Please be patient as the page loads
|
HYAL2_MOUSE
|
||||||
| SwissProt Accessions | O35632, O35631, Q99MS9, Q99MT0 | Gene names | Hyal2 | |||
|
Domain Architecture |

|
|||||
| Description | Hyaluronidase-2 precursor (EC 3.2.1.35) (Hyal-2) (Hyaluronoglucosaminidase-2) (LUCA-2). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
HYAL2_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.997381 (rank : 2) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q12891, O15177, Q9BW29 | Gene names | HYAL2, LUCA2 | |||
|
Domain Architecture |

|
|||||
| Description | Hyaluronidase-2 precursor (EC 3.2.1.35) (Hyal-2) (Hyaluronoglucosaminidase-2) (LUCA-2). | |||||
|
HYAL2_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O35632, O35631, Q99MS9, Q99MT0 | Gene names | Hyal2 | |||
|
Domain Architecture |

|
|||||
| Description | Hyaluronidase-2 precursor (EC 3.2.1.35) (Hyal-2) (Hyaluronoglucosaminidase-2) (LUCA-2). | |||||
|
HYAL1_HUMAN
|
||||||
| θ value | 1.14293e-95 (rank : 3) | NC score | 0.970764 (rank : 3) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q12794, Q6FH23, Q6PIZ6, Q7KYU2, Q7LE34, Q8NFK5, Q8NFK6, Q8NFK7, Q8NFK8, Q8NFK9, Q93013, Q9UKD5, Q9UNI8 | Gene names | HYAL1, LUCA1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hyaluronidase-1 precursor (EC 3.2.1.35) (Hyal-1) (Hyaluronoglucosaminidase-1) (LUCA-1). | |||||
|
HYALP_HUMAN
|
||||||
| θ value | 8.47616e-91 (rank : 4) | NC score | 0.968733 (rank : 4) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P38567 | Gene names | SPAM1, HYAL3, PH20 | |||
|
Domain Architecture |

|
|||||
| Description | Hyaluronidase PH-20 precursor (EC 3.2.1.35) (Hyal-PH20) (Sperm surface protein PH-20) (Sperm adhesion molecule 1). | |||||
|
HYALP_MOUSE
|
||||||
| θ value | 8.47616e-91 (rank : 5) | NC score | 0.966801 (rank : 6) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P48794, Q7TSD6, Q812F4, Q9DAQ1 | Gene names | Spam1, Ph20, Spam | |||
|
Domain Architecture |

|
|||||
| Description | Hyaluronidase PH-20 precursor (EC 3.2.1.35) (Hyal-PH20) (Sperm surface protein PH-20) (Sperm adhesion molecule 1). | |||||
|
HYAL1_MOUSE
|
||||||
| θ value | 1.10702e-90 (rank : 6) | NC score | 0.968692 (rank : 5) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q91ZJ9, O70229, Q8CE62, Q8QZX3, Q8VBW7, Q8VDK0 | Gene names | Hyal1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hyaluronidase-1 precursor (EC 3.2.1.35) (Hyal-1) (Hyaluronoglucosaminidase-1). | |||||
|
HYAL3_HUMAN
|
||||||
| θ value | 1.07224e-85 (rank : 7) | NC score | 0.957672 (rank : 7) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O43820, O60540, Q8NFK2, Q8NFK3, Q8NFK4, Q96E56, Q9BRW9 | Gene names | HYAL3, LUCA3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hyaluronidase-3 precursor (EC 3.2.1.35) (Hyal-3) (Hyaluronoglucosaminidase-3) (LUCA-3). | |||||
|
HYAL3_MOUSE
|
||||||
| θ value | 1.45253e-74 (rank : 8) | NC score | 0.949996 (rank : 8) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8VEI3, Q8VBX7, Q8VI77 | Gene names | Hyal3, Hyl3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hyaluronidase-3 precursor (EC 3.2.1.35) (Hyal-3) (Hyaluronoglucosaminidase-3). | |||||
|
PON1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 9) | NC score | 0.012852 (rank : 11) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P27169, Q16052 | Gene names | PON1, PON | |||
|
Domain Architecture |

|
|||||
| Description | Serum paraoxonase/arylesterase 1 (EC 3.1.1.2) (EC 3.1.8.1) (PON 1) (Serum aryldialkylphosphatase 1) (A-esterase 1) (Aromatic esterase 1) (K-45). | |||||
|
ITB8_HUMAN
|
||||||
| θ value | 5.27518 (rank : 10) | NC score | 0.013022 (rank : 10) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P26012 | Gene names | ITGB8 | |||
|
Domain Architecture |
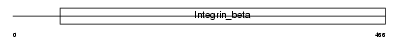
|
|||||
| Description | Integrin beta-8 precursor. | |||||
|
METH_HUMAN
|
||||||
| θ value | 5.27518 (rank : 11) | NC score | 0.018399 (rank : 9) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99707, Q99713, Q99723 | Gene names | MTR | |||
|
Domain Architecture |

