Please be patient as the page loads
|
NRIP1_MOUSE
|
||||||
| SwissProt Accessions | Q8CBD1, Q8C3Y8, Q9Z2K2 | Gene names | Nrip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor-interacting protein 1 (Nuclear factor RIP140) (Receptor-interacting protein 140). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
NRIP1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.963568 (rank : 2) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P48552, Q8IWE8 | Gene names | NRIP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor-interacting protein 1 (Nuclear factor RIP140) (Receptor-interacting protein 140). | |||||
|
NRIP1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 110 | |
| SwissProt Accessions | Q8CBD1, Q8C3Y8, Q9Z2K2 | Gene names | Nrip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor-interacting protein 1 (Nuclear factor RIP140) (Receptor-interacting protein 140). | |||||
|
SRBS1_MOUSE
|
||||||
| θ value | 0.00390308 (rank : 3) | NC score | 0.057960 (rank : 8) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q62417, Q80TF8, Q8BZI3, Q8K3Y2, Q921F8, Q9Z0Z8, Q9Z0Z9 | Gene names | Sorbs1, Kiaa1296, Sh3d5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sorbin and SH3 domain-containing protein 1 (Ponsin) (c-Cbl-associated protein) (CAP) (SH3 domain protein 5) (SH3P12). | |||||
|
SYNPO_HUMAN
|
||||||
| θ value | 0.00665767 (rank : 4) | NC score | 0.081441 (rank : 4) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8N3V7, O15271, Q9UPX1 | Gene names | SYNPO, KIAA1029 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptopodin. | |||||
|
DSPP_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 5) | NC score | 0.098988 (rank : 3) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9NZW4, O95815 | Gene names | DSPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
MLL2_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 6) | NC score | 0.043241 (rank : 24) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
AF9_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 7) | NC score | 0.072095 (rank : 5) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P42568 | Gene names | MLLT3, AF9, YEATS3 | |||
|
Domain Architecture |

|
|||||
| Description | Protein AF-9 (ALL1 fused gene from chromosome 9 protein) (Myeloid/lymphoid or mixed-lineage leukemia translocated to chromosome 3 protein) (YEATS domain-containing protein 3). | |||||
|
LR37A_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 8) | NC score | 0.061070 (rank : 6) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 720 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O60309, Q49A01, Q49A80, Q8NB33 | Gene names | LRRC37A, KIAA0563 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 37A. | |||||
|
LR37B_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 9) | NC score | 0.055545 (rank : 13) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96QE4, Q5YKG6 | Gene names | LRRC37B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 37B precursor (C66 SLIT-like testicular protein). | |||||
|
T3JAM_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 10) | NC score | 0.060758 (rank : 7) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 533 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9Y228, Q2YDB5, Q4VY06, Q7Z706 | Gene names | TRAF3IP3, T3JAM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TRAF3-interacting JNK-activating modulator (TRAF3-interacting protein 3). | |||||
|
K0460_HUMAN
|
||||||
| θ value | 0.125558 (rank : 11) | NC score | 0.056172 (rank : 12) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q5VT52, O75048, Q5VT53, Q6MZL4, Q86XD2 | Gene names | KIAA0460 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0460. | |||||
|
MPIP1_MOUSE
|
||||||
| θ value | 0.125558 (rank : 12) | NC score | 0.047114 (rank : 18) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P48964 | Gene names | Cdc25a, Cdc25m3 | |||
|
Domain Architecture |

|
|||||
| Description | M-phase inducer phosphatase 1 (EC 3.1.3.48) (Dual specificity phosphatase Cdc25A). | |||||
|
FA8_MOUSE
|
||||||
| θ value | 0.163984 (rank : 13) | NC score | 0.033699 (rank : 37) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q06194 | Gene names | F8, Cf8, F8c | |||
|
Domain Architecture |

|
|||||
| Description | Coagulation factor VIII precursor (Procoagulant component). | |||||
|
REPS2_HUMAN
|
||||||
| θ value | 0.163984 (rank : 14) | NC score | 0.045313 (rank : 21) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8NFH8, O43428, Q8NFI5 | Gene names | REPS2, POB1 | |||
|
Domain Architecture |

|
|||||
| Description | RalBP1-associated Eps domain-containing protein 2 (RalBP1-interacting protein 2) (Partner of RalBP1). | |||||
|
MARCS_HUMAN
|
||||||
| θ value | 0.21417 (rank : 15) | NC score | 0.056629 (rank : 11) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P29966, Q2LA83, Q5TDB7 | Gene names | MARCKS, MACS, PRKCSL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myristoylated alanine-rich C-kinase substrate (MARCKS) (Protein kinase C substrate, 80 kDa protein, light chain) (PKCSL) (80K-L protein). | |||||
|
PRG4_MOUSE
|
||||||
| θ value | 0.21417 (rank : 16) | NC score | 0.038400 (rank : 27) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 624 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9JM99, Q3UEL1, Q3V198 | Gene names | Prg4, Msf, Szp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
WDR22_MOUSE
|
||||||
| θ value | 0.21417 (rank : 17) | NC score | 0.047664 (rank : 17) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 302 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q80T85, Q80VT3, Q80ZW6, Q8BIP8, Q8BVK5, Q8K0Q6 | Gene names | Wdr22, Kiaa1824 | |||
|
Domain Architecture |
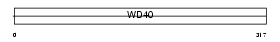
|
|||||
| Description | WD repeat protein 22. | |||||
|
APC_HUMAN
|
||||||
| θ value | 0.279714 (rank : 18) | NC score | 0.057525 (rank : 9) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 603 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P25054, Q15162, Q15163, Q93042 | Gene names | APC, DP2.5 | |||
|
Domain Architecture |

|
|||||
| Description | Adenomatous polyposis coli protein (Protein APC). | |||||
|
SRRM2_HUMAN
|
||||||
| θ value | 0.279714 (rank : 19) | NC score | 0.052969 (rank : 15) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
CO4A4_HUMAN
|
||||||
| θ value | 0.365318 (rank : 20) | NC score | 0.039696 (rank : 25) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 440 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P53420 | Gene names | COL4A4 | |||
|
Domain Architecture |
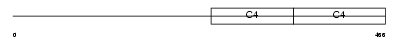
|
|||||
| Description | Collagen alpha-4(IV) chain precursor. | |||||
|
WDR22_HUMAN
|
||||||
| θ value | 0.365318 (rank : 21) | NC score | 0.045605 (rank : 20) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96JK2, O60559, Q8N3V3, Q8N3V5 | Gene names | WDR22, BCRG2, KIAA1824 | |||
|
Domain Architecture |
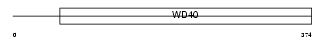
|
|||||
| Description | WD repeat protein 22 (Breakpoint cluster region protein 2) (BCRP2). | |||||
|
ANLN_MOUSE
|
||||||
| θ value | 0.47712 (rank : 22) | NC score | 0.043280 (rank : 23) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 297 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8K298, Q8BL79, Q8BLB3, Q8K2N0, Q9CUY0 | Gene names | Anln | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Actin-binding protein anillin. | |||||
|
CHD1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 23) | NC score | 0.028796 (rank : 52) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O14646 | Gene names | CHD1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
FOXP1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 24) | NC score | 0.023162 (rank : 64) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9H334, Q9H332, Q9H333, Q9P0R1 | Gene names | FOXP1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein P1. | |||||
|
CHD1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 25) | NC score | 0.027349 (rank : 58) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P40201 | Gene names | Chd1, Chd-1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
CO4A1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 26) | NC score | 0.036564 (rank : 30) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 450 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P02463 | Gene names | Col4a1 | |||
|
Domain Architecture |
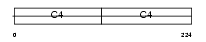
|
|||||
| Description | Collagen alpha-1(IV) chain precursor. | |||||
|
KAISO_HUMAN
|
||||||
| θ value | 1.06291 (rank : 27) | NC score | 0.007792 (rank : 95) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 840 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q86T24, O00319, Q7Z361, Q8IVP6, Q8N3P0 | Gene names | ZBTB33, KAISO, ZNF348 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcriptional regulator Kaiso (Zinc finger and BTB domain-containing protein 33). | |||||
|
BAZ2B_HUMAN
|
||||||
| θ value | 1.38821 (rank : 28) | NC score | 0.034666 (rank : 33) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 803 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9UIF8, Q96EA1, Q96SQ8, Q9P252, Q9Y4N8 | Gene names | BAZ2B, KIAA1476 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2B (hWALp4). | |||||
|
DSPP_MOUSE
|
||||||
| θ value | 1.38821 (rank : 29) | NC score | 0.057315 (rank : 10) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P97399, O70567 | Gene names | Dspp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor (Dentin matrix protein 3) (DMP-3) [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
GOGA2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 30) | NC score | 0.027634 (rank : 57) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 856 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q08379, Q9BRB0, Q9NYF9 | Gene names | GOLGA2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgin subfamily A member 2 (Cis-Golgi matrix protein GM130) (Gm130 autoantigen) (Golgin-95). | |||||
|
K0240_MOUSE
|
||||||
| θ value | 1.38821 (rank : 31) | NC score | 0.053503 (rank : 14) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8CHH5 | Gene names | Kiaa0240 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0240. | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 32) | NC score | 0.045055 (rank : 22) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
AP3B1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 33) | NC score | 0.022307 (rank : 69) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O00203, O00580, Q7Z393, Q9HD66 | Gene names | AP3B1, ADTB3A | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit beta-1 (Adapter-related protein complex 3 beta-1 subunit) (Beta3A-adaptin) (Adaptor protein complex AP-3 beta-1 subunit) (Clathrin assembly protein complex 3 beta-1 large chain). | |||||
|
APC_MOUSE
|
||||||
| θ value | 1.81305 (rank : 34) | NC score | 0.049266 (rank : 16) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 441 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q61315, Q62044 | Gene names | Apc | |||
|
Domain Architecture |

