Please be patient as the page loads
|
ADCY8_MOUSE
|
||||||
| SwissProt Accessions | P97490 | Gene names | Adcy8 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylate cyclase type 8 (EC 4.6.1.1) (Adenylate cyclase type VIII) (ATP pyrophosphate-lyase 8) (Ca(2+)/calmodulin-activated adenylyl cyclase). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
ADCY1_MOUSE
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.982307 (rank : 5) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O88444, Q5SS89 | Gene names | Adcy1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Adenylate cyclase type 1 (EC 4.6.1.1) (Adenylate cyclase type I) (ATP pyrophosphate-lyase 1) (Ca(2+)/calmodulin-activated adenylyl cyclase). | |||||
|
ADCY3_HUMAN
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.979628 (rank : 7) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O60266, Q9UDB1 | Gene names | ADCY3, KIAA0511 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylate cyclase type 3 (EC 4.6.1.1) (Adenylate cyclase type III) (Adenylate cyclase, olfactive type) (ATP pyrophosphate-lyase 3) (Adenylyl cyclase 3) (AC-III) (AC3). | |||||
|
ADCY3_MOUSE
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.978268 (rank : 9) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8VHH7 | Gene names | Adcy3 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylate cyclase type 3 (EC 4.6.1.1) (Adenylate cyclase type III) (Adenylate cyclase, olfactive type) (ATP pyrophosphate-lyase 3) (Adenylyl cyclase 3) (AC-III) (AC3). | |||||
|
ADCY6_HUMAN
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.985220 (rank : 3) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O43306, Q9NR75 | Gene names | ADCY6, KIAA0422 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylate cyclase type 6 (EC 4.6.1.1) (Adenylate cyclase type VI) (ATP pyrophosphate-lyase 6) (Ca(2+)-inhibitable adenylyl cyclase). | |||||
|
ADCY6_MOUSE
|
||||||
| θ value | 0 (rank : 5) | NC score | 0.984451 (rank : 4) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q01341 | Gene names | Adcy6 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylate cyclase type 6 (EC 4.6.1.1) (Adenylate cyclase type VI) (ATP pyrophosphate-lyase 6) (Ca(2+)-inhibitable adenylyl cyclase). | |||||
|
ADCY8_HUMAN
|
||||||
| θ value | 0 (rank : 6) | NC score | 0.999786 (rank : 2) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P40145 | Gene names | ADCY8 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylate cyclase type 8 (EC 4.6.1.1) (Adenylate cyclase type VIII) (ATP pyrophosphate-lyase 8) (Ca(2+)/calmodulin-activated adenylyl cyclase). | |||||
|
ADCY8_MOUSE
|
||||||
| θ value | 0 (rank : 7) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P97490 | Gene names | Adcy8 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylate cyclase type 8 (EC 4.6.1.1) (Adenylate cyclase type VIII) (ATP pyrophosphate-lyase 8) (Ca(2+)/calmodulin-activated adenylyl cyclase). | |||||
|
ADCY1_HUMAN
|
||||||
| θ value | 2.24492e-184 (rank : 8) | NC score | 0.979784 (rank : 6) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q08828, Q75MI1 | Gene names | ADCY1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Adenylate cyclase type 1 (EC 4.6.1.1) (Adenylate cyclase type I) (ATP pyrophosphate-lyase 1) (Ca(2+)/calmodulin-activated adenylyl cyclase). | |||||
|
ADCY2_MOUSE
|
||||||
| θ value | 4.1007e-178 (rank : 9) | NC score | 0.978469 (rank : 8) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q80TL1, Q8CFU6 | Gene names | Adcy2, Kiaa1060 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Adenylate cyclase type 2 (EC 4.6.1.1) (Adenylate cyclase type II) (ATP pyrophosphate-lyase 2) (Adenylyl cyclase 2). | |||||
|
ADCY2_HUMAN
|
||||||
| θ value | 2.84643e-171 (rank : 10) | NC score | 0.976397 (rank : 12) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q08462, Q9UPU2 | Gene names | ADCY2, KIAA1060 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylate cyclase type 2 (EC 4.6.1.1) (Adenylate cyclase type II) (ATP pyrophosphate-lyase 2) (Adenylyl cyclase 2). | |||||
|
ADCY4_HUMAN
|
||||||
| θ value | 1.15832e-164 (rank : 11) | NC score | 0.975192 (rank : 14) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8NFM4, Q96ML7 | Gene names | ADCY4 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylate cyclase type 4 (EC 4.6.1.1) (Adenylate cyclase type IV) (ATP pyrophosphate-lyase 4) (Adenylyl cyclase 4). | |||||
|
ADCY7_HUMAN
|
||||||
| θ value | 1.84928e-162 (rank : 12) | NC score | 0.976844 (rank : 11) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P51828 | Gene names | ADCY7, KIAA0037 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Adenylate cyclase type 7 (EC 4.6.1.1) (Adenylate cyclase type VII) (ATP pyrophosphate-lyase 7) (Adenylyl cyclase 7). | |||||
|
ADCY4_MOUSE
|
||||||
| θ value | 5.94893e-161 (rank : 13) | NC score | 0.975857 (rank : 13) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q91WF3 | Gene names | Adcy4 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylate cyclase type 4 (EC 4.6.1.1) (Adenylate cyclase type IV) (ATP pyrophosphate-lyase 4) (Adenylyl cyclase 4). | |||||
|
ADCY7_MOUSE
|
||||||
| θ value | 2.18956e-155 (rank : 14) | NC score | 0.977923 (rank : 10) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P51829 | Gene names | Adcy7 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylate cyclase type 7 (EC 4.6.1.1) (Adenylate cyclase type VII) (ATP pyrophosphate-lyase 7) (Adenylyl cyclase 7). | |||||
|
ADCY9_MOUSE
|
||||||
| θ value | 2.48917e-58 (rank : 15) | NC score | 0.906568 (rank : 15) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P51830, Q61279 | Gene names | Adcy9 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylate cyclase type 9 (EC 4.6.1.1) (Adenylate cyclase type IX) (ATP pyrophosphate-lyase 9) (Adenylyl cyclase 9) (Adenylyl cyclase type 10) (ACTP10). | |||||
|
ADCY9_HUMAN
|
||||||
| θ value | 4.24587e-58 (rank : 16) | NC score | 0.906395 (rank : 16) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O60503, O60273, Q4ZHT9, Q4ZIR5, Q9BWT4, Q9UGP2 | Gene names | ADCY9, KIAA0520 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylate cyclase type 9 (EC 4.6.1.1) (Adenylate cyclase type IX) (ATP pyrophosphate-lyase 9) (Adenylyl cyclase 9). | |||||
|
ANPRA_HUMAN
|
||||||
| θ value | 2.68423e-28 (rank : 17) | NC score | 0.446835 (rank : 28) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 612 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P16066, Q6P4Q3 | Gene names | NPR1, ANPRA | |||
|
Domain Architecture |
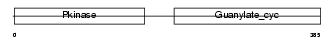
|
|||||
| Description | Atrial natriuretic peptide receptor A precursor (ANP-A) (ANPRA) (GC-A) (Guanylate cyclase) (EC 4.6.1.2) (NPR-A) (Atrial natriuretic peptide A-type receptor). | |||||
|
ANPRA_MOUSE
|
||||||
| θ value | 5.97985e-28 (rank : 18) | NC score | 0.447826 (rank : 27) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 600 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P18293 | Gene names | Npr1, Npra | |||
|
Domain Architecture |

