Please be patient as the page loads
|
TDRD3_HUMAN
|
||||||
| SwissProt Accessions | Q9H7E2, Q6P992 | Gene names | TDRD3 | |||
|
Domain Architecture |

|
|||||
| Description | Tudor domain-containing protein 3. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
TDRD3_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | Q9H7E2, Q6P992 | Gene names | TDRD3 | |||
|
Domain Architecture |

|
|||||
| Description | Tudor domain-containing protein 3. | |||||
|
TDRD3_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.980100 (rank : 2) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q91W18, Q8BZW6 | Gene names | Tdrd3 | |||
|
Domain Architecture |

|
|||||
| Description | Tudor domain-containing protein 3. | |||||
|
RMI1_HUMAN
|
||||||
| θ value | 1.06045e-08 (rank : 3) | NC score | 0.357747 (rank : 5) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H9A7, Q5SQG8, Q5SQG9, Q6P1Q4, Q6PI89, Q7Z6L6 | Gene names | RMI1, C9orf76 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein RMI1 homolog. | |||||
|
SPF30_HUMAN
|
||||||
| θ value | 1.99992e-07 (rank : 4) | NC score | 0.409881 (rank : 3) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O75940 | Gene names | SMNDC1, SMNR, SPF30 | |||
|
Domain Architecture |
|
|||||
| Description | Survival of motor neuron-related-splicing factor 30 (SMN-related protein) (30 kDa splicing factor SMNrp) (Survival motor neuron domain- containing protein 1). | |||||
|
SPF30_MOUSE
|
||||||
| θ value | 3.41135e-07 (rank : 5) | NC score | 0.406057 (rank : 4) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8BGT7, Q4QQL0 | Gene names | Smndc1, Smnr, Spf30 | |||
|
Domain Architecture |
|
|||||
| Description | Survival of motor neuron-related-splicing factor 30 (SMN-related protein) (30 kDa splicing factor SMNrp) (Survival motor neuron domain- containing protein 1). | |||||
|
RMI1_MOUSE
|
||||||
| θ value | 5.81887e-07 (rank : 6) | NC score | 0.341262 (rank : 6) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9D4G9, Q8C560, Q8CI20 | Gene names | Rmi1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein RMI1 homolog. | |||||
|
SMN_HUMAN
|
||||||
| θ value | 0.000461057 (rank : 7) | NC score | 0.313007 (rank : 7) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q16637, Q13119, Q96J51 | Gene names | SMN1, SMN, SMNT | |||
|
Domain Architecture |
|
|||||
| Description | Survival motor neuron protein (Component of gems 1) (Gemin-1). | |||||
|
SMN_MOUSE
|
||||||
| θ value | 0.000461057 (rank : 8) | NC score | 0.308621 (rank : 8) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P97801, O09092 | Gene names | Smn1, Smn | |||
|
Domain Architecture |
|
|||||
| Description | Survival motor neuron protein. | |||||
|
TDRD1_MOUSE
|
||||||
| θ value | 0.00134147 (rank : 9) | NC score | 0.192023 (rank : 9) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q99MV1 | Gene names | Tdrd1 | |||
|
Domain Architecture |

|
|||||
| Description | Tudor domain-containing protein 1. | |||||
|
TDRD1_HUMAN
|
||||||
| θ value | 0.00228821 (rank : 10) | NC score | 0.183457 (rank : 10) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9BXT4, Q9H7B3 | Gene names | TDRD1 | |||
|
Domain Architecture |

|
|||||
| Description | Tudor domain-containing protein 1. | |||||
|
NIPBL_HUMAN
|
||||||
| θ value | 0.00298849 (rank : 11) | NC score | 0.099272 (rank : 14) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
FOXGA_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 12) | NC score | 0.030501 (rank : 49) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P55316 | Gene names | FOXG1A, FKHL2 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein G1A (Forkhead-related protein FKHL2) (Transcription factor BF-2) (Brain factor 2) (BF2) (HFK2). | |||||
|
TDRD6_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 13) | NC score | 0.151390 (rank : 11) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P61407 | Gene names | Tdrd6 | |||
|
Domain Architecture |

|
|||||
| Description | Tudor domain-containing protein 6. | |||||
|
CHD1_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 14) | NC score | 0.044798 (rank : 33) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P40201 | Gene names | Chd1, Chd-1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
NPBL_MOUSE
|
||||||
| θ value | 0.125558 (rank : 15) | NC score | 0.096931 (rank : 15) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
UBDC1_MOUSE
|
||||||
| θ value | 0.125558 (rank : 16) | NC score | 0.127095 (rank : 12) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8VDI7, Q3TZH3 | Gene names | Ubadc1, Kpc2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-associated domain-containing protein 1 (E3 ubiquitin-protein ligase subunit KPC2) (Kip1 ubiquitination-promoting complex protein 2). | |||||
|
CHD1_HUMAN
|
||||||
| θ value | 0.163984 (rank : 17) | NC score | 0.045273 (rank : 31) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O14646 | Gene names | CHD1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
GNAS3_HUMAN
|
||||||
| θ value | 0.21417 (rank : 18) | NC score | 0.107727 (rank : 13) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O95467, O95417 | Gene names | GNAS, GNAS1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuroendocrine secretory protein 55 (NESP55) [Contains: LHAL tetrapeptide; GPIPIRRH peptide]. | |||||
|
PPIG_HUMAN
|
||||||
| θ value | 0.21417 (rank : 19) | NC score | 0.048275 (rank : 28) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q13427, O00706, Q96DG9 | Gene names | PPIG | |||
|
Domain Architecture |

