Please be patient as the page loads
|
GALT5_HUMAN
|
||||||
| SwissProt Accessions | Q7Z7M9, Q9UGK7, Q9UHL6 | Gene names | GALNT5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 5 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 5) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 5) (Polypeptide GalNAc transferase 5) (GalNAc-T5) (pp-GaNTase 5). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
GALT5_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 94 | |
| SwissProt Accessions | Q7Z7M9, Q9UGK7, Q9UHL6 | Gene names | GALNT5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 5 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 5) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 5) (Polypeptide GalNAc transferase 5) (GalNAc-T5) (pp-GaNTase 5). | |||||
|
GALT5_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.995904 (rank : 2) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8C102 | Gene names | Galnt5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 5 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 5) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 5) (Polypeptide GalNAc transferase 5) (GalNAc-T5) (pp-GaNTase 5). | |||||
|
GALT1_HUMAN
|
||||||
| θ value | 5.64607e-111 (rank : 3) | NC score | 0.981074 (rank : 6) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q10472, Q86TJ7, Q9UM86 | Gene names | GALNT1 | |||
|
Domain Architecture |

|
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 1 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 1) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 1) (Polypeptide GalNAc transferase 1) (GalNAc-T1) (pp-GaNTase 1) [Contains: Polypeptide N-acetylgalactosaminyltransferase 1 soluble form]. | |||||
|
GALT1_MOUSE
|
||||||
| θ value | 5.64607e-111 (rank : 4) | NC score | 0.981251 (rank : 5) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O08912, Q5BKS3, Q7TND1 | Gene names | Galnt1 | |||
|
Domain Architecture |

