Please be patient as the page loads
|
P4HA2_MOUSE
|
||||||
| SwissProt Accessions | Q60716, Q8VBU4 | Gene names | P4ha2 | |||
|
Domain Architecture |

|
|||||
| Description | Prolyl 4-hydroxylase alpha-2 subunit precursor (EC 1.14.11.2) (4-PH alpha-2) (Procollagen-proline,2-oxoglutarate-4-dioxygenase alpha-2 subunit). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
P4HA1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.989933 (rank : 3) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P13674, Q15082, Q15083, Q5VSQ5 | Gene names | P4HA1, P4HA | |||
|
Domain Architecture |

|
|||||
| Description | Prolyl 4-hydroxylase alpha-1 subunit precursor (EC 1.14.11.2) (4-PH alpha-1) (Procollagen-proline,2-oxoglutarate-4-dioxygenase alpha-1 subunit). | |||||
|
P4HA1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.989092 (rank : 4) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q60715, Q3TEB7, Q80T05, Q91VJ7 | Gene names | P4ha1 | |||
|
Domain Architecture |

|
|||||
| Description | Prolyl 4-hydroxylase alpha-1 subunit precursor (EC 1.14.11.2) (4-PH alpha-1) (Procollagen-proline,2-oxoglutarate-4-dioxygenase alpha-1 subunit). | |||||
|
P4HA2_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.997238 (rank : 2) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O15460, Q8WWN0 | Gene names | P4HA2 | |||
|
Domain Architecture |

|
|||||
| Description | Prolyl 4-hydroxylase alpha-2 subunit precursor (EC 1.14.11.2) (4-PH alpha-2) (Procollagen-proline,2-oxoglutarate-4-dioxygenase alpha-2 subunit). | |||||
|
P4HA2_MOUSE
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q60716, Q8VBU4 | Gene names | P4ha2 | |||
|
Domain Architecture |

|
|||||
| Description | Prolyl 4-hydroxylase alpha-2 subunit precursor (EC 1.14.11.2) (4-PH alpha-2) (Procollagen-proline,2-oxoglutarate-4-dioxygenase alpha-2 subunit). | |||||
|
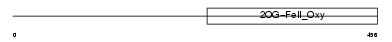
EGLX_MOUSE
|
||||||
| θ value | 2.0648e-12 (rank : 5) | NC score | 0.511549 (rank : 6) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8BG58, Q8CAF1, Q9D499 | Gene names | Ph4 | |||
|
Domain Architecture |
|
|||||
| Description | Putative HIF-prolyl hydroxylase PH-4 (EC 1.14.11.-) (Hypoxia-inducible factor prolyl 4-hydroxylase). | |||||
|
EGLX_HUMAN
|
||||||
| θ value | 3.52202e-12 (rank : 6) | NC score | 0.511806 (rank : 5) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9NXG6, Q6PAG6, Q8TCJ9, Q8WV55, Q96F22, Q9BW77 | Gene names | PH4 | |||
|
Domain Architecture |
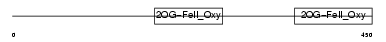
|
|||||
| Description | Putative HIF-prolyl hydroxylase PH-4 (EC 1.14.11.-) (Hypoxia-inducible factor prolyl 4-hydroxylase). | |||||
|
P3H1_HUMAN
|
||||||
| θ value | 0.00665767 (rank : 7) | NC score | 0.102102 (rank : 10) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q32P28, Q7KZR4, Q96BR8, Q96SK8, Q96SL5, Q96SN3, Q9H6K3, Q9HC86, Q9HC87 | Gene names | LEPRE1, GROS1, P3H1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prolyl 3-hydroxylase 1 precursor (EC 1.14.11.7) (Leucine- and proline- enriched proteoglycan 1) (Leprecan-1) (Growth suppressor 1). | |||||
|
P3H1_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 8) | NC score | 0.116125 (rank : 9) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q3V1T4, Q3TWX8, Q8BSV2, Q8CFL3, Q9CWK5, Q9QZT6, Q9QZT7 | Gene names | Lepre1, Gros1, P3h1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prolyl 3-hydroxylase 1 precursor (EC 1.14.11.7) (Leucine- and proline- enriched proteoglycan 1) (Leprecan-1) (Growth suppressor 1). | |||||
|
P3H2_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 9) | NC score | 0.120459 (rank : 8) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8IVL5, Q9NVI2 | Gene names | LEPREL1, MLAT4, P3H2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prolyl 3-hydroxylase 2 precursor (EC 1.14.11.7) (Leprecan-like protein 1) (Myxoid liposarcoma-associated protein 4). | |||||
|
P3H2_MOUSE
|
||||||
| θ value | 0.279714 (rank : 10) | NC score | 0.121824 (rank : 7) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8CG71, Q8C673 | Gene names | Leprel1, P3h2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prolyl 3-hydroxylase 2 precursor (EC 1.14.11.7) (Leprecan-like protein 1). | |||||
|
NO55_HUMAN
|
||||||
| θ value | 0.62314 (rank : 11) | NC score | 0.092680 (rank : 11) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q92791, Q9H4F6 | Gene names | SC65, NOL55 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleolar autoantigen No55. | |||||
|
PCYOX_MOUSE
|
||||||
| θ value | 0.813845 (rank : 12) | NC score | 0.081780 (rank : 12) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9CQF9, Q3UHV6, Q69ZW0, Q8BZX1 | Gene names | Pcyox1, Kiaa0908 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prenylcysteine oxidase precursor (EC 1.8.3.5). | |||||
|
AZI1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 13) | NC score | 0.007126 (rank : 21) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 990 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q62036 | Gene names | Azi1, Az1, Azi | |||
|
Domain Architecture |