|
|||||
| Description | Peptidyl-prolyl cis-trans isomerase G (EC 5.2.1.8) (Peptidyl-prolyl isomerase G) (PPIase G) (Rotamase G) (Cyclophilin G) (Clk-associating RS-cyclophilin) (CARS-cyclophilin) (CARS-Cyp) (SR-cyclophilin) (SRcyp) (SR-cyp) (CASP10). | |||||
|
TOPRS_HUMAN
|
||||||
| θ value | 0.21417 (rank : 20) | NC score | 0.056388 (rank : 23) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9NS56, O43273, Q6P987, Q9NS55, Q9UNR9 | Gene names | TOPORS, LUN, TP53BPL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein E3 ligase Topors (EC 6.3.2.-) (SUMO1-protein E3 ligase Topors) (Topoisomerase I-binding RING finger protein) (Topoisomerase I-binding arginine/serine-rich protein) (Tumor suppressor p53-binding protein 3) (p53-binding protein 3) (p53BP3). | |||||
|
TXND2_MOUSE
|
||||||
| θ value | 0.21417 (rank : 21) | NC score | 0.058487 (rank : 21) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
FILA_HUMAN
|
||||||
| θ value | 0.365318 (rank : 22) | NC score | 0.062329 (rank : 18) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 685 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P20930, Q01720, Q5T583 | Gene names | FLG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filaggrin. | |||||
|
RSRC1_MOUSE
|
||||||
| θ value | 0.365318 (rank : 23) | NC score | 0.081270 (rank : 16) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9DBU6, Q3TF06 | Gene names | Rsrc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine/serine-rich coiled coil protein 1. | |||||
|
ANR11_HUMAN
|
||||||
| θ value | 0.47712 (rank : 24) | NC score | 0.028685 (rank : 53) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1289 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q6UB99, Q6NTG1, Q6QMF8 | Gene names | ANKRD11, ANCO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 11 (Ankyrin repeat-containing cofactor 1). | |||||
|
WDR46_MOUSE
|
||||||
| θ value | 0.47712 (rank : 25) | NC score | 0.044984 (rank : 32) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Z0H1 | Gene names | Wdr46, Bing4 | |||
|
Domain Architecture |
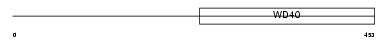
|
|||||
| Description | WD repeat protein 46 (WD repeat protein BING4). | |||||
|
DDX23_HUMAN
|
||||||
| θ value | 1.06291 (rank : 26) | NC score | 0.018696 (rank : 62) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 486 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9BUQ8, O43188 | Gene names | DDX23 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX23 (EC 3.6.1.-) (DEAD box protein 23) (100 kDa U5 snRNP-specific protein) (U5-100kD) (PRP28 homolog). | |||||
|
DMP1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 27) | NC score | 0.057157 (rank : 22) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O55188 | Gene names | Dmp1, Dmp | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1) (AG1). | |||||
|
NUB1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 28) | NC score | 0.060081 (rank : 20) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P54729, Q8K3U0 | Gene names | Nub1, Nyren18 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD8 ultimate buster 1 (Protein BS4). | |||||
|
SRCH_HUMAN
|
||||||
| θ value | 1.06291 (rank : 29) | NC score | 0.062318 (rank : 19) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P23327, Q504Y6 | Gene names | HRC, HCP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcoplasmic reticulum histidine-rich calcium-binding protein precursor. | |||||
|
RPA1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 30) | NC score | 0.026823 (rank : 55) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O35134, Q7TSA9 | Gene names | Polr1a, Rpa1, Rpo1-4 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-directed RNA polymerase I largest subunit (EC 2.7.7.6) (RNA polymerase I 194 kDa subunit) (RPA194). | |||||
|
UBP5_HUMAN
|
||||||
| θ value | 1.38821 (rank : 31) | NC score | 0.049454 (rank : 27) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P45974, Q96J22 | Gene names | USP5, ISOT | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 5 (EC 3.1.2.15) (Ubiquitin thioesterase 5) (Ubiquitin-specific-processing protease 5) (Deubiquitinating enzyme 5) (Isopeptidase T). | |||||
|
UBP5_MOUSE
|
||||||
| θ value | 1.38821 (rank : 32) | NC score | 0.049473 (rank : 26) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P56399 | Gene names | Usp5, Isot | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 5 (EC 3.1.2.15) (Ubiquitin thioesterase 5) (Ubiquitin-specific-processing protease 5) (Deubiquitinating enzyme 5) (Isopeptidase T). | |||||
|
GSLG1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 33) | NC score | 0.030142 (rank : 50) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q61543, Q9QZ40 | Gene names | Glg1, Esl1, Mg160, Selel | |||
|
Domain Architecture |
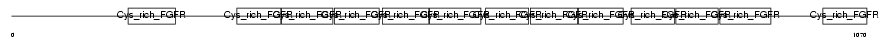
|
|||||
| Description | Golgi apparatus protein 1 precursor (Golgi sialoglycoprotein MG-160) (E-selectin ligand 1) (ESL-1) (Selel). | |||||
|
UBDC1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 34) | NC score | 0.069321 (rank : 17) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9BSL1, O75500, Q9UMW7 | Gene names | UBADC1, GBDR1, KPC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-associated domain-containing protein 1 (E3 ubiquitin-protein ligase subunit KPC2) (Kip1 ubiquitination-promoting complex protein 2) (Glialblastoma cell differentiation-related protein 1). | |||||
|
UBP13_HUMAN
|
||||||
| θ value | 1.81305 (rank : 35) | NC score | 0.050438 (rank : 25) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q92995 | Gene names | USP13, ISOT3 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 13 (EC 3.1.2.15) (Ubiquitin thioesterase 13) (Ubiquitin-specific-processing protease 13) (Deubiquitinating enzyme 13) (Isopeptidase T-3) (ISOT-3). | |||||
|
RSF1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 36) | NC score | 0.036350 (rank : 41) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 539 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q96T23, Q86X86, Q9H3L8, Q9NVZ8, Q9NYU0 | Gene names | RSF1, HBXAP, XAP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Remodeling and spacing factor 1 (Rsf-1) (Hepatitis B virus X- associated protein) (HBV pX-associated protein 8) (p325 subunit of RSF chromatin remodelling complex). | |||||
|
SFRS4_MOUSE
|
||||||
| θ value | 2.36792 (rank : 37) | NC score | 0.041976 (rank : 36) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8VE97, Q9JJC3 | Gene names | Sfrs4 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 4. | |||||
|
BRD4_HUMAN
|
||||||
| θ value | 3.0926 (rank : 38) | NC score | 0.019931 (rank : 59) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 851 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O60885, O60433, Q96PD3 | Gene names | BRD4, HUNK1 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 4 (HUNK1 protein). | |||||
|
LC7L2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 39) | NC score | 0.033033 (rank : 45) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 244 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q7TNC4, Q99LM5, Q99PC3 | Gene names | Luc7l2 | |||
|
Domain Architecture |
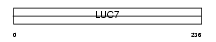
|
|||||
| Description | Putative RNA-binding protein Luc7-like 2 (CGI-74 homolog). | |||||
|
TOP2A_MOUSE
|
||||||
| θ value | 3.0926 (rank : 40) | NC score | 0.019905 (rank : 60) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q01320 | Gene names | Top2a, Top-2, Top2 | |||
|
Domain Architecture |