|
|||||
| Description | Atrial natriuretic peptide receptor A precursor (ANP-A) (ANPRA) (GC-A) (Guanylate cyclase) (EC 4.6.1.2) (NPR-A) (Atrial natriuretic peptide A-type receptor). | |||||
|
ANPRB_HUMAN
|
||||||
| θ value | 1.92365e-26 (rank : 19) | NC score | 0.442792 (rank : 29) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 646 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P20594, O60871, Q9UQ50 | Gene names | NPR2, ANPRB | |||
|
Domain Architecture |

|
|||||
| Description | Atrial natriuretic peptide receptor B precursor (ANP-B) (ANPRB) (GC-B) (Guanylate cyclase B) (EC 4.6.1.2) (NPR-B) (Atrial natriuretic peptide B-type receptor). | |||||
|
GUC2E_MOUSE
|
||||||
| θ value | 1.92365e-26 (rank : 20) | NC score | 0.495708 (rank : 23) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 692 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P52785 | Gene names | Gucy2e, Guc2e | |||
|
Domain Architecture |

|
|||||
| Description | Guanylyl cyclase GC-E precursor (EC 4.6.1.2) (Guanylate cyclase 2E). | |||||
|
ANPRB_MOUSE
|
||||||
| θ value | 7.30988e-26 (rank : 21) | NC score | 0.455251 (rank : 26) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 607 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q6VVW5, Q6VVW3, Q6VVW4, Q8CGA9 | Gene names | Npr2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Atrial natriuretic peptide receptor B precursor (ANP-B) (ANPRB) (GC-B) (Guanylate cyclase B) (EC 4.6.1.2) (NPR-B) (Atrial natriuretic peptide B-type receptor). | |||||
|
GUC2F_HUMAN
|
||||||
| θ value | 1.24688e-25 (rank : 22) | NC score | 0.419375 (rank : 31) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 740 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P51841, Q9UJF1 | Gene names | GUCY2F, GUC2F, RETGC2 | |||
|
Domain Architecture |