|
|||||
| Description | Adenomatous polyposis coli protein (Protein APC) (mAPC). | |||||
|
CO7A1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 35) | NC score | 0.024825 (rank : 60) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 807 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q02388, Q14054, Q16507 | Gene names | COL7A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(VII) chain precursor (Long-chain collagen) (LC collagen). | |||||
|
FOXP1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 36) | NC score | 0.020943 (rank : 71) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P58462 | Gene names | Foxp1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein P1 (Forkhead-related transcription factor 1). | |||||
|
K0240_HUMAN
|
||||||
| θ value | 1.81305 (rank : 37) | NC score | 0.046141 (rank : 19) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q6AI39, Q5TFZ3, Q92514 | Gene names | KIAA0240 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0240. | |||||
|
C1QC_MOUSE
|
||||||
| θ value | 2.36792 (rank : 38) | NC score | 0.028849 (rank : 51) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q02105 | Gene names | C1qc, C1qg | |||
|
Domain Architecture |
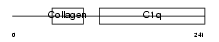
|
|||||
| Description | Complement C1q subcomponent subunit C precursor. | |||||
|
CO4A1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 39) | NC score | 0.033654 (rank : 38) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P02462 | Gene names | COL4A1 | |||
|
Domain Architecture |
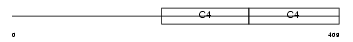
|
|||||
| Description | Collagen alpha-1(IV) chain precursor. | |||||
|
DGCR8_HUMAN
|
||||||
| θ value | 2.36792 (rank : 40) | NC score | 0.035850 (rank : 32) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8WYQ5, Q6DCB2, Q6MZE9, Q6Y2L0, Q96G39, Q96GP8, Q9H6L8, Q9H6T7, Q9NRW2 | Gene names | DGCR8, C22orf12, DGCRK6 | |||
|
Domain Architecture |
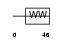
|
|||||
| Description | DGCR8 protein (DiGeorge syndrome critical region 8). | |||||
|
IF4B_HUMAN
|
||||||
| θ value | 2.36792 (rank : 41) | NC score | 0.030730 (rank : 45) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P23588 | Gene names | EIF4B | |||
|
Domain Architecture |
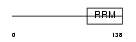
|
|||||
| Description | Eukaryotic translation initiation factor 4B (eIF-4B). | |||||
|
MUC5B_HUMAN
|
||||||
| θ value | 2.36792 (rank : 42) | NC score | 0.020275 (rank : 74) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1020 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9HC84, O00447, O00573, O14985, O15494, O95291, O95451, Q14881, Q99552, Q9UE28 | Gene names | MUC5B, MUC5 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-5B precursor (Mucin-5 subtype B, tracheobronchial) (High molecular weight salivary mucin MG1) (Sublingual gland mucin). | |||||
|
PO121_MOUSE
|
||||||
| θ value | 2.36792 (rank : 43) | NC score | 0.027670 (rank : 56) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 345 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8K3Z9, Q7TSH5 | Gene names | Pom121, Nup121 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear envelope pore membrane protein POM 121 (Pore membrane protein of 121 kDa). | |||||
|
RCOR2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 44) | NC score | 0.038182 (rank : 28) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8IZ40, Q96FP3 | Gene names | RCOR2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | REST corepressor 2. | |||||
|
RCOR2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 45) | NC score | 0.038121 (rank : 29) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8C796, Q9JMK4 | Gene names | Rcor2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | REST corepressor 2 (M-CoREST). | |||||
|
SCN5A_HUMAN
|
||||||
| θ value | 2.36792 (rank : 46) | NC score | 0.009876 (rank : 91) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14524 | Gene names | SCN5A | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 5 subunit alpha (Sodium channel protein type V subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.5) (Sodium channel protein, cardiac muscle alpha-subunit) (HH1). | |||||
|
TARA_MOUSE
|
||||||
| θ value | 2.36792 (rank : 47) | NC score | 0.030647 (rank : 47) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1133 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q99KW3, Q2PZW9, Q2Q402, Q2Q403, Q2Q404, Q6ZPK4, Q8C6T3, Q8CG90 | Gene names | Triobp, Kiaa1662, Tara | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
ECHB_MOUSE
|
||||||
| θ value | 3.0926 (rank : 48) | NC score | 0.023229 (rank : 63) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99JY0, Q3TEH9, Q8BJI5, Q8BJM0, Q8BK52 | Gene names | Hadhb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trifunctional enzyme subunit beta, mitochondrial precursor (TP-beta) [Includes: 3-ketoacyl-CoA thiolase (EC 2.3.1.16) (Acetyl-CoA acyltransferase) (Beta-ketothiolase)]. | |||||
|
HUWE1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 49) | NC score | 0.014407 (rank : 84) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q7TMY8, Q4G2Z1, Q5BMM7, Q6NS61, Q8BNJ7, Q8CFH2, Q8VD14, Q921M5, Q9R0P2 | Gene names | Huwe1, Kiaa0312, Ureb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT, UBA and WWE domain-containing protein 1 (EC 6.3.2.-) (E3 ubiquitin protein ligase URE-B1) (E3Histone). | |||||
|
NCOR1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 50) | NC score | 0.034404 (rank : 35) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 577 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O75376, Q9UPV5, Q9UQ18 | Gene names | NCOR1, KIAA1047 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 1 (N-CoR1) (N-CoR). | |||||
|
RIF1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 51) | NC score | 0.032303 (rank : 41) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q6PR54, Q6T8D9, Q7TPD8, Q8BTU2, Q8C408, Q9CS77 | Gene names | Rif1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Telomere-associated protein RIF1 (Rap1-interacting factor 1 homolog) (mRif1). | |||||
|
SRPK1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 52) | NC score | 0.012783 (rank : 86) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 706 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q96SB4, Q12890, Q5R364, Q5R365, Q8IY12 | Gene names | SRPK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SRPK1 (EC 2.7.11.1) (Serine/arginine- rich protein-specific kinase 1) (SR-protein-specific kinase 1) (SFRS protein kinase 1). | |||||
|
TNIK_HUMAN
|
||||||
| θ value | 3.0926 (rank : 53) | NC score | 0.004074 (rank : 102) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1570 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9UKE5, O60298, Q8WUY7, Q9UKD8, Q9UKD9, Q9UKE0, Q9UKE1, Q9UKE2, Q9UKE3, Q9UKE4 | Gene names | TNIK, KIAA0551 | |||
|
Domain Architecture |

|
|||||
| Description | TRAF2 and NCK-interacting protein kinase (EC 2.7.11.1). | |||||
|
UBP32_HUMAN
|
||||||
| θ value | 3.0926 (rank : 54) | NC score | 0.018471 (rank : 76) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8NFA0, Q9BX85, Q9Y591 | Gene names | USP32, USP10 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 32 (EC 3.1.2.15) (Ubiquitin thioesterase 32) (Ubiquitin-specific-processing protease 32) (Deubiquitinating enzyme 32) (NY-REN-60 antigen). | |||||
|
CADH1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 55) | NC score | 0.004218 (rank : 101) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P09803, Q61377 | Gene names | Cdh1 | |||
|
Domain Architecture |

|
|||||
| Description | Epithelial-cadherin precursor (E-cadherin) (Uvomorulin) (Cadherin-1) (ARC-1) (CD324 antigen) [Contains: E-Cad/CTF1; E-Cad/CTF2; E- Cad/CTF3]. | |||||
|
CO4A2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 56) | NC score | 0.031016 (rank : 44) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 454 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P08572 | Gene names | COL4A2 | |||
|
Domain Architecture |
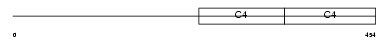
|
|||||
| Description | Collagen alpha-2(IV) chain precursor. | |||||
|
KCNA6_MOUSE
|
||||||
| θ value | 4.03905 (rank : 57) | NC score | 0.006948 (rank : 97) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q61923 | Gene names | Kcna6 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 6 (Voltage-gated potassium channel subunit Kv1.6) (MK1.6). | |||||
|
M3K2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 58) | NC score | 0.002846 (rank : 105) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 846 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q61083 | Gene names | Map3k2, Mekk2 | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase kinase kinase 2 (EC 2.7.11.25) (MAPK/ERK kinase kinase 2) (MEK kinase 2) (MEKK 2). | |||||
|
PRDM9_HUMAN
|
||||||
| θ value | 4.03905 (rank : 59) | NC score | 0.002018 (rank : 108) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 750 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9NQV7 | Gene names | PRDM9, PFM6 | |||
|
Domain Architecture |
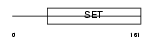
|
|||||
| Description | PR domain zinc finger protein 9 (PR domain-containing protein 9). | |||||
|
TGFR2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 60) | NC score | 0.005784 (rank : 98) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 748 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q62312, Q63947 | Gene names | Tgfbr2 | |||
|
Domain Architecture |