|
|||||
| Description | DNA topoisomerase 2-alpha (EC 5.99.1.3) (DNA topoisomerase II, alpha isozyme). | |||||
|
UBP53_MOUSE
|
||||||
| θ value | 3.0926 (rank : 41) | NC score | 0.029731 (rank : 51) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P15975, Q8CB37, Q8R251 | Gene names | Usp53, Phxr3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inactive ubiquitin carboxyl-terminal hydrolase 53 (Inactive ubiquitin- specific peptidase 53) (Per-hexamer repeat protein 3). | |||||
|
CABIN_HUMAN
|
||||||
| θ value | 4.03905 (rank : 42) | NC score | 0.024689 (rank : 57) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 490 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Y6J0, Q9Y460 | Gene names | CABIN1, KIAA0330 | |||
|
Domain Architecture |

|
|||||
| Description | Calcineurin-binding protein Cabin 1 (Calcineurin inhibitor) (CAIN). | |||||
|
CAC1B_MOUSE
|
||||||
| θ value | 4.03905 (rank : 43) | NC score | 0.010457 (rank : 73) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O55017, Q60609 | Gene names | Cacna1b, Cach5, Cacnl1a5, Cchn1a | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent N-type calcium channel subunit alpha-1B (Voltage- gated calcium channel subunit alpha Cav2.2) (Calcium channel, L type, alpha-1 polypeptide isoform 5) (Brain calcium channel III) (BIII). | |||||
|
IF3A_HUMAN
|
||||||
| θ value | 4.03905 (rank : 44) | NC score | 0.015938 (rank : 66) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 875 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q14152, O00653 | Gene names | EIF3S10, KIAA0139 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 3 subunit 10 (eIF-3 theta) (eIF3 p167) (eIF3 p180) (eIF3 p185) (eIF3a). | |||||
|
NUB1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 45) | NC score | 0.043015 (rank : 34) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y5A7, O95422, Q8IX22, Q9BXR2 | Gene names | NUB1, NYREN18 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD8 ultimate buster 1 (NY-REN-18 antigen). | |||||
|
SPRL1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 46) | NC score | 0.031142 (rank : 48) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q14515, Q14800 | Gene names | SPARCL1 | |||
|
Domain Architecture |

|
|||||
| Description | SPARC-like protein 1 precursor (High endothelial venule protein) (Hevin) (MAST 9). | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 47) | NC score | 0.033650 (rank : 44) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
TTLL7_HUMAN
|
||||||
| θ value | 4.03905 (rank : 48) | NC score | 0.018110 (rank : 63) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6ZT98, Q5TAX8, Q5TAX9, Q6P990, Q86YS1, Q9H5U4 | Gene names | TTLL7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tubulin--tyrosine ligase-like protein 7 (Protein NYD-SP30). | |||||
|
UBQL4_MOUSE
|
||||||
| θ value | 4.03905 (rank : 49) | NC score | 0.014315 (rank : 69) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99NB8, Q8BP88 | Gene names | Ubqln4, Ubin | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquilin-4 (Ataxin-1 ubiquitin-like-interacting protein A1U). | |||||
|
AKAP1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 50) | NC score | 0.046905 (rank : 29) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q92667, Q13320, Q9BW14 | Gene names | AKAP1, AKAP149 | |||
|
Domain Architecture |

|
|||||
| Description | A kinase anchor protein 1, mitochondrial precursor (Protein kinase A- anchoring protein 1) (PRKA1) (A-kinase anchor protein 149 kDa) (AKAP 149) (Dual specificity A-kinase-anchoring protein 1) (D-AKAP-1) (Spermatid A-kinase anchor protein 84) (S-AKAP84). | |||||
|
AKAP1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 51) | NC score | 0.045453 (rank : 30) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O08715, O08714, P97488 | Gene names | Akap1, Akap | |||
|
Domain Architecture |

|
|||||
| Description | A kinase anchor protein 1, mitochondrial precursor (Protein kinase A- anchoring protein 1) (PRKA1) (Dual specificity A-kinase-anchoring protein 1) (D-AKAP-1) (Spermatid A-kinase anchor protein) (S-AKAP). | |||||
|
ELOA1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 52) | NC score | 0.034548 (rank : 43) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q14241, Q8IXH1 | Gene names | TCEB3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription elongation factor B polypeptide 3 (RNA polymerase II transcription factor SIII subunit A1) (SIII p110) (Elongin-A) (EloA) (Elongin 110 kDa subunit). | |||||
|
FRM4A_MOUSE
|
||||||
| θ value | 5.27518 (rank : 53) | NC score | 0.011143 (rank : 72) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BIE6, Q6PDJ4, Q8CHA6 | Gene names | Frmd4a, Frmd4, Kiaa1294 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FERM domain-containing protein 4A. | |||||
|
SFRS4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 54) | NC score | 0.036972 (rank : 40) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q08170, Q5VXP1, Q9BUA4, Q9UEB5 | Gene names | SFRS4, SRP75 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 4 (Pre-mRNA-splicing factor SRP75) (SRP001LB). | |||||
|
TCPZ_HUMAN
|
||||||
| θ value | 5.27518 (rank : 55) | NC score | 0.014447 (rank : 67) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P40227, Q75LP4, Q96S46 | Gene names | CCT6A, CCT6, CCTZ | |||
|
Domain Architecture |
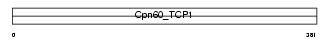
|
|||||
| Description | T-complex protein 1 subunit zeta (TCP-1-zeta) (CCT-zeta) (CCT-zeta-1) (Tcp20) (HTR3) (Acute morphine dependence-related protein 2). | |||||
|
TCPZ_MOUSE
|
||||||
| θ value | 5.27518 (rank : 56) | NC score | 0.014404 (rank : 68) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P80317 | Gene names | Cct6a, Cct6, Cctz, Cctz1 | |||
|
Domain Architecture |