|
|||||
| Description | 5-azacytidine-induced protein 1 (Pre-acrosome localization protein 1). | |||||
|
SPTB1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 14) | NC score | 0.004677 (rank : 22) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P15508 | Gene names | Sptb, Spnb-1, Spnb1, Sptb1 | |||
|
Domain Architecture |

|
|||||
| Description | Spectrin beta chain, erythrocyte (Beta-I spectrin). | |||||
|
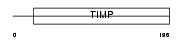
TIMP1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 15) | NC score | 0.025189 (rank : 18) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P12032, P20064, Q61720 | Gene names | Timp1, Timp, Timp-1 | |||
|
Domain Architecture |
|
|||||
| Description | Metalloproteinase inhibitor 1 precursor (TIMP-1) (Erythroid potentiating activity) (EPA) (Tissue inhibitor of metalloproteinases) (Collagenase inhibitor 16C8 fibroblast) (TPA-induced protein) (TPA- S1). | |||||
|
ANR11_HUMAN
|
||||||
| θ value | 5.27518 (rank : 16) | NC score | 0.009427 (rank : 20) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 1289 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6UB99, Q6NTG1, Q6QMF8 | Gene names | ANKRD11, ANCO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 11 (Ankyrin repeat-containing cofactor 1). | |||||
|
MTMRD_HUMAN
|
||||||
| θ value | 6.88961 (rank : 17) | NC score | 0.011885 (rank : 19) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q86WG5, Q3MJF0, Q68DQ3, Q6P459, Q6PJD1, Q7Z325, Q7Z621, Q86VE2, Q96FE2, Q9C097 | Gene names | SBF2, CMT4B2, KIAA1766, MTMR13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myotubularin-related protein 13 (SET-binding factor 2). | |||||
|
PCYOX_HUMAN
|
||||||
| θ value | 6.88961 (rank : 18) | NC score | 0.038228 (rank : 17) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UHG3, O94982, Q8N4N5, Q96QM8 | Gene names | PCYOX1, KIAA0908, PCL1 | |||
|
Domain Architecture |

|
|||||
| Description | Prenylcysteine oxidase precursor (EC 1.8.3.5) (PCL1). | |||||
|
CRTAP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 19) | NC score | 0.057190 (rank : 16) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O75718 | Gene names | CRTAP, CASP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cartilage-associated protein precursor. | |||||
|
CRTAP_MOUSE
|
||||||
| θ value | θ > 10 (rank : 20) | NC score | 0.060878 (rank : 15) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9CYD3, O88698 | Gene names | Crtap, Casp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cartilage-associated protein precursor. | |||||
|
P3H3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 21) | NC score | 0.062960 (rank : 13) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8IVL6, Q13512, Q15740, Q66K32, Q6NX61, Q7L2T1 | Gene names | LEPREL2, P3H3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prolyl 3-hydroxylase 3 precursor (EC 1.14.11.7) (Leprecan-like protein 2) (Protein B). | |||||
|
P3H3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 22) | NC score | 0.062291 (rank : 14) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8CG70, O88836 | Gene names | Leprel2, P3h3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prolyl 3-hydroxylase 3 precursor (EC 1.14.11.7) (Leprecan-like protein 2) (Protein B). | |||||
|
P4HA2_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q60716, Q8VBU4 | Gene names | P4ha2 | |||
|
Domain Architecture |