|
|||||
| Description | TGF-beta receptor type-2 precursor (EC 2.7.11.30) (TGF-beta receptor type II) (TGFR-2) (TGF-beta type II receptor). | |||||
|
UBE4B_HUMAN
|
||||||
| θ value | 4.03905 (rank : 61) | NC score | 0.022311 (rank : 68) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O95155, O75169, O95338, Q5SZ12, Q5SZ16, Q96QD4, Q9BYI7 | Gene names | UBE4B, HDNB1, KIAA0684, UFD2 | |||
|
Domain Architecture |
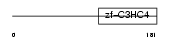
|
|||||
| Description | Ubiquitin conjugation factor E4 B (Ubiquitin fusion degradation protein 2) (Homozygously deleted in neuroblastoma 1). | |||||
|
CO4A3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 62) | NC score | 0.031750 (rank : 43) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 421 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q01955, Q9BQT2 | Gene names | COL4A3 | |||
|
Domain Architecture |
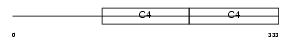
|
|||||
| Description | Collagen alpha-3(IV) chain precursor (Goodpasture antigen). | |||||
|
CO4A5_HUMAN
|
||||||
| θ value | 5.27518 (rank : 63) | NC score | 0.038900 (rank : 26) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 383 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P29400, Q16006, Q16126, Q6LD84 | Gene names | COL4A5 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-5(IV) chain precursor. | |||||
|
DACH1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 64) | NC score | 0.017739 (rank : 79) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9QYB2, O88716, Q8BPQ0, Q8C8D7, Q9QY47, Q9R218, Q9Z0Y5 | Gene names | Dach1, Dach | |||
|
Domain Architecture |

|
|||||
| Description | Dachshund homolog 1 (Dach1). | |||||
|
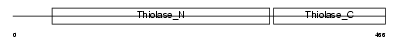
ECHB_HUMAN
|
||||||
| θ value | 5.27518 (rank : 65) | NC score | 0.020864 (rank : 72) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P55084, O14969, Q53TA6, Q96C77, Q9H3F5 | Gene names | HADHB | |||
|
Domain Architecture |
|
|||||
| Description | Trifunctional enzyme subunit beta, mitochondrial precursor (TP-beta) [Includes: 3-ketoacyl-CoA thiolase (EC 2.3.1.16) (Acetyl-CoA acyltransferase) (Beta-ketothiolase)]. | |||||
|
LIMA1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 66) | NC score | 0.015468 (rank : 81) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 309 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9UHB6, Q2TAN7, Q9BVF2, Q9H8J1, Q9HBN5, Q9NX96, Q9NXC3, Q9NXU6, Q9P0H8, Q9UHB5 | Gene names | LIMA1, EPLIN, SREBP3 | |||
|
Domain Architecture |

|
|||||
| Description | LIM domain and actin-binding protein 1 (Epithelial protein lost in neoplasm). | |||||
|
NCOR2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 67) | NC score | 0.026823 (rank : 59) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 790 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9WU42, Q9WU43, Q9WUC1 | Gene names | Ncor2, Smrt | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC). | |||||
|
PHF2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 68) | NC score | 0.013029 (rank : 85) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O75151, Q8N3K2, Q9Y6N4 | Gene names | PHF2, KIAA0662 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 2 (GRC5). | |||||
|
UBF1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 69) | NC score | 0.015356 (rank : 83) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 415 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P17480 | Gene names | UBTF, UBF, UBF1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar transcription factor 1 (Upstream-binding factor 1) (UBF-1) (Autoantigen NOR-90). | |||||
|
VATC_HUMAN
|
||||||
| θ value | 5.27518 (rank : 70) | NC score | 0.030652 (rank : 46) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P21283 | Gene names | ATP6V1C1, ATP6C, ATP6D, VATC | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar ATP synthase subunit C (EC 3.6.3.14) (V-ATPase C subunit) (Vacuolar proton pump C subunit). | |||||
|
VATC_MOUSE
|
||||||
| θ value | 5.27518 (rank : 71) | NC score | 0.030591 (rank : 48) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Z1G3 | Gene names | Atp6v1c1, Atp6c, Vatc | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar ATP synthase subunit C (EC 3.6.3.14) (V-ATPase C subunit) (Vacuolar proton pump C subunit). | |||||
|
WNK4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 72) | NC score | 0.004071 (rank : 103) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1043 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q96J92, Q8N8X3, Q8N8Z2, Q96DT8, Q9BYS5 | Gene names | WNK4, PRKWNK4 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK4 (EC 2.7.11.1) (Protein kinase with no lysine 4) (Protein kinase, lysine-deficient 4). | |||||
|
ZCH14_HUMAN
|
||||||
| θ value | 5.27518 (rank : 73) | NC score | 0.022430 (rank : 67) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8WYQ9, O60324, Q9UFP0 | Gene names | ZCCHC14, KIAA0579 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 14 (BDG-29). | |||||
|
CO4A6_HUMAN
|
||||||
| θ value | 6.88961 (rank : 74) | NC score | 0.030433 (rank : 49) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q14031, Q12823, Q14053, Q5JYH6, Q9NQM5, Q9NTX3, Q9UJ76, Q9UMG6, Q9Y4L4 | Gene names | COL4A6 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-6(IV) chain precursor. | |||||
|
COAA1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 75) | NC score | 0.028621 (rank : 53) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 316 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q03692 | Gene names | COL10A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(X) chain precursor. | |||||
|
FYCO1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 76) | NC score | 0.009853 (rank : 92) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1059 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8VDC1, Q3V385, Q7TMT0, Q8BJN2 | Gene names | Fyco1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE and coiled-coil domain-containing protein 1. | |||||
|
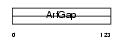
GIT2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 77) | NC score | 0.007531 (rank : 96) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9JLQ2 | Gene names | Git2 | |||
|
Domain Architecture |
|
|||||
| Description | ARF GTPase-activating protein GIT2 (G protein-coupled receptor kinase- interactor 2) (Cool-interacting tyrosine-phosphorylated protein 2) (CAT2) (CAT-2). | |||||
|
MACF1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 78) | NC score | 0.011418 (rank : 89) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1341 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9UPN3, O75053, Q5VW20, Q8WXY2, Q9H540, Q9UKP0, Q9ULG9 | Gene names | MACF1, ABP620, ACF7, KIAA0465, KIAA1251 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-actin cross-linking factor 1, isoforms 1/2/3/5 (Actin cross-linking family protein 7) (Macrophin-1) (Trabeculin-alpha) (620 kDa actin-binding protein) (ABP620). | |||||
|
MACF4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 79) | NC score | 0.015363 (rank : 82) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1321 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q96PK2, Q8WXY1 | Gene names | MACF1, ABP620, ACF7, KIAA0465, KIAA1251 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-actin cross-linking factor 1, isoform 4. | |||||
|
MACOI_HUMAN
|
||||||
| θ value | 6.88961 (rank : 80) | NC score | 0.011399 (rank : 90) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8N5G2, Q2TLX5, Q2TLX6, Q9NVG6 | Gene names | TMEM57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Macoilin (Transmembrane protein 57). | |||||
|
MAP1B_MOUSE
|
||||||
| θ value | 6.88961 (rank : 81) | NC score | 0.024159 (rank : 62) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 977 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P14873 | Gene names | Map1b, Mtap1b, Mtap5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) (MAP1.2) (MAP1(X)) [Contains: MAP1 light chain LC1]. | |||||
|
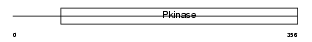
MK10_HUMAN
|
||||||
| θ value | 6.88961 (rank : 82) | NC score | 0.002674 (rank : 106) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 836 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P53779, Q15707 | Gene names | MAPK10, JNK3, JNK3A, PRKM10 | |||
|
Domain Architecture |
|
|||||
| Description | Mitogen-activated protein kinase 10 (EC 2.7.11.24) (Stress-activated protein kinase JNK3) (c-Jun N-terminal kinase 3) (MAP kinase p49 3F12). | |||||
|
NCOA2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 83) | NC score | 0.020486 (rank : 73) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q61026, O09001, P97759 | Gene names | Ncoa2, Grip1, Tif2 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 2 (NCoA-2) (Transcriptional intermediary factor 2) (Glucocorticoid receptor-interacting protein 1) (GRIP-1). | |||||
|
NIPBL_HUMAN
|
||||||
| θ value | 6.88961 (rank : 84) | NC score | 0.032366 (rank : 39) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
NOL8_MOUSE
|
||||||
| θ value | 6.88961 (rank : 85) | NC score | 0.036041 (rank : 31) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q3UHX0, Q504M4, Q8CDJ7, Q9CUR0 | Gene names | Nol8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleolar protein 8. | |||||
|
NPBL_MOUSE
|
||||||
| θ value | 6.88961 (rank : 86) | NC score | 0.034266 (rank : 36) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
NU214_HUMAN
|
||||||
| θ value | 6.88961 (rank : 87) | NC score | 0.028914 (rank : 50) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 657 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P35658 | Gene names | NUP214, CAIN, CAN | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup214 (Nucleoporin Nup214) (214 kDa nucleoporin) (CAN protein). | |||||
|
PEX19_MOUSE
|
||||||
| θ value | 6.88961 (rank : 88) | NC score | 0.019697 (rank : 75) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8VCI5, Q8CEE1, Q921H0, Q9CZC1, Q9QUQ1 | Gene names | Pex19, Pxf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisomal biogenesis factor 19 (Peroxin-19) (Peroxisomal farnesylated protein) (PxF). | |||||
|
PODXL_HUMAN
|
||||||
| θ value | 6.88961 (rank : 89) | NC score | 0.031883 (rank : 42) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O00592 | Gene names | PODXL, PCLP, PCLP1 | |||
|
Domain Architecture |