|
|||||
| Description | T-complex protein 1 subunit zeta (TCP-1-zeta) (CCT-zeta) (CCT-zeta-1). | |||||
|
XPC_MOUSE
|
||||||
| θ value | 5.27518 (rank : 57) | NC score | 0.023662 (rank : 58) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P51612, P54732, Q3TKI2, Q920M1, Q9DBW7 | Gene names | Xpc | |||
|
Domain Architecture |

|
|||||
| Description | DNA-repair protein complementing XP-C cells homolog (Xeroderma pigmentosum group C-complementing protein homolog) (p125). | |||||
|
AFF4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 58) | NC score | 0.038614 (rank : 39) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 611 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9UHB7, Q498B2, Q59FB3, Q6P592, Q8TDR1, Q9P0E4 | Gene names | AFF4, AF5Q31, MCEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4 (ALL1-fused gene from chromosome 5q31) (Major CDK9 elongation factor-associated protein). | |||||
|
CHD2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 59) | NC score | 0.035214 (rank : 42) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O14647 | Gene names | CHD2 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 2 (EC 3.6.1.-) (ATP- dependent helicase CHD2) (CHD-2). | |||||
|
DCP1B_HUMAN
|
||||||
| θ value | 6.88961 (rank : 60) | NC score | 0.018779 (rank : 61) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8IZD4, Q86XH9, Q96BP8, Q96MZ8 | Gene names | DCP1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | mRNA decapping enzyme 1B (EC 3.-.-.-). | |||||
|
IWS1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 61) | NC score | 0.039224 (rank : 37) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
K1210_HUMAN
|
||||||
| θ value | 6.88961 (rank : 62) | NC score | 0.028278 (rank : 54) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9ULL0, Q5JPN4 | Gene names | KIAA1210 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1210. | |||||
|
SFR11_HUMAN
|
||||||
| θ value | 6.88961 (rank : 63) | NC score | 0.053111 (rank : 24) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q05519 | Gene names | SFRS11 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor arginine/serine-rich 11 (Arginine-rich 54 kDa nuclear protein) (p54). | |||||
|
SON_HUMAN
|
||||||
| θ value | 6.88961 (rank : 64) | NC score | 0.016569 (rank : 64) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
SRRM1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 65) | NC score | 0.029170 (rank : 52) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
SYNPO_MOUSE
|
||||||
| θ value | 6.88961 (rank : 66) | NC score | 0.016035 (rank : 65) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8CC35, Q99JI0 | Gene names | Synpo, Kiaa1029 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptopodin. | |||||
|
TRDN_HUMAN
|
||||||
| θ value | 6.88961 (rank : 67) | NC score | 0.031507 (rank : 46) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 542 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q13061 | Gene names | TRDN | |||
|
Domain Architecture |

|
|||||
| Description | Triadin. | |||||
|
WWC1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 68) | NC score | 0.011369 (rank : 71) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5SXA9, Q571D0, Q8K1Y3, Q8VD17, Q922W3 | Gene names | Wwc1, Kiaa0869 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing protein 1 (Kidney and brain protein) (KIBRA). | |||||
|
AFF4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 69) | NC score | 0.038970 (rank : 38) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9ESC8, Q8C6K4, Q8C6W3, Q8CCH3 | Gene names | Aff4, Alf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4. | |||||
|
BCAR3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 70) | NC score | 0.008306 (rank : 74) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9QZK2, Q3TNC9, Q3UP10 | Gene names | Bcar3, And34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Breast cancer anti-estrogen resistance protein 3 (p130Cas-binding protein AND-34). | |||||
|
CCNL1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 71) | NC score | 0.025498 (rank : 56) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9UK58, Q6NVY9, Q6UWS7, Q8NI48, Q96QT0, Q9NZF3 | Gene names | CCNL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-L1 (Cyclin-L). | |||||
|
DEK_MOUSE
|
||||||
| θ value | 8.99809 (rank : 72) | NC score | 0.031353 (rank : 47) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q7TNV0, Q80VC5, Q8BZV6 | Gene names | Dek | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein DEK. | |||||
|
MMP1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 73) | NC score | 0.003900 (rank : 75) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P03956, P08156 | Gene names | MMP1, CLG | |||
|
Domain Architecture |

|
|||||
| Description | Interstitial collagenase precursor (EC 3.4.24.7) (Matrix metalloproteinase-1) (MMP-1) (Fibroblast collagenase) [Contains: 22 kDa interstitial collagenase; 27 kDa interstitial collagenase]. | |||||
|
MYOZ2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 74) | NC score | 0.013604 (rank : 70) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NPC6, O43415, Q9HB92 | Gene names | MYOZ2, C4orf5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myozenin-2 (Calsarcin-1) (FATZ-related protein 2). | |||||
|
RBBP6_HUMAN
|
||||||
| θ value | 8.99809 (rank : 75) | NC score | 0.042701 (rank : 35) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q7Z6E9, Q15290, Q6DKH4, Q6P4C2, Q6YNC9, Q7Z6E8, Q8N0V2, Q96PH3, Q9H3I8, Q9H5M5, Q9NPX4 | Gene names | RBBP6, P2PR, PACT, RBQ1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R) (Retinoblastoma-binding Q protein 1) (Protein RBQ-1). | |||||
|
TDRD3_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | Q9H7E2, Q6P992 | Gene names | TDRD3 | |||
|
Domain Architecture |

|
|||||
| Description | Tudor domain-containing protein 3. | |||||
|
TDRD3_MOUSE
|
||||||
| NC score | 0.980100 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q91W18, Q8BZW6 | Gene names | Tdrd3 | |||
|
Domain Architecture |

|
|||||
| Description | Tudor domain-containing protein 3. | |||||
|
SPF30_HUMAN
|
||||||
| NC score | 0.409881 (rank : 3) | θ value | 1.99992e-07 (rank : 4) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O75940 | Gene names | SMNDC1, SMNR, SPF30 | |||
|
Domain Architecture |

