Please be patient as the page loads
|
DGKA_MOUSE
|
||||||
| SwissProt Accessions | O88673, Q922X2 | Gene names | Dgka, Dagk1 | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase alpha (EC 2.7.1.107) (Diglyceride kinase alpha) (DGK-alpha) (DAG kinase alpha) (80 kDa diacylglycerol kinase). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
DGKA_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.982097 (rank : 2) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 114 | |
| SwissProt Accessions | P23743, O75481, O75482, O75483 | Gene names | DGKA, DAGK, DAGK1 | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase alpha (EC 2.7.1.107) (Diglyceride kinase alpha) (DGK-alpha) (DAG kinase alpha) (80 kDa diacylglycerol kinase). | |||||
|
DGKA_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 146 | |
| SwissProt Accessions | O88673, Q922X2 | Gene names | Dgka, Dagk1 | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase alpha (EC 2.7.1.107) (Diglyceride kinase alpha) (DGK-alpha) (DAG kinase alpha) (80 kDa diacylglycerol kinase). | |||||
|
DGKB_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.960040 (rank : 5) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 124 | |
| SwissProt Accessions | Q9Y6T7, O75241, Q75MF9, Q75MU7, Q86UI5, Q86UM9, Q9UQ29 | Gene names | DGKB, DAGK2, KIAA0718 | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase beta (EC 2.7.1.107) (Diglyceride kinase beta) (DGK-beta) (DAG kinase beta) (90 kDa diacylglycerol kinase). | |||||
|
DGKG_HUMAN
|
||||||
| θ value | 0 (rank : 4) | NC score | 0.963612 (rank : 4) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 104 | |
| SwissProt Accessions | P49619 | Gene names | DGKG, DAGK3 | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase gamma (EC 2.7.1.107) (Diglyceride kinase gamma) (DGK-gamma) (DAG kinase gamma). | |||||
|
DGKG_MOUSE
|
||||||
| θ value | 0 (rank : 5) | NC score | 0.966181 (rank : 3) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 110 | |
| SwissProt Accessions | Q91WG7 | Gene names | Dgkg, Dagk3 | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase gamma (EC 2.7.1.107) (Diglyceride kinase gamma) (DGK-gamma) (DAG kinase gamma) (88 kDa diacylglycerol kinase). | |||||
|
DGKI_HUMAN
|
||||||
| θ value | 5.1662e-72 (rank : 6) | NC score | 0.725792 (rank : 14) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 609 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O75912, Q9NZ49 | Gene names | DGKI | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase iota (EC 2.7.1.107) (Diglyceride kinase iota) (DGK-iota) (DAG kinase iota). | |||||
|
DGKZ_HUMAN
|
||||||
| θ value | 1.15091e-71 (rank : 7) | NC score | 0.733732 (rank : 12) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 551 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q13574, O00542 | Gene names | DGKZ, DAGK6 | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase zeta (EC 2.7.1.107) (Diglyceride kinase zeta) (DGK-zeta) (DAG kinase zeta). | |||||
|
DGKE_MOUSE
|
||||||
| θ value | 4.37348e-71 (rank : 8) | NC score | 0.887117 (rank : 7) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9R1C6 | Gene names | Dgke | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase epsilon (EC 2.7.1.107) (Diglyceride kinase epsilon) (DGK-epsilon) (DAG kinase epsilon). | |||||
|
DGKZ_MOUSE
|
||||||
| θ value | 4.37348e-71 (rank : 9) | NC score | 0.738001 (rank : 11) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 542 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q80UP3 | Gene names | Dgkz | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase zeta (EC 2.7.1.107) (Diglyceride kinase zeta) (DGK-zeta) (DAG kinase zeta). | |||||
|
DGKE_HUMAN
|
||||||
| θ value | 1.66191e-70 (rank : 10) | NC score | 0.888069 (rank : 6) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P52429, Q9UKQ3 | Gene names | DGKE, DAGK5 | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase epsilon (EC 2.7.1.107) (Diglyceride kinase epsilon) (DGK-epsilon) (DAG kinase epsilon). | |||||
|
DGKQ_HUMAN
|
||||||
| θ value | 2.84138e-62 (rank : 11) | NC score | 0.848716 (rank : 8) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P52824 | Gene names | DGKQ, DAGK4 | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase theta (EC 2.7.1.107) (Diglyceride kinase theta) (DGK-theta) (DAG kinase theta). | |||||
|
DGKH_HUMAN
|
||||||
| θ value | 3.2585e-50 (rank : 12) | NC score | 0.758871 (rank : 10) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q86XP1, Q5VZW0, Q6PI56, Q86XP2, Q8N3N0, Q8N7J9 | Gene names | DGKH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Diacylglycerol kinase eta (EC 2.7.1.107) (Diglyceride kinase eta) (DGK-eta) (DAG kinase eta). | |||||
|
DGKD_HUMAN
|
||||||
| θ value | 1.23823e-49 (rank : 13) | NC score | 0.764229 (rank : 9) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q16760, Q14158, Q6PK55, Q8NG53 | Gene names | DGKD, KIAA0145 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Diacylglycerol kinase delta (EC 2.7.1.107) (Diglyceride kinase delta) (DGK-delta) (DAG kinase delta) (130 kDa diacylglycerol kinase). | |||||
|
DGKK_HUMAN
|
||||||
| θ value | 3.15612e-45 (rank : 14) | NC score | 0.726448 (rank : 13) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 378 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q5KSL6 | Gene names | DGKK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Diacylglycerol kinase kappa (EC 2.7.1.107) (Diglyceride kinase kappa) (DGK-kappa) (DAG kinase kappa) (142 kDa diacylglycerol kinase). | |||||
|
HPCA_HUMAN
|
||||||
| θ value | 3.77169e-06 (rank : 15) | NC score | 0.320899 (rank : 21) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | P84074, P32076, P41211, P70510 | Gene names | HPCA, BDR2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuron-specific calcium-binding protein hippocalcin (Calcium-binding protein BDR-2). | |||||
|
HPCA_MOUSE
|
||||||
| θ value | 3.77169e-06 (rank : 16) | NC score | 0.320899 (rank : 22) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | P84075, P32076, P41211, P70510 | Gene names | Hpca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuron-specific calcium-binding protein hippocalcin. | |||||
|
HPCL1_MOUSE
|
||||||
| θ value | 3.77169e-06 (rank : 17) | NC score | 0.322392 (rank : 20) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | P62748, P35333 | Gene names | Hpcal1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hippocalcin-like protein 1 (Visinin-like protein 3) (VILIP-3) (Neural visinin-like protein 3) (NVL-3) (NVP-3). | |||||
|
HPCL1_HUMAN
|
||||||
| θ value | 8.40245e-06 (rank : 18) | NC score | 0.320732 (rank : 23) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | P37235, Q969S5 | Gene names | HPCAL1, BDR1 | |||
|
Domain Architecture |
|
|||||
| Description | Hippocalcin-like protein 1 (Visinin-like protein 3) (VILIP-3) (Calcium-binding protein BDR-1) (HLP2). | |||||
|
NCALD_HUMAN
|
||||||
| θ value | 1.09739e-05 (rank : 19) | NC score | 0.322661 (rank : 18) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 77 | |
| SwissProt Accessions | P61601, P29554, Q8IYC3, Q9H0W2 | Gene names | NCALD | |||
|
Domain Architecture |
|
|||||
| Description | Neurocalcin delta. | |||||
|
NCALD_MOUSE
|
||||||
| θ value | 1.09739e-05 (rank : 20) | NC score | 0.322661 (rank : 19) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 77 | |
| SwissProt Accessions | Q91X97, Q3TJS9, Q8BZN9 | Gene names | Ncald, D15Ertd412e | |||
|
Domain Architecture |
|
|||||
| Description | Neurocalcin delta. | |||||
|
CANB2_MOUSE
|
||||||
| θ value | 1.87187e-05 (rank : 21) | NC score | 0.304451 (rank : 27) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | Q63811 | Gene names | Ppp3r2 | |||
|
Domain Architecture |
|
|||||
| Description | Calcineurin subunit B isoform 2 (Protein phosphatase 2B regulatory subunit 2) (Protein phosphatase 3 regulatory subunit B beta isoform). | |||||
|
MRCKG_MOUSE
|
||||||
| θ value | 1.87187e-05 (rank : 22) | NC score | 0.077439 (rank : 145) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1466 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q80UW5 | Gene names | Cdc42bpg | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MRCK gamma (EC 2.7.11.1) (CDC42- binding protein kinase gamma) (Myotonic dystrophy kinase-related CDC42-binding kinase gamma) (Myotonic dystrophy protein kinase-like alpha) (MRCK gamma) (DMPK-like gamma). | |||||
|
CANB2_HUMAN
|
||||||
| θ value | 2.44474e-05 (rank : 23) | NC score | 0.298086 (rank : 29) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | Q96LZ3, Q7Z4V8, Q8WYJ4 | Gene names | PPP3R2, CBLP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcineurin subunit B isoform 2 (Protein phosphatase 2B regulatory subunit 2) (Protein phosphatase 3 regulatory subunit B beta isoform) (Calcineurin B-like protein) (CBLP) (CNBII). | |||||
|
GUC1B_MOUSE
|
||||||
| θ value | 2.44474e-05 (rank : 24) | NC score | 0.316686 (rank : 25) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q8VBV8 | Gene names | Guca1b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Guanylyl cyclase-activating protein 2 (GCAP 2) (Guanylate cyclase activator 1B). | |||||
|
MRCKG_HUMAN
|
||||||
| θ value | 3.19293e-05 (rank : 25) | NC score | 0.074945 (rank : 147) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1519 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q6DT37, O00565 | Gene names | CDC42BPG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MRCK gamma (EC 2.7.11.1) (CDC42- binding protein kinase gamma) (Myotonic dystrophy kinase-related CDC42-binding kinase gamma) (Myotonic dystrophy protein kinase-like alpha) (MRCK gamma) (DMPK-like gamma). | |||||
|
HPCL4_MOUSE
|
||||||
| θ value | 5.44631e-05 (rank : 26) | NC score | 0.297194 (rank : 30) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q8BGZ1 | Gene names | Hpcal4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hippocalcin-like protein 4 (Neural visinin-like protein 2) (NVP-2). | |||||
|
CHP2_MOUSE
|
||||||
| θ value | 7.1131e-05 (rank : 27) | NC score | 0.285621 (rank : 43) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q9D869 | Gene names | Chp2, Hca520 | |||
|
Domain Architecture |
|
|||||
| Description | Calcineurin B homologous protein 2 (Hepatocellular carcinoma- associated antigen 520 homolog). | |||||
|
HPCL4_HUMAN
|
||||||
| θ value | 7.1131e-05 (rank : 28) | NC score | 0.297002 (rank : 31) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q9UM19, Q5TG97, Q8N611 | Gene names | HPCAL4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hippocalcin-like protein 4 (HLP4). | |||||
|
GUC1B_HUMAN
|
||||||
| θ value | 0.000158464 (rank : 29) | NC score | 0.313813 (rank : 26) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q9UMX6, Q9NU15 | Gene names | GUCA1B, GCAP2 | |||
|
Domain Architecture |
|
|||||
| Description | Guanylyl cyclase-activating protein 2 (GCAP 2) (Guanylate cyclase activator 1B). | |||||
|
CANB1_HUMAN
|
||||||
| θ value | 0.00020696 (rank : 30) | NC score | 0.296401 (rank : 32) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | P63098, P06705, P15117, Q08044 | Gene names | PPP3R1, CNA2, CNB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcineurin subunit B isoform 1 (Protein phosphatase 2B regulatory subunit 1) (Protein phosphatase 3 regulatory subunit B alpha isoform 1). | |||||
|
CANB1_MOUSE
|
||||||
| θ value | 0.00020696 (rank : 31) | NC score | 0.296347 (rank : 33) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | Q63810 | Gene names | Ppp3r1, Cnb | |||
|
Domain Architecture |
|
|||||
| Description | Calcineurin subunit B isoform 1 (Protein phosphatase 2B regulatory subunit 1) (Protein phosphatase 3 regulatory subunit B alpha isoform 1). | |||||
|
GUC1A_MOUSE
|
||||||
| θ value | 0.00020696 (rank : 32) | NC score | 0.325436 (rank : 16) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | P43081 | Gene names | Guca1a, Gcap, Gcap1, Guca1 | |||
|
Domain Architecture |
|
|||||
| Description | Guanylyl cyclase-activating protein 1 (GCAP 1) (Guanylate cyclase activator 1A). | |||||
|
KCIP1_MOUSE
|
||||||
| θ value | 0.00020696 (rank : 33) | NC score | 0.291875 (rank : 34) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9JJ57, Q5SSA3, Q6DTJ1, Q8BGJ4, Q8C4K4, Q8CGL1, Q8K1U1, Q8K3M2 | Gene names | Kcnip1, Kchip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kv channel-interacting protein 1 (KChIP1) (A-type potassium channel modulatory protein 1) (Potassium channel-interacting protein 1). | |||||
|
CHP2_HUMAN
|
||||||
| θ value | 0.000270298 (rank : 34) | NC score | 0.272567 (rank : 48) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | O43745 | Gene names | CHP2, HCA520 | |||
|
Domain Architecture |
|
|||||
| Description | Calcineurin B homologous protein 2 (Hepatocellular carcinoma- associated antigen 520). | |||||
|
KCIP1_HUMAN
|
||||||
| θ value | 0.000270298 (rank : 35) | NC score | 0.290896 (rank : 35) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9NZI2, Q5U822 | Gene names | KCNIP1, KCHIP1, VABP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kv channel-interacting protein 1 (KChIP1) (A-type potassium channel modulatory protein 1) (Potassium channel-interacting protein 1) (Vesicle APC-binding protein). | |||||
|
VISL1_HUMAN
|
||||||
| θ value | 0.00035302 (rank : 36) | NC score | 0.289097 (rank : 36) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | P62760, P28677, P29103, P42323, Q9UM20 | Gene names | VSNL1, VISL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Visinin-like protein 1 (VILIP) (Hippocalcin-like protein 3) (HLP3). | |||||
