Please be patient as the page loads
|
P3H1_HUMAN
|
||||||
| SwissProt Accessions | Q32P28, Q7KZR4, Q96BR8, Q96SK8, Q96SL5, Q96SN3, Q9H6K3, Q9HC86, Q9HC87 | Gene names | LEPRE1, GROS1, P3H1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prolyl 3-hydroxylase 1 precursor (EC 1.14.11.7) (Leucine- and proline- enriched proteoglycan 1) (Leprecan-1) (Growth suppressor 1). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
P3H1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q32P28, Q7KZR4, Q96BR8, Q96SK8, Q96SL5, Q96SN3, Q9H6K3, Q9HC86, Q9HC87 | Gene names | LEPRE1, GROS1, P3H1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prolyl 3-hydroxylase 1 precursor (EC 1.14.11.7) (Leucine- and proline- enriched proteoglycan 1) (Leprecan-1) (Growth suppressor 1). | |||||
|
P3H1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.998206 (rank : 2) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q3V1T4, Q3TWX8, Q8BSV2, Q8CFL3, Q9CWK5, Q9QZT6, Q9QZT7 | Gene names | Lepre1, Gros1, P3h1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prolyl 3-hydroxylase 1 precursor (EC 1.14.11.7) (Leucine- and proline- enriched proteoglycan 1) (Leprecan-1) (Growth suppressor 1). | |||||
|
P3H2_HUMAN
|
||||||
| θ value | 4.1007e-178 (rank : 3) | NC score | 0.982657 (rank : 3) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8IVL5, Q9NVI2 | Gene names | LEPREL1, MLAT4, P3H2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prolyl 3-hydroxylase 2 precursor (EC 1.14.11.7) (Leprecan-like protein 1) (Myxoid liposarcoma-associated protein 4). | |||||
|
P3H2_MOUSE
|
||||||
| θ value | 3.5924e-174 (rank : 4) | NC score | 0.981997 (rank : 4) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8CG71, Q8C673 | Gene names | Leprel1, P3h2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prolyl 3-hydroxylase 2 precursor (EC 1.14.11.7) (Leprecan-like protein 1). | |||||
|
P3H3_HUMAN
|
||||||
| θ value | 4.57613e-145 (rank : 5) | NC score | 0.970050 (rank : 5) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8IVL6, Q13512, Q15740, Q66K32, Q6NX61, Q7L2T1 | Gene names | LEPREL2, P3H3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prolyl 3-hydroxylase 3 precursor (EC 1.14.11.7) (Leprecan-like protein 2) (Protein B). | |||||
|
P3H3_MOUSE
|
||||||
| θ value | 1.09172e-138 (rank : 6) | NC score | 0.968213 (rank : 6) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8CG70, O88836 | Gene names | Leprel2, P3h3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prolyl 3-hydroxylase 3 precursor (EC 1.14.11.7) (Leprecan-like protein 2) (Protein B). | |||||
|
CRTAP_HUMAN
|
||||||
| θ value | 2.57589e-55 (rank : 7) | NC score | 0.879559 (rank : 7) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O75718 | Gene names | CRTAP, CASP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cartilage-associated protein precursor. | |||||
|
CRTAP_MOUSE
|
||||||
| θ value | 1.51363e-47 (rank : 8) | NC score | 0.870456 (rank : 8) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9CYD3, O88698 | Gene names | Crtap, Casp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cartilage-associated protein precursor. | |||||
|
NO55_HUMAN
|
||||||
| θ value | 5.95217e-44 (rank : 9) | NC score | 0.862192 (rank : 9) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q92791, Q9H4F6 | Gene names | SC65, NOL55 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleolar autoantigen No55. | |||||
|
P4HA2_MOUSE
|
||||||
| θ value | 0.00665767 (rank : 10) | NC score | 0.102102 (rank : 10) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q60716, Q8VBU4 | Gene names | P4ha2 | |||
|
Domain Architecture |