|
|||||
| Description | Prolyl 4-hydroxylase alpha-2 subunit precursor (EC 1.14.11.2) (4-PH alpha-2) (Procollagen-proline,2-oxoglutarate-4-dioxygenase alpha-2 subunit). | |||||
|
P4HA2_HUMAN
|
||||||
| NC score | 0.997238 (rank : 2) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O15460, Q8WWN0 | Gene names | P4HA2 | |||
|
Domain Architecture |

|
|||||
| Description | Prolyl 4-hydroxylase alpha-2 subunit precursor (EC 1.14.11.2) (4-PH alpha-2) (Procollagen-proline,2-oxoglutarate-4-dioxygenase alpha-2 subunit). | |||||
|
P4HA1_HUMAN
|
||||||
| NC score | 0.989933 (rank : 3) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P13674, Q15082, Q15083, Q5VSQ5 | Gene names | P4HA1, P4HA | |||
|
Domain Architecture |

|
|||||
| Description | Prolyl 4-hydroxylase alpha-1 subunit precursor (EC 1.14.11.2) (4-PH alpha-1) (Procollagen-proline,2-oxoglutarate-4-dioxygenase alpha-1 subunit). | |||||
|
P4HA1_MOUSE
|
||||||
| NC score | 0.989092 (rank : 4) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q60715, Q3TEB7, Q80T05, Q91VJ7 | Gene names | P4ha1 | |||
|
Domain Architecture |

|
|||||
| Description | Prolyl 4-hydroxylase alpha-1 subunit precursor (EC 1.14.11.2) (4-PH alpha-1) (Procollagen-proline,2-oxoglutarate-4-dioxygenase alpha-1 subunit). | |||||
|
EGLX_HUMAN
|
||||||
| NC score | 0.511806 (rank : 5) | θ value | 3.52202e-12 (rank : 6) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9NXG6, Q6PAG6, Q8TCJ9, Q8WV55, Q96F22, Q9BW77 | Gene names | PH4 | |||
|
Domain Architecture |
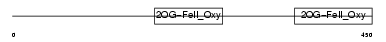
|
|||||
| Description | Putative HIF-prolyl hydroxylase PH-4 (EC 1.14.11.-) (Hypoxia-inducible factor prolyl 4-hydroxylase). | |||||
|
EGLX_MOUSE
|
||||||
| NC score | 0.511549 (rank : 6) | θ value | 2.0648e-12 (rank : 5) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8BG58, Q8CAF1, Q9D499 | Gene names | Ph4 | |||
|
Domain Architecture |