|
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 1 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 1) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 1) (Polypeptide GalNAc transferase 1) (GalNAc-T1) (pp-GaNTase 1) [Contains: Polypeptide N-acetylgalactosaminyltransferase 1 soluble form]. | |||||
|
GLT13_HUMAN
|
||||||
| θ value | 2.37213e-109 (rank : 5) | NC score | 0.981064 (rank : 8) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8IUC8, Q6ZWG1, Q96PX0, Q9UIE5 | Gene names | GALNT13, KIAA1918 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 13 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 13) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 13) (Polypeptide GalNAc transferase 13) (GalNAc-T13) (pp-GaNTase 13). | |||||
|
GLT13_MOUSE
|
||||||
| θ value | 3.0981e-109 (rank : 6) | NC score | 0.981072 (rank : 7) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8CF93, Q8BLE4, Q8BYT3 | Gene names | Galnt13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 13 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 13) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 13) (Polypeptide GalNAc transferase 13) (GalNAc-T13) (pp-GaNTase 13). | |||||
|
GLT10_HUMAN
|
||||||
| θ value | 7.14227e-106 (rank : 7) | NC score | 0.982419 (rank : 4) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q86SR1, Q6IN56, Q86VP8, Q8IXJ2, Q8TEJ2, Q96IV2, Q9H8E1, Q9Y4M4 | Gene names | GALNT10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 10 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 10) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 10) (Polypeptide GalNAc transferase 10) (GalNAc-T10) (pp-GaNTase 10). | |||||
|
GLT10_MOUSE
|
||||||
| θ value | 9.32813e-106 (rank : 8) | NC score | 0.982440 (rank : 3) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q6P9S7, Q6KAQ2, Q8BZU8, Q91YJ6 | Gene names | Galnt10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 10 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 10) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 10) (Polypeptide GalNAc transferase 10) (GalNAc-T10) (pp-GaNTase 10). | |||||
|
GALT3_MOUSE
|
||||||
| θ value | 1.10445e-98 (rank : 9) | NC score | 0.976837 (rank : 25) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P70419, Q80V55 | Gene names | Galnt3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 3 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 3) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 3) (Polypeptide GalNAc transferase 3) (GalNAc-T3) (pp-GaNTase 3). | |||||
|
GALT4_MOUSE
|
||||||
| θ value | 1.44246e-98 (rank : 10) | NC score | 0.979987 (rank : 10) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O08832, Q3U681 | Gene names | Galnt4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 4 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 4) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 4) (Polypeptide GalNAc transferase 4) (GalNAc-T4) (pp-GaNTase 4). | |||||
|
GALT4_HUMAN
|
||||||
| θ value | 1.88391e-98 (rank : 11) | NC score | 0.979928 (rank : 11) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8N4A0, O00208 | Gene names | GALNT4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 4 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 4) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 4) (Polypeptide GalNAc transferase 4) (GalNAc-T4) (pp-GaNTase 4). | |||||
|
GALT3_HUMAN
|
||||||
| θ value | 5.48135e-98 (rank : 12) | NC score | 0.976603 (rank : 27) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q14435, Q7Z476 | Gene names | GALNT3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 3 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 3) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 3) (Polypeptide GalNAc transferase 3) (GalNAc-T3) (pp-GaNTase 3). | |||||
|
GALT6_HUMAN
|
||||||
| θ value | 1.03374e-96 (rank : 13) | NC score | 0.977217 (rank : 22) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8NCL4, Q8IYH4, Q9H6G2, Q9UIV5 | Gene names | GALNT6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 6 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 6) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 6) (Polypeptide GalNAc transferase 6) (GalNAc-T6) (pp-GaNTase 6). | |||||
|
GALT6_MOUSE
|
||||||
| θ value | 2.81514e-94 (rank : 14) | NC score | 0.976751 (rank : 26) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8C7U7, Q3TWF0, Q8CED2, Q9QZ16 | Gene names | Galnt6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 6 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 6) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 6) (Polypeptide GalNAc transferase 6) (GalNAc-T6) (pp-GaNTase 6). | |||||
|
GLT11_HUMAN
|
||||||
| θ value | 1.54472e-92 (rank : 15) | NC score | 0.978489 (rank : 15) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8NCW6, Q6PCD1, Q9H6C2, Q9H6Z5, Q9UDR8 | Gene names | GALNT11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 11 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 11) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 11) (Polypeptide GalNAc transferase 11) (GalNAc-T11) (pp-GaNTase 11). | |||||
|
GLTL1_HUMAN
|
||||||
| θ value | 1.70789e-91 (rank : 16) | NC score | 0.977438 (rank : 21) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8N428, Q9ULT9 | Gene names | GALNTL1, KIAA1130 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative polypeptide N-acetylgalactosaminyltransferase-like protein 1 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase-like protein 1) (UDP-GalNAc:polypeptide N-acetylgalactosaminyltransferase- like protein 1) (Polypeptide GalNAc transferase-like protein 1) (GalNAc-T-like protein 1) (pp-GaNTase-like protein 1). | |||||
|
GLT12_HUMAN
|
||||||
| θ value | 2.23057e-91 (rank : 17) | NC score | 0.979809 (rank : 12) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q8IXK2, Q5TCF7, Q8NG54, Q96CT9, Q9H771 | Gene names | GALNT12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 12 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 12) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 12) (Polypeptide GalNAc transferase 12) (GalNAc-T12) (pp-GaNTase 12). | |||||
|
GLT12_MOUSE
|
||||||
| θ value | 3.80478e-91 (rank : 18) | NC score | 0.979632 (rank : 13) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8BGT9 | Gene names | Galnt12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 12 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 12) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 12) (Polypeptide GalNAc transferase 12) (GalNAc-T12) (pp-GaNTase 12). | |||||
|
GLT14_HUMAN
|
||||||
| θ value | 3.80478e-91 (rank : 19) | NC score | 0.977692 (rank : 20) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q96FL9, Q8IVI4, Q9BRH1, Q9H827, Q9H9J8 | Gene names | GALNT14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 14 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 14) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 14) (Polypeptide GalNAc transferase 14) (GalNAc-T14) (pp-GaNTase 14). | |||||
|
GALT2_HUMAN
|
||||||
| θ value | 1.88829e-90 (rank : 20) | NC score | 0.978381 (rank : 17) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q10471, Q9NPY4 | Gene names | GALNT2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 2 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 2) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 2) (Polypeptide GalNAc transferase 2) (GalNAc-T2) (pp-GaNTase 2) [Contains: Polypeptide N-acetylgalactosaminyltransferase 2 soluble form]. | |||||
|
GALT2_MOUSE
|
||||||
| θ value | 2.46618e-90 (rank : 21) | NC score | 0.978402 (rank : 16) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q6PB93, Q7TSI5, Q8BL27, Q922K5, Q99ME1 | Gene names | Galnt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 2 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 2) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 2) (Polypeptide GalNAc transferase 2) (GalNAc-T2) (pp-GaNTase 2) [Contains: Polypeptide N-acetylgalactosaminyltransferase 2 soluble form]. | |||||
|
GLT11_MOUSE
|
||||||
| θ value | 7.17548e-90 (rank : 22) | NC score | 0.978007 (rank : 18) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q921L8, Q3TT83, Q8BU26, Q8K032 | Gene names | Galnt11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 11 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 11) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 11) (Polypeptide GalNAc transferase 11) (GalNAc-T11) (pp-GaNTase 11). | |||||
|
GLTL1_MOUSE
|
||||||
| θ value | 3.56116e-89 (rank : 23) | NC score | 0.977198 (rank : 23) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9JJ61 | Gene names | Galntl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative polypeptide N-acetylgalactosaminyltransferase-like protein 1 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase-like protein 1) (UDP-GalNAc:polypeptide N-acetylgalactosaminyltransferase- like protein 1) (Polypeptide GalNAc transferase-like protein 1) (GalNAc-T-like protein 1) (pp-GaNTase-like protein 1). | |||||
|
GLT14_MOUSE
|
||||||
| θ value | 5.14231e-88 (rank : 24) | NC score | 0.976992 (rank : 24) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8BVG5, Q60GT2, Q8BTI6, Q9DCJ2 | Gene names | Galnt14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 14 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 14) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 14) (Polypeptide GalNAc transferase 14) (GalNAc-T14) (pp-GaNTase 14). | |||||
|
GLTL2_HUMAN
|
||||||
| θ value | 1.2666e-86 (rank : 25) | NC score | 0.977795 (rank : 19) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8N3T1, Q86T60, Q96C46, Q96DJ5 | Gene names | GALNTL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase-like protein 2 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase-like protein 2) (UDP-GalNAc:polypeptide N-acetylgalactosaminyltransferase- like protein 2) (Polypeptide GalNAc transferase-like protein 2) (GalNAc-T-like protein 2) (pp-GaNTase-like protein 2). | |||||
|
GLTL5_MOUSE
|
||||||
| θ value | 2.0926e-81 (rank : 26) | NC score | 0.980698 (rank : 9) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9D4M9, Q810S4, Q9CW06 | Gene names | Galntl5, Galnt15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative polypeptide N-acetylgalactosaminyltransferase-like protein 5 (EC 2.4.1.-) (Protein-UDP acetylgalactosaminyltransferase 15) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 15) (Polypeptide GalNAc transferase 15) (GalNAc-T15) (pp-GaNTase 15). | |||||
|
GALT7_MOUSE
|
||||||
| θ value | 8.79181e-80 (rank : 27) | NC score | 0.974925 (rank : 29) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q80VA0, Q8BY62, Q8BZ70, Q8BZW0, Q91VW6, Q99MD7 | Gene names | Galnt7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetylgalactosaminyltransferase 7 (EC 2.4.1.-) (Protein-UDP acetylgalactosaminyltransferase 7) (UDP-GalNAc:polypeptide N- acetylgalactosaminyltransferase 7) (Polypeptide GalNAc transferase 7) (GalNAc-T7) (pp-GaNTase 7). | |||||
|
GLTL3_HUMAN
|
||||||
| θ value | 1.49966e-79 (rank : 28) | NC score | 0.968970 (rank : 32) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q6IS24, Q8NFV9, Q9NTA8 | Gene names | WBSCR17, GALNTL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative polypeptide N-acetylgalactosaminyltransferase-like protein 3 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase-like protein 3) (UDP-GalNAc:polypeptide N-acetylgalactosaminyltransferase- like protein 3) (Polypeptide GalNAc transferase-like protein 3) (GalNAc-T-like protein 3) (pp-GaNTase-like protein 3) (Williams Beuren syndrome chromosome region 17 protein). | |||||
|
GLTL3_MOUSE
|
||||||
| θ value | 3.34088e-79 (rank : 29) | NC score | 0.968852 (rank : 33) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q7TT15, Q8BKN7, Q8BUY1, Q8BX73, Q8BZC8, Q8K483 | Gene names | Wbscr17, Galntl3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative polypeptide N-acetylgalactosaminyltransferase-like protein 3 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase-like protein 3) (UDP-GalNAc:polypeptide N-acetylgalactosaminyltransferase- like protein 3) (Polypeptide GalNAc transferase-like protein 3) (GalNAc-T-like protein 3) (pp-GaNTase-like protein 3) (Williams-Beuren syndrome chromosome region 17 protein homolog). | |||||
|
GALT7_HUMAN
|
||||||
| θ value | 1.65806e-78 (rank : 30) | NC score | 0.974461 (rank : 30) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q86SF2, Q7Z5W7, Q9UJ28 | Gene names | GALNT7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetylgalactosaminyltransferase 7 (EC 2.4.1.-) (Protein-UDP acetylgalactosaminyltransferase 7) (UDP-GalNAc:polypeptide N- acetylgalactosaminyltransferase 7) (Polypeptide GalNAc transferase 7) (GalNAc-T7) (pp-GaNTase 7). | |||||
|
GLTL5_HUMAN
|
||||||
| θ value | 6.30065e-78 (rank : 31) | NC score | 0.979424 (rank : 14) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q7Z4T8, Q75KN2, Q75MD3, Q8NCV4, Q8WW05, Q9UDR9 | Gene names | GALNTL5, GALNT15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative polypeptide N-acetylgalactosaminyltransferase-like protein 5 (EC 2.4.1.-) (Protein-UDP acetylgalactosaminyltransferase 15) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 15) (Polypeptide GalNAc transferase 15) (GalNAc-T15) (pp-GaNTase 15). | |||||
|
GALT9_HUMAN
|
||||||
| θ value | 1.5519e-76 (rank : 32) | NC score | 0.970362 (rank : 31) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9HCQ5, Q6NT54, Q8NFR1 | Gene names | GALNT9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 9 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 9) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 9) (Polypeptide GalNAc transferase 9) (GalNAc-T9) (pp-GaNTase 9). | |||||
|
GLTL2_MOUSE
|
||||||
| θ value | 2.02684e-76 (rank : 33) | NC score | 0.975849 (rank : 28) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9D2N8 | Gene names | Galntl2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase-like protein 2 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase-like protein 2) (UDP-GalNAc:polypeptide N-acetylgalactosaminyltransferase- like protein 2) (Polypeptide GalNAc transferase-like protein 2) (GalNAc-T-like protein 2) (pp-GaNTase-like protein 2). | |||||
|
GLTL4_HUMAN
|
||||||
| θ value | 7.70199e-76 (rank : 34) | NC score | 0.966725 (rank : 35) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q6P9A2, O95903, Q8NDY9 | Gene names | GALNTL4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative polypeptide N-acetylgalactosaminyltransferase-like protein 4 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase-like protein 4) (UDP-GalNAc:polypeptide N-acetylgalactosaminyltransferase- like protein 4) (Polypeptide GalNAc transferase-like protein 4) (GalNAc-T-like protein 4) (pp-GaNTase-like protein 4). | |||||
|
GLTL4_MOUSE
|
||||||
| θ value | 1.96316e-71 (rank : 35) | NC score | 0.965066 (rank : 36) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8K1B9 | Gene names | Galntl4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative polypeptide N-acetylgalactosaminyltransferase-like protein 4 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase-like protein 4) (UDP-GalNAc:polypeptide N-acetylgalactosaminyltransferase- like protein 4) (Polypeptide GalNAc transferase-like protein 4) (GalNAc-T-like protein 4) (pp-GaNTase-like protein 4). | |||||
|
GALT8_HUMAN
|
||||||
| θ value | 1.40689e-69 (rank : 36) | NC score | 0.968463 (rank : 34) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9NY28 | Gene names | GALNT8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable polypeptide N-acetylgalactosaminyltransferase 8 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 8) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 8) (Polypeptide GalNAc transferase 8) (GalNAc-T8) (pp-GaNTase 8). | |||||
|
CD2L5_HUMAN
|
||||||
| θ value | 0.00665767 (rank : 37) | NC score | 0.001700 (rank : 90) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 987 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q14004, Q6DKQ9, Q75MH4, Q75MH5, Q96JN4, Q9H4A0, Q9H4A1, Q9UDR4 | Gene names | CDC2L5, CDC2L, CHED, KIAA1791 | |||
|
Domain Architecture |
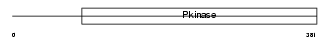
|
|||||
| Description | Cell division cycle 2-like protein kinase 5 (EC 2.7.11.22) (CDC2- related protein kinase 5) (Cholinesterase-related cell division controller). | |||||
|
DNMT1_HUMAN
|
||||||
| θ value | 0.0252991 (rank : 38) | NC score | 0.022683 (rank : 42) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P26358, Q9UHG5, Q9ULA2, Q9UMZ6 | Gene names | DNMT1, AIM, DNMT | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 1 (EC 2.1.1.37) (Dnmt1) (DNA methyltransferase HsaI) (DNA MTase HsaI) (MCMT) (M.HsaI). | |||||
|
ICAL_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 39) | NC score | 0.028553 (rank : 40) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P51125, Q9EQV4, Q9EQV5, Q9QXQ3, Q9QXQ4, Q9R0N1 | Gene names | Cast | |||
|
Domain Architecture |

