Please be patient as the page loads
|
KCNN1_MOUSE
|
||||||
| SwissProt Accessions | Q9EQR3, Q99JA9 | Gene names | Kcnn1, Sk1 | |||
|
Domain Architecture |

|
|||||
| Description | Small conductance calcium-activated potassium channel protein 1 (SK1). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
KCNN1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.991905 (rank : 2) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 85 | |
| SwissProt Accessions | Q92952 | Gene names | KCNN1, SK | |||
|
Domain Architecture |
|
|||||
| Description | Small conductance calcium-activated potassium channel protein 1 (SK1). | |||||
|
KCNN1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 117 | |
| SwissProt Accessions | Q9EQR3, Q99JA9 | Gene names | Kcnn1, Sk1 | |||
|
Domain Architecture |

|
|||||
| Description | Small conductance calcium-activated potassium channel protein 1 (SK1). | |||||
|
KCNN2_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.989156 (rank : 3) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | Q9H2S1 | Gene names | KCNN2 | |||
|
Domain Architecture |
|
|||||
| Description | Small conductance calcium-activated potassium channel protein 2 (SK2). | |||||
|
KCNN2_MOUSE
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.988273 (rank : 4) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | P58390 | Gene names | Kcnn2, Sk2 | |||
|
Domain Architecture |
|
|||||
| Description | Small conductance calcium-activated potassium channel protein 2 (SK2). | |||||
|
KCNN3_HUMAN
|
||||||
| θ value | 0 (rank : 5) | NC score | 0.974949 (rank : 6) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 78 | |
| SwissProt Accessions | Q9UGI6, O43517 | Gene names | KCNN3, K3 | |||
|
Domain Architecture |
|
|||||
| Description | Small conductance calcium-activated potassium channel protein 3 (SK3) (SKCa3). | |||||
|
KCNN3_MOUSE
|
||||||
| θ value | 0 (rank : 6) | NC score | 0.979850 (rank : 5) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | P58391 | Gene names | Kcnn3, Sk3 | |||
|
Domain Architecture |
|
|||||
| Description | Small conductance calcium-activated potassium channel protein 3 (SK3). | |||||
|
KCNN4_MOUSE
|
||||||
| θ value | 1.49966e-79 (rank : 7) | NC score | 0.951391 (rank : 7) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | O89109, Q3U1J8, Q9CY39 | Gene names | Kcnn4 | |||
|
Domain Architecture |

|
|||||
| Description | Intermediate conductance calcium-activated potassium channel protein 4 (SK4) (KCa4) (IK1). | |||||
|
KCNN4_HUMAN
|
||||||
| θ value | 6.30065e-78 (rank : 8) | NC score | 0.949220 (rank : 8) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | O15554 | Gene names | KCNN4, IK1, IKCA1, KCA4, SK4 | |||
|
Domain Architecture |

