Please be patient as the page loads
|
HYAL3_HUMAN
|
||||||
| SwissProt Accessions | O43820, O60540, Q8NFK2, Q8NFK3, Q8NFK4, Q96E56, Q9BRW9 | Gene names | HYAL3, LUCA3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hyaluronidase-3 precursor (EC 3.2.1.35) (Hyal-3) (Hyaluronoglucosaminidase-3) (LUCA-3). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
HYAL3_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O43820, O60540, Q8NFK2, Q8NFK3, Q8NFK4, Q96E56, Q9BRW9 | Gene names | HYAL3, LUCA3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hyaluronidase-3 precursor (EC 3.2.1.35) (Hyal-3) (Hyaluronoglucosaminidase-3) (LUCA-3). | |||||
|
HYAL3_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.996231 (rank : 2) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8VEI3, Q8VBX7, Q8VI77 | Gene names | Hyal3, Hyl3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hyaluronidase-3 precursor (EC 3.2.1.35) (Hyal-3) (Hyaluronoglucosaminidase-3). | |||||
|
HYAL2_MOUSE
|
||||||
| θ value | 1.07224e-85 (rank : 3) | NC score | 0.957672 (rank : 6) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O35632, O35631, Q99MS9, Q99MT0 | Gene names | Hyal2 | |||
|
Domain Architecture |

|
|||||
| Description | Hyaluronidase-2 precursor (EC 3.2.1.35) (Hyal-2) (Hyaluronoglucosaminidase-2) (LUCA-2). | |||||
|
HYAL2_HUMAN
|
||||||
| θ value | 3.22841e-82 (rank : 4) | NC score | 0.957724 (rank : 5) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q12891, O15177, Q9BW29 | Gene names | HYAL2, LUCA2 | |||
|
Domain Architecture |

|
|||||
| Description | Hyaluronidase-2 precursor (EC 3.2.1.35) (Hyal-2) (Hyaluronoglucosaminidase-2) (LUCA-2). | |||||
|
HYAL1_MOUSE
|
||||||
| θ value | 5.33382e-77 (rank : 5) | NC score | 0.959102 (rank : 4) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q91ZJ9, O70229, Q8CE62, Q8QZX3, Q8VBW7, Q8VDK0 | Gene names | Hyal1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hyaluronidase-1 precursor (EC 3.2.1.35) (Hyal-1) (Hyaluronoglucosaminidase-1). | |||||
|
HYAL1_HUMAN
|
||||||
| θ value | 1.5519e-76 (rank : 6) | NC score | 0.959222 (rank : 3) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q12794, Q6FH23, Q6PIZ6, Q7KYU2, Q7LE34, Q8NFK5, Q8NFK6, Q8NFK7, Q8NFK8, Q8NFK9, Q93013, Q9UKD5, Q9UNI8 | Gene names | HYAL1, LUCA1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hyaluronidase-1 precursor (EC 3.2.1.35) (Hyal-1) (Hyaluronoglucosaminidase-1) (LUCA-1). | |||||
|
HYALP_HUMAN
|
||||||
| θ value | 5.34618e-69 (rank : 7) | NC score | 0.949741 (rank : 7) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P38567 | Gene names | SPAM1, HYAL3, PH20 | |||
|
Domain Architecture |

|
|||||
| Description | Hyaluronidase PH-20 precursor (EC 3.2.1.35) (Hyal-PH20) (Sperm surface protein PH-20) (Sperm adhesion molecule 1). | |||||
|
HYALP_MOUSE
|
||||||
| θ value | 3.83135e-67 (rank : 8) | NC score | 0.947039 (rank : 8) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P48794, Q7TSD6, Q812F4, Q9DAQ1 | Gene names | Spam1, Ph20, Spam | |||
|
Domain Architecture |

|
|||||
| Description | Hyaluronidase PH-20 precursor (EC 3.2.1.35) (Hyal-PH20) (Sperm surface protein PH-20) (Sperm adhesion molecule 1). | |||||
|
NAGPA_MOUSE
|
||||||
| θ value | 0.00665767 (rank : 9) | NC score | 0.095595 (rank : 9) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 308 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8BJ48, Q3UUT5, Q8CHQ8, Q9QZE6 | Gene names | Nagpa | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetylglucosamine-1-phosphodiester alpha-N-acetylglucosaminidase precursor (EC 3.1.4.45) (Phosphodiester alpha-GlcNAcase) (Mannose 6- phosphate-uncovering enzyme). | |||||
|
JAG2_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 10) | NC score | 0.067651 (rank : 12) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9QYE5, O55139, O70219 | Gene names | Jag2 | |||
|
Domain Architecture |