|
|||||
| Description | Methionine synthase (EC 2.1.1.13) (5-methyltetrahydrofolate-- homocysteine methyltransferase) (Methionine synthase, vitamin-B12 dependent) (MS). | |||||
|
NOTC1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 12) | NC score | 0.007218 (rank : 13) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 916 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q01705, Q06007, Q61905, Q99JC2, Q9QW58, Q9R0X7 | Gene names | Notch1, Motch | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 1 precursor (Notch 1) (Motch A) (mT14) (p300) [Contains: Notch 1 extracellular truncation; Notch 1 intracellular domain]. | |||||
|
NOTC4_MOUSE
|
||||||
| θ value | 5.27518 (rank : 13) | NC score | 0.006300 (rank : 14) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 872 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P31695, O35442, O88314, O88316, Q62389, Q62390, Q9R1W9, Q9R1X0 | Gene names | Notch4, Int-3, Int3 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 4 precursor (Notch 4) [Contains: Transforming protein Int-3; Notch 4 extracellular truncation; Notch 4 intracellular domain]. | |||||
|
WIF1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 14) | NC score | 0.007389 (rank : 12) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y5W5, Q6UXI1, Q8WVG4 | Gene names | WIF1 | |||
|
Domain Architecture |
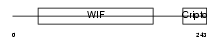
|
|||||
| Description | Wnt inhibitory factor 1 precursor (WIF-1). | |||||
|
CELR2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 15) | NC score | 0.003767 (rank : 15) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 584 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9R0M0, Q99K26, Q9Z2R4 | Gene names | Celsr2 | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin EGF LAG seven-pass G-type receptor 2 precursor (Flamingo 1) (mFmi1). | |||||
|
LAMA2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 16) | NC score | 0.002541 (rank : 16) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 1198 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q60675, Q05003, Q64061 | Gene names | Lama2 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-2 chain precursor (Laminin M chain) (Merosin heavy chain). | |||||
|
HYAL2_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O35632, O35631, Q99MS9, Q99MT0 | Gene names | Hyal2 | |||
|
Domain Architecture |

|
|||||
| Description | Hyaluronidase-2 precursor (EC 3.2.1.35) (Hyal-2) (Hyaluronoglucosaminidase-2) (LUCA-2). | |||||
|
HYAL2_HUMAN
|
||||||
| NC score | 0.997381 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q12891, O15177, Q9BW29 | Gene names | HYAL2, LUCA2 | |||
|
Domain Architecture |

|
|||||
| Description | Hyaluronidase-2 precursor (EC 3.2.1.35) (Hyal-2) (Hyaluronoglucosaminidase-2) (LUCA-2). | |||||
|
HYAL1_HUMAN
|
||||||
| NC score | 0.970764 (rank : 3) | θ value | 1.14293e-95 (rank : 3) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q12794, Q6FH23, Q6PIZ6, Q7KYU2, Q7LE34, Q8NFK5, Q8NFK6, Q8NFK7, Q8NFK8, Q8NFK9, Q93013, Q9UKD5, Q9UNI8 | Gene names | HYAL1, LUCA1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hyaluronidase-1 precursor (EC 3.2.1.35) (Hyal-1) (Hyaluronoglucosaminidase-1) (LUCA-1). | |||||
|
HYALP_HUMAN
|
||||||
| NC score | 0.968733 (rank : 4) | θ value | 8.47616e-91 (rank : 4) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P38567 | Gene names | SPAM1, HYAL3, PH20 | |||
|
Domain Architecture |

|
|||||
| Description | Hyaluronidase PH-20 precursor (EC 3.2.1.35) (Hyal-PH20) (Sperm surface protein PH-20) (Sperm adhesion molecule 1). | |||||
|
HYAL1_MOUSE
|
||||||
| NC score | 0.968692 (rank : 5) | θ value | 1.10702e-90 (rank : 6) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q91ZJ9, O70229, Q8CE62, Q8QZX3, Q8VBW7, Q8VDK0 | Gene names | Hyal1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hyaluronidase-1 precursor (EC 3.2.1.35) (Hyal-1) (Hyaluronoglucosaminidase-1). | |||||
|
HYALP_MOUSE
|
||||||
| NC score | 0.966801 (rank : 6) | θ value | 8.47616e-91 (rank : 5) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P48794, Q7TSD6, Q812F4, Q9DAQ1 | Gene names | Spam1, Ph20, Spam | |||
|
Domain Architecture |