|
|||||
| Description | Intermediate conductance calcium-activated potassium channel protein 4 (SK4) (KCa4) (IK1) (IKCa1) (Putative Gardos channel). | |||||
|
KCNQ5_HUMAN
|
||||||
| θ value | 8.40245e-06 (rank : 9) | NC score | 0.362020 (rank : 10) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q9NR82, Q9NRN0, Q9NYA6 | Gene names | KCNQ5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 5 (Voltage-gated potassium channel subunit Kv7.5) (Potassium channel subunit alpha KvLQT5) (KQT-like 5). | |||||
|
KCNQ4_HUMAN
|
||||||
| θ value | 1.43324e-05 (rank : 10) | NC score | 0.366981 (rank : 9) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | P56696, O96025 | Gene names | KCNQ4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 4 (Voltage-gated potassium channel subunit Kv7.4) (Potassium channel subunit alpha KvLQT4) (KQT-like 4). | |||||
|
KCNQ2_MOUSE
|
||||||
| θ value | 0.00020696 (rank : 11) | NC score | 0.344256 (rank : 11) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q9Z351, Q8R498, Q9QWN9, Q9Z342, Q9Z343, Q9Z344, Q9Z345, Q9Z346, Q9Z347, Q9Z348, Q9Z349, Q9Z350 | Gene names | Kcnq2, Kqt2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 2 (Voltage-gated potassium channel subunit Kv7.2) (Potassium channel subunit alpha KvLQT2) (KQT-like 2). | |||||
|
KCND1_MOUSE
|
||||||
| θ value | 0.000461057 (rank : 12) | NC score | 0.296664 (rank : 19) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q03719, Q8CC68 | Gene names | Kcnd1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 1 (Voltage-gated potassium channel subunit Kv4.1) (mShal). | |||||
|
KCND2_HUMAN
|
||||||
| θ value | 0.000461057 (rank : 13) | NC score | 0.296717 (rank : 17) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q9NZV8, O95012, O95021, Q9UBY7, Q9UN98, Q9UNH9 | Gene names | KCND2, KIAA1044 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 2 (Voltage-gated potassium channel subunit Kv4.2). | |||||
|
KCND2_MOUSE
|
||||||
| θ value | 0.000461057 (rank : 14) | NC score | 0.296717 (rank : 18) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q9Z0V2, Q8BSK3, Q8CHB7, Q9JJ60 | Gene names | Kcnd2, Kiaa1044 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 2 (Voltage-gated potassium channel subunit Kv4.2). | |||||
|
KCND1_HUMAN
|
||||||
| θ value | 0.000602161 (rank : 15) | NC score | 0.296653 (rank : 20) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q9NSA2, O75671 | Gene names | KCND1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 1 (Voltage-gated potassium channel subunit Kv4.1). | |||||
|
KCNQ2_HUMAN
|
||||||
| θ value | 0.000786445 (rank : 16) | NC score | 0.341467 (rank : 14) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | O43526, O43796, O75580, O95845, Q5VYT8, Q96J59, Q99454 | Gene names | KCNQ2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 2 (Voltage-gated potassium channel subunit Kv7.2) (Neuroblastoma-specific potassium channel subunit alpha KvLQT2) (KQT-like 2). | |||||
|
KCNQ3_HUMAN
|
||||||
| θ value | 0.00102713 (rank : 17) | NC score | 0.336119 (rank : 15) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | O43525 | Gene names | KCNQ3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 3 (Voltage-gated potassium channel subunit Kv7.3) (Potassium channel subunit alpha KvLQT3) (KQT-like 3). | |||||
|
KCND3_HUMAN
|
||||||
| θ value | 0.00134147 (rank : 18) | NC score | 0.292471 (rank : 22) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q9UK17, O60576, O60577, Q5T0M0, Q9UH85, Q9UH86, Q9UK16 | Gene names | KCND3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 3 (Voltage-gated potassium channel subunit Kv4.3). | |||||
|
KCND3_MOUSE
|
||||||
| θ value | 0.00134147 (rank : 19) | NC score | 0.292600 (rank : 21) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q9Z0V1, Q8CC44, Q9Z0V0 | Gene names | Kcnd3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 3 (Voltage-gated potassium channel subunit Kv4.3). | |||||
|
KCNQ3_MOUSE
|
||||||
| θ value | 0.00175202 (rank : 20) | NC score | 0.332863 (rank : 16) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q8K3F6 | Gene names | Kcnq3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 3 (Voltage-gated potassium channel subunit Kv7.3) (Potassium channel subunit alpha KvLQT3) (KQT-like 3). | |||||
|
KCNQ1_MOUSE
|
||||||
| θ value | 0.00228821 (rank : 21) | NC score | 0.341715 (rank : 13) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | P97414, O88702, Q7TNZ1, Q7TPL7, Q80VR7 | Gene names | Kcnq1, Kcna9, Kvlqt1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 1 (Voltage-gated potassium channel subunit Kv7.1) (IKs producing slow voltage-gated potassium channel subunit alpha KvLQT1) (KQT-like 1). | |||||
|
KCNQ1_HUMAN
|
||||||
| θ value | 0.00298849 (rank : 22) | NC score | 0.342836 (rank : 12) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | P51787, O00347, O60607, O94787, Q7Z6G9, Q92960, Q9UMN8, Q9UMN9 | Gene names | KCNQ1, KCNA8, KCNA9, KVLQT1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 1 (Voltage-gated potassium channel subunit Kv7.1) (IKs producing slow voltage-gated potassium channel subunit alpha KvLQT1) (KQT-like 1). | |||||
|
KCNB2_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 23) | NC score | 0.280444 (rank : 28) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q92953, Q9BXD3 | Gene names | KCNB2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily B member 2 (Voltage-gated potassium channel subunit Kv2.2). | |||||
|
KCNA6_HUMAN
|
||||||
| θ value | 0.00665767 (rank : 24) | NC score | 0.289035 (rank : 25) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | P17658 | Gene names | KCNA6 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 6 (Voltage-gated potassium channel subunit Kv1.6) (HBK2). | |||||
|
KCNA6_MOUSE
|
||||||
| θ value | 0.00869519 (rank : 25) | NC score | 0.286528 (rank : 26) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q61923 | Gene names | Kcna6 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 6 (Voltage-gated potassium channel subunit Kv1.6) (MK1.6). | |||||
|
KCNS2_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 26) | NC score | 0.290813 (rank : 24) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q9ULS6 | Gene names | KCNS2, KIAA1144 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily S member 2 (Voltage-gated potassium channel subunit Kv9.2) (Delayed-rectifier K(+) channel alpha subunit 2). | |||||
|
KCNS2_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 27) | NC score | 0.291765 (rank : 23) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | O35174, Q543P3 | Gene names | Kcns2 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium voltage-gated channel subfamily S member 2 (Voltage-gated potassium channel subunit Kv9.2) (Delayed-rectifier K(+) channel alpha subunit 2). | |||||
|
KCNB1_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 28) | NC score | 0.272626 (rank : 36) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q14721, Q14193 | Gene names | KCNB1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily B member 1 (Voltage-gated potassium channel subunit Kv2.1) (h-DRK1). | |||||
|
KCNB1_MOUSE
|
||||||
| θ value | 0.0193708 (rank : 29) | NC score | 0.272742 (rank : 35) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q03717 | Gene names | Kcnb1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily B member 1 (Voltage-gated potassium channel subunit Kv2.1) (mShab). | |||||
|
KCNA4_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 30) | NC score | 0.277448 (rank : 31) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | P22459 | Gene names | KCNA4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 4 (Voltage-gated potassium channel subunit Kv1.4) (HK1) (HPCN2) (HBK4) (HUKII). | |||||
|
KCNA5_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 31) | NC score | 0.283380 (rank : 27) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q61762 | Gene names | Kcna5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 5 (Voltage-gated potassium channel subunit Kv1.5) (KV1-5). | |||||
|
KCNA2_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 32) | NC score | 0.279516 (rank : 29) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | P16389 | Gene names | KCNA2 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium voltage-gated channel subfamily A member 2 (Voltage-gated potassium channel subunit Kv1.2) (HBK5) (NGK1) (HUKIV). | |||||
|
KCNG3_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 33) | NC score | 0.265816 (rank : 41) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q8TAE7 | Gene names | KCNG3 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium voltage-gated channel subfamily G member 3 (Voltage-gated potassium channel subunit Kv6.3) (Kv10.1). | |||||
|
KCNG3_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 34) | NC score | 0.265590 (rank : 42) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | P59053 | Gene names | Kcng3 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium voltage-gated channel subfamily G member 3 (Voltage-gated potassium channel subunit Kv6.3) (Kv10.1). | |||||
|
RAB3I_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 35) | NC score | 0.062146 (rank : 76) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q68EF0 | Gene names | Rab3ip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RAB3A-interacting protein (Rabin-3) (SSX2-interacting protein). | |||||
|
KCNA2_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 36) | NC score | 0.277224 (rank : 33) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | P63141, P15386, Q02010, Q8C8W4 | Gene names | Kcna2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium voltage-gated channel subfamily A member 2 (Voltage-gated potassium channel subunit Kv1.2) (MK2). | |||||
|
ROCK2_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 37) | NC score | 0.015206 (rank : 97) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 2073 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O75116, Q9UQN5 | Gene names | ROCK2, KIAA0619 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 2 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 2) (p164 ROCK-2) (Rho kinase 2). | |||||
|
KCNA3_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 38) | NC score | 0.277439 (rank : 32) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | P22001 | Gene names | KCNA3, HGK5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 3 (Voltage-gated potassium channel subunit Kv1.3) (HPCN3) (HGK5) (HuKIII) (HLK3). | |||||
|
KCNA3_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 39) | NC score | 0.276846 (rank : 34) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | P16390 | Gene names | Kcna3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 3 (Voltage-gated potassium channel subunit Kv1.3) (MK3). | |||||
|
KCNA4_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 40) | NC score | 0.272328 (rank : 38) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q61423 | Gene names | Kcna4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 4 (Voltage-gated potassium channel subunit Kv1.4). | |||||
|
KCNA5_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 41) | NC score | 0.278524 (rank : 30) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | P22460 | Gene names | KCNA5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 5 (Voltage-gated potassium channel subunit Kv1.5) (HK2) (HPCN1). | |||||
|
ROCK2_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 42) | NC score | 0.013746 (rank : 100) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 2072 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P70336, Q8BR64, Q8CBR0 | Gene names | Rock2 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 2 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 2) (p164 ROCK-2). | |||||
|
KCNA1_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 43) | NC score | 0.272423 (rank : 37) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q09470 | Gene names | KCNA1 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium voltage-gated channel subfamily A member 1 (Voltage-gated potassium channel subunit Kv1.1) (HUKI) (HBK1). | |||||
|
KCNA1_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 44) | NC score | 0.272180 (rank : 39) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | P16388 | Gene names | Kcna1 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium voltage-gated channel subfamily A member 1 (Voltage-gated potassium channel subunit Kv1.1) (MKI) (MBK1). | |||||
|
KCNK1_HUMAN
|
||||||
| θ value | 0.125558 (rank : 45) | NC score | 0.138466 (rank : 58) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O00180, Q13307 | Gene names | KCNK1, HOHO1, KCNO1, TWIK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 1 (Inward rectifying potassium channel protein TWIK-1) (Potassium channel KCNO1). | |||||
|
KCNS3_HUMAN
|
||||||
| θ value | 0.125558 (rank : 46) | NC score | 0.271204 (rank : 40) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9BQ31, O43651, Q96B56 | Gene names | KCNS3 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium voltage-gated channel subfamily S member 3 (Voltage-gated potassium channel subunit Kv9.3) (Delayed-rectifier K(+) channel alpha subunit 3). | |||||
|
ANGL7_MOUSE
|
||||||
| θ value | 0.163984 (rank : 47) | NC score | 0.033426 (rank : 89) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8R1Q3 | Gene names | Angptl7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiopoietin-related protein 7 precursor (Angiopoietin-like 7). | |||||
|
KCNG2_HUMAN
|
||||||
| θ value | 0.163984 (rank : 48) | NC score | 0.263530 (rank : 43) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9UJ96 | Gene names | KCNG2, KCNF2 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium voltage-gated channel subfamily G member 2 (Voltage-gated potassium channel subunit Kv6.2) (Cardiac potassium channel subunit). | |||||
|
ZN688_HUMAN
|
||||||
| θ value | 0.21417 (rank : 49) | NC score | -0.000510 (rank : 129) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 873 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8WV14, O75701, Q8IW91, Q96MN0 | Gene names | ZNF688 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 688. | |||||
|
KCNC3_MOUSE
|
||||||
| θ value | 0.279714 (rank : 50) | NC score | 0.248264 (rank : 52) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q63959, Q62088 | Gene names | Kcnc3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily C member 3 (Voltage-gated potassium channel subunit Kv3.3) (KSHIIID). | |||||
|
KCNG1_HUMAN
|
||||||
| θ value | 0.279714 (rank : 51) | NC score | 0.256548 (rank : 45) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q9UIX4, O43528, Q9BRC1 | Gene names | KCNG1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily G member 1 (Voltage-gated potassium channel subunit Kv6.1) (kH2). | |||||
|
KCNC1_HUMAN
|
||||||
| θ value | 0.365318 (rank : 52) | NC score | 0.248632 (rank : 50) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | P48547 | Gene names | KCNC1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily C member 1 (Voltage-gated potassium channel subunit Kv3.1) (Kv4) (NGK2). | |||||
|
KCNC1_MOUSE
|
||||||
| θ value | 0.365318 (rank : 53) | NC score | 0.248691 (rank : 49) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | P15388 | Gene names | Kcnc1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily C member 1 (Voltage-gated potassium channel subunit Kv3.1) (Kv4) (NGK2). | |||||
|
KCNC4_HUMAN
|
||||||
| θ value | 0.365318 (rank : 54) | NC score | 0.255227 (rank : 47) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q03721, Q3MIM4, Q5TBI6 | Gene names | KCNC4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily C member 4 (Voltage-gated potassium channel subunit Kv3.4) (KSHIIIC). | |||||
|
KCNC4_MOUSE
|
||||||
| θ value | 0.365318 (rank : 55) | NC score | 0.255883 (rank : 46) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q8R1C0 | Gene names | Kcnc4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium voltage-gated channel subfamily C member 4 (Voltage-gated potassium channel subunit Kv3.4). | |||||
|
KCNK1_MOUSE
|
||||||
| θ value | 0.365318 (rank : 56) | NC score | 0.123573 (rank : 60) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | O08581 | Gene names | Kcnk1 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium channel subfamily K member 1 (Inward rectifying potassium channel protein TWIK-1). | |||||
|
ROCK1_MOUSE
|
||||||
| θ value | 0.365318 (rank : 57) | NC score | 0.009942 (rank : 106) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 2126 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P70335, Q8C3G4, Q8C7H0 | Gene names | Rock1 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 1 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 1) (p160 ROCK-1) (p160ROCK). | |||||
|
DIAP3_HUMAN
|
||||||
| θ value | 0.47712 (rank : 58) | NC score | 0.015518 (rank : 94) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 209 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NSV4, Q5T2S7, Q5XKF6, Q6MZF0, Q6NUP0, Q86VS4, Q8NAV4 | Gene names | DIAPH3, DIAP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein diaphanous homolog 3 (Diaphanous-related formin-3) (DRF3). | |||||
|
KCMA1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 59) | NC score | 0.210084 (rank : 57) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q12791, Q12886, Q12917, Q12921, Q12960, Q13150, Q96LG8, Q9UBB0, Q9UQK6 | Gene names | KCNMA1, KCNMA | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-activated potassium channel subunit alpha 1 (Calcium-activated potassium channel, subfamily M subunit alpha 1) (Maxi K channel) (MaxiK) (BK channel) (K(VCA)alpha) (BKCA alpha) (KCa1.1) (Slowpoke homolog) (Slo homolog) (Slo-alpha) (Slo1) (hSlo). | |||||
|
KCMA1_MOUSE
|
||||||
| θ value | 0.47712 (rank : 60) | NC score | 0.210468 (rank : 56) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q08460, Q64703, Q8VHF1, Q9R196 | Gene names | Kcnma1, Kcnma | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-activated potassium channel subunit alpha 1 (Calcium-activated potassium channel, subfamily M subunit alpha 1) (Maxi K channel) (MaxiK) (BK channel) (K(VCA)alpha) (BKCA alpha) (KCa1.1) (Slowpoke homolog) (Slo homolog) (Slo-alpha) (Slo1) (mSlo) (mSlo1). | |||||
|
KCNC3_HUMAN
|
||||||
| θ value | 0.47712 (rank : 61) | NC score | 0.245026 (rank : 55) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 256 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q14003 | Gene names | KCNC3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily C member 3 (Voltage-gated potassium channel subunit Kv3.3) (KSHIIID). | |||||
|
KCNK6_HUMAN
|
||||||
| θ value | 0.47712 (rank : 62) | NC score | 0.137269 (rank : 59) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q9Y257, Q9HB47 | Gene names | KCNK6, TOSS, TWIK2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 6 (Inward rectifying potassium channel protein TWIK-2) (TWIK-originated similarity sequence). | |||||
|
KCNG4_HUMAN
|
||||||
| θ value | 0.62314 (rank : 63) | NC score | 0.248413 (rank : 51) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q8TDN1, Q96H24 | Gene names | KCNG4, KCNG3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily G member 4 (Voltage-gated potassium channel subunit Kv6.4). | |||||
|
GOGA4_HUMAN
|
||||||
| θ value | 0.813845 (rank : 64) | NC score | 0.019181 (rank : 93) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 2034 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q13439, Q13270, Q13654, Q14436 | Gene names | GOLGA4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (Trans-Golgi p230) (256 kDa golgin) (Golgin-245) (Protein 72.1). | |||||
|
KCNS1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 65) | NC score | 0.256616 (rank : 44) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q96KK3, O43652, Q6DJU6 | Gene names | KCNS1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily S member 1 (Voltage-gated potassium channel subunit Kv9.1) (Delayed-rectifier K(+) channel alpha subunit 1). | |||||
|
ERBB2_MOUSE
|
||||||
| θ value | 1.06291 (rank : 66) | NC score | 0.004448 (rank : 117) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 918 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P70424, Q61525, Q6ZPE0 | Gene names | Erbb2, Kiaa3023, Neu | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Receptor tyrosine-protein kinase erbB-2 precursor (EC 2.7.10.1) (p185erbB2) (C-erbB-2) (NEU proto-oncogene). | |||||
|
KCNS1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 67) | NC score | 0.246795 (rank : 53) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | O35173 | Gene names | Kcns1 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium voltage-gated channel subfamily S member 1 (Voltage-gated potassium channel subunit Kv9.1) (Delayed-rectifier K(+) channel alpha subunit 1). | |||||
|
CD2L6_MOUSE
|
||||||
| θ value | 1.38821 (rank : 68) | NC score | 0.003573 (rank : 122) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 851 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8BWD8, Q80TM1 | Gene names | Cdc2l6, Kiaa1028 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle 2-like protein kinase 6 (EC 2.7.11.22) (CDC2- related protein kinase 6). | |||||
|
CCD41_MOUSE
|
||||||
| θ value | 1.81305 (rank : 69) | NC score | 0.010254 (rank : 104) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9D5R3, Q3U7X7, Q3UX57, Q80VF0 | Gene names | Ccdc41 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 41. | |||||
|
CUTL2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 70) | NC score | 0.014045 (rank : 99) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 771 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P70298 | Gene names | Cutl2 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein cut-like 2 (Homeobox protein Cux-2) (Cut-like 2). | |||||
|
KCNF1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 71) | NC score | 0.250794 (rank : 48) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q9H3M0, O43527 | Gene names | KCNF1 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium voltage-gated channel subfamily F member 1 (Voltage-gated potassium channel subunit Kv5.1) (kH1). | |||||
|
KCNK4_MOUSE
|
||||||
| θ value | 1.81305 (rank : 72) | NC score | 0.084871 (rank : 70) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | O88454 | Gene names | Kcnk4, Traak | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 4 (TWIK-related arachidonic acid- stimulated potassium channel protein) (TRAAK). | |||||
|
KCNK9_HUMAN
|
||||||
| θ value | 1.81305 (rank : 73) | NC score | 0.111507 (rank : 64) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9NPC2, Q2M290, Q540F2 | Gene names | KCNK9, TASK3 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium channel subfamily K member 9 (Acid-sensitive potassium channel protein TASK-3) (TWIK-related acid-sensitive K(+) channel 3) (Two pore potassium channel KT3.2). | |||||
|
KCNK9_MOUSE
|
||||||
| θ value | 1.81305 (rank : 74) | NC score | 0.111136 (rank : 65) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q3LS21 | Gene names | Kcnk9, Task3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 9 (Acid-sensitive potassium channel protein TASK-3) (TWIK-related acid-sensitive K(+) channel 3). | |||||
|
RSMB_HUMAN
|
||||||
| θ value | 1.81305 (rank : 75) | NC score | 0.020195 (rank : 92) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P14678, Q15490, Q9UIS5 | Gene names | SNRPB, COD, SNRPB1 | |||
|
Domain Architecture |
|
|||||
| Description | Small nuclear ribonucleoprotein-associated proteins B and B' (snRNP-B) (Sm protein B/B') (Sm-B/Sm-B') (SmB/SmB'). | |||||
|
KCNV2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 76) | NC score | 0.245385 (rank : 54) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q8TDN2 | Gene names | KCNV2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily V member 2 (Voltage-gated potassium channel subunit Kv8.2). | |||||
|
RAB3I_HUMAN
|
||||||
| θ value | 2.36792 (rank : 77) | NC score | 0.036392 (rank : 88) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 270 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96QF0, Q6PCE4, Q96A24, Q96QE6, Q96QE7, Q96QE8, Q96QE9, Q96QF1, Q9H673 | Gene names | RAB3IP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RAB3A-interacting protein (Rabin-3) (SSX2-interacting protein). | |||||
|
ERBB2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 78) | NC score | 0.004015 (rank : 118) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 1073 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P04626, Q14256, Q6LDV1, Q9UMK4 | Gene names | ERBB2, HER2, NEU, NGL | |||
|
Domain Architecture |