|
|||||
| Description | Jagged-2 precursor (Jagged2). | |||||
|
JAG2_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 11) | NC score | 0.068052 (rank : 11) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9Y219, Q9UE17, Q9UE99, Q9UNK8, Q9Y6P9, Q9Y6Q0 | Gene names | JAG2 | |||
|
Domain Architecture |

|
|||||
| Description | Jagged-2 precursor (Jagged2) (HJ2). | |||||
|
NAGPA_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 12) | NC score | 0.091502 (rank : 10) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9UK23, Q96EJ8 | Gene names | NAGPA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetylglucosamine-1-phosphodiester alpha-N-acetylglucosaminidase precursor (EC 3.1.4.45) (Phosphodiester alpha-GlcNAcase) (Mannose 6- phosphate-uncovering enzyme). | |||||
|
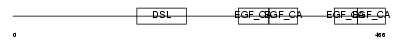
DLL1_MOUSE
|
||||||
| θ value | 0.279714 (rank : 13) | NC score | 0.056753 (rank : 18) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 456 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q61483 | Gene names | Dll1 | |||
|
Domain Architecture |
|
|||||
| Description | Delta-like protein 1 precursor (Drosophila Delta homolog 1) (Delta1). | |||||
|
JAG1_MOUSE
|
||||||
| θ value | 0.47712 (rank : 14) | NC score | 0.061876 (rank : 13) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 518 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9QXX0 | Gene names | Jag1 | |||
|
Domain Architecture |

|
|||||
| Description | Jagged-1 precursor (Jagged1) (CD339 antigen). | |||||
|
CRUM1_MOUSE
|
||||||
| θ value | 0.62314 (rank : 15) | NC score | 0.057524 (rank : 15) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 444 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8VHS2, Q6ST50, Q71JF2, Q8BGR4 | Gene names | Crb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Crumbs homolog 1 precursor. | |||||
|
JAG1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 16) | NC score | 0.061821 (rank : 14) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 514 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P78504, O14902, O15122, Q15816 | Gene names | JAG1, JAGL1 | |||
|
Domain Architecture |

|
|||||
| Description | Jagged-1 precursor (Jagged1) (hJ1) (CD339 antigen). | |||||
|
TENA_HUMAN
|
||||||
| θ value | 0.62314 (rank : 17) | NC score | 0.046260 (rank : 22) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 631 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P24821, Q14583, Q15567 | Gene names | TNC, HXB | |||
|
Domain Architecture |

|
|||||
| Description | Tenascin precursor (TN) (Tenascin-C) (TN-C) (Hexabrachion) (Cytotactin) (Neuronectin) (GMEM) (JI) (Myotendinous antigen) (Glioma- associated-extracellular matrix antigen) (GP 150-225). | |||||
|
ZAN_MOUSE
|
||||||
| θ value | 1.06291 (rank : 18) | NC score | 0.040087 (rank : 31) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 808 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O88799, O08647 | Gene names | Zan | |||
|
Domain Architecture |

|
|||||
| Description | Zonadhesin precursor. | |||||
|
CRUM2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 19) | NC score | 0.057138 (rank : 17) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 453 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q5IJ48, Q5JS41, Q5JS43, Q6ZTA9, Q6ZWI6 | Gene names | CRB2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Crumbs homolog 2 precursor (Crumbs-like protein 2). | |||||
|
NOTC1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 20) | NC score | 0.047019 (rank : 21) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 916 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q01705, Q06007, Q61905, Q99JC2, Q9QW58, Q9R0X7 | Gene names | Notch1, Motch | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 1 precursor (Notch 1) (Motch A) (mT14) (p300) [Contains: Notch 1 extracellular truncation; Notch 1 intracellular domain]. | |||||
|
NOTC4_MOUSE
|
||||||
| θ value | 2.36792 (rank : 21) | NC score | 0.045170 (rank : 24) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 872 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P31695, O35442, O88314, O88316, Q62389, Q62390, Q9R1W9, Q9R1X0 | Gene names | Notch4, Int-3, Int3 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 4 precursor (Notch 4) [Contains: Transforming protein Int-3; Notch 4 extracellular truncation; Notch 4 intracellular domain]. | |||||
|
STAB1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 22) | NC score | 0.041374 (rank : 27) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 546 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9NY15, Q8IUH0, Q8IUH1, Q93072 | Gene names | STAB1, FEEL1, KIAA0246 | |||
|
Domain Architecture |