|
|||||
| Description | Prolyl 4-hydroxylase alpha-2 subunit precursor (EC 1.14.11.2) (4-PH alpha-2) (Procollagen-proline,2-oxoglutarate-4-dioxygenase alpha-2 subunit). | |||||
|
NOP14_HUMAN
|
||||||
| θ value | 0.21417 (rank : 11) | NC score | 0.045576 (rank : 20) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 573 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P78316, Q7LGI5, Q7Z6K0, Q8TBR6 | Gene names | NOP14, C4orf9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable nucleolar complex protein 14. | |||||
|
EGLN2_MOUSE
|
||||||
| θ value | 0.47712 (rank : 12) | NC score | 0.085732 (rank : 11) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q91YE2, Q8C6I4, Q8CIL9, Q8VHJ1, Q99MI0 | Gene names | Egln2 | |||
|
Domain Architecture |
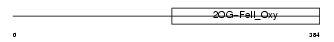
|
|||||
| Description | Egl nine homolog 2 (EC 1.14.11.-) (Hypoxia-inducible factor prolyl hydroxylase 1) (HIF-prolyl hydroxylase 1) (HIF-PH1) (Falkor). | |||||
|
EGLN1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 13) | NC score | 0.079066 (rank : 12) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9GZT9, Q8N3M8, Q9BZS8, Q9BZT0 | Gene names | EGLN1, C1orf12 | |||
|
Domain Architecture |
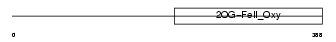
|
|||||
| Description | Egl nine homolog 1 (EC 1.14.11.-) (Hypoxia-inducible factor prolyl hydroxylase 2) (HIF-prolyl hydroxylase 2) (HIF-PH2) (HPH-2) (Prolyl hydroxylase domain-containing protein 2) (PHD2) (SM-20). | |||||
|
EGLN1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 14) | NC score | 0.077954 (rank : 13) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q91YE3, Q8VHJ2, Q922P3 | Gene names | Egln1 | |||
|
Domain Architecture |
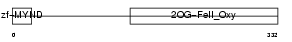
|
|||||
| Description | Egl nine homolog 1 (EC 1.14.11.-) (Hypoxia-inducible factor prolyl hydroxylase 2) (HIF-prolyl hydroxylase 2) (HIF-PH2) (HPH-2) (SM-20). | |||||
|
EGLN2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 15) | NC score | 0.076681 (rank : 15) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q96KS0, Q8WWY4, Q9BV14 | Gene names | EGLN2, EIT6 | |||
|
Domain Architecture |

|
|||||
| Description | Egl nine homolog 2 (EC 1.14.11.-) (Hypoxia-inducible factor prolyl hydroxylase 1) (HIF-prolyl hydroxylase 1) (HIF-PH1) (HPH-3) (Prolyl hydroxylase domain-containing protein 1) (PHD1) (Estrogen-induced tag 6). | |||||
|
NOP14_MOUSE
|
||||||
| θ value | 1.06291 (rank : 16) | NC score | 0.045350 (rank : 21) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 324 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8R3N1 | Gene names | Nop14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable nucleolar complex protein 14. | |||||
|
RBBP6_HUMAN
|
||||||
| θ value | 1.06291 (rank : 17) | NC score | 0.018600 (rank : 22) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q7Z6E9, Q15290, Q6DKH4, Q6P4C2, Q6YNC9, Q7Z6E8, Q8N0V2, Q96PH3, Q9H3I8, Q9H5M5, Q9NPX4 | Gene names | RBBP6, P2PR, PACT, RBQ1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R) (Retinoblastoma-binding Q protein 1) (Protein RBQ-1). | |||||
|
RAPH1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 18) | NC score | 0.013026 (rank : 25) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 593 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q70E73, Q96Q37, Q9C0I2 | Gene names | RAPH1, ALS2CR9, KIAA1681, LPD, PREL2, RMO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras-associated and pleckstrin homology domains-containing protein 1 (RAPH1) (Lamellipodin) (Proline-rich EVH1 ligand 2) (PREL-2) (Protein RMO1) (Amyotrophic lateral sclerosis 2 chromosomal region candidate 9 gene protein). | |||||
|
EP400_HUMAN
|
||||||
| θ value | 2.36792 (rank : 19) | NC score | 0.013680 (rank : 24) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 556 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96L91, O15411, Q6P2F5, Q8N8Q7, Q8NE05, Q96JK7, Q9P230 | Gene names | EP400, CAGH32, KIAA1498, KIAA1818, TNRC12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E1A-binding protein p400 (EC 3.6.1.-) (p400 kDa SWI2/SNF2-related protein) (Domino homolog) (hDomino) (CAG repeat protein 32) (Trinucleotide repeat-containing gene 12 protein). | |||||
|
RTN4_HUMAN
|
||||||
| θ value | 2.36792 (rank : 20) | NC score | 0.012495 (rank : 26) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NQC3, O94962, Q9BXG5, Q9H212, Q9H3I3, Q9UQ42, Q9Y293, Q9Y2Y7, Q9Y5U6 | Gene names | RTN4, KIAA0886, NOGO | |||
|
Domain Architecture |

