Please be patient as the page loads
|
ASTL_MOUSE
|
||||||
| SwissProt Accessions | Q6HA09, Q8BMA1 | Gene names | Astl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Astacin-like metalloendopeptidase precursor (EC 3.4.-.-) (Oocyte astacin) (Ovastacin). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
ASTL_MOUSE
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6HA09, Q8BMA1 | Gene names | Astl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Astacin-like metalloendopeptidase precursor (EC 3.4.-.-) (Oocyte astacin) (Ovastacin). | |||||
|
ASTL_HUMAN
|
||||||
| θ value | 1.7349e-152 (rank : 2) | NC score | 0.937536 (rank : 2) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 252 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q6HA08 | Gene names | ASTL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Astacin-like metalloendopeptidase precursor (EC 3.4.-.-) (Oocyte astacin) (Ovastacin). | |||||
|
MEP1B_HUMAN
|
||||||
| θ value | 1.32909e-35 (rank : 3) | NC score | 0.793493 (rank : 3) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q16820, Q670J1 | Gene names | MEP1B | |||
|
Domain Architecture |

|
|||||
| Description | Meprin A subunit beta precursor (EC 3.4.24.18) (Endopeptidase-2) (N- benzoyl-L-tyrosyl-P-amino-benzoic acid hydrolase subunit beta) (PABA peptide hydrolase) (PPH beta). | |||||
|
MEP1B_MOUSE
|
||||||
| θ value | 3.61944e-33 (rank : 4) | NC score | 0.783784 (rank : 4) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q61847 | Gene names | Mep1b, Mep-1b | |||
|
Domain Architecture |

|
|||||
| Description | Meprin A subunit beta precursor (EC 3.4.24.18) (Endopeptidase-2). | |||||
|
MEP1A_HUMAN
|
||||||
| θ value | 1.57365e-28 (rank : 5) | NC score | 0.749189 (rank : 5) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q16819 | Gene names | MEP1A | |||
|
Domain Architecture |

|
|||||
| Description | Meprin A subunit alpha precursor (EC 3.4.24.18) (Endopeptidase-2) (N- benzoyl-L-tyrosyl-P-amino-benzoic acid hydrolase subunit alpha) (PABA peptide hydrolase) (PPH alpha). | |||||
|
BMP1_MOUSE
|
||||||
| θ value | 2.27234e-27 (rank : 6) | NC score | 0.499101 (rank : 12) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P98063 | Gene names | Bmp1 | |||
|
Domain Architecture |

|
|||||
| Description | Bone morphogenetic protein 1 precursor (EC 3.4.24.19) (BMP-1) (Procollagen C-proteinase) (PCP) (Mammalian tolloid protein) (mTld). | |||||
|
MEP1A_MOUSE
|
||||||
| θ value | 2.27234e-27 (rank : 7) | NC score | 0.741882 (rank : 6) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P28825 | Gene names | Mep1a | |||
|
Domain Architecture |

|
|||||
| Description | Meprin A subunit alpha precursor (EC 3.4.24.18) (Endopeptidase-2) (MEP-1). | |||||
|
BMP1_HUMAN
|
||||||
| θ value | 3.87602e-27 (rank : 8) | NC score | 0.501405 (rank : 8) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P13497, Q13292, Q13872, Q14874, Q99421, Q99422, Q99423, Q9UL38 | Gene names | BMP1, PCOLC | |||
|
Domain Architecture |

|
|||||
| Description | Bone morphogenetic protein 1 precursor (EC 3.4.24.19) (BMP-1) (Procollagen C-proteinase) (PCP) (Mammalian tolloid protein) (mTld). | |||||
|
TLL2_MOUSE
|
||||||
| θ value | 5.06226e-27 (rank : 9) | NC score | 0.500482 (rank : 10) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9WVM6 | Gene names | Tll2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tolloid-like protein 2 precursor (EC 3.4.24.-). | |||||
|
TLL2_HUMAN
|
||||||
| θ value | 6.61148e-27 (rank : 10) | NC score | 0.502218 (rank : 7) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Y6L7, Q6PJN5, Q9UQ00 | Gene names | TLL2, KIAA0932 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tolloid-like protein 2 precursor (EC 3.4.24.-). | |||||
|
TLL1_HUMAN
|
||||||
| θ value | 2.51237e-26 (rank : 11) | NC score | 0.500945 (rank : 9) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O43897, Q96AN3, Q9NQS4 | Gene names | TLL1, TLL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tolloid-like protein 1 precursor (EC 3.4.24.-). | |||||
|
TLL1_MOUSE
|
||||||
| θ value | 5.59698e-26 (rank : 12) | NC score | 0.499143 (rank : 11) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 237 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q62381, Q3UTT9, Q8BNP5 | Gene names | Tll1, Tll | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tolloid-like protein 1 precursor (EC 3.4.24.-) (mTll). | |||||
|
PDCD7_HUMAN
|
||||||
| θ value | 1.06291 (rank : 13) | NC score | 0.040785 (rank : 106) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8N8D1, Q96AK8, Q9Y6D7 | Gene names | PDCD7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Programmed cell death protein 7 (ES18) (HES18). | |||||
|
NDE1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 14) | NC score | 0.010041 (rank : 110) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 823 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NXR1, Q49AQ2 | Gene names | NDE1, NUDE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear distribution protein nudE homolog 1 (NudE). | |||||
|
PDCD7_MOUSE
|
||||||
| θ value | 2.36792 (rank : 15) | NC score | 0.037937 (rank : 107) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9WTY1, Q8R5D9 | Gene names | Pdcd7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Programmed cell death protein 7 (ES18). | |||||
|
CS024_HUMAN
|
||||||
| θ value | 4.03905 (rank : 16) | NC score | 0.036461 (rank : 108) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9BVV8, Q9NWS2 | Gene names | C19orf24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C19orf24. | |||||
|
SMG7_HUMAN
|
||||||
| θ value | 4.03905 (rank : 17) | NC score | 0.016368 (rank : 109) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q92540, Q5T1Q0, Q6PIE0, Q7Z7H9, Q8IXC1, Q8IXC2 | Gene names | SMG7, C1orf16, EST1C, KIAA0250 | |||
|
Domain Architecture |

|
|||||
| Description | Protein SMG7 (SMG-7 homolog) (EST1-like protein C). | |||||
|
MLL2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 18) | NC score | 0.009227 (rank : 111) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
MAMC1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 19) | NC score | 0.054071 (rank : 96) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q7Z553 | Gene names | MAMDC1, MDGA2 | |||
|
Domain Architecture |

|
|||||
| Description | MAM domain-containing protein 1 precursor (MAM domain-containing glycosylphosphatidylinositol anchor protein 2). | |||||
|
MAMC1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 20) | NC score | 0.054712 (rank : 93) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P60755 | Gene names | Mamdc1, Mdga2 | |||
|
Domain Architecture |

|
|||||
| Description | MAM domain-containing protein 1 precursor (MAM domain-containing glycosylphosphatidylinositol anchor protein 2). | |||||
|
ATRN_HUMAN
|
||||||
| θ value | θ > 10 (rank : 21) | NC score | 0.109731 (rank : 47) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 525 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O75882, O60295, O95414, Q3MIT3, Q5TDA2, Q5TDA4, Q5VYW3, Q9NTQ3, Q9NTQ4, Q9NU01, Q9NZ57, Q9NZ58 | Gene names | ATRN, KIAA0548, MGCA | |||
|
Domain Architecture |

|
|||||
| Description | Attractin precursor (Mahogany homolog) (DPPT-L). | |||||
|
ATRN_MOUSE
|
||||||
| θ value | θ > 10 (rank : 22) | NC score | 0.107241 (rank : 51) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 519 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9WU60, Q9R263, Q9WU77 | Gene names | Atrn, mg, Mgca | |||
|
Domain Architecture |

|
|||||
| Description | Attractin precursor (Mahogany protein). | |||||
|
C1QR1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 23) | NC score | 0.062240 (rank : 80) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O89103, Q3U2X0 | Gene names | Cd93, Aa4, C1qr1, C1qrp, Ly68 | |||
|
Domain Architecture |

|
|||||
| Description | Complement component C1q receptor precursor (Complement component 1 q subcomponent receptor 1) (C1qRp) (C1qR(p)) (C1q/MBL/SPA receptor) (CD93 antigen) (Cell surface antigen AA4) (Lymphocyte antigen 68). | |||||
|
C1RA_MOUSE
|
||||||
| θ value | θ > 10 (rank : 24) | NC score | 0.104650 (rank : 54) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 379 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CG16, Q99KI6, Q9ET60 | Gene names | C1ra, C1r | |||
|
Domain Architecture |

|
|||||
| Description | Complement C1r-A subcomponent precursor (EC 3.4.21.41) (Complement component 1, r-A subcomponent) [Contains: Complement C1r-A subcomponent heavy chain; Complement C1r-A subcomponent light chain]. | |||||
|
C1RB_MOUSE
|
||||||
| θ value | θ > 10 (rank : 25) | NC score | 0.103346 (rank : 58) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CFG9 | Gene names | C1rb, C1r | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Complement C1r-B subcomponent precursor (EC 3.4.21.41) (Complement component 1, r-B subcomponent) [Contains: Complement C1r-B subcomponent heavy chain; Complement C1r-B subcomponent light chain]. | |||||
|
C1R_HUMAN
|
||||||
| θ value | θ > 10 (rank : 26) | NC score | 0.107900 (rank : 48) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P00736, Q8J012 | Gene names | C1R | |||
|
Domain Architecture |

|
|||||
| Description | Complement C1r subcomponent precursor (EC 3.4.21.41) (Complement component 1, r subcomponent) [Contains: Complement C1r subcomponent heavy chain; Complement C1r subcomponent light chain]. | |||||
|
C1S_HUMAN
|
||||||
| θ value | θ > 10 (rank : 27) | NC score | 0.102299 (rank : 61) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P09871, Q9UCU7, Q9UCU8, Q9UCU9, Q9UCV0, Q9UCV1, Q9UCV2, Q9UCV3, Q9UCV4, Q9UCV5, Q9UM14 | Gene names | C1S | |||
|
Domain Architecture |