|
|||||
| Description | Retinal guanylyl cyclase 2 precursor (EC 4.6.1.2) (Guanylate cyclase 2F, retinal) (RETGC-2) (Rod outer segment membrane guanylate cyclase 2) (ROS-GC2) (Guanylate cyclase F) (GC-F). | |||||
|
GCYB1_HUMAN
|
||||||
| θ value | 4.01107e-24 (rank : 23) | NC score | 0.778257 (rank : 22) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q02153, Q86WY5 | Gene names | GUCY1B3, GUC1B3, GUCSB3, GUCY1B1 | |||
|
Domain Architecture |

|
|||||
| Description | Guanylate cyclase soluble subunit beta-1 (EC 4.6.1.2) (GCS-beta-1) (Soluble guanylate cyclase small subunit) (GCS-beta-3). | |||||
|
GCYB2_HUMAN
|
||||||
| θ value | 4.01107e-24 (rank : 24) | NC score | 0.806589 (rank : 17) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O75343, Q9NZ64 | Gene names | GUCY1B2 | |||
|
Domain Architecture |

|
|||||
| Description | Guanylate cyclase soluble subunit beta-2 (EC 4.6.1.2) (GCS-beta-2). | |||||
|
GUC2D_HUMAN
|
||||||
| θ value | 5.23862e-24 (rank : 25) | NC score | 0.492111 (rank : 24) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 715 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q02846, Q6LEA7 | Gene names | GUCY2D, CORD6, GUC1A4, GUC2D, RETGC, RETGC1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinal guanylyl cyclase 1 precursor (EC 4.6.1.2) (Guanylate cyclase 2D, retinal) (RETGC-1) (Rod outer segment membrane guanylate cyclase) (ROS-GC). | |||||
|
GCYB1_MOUSE
|
||||||
| θ value | 6.84181e-24 (rank : 26) | NC score | 0.778445 (rank : 21) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O54865 | Gene names | Gucy1b3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Guanylate cyclase soluble subunit beta-1 (EC 4.6.1.2) (GCS-beta-1) (Soluble guanylate cyclase small subunit). | |||||
|
GUC2G_MOUSE
|
||||||
| θ value | 4.43474e-23 (rank : 27) | NC score | 0.462401 (rank : 25) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 644 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q6TL19, Q8BWU7 | Gene names | Gucy2g | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Guanylate cyclase 2G precursor (EC 4.6.1.2) (Guanylyl cyclase receptor G) (mGC-G). | |||||
|
GUC2C_HUMAN
|
||||||
| θ value | 3.75424e-22 (rank : 28) | NC score | 0.433912 (rank : 30) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 624 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P25092 | Gene names | GUCY2C, GUC2C, STAR | |||
|
Domain Architecture |

|
|||||
| Description | Heat-stable enterotoxin receptor precursor (GC-C) (Intestinal guanylate cyclase) (EC 4.6.1.2) (STA receptor) (hSTAR). | |||||
|
GCYA2_HUMAN
|
||||||
| θ value | 6.40375e-22 (rank : 29) | NC score | 0.783583 (rank : 18) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P33402 | Gene names | GUCY1A2, GUC1A2, GUCSA2 | |||
|
Domain Architecture |

|
|||||
| Description | Guanylate cyclase soluble subunit alpha-2 (EC 4.6.1.2) (GCS-alpha-2). | |||||
|
GCYA3_HUMAN
|
||||||
| θ value | 4.15078e-21 (rank : 30) | NC score | 0.780755 (rank : 19) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q02108, O43843, Q8TAH3 | Gene names | GUCY1A3, GUC1A3, GUCSA3, GUCY1A1 | |||
|
Domain Architecture |

|
|||||
| Description | Guanylate cyclase soluble subunit alpha-3 (EC 4.6.1.2) (GCS-alpha-3) (Soluble guanylate cyclase large subunit) (GCS-alpha-1). | |||||
|
GCYA3_MOUSE
|
||||||
| θ value | 7.0802e-21 (rank : 31) | NC score | 0.778795 (rank : 20) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9ERL9, Q9DBQ3 | Gene names | Gucy1a3, Gucy1a1 | |||
|
Domain Architecture |