|
|||||
| Description | Stabilin-1 precursor (FEEL-1 protein) (MS-1 antigen). | |||||
|
CELR2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 23) | NC score | 0.022734 (rank : 39) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 657 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9HCU4, Q92566 | Gene names | CELSR2, CDHF10, EGFL2, KIAA0279, MEGF3 | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin EGF LAG seven-pass G-type receptor 2 precursor (Epidermal growth factor-like 2) (Multiple epidermal growth factor-like domains 3) (Flamingo 1). | |||||
|
CELR2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 24) | NC score | 0.023439 (rank : 37) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 584 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9R0M0, Q99K26, Q9Z2R4 | Gene names | Celsr2 | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin EGF LAG seven-pass G-type receptor 2 precursor (Flamingo 1) (mFmi1). | |||||
|
CRUM1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 25) | NC score | 0.057495 (rank : 16) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 457 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P82279, Q5K3A6, Q5TC28, Q5VUT1, Q6N027, Q8WWY0 | Gene names | CRB1 | |||
|
Domain Architecture |

|
|||||
| Description | Crumbs homolog 1 precursor. | |||||
|
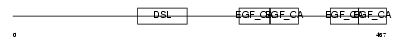
DLL1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 26) | NC score | 0.052809 (rank : 19) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O00548, Q9NU41, Q9UJV2 | Gene names | DLL1 | |||
|
Domain Architecture |
|
|||||
| Description | Delta-like protein 1 precursor (Drosophila Delta homolog 1) (Delta1) (H-Delta-1). | |||||
|
NOTC3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 27) | NC score | 0.044442 (rank : 25) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 1062 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9UM47, Q9UEB3, Q9UPL3, Q9Y6L8 | Gene names | NOTCH3 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 3 precursor (Notch 3) [Contains: Notch 3 extracellular truncation; Notch 3 intracellular domain]. | |||||
|
NOTC3_MOUSE
|
||||||
| θ value | 3.0926 (rank : 28) | NC score | 0.045728 (rank : 23) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q61982 | Gene names | Notch3 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 3 precursor (Notch 3) [Contains: Notch 3 extracellular truncation; Notch 3 intracellular domain]. | |||||
|
ADA19_MOUSE
|
||||||
| θ value | 4.03905 (rank : 29) | NC score | 0.022739 (rank : 38) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O35674 | Gene names | Adam19, Mltnb | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 19 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 19) (Meltrin beta). | |||||
|
ADA30_HUMAN
|
||||||
| θ value | 4.03905 (rank : 30) | NC score | 0.017778 (rank : 43) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 391 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9UKF2, Q9UKF1 | Gene names | ADAM30 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 30 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 30). | |||||
|
LYAM2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 31) | NC score | 0.028033 (rank : 35) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 420 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q00690 | Gene names | Sele, Elam-1 | |||
|
Domain Architecture |

|
|||||
| Description | E-selectin precursor (Endothelial leukocyte adhesion molecule 1) (ELAM-1) (Leukocyte-endothelial cell adhesion molecule 2) (LECAM2) (CD62E antigen). | |||||
|
SLIT1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 32) | NC score | 0.034434 (rank : 32) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 700 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O75093, Q8WWZ2, Q9UIL7 | Gene names | SLIT1, KIAA0813, MEGF4, SLIL1 | |||
|
Domain Architecture |