|
|||||
| Description | Complement C1s subcomponent precursor (EC 3.4.21.42) (C1 esterase) [Contains: Complement C1s subcomponent heavy chain; Complement C1s subcomponent light chain]. | |||||
|
CD248_HUMAN
|
||||||
| θ value | θ > 10 (rank : 28) | NC score | 0.051572 (rank : 99) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 679 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9HCU0, Q3SX55, Q96KB6 | Gene names | CD248, CD164L1, TEM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endosialin precursor (Tumor endothelial marker 1) (CD248 antigen). | |||||
|
CD248_MOUSE
|
||||||
| θ value | θ > 10 (rank : 29) | NC score | 0.055656 (rank : 90) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 578 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q91V98, Q3UMV6, Q91ZV1 | Gene names | Cd248, Tem1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endosialin precursor (Tumor endothelial marker 1) (CD248 antigen). | |||||
|
CLC14_HUMAN
|
||||||
| θ value | θ > 10 (rank : 30) | NC score | 0.057848 (rank : 86) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q86T13, Q6PWT6, Q8N5V5 | Gene names | CLEC14A, C14orf27, EGFR5 | |||
|
Domain Architecture |
|
|||||
| Description | C-type lectin domain family 14 member A precursor (Epidermal growth factor receptor 5) (EGFR-5). | |||||
|
CLC14_MOUSE
|
||||||
| θ value | θ > 10 (rank : 31) | NC score | 0.056078 (rank : 89) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8VCP9, Q3TP72, Q9CXA8, Q9D624, Q9DC55 | Gene names | Clec14a | |||
|
Domain Architecture |
|
|||||
| Description | C-type lectin domain family 14 member A precursor. | |||||
|
CS1A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 32) | NC score | 0.103560 (rank : 56) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CG14, Q8BJC4, Q8CH28, Q8VBY4 | Gene names | C1sa, C1s | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Complement C1s-A subcomponent precursor (EC 3.4.21.42) (C1 esterase) [Contains: Complement C1s-A subcomponent heavy chain; Complement C1s-A subcomponent light chain]. | |||||
|
CS1B_MOUSE
|
||||||
| θ value | θ > 10 (rank : 33) | NC score | 0.103275 (rank : 59) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 361 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CFG8 | Gene names | C1sb, C1s | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Complement C1s-B subcomponent precursor (EC 3.4.21.42) (C1 esterase) [Contains: Complement C1s-B subcomponent heavy chain; Complement C1s-B subcomponent light chain]. | |||||
|
CSMD1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 34) | NC score | 0.153883 (rank : 36) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96PZ7, Q96QU9, Q96RM4 | Gene names | CSMD1, KIAA1890 | |||
|
Domain Architecture |

|
|||||
| Description | CUB and sushi domain-containing protein 1 precursor (CUB and sushi multiple domains protein 1). | |||||
|
CSMD1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 35) | NC score | 0.152285 (rank : 39) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q923L3, Q8BUV1, Q8BYQ3 | Gene names | Csmd1 | |||
|
Domain Architecture |

|
|||||
| Description | CUB and sushi domain-containing protein 1 precursor (CUB and sushi multiple domains protein 1). | |||||
|
CSMD2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 36) | NC score | 0.157647 (rank : 35) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q7Z408, Q5VT59, Q8N963, Q96Q03, Q9H4V7, Q9H4V8, Q9H4V9, Q9H4W0, Q9H4W1, Q9H4W2, Q9H4W3, Q9H4W4, Q9HCY5, Q9HCY6, Q9HCY7 | Gene names | CSMD2, KIAA1884 | |||
|
Domain Architecture |

|
|||||
| Description | CUB and sushi domain-containing protein 2 (CUB and sushi multiple domains protein 2). | |||||
|
CSMD3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 37) | NC score | 0.153140 (rank : 38) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q7Z407, Q96PZ3 | Gene names | CSMD3, KIAA1894 | |||
|
Domain Architecture |

|
|||||
| Description | CUB and sushi domain-containing protein 3 precursor (CUB and sushi multiple domains protein 3). | |||||
|
CUBN_HUMAN
|
||||||
| θ value | θ > 10 (rank : 38) | NC score | 0.152230 (rank : 40) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 482 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O60494, Q5VTA6, Q96RU9 | Gene names | CUBN, IFCR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cubilin precursor (Intrinsic factor-cobalamin receptor) (Intrinsic factor-vitamin B12 receptor) (460 kDa receptor) (Intestinal intrinsic factor receptor). | |||||
|
CUBN_MOUSE
|
||||||
| θ value | θ > 10 (rank : 39) | NC score | 0.153460 (rank : 37) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 477 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9JLB4 | Gene names | Cubn, Ifcr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cubilin precursor (Intrinsic factor-cobalamin receptor). | |||||
|
CUZD1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 40) | NC score | 0.166402 (rank : 30) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q86UP6, Q7Z660, Q7Z661, Q86SG1, Q86UP5, Q9HAR7 | Gene names | CUZD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CUB and zona pellucida-like domain-containing protein 1 precursor (Transmembrane protein UO-44). | |||||
|
CUZD1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 41) | NC score | 0.160372 (rank : 34) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P70412, Q9CTZ7 | Gene names | Cuzd1, Itmap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CUB and zona pellucida-like domain-containing protein 1 precursor (Integral membrane-associated protein 1). | |||||
|
DCBD1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 42) | NC score | 0.103456 (rank : 57) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8N8Z6, Q5H992, Q8IYK5, Q8N7L9, Q96NH2 | Gene names | DCBLD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Discoidin, CUB and LCCL domain-containing protein 1 precursor. | |||||
|
DCBD1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 43) | NC score | 0.122622 (rank : 44) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9D4J3, Q8R327, Q9D696 | Gene names | Dcbld1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Discoidin, CUB and LCCL domain-containing protein 1 precursor. | |||||
|
DCBD2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 44) | NC score | 0.118628 (rank : 45) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96PD2, Q8N6M4, Q8TDX2 | Gene names | DCBLD2, CLCP1, ESDN | |||
|
Domain Architecture |
|
|||||
| Description | Discoidin, CUB and LCCL domain-containing protein 2 precursor (Endothelial and smooth muscle cell-derived neuropilin-like protein) (CUB, LCCL and coagulation factor V/VIII-homology domains protein 1). | |||||
|
DCBD2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 45) | NC score | 0.114086 (rank : 46) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q91ZV3, Q8BKI4 | Gene names | Dcbld2, Esdn | |||
|
Domain Architecture |
|
|||||
| Description | Discoidin, CUB and LCCL domain-containing protein 2 precursor (Endothelial and smooth muscle cell-derived neuropilin-like protein). | |||||
|
DMBT1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 46) | NC score | 0.138845 (rank : 43) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UGM3, Q59EX0, Q5JR26, Q6MZN4, Q96DU4, Q9UGM2, Q9UJ57, Q9UKJ4, Q9Y211, Q9Y4V9 | Gene names | DMBT1, GP340 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Deleted in malignant brain tumors 1 protein precursor (Glycoprotein 340) (Gp-340) (Surfactant pulmonary-associated D-binding protein). | |||||
|
DMBT1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 47) | NC score | 0.171091 (rank : 29) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q60997, Q80YC6, Q9JMJ9 | Gene names | Dmbt1, Crpd | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Deleted in malignant brain tumors 1 protein precursor (CRP-ductin) (Vomeroglandin). | |||||
|
EGFL7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 48) | NC score | 0.081007 (rank : 68) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UHF1, Q5M7Y5, Q5VUD5, Q96EG0 | Gene names | EGFL7, MEGF7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EGF-like domain-containing protein 7 precursor (Multiple EGF-like domain protein 7) (Multiple epidermal growth factor-like domain protein 7) (Vascular endothelial statin) (VE-statin) (NOTCH4-like protein) (ZNEU1). | |||||
|
EGFL7_MOUSE
|
||||||
| θ value | θ > 10 (rank : 49) | NC score | 0.051087 (rank : 102) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 287 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9QXT5, Q6XD35, Q9DCP5 | Gene names | Egfl7, Megf7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EGF-like domain-containing protein 7 precursor (Multiple EGF-like domain protein 7) (Multiple epidermal growth factor-like domain protein 7) (Vascular endothelial statin) (VE-statin) (NOTCH4-like protein) (Zneu1). | |||||
|
EGFL8_MOUSE
|
||||||
| θ value | θ > 10 (rank : 50) | NC score | 0.051039 (rank : 103) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q6GUQ1, O35447 | Gene names | Egfl8, Ng3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EGF-like domain-containing protein 8 precursor (Epidermal growth factor-like protein 8) (Multiple EGF-like domain protein 8). | |||||
|
EGF_MOUSE
|
||||||
| θ value | θ > 10 (rank : 51) | NC score | 0.051340 (rank : 101) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 358 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P01132, Q6P9J2 | Gene names | Egf | |||
|
Domain Architecture |

|
|||||
| Description | Pro-epidermal growth factor precursor (EGF) [Contains: Epidermal growth factor]. | |||||
|
ENTK_HUMAN
|
||||||
| θ value | θ > 10 (rank : 52) | NC score | 0.072353 (rank : 70) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P98073 | Gene names | PRSS7, ENTK | |||
|
Domain Architecture |

|
|||||
| Description | Enteropeptidase precursor (EC 3.4.21.9) (Enterokinase) [Contains: Enteropeptidase non-catalytic heavy chain; Enteropeptidase catalytic light chain]. | |||||
|
ENTK_MOUSE
|
||||||
| θ value | θ > 10 (rank : 53) | NC score | 0.079111 (rank : 69) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 338 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P97435 | Gene names | Prss7, Entk | |||
|
Domain Architecture |

|
|||||
| Description | Enteropeptidase (EC 3.4.21.9) (Enterokinase) [Contains: Enteropeptidase non-catalytic heavy chain; Enteropeptidase catalytic light chain]. | |||||
|
FBLN5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 54) | NC score | 0.056993 (rank : 87) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UBX5, O75966, Q6UWA3 | Gene names | FBLN5, DANCE | |||
|
Domain Architecture |

|
|||||
| Description | Fibulin-5 precursor (FIBL-5) (Developmental arteries and neural crest EGF-like protein) (Dance) (Urine p50 protein) (UP50). | |||||
|
FBLN5_MOUSE
|
||||||
| θ value | θ > 10 (rank : 55) | NC score | 0.054716 (rank : 92) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 317 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9WVH9, Q541Z7 | Gene names | Fbln5, Dance | |||
|
Domain Architecture |
|
|||||
| Description | Fibulin-5 precursor (FIBL-5) (Developmental arteries and neural crest EGF-like protein) (Dance). | |||||
|
FBN1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 56) | NC score | 0.052923 (rank : 98) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 557 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P35555, Q15972, Q75N87 | Gene names | FBN1, FBN | |||
|
Domain Architecture |

|
|||||
| Description | Fibrillin-1 precursor. | |||||
|
FBN1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 57) | NC score | 0.054886 (rank : 91) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 563 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q61554, Q60826 | Gene names | Fbn1, Fbn-1 | |||
|
Domain Architecture |