|
|||||
| Description | Receptor tyrosine-protein kinase erbB-2 precursor (EC 2.7.10.1) (p185erbB2) (C-erbB-2) (NEU proto-oncogene) (Tyrosine kinase-type cell surface receptor HER2) (MLN 19). | |||||
|
LMNB1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 79) | NC score | 0.015225 (rank : 95) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 719 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P14733, Q61791 | Gene names | Lmnb1 | |||
|
Domain Architecture |

|
|||||
| Description | Lamin-B1. | |||||
|
AAT1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 80) | NC score | 0.044149 (rank : 87) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q7Z4T9, Q68DX2, Q8TD41, Q96A45, Q96JE8 | Gene names | AAT1, C3orf15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AMY-1-associating protein expressed in testis 1 (AAT-1). | |||||
|
AZI1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 81) | NC score | 0.011757 (rank : 102) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 1000 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9UPN4, Q96F50 | Gene names | AZI1, CEP131, KIAA1118 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 5-azacytidine-induced protein 1 (Pre-acrosome localization protein 1) (Centrosomal protein of 131 kDa) (Cep131 protein). | |||||
|
BAZ2A_MOUSE
|
||||||
| θ value | 4.03905 (rank : 82) | NC score | 0.021634 (rank : 91) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q91YE5 | Gene names | Baz2a, Tip5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5). | |||||
|
LMNB1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 83) | NC score | 0.015130 (rank : 98) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 714 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P20700, Q3SYN7, Q96EI6 | Gene names | LMNB1, LMN2, LMNB | |||
|
Domain Architecture |

|
|||||
| Description | Lamin-B1. | |||||
|
SMCA4_HUMAN
|
||||||
| θ value | 4.03905 (rank : 84) | NC score | 0.006014 (rank : 115) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 597 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P51532, O95052, Q9HBD3 | Gene names | SMARCA4, BRG1, SNF2B, SNF2L4 | |||
|
Domain Architecture |

|
|||||
| Description | Probable global transcription activator SNF2L4 (EC 3.6.1.-) (ATP- dependent helicase SMARCA4) (SNF2-beta) (BRG-1 protein) (Mitotic growth and transcription activator) (Brahma protein homolog 1) (SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A member 4). | |||||
|
STK36_HUMAN
|
||||||
| θ value | 4.03905 (rank : 85) | NC score | 0.002986 (rank : 124) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9NRP7, Q8TC32, Q9H9N9, Q9UF35, Q9ULE2 | Gene names | STK36, KIAA1278 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase 36 (EC 2.7.11.1) (Fused homolog). | |||||
|
BAZ2A_HUMAN
|
||||||
| θ value | 5.27518 (rank : 86) | NC score | 0.012298 (rank : 101) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 780 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9UIF9, O00536, O15030, Q96H26 | Gene names | BAZ2A, KIAA0314, TIP5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5) (hWALp3). | |||||
|
CO6A2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 87) | NC score | 0.007145 (rank : 110) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P12110, Q13909, Q13910, Q13911, Q14049, Q16259, Q16597, Q9UML3 | Gene names | COL6A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(VI) chain precursor. | |||||
|
COBA2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 88) | NC score | 0.003806 (rank : 119) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 501 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q64739, Q61432, Q9Z1W0 | Gene names | Col11a2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(XI) chain precursor. | |||||
|
KCNK2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 89) | NC score | 0.082776 (rank : 72) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | P97438 | Gene names | Kcnk2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 2 (Outward rectifying potassium channel protein TREK-1) (Two-pore potassium channel TPKC1) (TREK-1 K(+) channel subunit). | |||||
|
KCNK3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 90) | NC score | 0.100016 (rank : 66) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | O14649 | Gene names | KCNK3, TASK, TASK1 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium channel subfamily K member 3 (Acid-sensitive potassium channel protein TASK-1) (TWIK-related acid-sensitive K(+) channel 1) (Two pore potassium channel KT3.1). | |||||
|
KCNK3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 91) | NC score | 0.099963 (rank : 67) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | O35111, O35163 | Gene names | Kcnk3, Ctbak, Task, Task1 | |||
|
Domain Architecture |
|
|||||
| Description | Potassium channel subfamily K member 3 (Acid-sensitive potassium channel protein TASK-1) (TWIK-related acid-sensitive K(+) channel 1) (Cardiac two-pore background K(+) channel) (cTBAK-1) (Two pore potassium channel KT3.1). | |||||
|
KCNK5_HUMAN
|
||||||
| θ value | 5.27518 (rank : 92) | NC score | 0.115484 (rank : 63) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | O95279 | Gene names | KCNK5, TASK2 | |||
|
Domain Architecture |
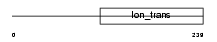
|
|||||
| Description | Potassium channel subfamily K member 5 (Acid-sensitive potassium channel protein TASK-2) (TWIK-related acid-sensitive K(+) channel 2). | |||||
|
KCNKD_HUMAN
|
||||||
| θ value | 5.27518 (rank : 93) | NC score | 0.069033 (rank : 74) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9HB14, Q96E79 | Gene names | KCNK13 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 13 (Tandem pore domain halothane- inhibited potassium channel 1) (THIK-1). | |||||
|
MSX1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 94) | NC score | 0.001176 (rank : 127) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P28360, Q96NY4 | Gene names | MSX1, HOX7 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein MSX-1 (Msh homeobox 1-like protein) (Hox-7). | |||||
|
REEP6_HUMAN
|
||||||
| θ value | 5.27518 (rank : 95) | NC score | 0.009595 (rank : 107) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96HR9, Q96LM0 | Gene names | REEP6, C19orf32, DP1L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Receptor expression-enhancing protein 6 (Polyposis locus protein 1- like 1). | |||||
|
TAOK1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 96) | NC score | 0.003656 (rank : 121) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 1734 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q7L7X3, Q96L75, Q9H2K7, Q9H7S5, Q9P2I6 | Gene names | TAOK1, KIAA1361, MAP3K16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase TAO1 (EC 2.7.11.1) (Thousand and one amino acid protein 1) (STE20-like kinase PSK2) (Kinase from chicken homolog B) (hKFC-B). | |||||
|
TAOK1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 97) | NC score | 0.003676 (rank : 120) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 1720 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q5F2E8, Q69ZL2, Q8JZX2, Q91VG7 | Gene names | Taok1, Kiaa1361 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase TAO1 (EC 2.7.11.1) (Thousand and one amino acid protein 1). | |||||
|
WDR33_HUMAN
|
||||||
| θ value | 5.27518 (rank : 98) | NC score | 0.002749 (rank : 126) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 638 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9C0J8 | Gene names | WDR33, WDC146 | |||
|
Domain Architecture |