|
|||||
| Description | Slit homolog 1 protein precursor (Slit-1) (Multiple epidermal growth factor-like domains 4). | |||||
|
ADA19_HUMAN
|
||||||
| θ value | 5.27518 (rank : 33) | NC score | 0.020313 (rank : 42) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 381 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9H013, Q9BZL5, Q9UHP2 | Gene names | ADAM19, MLTNB | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 19 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 19) (Meltrin beta) (Metalloprotease and disintegrin dentritic antigen marker) (MADDAM). | |||||
|
ADAM9_HUMAN
|
||||||
| θ value | 5.27518 (rank : 34) | NC score | 0.021307 (rank : 41) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q13443, Q10718, Q8NFM6 | Gene names | ADAM9, KIAA0021, MCMP, MDC9, MLTNG | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 9 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 9) (Metalloprotease/disintegrin/cysteine-rich protein 9) (Myeloma cell metalloproteinase) (Meltrin gamma) (Cellular disintegrin-related protein). | |||||
|
ADAM8_HUMAN
|
||||||
| θ value | 6.88961 (rank : 35) | NC score | 0.023894 (rank : 36) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 327 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P78325 | Gene names | ADAM8, MS2 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 8 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 8) (Cell surface antigen MS2) (CD156a antigen) (CD156). | |||||
|
CELR1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 36) | NC score | 0.021731 (rank : 40) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 613 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9NYQ6, O95722, Q5TH47, Q9BWQ5, Q9Y506, Q9Y526 | Gene names | CELSR1, CDHF9, FMI2 | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin EGF LAG seven-pass G-type receptor 1 precursor (Flamingo homolog 2) (hFmi2). | |||||
|
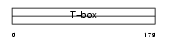
EOMES_MOUSE
|
||||||
| θ value | 6.88961 (rank : 37) | NC score | 0.004900 (rank : 46) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O54839, Q9QYG7 | Gene names | Eomes, Tbr2 | |||
|
Domain Architecture |
|
|||||
| Description | Eomesodermin homolog. | |||||
|
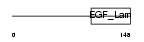
MEGF8_HUMAN
|
||||||
| θ value | 6.88961 (rank : 38) | NC score | 0.041678 (rank : 26) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 528 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q7Z7M0, O75097 | Gene names | MEGF8, EGFL4, KIAA0817 | |||
|
Domain Architecture |
|
|||||
| Description | Multiple epidermal growth factor-like domains 8 (EGF-like domain- containing protein 4) (Multiple EGF-like domain protein 4). | |||||
|
MEGF8_MOUSE
|
||||||
| θ value | 6.88961 (rank : 39) | NC score | 0.041044 (rank : 28) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 520 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P60882, Q80TR3, Q80V41, Q8BMN9, Q8JZW7, Q8K0J3 | Gene names | Megf8, Egfl4 | |||
|
Domain Architecture |

|
|||||
| Description | Multiple epidermal growth factor-like domains 8 (EGF-like domain- containing protein 4) (Multiple EGF-like domain protein 4). | |||||
|
SLIT1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 40) | NC score | 0.033738 (rank : 33) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 715 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q80TR4, Q9WVB5 | Gene names | Slit1, Kiaa0813 | |||
|
Domain Architecture |

|
|||||
| Description | Slit homolog 1 protein precursor (Slit-1). | |||||
|
TENX_HUMAN
|
||||||
| θ value | 6.88961 (rank : 41) | NC score | 0.040559 (rank : 30) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 789 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P22105, P78530, P78531, Q08424, Q9UMG7 | Gene names | TNXB, HXBL, TNX, XB | |||
|
Domain Architecture |

|
|||||
| Description | Tenascin-X precursor (TN-X) (Hexabrachion-like protein). | |||||
|
MEGF6_HUMAN
|
||||||
| θ value | 8.99809 (rank : 42) | NC score | 0.040562 (rank : 29) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O75095, Q5VV39 | Gene names | MEGF6, EGFL3, KIAA0815 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Multiple epidermal growth factor-like domains 6 precursor (EGF-like domain-containing protein 3) (Multiple EGF-like domain protein 3). | |||||
|
RAD54_HUMAN
|
||||||
| θ value | 8.99809 (rank : 43) | NC score | 0.006081 (rank : 45) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q92698, Q6IUY3 | Gene names | RAD54L, RAD54A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair and recombination protein RAD54-like (EC 3.6.1.-) (RAD54 homolog) (hRAD54) (hHR54). | |||||
|
RAD54_MOUSE
|
||||||
| θ value | 8.99809 (rank : 44) | NC score | 0.006125 (rank : 44) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P70270, Q8BSR5, Q8C2C4, Q8K3D4 | Gene names | Rad54l, Rad54 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair and recombination protein RAD54-like (EC 3.6.1.-) (RAD54 homolog) (mRAD54) (mHR54). | |||||
|
SLIT3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 45) | NC score | 0.032954 (rank : 34) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 682 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O75094, O95804, Q9UFH5 | Gene names | SLIT3, KIAA0814, MEGF5, SLIL2 | |||
|
Domain Architecture |