|
|||||
| Description | Fibrillin-1 precursor. | |||||
|
FBN3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 58) | NC score | 0.051500 (rank : 100) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q75N90, Q75N91, Q75N92, Q75N93, Q86SJ5, Q96JP8 | Gene names | FBN3, KIAA1776 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fibrillin-3 precursor. | |||||
|
GP126_HUMAN
|
||||||
| θ value | θ > 10 (rank : 59) | NC score | 0.104706 (rank : 53) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q86SQ4, Q6MZU7, Q8NC14, Q96JW0 | Gene names | GPR126 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 126 precursor. | |||||
|
KREM1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 60) | NC score | 0.094131 (rank : 65) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96MU8, Q5TIB9, Q6P3X6, Q9BY70, Q9UGS5, Q9UGU1 | Gene names | KREMEN1, KREMEN, KRM1 | |||
|
Domain Architecture |
|
|||||
| Description | Kremen protein 1 precursor (Kringle-containing protein marking the eye and the nose) (Kringle domain-containing transmembrane protein 1) (Dickkopf receptor). | |||||
|
KREM1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 61) | NC score | 0.082391 (rank : 67) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q99N43 | Gene names | Kremen1, Kremen | |||
|
Domain Architecture |
|
|||||
| Description | Kremen protein 1 precursor (Kringle-containing protein marking the eye and the nose) (Kringle domain-containing transmembrane protein 1) (Dickkopf receptor). | |||||
|
KREM2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 62) | NC score | 0.107555 (rank : 49) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8NCW0, Q8N2J4, Q8NCW1, Q96GL8, Q9BTP9 | Gene names | KREMEN2, KRM2 | |||
|
Domain Architecture |
|
|||||
| Description | Kremen protein 2 precursor (Kringle-containing protein marking the eye and the nose) (Kringle domain-containing transmembrane protein 2) (Dickkopf receptor 2). | |||||
|
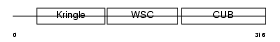
KREM2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 63) | NC score | 0.100527 (rank : 63) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8K1S7 | Gene names | Kremen2 | |||
|
Domain Architecture |
|
|||||
| Description | Kremen protein 2 precursor (Kringle-containing protein marking the eye and the nose) (Kringle domain-containing transmembrane protein 2) (Dickkopf receptor 2). | |||||
|
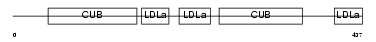
LRP12_HUMAN
|
||||||
| θ value | θ > 10 (rank : 64) | NC score | 0.146999 (rank : 42) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Y561 | Gene names | LRP12, ST7 | |||
|
Domain Architecture |
|
|||||
| Description | Low-density lipoprotein receptor-related protein 12 precursor (Suppressor of tumorigenicity protein 7). | |||||
|
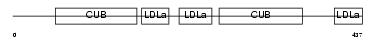
LRP12_MOUSE
|
||||||
| θ value | θ > 10 (rank : 65) | NC score | 0.147571 (rank : 41) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BUJ9, Q8BWM9 | Gene names | Lrp12 | |||
|
Domain Architecture |
|
|||||
| Description | Low-density lipoprotein receptor-related protein 12 precursor. | |||||
|
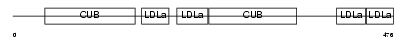
LRP3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 66) | NC score | 0.107407 (rank : 50) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O75074 | Gene names | LRP3 | |||
|
Domain Architecture |
|
|||||
| Description | Low-density lipoprotein receptor-related protein 3 precursor (hLRp105). | |||||
|
LTBP2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 67) | NC score | 0.053005 (rank : 97) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 617 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O08999, Q8C6W9 | Gene names | Ltbp2 | |||
|
Domain Architecture |

|
|||||
| Description | Latent-transforming growth factor beta-binding protein 2 precursor (LTBP-2). | |||||
|
LTBP3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 68) | NC score | 0.062661 (rank : 78) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 475 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q61810, Q8BNQ6 | Gene names | Ltbp3 | |||
|
Domain Architecture |

|
|||||
| Description | Latent-transforming growth factor beta-binding protein 3 precursor (LTBP-3). | |||||
|
MAMC2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 69) | NC score | 0.089846 (rank : 66) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q7Z304, Q5VW47, Q8WX43, Q96BM4 | Gene names | MAMDC2 | |||
|
Domain Architecture |

|
|||||
| Description | MAM domain-containing protein 2 precursor. | |||||
|
MAMC2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 70) | NC score | 0.098205 (rank : 64) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CG85, Q8BM24 | Gene names | Mamdc2 | |||
|
Domain Architecture |

|
|||||
| Description | MAM domain-containing protein 2 precursor. | |||||
|
MASP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 71) | NC score | 0.105645 (rank : 52) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 369 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P48740, O95570, Q9UF09 | Gene names | MASP1, CRARF, CRARF1, PRSS5 | |||
|
Domain Architecture |

|
|||||
| Description | Complement-activating component of Ra-reactive factor precursor (EC 3.4.21.-) (Ra-reactive factor serine protease p100) (RaRF) (Mannan-binding lectin serine protease 1) (Mannose-binding protein- associated serine protease) (MASP-1) [Contains: Complement-activating component of Ra-reactive factor heavy chain; Complement-activating component of Ra-reactive factor light chain]. | |||||
|
MASP1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 72) | NC score | 0.104055 (rank : 55) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P98064 | Gene names | Masp1, Crarf | |||
|
Domain Architecture |

|
|||||
| Description | Complement-activating component of Ra-reactive factor precursor (EC 3.4.21.-) (Ra-reactive factor serine protease p100) (RaRF) (Mannan-binding lectin serine protease 1) [Contains: Complement- activating component of Ra-reactive factor heavy chain; Complement- activating component of Ra-reactive factor light chain]. | |||||
|
MASP2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 73) | NC score | 0.102458 (rank : 60) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O00187, O75754, Q5TEQ5, Q5TER0, Q96QG4, Q9BZH0, Q9H498, Q9H499, Q9UBP3, Q9ULC7, Q9UMV3, Q9Y270 | Gene names | MASP2 | |||
|
Domain Architecture |

|
|||||
| Description | Mannan-binding lectin serine protease 2 precursor (EC 3.4.21.104) (Mannose-binding protein-associated serine protease 2) (MASP-2) (MBL- associated serine protease 2) [Contains: Mannan-binding lectin serine protease 2 A chain; Mannan-binding lectin serine protease 2 B chain]. | |||||
|
MASP2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 74) | NC score | 0.101205 (rank : 62) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 382 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q91WP0, Q9QXA4, Q9QXD2, Q9QXD5, Q9Z338 | Gene names | Masp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mannan-binding lectin serine protease 2 precursor (EC 3.4.21.104) (Mannose-binding protein-associated serine protease 2) (MASP-2) (MBL- associated serine protease 2) [Contains: Mannan-binding lectin serine protease 2 A chain; Mannan-binding lectin serine protease 2 B chain]. | |||||
|
MATN1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 75) | NC score | 0.056317 (rank : 88) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P21941 | Gene names | MATN1, CMP, CRTM | |||
|
Domain Architecture |
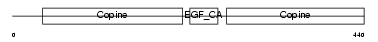
|
|||||
| Description | Cartilage matrix protein precursor (Matrilin-1). | |||||
|
MATN1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 76) | NC score | 0.057912 (rank : 85) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 186 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P51942 | Gene names | Matn1, Cmp, Crtm | |||
|
Domain Architecture |
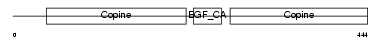
|
|||||
| Description | Cartilage matrix protein precursor (Matrilin-1). | |||||
|
MATN2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 77) | NC score | 0.060833 (rank : 82) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 365 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O00339, Q6UWA5, Q7Z5X1, Q8NDE6, Q96FT5, Q9NSZ1 | Gene names | MATN2 | |||
|
Domain Architecture |

|
|||||
| Description | Matrilin-2 precursor. | |||||
|
MATN2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 78) | NC score | 0.063549 (rank : 75) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 356 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O08746 | Gene names | Matn2 | |||
|
Domain Architecture |

|
|||||
| Description | Matrilin-2 precursor. | |||||
|
MATN4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 79) | NC score | 0.054705 (rank : 94) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 287 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O95460, Q5QPU2, Q5QPU3, Q5QPU4, Q8N2M5, Q8N2M7, Q9H1F8, Q9H1F9 | Gene names | MATN4 | |||
|
Domain Architecture |

|
|||||
| Description | Matrilin-4 precursor. | |||||
|
MDGA1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 80) | NC score | 0.063156 (rank : 77) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 358 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8NFP4, Q8NBE3 | Gene names | MDGA1, MAMDC3 | |||
|
Domain Architecture |

|
|||||
| Description | MAM domain-containing glycosylphosphatidylinositol anchor protein 1 precursor (MAM domain-containing protein 3) (GPI and MAM protein) (Glycosylphosphatidylinositol-MAM) (GPIM). | |||||
|
MEGF6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 81) | NC score | 0.050596 (rank : 104) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O75095, Q5VV39 | Gene names | MEGF6, EGFL3, KIAA0815 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Multiple epidermal growth factor-like domains 6 precursor (EGF-like domain-containing protein 3) (Multiple EGF-like domain protein 3). | |||||
|
MFRP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 82) | NC score | 0.184460 (rank : 25) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9BY79, Q335M3, Q96DQ9 | Gene names | MFRP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane frizzled-related protein (Membrane-type frizzled-related protein). | |||||
|
MFRP_MOUSE
|
||||||
| θ value | θ > 10 (rank : 83) | NC score | 0.180881 (rank : 27) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8K480, Q8BPP4 | Gene names | Mfrp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane frizzled-related protein (Membrane-type frizzled-related protein). | |||||
|
MUC13_MOUSE
|
||||||
| θ value | θ > 10 (rank : 84) | NC score | 0.054486 (rank : 95) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P19467 | Gene names | Muc13, Ly64 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-13 precursor (Cell surface antigen 114/A10) (Lymphocyte antigen 64). | |||||
|
NETO1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 85) | NC score | 0.189514 (rank : 22) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8TDF5, Q86W85, Q8ND78, Q8TDF4 | Gene names | NETO1, BTCL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuropilin and tolloid-like protein 1 precursor (Brain-specific transmembrane protein containing 2 CUB and 1 LDL-receptor class A domains protein 1). | |||||
|
NETO1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 86) | NC score | 0.188540 (rank : 23) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8R4I7, Q80X39, Q8C4S3, Q8CCM2 | Gene names | Neto1, Btcl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuropilin and tolloid-like protein 1 precursor (Brain-specific transmembrane protein containing 2 CUB and 1 LDL-receptor class A domains protein 1). | |||||
|
NETO2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 87) | NC score | 0.190381 (rank : 21) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8NC67, Q7Z381, Q8ND51, Q96SP4, Q9NVY8 | Gene names | NETO2, BTCL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuropilin and tolloid-like protein 2 precursor (Brain-specific transmembrane protein containing 2 CUB and 1 LDL-receptor class A domains protein 2). | |||||
|
NETO2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 88) | NC score | 0.191053 (rank : 20) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BNJ6, Q5VM49, Q8C4Q8 | Gene names | Neto2, Btcl2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuropilin and tolloid-like protein 2 precursor (Brain-specific transmembrane protein containing 2 CUB and 1 LDL-receptor class A domains protein 2). | |||||
|
NRP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 89) | NC score | 0.195616 (rank : 19) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O14786, O60461, Q96IH5 | Gene names | NRP1, NRP, VEGF165R | |||
|
Domain Architecture |