|
|||||
| Description | Guanylate cyclase soluble subunit alpha-3 (EC 4.6.1.2) (GCS-alpha-3) (Soluble guanylate cyclase large subunit) (GCS-alpha-1). | |||||
|
TM16H_MOUSE
|
||||||
| θ value | 0.00869519 (rank : 32) | NC score | 0.062397 (rank : 34) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6PB70, Q69ZE4 | Gene names | Tmem16h, Kiaa1623 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein 16H. | |||||
|
TM16H_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 33) | NC score | 0.053465 (rank : 35) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9HCE9 | Gene names | TMEM16H, KIAA1623 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein 16H. | |||||
|
COCA1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 34) | NC score | 0.005910 (rank : 40) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q60847, P70322 | Gene names | Col12a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XII) chain precursor. | |||||
|
E2AK4_MOUSE
|
||||||
| θ value | 1.06291 (rank : 35) | NC score | 0.014869 (rank : 36) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 984 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9QZ05, Q6GQT4, Q6ZPT5, Q8C5S0, Q8CIF5, Q9CT30, Q9CUV9, Q9ESB6, Q9ESB7, Q9ESB8 | Gene names | Eif2ak4, Gcn2, Kiaa1338 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 2-alpha kinase 4 (EC 2.7.11.1) (GCN2-like protein) (mGCN2). | |||||
|
TRHY_HUMAN
|
||||||
| θ value | 1.81305 (rank : 36) | NC score | 0.002503 (rank : 43) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 1385 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q07283 | Gene names | THH, THL, TRHY | |||
|
Domain Architecture |

|
|||||
| Description | Trichohyalin. | |||||
|
COCA1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 37) | NC score | 0.004877 (rank : 42) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99715, Q99716 | Gene names | COL12A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XII) chain precursor. | |||||
|
LY10_HUMAN
|
||||||
| θ value | 3.0926 (rank : 38) | NC score | 0.014076 (rank : 37) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q13342, Q13341, Q92881, Q96TG3 | Gene names | SP140, LYSP100 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear body protein SP140 (Nuclear autoantigen Sp-140) (Speckled 140 kDa) (LYSp100 protein) (Lymphoid-restricted homolog of Sp100). | |||||
|
RHBT1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 39) | NC score | 0.006243 (rank : 39) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O94844 | Gene names | RHOBTB1, KIAA0740 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-related BTB domain-containing protein 1. | |||||
|
PRDM1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 40) | NC score | -0.003348 (rank : 44) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 766 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q60636 | Gene names | Prdm1, Blimp1 | |||
|
Domain Architecture |

|
|||||
| Description | PR domain zinc finger protein 1 (PR domain-containing protein 1) (Beta-interferon gene positive-regulatory domain I-binding factor) (BLIMP-1). | |||||
|
PUR2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 41) | NC score | 0.006799 (rank : 38) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P22102 | Gene names | GART, PRGS | |||
|
Domain Architecture |

|
|||||
| Description | Trifunctional purine biosynthetic protein adenosine-3 [Includes: Phosphoribosylamine--glycine ligase (EC 6.3.4.13) (GARS) (Glycinamide ribonucleotide synthetase) (Phosphoribosylglycinamide synthetase); Phosphoribosylformylglycinamidine cyclo-ligase (EC 6.3.3.1) (AIRS) (Phosphoribosyl-aminoimidazole synthetase) (AIR synthase); Phosphoribosylglycinamide formyltransferase (EC 2.1.2.2) (GART) (GAR transformylase) (5'-phosphoribosylglycinamide transformylase)]. | |||||
|
LMIP_MOUSE
|
||||||
| θ value | 8.99809 (rank : 42) | NC score | 0.005881 (rank : 41) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P56563, Q3TNV8, Q99PA6 | Gene names | Lim2 | |||
|
Domain Architecture |

