Please be patient as the page loads
|
PGAM2_MOUSE
|
||||||
| SwissProt Accessions | O70250 | Gene names | Pgam2 | |||
|
Domain Architecture |
|
|||||
| Description | Phosphoglycerate mutase 2 (EC 5.4.2.1) (EC 5.4.2.4) (EC 3.1.3.13) (Phosphoglycerate mutase isozyme M) (PGAM-M) (BPG-dependent PGAM 2) (Muscle-specific phosphoglycerate mutase). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
PGAM2_MOUSE
|
||||||
| θ value | 4.88915e-147 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O70250 | Gene names | Pgam2 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphoglycerate mutase 2 (EC 5.4.2.1) (EC 5.4.2.4) (EC 3.1.3.13) (Phosphoglycerate mutase isozyme M) (PGAM-M) (BPG-dependent PGAM 2) (Muscle-specific phosphoglycerate mutase). | |||||
|
PGAM2_HUMAN
|
||||||
| θ value | 1.4755e-135 (rank : 2) | NC score | 0.999547 (rank : 2) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P15259 | Gene names | PGAM2, PGAMM | |||
|
Domain Architecture |
|
|||||
| Description | Phosphoglycerate mutase 2 (EC 5.4.2.1) (EC 5.4.2.4) (EC 3.1.3.13) (Phosphoglycerate mutase isozyme M) (PGAM-M) (BPG-dependent PGAM 2) (Muscle-specific phosphoglycerate mutase). | |||||
|
PGAM1_HUMAN
|
||||||
| θ value | 4.31301e-119 (rank : 3) | NC score | 0.998005 (rank : 3) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P18669, Q9BWC0 | Gene names | PGAM1 | |||
|
Domain Architecture |
|
|||||
| Description | Phosphoglycerate mutase 1 (EC 5.4.2.1) (EC 5.4.2.4) (EC 3.1.3.13) (Phosphoglycerate mutase isozyme B) (PGAM-B) (BPG-dependent PGAM 1). | |||||
|
PGAM1_MOUSE
|
||||||
| θ value | 1.2549e-118 (rank : 4) | NC score | 0.997985 (rank : 4) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9DBJ1, Q9D6W0, Q9ERT3 | Gene names | Pgam1 | |||
|
Domain Architecture |
|
|||||
| Description | Phosphoglycerate mutase 1 (EC 5.4.2.1) (EC 5.4.2.4) (EC 3.1.3.13) (Phosphoglycerate mutase isozyme B) (PGAM-B) (BPG-dependent PGAM 1). | |||||
|
PGAM4_HUMAN
|
||||||
| θ value | 2.29227e-112 (rank : 5) | NC score | 0.996576 (rank : 5) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8N0Y7, Q8NI24, Q8NI25, Q8NI26 | Gene names | PGAM4, PGAM3 | |||
|
Domain Architecture |
|
|||||
| Description | Probable phosphoglycerate mutase 4 (EC 5.4.2.1) (EC 5.4.2.4) (EC 3.1.3.13). | |||||
|
PMGE_HUMAN
|
||||||
| θ value | 7.44272e-79 (rank : 6) | NC score | 0.981935 (rank : 6) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P07738 | Gene names | BPGM | |||
|
Domain Architecture |
|
|||||
| Description | Bisphosphoglycerate mutase (EC 5.4.2.4) (2,3-bisphosphoglycerate mutase, erythrocyte) (2,3-bisphosphoglycerate synthase) (BPGM) (EC 5.4.2.1) (EC 3.1.3.13) (BPG-dependent PGAM). | |||||
|
PMGE_MOUSE
|
||||||
| θ value | 1.18825e-76 (rank : 7) | NC score | 0.979929 (rank : 7) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P15327, Q543Z6 | Gene names | Bpgm | |||
|
Domain Architecture |
|
|||||
| Description | Bisphosphoglycerate mutase (EC 5.4.2.4) (2,3-bisphosphoglycerate mutase, erythrocyte) (2,3-bisphosphoglycerate synthase) (BPGM) (EC 5.4.2.1) (EC 3.1.3.13) (BPG-dependent PGAM). | |||||
|
CL005_HUMAN
|
||||||
| θ value | 0.00665767 (rank : 8) | NC score | 0.260410 (rank : 8) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NQ88 | Gene names | C12orf5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C12orf5. | |||||
|
F264_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 9) | NC score | 0.148119 (rank : 10) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q16877 | Gene names | PFKFB4 | |||
|
Domain Architecture |