|
|||||
| Description | Podocalyxin-like protein 1 precursor. | |||||
|
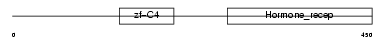
RXRG_HUMAN
|
||||||
| θ value | 6.88961 (rank : 90) | NC score | 0.003888 (rank : 104) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P48443 | Gene names | RXRG, NR2B3 | |||
|
Domain Architecture |
|
|||||
| Description | Retinoic acid receptor RXR-gamma (Retinoid X receptor gamma). | |||||
|
S6OS1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 91) | NC score | 0.016728 (rank : 80) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9CTN5, Q9D5F5, Q9D5Y9 | Gene names | Six6os1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein SIX6OS1 (Six6 opposite strand transcript 1). | |||||
|
SCEL_MOUSE
|
||||||
| θ value | 6.88961 (rank : 92) | NC score | 0.034574 (rank : 34) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9EQG3, Q9CTT9 | Gene names | Scel | |||
|
Domain Architecture |

|
|||||
| Description | Sciellin. | |||||
|
SFR17_HUMAN
|
||||||
| θ value | 6.88961 (rank : 93) | NC score | 0.024562 (rank : 61) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8TF01, Q5T064, Q6P2B4, Q6P2N4, Q6PJ93, Q6PK36, Q7Z640, Q8TF00, Q96K10, Q96SM5, Q9P076, Q9P0C0, Q9Y4N3 | Gene names | SRRP130, C6orf111 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, arginine/serine-rich 130 (Serine-arginine-rich- splicing regulatory protein 130) (SRrp130) (SR-rich protein) (SR- related protein). | |||||
|
AKAP6_HUMAN
|
||||||
| θ value | 8.99809 (rank : 94) | NC score | 0.032344 (rank : 40) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 263 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q13023, O15028 | Gene names | AKAP6, AKAP100, KIAA0311 | |||
|
Domain Architecture |

|
|||||
| Description | A-kinase anchor protein 6 (Protein kinase A-anchoring protein 6) (PRKA6) (A-kinase anchor protein 100 kDa) (AKAP 100) (mAKAP). | |||||
|
AP4E1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 95) | NC score | 0.009733 (rank : 93) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UPM8, Q9Y588 | Gene names | AP4E1 | |||
|
Domain Architecture |

|
|||||
| Description | AP-4 complex subunit epsilon-1 (Adapter-related protein complex 4 epsilon-1 subunit) (Epsilon subunit of AP-4) (AP-4 adapter complex epsilon subunit). | |||||
|
ARHGC_HUMAN
|
||||||
| θ value | 8.99809 (rank : 96) | NC score | 0.005717 (rank : 99) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NZN5, O15086 | Gene names | ARHGEF12, KIAA0382, LARG | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 12 (Leukemia-associated RhoGEF). | |||||
|
CO5A1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 97) | NC score | 0.027971 (rank : 55) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 434 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P20908, Q15094, Q5SUX4 | Gene names | COL5A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(V) chain precursor. | |||||
|
CO5A1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 98) | NC score | 0.027985 (rank : 54) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O88207 | Gene names | Col5a1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(V) chain precursor. | |||||
|
DJBP_MOUSE
|
||||||
| θ value | 8.99809 (rank : 99) | NC score | 0.011936 (rank : 87) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6P1E8, Q5DTV8, Q6PGK8, Q9D2D8 | Gene names | Djbp, Kiaa1672 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DJ-1-binding protein. | |||||
|
MAGC1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 100) | NC score | 0.011481 (rank : 88) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 431 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O60732, O75451, Q8TCV4 | Gene names | MAGEC1 | |||
|
Domain Architecture |

|
|||||
| Description | Melanoma-associated antigen C1 (MAGE-C1 antigen) (Cancer-testis antigen CT7). | |||||
|
MINT_MOUSE
|
||||||
| θ value | 8.99809 (rank : 101) | NC score | 0.018228 (rank : 77) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
MK10_MOUSE
|
||||||
| θ value | 8.99809 (rank : 102) | NC score | 0.002562 (rank : 107) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 849 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q61831, Q9R0U6 | Gene names | Mapk10, Jnk3, Prkm10, Serk2 | |||
|
Domain Architecture |
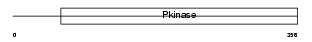
|
|||||
| Description | Mitogen-activated protein kinase 10 (EC 2.7.11.24) (Stress-activated protein kinase JNK3) (c-Jun N-terminal kinase 3) (MAP kinase p49 3F12). | |||||
|
MTF1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 103) | NC score | 0.000551 (rank : 110) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 814 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q14872, Q96CB1 | Gene names | MTF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metal-regulatory transcription factor 1 (Transcription factor MTF-1) (MRE-binding transcription factor). | |||||
|
NCOA2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 104) | NC score | 0.017984 (rank : 78) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q15596 | Gene names | NCOA2, TIF2 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 2 (NCoA-2) (Transcriptional intermediary factor 2). | |||||
|
NFH_MOUSE
|
||||||
| θ value | 8.99809 (rank : 105) | NC score | 0.023027 (rank : 65) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
OSTP_MOUSE
|
||||||
| θ value | 8.99809 (rank : 106) | NC score | 0.022733 (rank : 66) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P10923, P19008 | Gene names | Spp1, Eta-1, Op, Spp-1 | |||
|
Domain Architecture |

|
|||||
| Description | Osteopontin precursor (Bone sialoprotein-1) (Secreted phosphoprotein 1) (SPP-1) (Minopontin) (Early T-lymphocyte activation 1 protein) (2AR) (Calcium oxalate crystal growth inhibitor protein). | |||||
|
PLCB1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 107) | NC score | 0.005654 (rank : 100) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 531 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Z1B3, Q62075, Q8K5A5, Q8K5A6, Q9Z0E5, Q9Z2T5 | Gene names | Plcb1, Plcb | |||
|
Domain Architecture |

|
|||||
| Description | 1-phosphatidylinositol-4,5-bisphosphate phosphodiesterase beta 1 (EC 3.1.4.11) (Phosphoinositide phospholipase C) (Phospholipase C- beta-1) (PLC-beta-1) (PLC-I) (PLC-154). | |||||
|
SDC3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 108) | NC score | 0.021567 (rank : 70) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q64519, Q3UDD9, Q6ZQA4, Q7TQD4 | Gene names | Sdc3, Kiaa0468 | |||
|
Domain Architecture |
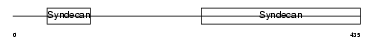
|
|||||
| Description | Syndecan-3 precursor (SYND3). | |||||
|
SP2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 109) | NC score | 0.000878 (rank : 109) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 747 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q02086 | Gene names | SP2, KIAA0048 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor Sp2. | |||||
|
SYN1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 110) | NC score | 0.009600 (rank : 94) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 363 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O88935, Q62279, Q8QZT8 | Gene names | Syn1, Syn-1 | |||
|
Domain Architecture |
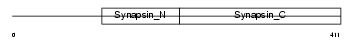
|
|||||
| Description | Synapsin-1 (Synapsin I). | |||||
|
NRIP1_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 110 | |
| SwissProt Accessions | Q8CBD1, Q8C3Y8, Q9Z2K2 | Gene names | Nrip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor-interacting protein 1 (Nuclear factor RIP140) (Receptor-interacting protein 140). | |||||
|
NRIP1_HUMAN
|
||||||
| NC score | 0.963568 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P48552, Q8IWE8 | Gene names | NRIP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear receptor-interacting protein 1 (Nuclear factor RIP140) (Receptor-interacting protein 140). | |||||
|
DSPP_HUMAN
|
||||||
| NC score | 0.098988 (rank : 3) | θ value | 0.0193708 (rank : 5) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9NZW4, O95815 | Gene names | DSPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
SYNPO_HUMAN
|
||||||
| NC score | 0.081441 (rank : 4) | θ value | 0.00665767 (rank : 4) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8N3V7, O15271, Q9UPX1 | Gene names | SYNPO, KIAA1029 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptopodin. | |||||
|
AF9_HUMAN
|
||||||
| NC score | 0.072095 (rank : 5) | θ value | 0.0736092 (rank : 7) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P42568 | Gene names | MLLT3, AF9, YEATS3 | |||
|
Domain Architecture |