|
|||||
| Description | Survival of motor neuron-related-splicing factor 30 (SMN-related protein) (30 kDa splicing factor SMNrp) (Survival motor neuron domain- containing protein 1). | |||||
|
SPF30_MOUSE
|
||||||
| NC score | 0.406057 (rank : 4) | θ value | 3.41135e-07 (rank : 5) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8BGT7, Q4QQL0 | Gene names | Smndc1, Smnr, Spf30 | |||
|
Domain Architecture |

|
|||||
| Description | Survival of motor neuron-related-splicing factor 30 (SMN-related protein) (30 kDa splicing factor SMNrp) (Survival motor neuron domain- containing protein 1). | |||||
|
RMI1_HUMAN
|
||||||
| NC score | 0.357747 (rank : 5) | θ value | 1.06045e-08 (rank : 3) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H9A7, Q5SQG8, Q5SQG9, Q6P1Q4, Q6PI89, Q7Z6L6 | Gene names | RMI1, C9orf76 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein RMI1 homolog. | |||||
|
RMI1_MOUSE
|
||||||
| NC score | 0.341262 (rank : 6) | θ value | 5.81887e-07 (rank : 6) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9D4G9, Q8C560, Q8CI20 | Gene names | Rmi1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein RMI1 homolog. | |||||
|
SMN_HUMAN
|
||||||
| NC score | 0.313007 (rank : 7) | θ value | 0.000461057 (rank : 7) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q16637, Q13119, Q96J51 | Gene names | SMN1, SMN, SMNT | |||
|
Domain Architecture |

|
|||||
| Description | Survival motor neuron protein (Component of gems 1) (Gemin-1). | |||||
|
SMN_MOUSE
|
||||||
| NC score | 0.308621 (rank : 8) | θ value | 0.000461057 (rank : 8) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P97801, O09092 | Gene names | Smn1, Smn | |||
|
Domain Architecture |

|
|||||
| Description | Survival motor neuron protein. | |||||
|
TDRD1_MOUSE
|
||||||
| NC score | 0.192023 (rank : 9) | θ value | 0.00134147 (rank : 9) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q99MV1 | Gene names | Tdrd1 | |||
|
Domain Architecture |

|
|||||
| Description | Tudor domain-containing protein 1. | |||||
|
TDRD1_HUMAN
|
||||||
| NC score | 0.183457 (rank : 10) | θ value | 0.00228821 (rank : 10) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9BXT4, Q9H7B3 | Gene names | TDRD1 | |||
|
Domain Architecture |

|
|||||
| Description | Tudor domain-containing protein 1. | |||||
|
TDRD6_MOUSE
|
||||||
| NC score | 0.151390 (rank : 11) | θ value | 0.0252991 (rank : 13) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P61407 | Gene names | Tdrd6 | |||
|
Domain Architecture |

|
|||||
| Description | Tudor domain-containing protein 6. | |||||
|
UBDC1_MOUSE
|
||||||
| NC score | 0.127095 (rank : 12) | θ value | 0.125558 (rank : 16) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8VDI7, Q3TZH3 | Gene names | Ubadc1, Kpc2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-associated domain-containing protein 1 (E3 ubiquitin-protein ligase subunit KPC2) (Kip1 ubiquitination-promoting complex protein 2). | |||||
|
GNAS3_HUMAN
|
||||||
| NC score | 0.107727 (rank : 13) | θ value | 0.21417 (rank : 18) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 71 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O95467, O95417 | Gene names | GNAS, GNAS1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuroendocrine secretory protein 55 (NESP55) [Contains: LHAL tetrapeptide; GPIPIRRH peptide]. | |||||
|
NIPBL_HUMAN
|
||||||
| NC score | 0.099272 (rank : 14) | θ value | 0.00298849 (rank : 11) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
NPBL_MOUSE
|
||||||
| NC score | 0.096931 (rank : 15) | θ value | 0.125558 (rank : 15) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
RSRC1_MOUSE
|
||||||
| NC score | 0.081270 (rank : 16) | θ value | 0.365318 (rank : 23) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9DBU6, Q3TF06 | Gene names | Rsrc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine/serine-rich coiled coil protein 1. | |||||
|
UBDC1_HUMAN
|
||||||
| NC score | 0.069321 (rank : 17) | θ value | 1.81305 (rank : 34) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9BSL1, O75500, Q9UMW7 | Gene names | UBADC1, GBDR1, KPC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-associated domain-containing protein 1 (E3 ubiquitin-protein ligase subunit KPC2) (Kip1 ubiquitination-promoting complex protein 2) (Glialblastoma cell differentiation-related protein 1). | |||||
|
FILA_HUMAN
|
||||||
| NC score | 0.062329 (rank : 18) | θ value | 0.365318 (rank : 22) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 685 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P20930, Q01720, Q5T583 | Gene names | FLG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filaggrin. | |||||
|
SRCH_HUMAN
|
||||||
| NC score | 0.062318 (rank : 19) | θ value | 1.06291 (rank : 29) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P23327, Q504Y6 | Gene names | HRC, HCP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcoplasmic reticulum histidine-rich calcium-binding protein precursor. | |||||
|
NUB1_MOUSE
|
||||||
| NC score | 0.060081 (rank : 20) | θ value | 1.06291 (rank : 28) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P54729, Q8K3U0 | Gene names | Nub1, Nyren18 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD8 ultimate buster 1 (Protein BS4). | |||||
|
TXND2_MOUSE
|
||||||
| NC score | 0.058487 (rank : 21) | θ value | 0.21417 (rank : 21) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
DMP1_MOUSE
|
||||||
| NC score | 0.057157 (rank : 22) | θ value | 1.06291 (rank : 27) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O55188 | Gene names | Dmp1, Dmp | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1) (AG1). | |||||
|
TOPRS_HUMAN
|
||||||
| NC score | 0.056388 (rank : 23) | θ value | 0.21417 (rank : 20) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9NS56, O43273, Q6P987, Q9NS55, Q9UNR9 | Gene names | TOPORS, LUN, TP53BPL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein E3 ligase Topors (EC 6.3.2.-) (SUMO1-protein E3 ligase Topors) (Topoisomerase I-binding RING finger protein) (Topoisomerase I-binding arginine/serine-rich protein) (Tumor suppressor p53-binding protein 3) (p53-binding protein 3) (p53BP3). | |||||
|
SFR11_HUMAN
|
||||||
| NC score | 0.053111 (rank : 24) | θ value | 6.88961 (rank : 63) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q05519 | Gene names | SFRS11 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor arginine/serine-rich 11 (Arginine-rich 54 kDa nuclear protein) (p54). | |||||
|
UBP13_HUMAN
|
||||||
| NC score | 0.050438 (rank : 25) | θ value | 1.81305 (rank : 35) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q92995 | Gene names | USP13, ISOT3 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 13 (EC 3.1.2.15) (Ubiquitin thioesterase 13) (Ubiquitin-specific-processing protease 13) (Deubiquitinating enzyme 13) (Isopeptidase T-3) (ISOT-3). | |||||
|
UBP5_MOUSE
|
||||||
| NC score | 0.049473 (rank : 26) | θ value | 1.38821 (rank : 32) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P56399 | Gene names | Usp5, Isot | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 5 (EC 3.1.2.15) (Ubiquitin thioesterase 5) (Ubiquitin-specific-processing protease 5) (Deubiquitinating enzyme 5) (Isopeptidase T). | |||||
|
UBP5_HUMAN
|
||||||
| NC score | 0.049454 (rank : 27) | θ value | 1.38821 (rank : 31) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P45974, Q96J22 | Gene names | USP5, ISOT | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 5 (EC 3.1.2.15) (Ubiquitin thioesterase 5) (Ubiquitin-specific-processing protease 5) (Deubiquitinating enzyme 5) (Isopeptidase T). | |||||
|
PPIG_HUMAN
|
||||||
| NC score | 0.048275 (rank : 28) | θ value | 0.21417 (rank : 19) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q13427, O00706, Q96DG9 | Gene names | PPIG | |||
|
Domain Architecture |

