Please be patient as the page loads
|
IP3KC_HUMAN
|
||||||
| SwissProt Accessions | Q96DU7, Q9UE25, Q9Y475 | Gene names | ITPKC, IP3KC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol-trisphosphate 3-kinase C (EC 2.7.1.127) (Inositol 1,4,5- trisphosphate 3-kinase C) (InsP 3-kinase C) (IP3K-C). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
IP3KC_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q96DU7, Q9UE25, Q9Y475 | Gene names | ITPKC, IP3KC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol-trisphosphate 3-kinase C (EC 2.7.1.127) (Inositol 1,4,5- trisphosphate 3-kinase C) (InsP 3-kinase C) (IP3K-C). | |||||
|
IP3KC_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.984772 (rank : 2) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q7TS72, Q3U384 | Gene names | Itpkc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol-trisphosphate 3-kinase C (EC 2.7.1.127) (Inositol 1,4,5- trisphosphate 3-kinase C) (IP3K-C). | |||||
|
IP3KB_HUMAN
|
||||||
| θ value | 1.21547e-113 (rank : 3) | NC score | 0.928394 (rank : 5) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P27987, Q5VWL9, Q5VWM0, Q96BZ2, Q96JS1, Q9UH47 | Gene names | ITPKB | |||
|
Domain Architecture |

|
|||||
| Description | Inositol-trisphosphate 3-kinase B (EC 2.7.1.127) (Inositol 1,4,5- trisphosphate 3-kinase B) (IP3K B) (IP3 3-kinase B) (IP3K-B). | |||||
|
IP3KA_HUMAN
|
||||||
| θ value | 3.00075e-104 (rank : 4) | NC score | 0.951304 (rank : 4) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P23677, Q8TAN3 | Gene names | ITPKA | |||
|
Domain Architecture |

|
|||||
| Description | Inositol-trisphosphate 3-kinase A (EC 2.7.1.127) (Inositol 1,4,5- trisphosphate 3-kinase A) (IP3K A) (IP3 3-kinase A). | |||||
|
IP3KA_MOUSE
|
||||||
| θ value | 3.00075e-104 (rank : 5) | NC score | 0.956819 (rank : 3) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8R071 | Gene names | Itpka | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol-trisphosphate 3-kinase A (EC 2.7.1.127) (Inositol 1,4,5- trisphosphate 3-kinase A) (IP3K A) (IP3 3-kinase A). | |||||
|
SSPO_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 6) | NC score | 0.023580 (rank : 20) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 800 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8CG65 | Gene names | Sspo | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SCO-spondin precursor. | |||||
|
CONA1_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 7) | NC score | 0.027370 (rank : 14) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q86Y22, Q8IVR4, Q9NT93 | Gene names | COL23A1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XXIII) chain. | |||||
|
MTMR3_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 8) | NC score | 0.026703 (rank : 17) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q13615, Q9NYN5, Q9NYN6, Q9UDX6, Q9UEG3 | Gene names | MTMR3, KIAA0371, ZFYVE10 | |||
|
Domain Architecture |