|
|||||
| Description | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 4 (6PF-2-K/Fru- 2,6-P2ASE testis-type isozyme) [Includes: 6-phosphofructo-2-kinase (EC 2.7.1.105); Fructose-2,6-bisphosphatase (EC 3.1.3.46)]. | |||||
|
CL005_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 10) | NC score | 0.244818 (rank : 9) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8BZA9 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C12orf5 homolog. | |||||
|
F262_HUMAN
|
||||||
| θ value | 0.62314 (rank : 11) | NC score | 0.086679 (rank : 12) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O60825, O60824, Q9H3P1 | Gene names | PFKFB2 | |||
|
Domain Architecture |

|
|||||
| Description | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 2 (6PF-2-K/Fru- 2,6-P2ASE heart-type isozyme) (PFK-2/FBPase-2) [Includes: 6- phosphofructo-2-kinase (EC 2.7.1.105); Fructose-2,6-bisphosphatase (EC 3.1.3.46)]. | |||||
|
F261_HUMAN
|
||||||
| θ value | 1.06291 (rank : 12) | NC score | 0.080712 (rank : 14) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P16118, Q99951 | Gene names | PFKFB1, F6PK, PFRX | |||
|
Domain Architecture |

|
|||||
| Description | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 1 (6PF-2-K/Fru- 2,6-P2ASE liver isozyme) [Includes: 6-phosphofructo-2-kinase (EC 2.7.1.105); Fructose-2,6-bisphosphatase (EC 3.1.3.46)]. | |||||
|
F262_MOUSE
|
||||||
| θ value | 1.06291 (rank : 13) | NC score | 0.077446 (rank : 15) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P70265, Q8VEI9 | Gene names | Pfkfb2 | |||
|
Domain Architecture |

|
|||||
| Description | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 2 (6PF-2-K/Fru- 2,6-P2ASE heart-type isozyme) (PFK-2/FBPase-2) [Includes: 6- phosphofructo-2-kinase (EC 2.7.1.105); Fructose-2,6-bisphosphatase (EC 3.1.3.46)]. | |||||
|
STS1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 14) | NC score | 0.096432 (rank : 11) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8TF42, Q53GT5, Q53GT8, Q8NBV7, Q96IG9, Q96NZ2 | Gene names | STS1, KIAA1959 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppressor of T-cell receptor signaling 1 (Sts-1) (Cbl-interacting protein p70). | |||||
|
STS1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 15) | NC score | 0.083219 (rank : 13) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8BGG7, Q3U8Z2, Q8BMW9 | Gene names | Sts1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppressor of T-cell receptor signaling 1 (Sts-1) (Cbl-interacting protein p70). | |||||
|
FUBP1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 16) | NC score | 0.011808 (rank : 16) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96AE4, Q12828 | Gene names | FUBP1 | |||
|
Domain Architecture |
|
|||||
| Description | Far upstream element-binding protein 1 (FUSE-binding protein 1) (FBP) (DNA helicase V) (HDH V). | |||||
|
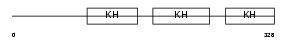
FUBP1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 17) | NC score | 0.011787 (rank : 17) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q91WJ8, Q8C0Y8 | Gene names | Fubp1, D3Ertd330e | |||
|
Domain Architecture |
|
|||||
| Description | Far upstream element-binding protein 1 (FUSE-binding protein 1) (FBP). | |||||
|
INOC1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 18) | NC score | 0.006800 (rank : 19) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6ZPV2, Q6P7V0, Q6PCP1, Q8C9T7 | Gene names | Inoc1, Kiaa1259 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative DNA helicase INO80 complex homolog 1 (EC 3.6.1.-). | |||||
|
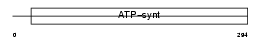
ATPG_HUMAN
|
||||||
| θ value | 8.99809 (rank : 19) | NC score | 0.011038 (rank : 18) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P36542, Q5VYP3, Q96AS8 | Gene names | ATP5C1, ATP5C | |||
|
Domain Architecture |
|
|||||
| Description | ATP synthase gamma chain, mitochondrial precursor (EC 3.6.3.14). | |||||
|
PGAM2_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 4.88915e-147 (rank : 1) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O70250 | Gene names | Pgam2 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphoglycerate mutase 2 (EC 5.4.2.1) (EC 5.4.2.4) (EC 3.1.3.13) (Phosphoglycerate mutase isozyme M) (PGAM-M) (BPG-dependent PGAM 2) (Muscle-specific phosphoglycerate mutase). | |||||
|
PGAM2_HUMAN
|
||||||
| NC score | 0.999547 (rank : 2) | θ value | 1.4755e-135 (rank : 2) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P15259 | Gene names | PGAM2, PGAMM | |||
|
Domain Architecture |