|
|||||
| Description | Neuropilin-1 precursor (Vascular endothelial cell growth factor 165 receptor) (CD304 antigen). | |||||
|
NRP1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 90) | NC score | 0.197351 (rank : 18) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P97333 | Gene names | Nrp1, Nrp | |||
|
Domain Architecture |

|
|||||
| Description | Neuropilin-1 precursor (A5 protein). | |||||
|
NRP2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 91) | NC score | 0.181185 (rank : 26) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O60462, O14820, O14821 | Gene names | NRP2, VEGF165R2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuropilin-2 precursor (Vascular endothelial cell growth factor 165 receptor 2). | |||||
|
NRP2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 92) | NC score | 0.171260 (rank : 28) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O35375, O35373, O35374, O35376, O35377, O35378 | Gene names | Nrp2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuropilin-2 precursor (Vascular endothelial cell growth factor 165 receptor 2). | |||||
|
PCOC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 93) | NC score | 0.214694 (rank : 14) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q15113, O14550 | Gene names | PCOLCE, PCPE1 | |||
|
Domain Architecture |
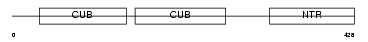
|
|||||
| Description | Procollagen C-endopeptidase enhancer 1 precursor (Procollagen COOH- terminal proteinase enhancer 1) (Procollagen C-proteinase enhancer 1) (PCPE-1) (Type I procollagen COOH-terminal proteinase enhancer) (Type 1 procollagen C-proteinase enhancer protein). | |||||
|
PCOC1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 94) | NC score | 0.215129 (rank : 13) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q61398, O35113 | Gene names | Pcolce, Pcpe1 | |||
|
Domain Architecture |
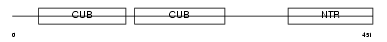
|
|||||
| Description | Procollagen C-endopeptidase enhancer 1 precursor (Procollagen COOH- terminal proteinase enhancer 1) (Procollagen C-proteinase enhancer 1) (PCPE-1) (Type I procollagen COOH-terminal proteinase enhancer) (Type 1 procollagen C-proteinase enhancer protein) (P14). | |||||
|
PCOC2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 95) | NC score | 0.212318 (rank : 16) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UKZ9, Q9BRH3 | Gene names | PCOLCE2, PCPE2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Procollagen C-endopeptidase enhancer 2 precursor (Procollagen COOH- terminal proteinase enhancer 2) (Procollagen C-proteinase enhancer 2) (PCPE-2). | |||||
|
PCOC2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 96) | NC score | 0.214148 (rank : 15) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8R4W6, Q3V1K6, Q9CX06 | Gene names | Pcolce2, Pcpe2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Procollagen C-endopeptidase enhancer 2 precursor (Procollagen COOH- terminal proteinase enhancer 2) (Procollagen C-proteinase enhancer 2) (PCPE-2). | |||||
|
PDGFD_HUMAN
|
||||||
| θ value | θ > 10 (rank : 97) | NC score | 0.187242 (rank : 24) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9GZP0, Q9BWV5 | Gene names | PDGFD, IEGF, SCDGFB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Platelet-derived growth factor D precursor (PDGF D) (Iris-expressed growth factor) (Spinal cord-derived growth factor B) (SCDGF-B). | |||||
|
PDGFD_MOUSE
|
||||||
| θ value | θ > 10 (rank : 98) | NC score | 0.197966 (rank : 17) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q925I7, Q3URF6, Q8K2L3, Q9D1L8 | Gene names | Pdgfd, Scdgfb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Platelet-derived growth factor chain precursor (PDGF D) (Spinal cord- derived growth factor B) (SCDGF-B). | |||||
|
PHXR5_MOUSE
|
||||||
| θ value | θ > 10 (rank : 99) | NC score | 0.060629 (rank : 83) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P08399 | Gene names | Phxr5, Per | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Per-hexamer repeat protein 5. | |||||
|
PROS_HUMAN
|
||||||
| θ value | θ > 10 (rank : 100) | NC score | 0.065801 (rank : 73) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 340 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P07225, Q15518, Q7Z715 | Gene names | PROS1, PROS | |||
|
Domain Architecture |

|
|||||
| Description | Vitamin K-dependent protein S precursor. | |||||
|
PROS_MOUSE
|
||||||
| θ value | θ > 10 (rank : 101) | NC score | 0.058650 (rank : 84) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q08761, P43483 | Gene names | Pros1, Pros | |||
|
Domain Architecture |

|
|||||
| Description | Vitamin K-dependent protein S precursor. | |||||
|
SCUB3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 102) | NC score | 0.063203 (rank : 76) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 428 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8IX30, Q5CZB3, Q86UZ9, Q8NAU9 | Gene names | SCUBE3, CEGF3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal peptide, CUB and EGF-like domain-containing protein 3 precursor. | |||||
|
SCUB3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 103) | NC score | 0.064349 (rank : 74) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 442 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q66PY1, Q68FG9 | Gene names | Scube3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal peptide, CUB and EGF-like domain-containing protein 3 precursor. | |||||
|
SEZ6L_HUMAN
|
||||||
| θ value | θ > 10 (rank : 104) | NC score | 0.164190 (rank : 32) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9BYH1, O95917, Q5THY5, Q6IBZ4, Q6UXD4, Q9NUI3, Q9NUI4, Q9NUI5, Q9Y2E1, Q9Y3J6 | Gene names | SEZ6L, KIAA0927 | |||
|
Domain Architecture |

|
|||||
| Description | Seizure 6-like protein precursor. | |||||
|
ST14_HUMAN
|
||||||
| θ value | θ > 10 (rank : 105) | NC score | 0.062477 (rank : 79) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Y5Y6, Q9BS01, Q9H3S0, Q9HB36, Q9HCA3 | Gene names | ST14, PRSS14, SNC19, TADG15 | |||
|
Domain Architecture |

|
|||||
| Description | Suppressor of tumorigenicity protein 14 (EC 3.4.21.-) (Serine protease 14) (Matriptase) (Membrane-type serine protease 1) (MT-SP1) (Prostamin) (Serine protease TADG-15) (Tumor-associated differentially-expressed gene 15 protein). | |||||
|
ST14_MOUSE
|
||||||
| θ value | θ > 10 (rank : 106) | NC score | 0.060905 (rank : 81) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P56677 | Gene names | St14, Prss14 | |||
|
Domain Architecture |

|
|||||
| Description | Suppressor of tumorigenicity protein 14 (EC 3.4.21.-) (Serine protease 14) (Epithin). | |||||
|
TRAF4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 107) | NC score | 0.067009 (rank : 72) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 373 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9BUZ4, O75615, Q14848, Q2PJN8 | Gene names | TRAF4, CART1, MLN62, RNF83 | |||
|
Domain Architecture |
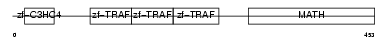
|
|||||
| Description | TNF receptor-associated factor 4 (Cysteine-rich domain associated with RING and Traf domains protein 1) (Malignant 62) (RING finger protein 83). | |||||
|
TRAF4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 108) | NC score | 0.070650 (rank : 71) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 338 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q61382 | Gene names | Traf4, Cart1 | |||
|
Domain Architecture |
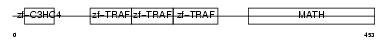
|
|||||
| Description | TNF receptor-associated factor 4 (Cysteine-rich motif associated to RING and Traf domains protein 1). | |||||
|
TSG6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 109) | NC score | 0.166225 (rank : 31) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P98066, Q8WWI9 | Gene names | TNFAIP6, TSG6 | |||
|
Domain Architecture |
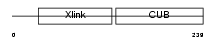
|
|||||
| Description | Tumor necrosis factor-inducible protein TSG-6 precursor (TNF- stimulated gene 6 protein) (Hyaluronate-binding protein). | |||||
|
TSG6_MOUSE
|
||||||
| θ value | θ > 10 (rank : 110) | NC score | 0.164137 (rank : 33) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O08859 | Gene names | Tnfaip6, Tnfip6, Tsg6 | |||
|
Domain Architecture |
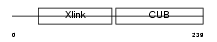
|
|||||
| Description | Tumor necrosis factor-inducible protein TSG-6 precursor (TNF- stimulated gene 6 protein). | |||||
|
UROL1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 111) | NC score | 0.050194 (rank : 105) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q5DID3, Q5DID1, Q5DID2 | Gene names | Umodl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uromodulin-like 1 precursor (Olfactorin). | |||||
|
ASTL_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6HA09, Q8BMA1 | Gene names | Astl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Astacin-like metalloendopeptidase precursor (EC 3.4.-.-) (Oocyte astacin) (Ovastacin). | |||||
|
ASTL_HUMAN
|
||||||
| NC score | 0.937536 (rank : 2) | θ value | 1.7349e-152 (rank : 2) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 252 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q6HA08 | Gene names | ASTL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Astacin-like metalloendopeptidase precursor (EC 3.4.-.-) (Oocyte astacin) (Ovastacin). | |||||
|
MEP1B_HUMAN
|
||||||
| NC score | 0.793493 (rank : 3) | θ value | 1.32909e-35 (rank : 3) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q16820, Q670J1 | Gene names | MEP1B | |||
|
Domain Architecture |

|
|||||
| Description | Meprin A subunit beta precursor (EC 3.4.24.18) (Endopeptidase-2) (N- benzoyl-L-tyrosyl-P-amino-benzoic acid hydrolase subunit beta) (PABA peptide hydrolase) (PPH beta). | |||||
|
MEP1B_MOUSE
|
||||||
| NC score | 0.783784 (rank : 4) | θ value | 3.61944e-33 (rank : 4) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q61847 | Gene names | Mep1b, Mep-1b | |||
|
Domain Architecture |

|
|||||
| Description | Meprin A subunit beta precursor (EC 3.4.24.18) (Endopeptidase-2). | |||||
|
MEP1A_HUMAN
|
||||||
| NC score | 0.749189 (rank : 5) | θ value | 1.57365e-28 (rank : 5) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q16819 | Gene names | MEP1A | |||
|
Domain Architecture |