|
|||||
| Description | Calpastatin (Calpain inhibitor). | |||||
|
H13_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 40) | NC score | 0.020538 (rank : 43) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P16402 | Gene names | HIST1H1D, H1F3 | |||
|
Domain Architecture |
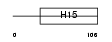
|
|||||
| Description | Histone H1.3 (Histone H1c). | |||||
|
TTBK2_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 41) | NC score | 0.004938 (rank : 79) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 822 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6IQ55, O94932, Q6ZN52, Q8IVV1 | Gene names | TTBK2, KIAA0847 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tau-tubulin kinase 2 (EC 2.7.11.1). | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 42) | NC score | 0.019259 (rank : 45) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
NOLC1_HUMAN
|
||||||
| θ value | 0.163984 (rank : 43) | NC score | 0.018675 (rank : 47) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q14978, Q15030 | Gene names | NOLC1, KIAA0035 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar phosphoprotein p130 (Nucleolar 130 kDa protein) (140 kDa nucleolar phosphoprotein) (Nopp140) (Nucleolar and coiled-body phosphoprotein 1). | |||||
|
SFRS4_MOUSE
|
||||||
| θ value | 0.163984 (rank : 44) | NC score | 0.010665 (rank : 61) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8VE97, Q9JJC3 | Gene names | Sfrs4 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 4. | |||||
|
SRRM2_HUMAN
|
||||||
| θ value | 0.21417 (rank : 45) | NC score | 0.023440 (rank : 41) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
K2C1B_MOUSE
|
||||||
| θ value | 0.279714 (rank : 46) | NC score | 0.002099 (rank : 88) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 503 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6IFZ6 | Gene names | Krt1b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin, type II cytoskeletal 1b (Type II keratin Kb39) (Embryonic type II keratin-1). | |||||
|
LARP4_MOUSE
|
||||||
| θ value | 0.279714 (rank : 47) | NC score | 0.018924 (rank : 46) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8BWW4, Q3UR69 | Gene names | Larp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | La-related protein 4 (La ribonucleoprotein domain family member 4). | |||||
|
MASTL_HUMAN
|
||||||
| θ value | 0.365318 (rank : 48) | NC score | -0.001067 (rank : 93) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 894 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96GX5, Q5T8D5, Q5T8D7, Q8NCD6, Q96SJ5 | Gene names | MASTL, THC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated serine/threonine-protein kinase-like (EC 2.7.11.1). | |||||
|
MYPT1_MOUSE
|
||||||
| θ value | 0.365318 (rank : 49) | NC score | 0.004951 (rank : 78) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 539 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9DBR7, Q8CBV2, Q99NB6 | Gene names | Ppp1r12a, Mypt1 | |||
|
Domain Architecture |
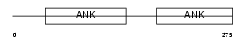
|
|||||
| Description | Protein phosphatase 1 regulatory subunit 12A (Myosin phosphatase- targeting subunit 1) (Myosin phosphatase target subunit 1). | |||||
|
TM108_MOUSE
|
||||||
| θ value | 0.365318 (rank : 50) | NC score | 0.019624 (rank : 44) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 336 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8BHE4, Q80WR9 | Gene names | Tmem108 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein 108 precursor. | |||||
|
CYLC1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 51) | NC score | 0.013150 (rank : 56) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P35663, Q5JQQ9 | Gene names | CYLC1, CYL, CYL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-1 (Cylicin I) (Multiple-band polypeptide I). | |||||
|
NACAM_MOUSE
|
||||||
| θ value | 0.47712 (rank : 52) | NC score | 0.010253 (rank : 62) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
TCOF_MOUSE
|
||||||
| θ value | 0.47712 (rank : 53) | NC score | 0.016217 (rank : 50) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 471 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O08784, O08857 | Gene names | Tcof1 | |||
|
Domain Architecture |

|
|||||
| Description | Treacle protein (Treacher Collins syndrome protein homolog). | |||||
|
SUHW3_HUMAN
|
||||||
| θ value | 0.813845 (rank : 54) | NC score | -0.000259 (rank : 92) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 695 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8ND82, Q9NXR3 | Gene names | SUHW3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppressor of hairy wing homolog 3. | |||||
|
TM131_MOUSE
|
||||||
| θ value | 0.813845 (rank : 55) | NC score | 0.018647 (rank : 48) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O70472 | Gene names | Tmem131, D1Bwg0491e, Rw1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein 131 (Protein RW1). | |||||
|
AMOL1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 56) | NC score | 0.007533 (rank : 68) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9D4H4, Q571F1 | Gene names | Amotl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin-like protein 1. | |||||
|
ATRX_HUMAN
|
||||||
| θ value | 1.06291 (rank : 57) | NC score | 0.005511 (rank : 75) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 842 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P46100, P51068, Q15886, Q59FB5, Q59H31, Q5H9A2, Q5JWI4, Q7Z2J1, Q9H0Z1, Q9NTS3 | Gene names | ATRX, RAD54L, XH2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ATRX (EC 3.6.1.-) (ATP-dependent helicase ATRX) (X-linked helicase II) (X-linked nuclear protein) (XNP) (Znf- HX). | |||||
|
CD2L5_MOUSE
|
||||||
| θ value | 1.06291 (rank : 58) | NC score | -0.000000 (rank : 91) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1011 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q69ZA1, Q80V11, Q8BZG1, Q8K0A4 | Gene names | Cdc2l5, Kiaa1791 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle 2-like protein kinase 5 (EC 2.7.11.22) (CDC2- related protein kinase 5). | |||||
|
HUWE1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 59) | NC score | 0.005508 (rank : 76) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q7TMY8, Q4G2Z1, Q5BMM7, Q6NS61, Q8BNJ7, Q8CFH2, Q8VD14, Q921M5, Q9R0P2 | Gene names | Huwe1, Kiaa0312, Ureb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT, UBA and WWE domain-containing protein 1 (EC 6.3.2.-) (E3 ubiquitin protein ligase URE-B1) (E3Histone). | |||||
|
NMDE1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 60) | NC score | 0.006660 (rank : 71) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P35436 | Gene names | Grin2a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate [NMDA] receptor subunit epsilon 1 precursor (N-methyl D- aspartate receptor subtype 2A) (NR2A) (NMDAR2A). | |||||
|
PAPOG_HUMAN
|
||||||
| θ value | 1.06291 (rank : 61) | NC score | 0.008918 (rank : 63) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9BWT3, Q969N1, Q9H8L2, Q9HAD0 | Gene names | PAPOLG, PAP2, PAPG | |||
|
Domain Architecture |

|
|||||
| Description | Poly(A) polymerase gamma (EC 2.7.7.19) (PAP gamma) (Polynucleotide adenylyltransferase gamma) (SRP RNA 3' adenylating enzyme) (Neo- poly(A) polymerase) (Neo-PAP). | |||||
|
BCLF1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 62) | NC score | 0.011686 (rank : 59) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8K019, Q8BNZ0, Q8C2E9, Q9CSW5 | Gene names | Bclaf1, Btf, Kiaa0164 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bcl-2-associated transcription factor 1 (Btf). | |||||
|
MYPT2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 63) | NC score | 0.003721 (rank : 82) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 707 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O60237, Q8N179, Q9HCB7, Q9HCB8 | Gene names | PPP1R12B, MYPT2 | |||
|
Domain Architecture |
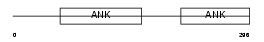
|
|||||
| Description | Protein phosphatase 1 regulatory subunit 12B (Myosin phosphatase- targeting subunit 2) (Myosin phosphatase target subunit 2). | |||||
|
SPTN2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 64) | NC score | 0.003013 (rank : 86) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 535 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O15020, O14872, O14873 | Gene names | SPTBN2, KIAA0302 | |||
|
Domain Architecture |

|
|||||
| Description | Spectrin beta chain, brain 2 (Spectrin, non-erythroid beta chain 2) (Beta-III spectrin). | |||||
|
PRG4_HUMAN
|
||||||
| θ value | 1.81305 (rank : 65) | NC score | 0.008494 (rank : 65) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q92954, Q6DNC4, Q6DNC5, Q6ZMZ5, Q9BX49 | Gene names | PRG4, MSF, SZP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
REST_MOUSE
|
||||||
| θ value | 1.81305 (rank : 66) | NC score | -0.002161 (rank : 94) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1506 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q922J3, Q571L7 | Gene names | Rsn, Kiaa4046 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Restin. | |||||
|
B4GN4_MOUSE
|
||||||
| θ value | 2.36792 (rank : 67) | NC score | 0.073879 (rank : 38) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q766D5, Q8CHV2 | Gene names | B4galnt4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyl-beta-glucosaminyl-glycoprotein 4-beta-N- acetylgalactosaminyltransferase 1 (EC 2.4.1.244) (NGalNAc-T1) (Beta- 1,4-N-acetylgalactosaminyltransferase IV) (Beta4GalNAc-T4) (Beta4GalNAcT4). | |||||
|
CKAP2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 68) | NC score | 0.013931 (rank : 53) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8WWK9, Q3KRA5, Q5VXB4, Q8IWV5, Q8IWV6, Q96FH9, Q9H012, Q9H0D0, Q9H988, Q9HC49, Q9NVG4 | Gene names | CKAP2, LB1, TMAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoskeleton-associated protein 2 (Tumor-associated microtubule- associated protein) (CTCL tumor antigen se20-10). | |||||
|
TBX1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 69) | NC score | 0.003343 (rank : 84) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P70323, Q60706, Q99MP0, Q99P22 | Gene names | Tbx1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | T-box transcription factor TBX1 (T-box protein 1) (Testis-specific T- box protein). | |||||
|
B4GN4_HUMAN
|
||||||
| θ value | 3.0926 (rank : 70) | NC score | 0.055014 (rank : 39) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q76KP1, Q96LV2 | Gene names | B4GALNT4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyl-beta-glucosaminyl-glycoprotein 4-beta-N- acetylgalactosaminyltransferase 1 (EC 2.4.1.244) (NGalNAc-T1) (Beta- 1,4-N-acetylgalactosaminyltransferase IV) (Beta4GalNAc-T4) (Beta4GalNAcT4). | |||||
|
CJ012_HUMAN
|
||||||
| θ value | 3.0926 (rank : 71) | NC score | 0.016600 (rank : 49) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8N655, Q9H945, Q9Y457 | Gene names | C10orf12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf12. | |||||
|
H15_HUMAN
|
||||||
| θ value | 3.0926 (rank : 72) | NC score | 0.012142 (rank : 58) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P16401, Q14529 | Gene names | HIST1H1B, H1F5 | |||
|
Domain Architecture |
|
|||||
| Description | Histone H1.5 (Histone H1a). | |||||
|
MILK1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 73) | NC score | 0.005571 (rank : 74) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 788 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8BGT6, Q8BJ60 | Gene names | Mirab13, D15Mit260, Kiaa1668 | |||
|
Domain Architecture |