|
|||||
| Description | Phosphoglycerate mutase 2 (EC 5.4.2.1) (EC 5.4.2.4) (EC 3.1.3.13) (Phosphoglycerate mutase isozyme M) (PGAM-M) (BPG-dependent PGAM 2) (Muscle-specific phosphoglycerate mutase). | |||||
|
PGAM1_HUMAN
|
||||||
| NC score | 0.998005 (rank : 3) | θ value | 4.31301e-119 (rank : 3) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P18669, Q9BWC0 | Gene names | PGAM1 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphoglycerate mutase 1 (EC 5.4.2.1) (EC 5.4.2.4) (EC 3.1.3.13) (Phosphoglycerate mutase isozyme B) (PGAM-B) (BPG-dependent PGAM 1). | |||||
|
PGAM1_MOUSE
|
||||||
| NC score | 0.997985 (rank : 4) | θ value | 1.2549e-118 (rank : 4) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9DBJ1, Q9D6W0, Q9ERT3 | Gene names | Pgam1 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphoglycerate mutase 1 (EC 5.4.2.1) (EC 5.4.2.4) (EC 3.1.3.13) (Phosphoglycerate mutase isozyme B) (PGAM-B) (BPG-dependent PGAM 1). | |||||
|
PGAM4_HUMAN
|
||||||
| NC score | 0.996576 (rank : 5) | θ value | 2.29227e-112 (rank : 5) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8N0Y7, Q8NI24, Q8NI25, Q8NI26 | Gene names | PGAM4, PGAM3 | |||
|
Domain Architecture |

|
|||||
| Description | Probable phosphoglycerate mutase 4 (EC 5.4.2.1) (EC 5.4.2.4) (EC 3.1.3.13). | |||||
|
PMGE_HUMAN
|
||||||
| NC score | 0.981935 (rank : 6) | θ value | 7.44272e-79 (rank : 6) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P07738 | Gene names | BPGM | |||
|
Domain Architecture |

|
|||||
| Description | Bisphosphoglycerate mutase (EC 5.4.2.4) (2,3-bisphosphoglycerate mutase, erythrocyte) (2,3-bisphosphoglycerate synthase) (BPGM) (EC 5.4.2.1) (EC 3.1.3.13) (BPG-dependent PGAM). | |||||
|
PMGE_MOUSE
|
||||||
| NC score | 0.979929 (rank : 7) | θ value | 1.18825e-76 (rank : 7) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P15327, Q543Z6 | Gene names | Bpgm | |||
|
Domain Architecture |

|
|||||
| Description | Bisphosphoglycerate mutase (EC 5.4.2.4) (2,3-bisphosphoglycerate mutase, erythrocyte) (2,3-bisphosphoglycerate synthase) (BPGM) (EC 5.4.2.1) (EC 3.1.3.13) (BPG-dependent PGAM). | |||||
|
CL005_HUMAN
|
||||||
| NC score | 0.260410 (rank : 8) | θ value | 0.00665767 (rank : 8) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NQ88 | Gene names | C12orf5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C12orf5. | |||||
|
CL005_MOUSE
|
||||||
| NC score | 0.244818 (rank : 9) | θ value | 0.0431538 (rank : 10) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8BZA9 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C12orf5 homolog. | |||||
|
F264_HUMAN
|
||||||
| NC score | 0.148119 (rank : 10) | θ value | 0.0252991 (rank : 9) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q16877 | Gene names | PFKFB4 | |||
|
Domain Architecture |