|
|||||
| Description | Protein AF-9 (ALL1 fused gene from chromosome 9 protein) (Myeloid/lymphoid or mixed-lineage leukemia translocated to chromosome 3 protein) (YEATS domain-containing protein 3). | |||||
|
LR37A_HUMAN
|
||||||
| NC score | 0.061070 (rank : 6) | θ value | 0.0736092 (rank : 8) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 720 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O60309, Q49A01, Q49A80, Q8NB33 | Gene names | LRRC37A, KIAA0563 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 37A. | |||||
|
T3JAM_HUMAN
|
||||||
| NC score | 0.060758 (rank : 7) | θ value | 0.0736092 (rank : 10) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 533 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9Y228, Q2YDB5, Q4VY06, Q7Z706 | Gene names | TRAF3IP3, T3JAM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TRAF3-interacting JNK-activating modulator (TRAF3-interacting protein 3). | |||||
|
SRBS1_MOUSE
|
||||||
| NC score | 0.057960 (rank : 8) | θ value | 0.00390308 (rank : 3) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q62417, Q80TF8, Q8BZI3, Q8K3Y2, Q921F8, Q9Z0Z8, Q9Z0Z9 | Gene names | Sorbs1, Kiaa1296, Sh3d5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sorbin and SH3 domain-containing protein 1 (Ponsin) (c-Cbl-associated protein) (CAP) (SH3 domain protein 5) (SH3P12). | |||||
|
APC_HUMAN
|
||||||
| NC score | 0.057525 (rank : 9) | θ value | 0.279714 (rank : 18) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 603 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P25054, Q15162, Q15163, Q93042 | Gene names | APC, DP2.5 | |||
|
Domain Architecture |

|
|||||
| Description | Adenomatous polyposis coli protein (Protein APC). | |||||
|
DSPP_MOUSE
|
||||||
| NC score | 0.057315 (rank : 10) | θ value | 1.38821 (rank : 29) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P97399, O70567 | Gene names | Dspp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor (Dentin matrix protein 3) (DMP-3) [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
MARCS_HUMAN
|
||||||
| NC score | 0.056629 (rank : 11) | θ value | 0.21417 (rank : 15) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P29966, Q2LA83, Q5TDB7 | Gene names | MARCKS, MACS, PRKCSL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myristoylated alanine-rich C-kinase substrate (MARCKS) (Protein kinase C substrate, 80 kDa protein, light chain) (PKCSL) (80K-L protein). | |||||
|
K0460_HUMAN
|
||||||
| NC score | 0.056172 (rank : 12) | θ value | 0.125558 (rank : 11) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q5VT52, O75048, Q5VT53, Q6MZL4, Q86XD2 | Gene names | KIAA0460 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0460. | |||||
|
LR37B_HUMAN
|
||||||
| NC score | 0.055545 (rank : 13) | θ value | 0.0736092 (rank : 9) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96QE4, Q5YKG6 | Gene names | LRRC37B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 37B precursor (C66 SLIT-like testicular protein). | |||||
|
K0240_MOUSE
|
||||||
| NC score | 0.053503 (rank : 14) | θ value | 1.38821 (rank : 31) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8CHH5 | Gene names | Kiaa0240 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0240. | |||||
|
SRRM2_HUMAN
|
||||||
| NC score | 0.052969 (rank : 15) | θ value | 0.279714 (rank : 19) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
APC_MOUSE
|
||||||
| NC score | 0.049266 (rank : 16) | θ value | 1.81305 (rank : 34) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 441 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q61315, Q62044 | Gene names | Apc | |||
|
Domain Architecture |

|
|||||
| Description | Adenomatous polyposis coli protein (Protein APC) (mAPC). | |||||
|
WDR22_MOUSE
|
||||||
| NC score | 0.047664 (rank : 17) | θ value | 0.21417 (rank : 17) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 302 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q80T85, Q80VT3, Q80ZW6, Q8BIP8, Q8BVK5, Q8K0Q6 | Gene names | Wdr22, Kiaa1824 | |||
|
Domain Architecture |

|
|||||
| Description | WD repeat protein 22. | |||||
|
MPIP1_MOUSE
|
||||||
| NC score | 0.047114 (rank : 18) | θ value | 0.125558 (rank : 12) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P48964 | Gene names | Cdc25a, Cdc25m3 | |||
|
Domain Architecture |

|
|||||
| Description | M-phase inducer phosphatase 1 (EC 3.1.3.48) (Dual specificity phosphatase Cdc25A). | |||||
|
K0240_HUMAN
|
||||||
| NC score | 0.046141 (rank : 19) | θ value | 1.81305 (rank : 37) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q6AI39, Q5TFZ3, Q92514 | Gene names | KIAA0240 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0240. | |||||
|
WDR22_HUMAN
|
||||||
| NC score | 0.045605 (rank : 20) | θ value | 0.365318 (rank : 21) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 328 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96JK2, O60559, Q8N3V3, Q8N3V5 | Gene names | WDR22, BCRG2, KIAA1824 | |||
|
Domain Architecture |

|
|||||
| Description | WD repeat protein 22 (Breakpoint cluster region protein 2) (BCRP2). | |||||
|
REPS2_HUMAN
|
||||||
| NC score | 0.045313 (rank : 21) | θ value | 0.163984 (rank : 14) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8NFH8, O43428, Q8NFI5 | Gene names | REPS2, POB1 | |||
|
Domain Architecture |

|
|||||
| Description | RalBP1-associated Eps domain-containing protein 2 (RalBP1-interacting protein 2) (Partner of RalBP1). | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.045055 (rank : 22) | θ value | 1.38821 (rank : 32) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
ANLN_MOUSE
|
||||||
| NC score | 0.043280 (rank : 23) | θ value | 0.47712 (rank : 22) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 297 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8K298, Q8BL79, Q8BLB3, Q8K2N0, Q9CUY0 | Gene names | Anln | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Actin-binding protein anillin. | |||||
|
MLL2_HUMAN
|
||||||
| NC score | 0.043241 (rank : 24) | θ value | 0.0431538 (rank : 6) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
CO4A4_HUMAN
|
||||||
| NC score | 0.039696 (rank : 25) | θ value | 0.365318 (rank : 20) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 440 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P53420 | Gene names | COL4A4 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-4(IV) chain precursor. | |||||
|
CO4A5_HUMAN
|
||||||
| NC score | 0.038900 (rank : 26) | θ value | 5.27518 (rank : 63) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 383 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P29400, Q16006, Q16126, Q6LD84 | Gene names | COL4A5 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-5(IV) chain precursor. | |||||
|
PRG4_MOUSE
|
||||||
| NC score | 0.038400 (rank : 27) | θ value | 0.21417 (rank : 16) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 624 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9JM99, Q3UEL1, Q3V198 | Gene names | Prg4, Msf, Szp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
RCOR2_HUMAN
|
||||||
| NC score | 0.038182 (rank : 28) | θ value | 2.36792 (rank : 44) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8IZ40, Q96FP3 | Gene names | RCOR2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | REST corepressor 2. | |||||
|
RCOR2_MOUSE
|
||||||
| NC score | 0.038121 (rank : 29) | θ value | 2.36792 (rank : 45) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8C796, Q9JMK4 | Gene names | Rcor2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | REST corepressor 2 (M-CoREST). | |||||
|
CO4A1_MOUSE
|
||||||
| NC score | 0.036564 (rank : 30) | θ value | 1.06291 (rank : 26) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 450 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P02463 | Gene names | Col4a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(IV) chain precursor. | |||||
|
NOL8_MOUSE
|
||||||
| NC score | 0.036041 (rank : 31) | θ value | 6.88961 (rank : 85) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q3UHX0, Q504M4, Q8CDJ7, Q9CUR0 | Gene names | Nol8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleolar protein 8. | |||||
|
DGCR8_HUMAN
|
||||||
| NC score | 0.035850 (rank : 32) | θ value | 2.36792 (rank : 40) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8WYQ5, Q6DCB2, Q6MZE9, Q6Y2L0, Q96G39, Q96GP8, Q9H6L8, Q9H6T7, Q9NRW2 | Gene names | DGCR8, C22orf12, DGCRK6 | |||
|
Domain Architecture |

|
|||||
| Description | DGCR8 protein (DiGeorge syndrome critical region 8). | |||||
|
BAZ2B_HUMAN
|
||||||
| NC score | 0.034666 (rank : 33) | θ value | 1.38821 (rank : 28) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 803 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9UIF8, Q96EA1, Q96SQ8, Q9P252, Q9Y4N8 | Gene names | BAZ2B, KIAA1476 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2B (hWALp4). | |||||
|
SCEL_MOUSE
|
||||||
| NC score | 0.034574 (rank : 34) | θ value | 6.88961 (rank : 92) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9EQG3, Q9CTT9 | Gene names | Scel | |||
|
Domain Architecture |