|
|||||
| Description | Putative HIF-prolyl hydroxylase PH-4 (EC 1.14.11.-) (Hypoxia-inducible factor prolyl 4-hydroxylase). | |||||
|
P3H2_MOUSE
|
||||||
| NC score | 0.121824 (rank : 7) | θ value | 0.279714 (rank : 10) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8CG71, Q8C673 | Gene names | Leprel1, P3h2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prolyl 3-hydroxylase 2 precursor (EC 1.14.11.7) (Leprecan-like protein 1). | |||||
|
P3H2_HUMAN
|
||||||
| NC score | 0.120459 (rank : 8) | θ value | 0.0961366 (rank : 9) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8IVL5, Q9NVI2 | Gene names | LEPREL1, MLAT4, P3H2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prolyl 3-hydroxylase 2 precursor (EC 1.14.11.7) (Leprecan-like protein 1) (Myxoid liposarcoma-associated protein 4). | |||||
|
P3H1_MOUSE
|
||||||
| NC score | 0.116125 (rank : 9) | θ value | 0.0113563 (rank : 8) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q3V1T4, Q3TWX8, Q8BSV2, Q8CFL3, Q9CWK5, Q9QZT6, Q9QZT7 | Gene names | Lepre1, Gros1, P3h1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prolyl 3-hydroxylase 1 precursor (EC 1.14.11.7) (Leucine- and proline- enriched proteoglycan 1) (Leprecan-1) (Growth suppressor 1). | |||||
|
P3H1_HUMAN
|
||||||
| NC score | 0.102102 (rank : 10) | θ value | 0.00665767 (rank : 7) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q32P28, Q7KZR4, Q96BR8, Q96SK8, Q96SL5, Q96SN3, Q9H6K3, Q9HC86, Q9HC87 | Gene names | LEPRE1, GROS1, P3H1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prolyl 3-hydroxylase 1 precursor (EC 1.14.11.7) (Leucine- and proline- enriched proteoglycan 1) (Leprecan-1) (Growth suppressor 1). | |||||
|
NO55_HUMAN
|
||||||
| NC score | 0.092680 (rank : 11) | θ value | 0.62314 (rank : 11) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q92791, Q9H4F6 | Gene names | SC65, NOL55 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleolar autoantigen No55. | |||||
|
PCYOX_MOUSE
|
||||||
| NC score | 0.081780 (rank : 12) | θ value | 0.813845 (rank : 12) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9CQF9, Q3UHV6, Q69ZW0, Q8BZX1 | Gene names | Pcyox1, Kiaa0908 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prenylcysteine oxidase precursor (EC 1.8.3.5). | |||||
|
P3H3_HUMAN
|
||||||
| NC score | 0.062960 (rank : 13) | θ value | θ > 10 (rank : 21) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8IVL6, Q13512, Q15740, Q66K32, Q6NX61, Q7L2T1 | Gene names | LEPREL2, P3H3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prolyl 3-hydroxylase 3 precursor (EC 1.14.11.7) (Leprecan-like protein 2) (Protein B). | |||||
|
P3H3_MOUSE
|
||||||
| NC score | 0.062291 (rank : 14) | θ value | θ > 10 (rank : 22) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8CG70, O88836 | Gene names | Leprel2, P3h3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prolyl 3-hydroxylase 3 precursor (EC 1.14.11.7) (Leprecan-like protein 2) (Protein B). | |||||
|
CRTAP_MOUSE
|
||||||
| NC score | 0.060878 (rank : 15) | θ value | θ > 10 (rank : 20) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9CYD3, O88698 | Gene names | Crtap, Casp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cartilage-associated protein precursor. | |||||
|
CRTAP_HUMAN
|
||||||
| NC score | 0.057190 (rank : 16) | θ value | θ > 10 (rank : 19) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O75718 | Gene names | CRTAP, CASP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cartilage-associated protein precursor. | |||||
|
PCYOX_HUMAN
|
||||||
| NC score | 0.038228 (rank : 17) | θ value | 6.88961 (rank : 18) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UHG3, O94982, Q8N4N5, Q96QM8 | Gene names | PCYOX1, KIAA0908, PCL1 | |||
|
Domain Architecture |

|
|||||
| Description | Prenylcysteine oxidase precursor (EC 1.8.3.5) (PCL1). | |||||
|
TIMP1_MOUSE
|
||||||
| NC score | 0.025189 (rank : 18) | θ value | 4.03905 (rank : 15) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P12032, P20064, Q61720 | Gene names | Timp1, Timp, Timp-1 | |||
|
Domain Architecture |

|
|||||
| Description | Metalloproteinase inhibitor 1 precursor (TIMP-1) (Erythroid potentiating activity) (EPA) (Tissue inhibitor of metalloproteinases) (Collagenase inhibitor 16C8 fibroblast) (TPA-induced protein) (TPA- S1). | |||||
|
MTMRD_HUMAN
|
||||||
| NC score | 0.011885 (rank : 19) | θ value | 6.88961 (rank : 17) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q86WG5, Q3MJF0, Q68DQ3, Q6P459, Q6PJD1, Q7Z325, Q7Z621, Q86VE2, Q96FE2, Q9C097 | Gene names | SBF2, CMT4B2, KIAA1766, MTMR13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myotubularin-related protein 13 (SET-binding factor 2). | |||||
|
ANR11_HUMAN
|
||||||
| NC score | 0.009427 (rank : 20) | θ value | 5.27518 (rank : 16) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 1289 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6UB99, Q6NTG1, Q6QMF8 | Gene names | ANKRD11, ANCO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 11 (Ankyrin repeat-containing cofactor 1). | |||||
|
AZI1_MOUSE
|
||||||
| NC score | 0.007126 (rank : 21) | θ value | 2.36792 (rank : 13) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 990 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q62036 | Gene names | Azi1, Az1, Azi | |||
|
Domain Architecture |

|
|||||
| Description | 5-azacytidine-induced protein 1 (Pre-acrosome localization protein 1). | |||||
|
SPTB1_MOUSE
|
||||||
| NC score | 0.004677 (rank : 22) | θ value | 4.03905 (rank : 14) | |||
| Query Neighborhood Hits | 18 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P15508 | Gene names | Sptb, Spnb-1, Spnb1, Sptb1 | |||
|
Domain Architecture |

|
|||||
| Description | Spectrin beta chain, erythrocyte (Beta-I spectrin). | |||||