|
VISL1_MOUSE
|
||||||
| θ value | 0.00035302 (rank : 37) | NC score | 0.289097 (rank : 37) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | P62761, P28677, P29103, P42323, Q9UM20 | Gene names | Vsnl1, Visl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Visinin-like protein 1 (VILIP) (Neural visinin-like protein 1) (NVL-1) (NVP-1). | |||||
|
GUC1A_HUMAN
|
||||||
| θ value | 0.000461057 (rank : 38) | NC score | 0.324543 (rank : 17) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | P43080, Q9NU14 | Gene names | GUCA1A, GCAP, GCAP1, GUCA1 | |||
|
Domain Architecture |
|
|||||
| Description | Guanylyl cyclase-activating protein 1 (GCAP 1) (Guanylate cyclase activator 1A). | |||||
|
CHP1_MOUSE
|
||||||
| θ value | 0.000602161 (rank : 39) | NC score | 0.272636 (rank : 47) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | P61022, Q62877, Q6ZWQ8 | Gene names | Chp, Sid470 | |||
|
Domain Architecture |
|
|||||
| Description | Calcium-binding protein p22 (Calcium-binding protein CHP) (Calcineurin homologous protein) (Sid 470). | |||||
|
KCIP2_HUMAN
|
||||||
| θ value | 0.000602161 (rank : 40) | NC score | 0.288557 (rank : 38) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q9NS61, Q7Z6F1, Q96K86, Q96T41, Q96T42, Q96T43, Q96T44, Q9H0N4, Q9HD10, Q9HD11, Q9NS60, Q9NY10, Q9NZI1 | Gene names | KCNIP2, KCHIP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kv channel-interacting protein 2 (KChIP2) (A-type potassium channel modulatory protein 2) (Potassium channel-interacting protein 2) (Cardiac voltage-gated potassium channel modulatory subunit). | |||||
|
CHP1_HUMAN
|
||||||
| θ value | 0.000786445 (rank : 41) | NC score | 0.269611 (rank : 50) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q99653 | Gene names | CHP | |||
|
Domain Architecture |
|
|||||
| Description | Calcium-binding protein p22 (Calcium-binding protein CHP) (Calcineurin homologous protein) (Calcineurin B homolog). | |||||
|
KCIP2_MOUSE
|
||||||
| θ value | 0.000786445 (rank : 42) | NC score | 0.286310 (rank : 40) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q9JJ69, Q6DTJ2, Q8K1T8, Q8K1T9, Q8K1U0, Q8VHN4, Q8VHN5, Q8VHN6, Q9JJ68 | Gene names | Kcnip2, Kchip2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kv channel-interacting protein 2 (KChIP2) (A-type potassium channel modulatory protein 2) (Potassium channel-interacting protein 2). | |||||
|
KPCG_HUMAN
|
||||||
| θ value | 0.00102713 (rank : 43) | NC score | 0.097843 (rank : 117) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1126 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P05129 | Gene names | PRKCG, PKCG | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C gamma type (EC 2.7.11.13) (PKC-gamma). | |||||
|
KPCG_MOUSE
|
||||||
| θ value | 0.00102713 (rank : 44) | NC score | 0.097507 (rank : 119) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1123 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P63318, P05697 | Gene names | Prkcg, Pkcc, Pkcg, Prkcc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein kinase C gamma type (EC 2.7.11.13) (PKC-gamma). | |||||
|
KPCZ_HUMAN
|
||||||
| θ value | 0.00102713 (rank : 45) | NC score | 0.082568 (rank : 139) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 884 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q05513, Q15207, Q5SYT5, Q969S4 | Gene names | PRKCZ, PKC2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C zeta type (EC 2.7.11.13) (nPKC-zeta). | |||||
|
KIP1_MOUSE
|
||||||
| θ value | 0.00175202 (rank : 46) | NC score | 0.330343 (rank : 15) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q9Z0F4, Q3TN80 | Gene names | Cib1, Cib, Kip, Prkdcip | |||
|
Domain Architecture |
|
|||||
| Description | Calcium and integrin-binding protein 1 (Calmyrin) (DNA-PKcs- interacting protein) (Kinase-interacting protein) (KIP) (CIB). | |||||
|
KPCL_HUMAN
|
||||||
| θ value | 0.00175202 (rank : 47) | NC score | 0.100366 (rank : 113) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 887 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P24723, Q16246 | Gene names | PRKCH, PKCL | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C eta type (EC 2.7.11.13) (nPKC-eta) (PKC-L). | |||||
|
KPCZ_MOUSE
|
||||||
| θ value | 0.00175202 (rank : 48) | NC score | 0.084032 (rank : 137) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 882 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q02956 | Gene names | Prkcz, Pkcz | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C zeta type (EC 2.7.11.13) (nPKC-zeta). | |||||
|
KPCE_MOUSE
|
||||||
| θ value | 0.00228821 (rank : 49) | NC score | 0.096303 (rank : 121) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P16054 | Gene names | Prkce, Pkce, Pkcea | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C epsilon type (EC 2.7.11.13) (nPKC-epsilon). | |||||
|
KPCI_HUMAN
|
||||||
| θ value | 0.00228821 (rank : 50) | NC score | 0.077699 (rank : 144) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 885 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P41743, Q8WW06 | Gene names | PRKCI, DXS1179E | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C iota type (EC 2.7.11.13) (nPKC-iota) (Atypical protein kinase C-lambda/iota) (aPKC-lambda/iota) (PRKC-lambda/iota). | |||||
|
KPCI_MOUSE
|
||||||
| θ value | 0.00228821 (rank : 51) | NC score | 0.077711 (rank : 143) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 888 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q62074 | Gene names | Prkci, Pkcl | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C iota type (EC 2.7.11.13) (nPKC-iota) (Atypical protein kinase C-lambda/iota) (aPKC-lambda/iota). | |||||
|
KPCD_HUMAN
|
||||||
| θ value | 0.00298849 (rank : 52) | NC score | 0.098261 (rank : 116) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 900 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q05655, Q15144 | Gene names | PRKCD | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C delta type (EC 2.7.11.13) (nPKC-delta). | |||||
|
KPCD_MOUSE
|
||||||
| θ value | 0.00298849 (rank : 53) | NC score | 0.098417 (rank : 115) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 888 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P28867, Q91V85, Q9Z333 | Gene names | Prkcd, Pkcd | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C delta type (EC 2.7.11.13) (nPKC-delta). | |||||
|
KPCE_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 54) | NC score | 0.095882 (rank : 123) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q02156, Q53SL4, Q53SM5, Q9UE81 | Gene names | PRKCE, PKCE | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C epsilon type (EC 2.7.11.13) (nPKC-epsilon). | |||||
|
KPCL_MOUSE
|
||||||
| θ value | 0.00390308 (rank : 55) | NC score | 0.097542 (rank : 118) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 891 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P23298 | Gene names | Prkch, Pkch | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C eta type (EC 2.7.11.13) (nPKC-eta) (PKC-L). | |||||
|
KIP1_HUMAN
|
||||||
| θ value | 0.00509761 (rank : 56) | NC score | 0.318805 (rank : 24) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q99828, O00693, O00735, Q6IB49, Q96J54, Q99971 | Gene names | CIB1, CIB, KIP, PRKDCIP | |||
|
Domain Architecture |
|
|||||
| Description | Calcium and integrin-binding protein 1 (Calmyrin) (DNA-PKcs- interacting protein) (Kinase-interacting protein) (KIP) (CIB) (SNK- interacting protein 2-28) (SIP2-28). | |||||
|
KPCB_HUMAN
|
||||||
| θ value | 0.00509761 (rank : 57) | NC score | 0.092428 (rank : 130) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1121 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P05771, O43744, P05127, Q15138, Q93060, Q9UE49, Q9UE50, Q9UEH8, Q9UJ30, Q9UJ33 | Gene names | PRKCB1, PKCB, PRKCB | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C beta type (EC 2.7.11.13) (PKC-beta) (PKC-B). | |||||
|
KPCB_MOUSE
|
||||||
| θ value | 0.00509761 (rank : 58) | NC score | 0.092602 (rank : 129) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1117 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P68404, P04410, P04411 | Gene names | Prkcb1, Pkcb, Prkcb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein kinase C beta type (EC 2.7.11.13) (PKC-beta) (PKC-B). | |||||
|
CALL3_HUMAN
|
||||||
| θ value | 0.00869519 (rank : 59) | NC score | 0.160292 (rank : 64) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | P27482 | Gene names | CALML3 | |||
|
Domain Architecture |
|
|||||
| Description | Calmodulin-like protein 3 (Calmodulin-related protein NB-1) (CaM-like protein) (CLP). | |||||
|
KCIP4_HUMAN
|
||||||
| θ value | 0.00869519 (rank : 60) | NC score | 0.285220 (rank : 45) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q6PIL6, Q4W5G8, Q8NEU0, Q9BWT2, Q9H294, Q9H2A4 | Gene names | KCNIP4, CALP, KCHIP4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kv channel-interacting protein 4 (KChIP4) (A-type potassium channel modulatory protein 4) (Potassium channel-interacting protein 4) (Calsenilin-like protein). | |||||
|
KCIP4_MOUSE
|
||||||
| θ value | 0.00869519 (rank : 61) | NC score | 0.285347 (rank : 44) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q6PHZ8, Q6DTJ3, Q8CAD0, Q8R4I2, Q9EQ01 | Gene names | Kcnip4, Calp, Kchip4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kv channel-interacting protein 4 (KChIP4) (A-type potassium channel modulatory protein 4) (Potassium channel-interacting protein 4) (Calsenilin-like protein). | |||||
|
NCS1_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 62) | NC score | 0.285782 (rank : 41) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | P62166, P36610, Q9UK26 | Gene names | FREQ, FLUP, NCS1 | |||
|
Domain Architecture |
|
|||||
| Description | Neuronal calcium sensor 1 (NCS-1) (Frequenin homolog) (Frequenin-like protein) (Frequenin-like ubiquitous protein). | |||||
|
NCS1_MOUSE
|
||||||
| θ value | 0.0148317 (rank : 63) | NC score | 0.285782 (rank : 42) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q8BNY6 | Gene names | Freq, Ncs1 | |||
|
Domain Architecture |
|
|||||
| Description | Neuronal calcium sensor 1 (NCS-1) (Frequenin homolog). | |||||
|
RECO_MOUSE
|
||||||
| θ value | 0.0148317 (rank : 64) | NC score | 0.288516 (rank : 39) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | P34057 | Gene names | Rcvrn, Rcv1 | |||
|
Domain Architecture |
|
|||||
| Description | Recoverin (Cancer-associated retinopathy protein) (Protein CAR) (23 kDa photoreceptor cell-specific protein). | |||||
|
GUC1C_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 65) | NC score | 0.302940 (rank : 28) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | O95843, O95844, Q9UNM0 | Gene names | GUCA1C, GCAP3 | |||
|
Domain Architecture |
|
|||||
| Description | Guanylyl cyclase-activating protein 3 (GCAP 3) (Guanylate cyclase activator 1C). | |||||
|
KPCA_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 66) | NC score | 0.088161 (rank : 135) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1067 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P20444 | Gene names | Prkca, Pkca | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C alpha type (EC 2.7.11.13) (PKC-alpha) (PKC-A). | |||||
|
KPCA_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 67) | NC score | 0.088381 (rank : 134) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1070 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P17252, Q15137, Q96RE4 | Gene names | PRKCA, PKCA, PRKACA | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C alpha type (EC 2.7.11.13) (PKC-alpha) (PKC-A). | |||||
|
CALM_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 68) | NC score | 0.161431 (rank : 62) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | P62158, P02593, P70667, P99014, Q13942, Q53S29, Q61379, Q61380, Q96HK3 | Gene names | CALM1, CALM, CAM, CAM1 | |||
|
Domain Architecture |
|
|||||
| Description | Calmodulin (CaM). | |||||
|
CALM_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 69) | NC score | 0.161431 (rank : 63) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | P62204, P02593, P70667, P99014, Q61379, Q61380 | Gene names | Calm1, Calm, Cam, Cam1 | |||
|
Domain Architecture |
|
|||||
| Description | Calmodulin (CaM). | |||||
|
CABP4_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 70) | NC score | 0.134905 (rank : 85) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 250 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | P57796, Q8N4Z2, Q8WWY5 | Gene names | CABP4 | |||
|
Domain Architecture |
|
|||||
| Description | Calcium-binding protein 4 (CaBP4). | |||||
|
CABP7_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 71) | NC score | 0.166235 (rank : 58) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q86V35 | Gene names | CABP7 | |||
|
Domain Architecture |
|
|||||
| Description | Calcium-binding protein 7 (CaBP7). | |||||
|
CABP7_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 72) | NC score | 0.166235 (rank : 59) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q91ZM8 | Gene names | Cabp7 | |||
|
Domain Architecture |
|
|||||
| Description | Calcium-binding protein 7 (CaBP7). | |||||
|
RAF1_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 73) | NC score | 0.044621 (rank : 172) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 886 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P04049 | Gene names | RAF1, RAF | |||
|
Domain Architecture |