|
|||||
| Description | Sciellin. | |||||
|
NCOR1_HUMAN
|
||||||
| NC score | 0.034404 (rank : 35) | θ value | 3.0926 (rank : 50) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 577 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | O75376, Q9UPV5, Q9UQ18 | Gene names | NCOR1, KIAA1047 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 1 (N-CoR1) (N-CoR). | |||||
|
NPBL_MOUSE
|
||||||
| NC score | 0.034266 (rank : 36) | θ value | 6.88961 (rank : 86) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
FA8_MOUSE
|
||||||
| NC score | 0.033699 (rank : 37) | θ value | 0.163984 (rank : 13) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q06194 | Gene names | F8, Cf8, F8c | |||
|
Domain Architecture |

|
|||||
| Description | Coagulation factor VIII precursor (Procoagulant component). | |||||
|
CO4A1_HUMAN
|
||||||
| NC score | 0.033654 (rank : 38) | θ value | 2.36792 (rank : 39) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P02462 | Gene names | COL4A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(IV) chain precursor. | |||||
|
NIPBL_HUMAN
|
||||||
| NC score | 0.032366 (rank : 39) | θ value | 6.88961 (rank : 84) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
AKAP6_HUMAN
|
||||||
| NC score | 0.032344 (rank : 40) | θ value | 8.99809 (rank : 94) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 263 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q13023, O15028 | Gene names | AKAP6, AKAP100, KIAA0311 | |||
|
Domain Architecture |

|
|||||
| Description | A-kinase anchor protein 6 (Protein kinase A-anchoring protein 6) (PRKA6) (A-kinase anchor protein 100 kDa) (AKAP 100) (mAKAP). | |||||
|
RIF1_MOUSE
|
||||||
| NC score | 0.032303 (rank : 41) | θ value | 3.0926 (rank : 51) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q6PR54, Q6T8D9, Q7TPD8, Q8BTU2, Q8C408, Q9CS77 | Gene names | Rif1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Telomere-associated protein RIF1 (Rap1-interacting factor 1 homolog) (mRif1). | |||||
|
PODXL_HUMAN
|
||||||
| NC score | 0.031883 (rank : 42) | θ value | 6.88961 (rank : 89) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O00592 | Gene names | PODXL, PCLP, PCLP1 | |||
|
Domain Architecture |

|
|||||
| Description | Podocalyxin-like protein 1 precursor. | |||||
|
CO4A3_HUMAN
|
||||||
| NC score | 0.031750 (rank : 43) | θ value | 5.27518 (rank : 62) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 421 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q01955, Q9BQT2 | Gene names | COL4A3 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-3(IV) chain precursor (Goodpasture antigen). | |||||
|
CO4A2_HUMAN
|
||||||
| NC score | 0.031016 (rank : 44) | θ value | 4.03905 (rank : 56) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 454 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P08572 | Gene names | COL4A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(IV) chain precursor. | |||||
|
IF4B_HUMAN
|
||||||
| NC score | 0.030730 (rank : 45) | θ value | 2.36792 (rank : 41) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 417 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P23588 | Gene names | EIF4B | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 4B (eIF-4B). | |||||
|
VATC_HUMAN
|
||||||
| NC score | 0.030652 (rank : 46) | θ value | 5.27518 (rank : 70) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P21283 | Gene names | ATP6V1C1, ATP6C, ATP6D, VATC | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar ATP synthase subunit C (EC 3.6.3.14) (V-ATPase C subunit) (Vacuolar proton pump C subunit). | |||||
|
TARA_MOUSE
|
||||||
| NC score | 0.030647 (rank : 47) | θ value | 2.36792 (rank : 47) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1133 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q99KW3, Q2PZW9, Q2Q402, Q2Q403, Q2Q404, Q6ZPK4, Q8C6T3, Q8CG90 | Gene names | Triobp, Kiaa1662, Tara | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
VATC_MOUSE
|
||||||
| NC score | 0.030591 (rank : 48) | θ value | 5.27518 (rank : 71) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Z1G3 | Gene names | Atp6v1c1, Atp6c, Vatc | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar ATP synthase subunit C (EC 3.6.3.14) (V-ATPase C subunit) (Vacuolar proton pump C subunit). | |||||
|
CO4A6_HUMAN
|
||||||
| NC score | 0.030433 (rank : 49) | θ value | 6.88961 (rank : 74) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q14031, Q12823, Q14053, Q5JYH6, Q9NQM5, Q9NTX3, Q9UJ76, Q9UMG6, Q9Y4L4 | Gene names | COL4A6 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-6(IV) chain precursor. | |||||
|
NU214_HUMAN
|
||||||
| NC score | 0.028914 (rank : 50) | θ value | 6.88961 (rank : 87) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 657 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P35658 | Gene names | NUP214, CAIN, CAN | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup214 (Nucleoporin Nup214) (214 kDa nucleoporin) (CAN protein). | |||||
|
C1QC_MOUSE
|
||||||
| NC score | 0.028849 (rank : 51) | θ value | 2.36792 (rank : 38) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q02105 | Gene names | C1qc, C1qg | |||
|
Domain Architecture |

|
|||||
| Description | Complement C1q subcomponent subunit C precursor. | |||||
|
CHD1_HUMAN
|
||||||
| NC score | 0.028796 (rank : 52) | θ value | 0.813845 (rank : 23) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O14646 | Gene names | CHD1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
COAA1_HUMAN
|
||||||
| NC score | 0.028621 (rank : 53) | θ value | 6.88961 (rank : 75) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 316 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q03692 | Gene names | COL10A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(X) chain precursor. | |||||
|
CO5A1_MOUSE
|
||||||
| NC score | 0.027985 (rank : 54) | θ value | 8.99809 (rank : 98) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O88207 | Gene names | Col5a1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(V) chain precursor. | |||||
|
CO5A1_HUMAN
|
||||||
| NC score | 0.027971 (rank : 55) | θ value | 8.99809 (rank : 97) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 434 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P20908, Q15094, Q5SUX4 | Gene names | COL5A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(V) chain precursor. | |||||
|
PO121_MOUSE
|
||||||
| NC score | 0.027670 (rank : 56) | θ value | 2.36792 (rank : 43) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 345 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8K3Z9, Q7TSH5 | Gene names | Pom121, Nup121 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear envelope pore membrane protein POM 121 (Pore membrane protein of 121 kDa). | |||||
|
GOGA2_HUMAN
|
||||||
| NC score | 0.027634 (rank : 57) | θ value | 1.38821 (rank : 30) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 856 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q08379, Q9BRB0, Q9NYF9 | Gene names | GOLGA2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgin subfamily A member 2 (Cis-Golgi matrix protein GM130) (Gm130 autoantigen) (Golgin-95). | |||||
|
CHD1_MOUSE
|
||||||
| NC score | 0.027349 (rank : 58) | θ value | 1.06291 (rank : 25) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P40201 | Gene names | Chd1, Chd-1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
NCOR2_MOUSE
|
||||||
| NC score | 0.026823 (rank : 59) | θ value | 5.27518 (rank : 67) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 790 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9WU42, Q9WU43, Q9WUC1 | Gene names | Ncor2, Smrt | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 2 (N-CoR2) (Silencing mediator of retinoic acid and thyroid hormone receptor) (SMRT) (SMRTe) (Thyroid-, retinoic-acid-receptor-associated corepressor) (T3 receptor- associating factor) (TRAC). | |||||
|
CO7A1_HUMAN
|
||||||
| NC score | 0.024825 (rank : 60) | θ value | 1.81305 (rank : 35) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 807 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q02388, Q14054, Q16507 | Gene names | COL7A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(VII) chain precursor (Long-chain collagen) (LC collagen). | |||||
|
SFR17_HUMAN
|
||||||
| NC score | 0.024562 (rank : 61) | θ value | 6.88961 (rank : 93) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8TF01, Q5T064, Q6P2B4, Q6P2N4, Q6PJ93, Q6PK36, Q7Z640, Q8TF00, Q96K10, Q96SM5, Q9P076, Q9P0C0, Q9Y4N3 | Gene names | SRRP130, C6orf111 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Splicing factor, arginine/serine-rich 130 (Serine-arginine-rich- splicing regulatory protein 130) (SRrp130) (SR-rich protein) (SR- related protein). | |||||
|
MAP1B_MOUSE
|
||||||
| NC score | 0.024159 (rank : 62) | θ value | 6.88961 (rank : 81) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 977 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P14873 | Gene names | Map1b, Mtap1b, Mtap5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) (MAP1.2) (MAP1(X)) [Contains: MAP1 light chain LC1]. | |||||
|
ECHB_MOUSE
|
||||||
| NC score | 0.023229 (rank : 63) | θ value | 3.0926 (rank : 48) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99JY0, Q3TEH9, Q8BJI5, Q8BJM0, Q8BK52 | Gene names | Hadhb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trifunctional enzyme subunit beta, mitochondrial precursor (TP-beta) [Includes: 3-ketoacyl-CoA thiolase (EC 2.3.1.16) (Acetyl-CoA acyltransferase) (Beta-ketothiolase)]. | |||||
|
FOXP1_HUMAN
|
||||||
| NC score | 0.023162 (rank : 64) | θ value | 0.813845 (rank : 24) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9H334, Q9H332, Q9H333, Q9P0R1 | Gene names | FOXP1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein P1. | |||||
|
NFH_MOUSE
|
||||||
| NC score | 0.023027 (rank : 65) | θ value | 8.99809 (rank : 105) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
OSTP_MOUSE
|
||||||
| NC score | 0.022733 (rank : 66) | θ value | 8.99809 (rank : 106) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P10923, P19008 | Gene names | Spp1, Eta-1, Op, Spp-1 | |||
|
Domain Architecture |