|
|||||
| Description | Lens fiber membrane intrinsic protein (MP17) (MP18) (MP19) (MP20). | |||||
|
ANPRC_HUMAN
|
||||||
| θ value | θ > 10 (rank : 43) | NC score | 0.157747 (rank : 32) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P17342 | Gene names | NPR3, ANPRC | |||
|
Domain Architecture |
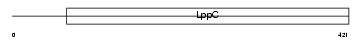
|
|||||
| Description | Atrial natriuretic peptide clearance receptor precursor (ANP-C) (ANPRC) (NPR-C) (Atrial natriuretic peptide C-type receptor). | |||||
|
ANPRC_MOUSE
|
||||||
| θ value | θ > 10 (rank : 44) | NC score | 0.154500 (rank : 33) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P70180, P97804, Q9R025, Q9R027, Q9R028 | Gene names | Npr3 | |||
|
Domain Architecture |
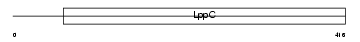
|
|||||
| Description | Atrial natriuretic peptide clearance receptor precursor (ANP-C) (ANPRC) (NPR-C) (Atrial natriuretic peptide C-type receptor) (EF-2). | |||||
|
ADCY8_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 7) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | P97490 | Gene names | Adcy8 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylate cyclase type 8 (EC 4.6.1.1) (Adenylate cyclase type VIII) (ATP pyrophosphate-lyase 8) (Ca(2+)/calmodulin-activated adenylyl cyclase). | |||||
|
ADCY8_HUMAN
|
||||||
| NC score | 0.999786 (rank : 2) | θ value | 0 (rank : 6) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P40145 | Gene names | ADCY8 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylate cyclase type 8 (EC 4.6.1.1) (Adenylate cyclase type VIII) (ATP pyrophosphate-lyase 8) (Ca(2+)/calmodulin-activated adenylyl cyclase). | |||||
|
ADCY6_HUMAN
|
||||||
| NC score | 0.985220 (rank : 3) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O43306, Q9NR75 | Gene names | ADCY6, KIAA0422 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylate cyclase type 6 (EC 4.6.1.1) (Adenylate cyclase type VI) (ATP pyrophosphate-lyase 6) (Ca(2+)-inhibitable adenylyl cyclase). | |||||
|
ADCY6_MOUSE
|
||||||
| NC score | 0.984451 (rank : 4) | θ value | 0 (rank : 5) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q01341 | Gene names | Adcy6 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylate cyclase type 6 (EC 4.6.1.1) (Adenylate cyclase type VI) (ATP pyrophosphate-lyase 6) (Ca(2+)-inhibitable adenylyl cyclase). | |||||
|
ADCY1_MOUSE
|
||||||
| NC score | 0.982307 (rank : 5) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O88444, Q5SS89 | Gene names | Adcy1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Adenylate cyclase type 1 (EC 4.6.1.1) (Adenylate cyclase type I) (ATP pyrophosphate-lyase 1) (Ca(2+)/calmodulin-activated adenylyl cyclase). | |||||
|
ADCY1_HUMAN
|
||||||
| NC score | 0.979784 (rank : 6) | θ value | 2.24492e-184 (rank : 8) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q08828, Q75MI1 | Gene names | ADCY1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Adenylate cyclase type 1 (EC 4.6.1.1) (Adenylate cyclase type I) (ATP pyrophosphate-lyase 1) (Ca(2+)/calmodulin-activated adenylyl cyclase). | |||||
|
ADCY3_HUMAN
|
||||||
| NC score | 0.979628 (rank : 7) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O60266, Q9UDB1 | Gene names | ADCY3, KIAA0511 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylate cyclase type 3 (EC 4.6.1.1) (Adenylate cyclase type III) (Adenylate cyclase, olfactive type) (ATP pyrophosphate-lyase 3) (Adenylyl cyclase 3) (AC-III) (AC3). | |||||
|
ADCY2_MOUSE
|
||||||
| NC score | 0.978469 (rank : 8) | θ value | 4.1007e-178 (rank : 9) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q80TL1, Q8CFU6 | Gene names | Adcy2, Kiaa1060 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Adenylate cyclase type 2 (EC 4.6.1.1) (Adenylate cyclase type II) (ATP pyrophosphate-lyase 2) (Adenylyl cyclase 2). | |||||
|
ADCY3_MOUSE
|
||||||
| NC score | 0.978268 (rank : 9) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8VHH7 | Gene names | Adcy3 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylate cyclase type 3 (EC 4.6.1.1) (Adenylate cyclase type III) (Adenylate cyclase, olfactive type) (ATP pyrophosphate-lyase 3) (Adenylyl cyclase 3) (AC-III) (AC3). | |||||
|
ADCY7_MOUSE
|
||||||
| NC score | 0.977923 (rank : 10) | θ value | 2.18956e-155 (rank : 14) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P51829 | Gene names | Adcy7 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylate cyclase type 7 (EC 4.6.1.1) (Adenylate cyclase type VII) (ATP pyrophosphate-lyase 7) (Adenylyl cyclase 7). | |||||
|
ADCY7_HUMAN
|
||||||
| NC score | 0.976844 (rank : 11) | θ value | 1.84928e-162 (rank : 12) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P51828 | Gene names | ADCY7, KIAA0037 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Adenylate cyclase type 7 (EC 4.6.1.1) (Adenylate cyclase type VII) (ATP pyrophosphate-lyase 7) (Adenylyl cyclase 7). | |||||
|
ADCY2_HUMAN
|
||||||
| NC score | 0.976397 (rank : 12) | θ value | 2.84643e-171 (rank : 10) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q08462, Q9UPU2 | Gene names | ADCY2, KIAA1060 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylate cyclase type 2 (EC 4.6.1.1) (Adenylate cyclase type II) (ATP pyrophosphate-lyase 2) (Adenylyl cyclase 2). | |||||
|
ADCY4_MOUSE
|
||||||
| NC score | 0.975857 (rank : 13) | θ value | 5.94893e-161 (rank : 13) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q91WF3 | Gene names | Adcy4 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylate cyclase type 4 (EC 4.6.1.1) (Adenylate cyclase type IV) (ATP pyrophosphate-lyase 4) (Adenylyl cyclase 4). | |||||
|
ADCY4_HUMAN
|
||||||
| NC score | 0.975192 (rank : 14) | θ value | 1.15832e-164 (rank : 11) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8NFM4, Q96ML7 | Gene names | ADCY4 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylate cyclase type 4 (EC 4.6.1.1) (Adenylate cyclase type IV) (ATP pyrophosphate-lyase 4) (Adenylyl cyclase 4). | |||||
|
ADCY9_MOUSE
|
||||||
| NC score | 0.906568 (rank : 15) | θ value | 2.48917e-58 (rank : 15) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P51830, Q61279 | Gene names | Adcy9 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylate cyclase type 9 (EC 4.6.1.1) (Adenylate cyclase type IX) (ATP pyrophosphate-lyase 9) (Adenylyl cyclase 9) (Adenylyl cyclase type 10) (ACTP10). | |||||
|
ADCY9_HUMAN
|
||||||
| NC score | 0.906395 (rank : 16) | θ value | 4.24587e-58 (rank : 16) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O60503, O60273, Q4ZHT9, Q4ZIR5, Q9BWT4, Q9UGP2 | Gene names | ADCY9, KIAA0520 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylate cyclase type 9 (EC 4.6.1.1) (Adenylate cyclase type IX) (ATP pyrophosphate-lyase 9) (Adenylyl cyclase 9). | |||||
|
GCYB2_HUMAN
|
||||||
| NC score | 0.806589 (rank : 17) | θ value | 4.01107e-24 (rank : 24) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O75343, Q9NZ64 | Gene names | GUCY1B2 | |||
|
Domain Architecture |