|
|||||
| Description | Reticulon-4 (Neurite outgrowth inhibitor) (Nogo protein) (Foocen) (Neuroendocrine-specific protein) (NSP) (Neuroendocrine-specific protein C homolog) (RTN-x) (Reticulon-5). | |||||
|
CLD8_MOUSE
|
||||||
| θ value | 4.03905 (rank : 21) | NC score | 0.005997 (rank : 32) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Z260 | Gene names | Cldn8 | |||
|
Domain Architecture |

|
|||||
| Description | Claudin-8. | |||||
|
FARP1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 22) | NC score | 0.004668 (rank : 33) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y4F1, Q6IQ29 | Gene names | FARP1, CDEP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FERM, RhoGEF and pleckstrin domain-containing protein 1 (Chondrocyte- derived ezrin-like protein). | |||||
|
K2002_MOUSE
|
||||||
| θ value | 4.03905 (rank : 23) | NC score | 0.011686 (rank : 27) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q69Z38, Q8R365 | Gene names | Kiaa2002 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized tyrosine-protein kinase KIAA2002 (EC 2.7.10.2). | |||||
|
P4HA1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 24) | NC score | 0.062703 (rank : 16) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P13674, Q15082, Q15083, Q5VSQ5 | Gene names | P4HA1, P4HA | |||
|
Domain Architecture |

|
|||||
| Description | Prolyl 4-hydroxylase alpha-1 subunit precursor (EC 1.14.11.2) (4-PH alpha-1) (Procollagen-proline,2-oxoglutarate-4-dioxygenase alpha-1 subunit). | |||||
|
4ET_HUMAN
|
||||||
| θ value | 5.27518 (rank : 25) | NC score | 0.016822 (rank : 23) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NRA8, Q8NCF2, Q9H708 | Gene names | EIF4ENIF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 4E transporter (eIF4E transporter) (4E-T) (Eukaryotic translation initiation factor 4E nuclear import factor 1). | |||||
|
EGLN3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 26) | NC score | 0.057273 (rank : 19) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9H6Z9 | Gene names | EGLN3 | |||
|
Domain Architecture |

|
|||||
| Description | Egl nine homolog 3 (EC 1.14.11.-) (Hypoxia-inducible factor prolyl hydroxylase 3) (HIF-prolyl hydroxylase 3) (HIF-PH3) (HPH-1) (Prolyl hydroxylase domain-containing protein 3) (PHD3). | |||||
|
EGLN3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 27) | NC score | 0.057279 (rank : 18) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q91UZ4, Q3TCG8, Q8C8M6, Q8CCA8, Q8R5C7 | Gene names | Egln3 | |||
|
Domain Architecture |

|
|||||
| Description | Egl nine homolog 3 (EC 1.14.11.-) (Hypoxia-inducible factor prolyl hydroxylase 3) (HIF-prolyl hydroxylase 3) (HIF-PH3) (SM-20). | |||||
|
PHF8_MOUSE
|
||||||
| θ value | 5.27518 (rank : 28) | NC score | 0.006922 (rank : 31) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q80TJ7, Q8BLX8, Q8BLY0, Q8BZ61, Q8CG26 | Gene names | Phf8, Kiaa1111 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 8. | |||||
|
UACA_MOUSE
|
||||||
| θ value | 5.27518 (rank : 29) | NC score | 0.001392 (rank : 35) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8CGB3, Q69ZG3, Q7TN77, Q8BJC8 | Gene names | Uaca, Kiaa1561 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uveal autoantigen with coiled-coil domains and ankyrin repeats protein (Nucling) (Nuclear membrane-binding protein). | |||||
|
YLPM1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 30) | NC score | 0.009033 (rank : 30) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 505 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9R0I7, Q7TMM4 | Gene names | Ylpm1, Zap, Zap3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3). | |||||
|
P4HA1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 31) | NC score | 0.057880 (rank : 17) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q60715, Q3TEB7, Q80T05, Q91VJ7 | Gene names | P4ha1 | |||
|
Domain Architecture |