|
|||||
| Description | Slit homolog 3 protein precursor (Slit-3) (Multiple epidermal growth factor-like domains 5). | |||||
|
DNER_HUMAN
|
||||||
| θ value | θ > 10 (rank : 46) | NC score | 0.050272 (rank : 20) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 460 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8NFT8, Q53R88, Q53TP7, Q53TQ5, Q8IYT0, Q8TB42, Q9NTF1, Q9UDM2 | Gene names | DNER, BET | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Delta and Notch-like epidermal growth factor-related receptor precursor. | |||||
|
HYAL3_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O43820, O60540, Q8NFK2, Q8NFK3, Q8NFK4, Q96E56, Q9BRW9 | Gene names | HYAL3, LUCA3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hyaluronidase-3 precursor (EC 3.2.1.35) (Hyal-3) (Hyaluronoglucosaminidase-3) (LUCA-3). | |||||
|
HYAL3_MOUSE
|
||||||
| NC score | 0.996231 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8VEI3, Q8VBX7, Q8VI77 | Gene names | Hyal3, Hyl3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hyaluronidase-3 precursor (EC 3.2.1.35) (Hyal-3) (Hyaluronoglucosaminidase-3). | |||||
|
HYAL1_HUMAN
|
||||||
| NC score | 0.959222 (rank : 3) | θ value | 1.5519e-76 (rank : 6) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q12794, Q6FH23, Q6PIZ6, Q7KYU2, Q7LE34, Q8NFK5, Q8NFK6, Q8NFK7, Q8NFK8, Q8NFK9, Q93013, Q9UKD5, Q9UNI8 | Gene names | HYAL1, LUCA1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hyaluronidase-1 precursor (EC 3.2.1.35) (Hyal-1) (Hyaluronoglucosaminidase-1) (LUCA-1). | |||||
|
HYAL1_MOUSE
|
||||||
| NC score | 0.959102 (rank : 4) | θ value | 5.33382e-77 (rank : 5) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q91ZJ9, O70229, Q8CE62, Q8QZX3, Q8VBW7, Q8VDK0 | Gene names | Hyal1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hyaluronidase-1 precursor (EC 3.2.1.35) (Hyal-1) (Hyaluronoglucosaminidase-1). | |||||
|
HYAL2_HUMAN
|
||||||
| NC score | 0.957724 (rank : 5) | θ value | 3.22841e-82 (rank : 4) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q12891, O15177, Q9BW29 | Gene names | HYAL2, LUCA2 | |||
|
Domain Architecture |

|
|||||
| Description | Hyaluronidase-2 precursor (EC 3.2.1.35) (Hyal-2) (Hyaluronoglucosaminidase-2) (LUCA-2). | |||||
|
HYAL2_MOUSE
|
||||||
| NC score | 0.957672 (rank : 6) | θ value | 1.07224e-85 (rank : 3) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O35632, O35631, Q99MS9, Q99MT0 | Gene names | Hyal2 | |||
|
Domain Architecture |

|
|||||
| Description | Hyaluronidase-2 precursor (EC 3.2.1.35) (Hyal-2) (Hyaluronoglucosaminidase-2) (LUCA-2). | |||||
|
HYALP_HUMAN
|
||||||
| NC score | 0.949741 (rank : 7) | θ value | 5.34618e-69 (rank : 7) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P38567 | Gene names | SPAM1, HYAL3, PH20 | |||
|
Domain Architecture |

|
|||||
| Description | Hyaluronidase PH-20 precursor (EC 3.2.1.35) (Hyal-PH20) (Sperm surface protein PH-20) (Sperm adhesion molecule 1). | |||||
|
HYALP_MOUSE
|
||||||
| NC score | 0.947039 (rank : 8) | θ value | 3.83135e-67 (rank : 8) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P48794, Q7TSD6, Q812F4, Q9DAQ1 | Gene names | Spam1, Ph20, Spam | |||
|
Domain Architecture |

|
|||||
| Description | Hyaluronidase PH-20 precursor (EC 3.2.1.35) (Hyal-PH20) (Sperm surface protein PH-20) (Sperm adhesion molecule 1). | |||||
|
NAGPA_MOUSE
|
||||||
| NC score | 0.095595 (rank : 9) | θ value | 0.00665767 (rank : 9) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 308 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8BJ48, Q3UUT5, Q8CHQ8, Q9QZE6 | Gene names | Nagpa | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetylglucosamine-1-phosphodiester alpha-N-acetylglucosaminidase precursor (EC 3.1.4.45) (Phosphodiester alpha-GlcNAcase) (Mannose 6- phosphate-uncovering enzyme). | |||||
|
NAGPA_HUMAN
|
||||||
| NC score | 0.091502 (rank : 10) | θ value | 0.0961366 (rank : 12) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 282 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9UK23, Q96EJ8 | Gene names | NAGPA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetylglucosamine-1-phosphodiester alpha-N-acetylglucosaminidase precursor (EC 3.1.4.45) (Phosphodiester alpha-GlcNAcase) (Mannose 6- phosphate-uncovering enzyme). | |||||
|
JAG2_HUMAN
|
||||||
| NC score | 0.068052 (rank : 11) | θ value | 0.0961366 (rank : 11) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9Y219, Q9UE17, Q9UE99, Q9UNK8, Q9Y6P9, Q9Y6Q0 | Gene names | JAG2 | |||
|
Domain Architecture |