|
|||||
| Description | Guanylate cyclase soluble subunit beta-2 (EC 4.6.1.2) (GCS-beta-2). | |||||
|
GCYA2_HUMAN
|
||||||
| NC score | 0.783583 (rank : 18) | θ value | 6.40375e-22 (rank : 29) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P33402 | Gene names | GUCY1A2, GUC1A2, GUCSA2 | |||
|
Domain Architecture |

|
|||||
| Description | Guanylate cyclase soluble subunit alpha-2 (EC 4.6.1.2) (GCS-alpha-2). | |||||
|
GCYA3_HUMAN
|
||||||
| NC score | 0.780755 (rank : 19) | θ value | 4.15078e-21 (rank : 30) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q02108, O43843, Q8TAH3 | Gene names | GUCY1A3, GUC1A3, GUCSA3, GUCY1A1 | |||
|
Domain Architecture |

|
|||||
| Description | Guanylate cyclase soluble subunit alpha-3 (EC 4.6.1.2) (GCS-alpha-3) (Soluble guanylate cyclase large subunit) (GCS-alpha-1). | |||||
|
GCYA3_MOUSE
|
||||||
| NC score | 0.778795 (rank : 20) | θ value | 7.0802e-21 (rank : 31) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9ERL9, Q9DBQ3 | Gene names | Gucy1a3, Gucy1a1 | |||
|
Domain Architecture |

|
|||||
| Description | Guanylate cyclase soluble subunit alpha-3 (EC 4.6.1.2) (GCS-alpha-3) (Soluble guanylate cyclase large subunit) (GCS-alpha-1). | |||||
|
GCYB1_MOUSE
|
||||||
| NC score | 0.778445 (rank : 21) | θ value | 6.84181e-24 (rank : 26) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O54865 | Gene names | Gucy1b3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Guanylate cyclase soluble subunit beta-1 (EC 4.6.1.2) (GCS-beta-1) (Soluble guanylate cyclase small subunit). | |||||
|
GCYB1_HUMAN
|
||||||
| NC score | 0.778257 (rank : 22) | θ value | 4.01107e-24 (rank : 23) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q02153, Q86WY5 | Gene names | GUCY1B3, GUC1B3, GUCSB3, GUCY1B1 | |||
|
Domain Architecture |

|
|||||
| Description | Guanylate cyclase soluble subunit beta-1 (EC 4.6.1.2) (GCS-beta-1) (Soluble guanylate cyclase small subunit) (GCS-beta-3). | |||||
|
GUC2E_MOUSE
|
||||||
| NC score | 0.495708 (rank : 23) | θ value | 1.92365e-26 (rank : 20) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 692 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P52785 | Gene names | Gucy2e, Guc2e | |||
|
Domain Architecture |