|
|||||
| Description | RAF proto-oncogene serine/threonine-protein kinase (EC 2.7.11.1) (Raf- 1) (C-RAF) (cRaf). | |||||
|
RAF1_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 74) | NC score | 0.045023 (rank : 171) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 886 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q99N57, Q91WH1 | Gene names | Raf1, Craf | |||
|
Domain Architecture |

|
|||||
| Description | RAF proto-oncogene serine/threonine-protein kinase (EC 2.7.11.1) (Raf- 1) (C-RAF) (cRaf). | |||||
|
UBP32_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 75) | NC score | 0.132935 (rank : 87) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q8NFA0, Q9BX85, Q9Y591 | Gene names | USP32, USP10 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 32 (EC 3.1.2.15) (Ubiquitin thioesterase 32) (Ubiquitin-specific-processing protease 32) (Deubiquitinating enzyme 32) (NY-REN-60 antigen). | |||||
|
RECO_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 76) | NC score | 0.277401 (rank : 46) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | P35243 | Gene names | RCVRN, RCV1 | |||
|
Domain Architecture |
|
|||||
| Description | Recoverin (Cancer-associated retinopathy protein) (Protein CAR). | |||||
|
RGNEF_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 77) | NC score | 0.052077 (rank : 169) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 641 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P97433 | Gene names | Rgnef, Rhoip2 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-guanine nucleotide exchange factor (Rho-interacting protein 2) (RhoGEF) (RIP2). | |||||
|
KPCD3_MOUSE
|
||||||
| θ value | 0.125558 (rank : 78) | NC score | 0.068769 (rank : 151) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 901 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q8K1Y2 | Gene names | Prkd3, Prkcn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase D3 (EC 2.7.11.13) (Protein kinase C nu type) (nPKC-nu). | |||||
|
NOX5_HUMAN
|
||||||
| θ value | 0.125558 (rank : 79) | NC score | 0.151028 (rank : 69) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q96PH1, Q8TEQ1, Q8TER4, Q96PH2, Q96PJ8, Q96PJ9, Q9H6E0, Q9HAM8 | Gene names | NOX5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NADPH oxidase 5 (EC 1.6.3.-). | |||||
|
CALB2_HUMAN
|
||||||
| θ value | 0.163984 (rank : 80) | NC score | 0.135661 (rank : 83) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | P22676, Q53HD2, Q96BK4 | Gene names | CALB2, CAB29 | |||
|
Domain Architecture |
|
|||||
| Description | Calretinin (CR) (29 kDa calbindin). | |||||
|
CALB2_MOUSE
|
||||||
| θ value | 0.163984 (rank : 81) | NC score | 0.138072 (rank : 81) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q08331, Q60964, Q9JM81 | Gene names | Calb2 | |||
|
Domain Architecture |