|
|||||
| Description | Prolyl 4-hydroxylase alpha-1 subunit precursor (EC 1.14.11.2) (4-PH alpha-1) (Procollagen-proline,2-oxoglutarate-4-dioxygenase alpha-1 subunit). | |||||
|
CHD6_HUMAN
|
||||||
| θ value | 8.99809 (rank : 32) | NC score | 0.004643 (rank : 34) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8TD26, Q5JYQ0, Q5TGZ9, Q5TH00, Q5TH01, Q8IZR2, Q8WTY0, Q9H4H6, Q9H6D4, Q9NTT7, Q9P2L1 | Gene names | CHD6, CHD5, KIAA1335, RIGB | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 6 (EC 3.6.1.-) (ATP- dependent helicase CHD6) (CHD-6) (Radiation-induced gene B protein). | |||||
|
DBF4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 33) | NC score | 0.009039 (rank : 29) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UBU7, O75226, Q75MS6, Q75N01, Q9Y2M6 | Gene names | DBF4, ASK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein DBF4 homolog (Activator of S phase Kinase). | |||||
|
ZNHI4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 34) | NC score | 0.010728 (rank : 28) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9C086 | Gene names | ZNHIT4, HMGA1L4, PAPA1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger HIT domain-containing protein 4 (PAP-1-associated protein 1) (PAPA-1) (High mobility group AT-hook 1-like 4). | |||||
|
P4HA2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 35) | NC score | 0.076927 (rank : 14) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O15460, Q8WWN0 | Gene names | P4HA2 | |||
|
Domain Architecture |

|
|||||
| Description | Prolyl 4-hydroxylase alpha-2 subunit precursor (EC 1.14.11.2) (4-PH alpha-2) (Procollagen-proline,2-oxoglutarate-4-dioxygenase alpha-2 subunit). | |||||
|
P3H1_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q32P28, Q7KZR4, Q96BR8, Q96SK8, Q96SL5, Q96SN3, Q9H6K3, Q9HC86, Q9HC87 | Gene names | LEPRE1, GROS1, P3H1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prolyl 3-hydroxylase 1 precursor (EC 1.14.11.7) (Leucine- and proline- enriched proteoglycan 1) (Leprecan-1) (Growth suppressor 1). | |||||
|
P3H1_MOUSE
|
||||||
| NC score | 0.998206 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q3V1T4, Q3TWX8, Q8BSV2, Q8CFL3, Q9CWK5, Q9QZT6, Q9QZT7 | Gene names | Lepre1, Gros1, P3h1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prolyl 3-hydroxylase 1 precursor (EC 1.14.11.7) (Leucine- and proline- enriched proteoglycan 1) (Leprecan-1) (Growth suppressor 1). | |||||
|
P3H2_HUMAN
|
||||||
| NC score | 0.982657 (rank : 3) | θ value | 4.1007e-178 (rank : 3) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8IVL5, Q9NVI2 | Gene names | LEPREL1, MLAT4, P3H2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prolyl 3-hydroxylase 2 precursor (EC 1.14.11.7) (Leprecan-like protein 1) (Myxoid liposarcoma-associated protein 4). | |||||
|
P3H2_MOUSE
|
||||||
| NC score | 0.981997 (rank : 4) | θ value | 3.5924e-174 (rank : 4) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8CG71, Q8C673 | Gene names | Leprel1, P3h2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prolyl 3-hydroxylase 2 precursor (EC 1.14.11.7) (Leprecan-like protein 1). | |||||
|
P3H3_HUMAN
|
||||||
| NC score | 0.970050 (rank : 5) | θ value | 4.57613e-145 (rank : 5) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8IVL6, Q13512, Q15740, Q66K32, Q6NX61, Q7L2T1 | Gene names | LEPREL2, P3H3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prolyl 3-hydroxylase 3 precursor (EC 1.14.11.7) (Leprecan-like protein 2) (Protein B). | |||||
|
P3H3_MOUSE
|
||||||
| NC score | 0.968213 (rank : 6) | θ value | 1.09172e-138 (rank : 6) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8CG70, O88836 | Gene names | Leprel2, P3h3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prolyl 3-hydroxylase 3 precursor (EC 1.14.11.7) (Leprecan-like protein 2) (Protein B). | |||||
|
CRTAP_HUMAN
|
||||||
| NC score | 0.879559 (rank : 7) | θ value | 2.57589e-55 (rank : 7) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O75718 | Gene names | CRTAP, CASP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cartilage-associated protein precursor. | |||||
|
CRTAP_MOUSE
|
||||||
| NC score | 0.870456 (rank : 8) | θ value | 1.51363e-47 (rank : 8) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9CYD3, O88698 | Gene names | Crtap, Casp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cartilage-associated protein precursor. | |||||
|
NO55_HUMAN
|
||||||
| NC score | 0.862192 (rank : 9) | θ value | 5.95217e-44 (rank : 9) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q92791, Q9H4F6 | Gene names | SC65, NOL55 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleolar autoantigen No55. | |||||
|
P4HA2_MOUSE
|
||||||
| NC score | 0.102102 (rank : 10) | θ value | 0.00665767 (rank : 10) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q60716, Q8VBU4 | Gene names | P4ha2 | |||
|
Domain Architecture |