|
|||||
| Description | Osteopontin precursor (Bone sialoprotein-1) (Secreted phosphoprotein 1) (SPP-1) (Minopontin) (Early T-lymphocyte activation 1 protein) (2AR) (Calcium oxalate crystal growth inhibitor protein). | |||||
|
ZCH14_HUMAN
|
||||||
| NC score | 0.022430 (rank : 67) | θ value | 5.27518 (rank : 73) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8WYQ9, O60324, Q9UFP0 | Gene names | ZCCHC14, KIAA0579 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCHC domain-containing protein 14 (BDG-29). | |||||
|
UBE4B_HUMAN
|
||||||
| NC score | 0.022311 (rank : 68) | θ value | 4.03905 (rank : 61) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O95155, O75169, O95338, Q5SZ12, Q5SZ16, Q96QD4, Q9BYI7 | Gene names | UBE4B, HDNB1, KIAA0684, UFD2 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin conjugation factor E4 B (Ubiquitin fusion degradation protein 2) (Homozygously deleted in neuroblastoma 1). | |||||
|
AP3B1_HUMAN
|
||||||
| NC score | 0.022307 (rank : 69) | θ value | 1.81305 (rank : 33) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O00203, O00580, Q7Z393, Q9HD66 | Gene names | AP3B1, ADTB3A | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit beta-1 (Adapter-related protein complex 3 beta-1 subunit) (Beta3A-adaptin) (Adaptor protein complex AP-3 beta-1 subunit) (Clathrin assembly protein complex 3 beta-1 large chain). | |||||
|
SDC3_MOUSE
|
||||||
| NC score | 0.021567 (rank : 70) | θ value | 8.99809 (rank : 108) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q64519, Q3UDD9, Q6ZQA4, Q7TQD4 | Gene names | Sdc3, Kiaa0468 | |||
|
Domain Architecture |

|
|||||
| Description | Syndecan-3 precursor (SYND3). | |||||
|
FOXP1_MOUSE
|
||||||
| NC score | 0.020943 (rank : 71) | θ value | 1.81305 (rank : 36) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P58462 | Gene names | Foxp1 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein P1 (Forkhead-related transcription factor 1). | |||||
|
ECHB_HUMAN
|
||||||
| NC score | 0.020864 (rank : 72) | θ value | 5.27518 (rank : 65) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P55084, O14969, Q53TA6, Q96C77, Q9H3F5 | Gene names | HADHB | |||
|
Domain Architecture |

|
|||||
| Description | Trifunctional enzyme subunit beta, mitochondrial precursor (TP-beta) [Includes: 3-ketoacyl-CoA thiolase (EC 2.3.1.16) (Acetyl-CoA acyltransferase) (Beta-ketothiolase)]. | |||||
|
NCOA2_MOUSE
|
||||||
| NC score | 0.020486 (rank : 73) | θ value | 6.88961 (rank : 83) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q61026, O09001, P97759 | Gene names | Ncoa2, Grip1, Tif2 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 2 (NCoA-2) (Transcriptional intermediary factor 2) (Glucocorticoid receptor-interacting protein 1) (GRIP-1). | |||||
|
MUC5B_HUMAN
|
||||||
| NC score | 0.020275 (rank : 74) | θ value | 2.36792 (rank : 42) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1020 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9HC84, O00447, O00573, O14985, O15494, O95291, O95451, Q14881, Q99552, Q9UE28 | Gene names | MUC5B, MUC5 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-5B precursor (Mucin-5 subtype B, tracheobronchial) (High molecular weight salivary mucin MG1) (Sublingual gland mucin). | |||||
|
PEX19_MOUSE
|
||||||
| NC score | 0.019697 (rank : 75) | θ value | 6.88961 (rank : 88) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8VCI5, Q8CEE1, Q921H0, Q9CZC1, Q9QUQ1 | Gene names | Pex19, Pxf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisomal biogenesis factor 19 (Peroxin-19) (Peroxisomal farnesylated protein) (PxF). | |||||
|
UBP32_HUMAN
|
||||||
| NC score | 0.018471 (rank : 76) | θ value | 3.0926 (rank : 54) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8NFA0, Q9BX85, Q9Y591 | Gene names | USP32, USP10 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 32 (EC 3.1.2.15) (Ubiquitin thioesterase 32) (Ubiquitin-specific-processing protease 32) (Deubiquitinating enzyme 32) (NY-REN-60 antigen). | |||||
|
MINT_MOUSE
|
||||||
| NC score | 0.018228 (rank : 77) | θ value | 8.99809 (rank : 101) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
NCOA2_HUMAN
|
||||||
| NC score | 0.017984 (rank : 78) | θ value | 8.99809 (rank : 104) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q15596 | Gene names | NCOA2, TIF2 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor coactivator 2 (NCoA-2) (Transcriptional intermediary factor 2). | |||||
|
DACH1_MOUSE
|
||||||
| NC score | 0.017739 (rank : 79) | θ value | 5.27518 (rank : 64) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9QYB2, O88716, Q8BPQ0, Q8C8D7, Q9QY47, Q9R218, Q9Z0Y5 | Gene names | Dach1, Dach | |||
|
Domain Architecture |

|
|||||
| Description | Dachshund homolog 1 (Dach1). | |||||
|
S6OS1_MOUSE
|
||||||
| NC score | 0.016728 (rank : 80) | θ value | 6.88961 (rank : 91) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9CTN5, Q9D5F5, Q9D5Y9 | Gene names | Six6os1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein SIX6OS1 (Six6 opposite strand transcript 1). | |||||
|
LIMA1_HUMAN
|
||||||
| NC score | 0.015468 (rank : 81) | θ value | 5.27518 (rank : 66) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 309 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9UHB6, Q2TAN7, Q9BVF2, Q9H8J1, Q9HBN5, Q9NX96, Q9NXC3, Q9NXU6, Q9P0H8, Q9UHB5 | Gene names | LIMA1, EPLIN, SREBP3 | |||
|
Domain Architecture |

|
|||||
| Description | LIM domain and actin-binding protein 1 (Epithelial protein lost in neoplasm). | |||||
|
MACF4_HUMAN
|
||||||
| NC score | 0.015363 (rank : 82) | θ value | 6.88961 (rank : 79) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1321 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q96PK2, Q8WXY1 | Gene names | MACF1, ABP620, ACF7, KIAA0465, KIAA1251 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-actin cross-linking factor 1, isoform 4. | |||||
|
UBF1_HUMAN
|
||||||
| NC score | 0.015356 (rank : 83) | θ value | 5.27518 (rank : 69) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 415 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P17480 | Gene names | UBTF, UBF, UBF1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar transcription factor 1 (Upstream-binding factor 1) (UBF-1) (Autoantigen NOR-90). | |||||
|
HUWE1_MOUSE
|
||||||
| NC score | 0.014407 (rank : 84) | θ value | 3.0926 (rank : 49) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q7TMY8, Q4G2Z1, Q5BMM7, Q6NS61, Q8BNJ7, Q8CFH2, Q8VD14, Q921M5, Q9R0P2 | Gene names | Huwe1, Kiaa0312, Ureb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT, UBA and WWE domain-containing protein 1 (EC 6.3.2.-) (E3 ubiquitin protein ligase URE-B1) (E3Histone). | |||||
|
PHF2_HUMAN
|
||||||
| NC score | 0.013029 (rank : 85) | θ value | 5.27518 (rank : 68) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O75151, Q8N3K2, Q9Y6N4 | Gene names | PHF2, KIAA0662 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 2 (GRC5). | |||||
|
SRPK1_HUMAN
|
||||||
| NC score | 0.012783 (rank : 86) | θ value | 3.0926 (rank : 52) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 706 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q96SB4, Q12890, Q5R364, Q5R365, Q8IY12 | Gene names | SRPK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SRPK1 (EC 2.7.11.1) (Serine/arginine- rich protein-specific kinase 1) (SR-protein-specific kinase 1) (SFRS protein kinase 1). | |||||
|
DJBP_MOUSE
|
||||||
| NC score | 0.011936 (rank : 87) | θ value | 8.99809 (rank : 99) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6P1E8, Q5DTV8, Q6PGK8, Q9D2D8 | Gene names | Djbp, Kiaa1672 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DJ-1-binding protein. | |||||
|
MAGC1_HUMAN
|
||||||
| NC score | 0.011481 (rank : 88) | θ value | 8.99809 (rank : 100) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 431 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O60732, O75451, Q8TCV4 | Gene names | MAGEC1 | |||
|
Domain Architecture |