|
|||||
| Description | Myotubularin-related protein 3 (EC 3.1.3.48) (FYVE domain-containing dual specificity protein phosphatase 1) (FYVE-DSP1) (Zinc finger FYVE domain-containing protein 10). | |||||
|
TR150_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 9) | NC score | 0.041037 (rank : 11) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 468 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q569Z6 | Gene names | Thrap3, Trap150 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid hormone receptor-associated protein 3 (Thyroid hormone receptor-associated protein complex 150 kDa component) (Trap150). | |||||
|
IP6K3_HUMAN
|
||||||
| θ value | 0.279714 (rank : 10) | NC score | 0.078066 (rank : 8) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96PC2, Q96MQ9 | Gene names | IHPK3, IP6K3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol hexaphosphate kinase 3 (EC 2.7.4.21) (InsP6 kinase 3) (Inositol hexakisphosphate kinase 3). | |||||
|
IPMK_HUMAN
|
||||||
| θ value | 0.47712 (rank : 11) | NC score | 0.136233 (rank : 6) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8NFU5 | Gene names | IPMK, IMPK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol polyphosphate multikinase (EC 2.7.1.151) (Inositol 1,3,4,6- tetrakisphosphate 5-kinase). | |||||
|
NSBP1_MOUSE
|
||||||
| θ value | 0.47712 (rank : 12) | NC score | 0.027159 (rank : 16) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9JL35, O88832, Q8VC71, Q9CUW1 | Gene names | Nsbp1, Garp45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1 (Nucleosome-binding protein 45) (NBP-45) (GARP45 protein). | |||||
|
LRC42_HUMAN
|
||||||
| θ value | 0.62314 (rank : 13) | NC score | 0.038495 (rank : 12) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y546, Q8N2Q8 | Gene names | LRRC42 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 42. | |||||
|
PCD18_HUMAN
|
||||||
| θ value | 1.06291 (rank : 14) | NC score | 0.005743 (rank : 48) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9HCL0 | Gene names | PCDH18, KIAA1562 | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin-18 precursor. | |||||
|
ARHG1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 15) | NC score | 0.015716 (rank : 28) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q92888, O00513, Q8N4J4, Q96BF4, Q96F17, Q9BSB1 | Gene names | ARHGEF1 | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 1 (p115-RhoGEF) (p115RhoGEF) (115 kDa guanine nucleotide exchange factor) (Sub1.5). | |||||
|
IPMK_MOUSE
|
||||||
| θ value | 1.38821 (rank : 16) | NC score | 0.131354 (rank : 7) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q7TT16, Q8BZ11, Q8BZA8, Q9CWM9 | Gene names | Ipmk, Impk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol polyphosphate multikinase (EC 2.7.1.151) (Inositol 1,3,4,6- tetrakisphosphate 5-kinase). | |||||
|
CO2A1_HUMAN
|
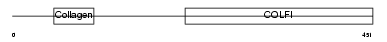
||||||
| θ value | 1.81305 (rank : 17) | NC score | 0.021868 (rank : 22) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 486 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P02458, Q12985, Q14009, Q14044, Q14046, Q14056, Q14058, Q16672, Q6LBY1, Q6LBY2, Q6LBY3, Q99227, Q9UE38, Q9UE39, Q9UE40, Q9UE41, Q9UE42, Q9UE43 | Gene names | COL2A1 | |||
|
Domain Architecture |
|
|||||
| Description | Collagen alpha-1(II) chain precursor [Contains: Chondrocalcin]. | |||||
|
CONA1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 18) | NC score | 0.023412 (rank : 21) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 254 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8K4G2, Q5SUQ0 | Gene names | Col23a1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XXIII) chain. | |||||
|
SORC3_HUMAN
|
||||||
| θ value | 1.81305 (rank : 19) | NC score | 0.014729 (rank : 29) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UPU3, Q5VXF9, Q9NQJ2 | Gene names | SORCS3, KIAA1059 | |||
|
Domain Architecture |

|
|||||
| Description | VPS10 domain-containing receptor SorCS3 precursor. | |||||
|
TP53B_MOUSE
|
||||||
| θ value | 1.81305 (rank : 20) | NC score | 0.020758 (rank : 24) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 644 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P70399, Q68FD0, Q8CI97, Q91YC9 | Gene names | Tp53bp1, Trp53bp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor suppressor p53-binding protein 1 (p53-binding protein 1) (p53BP1) (53BP1). | |||||
|
ZN385_HUMAN
|
||||||
| θ value | 1.81305 (rank : 21) | NC score | 0.021512 (rank : 23) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 614 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96PM9, Q5VH53, Q9H7R6, Q9UFU3 | Gene names | ZNF385, HZF, RZF | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 385 (Hematopoietic zinc finger protein) (Retinal zinc finger protein). | |||||
|
LUZP1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 22) | NC score | 0.010264 (rank : 39) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 940 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8R4U7, Q3UV18, Q7TS71, Q8BQW1, Q99NG3, Q9CSL6 | Gene names | Luzp1, Luzp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine zipper protein 1 (Leucine zipper motif-containing protein). | |||||
|
PMS2_HUMAN
|
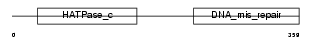
||||||
| θ value | 2.36792 (rank : 23) | NC score | 0.036845 (rank : 13) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P54278 | Gene names | PMS2, PMSL2 | |||
|
Domain Architecture |
|
|||||
| Description | PMS1 protein homolog 2 (DNA mismatch repair protein PMS2). | |||||
|
DEND_HUMAN
|
||||||
| θ value | 3.0926 (rank : 24) | NC score | 0.023911 (rank : 19) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O94850 | Gene names | DDN, KIAA0749 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dendrin. | |||||
|
FGF3_MOUSE
|
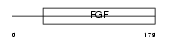
||||||
| θ value | 3.0926 (rank : 25) | NC score | 0.011120 (rank : 35) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P05524, Q61736 | Gene names | Fgf3, Fgf-3, Int-2 | |||
|
Domain Architecture |
|
|||||
| Description | INT-2 proto-oncogene protein precursor (Fibroblast growth factor 3) (FGF-3) (HBGF-3). | |||||
|
CSPG3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 26) | NC score | 0.005257 (rank : 49) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O14594, Q9UPK6 | Gene names | CSPG3, NCAN, NEUR | |||
|
Domain Architecture |