|
|||||
| Description | Molecule interacting with Rab13 (MIRab13) (MICAL-like protein 1). | |||||
|
SP17_HUMAN
|
||||||
| θ value | 3.0926 (rank : 74) | NC score | 0.013273 (rank : 54) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q15506, Q9BXF7 | Gene names | SPA17, SP17 | |||
|
Domain Architecture |
|
|||||
| Description | Sperm surface protein Sp17 (Sperm autoantigenic protein 17) (Sperm protein 17) (Sp17-1). | |||||
|
BASP_MOUSE
|
||||||
| θ value | 4.03905 (rank : 75) | NC score | 0.010824 (rank : 60) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q91XV3 | Gene names | Basp1, Nap22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
NFH_MOUSE
|
||||||
| θ value | 4.03905 (rank : 76) | NC score | 0.003320 (rank : 85) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
NMDE1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 77) | NC score | 0.005388 (rank : 77) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q12879, O00669 | Gene names | GRIN2A | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate [NMDA] receptor subunit epsilon 1 precursor (N-methyl D- aspartate receptor subtype 2A) (NR2A) (NMDAR2A) (hNR2A). | |||||
|
YETS2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 78) | NC score | 0.008515 (rank : 64) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9ULM3, Q641P6, Q9NW96 | Gene names | YEATS2, KIAA1197 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YEATS domain-containing protein 2. | |||||
|
LAP2B_MOUSE
|
||||||
| θ value | 5.27518 (rank : 79) | NC score | 0.008064 (rank : 66) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q61029, Q61030, Q61031, Q61032 | Gene names | Tmpo, Lap2 | |||
|
Domain Architecture |

|
|||||
| Description | Lamina-associated polypeptide 2 isoforms beta/delta/epsilon/gamma (Thymopoietin isoforms beta/delta/epsilon/gamma) (TP beta/delta/epsilon/gamma). | |||||
|
PARG_MOUSE
|
||||||
| θ value | 5.27518 (rank : 80) | NC score | 0.006889 (rank : 70) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O88622, Q80YQ6, Q8CB72 | Gene names | Parg | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly(ADP-ribose) glycohydrolase (EC 3.2.1.143). | |||||
|
SFRIP_HUMAN
|
||||||
| θ value | 5.27518 (rank : 81) | NC score | 0.012347 (rank : 57) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 530 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q99590, Q8IW59 | Gene names | SFRS2IP, CASP11, SIP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SFRS2-interacting protein (Splicing factor, arginine/serine-rich 2- interacting protein) (SC35-interacting protein 1) (CTD-associated SR protein 11) (Splicing regulatory protein 129) (SRrp129) (NY-REN-40 antigen). | |||||
|
AB1IP_MOUSE
|
||||||
| θ value | 6.88961 (rank : 82) | NC score | 0.013170 (rank : 55) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8R5A3, O35329, Q8BRU0, Q99KV8 | Gene names | Apbb1ip, Prel1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 precursor protein-binding family B member 1- interacting protein (APBB1-interacting protein 1) (Proline-rich EVH1 ligand 1) (PREL-1) (Proline-rich protein 48). | |||||
|
BMP2K_MOUSE
|
||||||
| θ value | 6.88961 (rank : 83) | NC score | -0.002790 (rank : 95) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 874 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q91Z96, Q8C8L7 | Gene names | Bmp2k, Bike | |||
|
Domain Architecture |
|
|||||
| Description | BMP-2-inducible protein kinase (EC 2.7.11.1) (BIKe). | |||||
|
BRPF1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 84) | NC score | 0.001727 (rank : 89) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P55201, Q9UHI0 | Gene names | BRPF1, BR140 | |||
|
Domain Architecture |

|
|||||
| Description | Peregrin (Bromodomain and PHD finger-containing protein 1) (BR140 protein). | |||||
|
CIC_HUMAN
|
||||||
| θ value | 6.88961 (rank : 85) | NC score | 0.002318 (rank : 87) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 992 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q96RK0, Q7LGI1, Q9UEG5, Q9Y6T1 | Gene names | CIC, KIAA0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
FILA_HUMAN
|
||||||
| θ value | 6.88961 (rank : 86) | NC score | 0.005905 (rank : 73) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 685 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P20930, Q01720, Q5T583 | Gene names | FLG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filaggrin. | |||||
|
PDZD6_HUMAN
|
||||||
| θ value | 6.88961 (rank : 87) | NC score | 0.007730 (rank : 67) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9ULD6, Q4W5I8, Q86V55 | Gene names | PDZD6, KIAA1284, PDZK6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing protein 6. | |||||
|
RBBP6_HUMAN
|
||||||
| θ value | 6.88961 (rank : 88) | NC score | 0.007300 (rank : 69) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q7Z6E9, Q15290, Q6DKH4, Q6P4C2, Q6YNC9, Q7Z6E8, Q8N0V2, Q96PH3, Q9H3I8, Q9H5M5, Q9NPX4 | Gene names | RBBP6, P2PR, PACT, RBQ1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R) (Retinoblastoma-binding Q protein 1) (Protein RBQ-1). | |||||
|
RTN4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 89) | NC score | 0.004533 (rank : 80) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9NQC3, O94962, Q9BXG5, Q9H212, Q9H3I3, Q9UQ42, Q9Y293, Q9Y2Y7, Q9Y5U6 | Gene names | RTN4, KIAA0886, NOGO | |||
|
Domain Architecture |

|
|||||
| Description | Reticulon-4 (Neurite outgrowth inhibitor) (Nogo protein) (Foocen) (Neuroendocrine-specific protein) (NSP) (Neuroendocrine-specific protein C homolog) (RTN-x) (Reticulon-5). | |||||
|
UTX_HUMAN
|
||||||
| θ value | 6.88961 (rank : 90) | NC score | 0.004058 (rank : 81) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O15550 | Gene names | UTX | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitously transcribed X chromosome tetratricopeptide repeat protein (Ubiquitously transcribed TPR protein on the X chromosome). | |||||
|
LAP2A_HUMAN
|
||||||
| θ value | 8.99809 (rank : 91) | NC score | 0.006012 (rank : 72) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P42166, P08918, P08919, Q14860, Q16295 | Gene names | TMPO, LAP2 | |||
|
Domain Architecture |

|
|||||
| Description | Lamina-associated polypeptide 2 isoform alpha (Thymopoietin isoform alpha) (TP alpha) (Thymopoietin-related peptide isoform alpha) (TPRP isoform alpha) [Contains: Thymopoietin (TP) (Splenin); Thymopentin (TP5)]. | |||||
|
NACA_HUMAN
|
||||||
| θ value | 8.99809 (rank : 92) | NC score | 0.013965 (rank : 52) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13765, Q53A18, Q53G46 | Gene names | NACA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha (NAC-alpha) (Alpha-NAC) (Hom s 2.02). | |||||
|
NACA_MOUSE
|
||||||
| θ value | 8.99809 (rank : 93) | NC score | 0.013974 (rank : 51) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q60817 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha (Alpha-NAC) (Alpha-NAC/1.9.2). | |||||
|
SI1L3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 94) | NC score | 0.003702 (rank : 83) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O60292, Q2TV87 | Gene names | SIPA1L3, KIAA0545, SPAL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal-induced proliferation-associated 1-like protein 3 (SPA-1-like protein 3). | |||||
|
DPM1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 95) | NC score | 0.078554 (rank : 37) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O70152, Q9D829 | Gene names | Dpm1 | |||
|
Domain Architecture |

|
|||||
| Description | Dolichol-phosphate mannosyltransferase (EC 2.4.1.83) (Dolichol- phosphate mannose synthase) (Dolichyl-phosphate beta-D- mannosyltransferase) (Mannose-P-dolichol synthase) (MPD synthase) (DPM synthase). | |||||
|
GALT5_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 94 | |
| SwissProt Accessions | Q7Z7M9, Q9UGK7, Q9UHL6 | Gene names | GALNT5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 5 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 5) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 5) (Polypeptide GalNAc transferase 5) (GalNAc-T5) (pp-GaNTase 5). | |||||
|
GALT5_MOUSE
|
||||||
| NC score | 0.995904 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8C102 | Gene names | Galnt5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 5 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 5) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 5) (Polypeptide GalNAc transferase 5) (GalNAc-T5) (pp-GaNTase 5). | |||||
|
GLT10_MOUSE
|
||||||
| NC score | 0.982440 (rank : 3) | θ value | 9.32813e-106 (rank : 8) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q6P9S7, Q6KAQ2, Q8BZU8, Q91YJ6 | Gene names | Galnt10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 10 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 10) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 10) (Polypeptide GalNAc transferase 10) (GalNAc-T10) (pp-GaNTase 10). | |||||
|
GLT10_HUMAN
|
||||||
| NC score | 0.982419 (rank : 4) | θ value | 7.14227e-106 (rank : 7) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q86SR1, Q6IN56, Q86VP8, Q8IXJ2, Q8TEJ2, Q96IV2, Q9H8E1, Q9Y4M4 | Gene names | GALNT10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 10 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 10) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 10) (Polypeptide GalNAc transferase 10) (GalNAc-T10) (pp-GaNTase 10). | |||||
|
GALT1_MOUSE
|
||||||
| NC score | 0.981251 (rank : 5) | θ value | 5.64607e-111 (rank : 4) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | O08912, Q5BKS3, Q7TND1 | Gene names | Galnt1 | |||
|
Domain Architecture |