|
|||||
| Description | Meprin A subunit alpha precursor (EC 3.4.24.18) (Endopeptidase-2) (N- benzoyl-L-tyrosyl-P-amino-benzoic acid hydrolase subunit alpha) (PABA peptide hydrolase) (PPH alpha). | |||||
|
MEP1A_MOUSE
|
||||||
| NC score | 0.741882 (rank : 6) | θ value | 2.27234e-27 (rank : 7) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P28825 | Gene names | Mep1a | |||
|
Domain Architecture |

|
|||||
| Description | Meprin A subunit alpha precursor (EC 3.4.24.18) (Endopeptidase-2) (MEP-1). | |||||
|
TLL2_HUMAN
|
||||||
| NC score | 0.502218 (rank : 7) | θ value | 6.61148e-27 (rank : 10) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 241 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9Y6L7, Q6PJN5, Q9UQ00 | Gene names | TLL2, KIAA0932 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tolloid-like protein 2 precursor (EC 3.4.24.-). | |||||
|
BMP1_HUMAN
|
||||||
| NC score | 0.501405 (rank : 8) | θ value | 3.87602e-27 (rank : 8) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P13497, Q13292, Q13872, Q14874, Q99421, Q99422, Q99423, Q9UL38 | Gene names | BMP1, PCOLC | |||
|
Domain Architecture |

|
|||||
| Description | Bone morphogenetic protein 1 precursor (EC 3.4.24.19) (BMP-1) (Procollagen C-proteinase) (PCP) (Mammalian tolloid protein) (mTld). | |||||
|
TLL1_HUMAN
|
||||||
| NC score | 0.500945 (rank : 9) | θ value | 2.51237e-26 (rank : 11) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O43897, Q96AN3, Q9NQS4 | Gene names | TLL1, TLL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tolloid-like protein 1 precursor (EC 3.4.24.-). | |||||
|
TLL2_MOUSE
|
||||||
| NC score | 0.500482 (rank : 10) | θ value | 5.06226e-27 (rank : 9) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 240 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9WVM6 | Gene names | Tll2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tolloid-like protein 2 precursor (EC 3.4.24.-). | |||||
|
TLL1_MOUSE
|
||||||
| NC score | 0.499143 (rank : 11) | θ value | 5.59698e-26 (rank : 12) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 237 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q62381, Q3UTT9, Q8BNP5 | Gene names | Tll1, Tll | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tolloid-like protein 1 precursor (EC 3.4.24.-) (mTll). | |||||
|
BMP1_MOUSE
|
||||||
| NC score | 0.499101 (rank : 12) | θ value | 2.27234e-27 (rank : 6) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P98063 | Gene names | Bmp1 | |||
|
Domain Architecture |

|
|||||
| Description | Bone morphogenetic protein 1 precursor (EC 3.4.24.19) (BMP-1) (Procollagen C-proteinase) (PCP) (Mammalian tolloid protein) (mTld). | |||||
|
PCOC1_MOUSE
|
||||||
| NC score | 0.215129 (rank : 13) | θ value | θ > 10 (rank : 94) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q61398, O35113 | Gene names | Pcolce, Pcpe1 | |||
|
Domain Architecture |

|
|||||
| Description | Procollagen C-endopeptidase enhancer 1 precursor (Procollagen COOH- terminal proteinase enhancer 1) (Procollagen C-proteinase enhancer 1) (PCPE-1) (Type I procollagen COOH-terminal proteinase enhancer) (Type 1 procollagen C-proteinase enhancer protein) (P14). | |||||
|
PCOC1_HUMAN
|
||||||
| NC score | 0.214694 (rank : 14) | θ value | θ > 10 (rank : 93) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q15113, O14550 | Gene names | PCOLCE, PCPE1 | |||
|
Domain Architecture |

|
|||||
| Description | Procollagen C-endopeptidase enhancer 1 precursor (Procollagen COOH- terminal proteinase enhancer 1) (Procollagen C-proteinase enhancer 1) (PCPE-1) (Type I procollagen COOH-terminal proteinase enhancer) (Type 1 procollagen C-proteinase enhancer protein). | |||||
|
PCOC2_MOUSE
|
||||||
| NC score | 0.214148 (rank : 15) | θ value | θ > 10 (rank : 96) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8R4W6, Q3V1K6, Q9CX06 | Gene names | Pcolce2, Pcpe2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Procollagen C-endopeptidase enhancer 2 precursor (Procollagen COOH- terminal proteinase enhancer 2) (Procollagen C-proteinase enhancer 2) (PCPE-2). | |||||
|
PCOC2_HUMAN
|
||||||
| NC score | 0.212318 (rank : 16) | θ value | θ > 10 (rank : 95) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UKZ9, Q9BRH3 | Gene names | PCOLCE2, PCPE2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Procollagen C-endopeptidase enhancer 2 precursor (Procollagen COOH- terminal proteinase enhancer 2) (Procollagen C-proteinase enhancer 2) (PCPE-2). | |||||
|
PDGFD_MOUSE
|
||||||
| NC score | 0.197966 (rank : 17) | θ value | θ > 10 (rank : 98) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q925I7, Q3URF6, Q8K2L3, Q9D1L8 | Gene names | Pdgfd, Scdgfb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Platelet-derived growth factor chain precursor (PDGF D) (Spinal cord- derived growth factor B) (SCDGF-B). | |||||
|
NRP1_MOUSE
|
||||||
| NC score | 0.197351 (rank : 18) | θ value | θ > 10 (rank : 90) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P97333 | Gene names | Nrp1, Nrp | |||
|
Domain Architecture |

|
|||||
| Description | Neuropilin-1 precursor (A5 protein). | |||||
|
NRP1_HUMAN
|
||||||
| NC score | 0.195616 (rank : 19) | θ value | θ > 10 (rank : 89) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O14786, O60461, Q96IH5 | Gene names | NRP1, NRP, VEGF165R | |||
|
Domain Architecture |

|
|||||
| Description | Neuropilin-1 precursor (Vascular endothelial cell growth factor 165 receptor) (CD304 antigen). | |||||
|
NETO2_MOUSE
|
||||||
| NC score | 0.191053 (rank : 20) | θ value | θ > 10 (rank : 88) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BNJ6, Q5VM49, Q8C4Q8 | Gene names | Neto2, Btcl2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuropilin and tolloid-like protein 2 precursor (Brain-specific transmembrane protein containing 2 CUB and 1 LDL-receptor class A domains protein 2). | |||||
|
NETO2_HUMAN
|
||||||
| NC score | 0.190381 (rank : 21) | θ value | θ > 10 (rank : 87) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8NC67, Q7Z381, Q8ND51, Q96SP4, Q9NVY8 | Gene names | NETO2, BTCL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuropilin and tolloid-like protein 2 precursor (Brain-specific transmembrane protein containing 2 CUB and 1 LDL-receptor class A domains protein 2). | |||||
|
NETO1_HUMAN
|
||||||
| NC score | 0.189514 (rank : 22) | θ value | θ > 10 (rank : 85) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8TDF5, Q86W85, Q8ND78, Q8TDF4 | Gene names | NETO1, BTCL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuropilin and tolloid-like protein 1 precursor (Brain-specific transmembrane protein containing 2 CUB and 1 LDL-receptor class A domains protein 1). | |||||
|
NETO1_MOUSE
|
||||||
| NC score | 0.188540 (rank : 23) | θ value | θ > 10 (rank : 86) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8R4I7, Q80X39, Q8C4S3, Q8CCM2 | Gene names | Neto1, Btcl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuropilin and tolloid-like protein 1 precursor (Brain-specific transmembrane protein containing 2 CUB and 1 LDL-receptor class A domains protein 1). | |||||
|
PDGFD_HUMAN
|
||||||
| NC score | 0.187242 (rank : 24) | θ value | θ > 10 (rank : 97) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9GZP0, Q9BWV5 | Gene names | PDGFD, IEGF, SCDGFB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Platelet-derived growth factor D precursor (PDGF D) (Iris-expressed growth factor) (Spinal cord-derived growth factor B) (SCDGF-B). | |||||
|
MFRP_HUMAN
|
||||||
| NC score | 0.184460 (rank : 25) | θ value | θ > 10 (rank : 82) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 165 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9BY79, Q335M3, Q96DQ9 | Gene names | MFRP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane frizzled-related protein (Membrane-type frizzled-related protein). | |||||
|
NRP2_HUMAN
|
||||||
| NC score | 0.181185 (rank : 26) | θ value | θ > 10 (rank : 91) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O60462, O14820, O14821 | Gene names | NRP2, VEGF165R2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuropilin-2 precursor (Vascular endothelial cell growth factor 165 receptor 2). | |||||
|
MFRP_MOUSE
|
||||||
| NC score | 0.180881 (rank : 27) | θ value | θ > 10 (rank : 83) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8K480, Q8BPP4 | Gene names | Mfrp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Membrane frizzled-related protein (Membrane-type frizzled-related protein). | |||||
|
NRP2_MOUSE
|
||||||
| NC score | 0.171260 (rank : 28) | θ value | θ > 10 (rank : 92) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O35375, O35373, O35374, O35376, O35377, O35378 | Gene names | Nrp2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuropilin-2 precursor (Vascular endothelial cell growth factor 165 receptor 2). | |||||
|
DMBT1_MOUSE
|
||||||
| NC score | 0.171091 (rank : 29) | θ value | θ > 10 (rank : 47) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q60997, Q80YC6, Q9JMJ9 | Gene names | Dmbt1, Crpd | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Deleted in malignant brain tumors 1 protein precursor (CRP-ductin) (Vomeroglandin). | |||||
|
CUZD1_HUMAN
|
||||||
| NC score | 0.166402 (rank : 30) | θ value | θ > 10 (rank : 40) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q86UP6, Q7Z660, Q7Z661, Q86SG1, Q86UP5, Q9HAR7 | Gene names | CUZD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CUB and zona pellucida-like domain-containing protein 1 precursor (Transmembrane protein UO-44). | |||||
|
TSG6_HUMAN
|
||||||
| NC score | 0.166225 (rank : 31) | θ value | θ > 10 (rank : 109) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P98066, Q8WWI9 | Gene names | TNFAIP6, TSG6 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor-inducible protein TSG-6 precursor (TNF- stimulated gene 6 protein) (Hyaluronate-binding protein). | |||||
|
SEZ6L_HUMAN
|
||||||
| NC score | 0.164190 (rank : 32) | θ value | θ > 10 (rank : 104) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9BYH1, O95917, Q5THY5, Q6IBZ4, Q6UXD4, Q9NUI3, Q9NUI4, Q9NUI5, Q9Y2E1, Q9Y3J6 | Gene names | SEZ6L, KIAA0927 | |||
|
Domain Architecture |