|
|||||
| Description | Peptidyl-prolyl cis-trans isomerase G (EC 5.2.1.8) (Peptidyl-prolyl isomerase G) (PPIase G) (Rotamase G) (Cyclophilin G) (Clk-associating RS-cyclophilin) (CARS-cyclophilin) (CARS-Cyp) (SR-cyclophilin) (SRcyp) (SR-cyp) (CASP10). | |||||
|
AKAP1_HUMAN
|
||||||
| NC score | 0.046905 (rank : 29) | θ value | 5.27518 (rank : 50) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q92667, Q13320, Q9BW14 | Gene names | AKAP1, AKAP149 | |||
|
Domain Architecture |

|
|||||
| Description | A kinase anchor protein 1, mitochondrial precursor (Protein kinase A- anchoring protein 1) (PRKA1) (A-kinase anchor protein 149 kDa) (AKAP 149) (Dual specificity A-kinase-anchoring protein 1) (D-AKAP-1) (Spermatid A-kinase anchor protein 84) (S-AKAP84). | |||||
|
AKAP1_MOUSE
|
||||||
| NC score | 0.045453 (rank : 30) | θ value | 5.27518 (rank : 51) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 198 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O08715, O08714, P97488 | Gene names | Akap1, Akap | |||
|
Domain Architecture |

|
|||||
| Description | A kinase anchor protein 1, mitochondrial precursor (Protein kinase A- anchoring protein 1) (PRKA1) (Dual specificity A-kinase-anchoring protein 1) (D-AKAP-1) (Spermatid A-kinase anchor protein) (S-AKAP). | |||||
|
CHD1_HUMAN
|
||||||
| NC score | 0.045273 (rank : 31) | θ value | 0.163984 (rank : 17) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O14646 | Gene names | CHD1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
WDR46_MOUSE
|
||||||
| NC score | 0.044984 (rank : 32) | θ value | 0.47712 (rank : 25) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Z0H1 | Gene names | Wdr46, Bing4 | |||
|
Domain Architecture |

|
|||||
| Description | WD repeat protein 46 (WD repeat protein BING4). | |||||
|
CHD1_MOUSE
|
||||||
| NC score | 0.044798 (rank : 33) | θ value | 0.0961366 (rank : 14) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P40201 | Gene names | Chd1, Chd-1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
NUB1_HUMAN
|
||||||
| NC score | 0.043015 (rank : 34) | θ value | 4.03905 (rank : 45) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y5A7, O95422, Q8IX22, Q9BXR2 | Gene names | NUB1, NYREN18 | |||
|
Domain Architecture |