|
|||||
| Description | Guanylyl cyclase GC-E precursor (EC 4.6.1.2) (Guanylate cyclase 2E). | |||||
|
GUC2D_HUMAN
|
||||||
| NC score | 0.492111 (rank : 24) | θ value | 5.23862e-24 (rank : 25) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 715 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q02846, Q6LEA7 | Gene names | GUCY2D, CORD6, GUC1A4, GUC2D, RETGC, RETGC1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinal guanylyl cyclase 1 precursor (EC 4.6.1.2) (Guanylate cyclase 2D, retinal) (RETGC-1) (Rod outer segment membrane guanylate cyclase) (ROS-GC). | |||||
|
GUC2G_MOUSE
|
||||||
| NC score | 0.462401 (rank : 25) | θ value | 4.43474e-23 (rank : 27) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 644 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q6TL19, Q8BWU7 | Gene names | Gucy2g | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Guanylate cyclase 2G precursor (EC 4.6.1.2) (Guanylyl cyclase receptor G) (mGC-G). | |||||
|
ANPRB_MOUSE
|
||||||
| NC score | 0.455251 (rank : 26) | θ value | 7.30988e-26 (rank : 21) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 607 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q6VVW5, Q6VVW3, Q6VVW4, Q8CGA9 | Gene names | Npr2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Atrial natriuretic peptide receptor B precursor (ANP-B) (ANPRB) (GC-B) (Guanylate cyclase B) (EC 4.6.1.2) (NPR-B) (Atrial natriuretic peptide B-type receptor). | |||||
|
ANPRA_MOUSE
|
||||||
| NC score | 0.447826 (rank : 27) | θ value | 5.97985e-28 (rank : 18) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 600 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P18293 | Gene names | Npr1, Npra | |||
|
Domain Architecture |

|
|||||
| Description | Atrial natriuretic peptide receptor A precursor (ANP-A) (ANPRA) (GC-A) (Guanylate cyclase) (EC 4.6.1.2) (NPR-A) (Atrial natriuretic peptide A-type receptor). | |||||
|
ANPRA_HUMAN
|
||||||
| NC score | 0.446835 (rank : 28) | θ value | 2.68423e-28 (rank : 17) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 612 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P16066, Q6P4Q3 | Gene names | NPR1, ANPRA | |||
|
Domain Architecture |

|
|||||
| Description | Atrial natriuretic peptide receptor A precursor (ANP-A) (ANPRA) (GC-A) (Guanylate cyclase) (EC 4.6.1.2) (NPR-A) (Atrial natriuretic peptide A-type receptor). | |||||
|
ANPRB_HUMAN
|
||||||
| NC score | 0.442792 (rank : 29) | θ value | 1.92365e-26 (rank : 19) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 646 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P20594, O60871, Q9UQ50 | Gene names | NPR2, ANPRB | |||
|
Domain Architecture |

|
|||||
| Description | Atrial natriuretic peptide receptor B precursor (ANP-B) (ANPRB) (GC-B) (Guanylate cyclase B) (EC 4.6.1.2) (NPR-B) (Atrial natriuretic peptide B-type receptor). | |||||
|
GUC2C_HUMAN
|
||||||
| NC score | 0.433912 (rank : 30) | θ value | 3.75424e-22 (rank : 28) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 624 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P25092 | Gene names | GUCY2C, GUC2C, STAR | |||
|
Domain Architecture |

|
|||||
| Description | Heat-stable enterotoxin receptor precursor (GC-C) (Intestinal guanylate cyclase) (EC 4.6.1.2) (STA receptor) (hSTAR). | |||||
|
GUC2F_HUMAN
|
||||||
| NC score | 0.419375 (rank : 31) | θ value | 1.24688e-25 (rank : 22) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 740 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P51841, Q9UJF1 | Gene names | GUCY2F, GUC2F, RETGC2 | |||
|
Domain Architecture |

|
|||||
| Description | Retinal guanylyl cyclase 2 precursor (EC 4.6.1.2) (Guanylate cyclase 2F, retinal) (RETGC-2) (Rod outer segment membrane guanylate cyclase 2) (ROS-GC2) (Guanylate cyclase F) (GC-F). | |||||
|
ANPRC_HUMAN
|
||||||
| NC score | 0.157747 (rank : 32) | θ value | θ > 10 (rank : 43) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P17342 | Gene names | NPR3, ANPRC | |||
|
Domain Architecture |