|
|||||
| Description | WD repeat protein 33 (WD repeat protein WDC146). | |||||
|
CO6A2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 99) | NC score | 0.006679 (rank : 112) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q02788, Q05505 | Gene names | Col6a2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(VI) chain precursor. | |||||
|
GPR84_MOUSE
|
||||||
| θ value | 6.88961 (rank : 100) | NC score | -0.000329 (rank : 128) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 703 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CIM5, Q99MX9 | Gene names | Gpr84 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 84. | |||||
|
KCNH4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 101) | NC score | 0.029297 (rank : 90) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9UQ05 | Gene names | KCNH4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 4 (Voltage-gated potassium channel subunit Kv12.3) (Ether-a-go-go-like potassium channel 1) (ELK channel 1) (ELK1) (Brain-specific eag-like channel 2) (BEC2). | |||||
|
KCNKA_HUMAN
|
||||||
| θ value | 6.88961 (rank : 102) | NC score | 0.117382 (rank : 61) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | P57789, Q8TDK7, Q8TDK8, Q9HB59 | Gene names | KCNK10, TREK2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 10 (Outward rectifying potassium channel protein TREK-2) (TREK-2 K(+) channel subunit). | |||||
|
KCNKD_MOUSE
|
||||||
| θ value | 6.88961 (rank : 103) | NC score | 0.061911 (rank : 77) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8R1P5 | Gene names | Kcnk13 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 13 (Tandem pore domain halothane- inhibited potassium channel 1) (THIK-1). | |||||
|
KCNKG_HUMAN
|
||||||
| θ value | 6.88961 (rank : 104) | NC score | 0.090422 (rank : 69) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q96T55, Q5TCF3, Q9H591 | Gene names | KCNK16, TALK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 16 (TWIK-related alkaline pH- activated K(+) channel 1) (2P domain potassium channel Talk-1). | |||||
|
METL6_MOUSE
|
||||||
| θ value | 6.88961 (rank : 105) | NC score | 0.010064 (rank : 105) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BVH9, Q9D9M6, Q9DAX6 | Gene names | Mettl6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyltransferase-like protein 6 (EC 2.1.1.-). | |||||
|
MYH9_HUMAN
|
||||||
| θ value | 6.88961 (rank : 106) | NC score | 0.006591 (rank : 113) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P35579, O60805 | Gene names | MYH9 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
MYH9_MOUSE
|
||||||
| θ value | 6.88961 (rank : 107) | NC score | 0.006335 (rank : 114) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 1829 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8VDD5 | Gene names | Myh9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
PCLO_MOUSE
|
||||||
| θ value | 6.88961 (rank : 108) | NC score | 0.010870 (rank : 103) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
PKP4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 109) | NC score | 0.004686 (rank : 116) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99569 | Gene names | PKP4 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-4 (p0071). | |||||
|
RSMB_MOUSE
|
||||||
| θ value | 6.88961 (rank : 110) | NC score | 0.015224 (rank : 96) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P27048, Q3UJT1 | Gene names | Snrpb | |||
|
Domain Architecture |
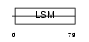
|
|||||
| Description | Small nuclear ribonucleoprotein-associated protein B (snRNP-B) (Sm protein B) (Sm-B) (SmB). | |||||
|
TPR_HUMAN
|
||||||
| θ value | 6.88961 (rank : 111) | NC score | 0.009059 (rank : 108) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P12270, Q15655 | Gene names | TPR | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoprotein TPR. | |||||
|
B3A3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 112) | NC score | 0.003194 (rank : 123) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P16283 | Gene names | Slc4a3, Ae3 | |||
|
Domain Architecture |

|
|||||
| Description | Anion exchange protein 3 (Neuronal band 3-like protein) (Solute carrier family 4 member 3). | |||||
|
KCNK2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 113) | NC score | 0.078082 (rank : 73) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | O95069, Q9UNE3 | Gene names | KCNK2, TREK, TREK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 2 (Outward rectifying potassium channel protein TREK-1) (TREK-1 K(+) channel subunit) (Two-pore potassium channel TPKC1). | |||||
|
KCNKF_HUMAN
|
||||||
| θ value | 8.99809 (rank : 114) | NC score | 0.116985 (rank : 62) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q9H427, Q9HBC8 | Gene names | KCNK15, TASK5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 15 (Acid-sensitive potassium channel protein TASK-5) (TWIK-related acid-sensitive K(+) channel 5) (Two pore potassium channel KT3.3). | |||||
|
RIMB1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 115) | NC score | 0.006815 (rank : 111) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 1376 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q7TNF8, Q5NCP7, Q80TV9, Q8BIH5 | Gene names | Bzrap1, Kiaa0612, Rbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Peripheral-type benzodiazepine receptor-associated protein 1 (PRAX-1) (Peripheral benzodiazepine receptor-interacting protein) (PBR-IP) (RIM-binding protein 1) (RIM-BP1). | |||||
|
SF3B4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 116) | NC score | 0.007415 (rank : 109) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q15427 | Gene names | SF3B4, SAP49 | |||
|
Domain Architecture |
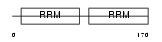
|
|||||
| Description | Splicing factor 3B subunit 4 (Spliceosome-associated protein 49) (SAP 49) (SF3b50) (Pre-mRNA-splicing factor SF3b 49 kDa subunit). | |||||
|
STK36_MOUSE
|
||||||
| θ value | 8.99809 (rank : 117) | NC score | 0.002758 (rank : 125) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 877 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q69ZM6, Q6PDI0, Q80XQ6 | Gene names | Stk36, Kiaa1278 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase 36 (EC 2.7.11.1) (Fused homolog). | |||||
|
KCD15_HUMAN
|
||||||
| θ value | θ > 10 (rank : 118) | NC score | 0.053561 (rank : 83) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q96SI1, Q9BVI6 | Gene names | KCTD15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein KCTD15. | |||||
|
KCD15_MOUSE
|
||||||
| θ value | θ > 10 (rank : 119) | NC score | 0.051320 (rank : 85) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8K0E1 | Gene names | Kctd15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein KCTD15. | |||||
|
KCNK4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 120) | NC score | 0.062355 (rank : 75) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9NYG8, Q96T94 | Gene names | KCNK4, TRAAK | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 4 (TWIK-related arachidonic acid- stimulated potassium channel protein) (TRAAK) (Two pore K(+) channel KT4.1). | |||||
|
KCNK7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 121) | NC score | 0.052023 (rank : 84) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9Y2U2, Q9Y2U3, Q9Y2U4 | Gene names | KCNK7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 7. | |||||
|
KCNK7_MOUSE
|
||||||
| θ value | θ > 10 (rank : 122) | NC score | 0.056195 (rank : 78) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9Z2T1, Q9QXY0, Q9QYE8, Q9R1V1, Q9R242 | Gene names | Kcnk7, Dpkch3, Kcnk6, Kcnk8, Knot1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 7 (Putative potassium channel DP3) (Double-pore K(+) channel 3) (Neuromuscular two p domain potassium channel). | |||||
|
KCNKH_HUMAN
|
||||||
| θ value | θ > 10 (rank : 123) | NC score | 0.091109 (rank : 68) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q96T54, Q5TCF4, Q8TAW4, Q9BXD1, Q9H592 | Gene names | KCNK17, TALK2, TASK4 | |||
|
Domain Architecture |
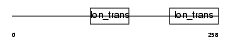
|
|||||
| Description | Potassium channel subfamily K member 17 (TWIK-related alkaline pH- activated K(+) channel 2) (2P domain potassium channel Talk-2) (TWIK- related acid-sensitive K(+) channel 4) (TASK-4). | |||||
|
KCNT1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 124) | NC score | 0.083112 (rank : 71) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q5JUK3, Q9P2C5 | Gene names | KCNT1, KIAA1422 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily T member 1 (KCa4.1). | |||||
|
KCTD1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 125) | NC score | 0.054823 (rank : 79) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q719H9 | Gene names | KCTD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein KCTD1. | |||||
|
KCTD1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 126) | NC score | 0.054823 (rank : 80) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q5M956, Q5EER9 | Gene names | Kctd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein KCTD1. | |||||
|
KCTD4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 127) | NC score | 0.050196 (rank : 86) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9D7X1, Q8CCQ3, Q9CYK4 | Gene names | Kctd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein KCTD4. | |||||
|
KCTD6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 128) | NC score | 0.054195 (rank : 82) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8NC69, Q8NBS6, Q8TCA6 | Gene names | KCTD6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein KCTD6. | |||||
|
KCTD6_MOUSE
|
||||||
| θ value | θ > 10 (rank : 129) | NC score | 0.054558 (rank : 81) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8BNL5, Q3TXX1 | Gene names | Kctd6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein KCTD6. | |||||
|
KCNN1_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 117 | |
| SwissProt Accessions | Q9EQR3, Q99JA9 | Gene names | Kcnn1, Sk1 | |||
|
Domain Architecture |

|
|||||
| Description | Small conductance calcium-activated potassium channel protein 1 (SK1). | |||||
|
KCNN1_HUMAN
|
||||||
| NC score | 0.991905 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 85 | |
| SwissProt Accessions | Q92952 | Gene names | KCNN1, SK | |||
|
Domain Architecture |

|
|||||
| Description | Small conductance calcium-activated potassium channel protein 1 (SK1). | |||||
|
KCNN2_HUMAN
|
||||||
| NC score | 0.989156 (rank : 3) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | Q9H2S1 | Gene names | KCNN2 | |||
|
Domain Architecture |

|
|||||
| Description | Small conductance calcium-activated potassium channel protein 2 (SK2). | |||||
|
KCNN2_MOUSE
|
||||||
| NC score | 0.988273 (rank : 4) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | P58390 | Gene names | Kcnn2, Sk2 | |||
|
Domain Architecture |

