 0.013131 (rank : 130) | θ value | 2.36792 (rank : 60) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 952 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9D952 | Gene names | Evpl | |||
|
Domain Architecture |

|
|||||
| Description | Envoplakin (p210) (210 kDa cornified envelope precursor protein). | |||||
|
INVS_HUMAN
|
||||||
| NC score | 0.011523 (rank : 131) | θ value | 4.03905 (rank : 76) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9Y283, Q5W0T6, Q8IVX8, Q9BRB9, Q9Y488, Q9Y498 | Gene names | INVS, INV, NPHP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inversin (Inversion of embryo turning homolog) (Nephrocystin-2). | |||||
|
EVPL_HUMAN
|
||||||
| NC score | 0.011511 (rank : 132) | θ value | 6.88961 (rank : 88) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 895 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q92817 | Gene names | EVPL | |||
|
Domain Architecture |

|
|||||
| Description | Envoplakin (210 kDa paraneoplastic pemphigus antigen) (p210) (210 kDa cornified envelope precursor protein). | |||||
|
FOXP4_HUMAN
|
||||||
| NC score | 0.010135 (rank : 133) | θ value | 6.88961 (rank : 89) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8IVH2, Q96E19 | Gene names | FOXP4, FKHLA | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein P4 (Fork head-related protein-like A). | |||||
|
AZI1_HUMAN
|
||||||
| NC score | 0.009271 (rank : 134) | θ value | 8.99809 (rank : 95) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 1000 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9UPN4, Q96F50 | Gene names | AZI1, CEP131, KIAA1118 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 5-azacytidine-induced protein 1 (Pre-acrosome localization protein 1) (Centrosomal protein of 131 kDa) (Cep131 protein). | |||||
|
FOXP4_MOUSE
|
||||||
| NC score | 0.009161 (rank : 135) | θ value | 8.99809 (rank : 97) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9DBY0, Q80V92, Q8CG10, Q8CIS1 | Gene names | Foxp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Forkhead box protein P4 (Fork head-related protein like A) (mFKHLA). | |||||
|
ANDR_HUMAN
|
||||||
| NC score | 0.005670 (rank : 136) | θ value | 6.88961 (rank : 86) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P10275 | Gene names | AR, DHTR, NR3C4 | |||
|
Domain Architecture |

|
|||||
| Description | Androgen receptor (Dihydrotestosterone receptor). | |||||
|
TMF1_HUMAN
|
||||||
| NC score | 0.004256 (rank : 137) | θ value | 8.99809 (rank : 104) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 987 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P82094 | Gene names | TMF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TATA element modulatory factor (TMF). | |||||
|
ATBF1_HUMAN
|
||||||
| NC score | 0.004233 (rank : 138) | θ value | 8.99809 (rank : 94) | |||
| Query Neighborhood Hits | 106 | Target Neighborhood Hits | 1547 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q15911, O15101, Q13719 | Gene names | ATBF1 | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-fetoprotein enhancer-binding protein (AT motif-binding factor) (AT-binding transcription factor 1). | |||||