|
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 1 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 1) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 1) (Polypeptide GalNAc transferase 1) (GalNAc-T1) (pp-GaNTase 1) [Contains: Polypeptide N-acetylgalactosaminyltransferase 1 soluble form]. | |||||
|
GALT1_HUMAN
|
||||||
| NC score | 0.981074 (rank : 6) | θ value | 5.64607e-111 (rank : 3) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q10472, Q86TJ7, Q9UM86 | Gene names | GALNT1 | |||
|
Domain Architecture |

|
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 1 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 1) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 1) (Polypeptide GalNAc transferase 1) (GalNAc-T1) (pp-GaNTase 1) [Contains: Polypeptide N-acetylgalactosaminyltransferase 1 soluble form]. | |||||
|
GLT13_MOUSE
|
||||||
| NC score | 0.981072 (rank : 7) | θ value | 3.0981e-109 (rank : 6) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8CF93, Q8BLE4, Q8BYT3 | Gene names | Galnt13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 13 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 13) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 13) (Polypeptide GalNAc transferase 13) (GalNAc-T13) (pp-GaNTase 13). | |||||
|
GLT13_HUMAN
|
||||||
| NC score | 0.981064 (rank : 8) | θ value | 2.37213e-109 (rank : 5) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8IUC8, Q6ZWG1, Q96PX0, Q9UIE5 | Gene names | GALNT13, KIAA1918 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 13 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 13) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 13) (Polypeptide GalNAc transferase 13) (GalNAc-T13) (pp-GaNTase 13). | |||||
|
GLTL5_MOUSE
|
||||||
| NC score | 0.980698 (rank : 9) | θ value | 2.0926e-81 (rank : 26) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9D4M9, Q810S4, Q9CW06 | Gene names | Galntl5, Galnt15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative polypeptide N-acetylgalactosaminyltransferase-like protein 5 (EC 2.4.1.-) (Protein-UDP acetylgalactosaminyltransferase 15) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 15) (Polypeptide GalNAc transferase 15) (GalNAc-T15) (pp-GaNTase 15). | |||||
|
GALT4_MOUSE
|
||||||
| NC score | 0.979987 (rank : 10) | θ value | 1.44246e-98 (rank : 10) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | O08832, Q3U681 | Gene names | Galnt4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 4 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 4) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 4) (Polypeptide GalNAc transferase 4) (GalNAc-T4) (pp-GaNTase 4). | |||||
|
GALT4_HUMAN
|
||||||
| NC score | 0.979928 (rank : 11) | θ value | 1.88391e-98 (rank : 11) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8N4A0, O00208 | Gene names | GALNT4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 4 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 4) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 4) (Polypeptide GalNAc transferase 4) (GalNAc-T4) (pp-GaNTase 4). | |||||
|
GLT12_HUMAN
|
||||||
| NC score | 0.979809 (rank : 12) | θ value | 2.23057e-91 (rank : 17) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q8IXK2, Q5TCF7, Q8NG54, Q96CT9, Q9H771 | Gene names | GALNT12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 12 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 12) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 12) (Polypeptide GalNAc transferase 12) (GalNAc-T12) (pp-GaNTase 12). | |||||
|
GLT12_MOUSE
|
||||||
| NC score | 0.979632 (rank : 13) | θ value | 3.80478e-91 (rank : 18) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8BGT9 | Gene names | Galnt12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 12 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 12) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 12) (Polypeptide GalNAc transferase 12) (GalNAc-T12) (pp-GaNTase 12). | |||||
|
GLTL5_HUMAN
|
||||||
| NC score | 0.979424 (rank : 14) | θ value | 6.30065e-78 (rank : 31) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q7Z4T8, Q75KN2, Q75MD3, Q8NCV4, Q8WW05, Q9UDR9 | Gene names | GALNTL5, GALNT15 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative polypeptide N-acetylgalactosaminyltransferase-like protein 5 (EC 2.4.1.-) (Protein-UDP acetylgalactosaminyltransferase 15) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 15) (Polypeptide GalNAc transferase 15) (GalNAc-T15) (pp-GaNTase 15). | |||||
|
GLT11_HUMAN
|
||||||
| NC score | 0.978489 (rank : 15) | θ value | 1.54472e-92 (rank : 15) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8NCW6, Q6PCD1, Q9H6C2, Q9H6Z5, Q9UDR8 | Gene names | GALNT11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 11 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 11) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 11) (Polypeptide GalNAc transferase 11) (GalNAc-T11) (pp-GaNTase 11). | |||||
|
GALT2_MOUSE
|
||||||
| NC score | 0.978402 (rank : 16) | θ value | 2.46618e-90 (rank : 21) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q6PB93, Q7TSI5, Q8BL27, Q922K5, Q99ME1 | Gene names | Galnt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 2 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 2) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 2) (Polypeptide GalNAc transferase 2) (GalNAc-T2) (pp-GaNTase 2) [Contains: Polypeptide N-acetylgalactosaminyltransferase 2 soluble form]. | |||||
|
GALT2_HUMAN
|
||||||
| NC score | 0.978381 (rank : 17) | θ value | 1.88829e-90 (rank : 20) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q10471, Q9NPY4 | Gene names | GALNT2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 2 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 2) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 2) (Polypeptide GalNAc transferase 2) (GalNAc-T2) (pp-GaNTase 2) [Contains: Polypeptide N-acetylgalactosaminyltransferase 2 soluble form]. | |||||
|
GLT11_MOUSE
|
||||||
| NC score | 0.978007 (rank : 18) | θ value | 7.17548e-90 (rank : 22) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q921L8, Q3TT83, Q8BU26, Q8K032 | Gene names | Galnt11 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 11 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 11) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 11) (Polypeptide GalNAc transferase 11) (GalNAc-T11) (pp-GaNTase 11). | |||||
|
GLTL2_HUMAN
|
||||||
| NC score | 0.977795 (rank : 19) | θ value | 1.2666e-86 (rank : 25) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8N3T1, Q86T60, Q96C46, Q96DJ5 | Gene names | GALNTL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase-like protein 2 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase-like protein 2) (UDP-GalNAc:polypeptide N-acetylgalactosaminyltransferase- like protein 2) (Polypeptide GalNAc transferase-like protein 2) (GalNAc-T-like protein 2) (pp-GaNTase-like protein 2). | |||||
|
GLT14_HUMAN
|
||||||
| NC score | 0.977692 (rank : 20) | θ value | 3.80478e-91 (rank : 19) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q96FL9, Q8IVI4, Q9BRH1, Q9H827, Q9H9J8 | Gene names | GALNT14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 14 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 14) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 14) (Polypeptide GalNAc transferase 14) (GalNAc-T14) (pp-GaNTase 14). | |||||
|
GLTL1_HUMAN
|
||||||
| NC score | 0.977438 (rank : 21) | θ value | 1.70789e-91 (rank : 16) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8N428, Q9ULT9 | Gene names | GALNTL1, KIAA1130 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative polypeptide N-acetylgalactosaminyltransferase-like protein 1 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase-like protein 1) (UDP-GalNAc:polypeptide N-acetylgalactosaminyltransferase- like protein 1) (Polypeptide GalNAc transferase-like protein 1) (GalNAc-T-like protein 1) (pp-GaNTase-like protein 1). | |||||
|
GALT6_HUMAN
|
||||||
| NC score | 0.977217 (rank : 22) | θ value | 1.03374e-96 (rank : 13) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8NCL4, Q8IYH4, Q9H6G2, Q9UIV5 | Gene names | GALNT6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 6 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 6) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 6) (Polypeptide GalNAc transferase 6) (GalNAc-T6) (pp-GaNTase 6). | |||||
|
GLTL1_MOUSE
|
||||||
| NC score | 0.977198 (rank : 23) | θ value | 3.56116e-89 (rank : 23) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9JJ61 | Gene names | Galntl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative polypeptide N-acetylgalactosaminyltransferase-like protein 1 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase-like protein 1) (UDP-GalNAc:polypeptide N-acetylgalactosaminyltransferase- like protein 1) (Polypeptide GalNAc transferase-like protein 1) (GalNAc-T-like protein 1) (pp-GaNTase-like protein 1). | |||||
|
GLT14_MOUSE
|
||||||
| NC score | 0.976992 (rank : 24) | θ value | 5.14231e-88 (rank : 24) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8BVG5, Q60GT2, Q8BTI6, Q9DCJ2 | Gene names | Galnt14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 14 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 14) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 14) (Polypeptide GalNAc transferase 14) (GalNAc-T14) (pp-GaNTase 14). | |||||
|
GALT3_MOUSE
|
||||||
| NC score | 0.976837 (rank : 25) | θ value | 1.10445e-98 (rank : 9) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P70419, Q80V55 | Gene names | Galnt3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 3 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 3) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 3) (Polypeptide GalNAc transferase 3) (GalNAc-T3) (pp-GaNTase 3). | |||||
|
GALT6_MOUSE
|
||||||
| NC score | 0.976751 (rank : 26) | θ value | 2.81514e-94 (rank : 14) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q8C7U7, Q3TWF0, Q8CED2, Q9QZ16 | Gene names | Galnt6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 6 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 6) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 6) (Polypeptide GalNAc transferase 6) (GalNAc-T6) (pp-GaNTase 6). | |||||
|
GALT3_HUMAN
|
||||||
| NC score | 0.976603 (rank : 27) | θ value | 5.48135e-98 (rank : 12) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q14435, Q7Z476 | Gene names | GALNT3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 3 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 3) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 3) (Polypeptide GalNAc transferase 3) (GalNAc-T3) (pp-GaNTase 3). | |||||
|
GLTL2_MOUSE
|
||||||
| NC score | 0.975849 (rank : 28) | θ value | 2.02684e-76 (rank : 33) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9D2N8 | Gene names | Galntl2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase-like protein 2 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase-like protein 2) (UDP-GalNAc:polypeptide N-acetylgalactosaminyltransferase- like protein 2) (Polypeptide GalNAc transferase-like protein 2) (GalNAc-T-like protein 2) (pp-GaNTase-like protein 2). | |||||
|
GALT7_MOUSE
|
||||||
| NC score | 0.974925 (rank : 29) | θ value | 8.79181e-80 (rank : 27) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q80VA0, Q8BY62, Q8BZ70, Q8BZW0, Q91VW6, Q99MD7 | Gene names | Galnt7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetylgalactosaminyltransferase 7 (EC 2.4.1.-) (Protein-UDP acetylgalactosaminyltransferase 7) (UDP-GalNAc:polypeptide N- acetylgalactosaminyltransferase 7) (Polypeptide GalNAc transferase 7) (GalNAc-T7) (pp-GaNTase 7). | |||||
|
GALT7_HUMAN
|
||||||
| NC score | 0.974461 (rank : 30) | θ value | 1.65806e-78 (rank : 30) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q86SF2, Q7Z5W7, Q9UJ28 | Gene names | GALNT7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetylgalactosaminyltransferase 7 (EC 2.4.1.-) (Protein-UDP acetylgalactosaminyltransferase 7) (UDP-GalNAc:polypeptide N- acetylgalactosaminyltransferase 7) (Polypeptide GalNAc transferase 7) (GalNAc-T7) (pp-GaNTase 7). | |||||
|
GALT9_HUMAN
|
||||||
| NC score | 0.970362 (rank : 31) | θ value | 1.5519e-76 (rank : 32) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9HCQ5, Q6NT54, Q8NFR1 | Gene names | GALNT9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polypeptide N-acetylgalactosaminyltransferase 9 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 9) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 9) (Polypeptide GalNAc transferase 9) (GalNAc-T9) (pp-GaNTase 9). | |||||
|
GLTL3_HUMAN
|
||||||
| NC score | 0.968970 (rank : 32) | θ value | 1.49966e-79 (rank : 28) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q6IS24, Q8NFV9, Q9NTA8 | Gene names | WBSCR17, GALNTL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative polypeptide N-acetylgalactosaminyltransferase-like protein 3 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase-like protein 3) (UDP-GalNAc:polypeptide N-acetylgalactosaminyltransferase- like protein 3) (Polypeptide GalNAc transferase-like protein 3) (GalNAc-T-like protein 3) (pp-GaNTase-like protein 3) (Williams Beuren syndrome chromosome region 17 protein). | |||||
|
GLTL3_MOUSE
|
||||||
| NC score | 0.968852 (rank : 33) | θ value | 3.34088e-79 (rank : 29) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q7TT15, Q8BKN7, Q8BUY1, Q8BX73, Q8BZC8, Q8K483 | Gene names | Wbscr17, Galntl3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative polypeptide N-acetylgalactosaminyltransferase-like protein 3 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase-like protein 3) (UDP-GalNAc:polypeptide N-acetylgalactosaminyltransferase- like protein 3) (Polypeptide GalNAc transferase-like protein 3) (GalNAc-T-like protein 3) (pp-GaNTase-like protein 3) (Williams-Beuren syndrome chromosome region 17 protein homolog). | |||||
|
GALT8_HUMAN
|
||||||
| NC score | 0.968463 (rank : 34) | θ value | 1.40689e-69 (rank : 36) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9NY28 | Gene names | GALNT8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable polypeptide N-acetylgalactosaminyltransferase 8 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase 8) (UDP- GalNAc:polypeptide N-acetylgalactosaminyltransferase 8) (Polypeptide GalNAc transferase 8) (GalNAc-T8) (pp-GaNTase 8). | |||||
|
GLTL4_HUMAN
|
||||||
| NC score | 0.966725 (rank : 35) | θ value | 7.70199e-76 (rank : 34) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q6P9A2, O95903, Q8NDY9 | Gene names | GALNTL4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative polypeptide N-acetylgalactosaminyltransferase-like protein 4 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase-like protein 4) (UDP-GalNAc:polypeptide N-acetylgalactosaminyltransferase- like protein 4) (Polypeptide GalNAc transferase-like protein 4) (GalNAc-T-like protein 4) (pp-GaNTase-like protein 4). | |||||
|
GLTL4_MOUSE
|
||||||
| NC score | 0.965066 (rank : 36) | θ value | 1.96316e-71 (rank : 35) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8K1B9 | Gene names | Galntl4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative polypeptide N-acetylgalactosaminyltransferase-like protein 4 (EC 2.4.1.41) (Protein-UDP acetylgalactosaminyltransferase-like protein 4) (UDP-GalNAc:polypeptide N-acetylgalactosaminyltransferase- like protein 4) (Polypeptide GalNAc transferase-like protein 4) (GalNAc-T-like protein 4) (pp-GaNTase-like protein 4). | |||||
|
DPM1_MOUSE
|
||||||
| NC score | 0.078554 (rank : 37) | θ value | θ > 10 (rank : 95) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O70152, Q9D829 | Gene names | Dpm1 | |||
|
Domain Architecture |