|
|||||
| Description | Jagged-2 precursor (Jagged2) (HJ2). | |||||
|
JAG2_MOUSE
|
||||||
| NC score | 0.067651 (rank : 12) | θ value | 0.0431538 (rank : 10) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9QYE5, O55139, O70219 | Gene names | Jag2 | |||
|
Domain Architecture |

|
|||||
| Description | Jagged-2 precursor (Jagged2). | |||||
|
JAG1_MOUSE
|
||||||
| NC score | 0.061876 (rank : 13) | θ value | 0.47712 (rank : 14) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 518 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9QXX0 | Gene names | Jag1 | |||
|
Domain Architecture |

|
|||||
| Description | Jagged-1 precursor (Jagged1) (CD339 antigen). | |||||
|
JAG1_HUMAN
|
||||||
| NC score | 0.061821 (rank : 14) | θ value | 0.62314 (rank : 16) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 514 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P78504, O14902, O15122, Q15816 | Gene names | JAG1, JAGL1 | |||
|
Domain Architecture |

|
|||||
| Description | Jagged-1 precursor (Jagged1) (hJ1) (CD339 antigen). | |||||
|
CRUM1_MOUSE
|
||||||
| NC score | 0.057524 (rank : 15) | θ value | 0.62314 (rank : 15) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 444 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8VHS2, Q6ST50, Q71JF2, Q8BGR4 | Gene names | Crb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Crumbs homolog 1 precursor. | |||||
|
CRUM1_HUMAN
|
||||||
| NC score | 0.057495 (rank : 16) | θ value | 3.0926 (rank : 25) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 457 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P82279, Q5K3A6, Q5TC28, Q5VUT1, Q6N027, Q8WWY0 | Gene names | CRB1 | |||
|
Domain Architecture |

|
|||||
| Description | Crumbs homolog 1 precursor. | |||||
|
CRUM2_HUMAN
|
||||||
| NC score | 0.057138 (rank : 17) | θ value | 1.81305 (rank : 19) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 453 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q5IJ48, Q5JS41, Q5JS43, Q6ZTA9, Q6ZWI6 | Gene names | CRB2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Crumbs homolog 2 precursor (Crumbs-like protein 2). | |||||
|
DLL1_MOUSE
|
||||||
| NC score | 0.056753 (rank : 18) | θ value | 0.279714 (rank : 13) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 456 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q61483 | Gene names | Dll1 | |||
|
Domain Architecture |

|
|||||
| Description | Delta-like protein 1 precursor (Drosophila Delta homolog 1) (Delta1). | |||||
|
DLL1_HUMAN
|
||||||
| NC score | 0.052809 (rank : 19) | θ value | 3.0926 (rank : 26) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O00548, Q9NU41, Q9UJV2 | Gene names | DLL1 | |||
|
Domain Architecture |

|
|||||
| Description | Delta-like protein 1 precursor (Drosophila Delta homolog 1) (Delta1) (H-Delta-1). | |||||
|
DNER_HUMAN
|
||||||
| NC score | 0.050272 (rank : 20) | θ value | θ > 10 (rank : 46) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 460 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q8NFT8, Q53R88, Q53TP7, Q53TQ5, Q8IYT0, Q8TB42, Q9NTF1, Q9UDM2 | Gene names | DNER, BET | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Delta and Notch-like epidermal growth factor-related receptor precursor. | |||||
|
NOTC1_MOUSE
|
||||||
| NC score | 0.047019 (rank : 21) | θ value | 2.36792 (rank : 20) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 916 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q01705, Q06007, Q61905, Q99JC2, Q9QW58, Q9R0X7 | Gene names | Notch1, Motch | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 1 precursor (Notch 1) (Motch A) (mT14) (p300) [Contains: Notch 1 extracellular truncation; Notch 1 intracellular domain]. | |||||
|
TENA_HUMAN
|
||||||
| NC score | 0.046260 (rank : 22) | θ value | 0.62314 (rank : 17) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 631 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P24821, Q14583, Q15567 | Gene names | TNC, HXB | |||
|
Domain Architecture |