|
|||||
| Description | Neurocan core protein precursor (Chondroitin sulfate proteoglycan 3). | |||||
|
EMIL1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 27) | NC score | 0.015951 (rank : 26) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9Y6C2, Q96G58, Q9UG76 | Gene names | EMILIN1, EMI | |||
|
Domain Architecture |

|
|||||
| Description | EMILIN-1 precursor (Elastin microfibril interface-located protein 1) (Elastin microfibril interfacer 1). | |||||
|
JADE3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 28) | NC score | 0.008296 (rank : 45) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6IE82, Q6ZQG2 | Gene names | Phf16, Jade3, Kiaa0215 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Jade-3 (PHD finger protein 16). | |||||
|
MDC1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 29) | NC score | 0.009924 (rank : 41) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
NFH_MOUSE
|
||||||
| θ value | 5.27518 (rank : 30) | NC score | 0.011128 (rank : 34) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
TARA_MOUSE
|
||||||
| θ value | 5.27518 (rank : 31) | NC score | 0.010276 (rank : 38) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1133 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q99KW3, Q2PZW9, Q2Q402, Q2Q403, Q2Q404, Q6ZPK4, Q8C6T3, Q8CG90 | Gene names | Triobp, Kiaa1662, Tara | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
AP3B2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 32) | NC score | 0.008835 (rank : 44) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9JME5, Q6QR53, Q8R1E5 | Gene names | Ap3b2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AP-3 complex subunit beta-2 (Adapter-related protein complex 3 beta-2 subunit) (Beta3B-adaptin) (Adaptor protein complex AP-3 beta-2 subunit) (Clathrin assembly protein complex 3 beta-2 large chain). | |||||
|
DIDO1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 33) | NC score | 0.013646 (rank : 30) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9BTC0, O15043, Q3ZTL7, Q3ZTL8, Q4VXS1, Q4VXS2, Q96D72, Q9BQW0, Q9BW03, Q9H4G6, Q9H4G7, Q9NTU8, Q9NUM8, Q9UFB6 | Gene names | DIDO1, C20orf158, DATF1, KIAA0333 | |||
|
Domain Architecture |

|
|||||
| Description | Death-inducer obliterator 1 (DIO-1) (Death-associated transcription factor 1) (DATF-1) (hDido1). | |||||
|
DMD_MOUSE
|
||||||
| θ value | 6.88961 (rank : 34) | NC score | 0.002789 (rank : 52) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1111 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P11531, O35653, Q60703 | Gene names | Dmd | |||
|
Domain Architecture |

|
|||||
| Description | Dystrophin. | |||||
|
FUBP2_HUMAN
|
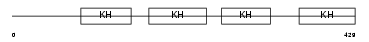
||||||
| θ value | 6.88961 (rank : 35) | NC score | 0.010691 (rank : 36) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q92945, O00301, Q9UNT5, Q9UQH5 | Gene names | KHSRP, FUBP2 | |||
|
Domain Architecture |
|
|||||
| Description | Far upstream element-binding protein 2 (FUSE-binding protein 2) (KH type-splicing regulatory protein) (KSRP) (p75). | |||||
|
LPIN3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 36) | NC score | 0.012693 (rank : 31) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9BQK8, Q5TDB9, Q9NPY8, Q9UJE5 | Gene names | LPIN3, LIPN3L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lipin-3 (Lipin 3-like). | |||||
|
PINK1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 37) | NC score | 0.003801 (rank : 50) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 741 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9BXM7, Q8N6T9, Q8NBU3, Q96DE4 | Gene names | PINK1 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase PINK1, mitochondrial precursor (EC 2.7.11.1) (PTEN-induced putative kinase protein 1) (BRPK). | |||||
|
SCND1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 38) | NC score | 0.006803 (rank : 47) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P57086 | Gene names | SCAND1, SDP1 | |||
|
Domain Architecture |
|
|||||
| Description | SCAN domain-containing protein 1. | |||||
|
SHC3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 39) | NC score | 0.010260 (rank : 40) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q61120 | Gene names | Shc3, Nshc, ShcC | |||
|
Domain Architecture |