|
|||||
| Description | Calretinin (CR). | |||||
|
DUOX2_HUMAN
|
||||||
| θ value | 0.163984 (rank : 82) | NC score | 0.161806 (rank : 61) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q9NRD8, Q9NR02, Q9UHF9 | Gene names | DUOX2, LNOX2, THOX2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dual oxidase 2 precursor (EC 1.6.3.1) (EC 1.11.1.-) (NADPH oxidase/peroxidase DUOX2) (NADPH thyroid oxidase 2) (Thyroid oxidase 2) (NADH/NADPH thyroid oxidase p138-tox) (p138 thyroid oxidase) (Large NOX 2) (Long NOX 2). | |||||
|
LAMC3_HUMAN
|
||||||
| θ value | 0.163984 (rank : 83) | NC score | 0.012760 (rank : 181) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 588 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9Y6N6 | Gene names | LAMC3 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin gamma-3 chain precursor (Laminin 12 gamma 3 subunit). | |||||
|
STAC3_HUMAN
|
||||||
| θ value | 0.163984 (rank : 84) | NC score | 0.149430 (rank : 71) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q96MF2, Q96HU5 | Gene names | STAC3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH3 and cysteine-rich domain-containing protein 3. | |||||
|
STAC3_MOUSE
|
||||||
| θ value | 0.163984 (rank : 85) | NC score | 0.149253 (rank : 72) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8BZ71 | Gene names | Stac3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH3 and cysteine-rich domain-containing protein 3. | |||||
|
DUOX1_HUMAN
|
||||||
| θ value | 0.21417 (rank : 86) | NC score | 0.162433 (rank : 60) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q9NRD9, Q6ZMB3, Q6ZR09, Q9NZC1 | Gene names | DUOX1, DUOX, LNOX1, THOX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dual oxidase 1 precursor (EC 1.6.3.1) (EC 1.11.1.-) (NADPH thyroid oxidase 1) (Thyroid oxidase 1) (Large NOX 1) (Long NOX 1). | |||||
|
EFHD1_HUMAN
|
||||||
| θ value | 0.21417 (rank : 87) | NC score | 0.135157 (rank : 84) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9BUP0, Q9BTF8, Q9H8I2, Q9HBQ0 | Gene names | EFHD1, SWS2 | |||
|
Domain Architecture |
|
|||||
| Description | EF-hand domain-containing protein 1 (Swiprosin-2). | |||||
|
CABP4_MOUSE
|
||||||
| θ value | 0.279714 (rank : 88) | NC score | 0.129137 (rank : 93) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q8VHC5 | Gene names | Cabp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcium-binding protein 4 (CaBP4). | |||||
|
CETN2_HUMAN
|
||||||
| θ value | 0.279714 (rank : 89) | NC score | 0.139698 (rank : 80) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | P41208, Q53XW1 | Gene names | CETN2, CALT, CEN2 | |||
|
Domain Architecture |
|
|||||
| Description | Centrin-2 (Caltractin isoform 1). | |||||
|
KPCD3_HUMAN
|
||||||
| θ value | 0.279714 (rank : 90) | NC score | 0.068632 (rank : 152) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 895 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O94806 | Gene names | PRKD3, EPK2, PRKCN | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase D3 (EC 2.7.11.13) (Protein kinase C nu type) (nPKC-nu) (Protein kinase EPK2). | |||||
|
KPCT_HUMAN
|
||||||
| θ value | 0.279714 (rank : 91) | NC score | 0.093711 (rank : 128) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 897 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q04759, Q3MJF1, Q64FY5, Q9H508, Q9H549 | Gene names | PRKCQ, PRKCT | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C theta type (EC 2.7.11.13) (nPKC-theta). | |||||
|
KPCT_MOUSE
|
||||||
| θ value | 0.279714 (rank : 92) | NC score | 0.094035 (rank : 126) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 895 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q02111 | Gene names | Prkcq, Pkcq | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C theta type (EC 2.7.11.13) (nPKC-theta). | |||||
|
CSEN_HUMAN
|
||||||
| θ value | 0.365318 (rank : 93) | NC score | 0.271324 (rank : 49) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q9Y2W7, Q53TJ5, Q96T40, Q9UJ84, Q9UJ85 | Gene names | KCNIP3, CSEN, DREAM, KCHIP3 | |||
|
Domain Architecture |

|
|||||
| Description | Calsenilin (DRE-antagonist modulator) (DREAM) (Kv channel-interacting protein 3) (KChIP3) (A-type potassium channel modulatory protein 3). | |||||
|
CSEN_MOUSE
|
||||||
| θ value | 0.365318 (rank : 94) | NC score | 0.268007 (rank : 52) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q9QXT8, Q924L0, Q99PH9, Q99PI0, Q99PI2, Q99PI3, Q9JHZ5 | Gene names | Kcnip3, Csen, Dream, Kchip3 | |||
|
Domain Architecture |