|
|||||
| Description | Tenascin precursor (TN) (Tenascin-C) (TN-C) (Hexabrachion) (Cytotactin) (Neuronectin) (GMEM) (JI) (Myotendinous antigen) (Glioma- associated-extracellular matrix antigen) (GP 150-225). | |||||
|
NOTC3_MOUSE
|
||||||
| NC score | 0.045728 (rank : 23) | θ value | 3.0926 (rank : 28) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q61982 | Gene names | Notch3 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 3 precursor (Notch 3) [Contains: Notch 3 extracellular truncation; Notch 3 intracellular domain]. | |||||
|
NOTC4_MOUSE
|
||||||
| NC score | 0.045170 (rank : 24) | θ value | 2.36792 (rank : 21) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 872 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | P31695, O35442, O88314, O88316, Q62389, Q62390, Q9R1W9, Q9R1X0 | Gene names | Notch4, Int-3, Int3 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 4 precursor (Notch 4) [Contains: Transforming protein Int-3; Notch 4 extracellular truncation; Notch 4 intracellular domain]. | |||||
|
NOTC3_HUMAN
|
||||||
| NC score | 0.044442 (rank : 25) | θ value | 3.0926 (rank : 27) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 1062 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q9UM47, Q9UEB3, Q9UPL3, Q9Y6L8 | Gene names | NOTCH3 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 3 precursor (Notch 3) [Contains: Notch 3 extracellular truncation; Notch 3 intracellular domain]. | |||||
|
MEGF8_HUMAN
|
||||||
| NC score | 0.041678 (rank : 26) | θ value | 6.88961 (rank : 38) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 528 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q7Z7M0, O75097 | Gene names | MEGF8, EGFL4, KIAA0817 | |||
|
Domain Architecture |

|
|||||
| Description | Multiple epidermal growth factor-like domains 8 (EGF-like domain- containing protein 4) (Multiple EGF-like domain protein 4). | |||||
|
STAB1_HUMAN
|
||||||
| NC score | 0.041374 (rank : 27) | θ value | 2.36792 (rank : 22) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 546 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9NY15, Q8IUH0, Q8IUH1, Q93072 | Gene names | STAB1, FEEL1, KIAA0246 | |||
|
Domain Architecture |

|
|||||
| Description | Stabilin-1 precursor (FEEL-1 protein) (MS-1 antigen). | |||||
|
MEGF8_MOUSE
|
||||||
| NC score | 0.041044 (rank : 28) | θ value | 6.88961 (rank : 39) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 520 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | P60882, Q80TR3, Q80V41, Q8BMN9, Q8JZW7, Q8K0J3 | Gene names | Megf8, Egfl4 | |||
|
Domain Architecture |

|
|||||
| Description | Multiple epidermal growth factor-like domains 8 (EGF-like domain- containing protein 4) (Multiple EGF-like domain protein 4). | |||||
|
MEGF6_HUMAN
|
||||||
| NC score | 0.040562 (rank : 29) | θ value | 8.99809 (rank : 42) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O75095, Q5VV39 | Gene names | MEGF6, EGFL3, KIAA0815 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Multiple epidermal growth factor-like domains 6 precursor (EGF-like domain-containing protein 3) (Multiple EGF-like domain protein 3). | |||||
|
TENX_HUMAN
|
||||||
| NC score | 0.040559 (rank : 30) | θ value | 6.88961 (rank : 41) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 789 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P22105, P78530, P78531, Q08424, Q9UMG7 | Gene names | TNXB, HXBL, TNX, XB | |||
|
Domain Architecture |

|
|||||
| Description | Tenascin-X precursor (TN-X) (Hexabrachion-like protein). | |||||
|
ZAN_MOUSE
|
||||||
| NC score | 0.040087 (rank : 31) | θ value | 1.06291 (rank : 18) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 808 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | O88799, O08647 | Gene names | Zan | |||
|
Domain Architecture |

|
|||||
| Description | Zonadhesin precursor. | |||||
|
SLIT1_HUMAN
|
||||||
| NC score | 0.034434 (rank : 32) | θ value | 4.03905 (rank : 32) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 700 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | O75093, Q8WWZ2, Q9UIL7 | Gene names | SLIT1, KIAA0813, MEGF4, SLIL1 | |||
|
Domain Architecture |

|
|||||
| Description | Slit homolog 1 protein precursor (Slit-1) (Multiple epidermal growth factor-like domains 4). | |||||
|
SLIT1_MOUSE
|
||||||
| NC score | 0.033738 (rank : 33) | θ value | 6.88961 (rank : 40) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 715 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q80TR4, Q9WVB5 | Gene names | Slit1, Kiaa0813 | |||
|
Domain Architecture |

|
|||||
| Description | Slit homolog 1 protein precursor (Slit-1). | |||||
|
SLIT3_HUMAN
|
||||||
| NC score | 0.032954 (rank : 34) | θ value | 8.99809 (rank : 45) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 682 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O75094, O95804, Q9UFH5 | Gene names | SLIT3, KIAA0814, MEGF5, SLIL2 | |||
|
Domain Architecture |