|
|||||
| Description | Hyaluronidase PH-20 precursor (EC 3.2.1.35) (Hyal-PH20) (Sperm surface protein PH-20) (Sperm adhesion molecule 1). | |||||
|
HYAL3_HUMAN
|
||||||
| NC score | 0.957672 (rank : 7) | θ value | 1.07224e-85 (rank : 7) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O43820, O60540, Q8NFK2, Q8NFK3, Q8NFK4, Q96E56, Q9BRW9 | Gene names | HYAL3, LUCA3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hyaluronidase-3 precursor (EC 3.2.1.35) (Hyal-3) (Hyaluronoglucosaminidase-3) (LUCA-3). | |||||
|
HYAL3_MOUSE
|
||||||
| NC score | 0.949996 (rank : 8) | θ value | 1.45253e-74 (rank : 8) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8VEI3, Q8VBX7, Q8VI77 | Gene names | Hyal3, Hyl3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hyaluronidase-3 precursor (EC 3.2.1.35) (Hyal-3) (Hyaluronoglucosaminidase-3). | |||||
|
METH_HUMAN
|
||||||
| NC score | 0.018399 (rank : 9) | θ value | 5.27518 (rank : 11) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99707, Q99713, Q99723 | Gene names | MTR | |||
|
Domain Architecture |

|
|||||
| Description | Methionine synthase (EC 2.1.1.13) (5-methyltetrahydrofolate-- homocysteine methyltransferase) (Methionine synthase, vitamin-B12 dependent) (MS). | |||||
|
ITB8_HUMAN
|
||||||
| NC score | 0.013022 (rank : 10) | θ value | 5.27518 (rank : 10) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P26012 | Gene names | ITGB8 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-8 precursor. | |||||
|
PON1_HUMAN
|
||||||
| NC score | 0.012852 (rank : 11) | θ value | 3.0926 (rank : 9) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P27169, Q16052 | Gene names | PON1, PON | |||
|
Domain Architecture |

|
|||||
| Description | Serum paraoxonase/arylesterase 1 (EC 3.1.1.2) (EC 3.1.8.1) (PON 1) (Serum aryldialkylphosphatase 1) (A-esterase 1) (Aromatic esterase 1) (K-45). | |||||
|
WIF1_HUMAN
|
||||||
| NC score | 0.007389 (rank : 12) | θ value | 5.27518 (rank : 14) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y5W5, Q6UXI1, Q8WVG4 | Gene names | WIF1 | |||
|
Domain Architecture |

|
|||||
| Description | Wnt inhibitory factor 1 precursor (WIF-1). | |||||
|
NOTC1_MOUSE
|
||||||
| NC score | 0.007218 (rank : 13) | θ value | 5.27518 (rank : 12) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 916 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q01705, Q06007, Q61905, Q99JC2, Q9QW58, Q9R0X7 | Gene names | Notch1, Motch | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 1 precursor (Notch 1) (Motch A) (mT14) (p300) [Contains: Notch 1 extracellular truncation; Notch 1 intracellular domain]. | |||||
|
NOTC4_MOUSE
|
||||||
| NC score | 0.006300 (rank : 14) | θ value | 5.27518 (rank : 13) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 872 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P31695, O35442, O88314, O88316, Q62389, Q62390, Q9R1W9, Q9R1X0 | Gene names | Notch4, Int-3, Int3 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 4 precursor (Notch 4) [Contains: Transforming protein Int-3; Notch 4 extracellular truncation; Notch 4 intracellular domain]. | |||||
|
CELR2_MOUSE
|
||||||
| NC score | 0.003767 (rank : 15) | θ value | 6.88961 (rank : 15) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 584 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9R0M0, Q99K26, Q9Z2R4 | Gene names | Celsr2 | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin EGF LAG seven-pass G-type receptor 2 precursor (Flamingo 1) (mFmi1). | |||||
|
LAMA2_MOUSE
|
||||||
| NC score | 0.002541 (rank : 16) | θ value | 8.99809 (rank : 16) | |||
| Query Neighborhood Hits | 16 | Target Neighborhood Hits | 1198 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q60675, Q05003, Q64061 | Gene names | Lama2 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-2 chain precursor (Laminin M chain) (Merosin heavy chain). | |||||