|
|||||
| Description | Melanoma-associated antigen C1 (MAGE-C1 antigen) (Cancer-testis antigen CT7). | |||||
|
MACF1_HUMAN
|
||||||
| NC score | 0.011418 (rank : 89) | θ value | 6.88961 (rank : 78) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1341 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9UPN3, O75053, Q5VW20, Q8WXY2, Q9H540, Q9UKP0, Q9ULG9 | Gene names | MACF1, ABP620, ACF7, KIAA0465, KIAA1251 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-actin cross-linking factor 1, isoforms 1/2/3/5 (Actin cross-linking family protein 7) (Macrophin-1) (Trabeculin-alpha) (620 kDa actin-binding protein) (ABP620). | |||||
|
MACOI_HUMAN
|
||||||
| NC score | 0.011399 (rank : 90) | θ value | 6.88961 (rank : 80) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8N5G2, Q2TLX5, Q2TLX6, Q9NVG6 | Gene names | TMEM57 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Macoilin (Transmembrane protein 57). | |||||
|
SCN5A_HUMAN
|
||||||
| NC score | 0.009876 (rank : 91) | θ value | 2.36792 (rank : 46) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14524 | Gene names | SCN5A | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 5 subunit alpha (Sodium channel protein type V subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.5) (Sodium channel protein, cardiac muscle alpha-subunit) (HH1). | |||||
|
FYCO1_MOUSE
|
||||||
| NC score | 0.009853 (rank : 92) | θ value | 6.88961 (rank : 76) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1059 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q8VDC1, Q3V385, Q7TMT0, Q8BJN2 | Gene names | Fyco1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE and coiled-coil domain-containing protein 1. | |||||
|
AP4E1_HUMAN
|
||||||
| NC score | 0.009733 (rank : 93) | θ value | 8.99809 (rank : 95) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UPM8, Q9Y588 | Gene names | AP4E1 | |||
|
Domain Architecture |

|
|||||
| Description | AP-4 complex subunit epsilon-1 (Adapter-related protein complex 4 epsilon-1 subunit) (Epsilon subunit of AP-4) (AP-4 adapter complex epsilon subunit). | |||||
|
SYN1_MOUSE
|
||||||
| NC score | 0.009600 (rank : 94) | θ value | 8.99809 (rank : 110) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 363 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O88935, Q62279, Q8QZT8 | Gene names | Syn1, Syn-1 | |||
|
Domain Architecture |

|
|||||
| Description | Synapsin-1 (Synapsin I). | |||||
|
KAISO_HUMAN
|
||||||
| NC score | 0.007792 (rank : 95) | θ value | 1.06291 (rank : 27) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 840 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q86T24, O00319, Q7Z361, Q8IVP6, Q8N3P0 | Gene names | ZBTB33, KAISO, ZNF348 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcriptional regulator Kaiso (Zinc finger and BTB domain-containing protein 33). | |||||
|
GIT2_MOUSE
|
||||||
| NC score | 0.007531 (rank : 96) | θ value | 6.88961 (rank : 77) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9JLQ2 | Gene names | Git2 | |||
|
Domain Architecture |

|
|||||
| Description | ARF GTPase-activating protein GIT2 (G protein-coupled receptor kinase- interactor 2) (Cool-interacting tyrosine-phosphorylated protein 2) (CAT2) (CAT-2). | |||||
|
KCNA6_MOUSE
|
||||||
| NC score | 0.006948 (rank : 97) | θ value | 4.03905 (rank : 57) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q61923 | Gene names | Kcna6 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 6 (Voltage-gated potassium channel subunit Kv1.6) (MK1.6). | |||||
|
TGFR2_MOUSE
|
||||||
| NC score | 0.005784 (rank : 98) | θ value | 4.03905 (rank : 60) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 748 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q62312, Q63947 | Gene names | Tgfbr2 | |||
|
Domain Architecture |

|
|||||
| Description | TGF-beta receptor type-2 precursor (EC 2.7.11.30) (TGF-beta receptor type II) (TGFR-2) (TGF-beta type II receptor). | |||||
|
ARHGC_HUMAN
|
||||||
| NC score | 0.005717 (rank : 99) | θ value | 8.99809 (rank : 96) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NZN5, O15086 | Gene names | ARHGEF12, KIAA0382, LARG | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 12 (Leukemia-associated RhoGEF). | |||||
|
PLCB1_MOUSE
|
||||||
| NC score | 0.005654 (rank : 100) | θ value | 8.99809 (rank : 107) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 531 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Z1B3, Q62075, Q8K5A5, Q8K5A6, Q9Z0E5, Q9Z2T5 | Gene names | Plcb1, Plcb | |||
|
Domain Architecture |

|
|||||
| Description | 1-phosphatidylinositol-4,5-bisphosphate phosphodiesterase beta 1 (EC 3.1.4.11) (Phosphoinositide phospholipase C) (Phospholipase C- beta-1) (PLC-beta-1) (PLC-I) (PLC-154). | |||||
|
CADH1_MOUSE
|
||||||
| NC score | 0.004218 (rank : 101) | θ value | 4.03905 (rank : 55) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P09803, Q61377 | Gene names | Cdh1 | |||
|
Domain Architecture |

|
|||||
| Description | Epithelial-cadherin precursor (E-cadherin) (Uvomorulin) (Cadherin-1) (ARC-1) (CD324 antigen) [Contains: E-Cad/CTF1; E-Cad/CTF2; E- Cad/CTF3]. | |||||
|
TNIK_HUMAN
|
||||||
| NC score | 0.004074 (rank : 102) | θ value | 3.0926 (rank : 53) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1570 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q9UKE5, O60298, Q8WUY7, Q9UKD8, Q9UKD9, Q9UKE0, Q9UKE1, Q9UKE2, Q9UKE3, Q9UKE4 | Gene names | TNIK, KIAA0551 | |||
|
Domain Architecture |

|
|||||
| Description | TRAF2 and NCK-interacting protein kinase (EC 2.7.11.1). | |||||
|
WNK4_HUMAN
|
||||||
| NC score | 0.004071 (rank : 103) | θ value | 5.27518 (rank : 72) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 1043 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q96J92, Q8N8X3, Q8N8Z2, Q96DT8, Q9BYS5 | Gene names | WNK4, PRKWNK4 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK4 (EC 2.7.11.1) (Protein kinase with no lysine 4) (Protein kinase, lysine-deficient 4). | |||||
|
RXRG_HUMAN
|
||||||
| NC score | 0.003888 (rank : 104) | θ value | 6.88961 (rank : 90) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P48443 | Gene names | RXRG, NR2B3 | |||
|
Domain Architecture |

|
|||||
| Description | Retinoic acid receptor RXR-gamma (Retinoid X receptor gamma). | |||||
|
M3K2_MOUSE
|
||||||
| NC score | 0.002846 (rank : 105) | θ value | 4.03905 (rank : 58) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 846 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q61083 | Gene names | Map3k2, Mekk2 | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase kinase kinase 2 (EC 2.7.11.25) (MAPK/ERK kinase kinase 2) (MEK kinase 2) (MEKK 2). | |||||
|
MK10_HUMAN
|
||||||
| NC score | 0.002674 (rank : 106) | θ value | 6.88961 (rank : 82) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 836 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P53779, Q15707 | Gene names | MAPK10, JNK3, JNK3A, PRKM10 | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase 10 (EC 2.7.11.24) (Stress-activated protein kinase JNK3) (c-Jun N-terminal kinase 3) (MAP kinase p49 3F12). | |||||
|
MK10_MOUSE
|
||||||
| NC score | 0.002562 (rank : 107) | θ value | 8.99809 (rank : 102) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 849 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q61831, Q9R0U6 | Gene names | Mapk10, Jnk3, Prkm10, Serk2 | |||
|
Domain Architecture |

|
|||||
| Description | Mitogen-activated protein kinase 10 (EC 2.7.11.24) (Stress-activated protein kinase JNK3) (c-Jun N-terminal kinase 3) (MAP kinase p49 3F12). | |||||
|
PRDM9_HUMAN
|
||||||
| NC score | 0.002018 (rank : 108) | θ value | 4.03905 (rank : 59) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 750 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9NQV7 | Gene names | PRDM9, PFM6 | |||
|
Domain Architecture |

|
|||||
| Description | PR domain zinc finger protein 9 (PR domain-containing protein 9). | |||||
|
SP2_HUMAN
|
||||||
| NC score | 0.000878 (rank : 109) | θ value | 8.99809 (rank : 109) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 747 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q02086 | Gene names | SP2, KIAA0048 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor Sp2. | |||||
|
MTF1_HUMAN
|
||||||
| NC score | 0.000551 (rank : 110) | θ value | 8.99809 (rank : 103) | |||
| Query Neighborhood Hits | 110 | Target Neighborhood Hits | 814 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q14872, Q96CB1 | Gene names | MTF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metal-regulatory transcription factor 1 (Transcription factor MTF-1) (MRE-binding transcription factor). | |||||