|
|||||
| Description | Slit homolog 3 protein precursor (Slit-3) (Multiple epidermal growth factor-like domains 5). | |||||
|
LYAM2_MOUSE
|
||||||
| NC score | 0.028033 (rank : 35) | θ value | 4.03905 (rank : 31) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 420 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q00690 | Gene names | Sele, Elam-1 | |||
|
Domain Architecture |

|
|||||
| Description | E-selectin precursor (Endothelial leukocyte adhesion molecule 1) (ELAM-1) (Leukocyte-endothelial cell adhesion molecule 2) (LECAM2) (CD62E antigen). | |||||
|
ADAM8_HUMAN
|
||||||
| NC score | 0.023894 (rank : 36) | θ value | 6.88961 (rank : 35) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 327 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P78325 | Gene names | ADAM8, MS2 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 8 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 8) (Cell surface antigen MS2) (CD156a antigen) (CD156). | |||||
|
CELR2_MOUSE
|
||||||
| NC score | 0.023439 (rank : 37) | θ value | 3.0926 (rank : 24) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 584 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9R0M0, Q99K26, Q9Z2R4 | Gene names | Celsr2 | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin EGF LAG seven-pass G-type receptor 2 precursor (Flamingo 1) (mFmi1). | |||||
|
ADA19_MOUSE
|
||||||
| NC score | 0.022739 (rank : 38) | θ value | 4.03905 (rank : 29) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 315 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | O35674 | Gene names | Adam19, Mltnb | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 19 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 19) (Meltrin beta). | |||||
|
CELR2_HUMAN
|
||||||
| NC score | 0.022734 (rank : 39) | θ value | 3.0926 (rank : 23) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 657 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9HCU4, Q92566 | Gene names | CELSR2, CDHF10, EGFL2, KIAA0279, MEGF3 | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin EGF LAG seven-pass G-type receptor 2 precursor (Epidermal growth factor-like 2) (Multiple epidermal growth factor-like domains 3) (Flamingo 1). | |||||
|
CELR1_HUMAN
|
||||||
| NC score | 0.021731 (rank : 40) | θ value | 6.88961 (rank : 36) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 613 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9NYQ6, O95722, Q5TH47, Q9BWQ5, Q9Y506, Q9Y526 | Gene names | CELSR1, CDHF9, FMI2 | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin EGF LAG seven-pass G-type receptor 1 precursor (Flamingo homolog 2) (hFmi2). | |||||
|
ADAM9_HUMAN
|
||||||
| NC score | 0.021307 (rank : 41) | θ value | 5.27518 (rank : 34) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q13443, Q10718, Q8NFM6 | Gene names | ADAM9, KIAA0021, MCMP, MDC9, MLTNG | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 9 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 9) (Metalloprotease/disintegrin/cysteine-rich protein 9) (Myeloma cell metalloproteinase) (Meltrin gamma) (Cellular disintegrin-related protein). | |||||
|
ADA19_HUMAN
|
||||||
| NC score | 0.020313 (rank : 42) | θ value | 5.27518 (rank : 33) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 381 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9H013, Q9BZL5, Q9UHP2 | Gene names | ADAM19, MLTNB | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 19 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 19) (Meltrin beta) (Metalloprotease and disintegrin dentritic antigen marker) (MADDAM). | |||||
|
ADA30_HUMAN
|
||||||
| NC score | 0.017778 (rank : 43) | θ value | 4.03905 (rank : 30) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 391 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9UKF2, Q9UKF1 | Gene names | ADAM30 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 30 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 30). | |||||
|
RAD54_MOUSE
|
||||||
| NC score | 0.006125 (rank : 44) | θ value | 8.99809 (rank : 44) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P70270, Q8BSR5, Q8C2C4, Q8K3D4 | Gene names | Rad54l, Rad54 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair and recombination protein RAD54-like (EC 3.6.1.-) (RAD54 homolog) (mRAD54) (mHR54). | |||||
|
RAD54_HUMAN
|
||||||
| NC score | 0.006081 (rank : 45) | θ value | 8.99809 (rank : 43) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q92698, Q6IUY3 | Gene names | RAD54L, RAD54A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair and recombination protein RAD54-like (EC 3.6.1.-) (RAD54 homolog) (hRAD54) (hHR54). | |||||
|
EOMES_MOUSE
|
||||||
| NC score | 0.004900 (rank : 46) | θ value | 6.88961 (rank : 37) | |||
| Query Neighborhood Hits | 45 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O54839, Q9QYG7 | Gene names | Eomes, Tbr2 | |||
|
Domain Architecture |

|
|||||
| Description | Eomesodermin homolog. | |||||