|
|||||
| Description | Seizure 6-like protein precursor. | |||||
|
TSG6_MOUSE
|
||||||
| NC score | 0.164137 (rank : 33) | θ value | θ > 10 (rank : 110) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O08859 | Gene names | Tnfaip6, Tnfip6, Tsg6 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor-inducible protein TSG-6 precursor (TNF- stimulated gene 6 protein). | |||||
|
CUZD1_MOUSE
|
||||||
| NC score | 0.160372 (rank : 34) | θ value | θ > 10 (rank : 41) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P70412, Q9CTZ7 | Gene names | Cuzd1, Itmap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CUB and zona pellucida-like domain-containing protein 1 precursor (Integral membrane-associated protein 1). | |||||
|
CSMD2_HUMAN
|
||||||
| NC score | 0.157647 (rank : 35) | θ value | θ > 10 (rank : 36) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q7Z408, Q5VT59, Q8N963, Q96Q03, Q9H4V7, Q9H4V8, Q9H4V9, Q9H4W0, Q9H4W1, Q9H4W2, Q9H4W3, Q9H4W4, Q9HCY5, Q9HCY6, Q9HCY7 | Gene names | CSMD2, KIAA1884 | |||
|
Domain Architecture |

|
|||||
| Description | CUB and sushi domain-containing protein 2 (CUB and sushi multiple domains protein 2). | |||||
|
CSMD1_HUMAN
|
||||||
| NC score | 0.153883 (rank : 36) | θ value | θ > 10 (rank : 34) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96PZ7, Q96QU9, Q96RM4 | Gene names | CSMD1, KIAA1890 | |||
|
Domain Architecture |

|
|||||
| Description | CUB and sushi domain-containing protein 1 precursor (CUB and sushi multiple domains protein 1). | |||||
|
CUBN_MOUSE
|
||||||
| NC score | 0.153460 (rank : 37) | θ value | θ > 10 (rank : 39) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 477 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9JLB4 | Gene names | Cubn, Ifcr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cubilin precursor (Intrinsic factor-cobalamin receptor). | |||||
|
CSMD3_HUMAN
|
||||||
| NC score | 0.153140 (rank : 38) | θ value | θ > 10 (rank : 37) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 199 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q7Z407, Q96PZ3 | Gene names | CSMD3, KIAA1894 | |||
|
Domain Architecture |

|
|||||
| Description | CUB and sushi domain-containing protein 3 precursor (CUB and sushi multiple domains protein 3). | |||||
|
CSMD1_MOUSE
|
||||||
| NC score | 0.152285 (rank : 39) | θ value | θ > 10 (rank : 35) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q923L3, Q8BUV1, Q8BYQ3 | Gene names | Csmd1 | |||
|
Domain Architecture |

|
|||||
| Description | CUB and sushi domain-containing protein 1 precursor (CUB and sushi multiple domains protein 1). | |||||
|
CUBN_HUMAN
|
||||||
| NC score | 0.152230 (rank : 40) | θ value | θ > 10 (rank : 38) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 482 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O60494, Q5VTA6, Q96RU9 | Gene names | CUBN, IFCR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cubilin precursor (Intrinsic factor-cobalamin receptor) (Intrinsic factor-vitamin B12 receptor) (460 kDa receptor) (Intestinal intrinsic factor receptor). | |||||
|
LRP12_MOUSE
|
||||||
| NC score | 0.147571 (rank : 41) | θ value | θ > 10 (rank : 65) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BUJ9, Q8BWM9 | Gene names | Lrp12 | |||
|
Domain Architecture |

|
|||||
| Description | Low-density lipoprotein receptor-related protein 12 precursor. | |||||
|
LRP12_HUMAN
|
||||||
| NC score | 0.146999 (rank : 42) | θ value | θ > 10 (rank : 64) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Y561 | Gene names | LRP12, ST7 | |||
|
Domain Architecture |

|
|||||
| Description | Low-density lipoprotein receptor-related protein 12 precursor (Suppressor of tumorigenicity protein 7). | |||||
|
DMBT1_HUMAN
|
||||||
| NC score | 0.138845 (rank : 43) | θ value | θ > 10 (rank : 46) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UGM3, Q59EX0, Q5JR26, Q6MZN4, Q96DU4, Q9UGM2, Q9UJ57, Q9UKJ4, Q9Y211, Q9Y4V9 | Gene names | DMBT1, GP340 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Deleted in malignant brain tumors 1 protein precursor (Glycoprotein 340) (Gp-340) (Surfactant pulmonary-associated D-binding protein). | |||||
|
DCBD1_MOUSE
|
||||||
| NC score | 0.122622 (rank : 44) | θ value | θ > 10 (rank : 43) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9D4J3, Q8R327, Q9D696 | Gene names | Dcbld1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Discoidin, CUB and LCCL domain-containing protein 1 precursor. | |||||
|
DCBD2_HUMAN
|
||||||
| NC score | 0.118628 (rank : 45) | θ value | θ > 10 (rank : 44) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96PD2, Q8N6M4, Q8TDX2 | Gene names | DCBLD2, CLCP1, ESDN | |||
|
Domain Architecture |

|
|||||
| Description | Discoidin, CUB and LCCL domain-containing protein 2 precursor (Endothelial and smooth muscle cell-derived neuropilin-like protein) (CUB, LCCL and coagulation factor V/VIII-homology domains protein 1). | |||||
|
DCBD2_MOUSE
|
||||||
| NC score | 0.114086 (rank : 46) | θ value | θ > 10 (rank : 45) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q91ZV3, Q8BKI4 | Gene names | Dcbld2, Esdn | |||
|
Domain Architecture |

|
|||||
| Description | Discoidin, CUB and LCCL domain-containing protein 2 precursor (Endothelial and smooth muscle cell-derived neuropilin-like protein). | |||||
|
ATRN_HUMAN
|
||||||
| NC score | 0.109731 (rank : 47) | θ value | θ > 10 (rank : 21) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 525 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O75882, O60295, O95414, Q3MIT3, Q5TDA2, Q5TDA4, Q5VYW3, Q9NTQ3, Q9NTQ4, Q9NU01, Q9NZ57, Q9NZ58 | Gene names | ATRN, KIAA0548, MGCA | |||
|
Domain Architecture |

|
|||||
| Description | Attractin precursor (Mahogany homolog) (DPPT-L). | |||||
|
C1R_HUMAN
|
||||||
| NC score | 0.107900 (rank : 48) | θ value | θ > 10 (rank : 26) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P00736, Q8J012 | Gene names | C1R | |||
|
Domain Architecture |

|
|||||
| Description | Complement C1r subcomponent precursor (EC 3.4.21.41) (Complement component 1, r subcomponent) [Contains: Complement C1r subcomponent heavy chain; Complement C1r subcomponent light chain]. | |||||
|
KREM2_HUMAN
|
||||||
| NC score | 0.107555 (rank : 49) | θ value | θ > 10 (rank : 62) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8NCW0, Q8N2J4, Q8NCW1, Q96GL8, Q9BTP9 | Gene names | KREMEN2, KRM2 | |||
|
Domain Architecture |

|
|||||
| Description | Kremen protein 2 precursor (Kringle-containing protein marking the eye and the nose) (Kringle domain-containing transmembrane protein 2) (Dickkopf receptor 2). | |||||
|
LRP3_HUMAN
|
||||||
| NC score | 0.107407 (rank : 50) | θ value | θ > 10 (rank : 66) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O75074 | Gene names | LRP3 | |||
|
Domain Architecture |

|
|||||
| Description | Low-density lipoprotein receptor-related protein 3 precursor (hLRp105). | |||||
|
ATRN_MOUSE
|
||||||
| NC score | 0.107241 (rank : 51) | θ value | θ > 10 (rank : 22) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 519 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9WU60, Q9R263, Q9WU77 | Gene names | Atrn, mg, Mgca | |||
|
Domain Architecture |

|
|||||
| Description | Attractin precursor (Mahogany protein). | |||||
|
MASP1_HUMAN
|
||||||
| NC score | 0.105645 (rank : 52) | θ value | θ > 10 (rank : 71) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 369 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P48740, O95570, Q9UF09 | Gene names | MASP1, CRARF, CRARF1, PRSS5 | |||
|
Domain Architecture |

|
|||||
| Description | Complement-activating component of Ra-reactive factor precursor (EC 3.4.21.-) (Ra-reactive factor serine protease p100) (RaRF) (Mannan-binding lectin serine protease 1) (Mannose-binding protein- associated serine protease) (MASP-1) [Contains: Complement-activating component of Ra-reactive factor heavy chain; Complement-activating component of Ra-reactive factor light chain]. | |||||
|
GP126_HUMAN
|
||||||
| NC score | 0.104706 (rank : 53) | θ value | θ > 10 (rank : 59) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q86SQ4, Q6MZU7, Q8NC14, Q96JW0 | Gene names | GPR126 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 126 precursor. | |||||
|
C1RA_MOUSE
|
||||||
| NC score | 0.104650 (rank : 54) | θ value | θ > 10 (rank : 24) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 379 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CG16, Q99KI6, Q9ET60 | Gene names | C1ra, C1r | |||
|
Domain Architecture |

|
|||||
| Description | Complement C1r-A subcomponent precursor (EC 3.4.21.41) (Complement component 1, r-A subcomponent) [Contains: Complement C1r-A subcomponent heavy chain; Complement C1r-A subcomponent light chain]. | |||||
|
MASP1_MOUSE
|
||||||
| NC score | 0.104055 (rank : 55) | θ value | θ > 10 (rank : 72) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P98064 | Gene names | Masp1, Crarf | |||
|
Domain Architecture |