|
|||||
| Description | Prolyl 4-hydroxylase alpha-2 subunit precursor (EC 1.14.11.2) (4-PH alpha-2) (Procollagen-proline,2-oxoglutarate-4-dioxygenase alpha-2 subunit). | |||||
|
EGLN2_MOUSE
|
||||||
| NC score | 0.085732 (rank : 11) | θ value | 0.47712 (rank : 12) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q91YE2, Q8C6I4, Q8CIL9, Q8VHJ1, Q99MI0 | Gene names | Egln2 | |||
|
Domain Architecture |

|
|||||
| Description | Egl nine homolog 2 (EC 1.14.11.-) (Hypoxia-inducible factor prolyl hydroxylase 1) (HIF-prolyl hydroxylase 1) (HIF-PH1) (Falkor). | |||||
|
EGLN1_HUMAN
|
||||||
| NC score | 0.079066 (rank : 12) | θ value | 0.813845 (rank : 13) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9GZT9, Q8N3M8, Q9BZS8, Q9BZT0 | Gene names | EGLN1, C1orf12 | |||
|
Domain Architecture |

|
|||||
| Description | Egl nine homolog 1 (EC 1.14.11.-) (Hypoxia-inducible factor prolyl hydroxylase 2) (HIF-prolyl hydroxylase 2) (HIF-PH2) (HPH-2) (Prolyl hydroxylase domain-containing protein 2) (PHD2) (SM-20). | |||||
|
EGLN1_MOUSE
|
||||||
| NC score | 0.077954 (rank : 13) | θ value | 0.813845 (rank : 14) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q91YE3, Q8VHJ2, Q922P3 | Gene names | Egln1 | |||
|
Domain Architecture |

|
|||||
| Description | Egl nine homolog 1 (EC 1.14.11.-) (Hypoxia-inducible factor prolyl hydroxylase 2) (HIF-prolyl hydroxylase 2) (HIF-PH2) (HPH-2) (SM-20). | |||||
|
P4HA2_HUMAN
|
||||||
| NC score | 0.076927 (rank : 14) | θ value | θ > 10 (rank : 35) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O15460, Q8WWN0 | Gene names | P4HA2 | |||
|
Domain Architecture |

|
|||||
| Description | Prolyl 4-hydroxylase alpha-2 subunit precursor (EC 1.14.11.2) (4-PH alpha-2) (Procollagen-proline,2-oxoglutarate-4-dioxygenase alpha-2 subunit). | |||||
|
EGLN2_HUMAN
|
||||||
| NC score | 0.076681 (rank : 15) | θ value | 0.813845 (rank : 15) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q96KS0, Q8WWY4, Q9BV14 | Gene names | EGLN2, EIT6 | |||
|
Domain Architecture |

|
|||||
| Description | Egl nine homolog 2 (EC 1.14.11.-) (Hypoxia-inducible factor prolyl hydroxylase 1) (HIF-prolyl hydroxylase 1) (HIF-PH1) (HPH-3) (Prolyl hydroxylase domain-containing protein 1) (PHD1) (Estrogen-induced tag 6). | |||||
|
P4HA1_HUMAN
|
||||||
| NC score | 0.062703 (rank : 16) | θ value | 4.03905 (rank : 24) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P13674, Q15082, Q15083, Q5VSQ5 | Gene names | P4HA1, P4HA | |||
|
Domain Architecture |