|
|||||
| Description | Dolichol-phosphate mannosyltransferase (EC 2.4.1.83) (Dolichol- phosphate mannose synthase) (Dolichyl-phosphate beta-D- mannosyltransferase) (Mannose-P-dolichol synthase) (MPD synthase) (DPM synthase). | |||||
|
B4GN4_MOUSE
|
||||||
| NC score | 0.073879 (rank : 38) | θ value | 2.36792 (rank : 67) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q766D5, Q8CHV2 | Gene names | B4galnt4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyl-beta-glucosaminyl-glycoprotein 4-beta-N- acetylgalactosaminyltransferase 1 (EC 2.4.1.244) (NGalNAc-T1) (Beta- 1,4-N-acetylgalactosaminyltransferase IV) (Beta4GalNAc-T4) (Beta4GalNAcT4). | |||||
|
B4GN4_HUMAN
|
||||||
| NC score | 0.055014 (rank : 39) | θ value | 3.0926 (rank : 70) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q76KP1, Q96LV2 | Gene names | B4GALNT4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyl-beta-glucosaminyl-glycoprotein 4-beta-N- acetylgalactosaminyltransferase 1 (EC 2.4.1.244) (NGalNAc-T1) (Beta- 1,4-N-acetylgalactosaminyltransferase IV) (Beta4GalNAc-T4) (Beta4GalNAcT4). | |||||
|
ICAL_MOUSE
|
||||||
| NC score | 0.028553 (rank : 40) | θ value | 0.0252991 (rank : 39) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P51125, Q9EQV4, Q9EQV5, Q9QXQ3, Q9QXQ4, Q9R0N1 | Gene names | Cast | |||
|
Domain Architecture |

|
|||||
| Description | Calpastatin (Calpain inhibitor). | |||||
|
SRRM2_HUMAN
|
||||||
| NC score | 0.023440 (rank : 41) | θ value | 0.21417 (rank : 45) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
DNMT1_HUMAN
|
||||||
| NC score | 0.022683 (rank : 42) | θ value | 0.0252991 (rank : 38) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 401 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P26358, Q9UHG5, Q9ULA2, Q9UMZ6 | Gene names | DNMT1, AIM, DNMT | |||
|
Domain Architecture |

|
|||||
| Description | DNA (cytosine-5)-methyltransferase 1 (EC 2.1.1.37) (Dnmt1) (DNA methyltransferase HsaI) (DNA MTase HsaI) (MCMT) (M.HsaI). | |||||
|
H13_HUMAN
|
||||||
| NC score | 0.020538 (rank : 43) | θ value | 0.0736092 (rank : 40) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P16402 | Gene names | HIST1H1D, H1F3 | |||
|
Domain Architecture |