|
|||||
| Description | Complement-activating component of Ra-reactive factor precursor (EC 3.4.21.-) (Ra-reactive factor serine protease p100) (RaRF) (Mannan-binding lectin serine protease 1) [Contains: Complement- activating component of Ra-reactive factor heavy chain; Complement- activating component of Ra-reactive factor light chain]. | |||||
|
CS1A_MOUSE
|
||||||
| NC score | 0.103560 (rank : 56) | θ value | θ > 10 (rank : 32) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CG14, Q8BJC4, Q8CH28, Q8VBY4 | Gene names | C1sa, C1s | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Complement C1s-A subcomponent precursor (EC 3.4.21.42) (C1 esterase) [Contains: Complement C1s-A subcomponent heavy chain; Complement C1s-A subcomponent light chain]. | |||||
|
DCBD1_HUMAN
|
||||||
| NC score | 0.103456 (rank : 57) | θ value | θ > 10 (rank : 42) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8N8Z6, Q5H992, Q8IYK5, Q8N7L9, Q96NH2 | Gene names | DCBLD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Discoidin, CUB and LCCL domain-containing protein 1 precursor. | |||||
|
C1RB_MOUSE
|
||||||
| NC score | 0.103346 (rank : 58) | θ value | θ > 10 (rank : 25) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CFG9 | Gene names | C1rb, C1r | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Complement C1r-B subcomponent precursor (EC 3.4.21.41) (Complement component 1, r-B subcomponent) [Contains: Complement C1r-B subcomponent heavy chain; Complement C1r-B subcomponent light chain]. | |||||
|
CS1B_MOUSE
|
||||||
| NC score | 0.103275 (rank : 59) | θ value | θ > 10 (rank : 33) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 361 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CFG8 | Gene names | C1sb, C1s | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Complement C1s-B subcomponent precursor (EC 3.4.21.42) (C1 esterase) [Contains: Complement C1s-B subcomponent heavy chain; Complement C1s-B subcomponent light chain]. | |||||
|
MASP2_HUMAN
|
||||||
| NC score | 0.102458 (rank : 60) | θ value | θ > 10 (rank : 73) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O00187, O75754, Q5TEQ5, Q5TER0, Q96QG4, Q9BZH0, Q9H498, Q9H499, Q9UBP3, Q9ULC7, Q9UMV3, Q9Y270 | Gene names | MASP2 | |||
|
Domain Architecture |

|
|||||
| Description | Mannan-binding lectin serine protease 2 precursor (EC 3.4.21.104) (Mannose-binding protein-associated serine protease 2) (MASP-2) (MBL- associated serine protease 2) [Contains: Mannan-binding lectin serine protease 2 A chain; Mannan-binding lectin serine protease 2 B chain]. | |||||
|
C1S_HUMAN
|
||||||
| NC score | 0.102299 (rank : 61) | θ value | θ > 10 (rank : 27) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 359 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P09871, Q9UCU7, Q9UCU8, Q9UCU9, Q9UCV0, Q9UCV1, Q9UCV2, Q9UCV3, Q9UCV4, Q9UCV5, Q9UM14 | Gene names | C1S | |||
|
Domain Architecture |

|
|||||
| Description | Complement C1s subcomponent precursor (EC 3.4.21.42) (C1 esterase) [Contains: Complement C1s subcomponent heavy chain; Complement C1s subcomponent light chain]. | |||||
|
MASP2_MOUSE
|
||||||
| NC score | 0.101205 (rank : 62) | θ value | θ > 10 (rank : 74) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 382 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q91WP0, Q9QXA4, Q9QXD2, Q9QXD5, Q9Z338 | Gene names | Masp2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mannan-binding lectin serine protease 2 precursor (EC 3.4.21.104) (Mannose-binding protein-associated serine protease 2) (MASP-2) (MBL- associated serine protease 2) [Contains: Mannan-binding lectin serine protease 2 A chain; Mannan-binding lectin serine protease 2 B chain]. | |||||
|
KREM2_MOUSE
|
||||||
| NC score | 0.100527 (rank : 63) | θ value | θ > 10 (rank : 63) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8K1S7 | Gene names | Kremen2 | |||
|
Domain Architecture |

|
|||||
| Description | Kremen protein 2 precursor (Kringle-containing protein marking the eye and the nose) (Kringle domain-containing transmembrane protein 2) (Dickkopf receptor 2). | |||||
|
MAMC2_MOUSE
|
||||||
| NC score | 0.098205 (rank : 64) | θ value | θ > 10 (rank : 70) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CG85, Q8BM24 | Gene names | Mamdc2 | |||
|
Domain Architecture |

|
|||||
| Description | MAM domain-containing protein 2 precursor. | |||||
|
KREM1_HUMAN
|
||||||
| NC score | 0.094131 (rank : 65) | θ value | θ > 10 (rank : 60) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96MU8, Q5TIB9, Q6P3X6, Q9BY70, Q9UGS5, Q9UGU1 | Gene names | KREMEN1, KREMEN, KRM1 | |||
|
Domain Architecture |

|
|||||
| Description | Kremen protein 1 precursor (Kringle-containing protein marking the eye and the nose) (Kringle domain-containing transmembrane protein 1) (Dickkopf receptor). | |||||
|
MAMC2_HUMAN
|
||||||
| NC score | 0.089846 (rank : 66) | θ value | θ > 10 (rank : 69) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q7Z304, Q5VW47, Q8WX43, Q96BM4 | Gene names | MAMDC2 | |||
|
Domain Architecture |

|
|||||
| Description | MAM domain-containing protein 2 precursor. | |||||
|
KREM1_MOUSE
|
||||||
| NC score | 0.082391 (rank : 67) | θ value | θ > 10 (rank : 61) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q99N43 | Gene names | Kremen1, Kremen | |||
|
Domain Architecture |

|
|||||
| Description | Kremen protein 1 precursor (Kringle-containing protein marking the eye and the nose) (Kringle domain-containing transmembrane protein 1) (Dickkopf receptor). | |||||
|
EGFL7_HUMAN
|
||||||
| NC score | 0.081007 (rank : 68) | θ value | θ > 10 (rank : 48) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UHF1, Q5M7Y5, Q5VUD5, Q96EG0 | Gene names | EGFL7, MEGF7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EGF-like domain-containing protein 7 precursor (Multiple EGF-like domain protein 7) (Multiple epidermal growth factor-like domain protein 7) (Vascular endothelial statin) (VE-statin) (NOTCH4-like protein) (ZNEU1). | |||||
|
ENTK_MOUSE
|
||||||
| NC score | 0.079111 (rank : 69) | θ value | θ > 10 (rank : 53) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 338 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P97435 | Gene names | Prss7, Entk | |||
|
Domain Architecture |

|
|||||
| Description | Enteropeptidase (EC 3.4.21.9) (Enterokinase) [Contains: Enteropeptidase non-catalytic heavy chain; Enteropeptidase catalytic light chain]. | |||||
|
ENTK_HUMAN
|
||||||
| NC score | 0.072353 (rank : 70) | θ value | θ > 10 (rank : 52) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 310 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P98073 | Gene names | PRSS7, ENTK | |||
|
Domain Architecture |

|
|||||
| Description | Enteropeptidase precursor (EC 3.4.21.9) (Enterokinase) [Contains: Enteropeptidase non-catalytic heavy chain; Enteropeptidase catalytic light chain]. | |||||
|
TRAF4_MOUSE
|
||||||
| NC score | 0.070650 (rank : 71) | θ value | θ > 10 (rank : 108) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 338 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q61382 | Gene names | Traf4, Cart1 | |||
|
Domain Architecture |

|
|||||
| Description | TNF receptor-associated factor 4 (Cysteine-rich motif associated to RING and Traf domains protein 1). | |||||
|
TRAF4_HUMAN
|
||||||
| NC score | 0.067009 (rank : 72) | θ value | θ > 10 (rank : 107) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 373 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9BUZ4, O75615, Q14848, Q2PJN8 | Gene names | TRAF4, CART1, MLN62, RNF83 | |||
|
Domain Architecture |

|
|||||
| Description | TNF receptor-associated factor 4 (Cysteine-rich domain associated with RING and Traf domains protein 1) (Malignant 62) (RING finger protein 83). | |||||
|
PROS_HUMAN
|
||||||
| NC score | 0.065801 (rank : 73) | θ value | θ > 10 (rank : 100) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 340 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P07225, Q15518, Q7Z715 | Gene names | PROS1, PROS | |||
|
Domain Architecture |

|
|||||
| Description | Vitamin K-dependent protein S precursor. | |||||
|
SCUB3_MOUSE
|
||||||
| NC score | 0.064349 (rank : 74) | θ value | θ > 10 (rank : 103) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 442 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q66PY1, Q68FG9 | Gene names | Scube3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal peptide, CUB and EGF-like domain-containing protein 3 precursor. | |||||
|
MATN2_MOUSE
|
||||||
| NC score | 0.063549 (rank : 75) | θ value | θ > 10 (rank : 78) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 356 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O08746 | Gene names | Matn2 | |||
|
Domain Architecture |

|
|||||
| Description | Matrilin-2 precursor. | |||||
|
SCUB3_HUMAN
|
||||||
| NC score | 0.063203 (rank : 76) | θ value | θ > 10 (rank : 102) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 428 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8IX30, Q5CZB3, Q86UZ9, Q8NAU9 | Gene names | SCUBE3, CEGF3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal peptide, CUB and EGF-like domain-containing protein 3 precursor. | |||||
|
MDGA1_HUMAN
|
||||||
| NC score | 0.063156 (rank : 77) | θ value | θ > 10 (rank : 80) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 358 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8NFP4, Q8NBE3 | Gene names | MDGA1, MAMDC3 | |||
|
Domain Architecture |

|
|||||
| Description | MAM domain-containing glycosylphosphatidylinositol anchor protein 1 precursor (MAM domain-containing protein 3) (GPI and MAM protein) (Glycosylphosphatidylinositol-MAM) (GPIM). | |||||
|
LTBP3_MOUSE
|
||||||
| NC score | 0.062661 (rank : 78) | θ value | θ > 10 (rank : 68) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 475 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q61810, Q8BNQ6 | Gene names | Ltbp3 | |||
|
Domain Architecture |

|
|||||
| Description | Latent-transforming growth factor beta-binding protein 3 precursor (LTBP-3). | |||||
|
ST14_HUMAN
|
||||||
| NC score | 0.062477 (rank : 79) | θ value | θ > 10 (rank : 105) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 409 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Y5Y6, Q9BS01, Q9H3S0, Q9HB36, Q9HCA3 | Gene names | ST14, PRSS14, SNC19, TADG15 | |||
|
Domain Architecture |

|
|||||
| Description | Suppressor of tumorigenicity protein 14 (EC 3.4.21.-) (Serine protease 14) (Matriptase) (Membrane-type serine protease 1) (MT-SP1) (Prostamin) (Serine protease TADG-15) (Tumor-associated differentially-expressed gene 15 protein). | |||||
|
C1QR1_MOUSE
|
||||||
| NC score | 0.062240 (rank : 80) | θ value | θ > 10 (rank : 23) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O89103, Q3U2X0 | Gene names | Cd93, Aa4, C1qr1, C1qrp, Ly68 | |||
|
Domain Architecture |

|
|||||
| Description | Complement component C1q receptor precursor (Complement component 1 q subcomponent receptor 1) (C1qRp) (C1qR(p)) (C1q/MBL/SPA receptor) (CD93 antigen) (Cell surface antigen AA4) (Lymphocyte antigen 68). | |||||
|
ST14_MOUSE
|
||||||
| NC score | 0.060905 (rank : 81) | θ value | θ > 10 (rank : 106) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P56677 | Gene names | St14, Prss14 | |||
|
Domain Architecture |