|
|||||
| Description | NEDD8 ultimate buster 1 (NY-REN-18 antigen). | |||||
|
RBBP6_HUMAN
|
||||||
| NC score | 0.042701 (rank : 35) | θ value | 8.99809 (rank : 75) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q7Z6E9, Q15290, Q6DKH4, Q6P4C2, Q6YNC9, Q7Z6E8, Q8N0V2, Q96PH3, Q9H3I8, Q9H5M5, Q9NPX4 | Gene names | RBBP6, P2PR, PACT, RBQ1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R) (Retinoblastoma-binding Q protein 1) (Protein RBQ-1). | |||||
|
SFRS4_MOUSE
|
||||||
| NC score | 0.041976 (rank : 36) | θ value | 2.36792 (rank : 37) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8VE97, Q9JJC3 | Gene names | Sfrs4 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 4. | |||||
|
IWS1_HUMAN
|
||||||
| NC score | 0.039224 (rank : 37) | θ value | 6.88961 (rank : 61) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
AFF4_MOUSE
|
||||||
| NC score | 0.038970 (rank : 38) | θ value | 8.99809 (rank : 69) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9ESC8, Q8C6K4, Q8C6W3, Q8CCH3 | Gene names | Aff4, Alf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4. | |||||
|
AFF4_HUMAN
|
||||||
| NC score | 0.038614 (rank : 39) | θ value | 6.88961 (rank : 58) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 611 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q9UHB7, Q498B2, Q59FB3, Q6P592, Q8TDR1, Q9P0E4 | Gene names | AFF4, AF5Q31, MCEF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4 (ALL1-fused gene from chromosome 5q31) (Major CDK9 elongation factor-associated protein). | |||||
|
SFRS4_HUMAN
|
||||||
| NC score | 0.036972 (rank : 40) | θ value | 5.27518 (rank : 54) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q08170, Q5VXP1, Q9BUA4, Q9UEB5 | Gene names | SFRS4, SRP75 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 4 (Pre-mRNA-splicing factor SRP75) (SRP001LB). | |||||
|
RSF1_HUMAN
|
||||||
| NC score | 0.036350 (rank : 41) | θ value | 2.36792 (rank : 36) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 539 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q96T23, Q86X86, Q9H3L8, Q9NVZ8, Q9NYU0 | Gene names | RSF1, HBXAP, XAP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Remodeling and spacing factor 1 (Rsf-1) (Hepatitis B virus X- associated protein) (HBV pX-associated protein 8) (p325 subunit of RSF chromatin remodelling complex). | |||||
|
CHD2_HUMAN
|
||||||
| NC score | 0.035214 (rank : 42) | θ value | 6.88961 (rank : 59) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 360 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O14647 | Gene names | CHD2 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 2 (EC 3.6.1.-) (ATP- dependent helicase CHD2) (CHD-2). | |||||
|
ELOA1_HUMAN
|
||||||
| NC score | 0.034548 (rank : 43) | θ value | 5.27518 (rank : 52) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q14241, Q8IXH1 | Gene names | TCEB3 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription elongation factor B polypeptide 3 (RNA polymerase II transcription factor SIII subunit A1) (SIII p110) (Elongin-A) (EloA) (Elongin 110 kDa subunit). | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.033650 (rank : 44) | θ value | 4.03905 (rank : 47) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
LC7L2_MOUSE
|
||||||
| NC score | 0.033033 (rank : 45) | θ value | 3.0926 (rank : 39) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 244 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q7TNC4, Q99LM5, Q99PC3 | Gene names | Luc7l2 | |||
|
Domain Architecture |

|
|||||
| Description | Putative RNA-binding protein Luc7-like 2 (CGI-74 homolog). | |||||
|
TRDN_HUMAN
|
||||||
| NC score | 0.031507 (rank : 46) | θ value | 6.88961 (rank : 67) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 542 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q13061 | Gene names | TRDN | |||
|
Domain Architecture |

|
|||||
| Description | Triadin. | |||||
|
DEK_MOUSE
|
||||||
| NC score | 0.031353 (rank : 47) | θ value | 8.99809 (rank : 72) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q7TNV0, Q80VC5, Q8BZV6 | Gene names | Dek | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein DEK. | |||||
|
SPRL1_HUMAN
|
||||||
| NC score | 0.031142 (rank : 48) | θ value | 4.03905 (rank : 46) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q14515, Q14800 | Gene names | SPARCL1 | |||
|
Domain Architecture |

|
|||||
| Description | SPARC-like protein 1 precursor (High endothelial venule protein) (Hevin) (MAST 9). | |||||
|
FOXGA_HUMAN
|
||||||
| NC score | 0.030501 (rank : 49) | θ value | 0.0113563 (rank : 12) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P55316 | Gene names | FOXG1A, FKHL2 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein G1A (Forkhead-related protein FKHL2) (Transcription factor BF-2) (Brain factor 2) (BF2) (HFK2). | |||||
|
GSLG1_MOUSE
|
||||||
| NC score | 0.030142 (rank : 50) | θ value | 1.81305 (rank : 33) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q61543, Q9QZ40 | Gene names | Glg1, Esl1, Mg160, Selel | |||
|
Domain Architecture |
|
|||||
| Description | Golgi apparatus protein 1 precursor (Golgi sialoglycoprotein MG-160) (E-selectin ligand 1) (ESL-1) (Selel). | |||||
|
UBP53_MOUSE
|
||||||
| NC score | 0.029731 (rank : 51) | θ value | 3.0926 (rank : 41) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P15975, Q8CB37, Q8R251 | Gene names | Usp53, Phxr3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inactive ubiquitin carboxyl-terminal hydrolase 53 (Inactive ubiquitin- specific peptidase 53) (Per-hexamer repeat protein 3). | |||||
|
SRRM1_MOUSE
|
||||||
| NC score | 0.029170 (rank : 52) | θ value | 6.88961 (rank : 65) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
ANR11_HUMAN
|
||||||
| NC score | 0.028685 (rank : 53) | θ value | 0.47712 (rank : 24) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1289 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q6UB99, Q6NTG1, Q6QMF8 | Gene names | ANKRD11, ANCO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 11 (Ankyrin repeat-containing cofactor 1). | |||||
|
K1210_HUMAN
|
||||||
| NC score | 0.028278 (rank : 54) | θ value | 6.88961 (rank : 62) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9ULL0, Q5JPN4 | Gene names | KIAA1210 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA1210. | |||||
|
RPA1_MOUSE
|
||||||
| NC score | 0.026823 (rank : 55) | θ value | 1.38821 (rank : 30) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O35134, Q7TSA9 | Gene names | Polr1a, Rpa1, Rpo1-4 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-directed RNA polymerase I largest subunit (EC 2.7.7.6) (RNA polymerase I 194 kDa subunit) (RPA194). | |||||
|
CCNL1_HUMAN
|
||||||
| NC score | 0.025498 (rank : 56) | θ value | 8.99809 (rank : 71) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9UK58, Q6NVY9, Q6UWS7, Q8NI48, Q96QT0, Q9NZF3 | Gene names | CCNL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-L1 (Cyclin-L). | |||||
|
CABIN_HUMAN
|
||||||
| NC score | 0.024689 (rank : 57) | θ value | 4.03905 (rank : 42) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 490 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Y6J0, Q9Y460 | Gene names | CABIN1, KIAA0330 | |||
|
Domain Architecture |

|
|||||
| Description | Calcineurin-binding protein Cabin 1 (Calcineurin inhibitor) (CAIN). | |||||
|
XPC_MOUSE
|
||||||
| NC score | 0.023662 (rank : 58) | θ value | 5.27518 (rank : 57) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P51612, P54732, Q3TKI2, Q920M1, Q9DBW7 | Gene names | Xpc | |||
|
Domain Architecture |