|
|||||
| Description | Small conductance calcium-activated potassium channel protein 2 (SK2). | |||||
|
KCNN3_MOUSE
|
||||||
| NC score | 0.979850 (rank : 5) | θ value | 0 (rank : 6) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | P58391 | Gene names | Kcnn3, Sk3 | |||
|
Domain Architecture |
|
|||||
| Description | Small conductance calcium-activated potassium channel protein 3 (SK3). | |||||
|
KCNN3_HUMAN
|
||||||
| NC score | 0.974949 (rank : 6) | θ value | 0 (rank : 5) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 78 | |
| SwissProt Accessions | Q9UGI6, O43517 | Gene names | KCNN3, K3 | |||
|
Domain Architecture |
|
|||||
| Description | Small conductance calcium-activated potassium channel protein 3 (SK3) (SKCa3). | |||||
|
KCNN4_MOUSE
|
||||||
| NC score | 0.951391 (rank : 7) | θ value | 1.49966e-79 (rank : 7) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | O89109, Q3U1J8, Q9CY39 | Gene names | Kcnn4 | |||
|
Domain Architecture |

|
|||||
| Description | Intermediate conductance calcium-activated potassium channel protein 4 (SK4) (KCa4) (IK1). | |||||
|
KCNN4_HUMAN
|
||||||
| NC score | 0.949220 (rank : 8) | θ value | 6.30065e-78 (rank : 8) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | O15554 | Gene names | KCNN4, IK1, IKCA1, KCA4, SK4 | |||
|
Domain Architecture |