|
|||||
| Description | SHC-transforming protein 3 (SH2 domain protein C3) (Src homology 2 domain-containing-transforming protein C3) (Neuronal Shc) (N-Shc). | |||||
|
SIX5_HUMAN
|
||||||
| θ value | 6.88961 (rank : 40) | NC score | 0.006886 (rank : 46) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 386 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8N196 | Gene names | SIX5, DMAHP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein SIX5 (DM locus-associated homeodomain protein). | |||||
|
SRRM2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 41) | NC score | 0.027318 (rank : 15) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
SRTD2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 42) | NC score | 0.025793 (rank : 18) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9JJG5, Q8C609, Q91WL3, Q925E5 | Gene names | Sertad2, Kiaa0127 | |||
|
Domain Architecture |
|
|||||
| Description | SERTA domain-containing protein 2 (Transcriptional regulator interacting with the PHD-bromodomain 2) (TRIP-Br2). | |||||
|
CO2A1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 43) | NC score | 0.020165 (rank : 25) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 483 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P28481 | Gene names | Col2a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(II) chain precursor [Contains: Chondrocalcin]. | |||||
|
DEPP_HUMAN
|
||||||
| θ value | 8.99809 (rank : 44) | NC score | 0.015883 (rank : 27) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NTK1, O94997, Q5T735, Q76MX8 | Gene names | DEPP, C10orf10, FIG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein DEPP (Decidual protein induced by progesterone) (Fasting- induced gene protein) (FIG). | |||||
|
LPIN3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 45) | NC score | 0.011895 (rank : 32) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99PI4, Q3TQ75, Q8C7R9 | Gene names | Lpin3 | |||
|
Domain Architecture |
|
|||||
| Description | Lipin-3. | |||||
|
PRGC2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 46) | NC score | 0.010664 (rank : 37) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 272 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q86YN6, Q86YN3, Q86YN4, Q86YN5, Q8N1N9, Q8TDE4, Q8TDE5 | Gene names | PPARGC1B, PERC, PGC1, PGC1B, PPARGC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor gamma coactivator 1-beta (PPAR gamma coactivator-1beta) (PGC-1-related estrogen receptor alpha coactivator) (PPARGC-1-beta) (PGC-1-beta). | |||||
|
RFXK_HUMAN
|
||||||
| θ value | 8.99809 (rank : 47) | NC score | 0.003233 (rank : 51) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O14593, O95839 | Gene names | RFXANK, ANKRA1, RFXB | |||
|
Domain Architecture |
|
|||||
| Description | DNA-binding protein RFXANK (Regulatory factor X subunit B) (RFX-B) (Ankyrin repeat family A protein 1). | |||||
|
RGS3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 48) | NC score | 0.009391 (rank : 43) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
RNF17_MOUSE
|
||||||
| θ value | 8.99809 (rank : 49) | NC score | 0.011229 (rank : 33) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99MV7 | Gene names | Rnf17 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 17. | |||||
|
SHC3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 50) | NC score | 0.009564 (rank : 42) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q92529, Q8TAP2, Q9UCX5 | Gene names | SHC3, NSHC, SHCC | |||
|
Domain Architecture |

|
|||||
| Description | SHC-transforming protein 3 (SH2 domain protein C3) (Src homology 2 domain-containing-transforming protein C3) (Neuronal Shc) (N-Shc) (Protein Rai). | |||||
|
IP6K2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 51) | NC score | 0.050071 (rank : 10) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UHH9, Q6P0N8, Q9BSZ6, Q9BUW3, Q9H4P7, Q9NT63, Q9UFU6 | Gene names | IHPK2, IP6K2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol hexakisphosphate kinase 2 (EC 2.7.4.21) (InsP6 kinase 2) (Inositol hexakisphosphate kinase 2) (P(i)-uptake stimulator) (PiUS). | |||||
|
IP6K3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 52) | NC score | 0.062905 (rank : 9) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BWD2 | Gene names | Ihpk3, Ip6k3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol hexaphosphate kinase 3 (EC 2.7.4.21) (InsP6 kinase 3) (Inositol hexakisphosphate kinase 3). | |||||
|
IP3KC_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q96DU7, Q9UE25, Q9Y475 | Gene names | ITPKC, IP3KC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol-trisphosphate 3-kinase C (EC 2.7.1.127) (Inositol 1,4,5- trisphosphate 3-kinase C) (InsP 3-kinase C) (IP3K-C). | |||||
|
IP3KC_MOUSE
|
||||||
| NC score | 0.984772 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q7TS72, Q3U384 | Gene names | Itpkc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol-trisphosphate 3-kinase C (EC 2.7.1.127) (Inositol 1,4,5- trisphosphate 3-kinase C) (IP3K-C). | |||||
|
IP3KA_MOUSE
|
||||||
| NC score | 0.956819 (rank : 3) | θ value | 3.00075e-104 (rank : 5) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8R071 | Gene names | Itpka | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol-trisphosphate 3-kinase A (EC 2.7.1.127) (Inositol 1,4,5- trisphosphate 3-kinase A) (IP3K A) (IP3 3-kinase A). | |||||
|
IP3KA_HUMAN
|
||||||
| NC score | 0.951304 (rank : 4) | θ value | 3.00075e-104 (rank : 4) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P23677, Q8TAN3 | Gene names | ITPKA | |||
|
Domain Architecture |