|
|||||
| Description | DNA-repair protein complementing XP-C cells homolog (Xeroderma pigmentosum group C-complementing protein homolog) (p125). | |||||
|
BRD4_HUMAN
|
||||||
| NC score | 0.019931 (rank : 59) | θ value | 3.0926 (rank : 38) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 851 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O60885, O60433, Q96PD3 | Gene names | BRD4, HUNK1 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 4 (HUNK1 protein). | |||||
|
TOP2A_MOUSE
|
||||||
| NC score | 0.019905 (rank : 60) | θ value | 3.0926 (rank : 40) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q01320 | Gene names | Top2a, Top-2, Top2 | |||
|
Domain Architecture |

|
|||||
| Description | DNA topoisomerase 2-alpha (EC 5.99.1.3) (DNA topoisomerase II, alpha isozyme). | |||||
|
DCP1B_HUMAN
|
||||||
| NC score | 0.018779 (rank : 61) | θ value | 6.88961 (rank : 60) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8IZD4, Q86XH9, Q96BP8, Q96MZ8 | Gene names | DCP1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | mRNA decapping enzyme 1B (EC 3.-.-.-). | |||||
|
DDX23_HUMAN
|
||||||
| NC score | 0.018696 (rank : 62) | θ value | 1.06291 (rank : 26) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 486 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9BUQ8, O43188 | Gene names | DDX23 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ATP-dependent RNA helicase DDX23 (EC 3.6.1.-) (DEAD box protein 23) (100 kDa U5 snRNP-specific protein) (U5-100kD) (PRP28 homolog). | |||||
|
TTLL7_HUMAN
|
||||||
| NC score | 0.018110 (rank : 63) | θ value | 4.03905 (rank : 48) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6ZT98, Q5TAX8, Q5TAX9, Q6P990, Q86YS1, Q9H5U4 | Gene names | TTLL7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tubulin--tyrosine ligase-like protein 7 (Protein NYD-SP30). | |||||
|
SON_HUMAN
|
||||||
| NC score | 0.016569 (rank : 64) | θ value | 6.88961 (rank : 64) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
SYNPO_MOUSE
|
||||||
| NC score | 0.016035 (rank : 65) | θ value | 6.88961 (rank : 66) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8CC35, Q99JI0 | Gene names | Synpo, Kiaa1029 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptopodin. | |||||
|
IF3A_HUMAN
|
||||||
| NC score | 0.015938 (rank : 66) | θ value | 4.03905 (rank : 44) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 875 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q14152, O00653 | Gene names | EIF3S10, KIAA0139 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 3 subunit 10 (eIF-3 theta) (eIF3 p167) (eIF3 p180) (eIF3 p185) (eIF3a). | |||||
|
TCPZ_HUMAN
|
||||||
| NC score | 0.014447 (rank : 67) | θ value | 5.27518 (rank : 55) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P40227, Q75LP4, Q96S46 | Gene names | CCT6A, CCT6, CCTZ | |||
|
Domain Architecture |

|
|||||
| Description | T-complex protein 1 subunit zeta (TCP-1-zeta) (CCT-zeta) (CCT-zeta-1) (Tcp20) (HTR3) (Acute morphine dependence-related protein 2). | |||||
|
TCPZ_MOUSE
|
||||||
| NC score | 0.014404 (rank : 68) | θ value | 5.27518 (rank : 56) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P80317 | Gene names | Cct6a, Cct6, Cctz, Cctz1 | |||
|
Domain Architecture |

|
|||||
| Description | T-complex protein 1 subunit zeta (TCP-1-zeta) (CCT-zeta) (CCT-zeta-1). | |||||
|
UBQL4_MOUSE
|
||||||
| NC score | 0.014315 (rank : 69) | θ value | 4.03905 (rank : 49) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99NB8, Q8BP88 | Gene names | Ubqln4, Ubin | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquilin-4 (Ataxin-1 ubiquitin-like-interacting protein A1U). | |||||
|
MYOZ2_HUMAN
|
||||||
| NC score | 0.013604 (rank : 70) | θ value | 8.99809 (rank : 74) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NPC6, O43415, Q9HB92 | Gene names | MYOZ2, C4orf5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myozenin-2 (Calsarcin-1) (FATZ-related protein 2). | |||||
|
WWC1_MOUSE
|
||||||
| NC score | 0.011369 (rank : 71) | θ value | 6.88961 (rank : 68) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 290 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5SXA9, Q571D0, Q8K1Y3, Q8VD17, Q922W3 | Gene names | Wwc1, Kiaa0869 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | WW domain-containing protein 1 (Kidney and brain protein) (KIBRA). | |||||
|
FRM4A_MOUSE
|
||||||
| NC score | 0.011143 (rank : 72) | θ value | 5.27518 (rank : 53) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BIE6, Q6PDJ4, Q8CHA6 | Gene names | Frmd4a, Frmd4, Kiaa1294 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FERM domain-containing protein 4A. | |||||
|
CAC1B_MOUSE
|
||||||
| NC score | 0.010457 (rank : 73) | θ value | 4.03905 (rank : 43) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O55017, Q60609 | Gene names | Cacna1b, Cach5, Cacnl1a5, Cchn1a | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent N-type calcium channel subunit alpha-1B (Voltage- gated calcium channel subunit alpha Cav2.2) (Calcium channel, L type, alpha-1 polypeptide isoform 5) (Brain calcium channel III) (BIII). | |||||
|
BCAR3_MOUSE
|
||||||
| NC score | 0.008306 (rank : 74) | θ value | 8.99809 (rank : 70) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 127 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9QZK2, Q3TNC9, Q3UP10 | Gene names | Bcar3, And34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Breast cancer anti-estrogen resistance protein 3 (p130Cas-binding protein AND-34). | |||||
|
MMP1_HUMAN
|
||||||
| NC score | 0.003900 (rank : 75) | θ value | 8.99809 (rank : 73) | |||
| Query Neighborhood Hits | 75 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P03956, P08156 | Gene names | MMP1, CLG | |||
|
Domain Architecture |

|
|||||
| Description | Interstitial collagenase precursor (EC 3.4.24.7) (Matrix metalloproteinase-1) (MMP-1) (Fibroblast collagenase) [Contains: 22 kDa interstitial collagenase; 27 kDa interstitial collagenase]. | |||||