|
|||||
| Description | Calsenilin (DRE-antagonist modulator) (DREAM) (Kv channel-interacting protein 3) (A-type potassium channel modulatory protein 3) (KChIP3). | |||||
|
CETN3_HUMAN
|
||||||
| θ value | 0.47712 (rank : 95) | NC score | 0.132249 (rank : 89) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | O15182, Q9BS23 | Gene names | CETN3, CEN3 | |||
|
Domain Architecture |
|
|||||
| Description | Centrin-3. | |||||
|
CETN3_MOUSE
|
||||||
| θ value | 0.47712 (rank : 96) | NC score | 0.132638 (rank : 88) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | O35648 | Gene names | Cetn3, Cen3 | |||
|
Domain Architecture |
|
|||||
| Description | Centrin-3. | |||||
|
SSPO_MOUSE
|
||||||
| θ value | 0.47712 (rank : 97) | NC score | 0.006695 (rank : 188) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 800 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8CG65 | Gene names | Sspo | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SCO-spondin precursor. | |||||
|
CETN2_MOUSE
|
||||||
| θ value | 0.62314 (rank : 98) | NC score | 0.137968 (rank : 82) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q9R1K9, Q3UBB4, Q9CWM0 | Gene names | Cetn2, Calt | |||
|
Domain Architecture |
|
|||||
| Description | Centrin-2 (Caltractin isoform 1). | |||||
|
FSTL5_MOUSE
|
||||||
| θ value | 0.62314 (rank : 99) | NC score | 0.036357 (rank : 176) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 355 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8BFR2, Q80TG3, Q8C4T3 | Gene names | Fstl5, Kiaa1263 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Follistatin-related protein 5 precursor (Follistatin-like 5) (m- D/Bsp120I 1-2). | |||||
|
KPCD2_HUMAN
|
||||||
| θ value | 0.62314 (rank : 100) | NC score | 0.064952 (rank : 156) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 910 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9BZL6, Q8TB08, Q9P0T6, Q9Y3X8 | Gene names | PRKD2, PKD2 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase D2 (EC 2.7.11.13) (nPKC-D2). | |||||
|
MLL3_MOUSE
|
||||||
| θ value | 0.62314 (rank : 101) | NC score | 0.015731 (rank : 179) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
TNNC1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 102) | NC score | 0.155501 (rank : 67) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | P63316, O14800, P02590, P04463 | Gene names | TNNC1, TNNC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Troponin C, slow skeletal and cardiac muscles (TN-C). | |||||
|
CALN1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 103) | NC score | 0.119417 (rank : 95) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q9BXU9 | Gene names | CALN1, CABP8 | |||
|
Domain Architecture |
|
|||||
| Description | Calneuron-1 (Calcium-binding protein CaBP8). | |||||
|
CALN1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 104) | NC score | 0.119189 (rank : 96) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q9JJG7 | Gene names | Caln1 | |||
|
Domain Architecture |
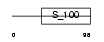
|
|||||
| Description | Calneuron-1. | |||||
|
ITB6_MOUSE
|
||||||
| θ value | 0.813845 (rank : 105) | NC score | 0.013142 (rank : 180) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9Z0T9 | Gene names | Itgb6 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-6 precursor. | |||||
|
KIP2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 106) | NC score | 0.199152 (rank : 54) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | O75838 | Gene names | CIB2, KIP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcium and integrin-binding protein 2 (Kinase-interacting protein 2) (KIP 2). | |||||
|
KPCD1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 107) | NC score | 0.065283 (rank : 154) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 891 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q62101 | Gene names | Prkd1, Pkcm, Pkd, Prkcm | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase D1 (EC 2.7.11.13) (nPKC-D1) (Protein kinase D) (Protein kinase C mu type) (nPKC-mu). | |||||
|
CETN1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 108) | NC score | 0.134078 (rank : 86) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q12798 | Gene names | CETN1, CEN1, CETN | |||
|
Domain Architecture |
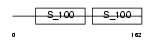
|
|||||
| Description | Centrin-1 (Caltractin isoform 2). | |||||
|
EFHD1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 109) | NC score | 0.118319 (rank : 98) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9D4J1, Q543U4 | Gene names | Efhd1, Sws2 | |||
|
Domain Architecture |
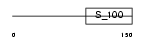
|
|||||
| Description | EF-hand domain-containing protein 1 (Swiprosin-2). | |||||
|
KPCD1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 110) | NC score | 0.064482 (rank : 157) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 901 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q15139 | Gene names | PRKD1, PKD, PKD1, PRKCM | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase D1 (EC 2.7.11.13) (nPKC-D1) (Protein kinase D) (Protein kinase C mu type) (nPKC-mu). | |||||
|
NUCB2_MOUSE
|
||||||
| θ value | 1.06291 (rank : 111) | NC score | 0.065301 (rank : 153) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 459 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P81117 | Gene names | Nucb2, Nefa | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleobindin-2 precursor (DNA-binding protein NEFA). | |||||
|
STAC_MOUSE
|
||||||
| θ value | 1.06291 (rank : 112) | NC score | 0.157756 (rank : 66) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 268 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P97306 | Gene names | Stac | |||
|
Domain Architecture |
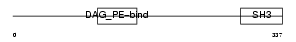
|
|||||
| Description | SH3 and cysteine-rich domain-containing protein (SRC homology 3 and cysteine-rich domain protein). | |||||
|
CETN1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 113) | NC score | 0.130710 (rank : 92) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | P41209, Q3V119, Q9D9G9, Q9DAL6 | Gene names | Cetn1, Calt | |||
|
Domain Architecture |
|
|||||
| Description | Centrin-1 (Caltractin). | |||||
|
PLST_HUMAN
|
||||||
| θ value | 1.38821 (rank : 114) | NC score | 0.051741 (rank : 170) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P13797, Q86YI6 | Gene names | PLS3 | |||
|
Domain Architecture |

|
|||||
| Description | Plastin-3 (T-plastin). | |||||
|
TNNC1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 115) | NC score | 0.154896 (rank : 68) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | P19123 | Gene names | Tnnc1, Tncc | |||
|
Domain Architecture |
|
|||||
| Description | Troponin C, slow skeletal and cardiac muscles (TN-C). | |||||
|
KIP3_HUMAN
|
||||||
| θ value | 2.36792 (rank : 116) | NC score | 0.177703 (rank : 56) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q96Q77 | Gene names | CIB3, KIP3 | |||
|
Domain Architecture |

|
|||||
| Description | Calcium and integrin-binding protein 3 (Kinase-interacting protein 3) (KIP 3). | |||||
|
LAMB2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 117) | NC score | 0.012735 (rank : 182) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 722 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q61292, Q62182 | Gene names | Lamb2, Lams | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-2 chain precursor (S-laminin) (S-LAM). | |||||
|
TCF8_MOUSE
|
||||||
| θ value | 2.36792 (rank : 118) | NC score | 0.001875 (rank : 191) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1067 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q64318, Q62519 | Gene names | Tcf8, Zfhx1a, Zfx1a, Zfx1ha | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 8 (Zinc finger homeobox protein 1a) (MEB1) (Delta EF1). | |||||
|
ZN545_HUMAN
|
||||||
| θ value | 2.36792 (rank : 119) | NC score | -0.001966 (rank : 194) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8N141, Q8NC63, Q8TF53 | Gene names | ZNF545, KIAA1948 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 545. | |||||
|
EFCB1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 120) | NC score | 0.268985 (rank : 51) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q9HAE3 | Gene names | EFCAB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EF-hand calcium-binding domain-containing protein 1. | |||||
|
PDZD8_HUMAN
|
||||||
| θ value | 3.0926 (rank : 121) | NC score | 0.118866 (rank : 97) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 185 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8NEN9, Q86WE0, Q86WE5, Q9UFF1 | Gene names | PDZD8, PDZK8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing protein 8 (Sarcoma antigen NY-SAR-84/NY-SAR- 104). | |||||
|
ARAF_HUMAN
|
||||||
| θ value | 4.03905 (rank : 122) | NC score | 0.037079 (rank : 175) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 924 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P10398, P07557, Q5H9B3 | Gene names | ARAF, ARAF1, PKS, PKS2 | |||
|
Domain Architecture |

|
|||||
| Description | A-Raf proto-oncogene serine/threonine-protein kinase (EC 2.7.11.1) (A- raf-1) (Proto-oncogene Pks). | |||||
|
ARAF_MOUSE
|
||||||
| θ value | 4.03905 (rank : 123) | NC score | 0.038013 (rank : 174) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 874 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P04627, Q99J44, Q9CTT5, Q9D6R6, Q9DBU7 | Gene names | Araf, A-raf, Araf1 | |||
|
Domain Architecture |

|
|||||
| Description | A-Raf proto-oncogene serine/threonine-protein kinase (EC 2.7.11.1). | |||||
|
CALL6_HUMAN
|
||||||
| θ value | 4.03905 (rank : 124) | NC score | 0.095026 (rank : 124) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q8TD86, Q6Q2C4 | Gene names | CALML6, CAGLP, CALGP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calmodulin-like protein 6 (Calglandulin-like protein). | |||||
|
NUCB2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 125) | NC score | 0.064000 (rank : 158) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P80303, Q8NFT5 | Gene names | NUCB2, NEFA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleobindin-2 precursor (DNA-binding protein NEFA) (Gastric cancer antigen Zg4). | |||||
|
PRDM4_HUMAN
|
||||||
| θ value | 4.03905 (rank : 126) | NC score | -0.002290 (rank : 196) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 749 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9UKN5, Q9UFA6 | Gene names | PRDM4, PFM1 | |||
|
Domain Architecture |