|
|||||
| Description | Prolyl 4-hydroxylase alpha-1 subunit precursor (EC 1.14.11.2) (4-PH alpha-1) (Procollagen-proline,2-oxoglutarate-4-dioxygenase alpha-1 subunit). | |||||
|
P4HA1_MOUSE
|
||||||
| NC score | 0.057880 (rank : 17) | θ value | 6.88961 (rank : 31) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q60715, Q3TEB7, Q80T05, Q91VJ7 | Gene names | P4ha1 | |||
|
Domain Architecture |

|
|||||
| Description | Prolyl 4-hydroxylase alpha-1 subunit precursor (EC 1.14.11.2) (4-PH alpha-1) (Procollagen-proline,2-oxoglutarate-4-dioxygenase alpha-1 subunit). | |||||
|
EGLN3_MOUSE
|
||||||
| NC score | 0.057279 (rank : 18) | θ value | 5.27518 (rank : 27) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q91UZ4, Q3TCG8, Q8C8M6, Q8CCA8, Q8R5C7 | Gene names | Egln3 | |||
|
Domain Architecture |

|
|||||
| Description | Egl nine homolog 3 (EC 1.14.11.-) (Hypoxia-inducible factor prolyl hydroxylase 3) (HIF-prolyl hydroxylase 3) (HIF-PH3) (SM-20). | |||||
|
EGLN3_HUMAN
|
||||||
| NC score | 0.057273 (rank : 19) | θ value | 5.27518 (rank : 26) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9H6Z9 | Gene names | EGLN3 | |||
|
Domain Architecture |

|
|||||
| Description | Egl nine homolog 3 (EC 1.14.11.-) (Hypoxia-inducible factor prolyl hydroxylase 3) (HIF-prolyl hydroxylase 3) (HIF-PH3) (HPH-1) (Prolyl hydroxylase domain-containing protein 3) (PHD3). | |||||
|
NOP14_HUMAN
|
||||||
| NC score | 0.045576 (rank : 20) | θ value | 0.21417 (rank : 11) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 573 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P78316, Q7LGI5, Q7Z6K0, Q8TBR6 | Gene names | NOP14, C4orf9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable nucleolar complex protein 14. | |||||
|
NOP14_MOUSE
|
||||||
| NC score | 0.045350 (rank : 21) | θ value | 1.06291 (rank : 16) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 324 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8R3N1 | Gene names | Nop14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable nucleolar complex protein 14. | |||||
|
RBBP6_HUMAN
|
||||||
| NC score | 0.018600 (rank : 22) | θ value | 1.06291 (rank : 17) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q7Z6E9, Q15290, Q6DKH4, Q6P4C2, Q6YNC9, Q7Z6E8, Q8N0V2, Q96PH3, Q9H3I8, Q9H5M5, Q9NPX4 | Gene names | RBBP6, P2PR, PACT, RBQ1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R) (Retinoblastoma-binding Q protein 1) (Protein RBQ-1). | |||||
|
4ET_HUMAN
|
||||||
| NC score | 0.016822 (rank : 23) | θ value | 5.27518 (rank : 25) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NRA8, Q8NCF2, Q9H708 | Gene names | EIF4ENIF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 4E transporter (eIF4E transporter) (4E-T) (Eukaryotic translation initiation factor 4E nuclear import factor 1). | |||||
|
EP400_HUMAN
|
||||||
| NC score | 0.013680 (rank : 24) | θ value | 2.36792 (rank : 19) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 556 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96L91, O15411, Q6P2F5, Q8N8Q7, Q8NE05, Q96JK7, Q9P230 | Gene names | EP400, CAGH32, KIAA1498, KIAA1818, TNRC12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | E1A-binding protein p400 (EC 3.6.1.-) (p400 kDa SWI2/SNF2-related protein) (Domino homolog) (hDomino) (CAG repeat protein 32) (Trinucleotide repeat-containing gene 12 protein). | |||||
|
RAPH1_HUMAN
|
||||||
| NC score | 0.013026 (rank : 25) | θ value | 1.38821 (rank : 18) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 593 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q70E73, Q96Q37, Q9C0I2 | Gene names | RAPH1, ALS2CR9, KIAA1681, LPD, PREL2, RMO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras-associated and pleckstrin homology domains-containing protein 1 (RAPH1) (Lamellipodin) (Proline-rich EVH1 ligand 2) (PREL-2) (Protein RMO1) (Amyotrophic lateral sclerosis 2 chromosomal region candidate 9 gene protein). | |||||
|
RTN4_HUMAN
|
||||||
| NC score | 0.012495 (rank : 26) | θ value | 2.36792 (rank : 20) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NQC3, O94962, Q9BXG5, Q9H212, Q9H3I3, Q9UQ42, Q9Y293, Q9Y2Y7, Q9Y5U6 | Gene names | RTN4, KIAA0886, NOGO | |||
|
Domain Architecture |