|
|||||
| Description | Intermediate conductance calcium-activated potassium channel protein 4 (SK4) (KCa4) (IK1) (IKCa1) (Putative Gardos channel). | |||||
|
KCNQ4_HUMAN
|
||||||
| NC score | 0.366981 (rank : 9) | θ value | 1.43324e-05 (rank : 10) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | P56696, O96025 | Gene names | KCNQ4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 4 (Voltage-gated potassium channel subunit Kv7.4) (Potassium channel subunit alpha KvLQT4) (KQT-like 4). | |||||
|
KCNQ5_HUMAN
|
||||||
| NC score | 0.362020 (rank : 10) | θ value | 8.40245e-06 (rank : 9) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q9NR82, Q9NRN0, Q9NYA6 | Gene names | KCNQ5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 5 (Voltage-gated potassium channel subunit Kv7.5) (Potassium channel subunit alpha KvLQT5) (KQT-like 5). | |||||
|
KCNQ2_MOUSE
|
||||||
| NC score | 0.344256 (rank : 11) | θ value | 0.00020696 (rank : 11) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q9Z351, Q8R498, Q9QWN9, Q9Z342, Q9Z343, Q9Z344, Q9Z345, Q9Z346, Q9Z347, Q9Z348, Q9Z349, Q9Z350 | Gene names | Kcnq2, Kqt2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 2 (Voltage-gated potassium channel subunit Kv7.2) (Potassium channel subunit alpha KvLQT2) (KQT-like 2). | |||||
|
KCNQ1_HUMAN
|
||||||
| NC score | 0.342836 (rank : 12) | θ value | 0.00298849 (rank : 22) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | P51787, O00347, O60607, O94787, Q7Z6G9, Q92960, Q9UMN8, Q9UMN9 | Gene names | KCNQ1, KCNA8, KCNA9, KVLQT1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 1 (Voltage-gated potassium channel subunit Kv7.1) (IKs producing slow voltage-gated potassium channel subunit alpha KvLQT1) (KQT-like 1). | |||||
|
KCNQ1_MOUSE
|
||||||
| NC score | 0.341715 (rank : 13) | θ value | 0.00228821 (rank : 21) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | P97414, O88702, Q7TNZ1, Q7TPL7, Q80VR7 | Gene names | Kcnq1, Kcna9, Kvlqt1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 1 (Voltage-gated potassium channel subunit Kv7.1) (IKs producing slow voltage-gated potassium channel subunit alpha KvLQT1) (KQT-like 1). | |||||
|
KCNQ2_HUMAN
|
||||||
| NC score | 0.341467 (rank : 14) | θ value | 0.000786445 (rank : 16) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | O43526, O43796, O75580, O95845, Q5VYT8, Q96J59, Q99454 | Gene names | KCNQ2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 2 (Voltage-gated potassium channel subunit Kv7.2) (Neuroblastoma-specific potassium channel subunit alpha KvLQT2) (KQT-like 2). | |||||
|
KCNQ3_HUMAN
|
||||||
| NC score | 0.336119 (rank : 15) | θ value | 0.00102713 (rank : 17) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | O43525 | Gene names | KCNQ3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 3 (Voltage-gated potassium channel subunit Kv7.3) (Potassium channel subunit alpha KvLQT3) (KQT-like 3). | |||||
|
KCNQ3_MOUSE
|
||||||
| NC score | 0.332863 (rank : 16) | θ value | 0.00175202 (rank : 20) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q8K3F6 | Gene names | Kcnq3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium voltage-gated channel subfamily KQT member 3 (Voltage-gated potassium channel subunit Kv7.3) (Potassium channel subunit alpha KvLQT3) (KQT-like 3). | |||||
|
KCND2_HUMAN
|
||||||
| NC score | 0.296717 (rank : 17) | θ value | 0.000461057 (rank : 13) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q9NZV8, O95012, O95021, Q9UBY7, Q9UN98, Q9UNH9 | Gene names | KCND2, KIAA1044 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 2 (Voltage-gated potassium channel subunit Kv4.2). | |||||
|
KCND2_MOUSE
|
||||||
| NC score | 0.296717 (rank : 18) | θ value | 0.000461057 (rank : 14) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q9Z0V2, Q8BSK3, Q8CHB7, Q9JJ60 | Gene names | Kcnd2, Kiaa1044 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 2 (Voltage-gated potassium channel subunit Kv4.2). | |||||
|
KCND1_MOUSE
|
||||||
| NC score | 0.296664 (rank : 19) | θ value | 0.000461057 (rank : 12) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q03719, Q8CC68 | Gene names | Kcnd1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 1 (Voltage-gated potassium channel subunit Kv4.1) (mShal). | |||||
|
KCND1_HUMAN
|
||||||
| NC score | 0.296653 (rank : 20) | θ value | 0.000602161 (rank : 15) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 160 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q9NSA2, O75671 | Gene names | KCND1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 1 (Voltage-gated potassium channel subunit Kv4.1). | |||||
|
KCND3_MOUSE
|
||||||
| NC score | 0.292600 (rank : 21) | θ value | 0.00134147 (rank : 19) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q9Z0V1, Q8CC44, Q9Z0V0 | Gene names | Kcnd3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 3 (Voltage-gated potassium channel subunit Kv4.3). | |||||
|
KCND3_HUMAN
|
||||||
| NC score | 0.292471 (rank : 22) | θ value | 0.00134147 (rank : 18) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q9UK17, O60576, O60577, Q5T0M0, Q9UH85, Q9UH86, Q9UK16 | Gene names | KCND3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily D member 3 (Voltage-gated potassium channel subunit Kv4.3). | |||||
|
KCNS2_MOUSE
|
||||||
| NC score | 0.291765 (rank : 23) | θ value | 0.0113563 (rank : 27) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | O35174, Q543P3 | Gene names | Kcns2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily S member 2 (Voltage-gated potassium channel subunit Kv9.2) (Delayed-rectifier K(+) channel alpha subunit 2). | |||||
|
KCNS2_HUMAN
|
||||||
| NC score | 0.290813 (rank : 24) | θ value | 0.0113563 (rank : 26) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q9ULS6 | Gene names | KCNS2, KIAA1144 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily S member 2 (Voltage-gated potassium channel subunit Kv9.2) (Delayed-rectifier K(+) channel alpha subunit 2). | |||||
|
KCNA6_HUMAN
|
||||||
| NC score | 0.289035 (rank : 25) | θ value | 0.00665767 (rank : 24) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | P17658 | Gene names | KCNA6 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 6 (Voltage-gated potassium channel subunit Kv1.6) (HBK2). | |||||
|
KCNA6_MOUSE
|
||||||
| NC score | 0.286528 (rank : 26) | θ value | 0.00869519 (rank : 25) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q61923 | Gene names | Kcna6 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 6 (Voltage-gated potassium channel subunit Kv1.6) (MK1.6). | |||||
|
KCNA5_MOUSE
|
||||||
| NC score | 0.283380 (rank : 27) | θ value | 0.0252991 (rank : 31) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q61762 | Gene names | Kcna5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 5 (Voltage-gated potassium channel subunit Kv1.5) (KV1-5). | |||||
|
KCNB2_HUMAN
|
||||||
| NC score | 0.280444 (rank : 28) | θ value | 0.00390308 (rank : 23) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q92953, Q9BXD3 | Gene names | KCNB2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily B member 2 (Voltage-gated potassium channel subunit Kv2.2). | |||||
|
KCNA2_HUMAN
|
||||||
| NC score | 0.279516 (rank : 29) | θ value | 0.0330416 (rank : 32) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | P16389 | Gene names | KCNA2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 2 (Voltage-gated potassium channel subunit Kv1.2) (HBK5) (NGK1) (HUKIV). | |||||
|
KCNA5_HUMAN
|
||||||
| NC score | 0.278524 (rank : 30) | θ value | 0.0563607 (rank : 41) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | P22460 | Gene names | KCNA5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 5 (Voltage-gated potassium channel subunit Kv1.5) (HK2) (HPCN1). | |||||
|
KCNA4_HUMAN
|
||||||
| NC score | 0.277448 (rank : 31) | θ value | 0.0252991 (rank : 30) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | P22459 | Gene names | KCNA4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 4 (Voltage-gated potassium channel subunit Kv1.4) (HK1) (HPCN2) (HBK4) (HUKII). | |||||
|
KCNA3_HUMAN
|
||||||
| NC score | 0.277439 (rank : 32) | θ value | 0.0563607 (rank : 38) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | P22001 | Gene names | KCNA3, HGK5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 3 (Voltage-gated potassium channel subunit Kv1.3) (HPCN3) (HGK5) (HuKIII) (HLK3). | |||||
|
KCNA2_MOUSE
|
||||||
| NC score | 0.277224 (rank : 33) | θ value | 0.0431538 (rank : 36) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | P63141, P15386, Q02010, Q8C8W4 | Gene names | Kcna2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium voltage-gated channel subfamily A member 2 (Voltage-gated potassium channel subunit Kv1.2) (MK2). | |||||
|
KCNA3_MOUSE
|
||||||
| NC score | 0.276846 (rank : 34) | θ value | 0.0563607 (rank : 39) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | P16390 | Gene names | Kcna3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 3 (Voltage-gated potassium channel subunit Kv1.3) (MK3). | |||||
|
KCNB1_MOUSE
|
||||||
| NC score | 0.272742 (rank : 35) | θ value | 0.0193708 (rank : 29) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q03717 | Gene names | Kcnb1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily B member 1 (Voltage-gated potassium channel subunit Kv2.1) (mShab). | |||||
|
KCNB1_HUMAN
|
||||||
| NC score | 0.272626 (rank : 36) | θ value | 0.0193708 (rank : 28) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q14721, Q14193 | Gene names | KCNB1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily B member 1 (Voltage-gated potassium channel subunit Kv2.1) (h-DRK1). | |||||
|
KCNA1_HUMAN
|
||||||
| NC score | 0.272423 (rank : 37) | θ value | 0.0961366 (rank : 43) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q09470 | Gene names | KCNA1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 1 (Voltage-gated potassium channel subunit Kv1.1) (HUKI) (HBK1). | |||||
|
KCNA4_MOUSE
|
||||||
| NC score | 0.272328 (rank : 38) | θ value | 0.0563607 (rank : 40) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q61423 | Gene names | Kcna4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 4 (Voltage-gated potassium channel subunit Kv1.4). | |||||
|
KCNA1_MOUSE
|
||||||
| NC score | 0.272180 (rank : 39) | θ value | 0.0961366 (rank : 44) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | P16388 | Gene names | Kcna1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily A member 1 (Voltage-gated potassium channel subunit Kv1.1) (MKI) (MBK1). | |||||
|
KCNS3_HUMAN
|
||||||
| NC score | 0.271204 (rank : 40) | θ value | 0.125558 (rank : 46) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9BQ31, O43651, Q96B56 | Gene names | KCNS3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily S member 3 (Voltage-gated potassium channel subunit Kv9.3) (Delayed-rectifier K(+) channel alpha subunit 3). | |||||
|
KCNG3_HUMAN
|
||||||
| NC score | 0.265816 (rank : 41) | θ value | 0.0330416 (rank : 33) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q8TAE7 | Gene names | KCNG3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily G member 3 (Voltage-gated potassium channel subunit Kv6.3) (Kv10.1). | |||||
|
KCNG3_MOUSE
|
||||||
| NC score | 0.265590 (rank : 42) | θ value | 0.0330416 (rank : 34) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | P59053 | Gene names | Kcng3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily G member 3 (Voltage-gated potassium channel subunit Kv6.3) (Kv10.1). | |||||
|
KCNG2_HUMAN
|
||||||
| NC score | 0.263530 (rank : 43) | θ value | 0.163984 (rank : 48) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9UJ96 | Gene names | KCNG2, KCNF2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily G member 2 (Voltage-gated potassium channel subunit Kv6.2) (Cardiac potassium channel subunit). | |||||
|
KCNS1_HUMAN
|
||||||
| NC score | 0.256616 (rank : 44) | θ value | 0.813845 (rank : 65) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q96KK3, O43652, Q6DJU6 | Gene names | KCNS1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily S member 1 (Voltage-gated potassium channel subunit Kv9.1) (Delayed-rectifier K(+) channel alpha subunit 1). | |||||
|
KCNG1_HUMAN
|
||||||
| NC score | 0.256548 (rank : 45) | θ value | 0.279714 (rank : 51) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q9UIX4, O43528, Q9BRC1 | Gene names | KCNG1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily G member 1 (Voltage-gated potassium channel subunit Kv6.1) (kH2). | |||||
|
KCNC4_MOUSE
|
||||||
| NC score | 0.255883 (rank : 46) | θ value | 0.365318 (rank : 55) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q8R1C0 | Gene names | Kcnc4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium voltage-gated channel subfamily C member 4 (Voltage-gated potassium channel subunit Kv3.4). | |||||
|
KCNC4_HUMAN
|
||||||
| NC score | 0.255227 (rank : 47) | θ value | 0.365318 (rank : 54) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q03721, Q3MIM4, Q5TBI6 | Gene names | KCNC4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily C member 4 (Voltage-gated potassium channel subunit Kv3.4) (KSHIIIC). | |||||
|
KCNF1_HUMAN
|
||||||
| NC score | 0.250794 (rank : 48) | θ value | 1.81305 (rank : 71) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q9H3M0, O43527 | Gene names | KCNF1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily F member 1 (Voltage-gated potassium channel subunit Kv5.1) (kH1). | |||||
|
KCNC1_MOUSE
|
||||||
| NC score | 0.248691 (rank : 49) | θ value | 0.365318 (rank : 53) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | P15388 | Gene names | Kcnc1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily C member 1 (Voltage-gated potassium channel subunit Kv3.1) (Kv4) (NGK2). | |||||
|
KCNC1_HUMAN
|
||||||
| NC score | 0.248632 (rank : 50) | θ value | 0.365318 (rank : 52) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | P48547 | Gene names | KCNC1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily C member 1 (Voltage-gated potassium channel subunit Kv3.1) (Kv4) (NGK2). | |||||
|
KCNG4_HUMAN
|
||||||
| NC score | 0.248413 (rank : 51) | θ value | 0.62314 (rank : 63) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q8TDN1, Q96H24 | Gene names | KCNG4, KCNG3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily G member 4 (Voltage-gated potassium channel subunit Kv6.4). | |||||
|
KCNC3_MOUSE
|
||||||
| NC score | 0.248264 (rank : 52) | θ value | 0.279714 (rank : 50) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q63959, Q62088 | Gene names | Kcnc3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily C member 3 (Voltage-gated potassium channel subunit Kv3.3) (KSHIIID). | |||||
|
KCNS1_MOUSE
|
||||||
| NC score | 0.246795 (rank : 53) | θ value | 1.06291 (rank : 67) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | O35173 | Gene names | Kcns1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily S member 1 (Voltage-gated potassium channel subunit Kv9.1) (Delayed-rectifier K(+) channel alpha subunit 1). | |||||
|
KCNV2_HUMAN
|
||||||
| NC score | 0.245385 (rank : 54) | θ value | 2.36792 (rank : 76) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q8TDN2 | Gene names | KCNV2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily V member 2 (Voltage-gated potassium channel subunit Kv8.2). | |||||
|
KCNC3_HUMAN
|
||||||
| NC score | 0.245026 (rank : 55) | θ value | 0.47712 (rank : 61) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 256 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q14003 | Gene names | KCNC3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily C member 3 (Voltage-gated potassium channel subunit Kv3.3) (KSHIIID). | |||||
|
KCMA1_MOUSE
|
||||||
| NC score | 0.210468 (rank : 56) | θ value | 0.47712 (rank : 60) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q08460, Q64703, Q8VHF1, Q9R196 | Gene names | Kcnma1, Kcnma | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-activated potassium channel subunit alpha 1 (Calcium-activated potassium channel, subfamily M subunit alpha 1) (Maxi K channel) (MaxiK) (BK channel) (K(VCA)alpha) (BKCA alpha) (KCa1.1) (Slowpoke homolog) (Slo homolog) (Slo-alpha) (Slo1) (mSlo) (mSlo1). | |||||
|
KCMA1_HUMAN
|
||||||
| NC score | 0.210084 (rank : 57) | θ value | 0.47712 (rank : 59) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q12791, Q12886, Q12917, Q12921, Q12960, Q13150, Q96LG8, Q9UBB0, Q9UQK6 | Gene names | KCNMA1, KCNMA | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-activated potassium channel subunit alpha 1 (Calcium-activated potassium channel, subfamily M subunit alpha 1) (Maxi K channel) (MaxiK) (BK channel) (K(VCA)alpha) (BKCA alpha) (KCa1.1) (Slowpoke homolog) (Slo homolog) (Slo-alpha) (Slo1) (hSlo). | |||||
|
KCNK1_HUMAN
|
||||||
| NC score | 0.138466 (rank : 58) | θ value | 0.125558 (rank : 45) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O00180, Q13307 | Gene names | KCNK1, HOHO1, KCNO1, TWIK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 1 (Inward rectifying potassium channel protein TWIK-1) (Potassium channel KCNO1). | |||||
|
KCNK6_HUMAN
|
||||||
| NC score | 0.137269 (rank : 59) | θ value | 0.47712 (rank : 62) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q9Y257, Q9HB47 | Gene names | KCNK6, TOSS, TWIK2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 6 (Inward rectifying potassium channel protein TWIK-2) (TWIK-originated similarity sequence). | |||||
|
KCNK1_MOUSE
|
||||||
| NC score | 0.123573 (rank : 60) | θ value | 0.365318 (rank : 56) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | O08581 | Gene names | Kcnk1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 1 (Inward rectifying potassium channel protein TWIK-1). | |||||
|
KCNKA_HUMAN
|
||||||
| NC score | 0.117382 (rank : 61) | θ value | 6.88961 (rank : 102) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | P57789, Q8TDK7, Q8TDK8, Q9HB59 | Gene names | KCNK10, TREK2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 10 (Outward rectifying potassium channel protein TREK-2) (TREK-2 K(+) channel subunit). | |||||
|
KCNKF_HUMAN
|
||||||
| NC score | 0.116985 (rank : 62) | θ value | 8.99809 (rank : 114) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q9H427, Q9HBC8 | Gene names | KCNK15, TASK5 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 15 (Acid-sensitive potassium channel protein TASK-5) (TWIK-related acid-sensitive K(+) channel 5) (Two pore potassium channel KT3.3). | |||||
|
KCNK5_HUMAN
|
||||||
| NC score | 0.115484 (rank : 63) | θ value | 5.27518 (rank : 92) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | O95279 | Gene names | KCNK5, TASK2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 5 (Acid-sensitive potassium channel protein TASK-2) (TWIK-related acid-sensitive K(+) channel 2). | |||||
|
KCNK9_HUMAN
|
||||||
| NC score | 0.111507 (rank : 64) | θ value | 1.81305 (rank : 73) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9NPC2, Q2M290, Q540F2 | Gene names | KCNK9, TASK3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 9 (Acid-sensitive potassium channel protein TASK-3) (TWIK-related acid-sensitive K(+) channel 3) (Two pore potassium channel KT3.2). | |||||
|
KCNK9_MOUSE
|
||||||
| NC score | 0.111136 (rank : 65) | θ value | 1.81305 (rank : 74) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q3LS21 | Gene names | Kcnk9, Task3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 9 (Acid-sensitive potassium channel protein TASK-3) (TWIK-related acid-sensitive K(+) channel 3). | |||||
|
KCNK3_HUMAN
|
||||||
| NC score | 0.100016 (rank : 66) | θ value | 5.27518 (rank : 90) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | O14649 | Gene names | KCNK3, TASK, TASK1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 3 (Acid-sensitive potassium channel protein TASK-1) (TWIK-related acid-sensitive K(+) channel 1) (Two pore potassium channel KT3.1). | |||||
|
KCNK3_MOUSE
|
||||||
| NC score | 0.099963 (rank : 67) | θ value | 5.27518 (rank : 91) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | O35111, O35163 | Gene names | Kcnk3, Ctbak, Task, Task1 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 3 (Acid-sensitive potassium channel protein TASK-1) (TWIK-related acid-sensitive K(+) channel 1) (Cardiac two-pore background K(+) channel) (cTBAK-1) (Two pore potassium channel KT3.1). | |||||
|
KCNKH_HUMAN
|
||||||
| NC score | 0.091109 (rank : 68) | θ value | θ > 10 (rank : 123) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q96T54, Q5TCF4, Q8TAW4, Q9BXD1, Q9H592 | Gene names | KCNK17, TALK2, TASK4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 17 (TWIK-related alkaline pH- activated K(+) channel 2) (2P domain potassium channel Talk-2) (TWIK- related acid-sensitive K(+) channel 4) (TASK-4). | |||||
|
KCNKG_HUMAN
|
||||||
| NC score | 0.090422 (rank : 69) | θ value | 6.88961 (rank : 104) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q96T55, Q5TCF3, Q9H591 | Gene names | KCNK16, TALK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 16 (TWIK-related alkaline pH- activated K(+) channel 1) (2P domain potassium channel Talk-1). | |||||
|
KCNK4_MOUSE
|
||||||
| NC score | 0.084871 (rank : 70) | θ value | 1.81305 (rank : 72) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | O88454 | Gene names | Kcnk4, Traak | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 4 (TWIK-related arachidonic acid- stimulated potassium channel protein) (TRAAK). | |||||
|
KCNT1_HUMAN
|
||||||
| NC score | 0.083112 (rank : 71) | θ value | θ > 10 (rank : 124) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q5JUK3, Q9P2C5 | Gene names | KCNT1, KIAA1422 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily T member 1 (KCa4.1). | |||||
|
KCNK2_MOUSE
|
||||||
| NC score | 0.082776 (rank : 72) | θ value | 5.27518 (rank : 89) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | P97438 | Gene names | Kcnk2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 2 (Outward rectifying potassium channel protein TREK-1) (Two-pore potassium channel TPKC1) (TREK-1 K(+) channel subunit). | |||||
|
KCNK2_HUMAN
|
||||||
| NC score | 0.078082 (rank : 73) | θ value | 8.99809 (rank : 113) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | O95069, Q9UNE3 | Gene names | KCNK2, TREK, TREK1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 2 (Outward rectifying potassium channel protein TREK-1) (TREK-1 K(+) channel subunit) (Two-pore potassium channel TPKC1). | |||||
|
KCNKD_HUMAN
|
||||||
| NC score | 0.069033 (rank : 74) | θ value | 5.27518 (rank : 93) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9HB14, Q96E79 | Gene names | KCNK13 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 13 (Tandem pore domain halothane- inhibited potassium channel 1) (THIK-1). | |||||
|
KCNK4_HUMAN
|
||||||
| NC score | 0.062355 (rank : 75) | θ value | θ > 10 (rank : 120) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q9NYG8, Q96T94 | Gene names | KCNK4, TRAAK | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 4 (TWIK-related arachidonic acid- stimulated potassium channel protein) (TRAAK) (Two pore K(+) channel KT4.1). | |||||
|
RAB3I_MOUSE
|
||||||
| NC score | 0.062146 (rank : 76) | θ value | 0.0330416 (rank : 35) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q68EF0 | Gene names | Rab3ip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RAB3A-interacting protein (Rabin-3) (SSX2-interacting protein). | |||||
|
KCNKD_MOUSE
|
||||||
| NC score | 0.061911 (rank : 77) | θ value | 6.88961 (rank : 103) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8R1P5 | Gene names | Kcnk13 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium channel subfamily K member 13 (Tandem pore domain halothane- inhibited potassium channel 1) (THIK-1). | |||||
|
KCNK7_MOUSE
|
||||||
| NC score | 0.056195 (rank : 78) | θ value | θ > 10 (rank : 122) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9Z2T1, Q9QXY0, Q9QYE8, Q9R1V1, Q9R242 | Gene names | Kcnk7, Dpkch3, Kcnk6, Kcnk8, Knot1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 7 (Putative potassium channel DP3) (Double-pore K(+) channel 3) (Neuromuscular two p domain potassium channel). | |||||
|
KCTD1_HUMAN
|
||||||
| NC score | 0.054823 (rank : 79) | θ value | θ > 10 (rank : 125) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q719H9 | Gene names | KCTD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein KCTD1. | |||||
|
KCTD1_MOUSE
|
||||||
| NC score | 0.054823 (rank : 80) | θ value | θ > 10 (rank : 126) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q5M956, Q5EER9 | Gene names | Kctd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein KCTD1. | |||||
|
KCTD6_MOUSE
|
||||||
| NC score | 0.054558 (rank : 81) | θ value | θ > 10 (rank : 129) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8BNL5, Q3TXX1 | Gene names | Kctd6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein KCTD6. | |||||
|
KCTD6_HUMAN
|
||||||
| NC score | 0.054195 (rank : 82) | θ value | θ > 10 (rank : 128) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q8NC69, Q8NBS6, Q8TCA6 | Gene names | KCTD6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein KCTD6. | |||||
|
KCD15_HUMAN
|
||||||
| NC score | 0.053561 (rank : 83) | θ value | θ > 10 (rank : 118) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q96SI1, Q9BVI6 | Gene names | KCTD15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein KCTD15. | |||||
|
KCNK7_HUMAN
|
||||||
| NC score | 0.052023 (rank : 84) | θ value | θ > 10 (rank : 121) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9Y2U2, Q9Y2U3, Q9Y2U4 | Gene names | KCNK7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 7. | |||||
|
KCD15_MOUSE
|
||||||
| NC score | 0.051320 (rank : 85) | θ value | θ > 10 (rank : 119) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8K0E1 | Gene names | Kctd15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein KCTD15. | |||||
|
KCTD4_MOUSE
|
||||||
| NC score | 0.050196 (rank : 86) | θ value | θ > 10 (rank : 127) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9D7X1, Q8CCQ3, Q9CYK4 | Gene names | Kctd4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein KCTD4. | |||||
|
AAT1_HUMAN
|
||||||
| NC score | 0.044149 (rank : 87) | θ value | 4.03905 (rank : 80) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q7Z4T9, Q68DX2, Q8TD41, Q96A45, Q96JE8 | Gene names | AAT1, C3orf15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AMY-1-associating protein expressed in testis 1 (AAT-1). | |||||
|
RAB3I_HUMAN
|
||||||
| NC score | 0.036392 (rank : 88) | θ value | 2.36792 (rank : 77) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 270 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q96QF0, Q6PCE4, Q96A24, Q96QE6, Q96QE7, Q96QE8, Q96QE9, Q96QF1, Q9H673 | Gene names | RAB3IP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RAB3A-interacting protein (Rabin-3) (SSX2-interacting protein). | |||||
|
ANGL7_MOUSE
|
||||||
| NC score | 0.033426 (rank : 89) | θ value | 0.163984 (rank : 47) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8R1Q3 | Gene names | Angptl7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiopoietin-related protein 7 precursor (Angiopoietin-like 7). | |||||
|
KCNH4_HUMAN
|
||||||
| NC score | 0.029297 (rank : 90) | θ value | 6.88961 (rank : 101) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9UQ05 | Gene names | KCNH4 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 4 (Voltage-gated potassium channel subunit Kv12.3) (Ether-a-go-go-like potassium channel 1) (ELK channel 1) (ELK1) (Brain-specific eag-like channel 2) (BEC2). | |||||
|
BAZ2A_MOUSE
|
||||||
| NC score | 0.021634 (rank : 91) | θ value | 4.03905 (rank : 82) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q91YE5 | Gene names | Baz2a, Tip5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5). | |||||
|
RSMB_HUMAN
|
||||||
| NC score | 0.020195 (rank : 92) | θ value | 1.81305 (rank : 75) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P14678, Q15490, Q9UIS5 | Gene names | SNRPB, COD, SNRPB1 | |||
|
Domain Architecture |