|
|||||
| Description | PR domain zinc finger protein 4 (PR domain-containing protein 4). | |||||
|
PRDM4_MOUSE
|
||||||
| θ value | 4.03905 (rank : 127) | NC score | -0.002279 (rank : 195) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q80V63 | Gene names | Prdm4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PR domain zinc finger protein 4 (PR domain-containing protein 4). | |||||
|
RN183_HUMAN
|
||||||
| θ value | 4.03905 (rank : 128) | NC score | 0.010556 (rank : 185) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96D59 | Gene names | RNF183 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 183. | |||||
|
STAC_HUMAN
|
||||||
| θ value | 4.03905 (rank : 129) | NC score | 0.150431 (rank : 70) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 276 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q99469 | Gene names | STAC | |||
|
Domain Architecture |
|
|||||
| Description | SH3 and cysteine-rich domain-containing protein (SRC homology 3 and cysteine-rich domain protein). | |||||
|
ZN545_MOUSE
|
||||||
| θ value | 4.03905 (rank : 130) | NC score | -0.002774 (rank : 197) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 739 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q6P9Y7, Q6ZPG0 | Gene names | Znf545, Kiaa1948, Zfp82 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 545 (Zinc finger protein 82). | |||||
|
AYTL1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 131) | NC score | 0.141986 (rank : 78) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q7L5N7, Q6MZJ6, Q9NX23 | Gene names | AYTL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acyltransferase-like 1 (EC 2.3.1.-). | |||||
|
CALL5_HUMAN
|
||||||
| θ value | 5.27518 (rank : 132) | NC score | 0.147255 (rank : 75) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q9NZT1, Q8IXU8 | Gene names | CALML5, CLSP | |||
|
Domain Architecture |
|
|||||
| Description | Calmodulin-like protein 5 (Calmodulin-like skin protein). | |||||
|
FKBP7_HUMAN
|
||||||
| θ value | 5.27518 (rank : 133) | NC score | 0.018994 (rank : 178) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Y680, Q86U65, Q96DA4, Q9Y6B0 | Gene names | FKBP7, FKBP23 | |||
|
Domain Architecture |
|
|||||
| Description | FK506-binding protein 7 precursor (EC 5.2.1.8) (Peptidyl-prolyl cis- trans isomerase) (PPIase) (Rotamase) (FKBP-23). | |||||
|
LAMB2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 134) | NC score | 0.011989 (rank : 183) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 692 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P55268, Q16321 | Gene names | LAMB2, LAMS | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-2 chain precursor (S-laminin) (Laminin B1s chain). | |||||
|
RASF1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 135) | NC score | 0.159524 (rank : 65) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q99MK9, Q9WUF5 | Gene names | Rassf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras association domain-containing protein 1 (Protein 123F2). | |||||
|
ADAM8_MOUSE
|
||||||
| θ value | 6.88961 (rank : 136) | NC score | 0.001766 (rank : 192) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 326 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q05910 | Gene names | Adam8, Ms2 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 8 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 8) (Cell surface antigen MS2) (Macrophage cysteine-rich glycoprotein) (CD156 antigen). | |||||
|
KIP2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 137) | NC score | 0.190864 (rank : 55) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q9Z309 | Gene names | Cib2, Kip2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcium and integrin-binding protein 2 (Kinase-interacting protein 2) (KIP 2). | |||||
|
LAMB1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 138) | NC score | 0.009723 (rank : 186) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1018 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P07942 | Gene names | LAMB1 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-1 chain precursor (Laminin B1 chain). | |||||
|
MRCKA_HUMAN
|
||||||
| θ value | 6.88961 (rank : 139) | NC score | 0.063348 (rank : 159) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1999 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q5VT25, Q59GZ1, Q5H9N9, Q5T797, Q5VT26, Q5VT27, Q86XX2, Q86XX3, Q99646 | Gene names | CDC42BPA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MRCK alpha (EC 2.7.11.1) (CDC42- binding protein kinase alpha) (Myotonic dystrophy kinase-related CDC42-binding kinase alpha) (Myotonic dystrophy protein kinase-like alpha) (MRCK alpha) (DMPK-like alpha). | |||||
|
PRD15_HUMAN
|
||||||
| θ value | 6.88961 (rank : 140) | NC score | -0.000314 (rank : 193) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 805 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P57071, Q8N0X3, Q8NEX0, Q9NQV3 | Gene names | PRDM15, C21orf83, ZNF298 | |||
|
Domain Architecture |

|
|||||
| Description | PR domain zinc finger protein 15 (PR domain-containing protein 15) (Zinc finger protein 298). | |||||
|
SPHK2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 141) | NC score | 0.043896 (rank : 173) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9NRA0, Q9BRN1, Q9H0Q2, Q9NWU7 | Gene names | SPHK2 | |||
|
Domain Architecture |
|
|||||
| Description | Sphingosine kinase 2 (EC 2.7.1.-) (SK 2) (SPK 2). | |||||
|
ADA20_HUMAN
|
||||||
| θ value | 8.99809 (rank : 142) | NC score | 0.002607 (rank : 190) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O43506, Q9UKJ9 | Gene names | ADAM20 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 20 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 20). | |||||
|
ADA29_HUMAN
|
||||||
| θ value | 8.99809 (rank : 143) | NC score | 0.003596 (rank : 189) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9UKF5, Q9UHP1, Q9UKF3, Q9UKF4 | Gene names | ADAM29 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 29 precursor (A disintegrin and metalloproteinase domain 29). | |||||
|
KSR2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 144) | NC score | 0.023249 (rank : 177) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 920 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q6VAB6, Q8N775 | Gene names | KSR2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinase suppressor of ras-2 (hKSR2). | |||||
|
LAMA2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 145) | NC score | 0.008312 (rank : 187) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1198 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q60675, Q05003, Q64061 | Gene names | Lama2 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-2 chain precursor (Laminin M chain) (Merosin heavy chain). | |||||
|
MEGF9_MOUSE
|
||||||
| θ value | 8.99809 (rank : 146) | NC score | 0.011833 (rank : 184) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8BH27, Q8BWI4 | Gene names | Megf9, Egfl5, Kiaa0818 | |||
|
Domain Architecture |