|
|||||
| Description | Suppressor of tumorigenicity protein 14 (EC 3.4.21.-) (Serine protease 14) (Epithin). | |||||
|
MATN2_HUMAN
|
||||||
| NC score | 0.060833 (rank : 82) | θ value | θ > 10 (rank : 77) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 365 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O00339, Q6UWA5, Q7Z5X1, Q8NDE6, Q96FT5, Q9NSZ1 | Gene names | MATN2 | |||
|
Domain Architecture |

|
|||||
| Description | Matrilin-2 precursor. | |||||
|
PHXR5_MOUSE
|
||||||
| NC score | 0.060629 (rank : 83) | θ value | θ > 10 (rank : 99) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P08399 | Gene names | Phxr5, Per | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Per-hexamer repeat protein 5. | |||||
|
PROS_MOUSE
|
||||||
| NC score | 0.058650 (rank : 84) | θ value | θ > 10 (rank : 101) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 330 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q08761, P43483 | Gene names | Pros1, Pros | |||
|
Domain Architecture |

|
|||||
| Description | Vitamin K-dependent protein S precursor. | |||||
|
MATN1_MOUSE
|
||||||
| NC score | 0.057912 (rank : 85) | θ value | θ > 10 (rank : 76) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 186 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P51942 | Gene names | Matn1, Cmp, Crtm | |||
|
Domain Architecture |

|
|||||
| Description | Cartilage matrix protein precursor (Matrilin-1). | |||||
|
CLC14_HUMAN
|
||||||
| NC score | 0.057848 (rank : 86) | θ value | θ > 10 (rank : 30) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q86T13, Q6PWT6, Q8N5V5 | Gene names | CLEC14A, C14orf27, EGFR5 | |||
|
Domain Architecture |

|
|||||
| Description | C-type lectin domain family 14 member A precursor (Epidermal growth factor receptor 5) (EGFR-5). | |||||
|
FBLN5_HUMAN
|
||||||
| NC score | 0.056993 (rank : 87) | θ value | θ > 10 (rank : 54) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 329 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UBX5, O75966, Q6UWA3 | Gene names | FBLN5, DANCE | |||
|
Domain Architecture |

|
|||||
| Description | Fibulin-5 precursor (FIBL-5) (Developmental arteries and neural crest EGF-like protein) (Dance) (Urine p50 protein) (UP50). | |||||
|
MATN1_HUMAN
|
||||||
| NC score | 0.056317 (rank : 88) | θ value | θ > 10 (rank : 75) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P21941 | Gene names | MATN1, CMP, CRTM | |||
|
Domain Architecture |

|
|||||
| Description | Cartilage matrix protein precursor (Matrilin-1). | |||||
|
CLC14_MOUSE
|
||||||
| NC score | 0.056078 (rank : 89) | θ value | θ > 10 (rank : 31) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8VCP9, Q3TP72, Q9CXA8, Q9D624, Q9DC55 | Gene names | Clec14a | |||
|
Domain Architecture |

|
|||||
| Description | C-type lectin domain family 14 member A precursor. | |||||
|
CD248_MOUSE
|
||||||
| NC score | 0.055656 (rank : 90) | θ value | θ > 10 (rank : 29) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 578 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q91V98, Q3UMV6, Q91ZV1 | Gene names | Cd248, Tem1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endosialin precursor (Tumor endothelial marker 1) (CD248 antigen). | |||||
|
FBN1_MOUSE
|
||||||
| NC score | 0.054886 (rank : 91) | θ value | θ > 10 (rank : 57) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 563 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q61554, Q60826 | Gene names | Fbn1, Fbn-1 | |||
|
Domain Architecture |

|
|||||
| Description | Fibrillin-1 precursor. | |||||
|
FBLN5_MOUSE
|
||||||
| NC score | 0.054716 (rank : 92) | θ value | θ > 10 (rank : 55) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 317 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9WVH9, Q541Z7 | Gene names | Fbln5, Dance | |||
|
Domain Architecture |

|
|||||
| Description | Fibulin-5 precursor (FIBL-5) (Developmental arteries and neural crest EGF-like protein) (Dance). | |||||
|
MAMC1_MOUSE
|
||||||
| NC score | 0.054712 (rank : 93) | θ value | 8.99809 (rank : 20) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P60755 | Gene names | Mamdc1, Mdga2 | |||
|
Domain Architecture |

|
|||||
| Description | MAM domain-containing protein 1 precursor (MAM domain-containing glycosylphosphatidylinositol anchor protein 2). | |||||
|
MATN4_HUMAN
|
||||||
| NC score | 0.054705 (rank : 94) | θ value | θ > 10 (rank : 79) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 287 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O95460, Q5QPU2, Q5QPU3, Q5QPU4, Q8N2M5, Q8N2M7, Q9H1F8, Q9H1F9 | Gene names | MATN4 | |||
|
Domain Architecture |

|
|||||
| Description | Matrilin-4 precursor. | |||||
|
MUC13_MOUSE
|
||||||
| NC score | 0.054486 (rank : 95) | θ value | θ > 10 (rank : 84) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P19467 | Gene names | Muc13, Ly64 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-13 precursor (Cell surface antigen 114/A10) (Lymphocyte antigen 64). | |||||
|
MAMC1_HUMAN
|
||||||
| NC score | 0.054071 (rank : 96) | θ value | 8.99809 (rank : 19) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q7Z553 | Gene names | MAMDC1, MDGA2 | |||
|
Domain Architecture |

|
|||||
| Description | MAM domain-containing protein 1 precursor (MAM domain-containing glycosylphosphatidylinositol anchor protein 2). | |||||
|
LTBP2_MOUSE
|
||||||
| NC score | 0.053005 (rank : 97) | θ value | θ > 10 (rank : 67) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 617 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O08999, Q8C6W9 | Gene names | Ltbp2 | |||
|
Domain Architecture |

|
|||||
| Description | Latent-transforming growth factor beta-binding protein 2 precursor (LTBP-2). | |||||
|
FBN1_HUMAN
|
||||||
| NC score | 0.052923 (rank : 98) | θ value | θ > 10 (rank : 56) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 557 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P35555, Q15972, Q75N87 | Gene names | FBN1, FBN | |||
|
Domain Architecture |

|
|||||
| Description | Fibrillin-1 precursor. | |||||
|
CD248_HUMAN
|
||||||
| NC score | 0.051572 (rank : 99) | θ value | θ > 10 (rank : 28) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 679 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9HCU0, Q3SX55, Q96KB6 | Gene names | CD248, CD164L1, TEM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endosialin precursor (Tumor endothelial marker 1) (CD248 antigen). | |||||
|
FBN3_HUMAN
|
||||||
| NC score | 0.051500 (rank : 100) | θ value | θ > 10 (rank : 58) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q75N90, Q75N91, Q75N92, Q75N93, Q86SJ5, Q96JP8 | Gene names | FBN3, KIAA1776 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fibrillin-3 precursor. | |||||
|
EGF_MOUSE
|
||||||
| NC score | 0.051340 (rank : 101) | θ value | θ > 10 (rank : 51) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 358 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P01132, Q6P9J2 | Gene names | Egf | |||
|
Domain Architecture |

|
|||||
| Description | Pro-epidermal growth factor precursor (EGF) [Contains: Epidermal growth factor]. | |||||
|
EGFL7_MOUSE
|
||||||
| NC score | 0.051087 (rank : 102) | θ value | θ > 10 (rank : 49) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 287 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9QXT5, Q6XD35, Q9DCP5 | Gene names | Egfl7, Megf7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EGF-like domain-containing protein 7 precursor (Multiple EGF-like domain protein 7) (Multiple epidermal growth factor-like domain protein 7) (Vascular endothelial statin) (VE-statin) (NOTCH4-like protein) (Zneu1). | |||||
|
EGFL8_MOUSE
|
||||||
| NC score | 0.051039 (rank : 103) | θ value | θ > 10 (rank : 50) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q6GUQ1, O35447 | Gene names | Egfl8, Ng3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EGF-like domain-containing protein 8 precursor (Epidermal growth factor-like protein 8) (Multiple EGF-like domain protein 8). | |||||
|
MEGF6_HUMAN
|
||||||
| NC score | 0.050596 (rank : 104) | θ value | θ > 10 (rank : 81) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O75095, Q5VV39 | Gene names | MEGF6, EGFL3, KIAA0815 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Multiple epidermal growth factor-like domains 6 precursor (EGF-like domain-containing protein 3) (Multiple EGF-like domain protein 3). | |||||
|
UROL1_MOUSE
|
||||||
| NC score | 0.050194 (rank : 105) | θ value | θ > 10 (rank : 111) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q5DID3, Q5DID1, Q5DID2 | Gene names | Umodl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uromodulin-like 1 precursor (Olfactorin). | |||||
|
PDCD7_HUMAN
|
||||||
| NC score | 0.040785 (rank : 106) | θ value | 1.06291 (rank : 13) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8N8D1, Q96AK8, Q9Y6D7 | Gene names | PDCD7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Programmed cell death protein 7 (ES18) (HES18). | |||||
|
PDCD7_MOUSE
|
||||||
| NC score | 0.037937 (rank : 107) | θ value | 2.36792 (rank : 15) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 214 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9WTY1, Q8R5D9 | Gene names | Pdcd7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Programmed cell death protein 7 (ES18). | |||||
|
CS024_HUMAN
|
||||||
| NC score | 0.036461 (rank : 108) | θ value | 4.03905 (rank : 16) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9BVV8, Q9NWS2 | Gene names | C19orf24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C19orf24. | |||||
|
SMG7_HUMAN
|
||||||
| NC score | 0.016368 (rank : 109) | θ value | 4.03905 (rank : 17) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q92540, Q5T1Q0, Q6PIE0, Q7Z7H9, Q8IXC1, Q8IXC2 | Gene names | SMG7, C1orf16, EST1C, KIAA0250 | |||
|
Domain Architecture |

|
|||||
| Description | Protein SMG7 (SMG-7 homolog) (EST1-like protein C). | |||||
|
NDE1_HUMAN
|
||||||
| NC score | 0.010041 (rank : 110) | θ value | 1.81305 (rank : 14) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 823 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NXR1, Q49AQ2 | Gene names | NDE1, NUDE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear distribution protein nudE homolog 1 (NudE). | |||||
|
MLL2_HUMAN
|
||||||
| NC score | 0.009227 (rank : 111) | θ value | 5.27518 (rank : 18) | |||
| Query Neighborhood Hits | 20 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||