|
|||||
| Description | Atrial natriuretic peptide clearance receptor precursor (ANP-C) (ANPRC) (NPR-C) (Atrial natriuretic peptide C-type receptor). | |||||
|
ANPRC_MOUSE
|
||||||
| NC score | 0.154500 (rank : 33) | θ value | θ > 10 (rank : 44) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P70180, P97804, Q9R025, Q9R027, Q9R028 | Gene names | Npr3 | |||
|
Domain Architecture |

|
|||||
| Description | Atrial natriuretic peptide clearance receptor precursor (ANP-C) (ANPRC) (NPR-C) (Atrial natriuretic peptide C-type receptor) (EF-2). | |||||
|
TM16H_MOUSE
|
||||||
| NC score | 0.062397 (rank : 34) | θ value | 0.00869519 (rank : 32) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6PB70, Q69ZE4 | Gene names | Tmem16h, Kiaa1623 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein 16H. | |||||
|
TM16H_HUMAN
|
||||||
| NC score | 0.053465 (rank : 35) | θ value | 0.0563607 (rank : 33) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9HCE9 | Gene names | TMEM16H, KIAA1623 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein 16H. | |||||
|
E2AK4_MOUSE
|
||||||
| NC score | 0.014869 (rank : 36) | θ value | 1.06291 (rank : 35) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 984 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9QZ05, Q6GQT4, Q6ZPT5, Q8C5S0, Q8CIF5, Q9CT30, Q9CUV9, Q9ESB6, Q9ESB7, Q9ESB8 | Gene names | Eif2ak4, Gcn2, Kiaa1338 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 2-alpha kinase 4 (EC 2.7.11.1) (GCN2-like protein) (mGCN2). | |||||
|
LY10_HUMAN
|
||||||
| NC score | 0.014076 (rank : 37) | θ value | 3.0926 (rank : 38) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q13342, Q13341, Q92881, Q96TG3 | Gene names | SP140, LYSP100 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear body protein SP140 (Nuclear autoantigen Sp-140) (Speckled 140 kDa) (LYSp100 protein) (Lymphoid-restricted homolog of Sp100). | |||||
|
PUR2_HUMAN
|
||||||
| NC score | 0.006799 (rank : 38) | θ value | 6.88961 (rank : 41) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P22102 | Gene names | GART, PRGS | |||
|
Domain Architecture |

|
|||||
| Description | Trifunctional purine biosynthetic protein adenosine-3 [Includes: Phosphoribosylamine--glycine ligase (EC 6.3.4.13) (GARS) (Glycinamide ribonucleotide synthetase) (Phosphoribosylglycinamide synthetase); Phosphoribosylformylglycinamidine cyclo-ligase (EC 6.3.3.1) (AIRS) (Phosphoribosyl-aminoimidazole synthetase) (AIR synthase); Phosphoribosylglycinamide formyltransferase (EC 2.1.2.2) (GART) (GAR transformylase) (5'-phosphoribosylglycinamide transformylase)]. | |||||
|
RHBT1_HUMAN
|
||||||
| NC score | 0.006243 (rank : 39) | θ value | 5.27518 (rank : 39) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 262 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O94844 | Gene names | RHOBTB1, KIAA0740 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-related BTB domain-containing protein 1. | |||||
|
COCA1_MOUSE
|
||||||
| NC score | 0.005910 (rank : 40) | θ value | 0.813845 (rank : 34) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q60847, P70322 | Gene names | Col12a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XII) chain precursor. | |||||
|
LMIP_MOUSE
|
||||||
| NC score | 0.005881 (rank : 41) | θ value | 8.99809 (rank : 42) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P56563, Q3TNV8, Q99PA6 | Gene names | Lim2 | |||
|
Domain Architecture |

|
|||||
| Description | Lens fiber membrane intrinsic protein (MP17) (MP18) (MP19) (MP20). | |||||
|
COCA1_HUMAN
|
||||||
| NC score | 0.004877 (rank : 42) | θ value | 3.0926 (rank : 37) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99715, Q99716 | Gene names | COL12A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(XII) chain precursor. | |||||
|
TRHY_HUMAN
|
||||||
| NC score | 0.002503 (rank : 43) | θ value | 1.81305 (rank : 36) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 1385 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q07283 | Gene names | THH, THL, TRHY | |||
|
Domain Architecture |

|
|||||
| Description | Trichohyalin. | |||||
|
PRDM1_MOUSE
|
||||||
| NC score | -0.003348 (rank : 44) | θ value | 6.88961 (rank : 40) | |||
| Query Neighborhood Hits | 42 | Target Neighborhood Hits | 766 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q60636 | Gene names | Prdm1, Blimp1 | |||
|
Domain Architecture |

|
|||||
| Description | PR domain zinc finger protein 1 (PR domain-containing protein 1) (Beta-interferon gene positive-regulatory domain I-binding factor) (BLIMP-1). | |||||