|
|||||
| Description | Inositol-trisphosphate 3-kinase A (EC 2.7.1.127) (Inositol 1,4,5- trisphosphate 3-kinase A) (IP3K A) (IP3 3-kinase A). | |||||
|
IP3KB_HUMAN
|
||||||
| NC score | 0.928394 (rank : 5) | θ value | 1.21547e-113 (rank : 3) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P27987, Q5VWL9, Q5VWM0, Q96BZ2, Q96JS1, Q9UH47 | Gene names | ITPKB | |||
|
Domain Architecture |

|
|||||
| Description | Inositol-trisphosphate 3-kinase B (EC 2.7.1.127) (Inositol 1,4,5- trisphosphate 3-kinase B) (IP3K B) (IP3 3-kinase B) (IP3K-B). | |||||
|
IPMK_HUMAN
|
||||||
| NC score | 0.136233 (rank : 6) | θ value | 0.47712 (rank : 11) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8NFU5 | Gene names | IPMK, IMPK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol polyphosphate multikinase (EC 2.7.1.151) (Inositol 1,3,4,6- tetrakisphosphate 5-kinase). | |||||
|
IPMK_MOUSE
|
||||||
| NC score | 0.131354 (rank : 7) | θ value | 1.38821 (rank : 16) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q7TT16, Q8BZ11, Q8BZA8, Q9CWM9 | Gene names | Ipmk, Impk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol polyphosphate multikinase (EC 2.7.1.151) (Inositol 1,3,4,6- tetrakisphosphate 5-kinase). | |||||
|
IP6K3_HUMAN
|
||||||
| NC score | 0.078066 (rank : 8) | θ value | 0.279714 (rank : 10) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q96PC2, Q96MQ9 | Gene names | IHPK3, IP6K3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol hexaphosphate kinase 3 (EC 2.7.4.21) (InsP6 kinase 3) (Inositol hexakisphosphate kinase 3). | |||||
|
IP6K3_MOUSE
|
||||||
| NC score | 0.062905 (rank : 9) | θ value | θ > 10 (rank : 52) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BWD2 | Gene names | Ihpk3, Ip6k3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol hexaphosphate kinase 3 (EC 2.7.4.21) (InsP6 kinase 3) (Inositol hexakisphosphate kinase 3). | |||||
|
IP6K2_HUMAN
|
||||||
| NC score | 0.050071 (rank : 10) | θ value | θ > 10 (rank : 51) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UHH9, Q6P0N8, Q9BSZ6, Q9BUW3, Q9H4P7, Q9NT63, Q9UFU6 | Gene names | IHPK2, IP6K2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inositol hexakisphosphate kinase 2 (EC 2.7.4.21) (InsP6 kinase 2) (Inositol hexakisphosphate kinase 2) (P(i)-uptake stimulator) (PiUS). | |||||
|
TR150_MOUSE
|
||||||
| NC score | 0.041037 (rank : 11) | θ value | 0.0736092 (rank : 9) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 468 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q569Z6 | Gene names | Thrap3, Trap150 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid hormone receptor-associated protein 3 (Thyroid hormone receptor-associated protein complex 150 kDa component) (Trap150). | |||||
|
LRC42_HUMAN
|
||||||
| NC score | 0.038495 (rank : 12) | θ value | 0.62314 (rank : 13) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y546, Q8N2Q8 | Gene names | LRRC42 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat-containing protein 42. | |||||
|
PMS2_HUMAN
|
||||||
| NC score | 0.036845 (rank : 13) | θ value | 2.36792 (rank : 23) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P54278 | Gene names | PMS2, PMSL2 | |||
|
Domain Architecture |