|
|||||
| Description | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 4 (6PF-2-K/Fru- 2,6-P2ASE testis-type isozyme) [Includes: 6-phosphofructo-2-kinase (EC 2.7.1.105); Fructose-2,6-bisphosphatase (EC 3.1.3.46)]. | |||||
|
STS1_HUMAN
|
||||||
| NC score | 0.096432 (rank : 11) | θ value | 2.36792 (rank : 14) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8TF42, Q53GT5, Q53GT8, Q8NBV7, Q96IG9, Q96NZ2 | Gene names | STS1, KIAA1959 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppressor of T-cell receptor signaling 1 (Sts-1) (Cbl-interacting protein p70). | |||||
|
F262_HUMAN
|
||||||
| NC score | 0.086679 (rank : 12) | θ value | 0.62314 (rank : 11) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O60825, O60824, Q9H3P1 | Gene names | PFKFB2 | |||
|
Domain Architecture |

|
|||||
| Description | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 2 (6PF-2-K/Fru- 2,6-P2ASE heart-type isozyme) (PFK-2/FBPase-2) [Includes: 6- phosphofructo-2-kinase (EC 2.7.1.105); Fructose-2,6-bisphosphatase (EC 3.1.3.46)]. | |||||
|
STS1_MOUSE
|
||||||
| NC score | 0.083219 (rank : 13) | θ value | 5.27518 (rank : 15) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 104 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8BGG7, Q3U8Z2, Q8BMW9 | Gene names | Sts1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppressor of T-cell receptor signaling 1 (Sts-1) (Cbl-interacting protein p70). | |||||
|
F261_HUMAN
|
||||||
| NC score | 0.080712 (rank : 14) | θ value | 1.06291 (rank : 12) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P16118, Q99951 | Gene names | PFKFB1, F6PK, PFRX | |||
|
Domain Architecture |

|
|||||
| Description | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 1 (6PF-2-K/Fru- 2,6-P2ASE liver isozyme) [Includes: 6-phosphofructo-2-kinase (EC 2.7.1.105); Fructose-2,6-bisphosphatase (EC 3.1.3.46)]. | |||||
|
F262_MOUSE
|
||||||
| NC score | 0.077446 (rank : 15) | θ value | 1.06291 (rank : 13) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P70265, Q8VEI9 | Gene names | Pfkfb2 | |||
|
Domain Architecture |

|
|||||
| Description | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 2 (6PF-2-K/Fru- 2,6-P2ASE heart-type isozyme) (PFK-2/FBPase-2) [Includes: 6- phosphofructo-2-kinase (EC 2.7.1.105); Fructose-2,6-bisphosphatase (EC 3.1.3.46)]. | |||||
|
FUBP1_HUMAN
|
||||||
| NC score | 0.011808 (rank : 16) | θ value | 6.88961 (rank : 16) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96AE4, Q12828 | Gene names | FUBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Far upstream element-binding protein 1 (FUSE-binding protein 1) (FBP) (DNA helicase V) (HDH V). | |||||
|
FUBP1_MOUSE
|
||||||
| NC score | 0.011787 (rank : 17) | θ value | 6.88961 (rank : 17) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q91WJ8, Q8C0Y8 | Gene names | Fubp1, D3Ertd330e | |||
|
Domain Architecture |

|
|||||
| Description | Far upstream element-binding protein 1 (FUSE-binding protein 1) (FBP). | |||||
|
ATPG_HUMAN
|
||||||
| NC score | 0.011038 (rank : 18) | θ value | 8.99809 (rank : 19) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P36542, Q5VYP3, Q96AS8 | Gene names | ATP5C1, ATP5C | |||
|
Domain Architecture |

|
|||||
| Description | ATP synthase gamma chain, mitochondrial precursor (EC 3.6.3.14). | |||||
|
INOC1_MOUSE
|
||||||
| NC score | 0.006800 (rank : 19) | θ value | 6.88961 (rank : 18) | |||
| Query Neighborhood Hits | 19 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6ZPV2, Q6P7V0, Q6PCP1, Q8C9T7 | Gene names | Inoc1, Kiaa1259 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative DNA helicase INO80 complex homolog 1 (EC 3.6.1.-). | |||||