|
|||||
| Description | Histone H1.3 (Histone H1c). | |||||
|
TM108_MOUSE
|
||||||
| NC score | 0.019624 (rank : 44) | θ value | 0.365318 (rank : 50) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 336 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8BHE4, Q80WR9 | Gene names | Tmem108 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein 108 precursor. | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.019259 (rank : 45) | θ value | 0.0961366 (rank : 42) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
LARP4_MOUSE
|
||||||
| NC score | 0.018924 (rank : 46) | θ value | 0.279714 (rank : 47) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8BWW4, Q3UR69 | Gene names | Larp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | La-related protein 4 (La ribonucleoprotein domain family member 4). | |||||
|
NOLC1_HUMAN
|
||||||
| NC score | 0.018675 (rank : 47) | θ value | 0.163984 (rank : 43) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q14978, Q15030 | Gene names | NOLC1, KIAA0035 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar phosphoprotein p130 (Nucleolar 130 kDa protein) (140 kDa nucleolar phosphoprotein) (Nopp140) (Nucleolar and coiled-body phosphoprotein 1). | |||||
|
TM131_MOUSE
|
||||||
| NC score | 0.018647 (rank : 48) | θ value | 0.813845 (rank : 55) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O70472 | Gene names | Tmem131, D1Bwg0491e, Rw1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein 131 (Protein RW1). | |||||
|
CJ012_HUMAN
|
||||||
| NC score | 0.016600 (rank : 49) | θ value | 3.0926 (rank : 71) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 467 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8N655, Q9H945, Q9Y457 | Gene names | C10orf12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C10orf12. | |||||
|
TCOF_MOUSE
|
||||||
| NC score | 0.016217 (rank : 50) | θ value | 0.47712 (rank : 53) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 471 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O08784, O08857 | Gene names | Tcof1 | |||
|
Domain Architecture |

|
|||||
| Description | Treacle protein (Treacher Collins syndrome protein homolog). | |||||
|
NACA_MOUSE
|
||||||
| NC score | 0.013974 (rank : 51) | θ value | 8.99809 (rank : 93) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q60817 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha (Alpha-NAC) (Alpha-NAC/1.9.2). | |||||
|
NACA_HUMAN
|
||||||
| NC score | 0.013965 (rank : 52) | θ value | 8.99809 (rank : 92) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13765, Q53A18, Q53G46 | Gene names | NACA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha (NAC-alpha) (Alpha-NAC) (Hom s 2.02). | |||||
|
CKAP2_HUMAN
|
||||||
| NC score | 0.013931 (rank : 53) | θ value | 2.36792 (rank : 68) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8WWK9, Q3KRA5, Q5VXB4, Q8IWV5, Q8IWV6, Q96FH9, Q9H012, Q9H0D0, Q9H988, Q9HC49, Q9NVG4 | Gene names | CKAP2, LB1, TMAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoskeleton-associated protein 2 (Tumor-associated microtubule- associated protein) (CTCL tumor antigen se20-10). | |||||
|
SP17_HUMAN
|
||||||
| NC score | 0.013273 (rank : 54) | θ value | 3.0926 (rank : 74) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q15506, Q9BXF7 | Gene names | SPA17, SP17 | |||
|
Domain Architecture |

|
|||||
| Description | Sperm surface protein Sp17 (Sperm autoantigenic protein 17) (Sperm protein 17) (Sp17-1). | |||||
|
AB1IP_MOUSE
|
||||||
| NC score | 0.013170 (rank : 55) | θ value | 6.88961 (rank : 82) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8R5A3, O35329, Q8BRU0, Q99KV8 | Gene names | Apbb1ip, Prel1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 precursor protein-binding family B member 1- interacting protein (APBB1-interacting protein 1) (Proline-rich EVH1 ligand 1) (PREL-1) (Proline-rich protein 48). | |||||
|
CYLC1_HUMAN
|
||||||
| NC score | 0.013150 (rank : 56) | θ value | 0.47712 (rank : 51) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P35663, Q5JQQ9 | Gene names | CYLC1, CYL, CYL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-1 (Cylicin I) (Multiple-band polypeptide I). | |||||
|
SFRIP_HUMAN
|
||||||
| NC score | 0.012347 (rank : 57) | θ value | 5.27518 (rank : 81) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 530 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q99590, Q8IW59 | Gene names | SFRS2IP, CASP11, SIP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SFRS2-interacting protein (Splicing factor, arginine/serine-rich 2- interacting protein) (SC35-interacting protein 1) (CTD-associated SR protein 11) (Splicing regulatory protein 129) (SRrp129) (NY-REN-40 antigen). | |||||
|
H15_HUMAN
|
||||||
| NC score | 0.012142 (rank : 58) | θ value | 3.0926 (rank : 72) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P16401, Q14529 | Gene names | HIST1H1B, H1F5 | |||
|
Domain Architecture |

|
|||||
| Description | Histone H1.5 (Histone H1a). | |||||
|
BCLF1_MOUSE
|
||||||
| NC score | 0.011686 (rank : 59) | θ value | 1.38821 (rank : 62) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8K019, Q8BNZ0, Q8C2E9, Q9CSW5 | Gene names | Bclaf1, Btf, Kiaa0164 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bcl-2-associated transcription factor 1 (Btf). | |||||
|
BASP_MOUSE
|
||||||
| NC score | 0.010824 (rank : 60) | θ value | 4.03905 (rank : 75) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q91XV3 | Gene names | Basp1, Nap22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
SFRS4_MOUSE
|
||||||
| NC score | 0.010665 (rank : 61) | θ value | 0.163984 (rank : 44) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8VE97, Q9JJC3 | Gene names | Sfrs4 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 4. | |||||
|
NACAM_MOUSE
|
||||||
| NC score | 0.010253 (rank : 62) | θ value | 0.47712 (rank : 52) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
PAPOG_HUMAN
|
||||||
| NC score | 0.008918 (rank : 63) | θ value | 1.06291 (rank : 61) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9BWT3, Q969N1, Q9H8L2, Q9HAD0 | Gene names | PAPOLG, PAP2, PAPG | |||
|
Domain Architecture |

|
|||||
| Description | Poly(A) polymerase gamma (EC 2.7.7.19) (PAP gamma) (Polynucleotide adenylyltransferase gamma) (SRP RNA 3' adenylating enzyme) (Neo- poly(A) polymerase) (Neo-PAP). | |||||
|
YETS2_HUMAN
|
||||||
| NC score | 0.008515 (rank : 64) | θ value | 4.03905 (rank : 78) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9ULM3, Q641P6, Q9NW96 | Gene names | YEATS2, KIAA1197 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YEATS domain-containing protein 2. | |||||
|
PRG4_HUMAN
|
||||||
| NC score | 0.008494 (rank : 65) | θ value | 1.81305 (rank : 65) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q92954, Q6DNC4, Q6DNC5, Q6ZMZ5, Q9BX49 | Gene names | PRG4, MSF, SZP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
LAP2B_MOUSE
|
||||||
| NC score | 0.008064 (rank : 66) | θ value | 5.27518 (rank : 79) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q61029, Q61030, Q61031, Q61032 | Gene names | Tmpo, Lap2 | |||
|
Domain Architecture |

|
|||||
| Description | Lamina-associated polypeptide 2 isoforms beta/delta/epsilon/gamma (Thymopoietin isoforms beta/delta/epsilon/gamma) (TP beta/delta/epsilon/gamma). | |||||
|
PDZD6_HUMAN
|
||||||
| NC score | 0.007730 (rank : 67) | θ value | 6.88961 (rank : 87) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9ULD6, Q4W5I8, Q86V55 | Gene names | PDZD6, KIAA1284, PDZK6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing protein 6. | |||||
|
AMOL1_MOUSE
|
||||||
| NC score | 0.007533 (rank : 68) | θ value | 1.06291 (rank : 56) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9D4H4, Q571F1 | Gene names | Amotl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin-like protein 1. | |||||
|
RBBP6_HUMAN
|
||||||
| NC score | 0.007300 (rank : 69) | θ value | 6.88961 (rank : 88) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q7Z6E9, Q15290, Q6DKH4, Q6P4C2, Q6YNC9, Q7Z6E8, Q8N0V2, Q96PH3, Q9H3I8, Q9H5M5, Q9NPX4 | Gene names | RBBP6, P2PR, PACT, RBQ1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Retinoblastoma-binding protein 6 (p53-associated cellular protein of testis) (Proliferation potential-related protein) (Protein P2P-R) (Retinoblastoma-binding Q protein 1) (Protein RBQ-1). | |||||
|
PARG_MOUSE
|
||||||
| NC score | 0.006889 (rank : 70) | θ value | 5.27518 (rank : 80) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O88622, Q80YQ6, Q8CB72 | Gene names | Parg | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly(ADP-ribose) glycohydrolase (EC 3.2.1.143). | |||||
|
NMDE1_MOUSE
|
||||||
| NC score | 0.006660 (rank : 71) | θ value | 1.06291 (rank : 60) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P35436 | Gene names | Grin2a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate [NMDA] receptor subunit epsilon 1 precursor (N-methyl D- aspartate receptor subtype 2A) (NR2A) (NMDAR2A). | |||||
|
LAP2A_HUMAN
|
||||||
| NC score | 0.006012 (rank : 72) | θ value | 8.99809 (rank : 91) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P42166, P08918, P08919, Q14860, Q16295 | Gene names | TMPO, LAP2 | |||
|
Domain Architecture |