|
|||||
| Description | PMS1 protein homolog 2 (DNA mismatch repair protein PMS2). | |||||
|
CONA1_HUMAN
|
||||||
| NC score | 0.027370 (rank : 14) | θ value | 0.0431538 (rank : 7) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 281 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q86Y22, Q8IVR4, Q9NT93 | Gene names | COL23A1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XXIII) chain. | |||||
|
SRRM2_HUMAN
|
||||||
| NC score | 0.027318 (rank : 15) | θ value | 6.88961 (rank : 41) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
NSBP1_MOUSE
|
||||||
| NC score | 0.027159 (rank : 16) | θ value | 0.47712 (rank : 12) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9JL35, O88832, Q8VC71, Q9CUW1 | Gene names | Nsbp1, Garp45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1 (Nucleosome-binding protein 45) (NBP-45) (GARP45 protein). | |||||
|
MTMR3_HUMAN
|
||||||
| NC score | 0.026703 (rank : 17) | θ value | 0.0736092 (rank : 8) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q13615, Q9NYN5, Q9NYN6, Q9UDX6, Q9UEG3 | Gene names | MTMR3, KIAA0371, ZFYVE10 | |||
|
Domain Architecture |

|
|||||
| Description | Myotubularin-related protein 3 (EC 3.1.3.48) (FYVE domain-containing dual specificity protein phosphatase 1) (FYVE-DSP1) (Zinc finger FYVE domain-containing protein 10). | |||||
|
SRTD2_MOUSE
|
||||||
| NC score | 0.025793 (rank : 18) | θ value | 6.88961 (rank : 42) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9JJG5, Q8C609, Q91WL3, Q925E5 | Gene names | Sertad2, Kiaa0127 | |||
|
Domain Architecture |

|
|||||
| Description | SERTA domain-containing protein 2 (Transcriptional regulator interacting with the PHD-bromodomain 2) (TRIP-Br2). | |||||
|
DEND_HUMAN
|
||||||
| NC score | 0.023911 (rank : 19) | θ value | 3.0926 (rank : 24) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O94850 | Gene names | DDN, KIAA0749 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dendrin. | |||||
|
SSPO_MOUSE
|
||||||
| NC score | 0.023580 (rank : 20) | θ value | 0.0252991 (rank : 6) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 800 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8CG65 | Gene names | Sspo | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SCO-spondin precursor. | |||||
|
CONA1_MOUSE
|
||||||
| NC score | 0.023412 (rank : 21) | θ value | 1.81305 (rank : 18) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 254 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8K4G2, Q5SUQ0 | Gene names | Col23a1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XXIII) chain. | |||||
|
CO2A1_HUMAN
|
||||||
| NC score | 0.021868 (rank : 22) | θ value | 1.81305 (rank : 17) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 486 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P02458, Q12985, Q14009, Q14044, Q14046, Q14056, Q14058, Q16672, Q6LBY1, Q6LBY2, Q6LBY3, Q99227, Q9UE38, Q9UE39, Q9UE40, Q9UE41, Q9UE42, Q9UE43 | Gene names | COL2A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(II) chain precursor [Contains: Chondrocalcin]. | |||||
|
ZN385_HUMAN
|
||||||
| NC score | 0.021512 (rank : 23) | θ value | 1.81305 (rank : 21) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 614 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96PM9, Q5VH53, Q9H7R6, Q9UFU3 | Gene names | ZNF385, HZF, RZF | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 385 (Hematopoietic zinc finger protein) (Retinal zinc finger protein). | |||||
|
TP53B_MOUSE
|
||||||
| NC score | 0.020758 (rank : 24) | θ value | 1.81305 (rank : 20) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 644 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P70399, Q68FD0, Q8CI97, Q91YC9 | Gene names | Tp53bp1, Trp53bp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tumor suppressor p53-binding protein 1 (p53-binding protein 1) (p53BP1) (53BP1). | |||||
|
CO2A1_MOUSE
|
||||||
| NC score | 0.020165 (rank : 25) | θ value | 8.99809 (rank : 43) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 483 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P28481 | Gene names | Col2a1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(II) chain precursor [Contains: Chondrocalcin]. | |||||
|
EMIL1_HUMAN
|
||||||
| NC score | 0.015951 (rank : 26) | θ value | 5.27518 (rank : 27) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 239 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9Y6C2, Q96G58, Q9UG76 | Gene names | EMILIN1, EMI | |||
|
Domain Architecture |