|
|||||
| Description | Small nuclear ribonucleoprotein-associated proteins B and B' (snRNP-B) (Sm protein B/B') (Sm-B/Sm-B') (SmB/SmB'). | |||||
|
GOGA4_HUMAN
|
||||||
| NC score | 0.019181 (rank : 93) | θ value | 0.813845 (rank : 64) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 2034 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q13439, Q13270, Q13654, Q14436 | Gene names | GOLGA4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (Trans-Golgi p230) (256 kDa golgin) (Golgin-245) (Protein 72.1). | |||||
|
DIAP3_HUMAN
|
||||||
| NC score | 0.015518 (rank : 94) | θ value | 0.47712 (rank : 58) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 209 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NSV4, Q5T2S7, Q5XKF6, Q6MZF0, Q6NUP0, Q86VS4, Q8NAV4 | Gene names | DIAPH3, DIAP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein diaphanous homolog 3 (Diaphanous-related formin-3) (DRF3). | |||||
|
LMNB1_MOUSE
|
||||||
| NC score | 0.015225 (rank : 95) | θ value | 3.0926 (rank : 79) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 719 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P14733, Q61791 | Gene names | Lmnb1 | |||
|
Domain Architecture |

|
|||||
| Description | Lamin-B1. | |||||
|
RSMB_MOUSE
|
||||||
| NC score | 0.015224 (rank : 96) | θ value | 6.88961 (rank : 110) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P27048, Q3UJT1 | Gene names | Snrpb | |||
|
Domain Architecture |

|
|||||
| Description | Small nuclear ribonucleoprotein-associated protein B (snRNP-B) (Sm protein B) (Sm-B) (SmB). | |||||
|
ROCK2_HUMAN
|
||||||
| NC score | 0.015206 (rank : 97) | θ value | 0.0431538 (rank : 37) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 2073 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O75116, Q9UQN5 | Gene names | ROCK2, KIAA0619 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 2 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 2) (p164 ROCK-2) (Rho kinase 2). | |||||
|
LMNB1_HUMAN
|
||||||
| NC score | 0.015130 (rank : 98) | θ value | 4.03905 (rank : 83) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 714 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P20700, Q3SYN7, Q96EI6 | Gene names | LMNB1, LMN2, LMNB | |||
|
Domain Architecture |