|
|||||
| Description | Reticulon-4 (Neurite outgrowth inhibitor) (Nogo protein) (Foocen) (Neuroendocrine-specific protein) (NSP) (Neuroendocrine-specific protein C homolog) (RTN-x) (Reticulon-5). | |||||
|
K2002_MOUSE
|
||||||
| NC score | 0.011686 (rank : 27) | θ value | 4.03905 (rank : 23) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q69Z38, Q8R365 | Gene names | Kiaa2002 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized tyrosine-protein kinase KIAA2002 (EC 2.7.10.2). | |||||
|
ZNHI4_HUMAN
|
||||||
| NC score | 0.010728 (rank : 28) | θ value | 8.99809 (rank : 34) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9C086 | Gene names | ZNHIT4, HMGA1L4, PAPA1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger HIT domain-containing protein 4 (PAP-1-associated protein 1) (PAPA-1) (High mobility group AT-hook 1-like 4). | |||||
|
DBF4_HUMAN
|
||||||
| NC score | 0.009039 (rank : 29) | θ value | 8.99809 (rank : 33) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UBU7, O75226, Q75MS6, Q75N01, Q9Y2M6 | Gene names | DBF4, ASK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein DBF4 homolog (Activator of S phase Kinase). | |||||
|
YLPM1_MOUSE
|
||||||
| NC score | 0.009033 (rank : 30) | θ value | 5.27518 (rank : 30) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 505 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9R0I7, Q7TMM4 | Gene names | Ylpm1, Zap, Zap3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YLP motif-containing protein 1 (Nuclear protein ZAP3). | |||||
|
PHF8_MOUSE
|
||||||
| NC score | 0.006922 (rank : 31) | θ value | 5.27518 (rank : 28) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q80TJ7, Q8BLX8, Q8BLY0, Q8BZ61, Q8CG26 | Gene names | Phf8, Kiaa1111 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 8. | |||||
|
CLD8_MOUSE
|
||||||
| NC score | 0.005997 (rank : 32) | θ value | 4.03905 (rank : 21) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Z260 | Gene names | Cldn8 | |||
|
Domain Architecture |

|
|||||
| Description | Claudin-8. | |||||
|
FARP1_HUMAN
|
||||||
| NC score | 0.004668 (rank : 33) | θ value | 4.03905 (rank : 22) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y4F1, Q6IQ29 | Gene names | FARP1, CDEP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FERM, RhoGEF and pleckstrin domain-containing protein 1 (Chondrocyte- derived ezrin-like protein). | |||||
|
CHD6_HUMAN
|
||||||
| NC score | 0.004643 (rank : 34) | θ value | 8.99809 (rank : 32) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8TD26, Q5JYQ0, Q5TGZ9, Q5TH00, Q5TH01, Q8IZR2, Q8WTY0, Q9H4H6, Q9H6D4, Q9NTT7, Q9P2L1 | Gene names | CHD6, CHD5, KIAA1335, RIGB | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 6 (EC 3.6.1.-) (ATP- dependent helicase CHD6) (CHD-6) (Radiation-induced gene B protein). | |||||
|
UACA_MOUSE
|
||||||
| NC score | 0.001392 (rank : 35) | θ value | 5.27518 (rank : 29) | |||
| Query Neighborhood Hits | 34 | Target Neighborhood Hits | 1620 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8CGB3, Q69ZG3, Q7TN77, Q8BJC8 | Gene names | Uaca, Kiaa1561 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uveal autoantigen with coiled-coil domains and ankyrin repeats protein (Nucling) (Nuclear membrane-binding protein). | |||||