|
|||||
| Description | EMILIN-1 precursor (Elastin microfibril interface-located protein 1) (Elastin microfibril interfacer 1). | |||||
|
DEPP_HUMAN
|
||||||
| NC score | 0.015883 (rank : 27) | θ value | 8.99809 (rank : 44) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NTK1, O94997, Q5T735, Q76MX8 | Gene names | DEPP, C10orf10, FIG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein DEPP (Decidual protein induced by progesterone) (Fasting- induced gene protein) (FIG). | |||||
|
ARHG1_HUMAN
|
||||||
| NC score | 0.015716 (rank : 28) | θ value | 1.38821 (rank : 15) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q92888, O00513, Q8N4J4, Q96BF4, Q96F17, Q9BSB1 | Gene names | ARHGEF1 | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 1 (p115-RhoGEF) (p115RhoGEF) (115 kDa guanine nucleotide exchange factor) (Sub1.5). | |||||
|
SORC3_HUMAN
|
||||||
| NC score | 0.014729 (rank : 29) | θ value | 1.81305 (rank : 19) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UPU3, Q5VXF9, Q9NQJ2 | Gene names | SORCS3, KIAA1059 | |||
|
Domain Architecture |

|
|||||
| Description | VPS10 domain-containing receptor SorCS3 precursor. | |||||
|
DIDO1_HUMAN
|
||||||
| NC score | 0.013646 (rank : 30) | θ value | 6.88961 (rank : 33) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 960 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9BTC0, O15043, Q3ZTL7, Q3ZTL8, Q4VXS1, Q4VXS2, Q96D72, Q9BQW0, Q9BW03, Q9H4G6, Q9H4G7, Q9NTU8, Q9NUM8, Q9UFB6 | Gene names | DIDO1, C20orf158, DATF1, KIAA0333 | |||
|
Domain Architecture |

|
|||||
| Description | Death-inducer obliterator 1 (DIO-1) (Death-associated transcription factor 1) (DATF-1) (hDido1). | |||||
|
LPIN3_HUMAN
|
||||||
| NC score | 0.012693 (rank : 31) | θ value | 6.88961 (rank : 36) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9BQK8, Q5TDB9, Q9NPY8, Q9UJE5 | Gene names | LPIN3, LIPN3L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lipin-3 (Lipin 3-like). | |||||
|
LPIN3_MOUSE
|
||||||
| NC score | 0.011895 (rank : 32) | θ value | 8.99809 (rank : 45) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99PI4, Q3TQ75, Q8C7R9 | Gene names | Lpin3 | |||
|
Domain Architecture |

|
|||||
| Description | Lipin-3. | |||||
|
RNF17_MOUSE
|
||||||
| NC score | 0.011229 (rank : 33) | θ value | 8.99809 (rank : 49) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99MV7 | Gene names | Rnf17 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 17. | |||||
|
NFH_MOUSE
|
||||||
| NC score | 0.011128 (rank : 34) | θ value | 5.27518 (rank : 30) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
FGF3_MOUSE
|
||||||
| NC score | 0.011120 (rank : 35) | θ value | 3.0926 (rank : 25) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P05524, Q61736 | Gene names | Fgf3, Fgf-3, Int-2 | |||
|
Domain Architecture |

|
|||||
| Description | INT-2 proto-oncogene protein precursor (Fibroblast growth factor 3) (FGF-3) (HBGF-3). | |||||
|
FUBP2_HUMAN
|
||||||
| NC score | 0.010691 (rank : 36) | θ value | 6.88961 (rank : 35) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q92945, O00301, Q9UNT5, Q9UQH5 | Gene names | KHSRP, FUBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Far upstream element-binding protein 2 (FUSE-binding protein 2) (KH type-splicing regulatory protein) (KSRP) (p75). | |||||
|
PRGC2_HUMAN
|
||||||
| NC score | 0.010664 (rank : 37) | θ value | 8.99809 (rank : 46) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 272 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q86YN6, Q86YN3, Q86YN4, Q86YN5, Q8N1N9, Q8TDE4, Q8TDE5 | Gene names | PPARGC1B, PERC, PGC1, PGC1B, PPARGC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor gamma coactivator 1-beta (PPAR gamma coactivator-1beta) (PGC-1-related estrogen receptor alpha coactivator) (PPARGC-1-beta) (PGC-1-beta). | |||||
|
TARA_MOUSE
|
||||||
| NC score | 0.010276 (rank : 38) | θ value | 5.27518 (rank : 31) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1133 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q99KW3, Q2PZW9, Q2Q402, Q2Q403, Q2Q404, Q6ZPK4, Q8C6T3, Q8CG90 | Gene names | Triobp, Kiaa1662, Tara | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
LUZP1_MOUSE
|
||||||
| NC score | 0.010264 (rank : 39) | θ value | 2.36792 (rank : 22) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 940 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8R4U7, Q3UV18, Q7TS71, Q8BQW1, Q99NG3, Q9CSL6 | Gene names | Luzp1, Luzp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine zipper protein 1 (Leucine zipper motif-containing protein). | |||||
|
SHC3_MOUSE
|
||||||
| NC score | 0.010260 (rank : 40) | θ value | 6.88961 (rank : 39) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q61120 | Gene names | Shc3, Nshc, ShcC | |||
|
Domain Architecture |