|
|||||
| Description | Lamina-associated polypeptide 2 isoform alpha (Thymopoietin isoform alpha) (TP alpha) (Thymopoietin-related peptide isoform alpha) (TPRP isoform alpha) [Contains: Thymopoietin (TP) (Splenin); Thymopentin (TP5)]. | |||||
|
FILA_HUMAN
|
||||||
| NC score | 0.005905 (rank : 73) | θ value | 6.88961 (rank : 86) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 685 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P20930, Q01720, Q5T583 | Gene names | FLG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filaggrin. | |||||
|
MILK1_MOUSE
|
||||||
| NC score | 0.005571 (rank : 74) | θ value | 3.0926 (rank : 73) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 788 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8BGT6, Q8BJ60 | Gene names | Mirab13, D15Mit260, Kiaa1668 | |||
|
Domain Architecture |

|
|||||
| Description | Molecule interacting with Rab13 (MIRab13) (MICAL-like protein 1). | |||||
|
ATRX_HUMAN
|
||||||
| NC score | 0.005511 (rank : 75) | θ value | 1.06291 (rank : 57) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 842 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P46100, P51068, Q15886, Q59FB5, Q59H31, Q5H9A2, Q5JWI4, Q7Z2J1, Q9H0Z1, Q9NTS3 | Gene names | ATRX, RAD54L, XH2 | |||
|
Domain Architecture |

|
|||||
| Description | Transcriptional regulator ATRX (EC 3.6.1.-) (ATP-dependent helicase ATRX) (X-linked helicase II) (X-linked nuclear protein) (XNP) (Znf- HX). | |||||
|
HUWE1_MOUSE
|
||||||
| NC score | 0.005508 (rank : 76) | θ value | 1.06291 (rank : 59) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q7TMY8, Q4G2Z1, Q5BMM7, Q6NS61, Q8BNJ7, Q8CFH2, Q8VD14, Q921M5, Q9R0P2 | Gene names | Huwe1, Kiaa0312, Ureb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT, UBA and WWE domain-containing protein 1 (EC 6.3.2.-) (E3 ubiquitin protein ligase URE-B1) (E3Histone). | |||||
|
NMDE1_HUMAN
|
||||||
| NC score | 0.005388 (rank : 77) | θ value | 4.03905 (rank : 77) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q12879, O00669 | Gene names | GRIN2A | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate [NMDA] receptor subunit epsilon 1 precursor (N-methyl D- aspartate receptor subtype 2A) (NR2A) (NMDAR2A) (hNR2A). | |||||
|
MYPT1_MOUSE
|
||||||
| NC score | 0.004951 (rank : 78) | θ value | 0.365318 (rank : 49) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 539 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9DBR7, Q8CBV2, Q99NB6 | Gene names | Ppp1r12a, Mypt1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein phosphatase 1 regulatory subunit 12A (Myosin phosphatase- targeting subunit 1) (Myosin phosphatase target subunit 1). | |||||
|
TTBK2_HUMAN
|
||||||
| NC score | 0.004938 (rank : 79) | θ value | 0.0736092 (rank : 41) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 822 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6IQ55, O94932, Q6ZN52, Q8IVV1 | Gene names | TTBK2, KIAA0847 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tau-tubulin kinase 2 (EC 2.7.11.1). | |||||
|
RTN4_HUMAN
|
||||||
| NC score | 0.004533 (rank : 80) | θ value | 6.88961 (rank : 89) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9NQC3, O94962, Q9BXG5, Q9H212, Q9H3I3, Q9UQ42, Q9Y293, Q9Y2Y7, Q9Y5U6 | Gene names | RTN4, KIAA0886, NOGO | |||
|
Domain Architecture |

|
|||||
| Description | Reticulon-4 (Neurite outgrowth inhibitor) (Nogo protein) (Foocen) (Neuroendocrine-specific protein) (NSP) (Neuroendocrine-specific protein C homolog) (RTN-x) (Reticulon-5). | |||||
|
UTX_HUMAN
|
||||||
| NC score | 0.004058 (rank : 81) | θ value | 6.88961 (rank : 90) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O15550 | Gene names | UTX | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitously transcribed X chromosome tetratricopeptide repeat protein (Ubiquitously transcribed TPR protein on the X chromosome). | |||||
|
MYPT2_HUMAN
|
||||||
| NC score | 0.003721 (rank : 82) | θ value | 1.38821 (rank : 63) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 707 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O60237, Q8N179, Q9HCB7, Q9HCB8 | Gene names | PPP1R12B, MYPT2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein phosphatase 1 regulatory subunit 12B (Myosin phosphatase- targeting subunit 2) (Myosin phosphatase target subunit 2). | |||||
|
SI1L3_HUMAN
|
||||||
| NC score | 0.003702 (rank : 83) | θ value | 8.99809 (rank : 94) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O60292, Q2TV87 | Gene names | SIPA1L3, KIAA0545, SPAL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Signal-induced proliferation-associated 1-like protein 3 (SPA-1-like protein 3). | |||||
|
TBX1_MOUSE
|
||||||
| NC score | 0.003343 (rank : 84) | θ value | 2.36792 (rank : 69) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P70323, Q60706, Q99MP0, Q99P22 | Gene names | Tbx1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | T-box transcription factor TBX1 (T-box protein 1) (Testis-specific T- box protein). | |||||
|
NFH_MOUSE
|
||||||
| NC score | 0.003320 (rank : 85) | θ value | 4.03905 (rank : 76) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
SPTN2_HUMAN
|
||||||
| NC score | 0.003013 (rank : 86) | θ value | 1.38821 (rank : 64) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 535 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O15020, O14872, O14873 | Gene names | SPTBN2, KIAA0302 | |||
|
Domain Architecture |

|
|||||
| Description | Spectrin beta chain, brain 2 (Spectrin, non-erythroid beta chain 2) (Beta-III spectrin). | |||||
|
CIC_HUMAN
|
||||||
| NC score | 0.002318 (rank : 87) | θ value | 6.88961 (rank : 85) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 992 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q96RK0, Q7LGI1, Q9UEG5, Q9Y6T1 | Gene names | CIC, KIAA0306 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein capicua homolog. | |||||
|
K2C1B_MOUSE
|
||||||
| NC score | 0.002099 (rank : 88) | θ value | 0.279714 (rank : 46) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 503 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6IFZ6 | Gene names | Krt1b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Keratin, type II cytoskeletal 1b (Type II keratin Kb39) (Embryonic type II keratin-1). | |||||
|
BRPF1_HUMAN
|
||||||
| NC score | 0.001727 (rank : 89) | θ value | 6.88961 (rank : 84) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P55201, Q9UHI0 | Gene names | BRPF1, BR140 | |||
|
Domain Architecture |

|
|||||
| Description | Peregrin (Bromodomain and PHD finger-containing protein 1) (BR140 protein). | |||||
|
CD2L5_HUMAN
|
||||||
| NC score | 0.001700 (rank : 90) | θ value | 0.00665767 (rank : 37) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 987 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q14004, Q6DKQ9, Q75MH4, Q75MH5, Q96JN4, Q9H4A0, Q9H4A1, Q9UDR4 | Gene names | CDC2L5, CDC2L, CHED, KIAA1791 | |||
|
Domain Architecture |

|
|||||
| Description | Cell division cycle 2-like protein kinase 5 (EC 2.7.11.22) (CDC2- related protein kinase 5) (Cholinesterase-related cell division controller). | |||||
|
CD2L5_MOUSE
|
||||||
| NC score | -0.000000 (rank : 91) | θ value | 1.06291 (rank : 58) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1011 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q69ZA1, Q80V11, Q8BZG1, Q8K0A4 | Gene names | Cdc2l5, Kiaa1791 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle 2-like protein kinase 5 (EC 2.7.11.22) (CDC2- related protein kinase 5). | |||||
|
SUHW3_HUMAN
|
||||||
| NC score | -0.000259 (rank : 92) | θ value | 0.813845 (rank : 54) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 695 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8ND82, Q9NXR3 | Gene names | SUHW3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppressor of hairy wing homolog 3. | |||||
|
MASTL_HUMAN
|
||||||
| NC score | -0.001067 (rank : 93) | θ value | 0.365318 (rank : 48) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 894 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q96GX5, Q5T8D5, Q5T8D7, Q8NCD6, Q96SJ5 | Gene names | MASTL, THC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated serine/threonine-protein kinase-like (EC 2.7.11.1). | |||||
|
REST_MOUSE
|
||||||
| NC score | -0.002161 (rank : 94) | θ value | 1.81305 (rank : 66) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 1506 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q922J3, Q571L7 | Gene names | Rsn, Kiaa4046 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Restin. | |||||
|
BMP2K_MOUSE
|
||||||
| NC score | -0.002790 (rank : 95) | θ value | 6.88961 (rank : 83) | |||
| Query Neighborhood Hits | 94 | Target Neighborhood Hits | 874 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q91Z96, Q8C8L7 | Gene names | Bmp2k, Bike | |||
|
Domain Architecture |

|
|||||
| Description | BMP-2-inducible protein kinase (EC 2.7.11.1) (BIKe). | |||||