|
|||||
| Description | Multiple epidermal growth factor-like domains 9 precursor (EGF-like domain-containing protein 5) (Multiple EGF-like domain protein 5). | |||||
|
AYT1A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 147) | NC score | 0.131187 (rank : 91) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q8BYI6 | Gene names | Aytl1a, Aytl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acyltransferase-like 1-A (EC 2.3.1.-). | |||||
|
AYT1B_MOUSE
|
||||||
| θ value | θ > 10 (rank : 148) | NC score | 0.131842 (rank : 90) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q9D5U0 | Gene names | Aytl1b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acyltransferase-like 1-B (EC 2.3.1.-). | |||||
|
CABP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 149) | NC score | 0.104120 (rank : 110) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9NZU7, O95663, Q9NZU8 | Gene names | CABP1 | |||
|
Domain Architecture |
|
|||||
| Description | Calcium-binding protein 1 (CaBP1) (Calbrain). | |||||
|
CABP1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 150) | NC score | 0.104138 (rank : 109) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9JLK7, Q9JLK6 | Gene names | Cabp1 | |||
|
Domain Architecture |
|
|||||
| Description | Calcium-binding protein 1 (CaBP1). | |||||
|
CABP2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 151) | NC score | 0.110334 (rank : 105) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9NPB3 | Gene names | CABP2 | |||
|
Domain Architecture |
|
|||||
| Description | Calcium-binding protein 2 (CaBP2). | |||||
|
CABP2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 152) | NC score | 0.112215 (rank : 103) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q9JLK4, Q9JLK5 | Gene names | Cabp2 | |||
|
Domain Architecture |
|
|||||
| Description | Calcium-binding protein 2 (CaBP2). | |||||
|
CABP3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 153) | NC score | 0.121082 (rank : 94) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9NZU6, Q9NYY0, Q9NZT9 | Gene names | CABP3 | |||
|
Domain Architecture |
|
|||||
| Description | Calcium-binding protein 3 (CaBP3). | |||||
|
CABP5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 154) | NC score | 0.106896 (rank : 108) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q9NP86 | Gene names | CABP5 | |||
|
Domain Architecture |
|
|||||
| Description | Calcium-binding protein 5 (CaBP5). | |||||
|
CABP5_MOUSE
|
||||||
| θ value | θ > 10 (rank : 155) | NC score | 0.107886 (rank : 106) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q9JLK3 | Gene names | Cabp5 | |||
|
Domain Architecture |
|
|||||
| Description | Calcium-binding protein 5 (CaBP5). | |||||
|
CALB1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 156) | NC score | 0.114323 (rank : 99) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | P05937 | Gene names | CALB1, CAB27 | |||
|
Domain Architecture |
|
|||||
| Description | Calbindin (Vitamin D-dependent calcium-binding protein, avian-type) (Calbindin D28) (D-28K). | |||||
|
CALB1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 157) | NC score | 0.113412 (rank : 100) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | P12658, Q545N6 | Gene names | Calb1 | |||
|
Domain Architecture |
|
|||||
| Description | Calbindin (Vitamin D-dependent calcium-binding protein, avian-type) (Calbindin D28) (D-28K) (Spot 35 protein) (PCD-29). | |||||
|
CALM4_MOUSE
|
||||||
| θ value | θ > 10 (rank : 158) | NC score | 0.149247 (rank : 73) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 229 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9JM83, Q9CR31, Q9D1E9 | Gene names | Calm4, Dd112 | |||
|
Domain Architecture |
|
|||||
| Description | Calmodulin-4 (Calcium-binding protein Dd112). | |||||
|
CAYP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 159) | NC score | 0.112860 (rank : 101) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q13938, Q3MJA7, Q8WUZ3 | Gene names | CAPS | |||
|
Domain Architecture |
|
|||||
| Description | Calcyphosin (Calcyphosine). | |||||
|
CHIN_HUMAN
|
||||||
| θ value | θ > 10 (rank : 160) | NC score | 0.099946 (rank : 114) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P15882, Q53SD6, Q53SH5, Q96FB0 | Gene names | CHN1, ARHGAP2, CHN | |||
|
Domain Architecture |
|
|||||
| Description | N-chimaerin (NC) (N-chimerin) (Alpha chimerin) (A-chimaerin) (Rho- GTPase-activating protein 2). | |||||
|
CHIN_MOUSE
|
||||||
| θ value | θ > 10 (rank : 161) | NC score | 0.094972 (rank : 125) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 229 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q91V57, Q3UGY3, Q7TQE5, Q8BWU6, Q9D9B3 | Gene names | Chn1, Arhgap2 | |||
|
Domain Architecture |
|
|||||
| Description | N-chimaerin (NC) (N-chimerin) (Alpha chimerin) (A-chimaerin) (Rho- GTPase-activating protein 2). | |||||
|
CHIO_HUMAN
|
||||||
| θ value | θ > 10 (rank : 162) | NC score | 0.101560 (rank : 112) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 220 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P52757, Q75MM2 | Gene names | CHN2, ARHGAP3, BCH | |||
|
Domain Architecture |
|
|||||
| Description | Beta-chimaerin (Beta-chimerin) (Rho-GTPase-activating protein 3). | |||||
|
CHIO_MOUSE
|
||||||
| θ value | θ > 10 (rank : 163) | NC score | 0.107371 (rank : 107) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q80XD1, Q9D9W2, Q9ER57 | Gene names | Chn2, Arhgap3, Bch | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-chimaerin (Beta-chimerin) (Rho-GTPase-activating protein 3). | |||||
|
EFCB1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 164) | NC score | 0.250127 (rank : 53) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q9D3N2, Q8BSW3, Q8C982, Q8CAI0, Q9D527, Q9D5E7 | Gene names | Efcab1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EF-hand calcium-binding domain-containing protein 1. | |||||
|
EFHB_HUMAN
|
||||||
| θ value | θ > 10 (rank : 165) | NC score | 0.056296 (rank : 165) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q8N7U6, Q6ZWK9, Q8IV58, Q96LQ6 | Gene names | EFHB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EF-hand domain-containing family member B. | |||||
|
EFHD2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 166) | NC score | 0.073814 (rank : 149) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q96C19 | Gene names | EFHD2, SWS1 | |||
|
Domain Architecture |
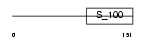
|
|||||
| Description | EF-hand domain-containing protein 2 (Swiprosin-1). | |||||
|
EFHD2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 167) | NC score | 0.080758 (rank : 142) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9D8Y0, Q9D0P4 | Gene names | Efhd2, Sws1 | |||
|
Domain Architecture |
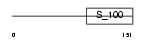
|
|||||
| Description | EF-hand domain-containing protein 2 (Swiprosin-1). | |||||
|
GRP3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 168) | NC score | 0.168572 (rank : 57) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q8IV61, O94931 | Gene names | RASGRP3, GRP3, KIAA0846 | |||
|
Domain Architecture |

|
|||||
| Description | RAS guanyl-releasing protein 3 (Calcium and DAG-regulated guanine nucleotide exchange factor III) (Guanine nucleotide exchange factor for Rap1). | |||||
|
MCFD2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 169) | NC score | 0.095896 (rank : 122) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q8NI22, Q8N3M5 | Gene names | MCFD2, SDNSF | |||
|
Domain Architecture |
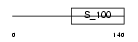
|
|||||
| Description | Multiple coagulation factor deficiency protein 2 precursor (Neural stem cell-derived neuronal survival protein). | |||||
|
MCFD2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 170) | NC score | 0.091981 (rank : 131) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8K5B2, Q3U9G5 | Gene names | Mcfd2, Sdnsf | |||
|
Domain Architecture |
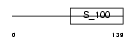
|
|||||
| Description | Multiple coagulation factor deficiency protein 2 homolog precursor (Neural stem cell-derived neuronal survival protein). | |||||
|
MLRM_HUMAN
|
||||||
| θ value | θ > 10 (rank : 171) | NC score | 0.054493 (rank : 167) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P19105 | Gene names | MRLC2, MLCB, RLC | |||
|
Domain Architecture |
|
|||||
| Description | Myosin regulatory light chain 2, nonsarcomeric (Myosin RLC). | |||||
|
MLRN_HUMAN
|
||||||
| θ value | θ > 10 (rank : 172) | NC score | 0.056684 (rank : 164) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P24844, Q9H136 | Gene names | MYL9, MLC2, MRLC1, MYRL2 | |||
|
Domain Architecture |
|
|||||
| Description | Myosin regulatory light chain 2, smooth muscle isoform (Myosin RLC) (Myosin regulatory light chain 9) (LC20). | |||||
|
MLRN_MOUSE
|
||||||
| θ value | θ > 10 (rank : 173) | NC score | 0.055620 (rank : 166) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9CQ19, Q80X77 | Gene names | Myl9, Myrl2 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin regulatory light chain 2, smooth muscle isoform (Myosin RLC) (Myosin regulatory light chain 9). | |||||
|
MRCKB_HUMAN
|
||||||
| θ value | θ > 10 (rank : 174) | NC score | 0.058524 (rank : 163) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 2239 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9Y5S2, Q2L7A5, Q86TJ1, Q9ULU5 | Gene names | CDC42BPB, KIAA1124 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MRCK beta (EC 2.7.11.1) (CDC42-binding protein kinase beta) (Myotonic dystrophy kinase-related CDC42-binding kinase beta) (Myotonic dystrophy protein kinase-like beta) (MRCK beta) (DMPK-like beta). | |||||
|
MRCKB_MOUSE
|
||||||
| θ value | θ > 10 (rank : 175) | NC score | 0.060127 (rank : 161) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 2180 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q7TT50, Q69ZR5, Q80W33 | Gene names | Cdc42bpb, Kiaa1124 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MRCK beta (EC 2.7.11.1) (CDC42-binding protein kinase beta) (Myotonic dystrophy kinase-related CDC42-binding kinase beta) (Myotonic dystrophy protein kinase-like beta) (MRCK beta) (DMPK-like beta). | |||||
|
ONCO_HUMAN
|
||||||
| θ value | θ > 10 (rank : 176) | NC score | 0.081221 (rank : 141) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P32930 | Gene names | OCM, OCMN | |||
|
Domain Architecture |
|
|||||
| Description | Oncomodulin (OM) (Parvalbumin beta). | |||||
|
ONCO_MOUSE
|
||||||
| θ value | θ > 10 (rank : 177) | NC score | 0.082269 (rank : 140) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P51879, Q62004 | Gene names | Ocm | |||
|
Domain Architecture |
|
|||||
| Description | Oncomodulin (OM) (Parvalbumin beta). | |||||
|
PCAT1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 178) | NC score | 0.093792 (rank : 127) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q8NF37, Q1HAQ1, Q7Z4G6, Q8N3U7, Q8WUL8, Q9GZW6 | Gene names | AYTL2, LPCAT1, PFAAP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 1-acylglycerophosphocholine O-acyltransferase 1 (EC 2.3.1.23) (Lung- type acyl-coa:lysophosphatidylcholine acyltransferase 1) (Acyltransferase-like 2) (Phosphonoformate immuno-associated protein 3). | |||||
|
PCAT1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 179) | NC score | 0.087740 (rank : 136) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q3TFD2, Q3TAX4, Q6NXZ6, Q8BG23, Q8BUX7, Q99JU6 | Gene names | Aytl2, Lpcat1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 1-acylglycerophosphocholine O-acyltransferase 1 (EC 2.3.1.23) (Lung- type acyl-coa:lysophosphatidylcholine acyltransferase 1) (mLPCAT1) (Acyltransferase-like 2). | |||||
|
PDCD6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 180) | NC score | 0.089516 (rank : 133) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | O75340 | Gene names | PDCD6, ALG2 | |||
|
Domain Architecture |
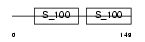
|
|||||
| Description | Programmed cell death protein 6 (Probable calcium-binding protein ALG- 2). | |||||
|
PDCD6_MOUSE
|
||||||
| θ value | θ > 10 (rank : 181) | NC score | 0.089841 (rank : 132) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | P12815, Q545I0, Q61145 | Gene names | Pdcd6, Alg2 | |||
|
Domain Architecture |
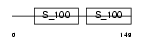
|
|||||
| Description | Programmed cell death protein 6 (Probable calcium-binding protein ALG- 2) (PMP41) (ALG-257). | |||||
|
PLSI_HUMAN
|
||||||
| θ value | θ > 10 (rank : 182) | NC score | 0.072025 (rank : 150) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 209 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q14651 | Gene names | PLS1 | |||
|
Domain Architecture |