|
|||||
| Description | SHC-transforming protein 3 (SH2 domain protein C3) (Src homology 2 domain-containing-transforming protein C3) (Neuronal Shc) (N-Shc). | |||||
|
MDC1_MOUSE
|
||||||
| NC score | 0.009924 (rank : 41) | θ value | 5.27518 (rank : 29) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 998 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q5PSV9, Q5U4D3, Q6ZQH7 | Gene names | Mdc1, Kiaa0170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1. | |||||
|
SHC3_HUMAN
|
||||||
| NC score | 0.009564 (rank : 42) | θ value | 8.99809 (rank : 50) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q92529, Q8TAP2, Q9UCX5 | Gene names | SHC3, NSHC, SHCC | |||
|
Domain Architecture |

|
|||||
| Description | SHC-transforming protein 3 (SH2 domain protein C3) (Src homology 2 domain-containing-transforming protein C3) (Neuronal Shc) (N-Shc) (Protein Rai). | |||||
|
RGS3_HUMAN
|
||||||
| NC score | 0.009391 (rank : 43) | θ value | 8.99809 (rank : 48) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
AP3B2_MOUSE
|
||||||
| NC score | 0.008835 (rank : 44) | θ value | 6.88961 (rank : 32) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9JME5, Q6QR53, Q8R1E5 | Gene names | Ap3b2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AP-3 complex subunit beta-2 (Adapter-related protein complex 3 beta-2 subunit) (Beta3B-adaptin) (Adaptor protein complex AP-3 beta-2 subunit) (Clathrin assembly protein complex 3 beta-2 large chain). | |||||
|
JADE3_MOUSE
|
||||||
| NC score | 0.008296 (rank : 45) | θ value | 5.27518 (rank : 28) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6IE82, Q6ZQG2 | Gene names | Phf16, Jade3, Kiaa0215 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Jade-3 (PHD finger protein 16). | |||||
|
SIX5_HUMAN
|
||||||
| NC score | 0.006886 (rank : 46) | θ value | 6.88961 (rank : 40) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 386 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8N196 | Gene names | SIX5, DMAHP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Homeobox protein SIX5 (DM locus-associated homeodomain protein). | |||||
|
SCND1_HUMAN
|
||||||
| NC score | 0.006803 (rank : 47) | θ value | 6.88961 (rank : 38) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P57086 | Gene names | SCAND1, SDP1 | |||
|
Domain Architecture |

|
|||||
| Description | SCAN domain-containing protein 1. | |||||
|
PCD18_HUMAN
|
||||||
| NC score | 0.005743 (rank : 48) | θ value | 1.06291 (rank : 14) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9HCL0 | Gene names | PCDH18, KIAA1562 | |||
|
Domain Architecture |

|
|||||
| Description | Protocadherin-18 precursor. | |||||
|
CSPG3_HUMAN
|
||||||
| NC score | 0.005257 (rank : 49) | θ value | 4.03905 (rank : 26) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O14594, Q9UPK6 | Gene names | CSPG3, NCAN, NEUR | |||
|
Domain Architecture |

|
|||||
| Description | Neurocan core protein precursor (Chondroitin sulfate proteoglycan 3). | |||||
|
PINK1_HUMAN
|
||||||
| NC score | 0.003801 (rank : 50) | θ value | 6.88961 (rank : 37) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 741 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9BXM7, Q8N6T9, Q8NBU3, Q96DE4 | Gene names | PINK1 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase PINK1, mitochondrial precursor (EC 2.7.11.1) (PTEN-induced putative kinase protein 1) (BRPK). | |||||
|
RFXK_HUMAN
|
||||||
| NC score | 0.003233 (rank : 51) | θ value | 8.99809 (rank : 47) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O14593, O95839 | Gene names | RFXANK, ANKRA1, RFXB | |||
|
Domain Architecture |

|
|||||
| Description | DNA-binding protein RFXANK (Regulatory factor X subunit B) (RFX-B) (Ankyrin repeat family A protein 1). | |||||
|
DMD_MOUSE
|
||||||
| NC score | 0.002789 (rank : 52) | θ value | 6.88961 (rank : 34) | |||
| Query Neighborhood Hits | 50 | Target Neighborhood Hits | 1111 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P11531, O35653, Q60703 | Gene names | Dmd | |||
|
Domain Architecture |

|
|||||
| Description | Dystrophin. | |||||