|
|||||
| Description | Lamin-B1. | |||||
|
CUTL2_MOUSE
|
||||||
| NC score | 0.014045 (rank : 99) | θ value | 1.81305 (rank : 70) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 771 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P70298 | Gene names | Cutl2 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein cut-like 2 (Homeobox protein Cux-2) (Cut-like 2). | |||||
|
ROCK2_MOUSE
|
||||||
| NC score | 0.013746 (rank : 100) | θ value | 0.0563607 (rank : 42) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 2072 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P70336, Q8BR64, Q8CBR0 | Gene names | Rock2 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 2 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 2) (p164 ROCK-2). | |||||
|
BAZ2A_HUMAN
|
||||||
| NC score | 0.012298 (rank : 101) | θ value | 5.27518 (rank : 86) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 780 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9UIF9, O00536, O15030, Q96H26 | Gene names | BAZ2A, KIAA0314, TIP5 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain adjacent to zinc finger domain 2A (Transcription termination factor I-interacting protein 5) (TTF-I-interacting protein 5) (Tip5) (hWALp3). | |||||
|
AZI1_HUMAN
|
||||||
| NC score | 0.011757 (rank : 102) | θ value | 4.03905 (rank : 81) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 1000 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9UPN4, Q96F50 | Gene names | AZI1, CEP131, KIAA1118 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 5-azacytidine-induced protein 1 (Pre-acrosome localization protein 1) (Centrosomal protein of 131 kDa) (Cep131 protein). | |||||
|
PCLO_MOUSE
|
||||||
| NC score | 0.010870 (rank : 103) | θ value | 6.88961 (rank : 108) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
CCD41_MOUSE
|
||||||
| NC score | 0.010254 (rank : 104) | θ value | 1.81305 (rank : 69) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9D5R3, Q3U7X7, Q3UX57, Q80VF0 | Gene names | Ccdc41 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 41. | |||||
|
METL6_MOUSE
|
||||||
| NC score | 0.010064 (rank : 105) | θ value | 6.88961 (rank : 105) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BVH9, Q9D9M6, Q9DAX6 | Gene names | Mettl6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyltransferase-like protein 6 (EC 2.1.1.-). | |||||
|
ROCK1_MOUSE
|
||||||
| NC score | 0.009942 (rank : 106) | θ value | 0.365318 (rank : 57) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 2126 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P70335, Q8C3G4, Q8C7H0 | Gene names | Rock1 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 1 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 1) (p160 ROCK-1) (p160ROCK). | |||||
|
REEP6_HUMAN
|
||||||
| NC score | 0.009595 (rank : 107) | θ value | 5.27518 (rank : 95) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96HR9, Q96LM0 | Gene names | REEP6, C19orf32, DP1L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Receptor expression-enhancing protein 6 (Polyposis locus protein 1- like 1). | |||||
|
TPR_HUMAN
|
||||||
| NC score | 0.009059 (rank : 108) | θ value | 6.88961 (rank : 111) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 1921 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P12270, Q15655 | Gene names | TPR | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoprotein TPR. | |||||
|
SF3B4_HUMAN
|
||||||
| NC score | 0.007415 (rank : 109) | θ value | 8.99809 (rank : 116) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q15427 | Gene names | SF3B4, SAP49 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 3B subunit 4 (Spliceosome-associated protein 49) (SAP 49) (SF3b50) (Pre-mRNA-splicing factor SF3b 49 kDa subunit). | |||||
|
CO6A2_HUMAN
|
||||||
| NC score | 0.007145 (rank : 110) | θ value | 5.27518 (rank : 87) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P12110, Q13909, Q13910, Q13911, Q14049, Q16259, Q16597, Q9UML3 | Gene names | COL6A2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(VI) chain precursor. | |||||
|
RIMB1_MOUSE
|
||||||
| NC score | 0.006815 (rank : 111) | θ value | 8.99809 (rank : 115) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 1376 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q7TNF8, Q5NCP7, Q80TV9, Q8BIH5 | Gene names | Bzrap1, Kiaa0612, Rbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Peripheral-type benzodiazepine receptor-associated protein 1 (PRAX-1) (Peripheral benzodiazepine receptor-interacting protein) (PBR-IP) (RIM-binding protein 1) (RIM-BP1). | |||||
|
CO6A2_MOUSE
|
||||||
| NC score | 0.006679 (rank : 112) | θ value | 6.88961 (rank : 99) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q02788, Q05505 | Gene names | Col6a2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(VI) chain precursor. | |||||
|
MYH9_HUMAN
|
||||||
| NC score | 0.006591 (rank : 113) | θ value | 6.88961 (rank : 106) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P35579, O60805 | Gene names | MYH9 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
MYH9_MOUSE
|
||||||
| NC score | 0.006335 (rank : 114) | θ value | 6.88961 (rank : 107) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 1829 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8VDD5 | Gene names | Myh9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
SMCA4_HUMAN
|
||||||
| NC score | 0.006014 (rank : 115) | θ value | 4.03905 (rank : 84) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 597 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P51532, O95052, Q9HBD3 | Gene names | SMARCA4, BRG1, SNF2B, SNF2L4 | |||
|
Domain Architecture |

|
|||||
| Description | Probable global transcription activator SNF2L4 (EC 3.6.1.-) (ATP- dependent helicase SMARCA4) (SNF2-beta) (BRG-1 protein) (Mitotic growth and transcription activator) (Brahma protein homolog 1) (SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A member 4). | |||||
|
PKP4_HUMAN
|
||||||
| NC score | 0.004686 (rank : 116) | θ value | 6.88961 (rank : 109) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99569 | Gene names | PKP4 | |||
|
Domain Architecture |

|
|||||
| Description | Plakophilin-4 (p0071). | |||||
|
ERBB2_MOUSE
|
||||||
| NC score | 0.004448 (rank : 117) | θ value | 1.06291 (rank : 66) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 918 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P70424, Q61525, Q6ZPE0 | Gene names | Erbb2, Kiaa3023, Neu | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Receptor tyrosine-protein kinase erbB-2 precursor (EC 2.7.10.1) (p185erbB2) (C-erbB-2) (NEU proto-oncogene). | |||||
|
ERBB2_HUMAN
|
||||||
| NC score | 0.004015 (rank : 118) | θ value | 3.0926 (rank : 78) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 1073 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P04626, Q14256, Q6LDV1, Q9UMK4 | Gene names | ERBB2, HER2, NEU, NGL | |||
|
Domain Architecture |

|
|||||
| Description | Receptor tyrosine-protein kinase erbB-2 precursor (EC 2.7.10.1) (p185erbB2) (C-erbB-2) (NEU proto-oncogene) (Tyrosine kinase-type cell surface receptor HER2) (MLN 19). | |||||
|
COBA2_MOUSE
|
||||||
| NC score | 0.003806 (rank : 119) | θ value | 5.27518 (rank : 88) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 501 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q64739, Q61432, Q9Z1W0 | Gene names | Col11a2 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-2(XI) chain precursor. | |||||
|
TAOK1_MOUSE
|
||||||
| NC score | 0.003676 (rank : 120) | θ value | 5.27518 (rank : 97) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 1720 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q5F2E8, Q69ZL2, Q8JZX2, Q91VG7 | Gene names | Taok1, Kiaa1361 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase TAO1 (EC 2.7.11.1) (Thousand and one amino acid protein 1). | |||||
|
TAOK1_HUMAN
|
||||||
| NC score | 0.003656 (rank : 121) | θ value | 5.27518 (rank : 96) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 1734 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q7L7X3, Q96L75, Q9H2K7, Q9H7S5, Q9P2I6 | Gene names | TAOK1, KIAA1361, MAP3K16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase TAO1 (EC 2.7.11.1) (Thousand and one amino acid protein 1) (STE20-like kinase PSK2) (Kinase from chicken homolog B) (hKFC-B). | |||||
|
CD2L6_MOUSE
|
||||||
| NC score | 0.003573 (rank : 122) | θ value | 1.38821 (rank : 68) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 851 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8BWD8, Q80TM1 | Gene names | Cdc2l6, Kiaa1028 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle 2-like protein kinase 6 (EC 2.7.11.22) (CDC2- related protein kinase 6). | |||||
|
B3A3_MOUSE
|
||||||
| NC score | 0.003194 (rank : 123) | θ value | 8.99809 (rank : 112) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P16283 | Gene names | Slc4a3, Ae3 | |||
|
Domain Architecture |

|
|||||
| Description | Anion exchange protein 3 (Neuronal band 3-like protein) (Solute carrier family 4 member 3). | |||||
|
STK36_HUMAN
|
||||||
| NC score | 0.002986 (rank : 124) | θ value | 4.03905 (rank : 85) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9NRP7, Q8TC32, Q9H9N9, Q9UF35, Q9ULE2 | Gene names | STK36, KIAA1278 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase 36 (EC 2.7.11.1) (Fused homolog). | |||||
|
STK36_MOUSE
|
||||||
| NC score | 0.002758 (rank : 125) | θ value | 8.99809 (rank : 117) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 877 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q69ZM6, Q6PDI0, Q80XQ6 | Gene names | Stk36, Kiaa1278 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase 36 (EC 2.7.11.1) (Fused homolog). | |||||
|
WDR33_HUMAN
|
||||||
| NC score | 0.002749 (rank : 126) | θ value | 5.27518 (rank : 98) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 638 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9C0J8 | Gene names | WDR33, WDC146 | |||
|
Domain Architecture |

|
|||||
| Description | WD repeat protein 33 (WD repeat protein WDC146). | |||||
|
MSX1_HUMAN
|
||||||
| NC score | 0.001176 (rank : 127) | θ value | 5.27518 (rank : 94) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 374 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P28360, Q96NY4 | Gene names | MSX1, HOX7 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein MSX-1 (Msh homeobox 1-like protein) (Hox-7). | |||||
|
GPR84_MOUSE
|
||||||
| NC score | -0.000329 (rank : 128) | θ value | 6.88961 (rank : 100) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 703 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CIM5, Q99MX9 | Gene names | Gpr84 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 84. | |||||
|
ZN688_HUMAN
|
||||||
| NC score | -0.000510 (rank : 129) | θ value | 0.21417 (rank : 49) | |||
| Query Neighborhood Hits | 117 | Target Neighborhood Hits | 873 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8WV14, O75701, Q8IW91, Q96MN0 | Gene names | ZNF688 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 688. | |||||