|
|||||
| Description | Plastin-1 (I-plastin) (Intestine-specific plastin). | |||||
|
PLSL_HUMAN
|
||||||
| θ value | θ > 10 (rank : 183) | NC score | 0.059304 (rank : 162) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P13796 | Gene names | LCP1, PLS2 | |||
|
Domain Architecture |

|
|||||
| Description | Plastin-2 (L-plastin) (Lymphocyte cytosolic protein 1) (LCP-1) (LC64P). | |||||
|
PLSL_MOUSE
|
||||||
| θ value | θ > 10 (rank : 184) | NC score | 0.060309 (rank : 160) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q61233, Q3UE24, Q8R1X3, Q9CV77 | Gene names | Lcp1, Pls2 | |||
|
Domain Architecture |

|
|||||
| Description | Plastin-2 (L-plastin) (Lymphocyte cytosolic protein 1) (LCP-1) (65 kDa macrophage protein) (pp65). | |||||
|
PRVA_HUMAN
|
||||||
| θ value | θ > 10 (rank : 185) | NC score | 0.065177 (rank : 155) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P20472, P78378, Q4VB78, Q5R3Q9 | Gene names | PVALB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Parvalbumin alpha. | |||||
|
PRVA_MOUSE
|
||||||
| θ value | θ > 10 (rank : 186) | NC score | 0.075435 (rank : 146) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P32848 | Gene names | Pvalb, Pva | |||
|
Domain Architecture |
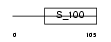
|
|||||
| Description | Parvalbumin alpha. | |||||
|
RASF1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 187) | NC score | 0.096671 (rank : 120) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9NS23, O14571, O60539, O60710, Q9HB04, Q9HB18, Q9NS22, Q9UND4, Q9UND5 | Gene names | RASSF1, RDA32 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras association domain-containing protein 1. | |||||
|
RCN2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 188) | NC score | 0.054037 (rank : 168) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q8BP92, O70341 | Gene names | Rcn2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Reticulocalbin-2 precursor (Taipoxin-associated calcium-binding protein 49) (TCBP-49). | |||||
|
SEGN_HUMAN
|
||||||
| θ value | θ > 10 (rank : 189) | NC score | 0.083472 (rank : 138) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O76038, Q5VV44, Q96QV7, Q9UJF6 | Gene names | SCGN, SECRET | |||
|
Domain Architecture |
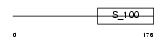
|
|||||
| Description | Secretagogin. | |||||
|
SEGN_MOUSE
|
||||||
| θ value | θ > 10 (rank : 190) | NC score | 0.074575 (rank : 148) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q91WD9 | Gene names | Scgn | |||
|
Domain Architecture |

|
|||||
| Description | Secretagogin. | |||||
|
TESC_HUMAN
|
||||||
| θ value | θ > 10 (rank : 191) | NC score | 0.145627 (rank : 76) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q96BS2 | Gene names | TESC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tescalcin (TSC). | |||||
|
TESC_MOUSE
|
||||||
| θ value | θ > 10 (rank : 192) | NC score | 0.140677 (rank : 79) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9JKL5 | Gene names | Tesc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tescalcin (TSC). | |||||
|
TNNC2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 193) | NC score | 0.145598 (rank : 77) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 234 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | P02585 | Gene names | TNNC2 | |||
|
Domain Architecture |
|
|||||
| Description | Troponin C, skeletal muscle. | |||||
|
TNNC2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 194) | NC score | 0.147941 (rank : 74) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | P20801 | Gene names | Tnnc2, Tncs | |||
|
Domain Architecture |
|
|||||
| Description | Troponin C, skeletal muscle (STNC). | |||||
|
UN13A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 195) | NC score | 0.111491 (rank : 104) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9UPW8 | Gene names | UNC13A, KIAA1032 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Unc-13 homolog A (Munc13-1). | |||||
|
UN13B_HUMAN
|
||||||
| θ value | θ > 10 (rank : 196) | NC score | 0.112218 (rank : 102) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O14795 | Gene names | UNC13B, UNC13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Unc-13 homolog B (Munc13-2) (munc13). | |||||
|
UN13B_MOUSE
|
||||||
| θ value | θ > 10 (rank : 197) | NC score | 0.102754 (rank : 111) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9Z1N9 | Gene names | Unc13b, Unc13a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Unc-13 homolog B (Munc13-2) (munc13). | |||||
|
DGKA_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 146 | |
| SwissProt Accessions | O88673, Q922X2 | Gene names | Dgka, Dagk1 | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase alpha (EC 2.7.1.107) (Diglyceride kinase alpha) (DGK-alpha) (DAG kinase alpha) (80 kDa diacylglycerol kinase). | |||||
|
DGKA_HUMAN
|
||||||
| NC score | 0.982097 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 114 | |
| SwissProt Accessions | P23743, O75481, O75482, O75483 | Gene names | DGKA, DAGK, DAGK1 | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase alpha (EC 2.7.1.107) (Diglyceride kinase alpha) (DGK-alpha) (DAG kinase alpha) (80 kDa diacylglycerol kinase). | |||||
|
DGKG_MOUSE
|
||||||
| NC score | 0.966181 (rank : 3) | θ value | 0 (rank : 5) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 110 | |
| SwissProt Accessions | Q91WG7 | Gene names | Dgkg, Dagk3 | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase gamma (EC 2.7.1.107) (Diglyceride kinase gamma) (DGK-gamma) (DAG kinase gamma) (88 kDa diacylglycerol kinase). | |||||
|
DGKG_HUMAN
|
||||||
| NC score | 0.963612 (rank : 4) | θ value | 0 (rank : 4) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 104 | |
| SwissProt Accessions | P49619 | Gene names | DGKG, DAGK3 | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase gamma (EC 2.7.1.107) (Diglyceride kinase gamma) (DGK-gamma) (DAG kinase gamma). | |||||
|
DGKB_HUMAN
|
||||||
| NC score | 0.960040 (rank : 5) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 124 | |
| SwissProt Accessions | Q9Y6T7, O75241, Q75MF9, Q75MU7, Q86UI5, Q86UM9, Q9UQ29 | Gene names | DGKB, DAGK2, KIAA0718 | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase beta (EC 2.7.1.107) (Diglyceride kinase beta) (DGK-beta) (DAG kinase beta) (90 kDa diacylglycerol kinase). | |||||
|
DGKE_HUMAN
|
||||||
| NC score | 0.888069 (rank : 6) | θ value | 1.66191e-70 (rank : 10) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P52429, Q9UKQ3 | Gene names | DGKE, DAGK5 | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase epsilon (EC 2.7.1.107) (Diglyceride kinase epsilon) (DGK-epsilon) (DAG kinase epsilon). | |||||
|
DGKE_MOUSE
|
||||||
| NC score | 0.887117 (rank : 7) | θ value | 4.37348e-71 (rank : 8) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9R1C6 | Gene names | Dgke | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase epsilon (EC 2.7.1.107) (Diglyceride kinase epsilon) (DGK-epsilon) (DAG kinase epsilon). | |||||
|
DGKQ_HUMAN
|
||||||
| NC score | 0.848716 (rank : 8) | θ value | 2.84138e-62 (rank : 11) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P52824 | Gene names | DGKQ, DAGK4 | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase theta (EC 2.7.1.107) (Diglyceride kinase theta) (DGK-theta) (DAG kinase theta). | |||||
|
DGKD_HUMAN
|
||||||
| NC score | 0.764229 (rank : 9) | θ value | 1.23823e-49 (rank : 13) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q16760, Q14158, Q6PK55, Q8NG53 | Gene names | DGKD, KIAA0145 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Diacylglycerol kinase delta (EC 2.7.1.107) (Diglyceride kinase delta) (DGK-delta) (DAG kinase delta) (130 kDa diacylglycerol kinase). | |||||
|
DGKH_HUMAN
|
||||||
| NC score | 0.758871 (rank : 10) | θ value | 3.2585e-50 (rank : 12) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q86XP1, Q5VZW0, Q6PI56, Q86XP2, Q8N3N0, Q8N7J9 | Gene names | DGKH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Diacylglycerol kinase eta (EC 2.7.1.107) (Diglyceride kinase eta) (DGK-eta) (DAG kinase eta). | |||||
|
DGKZ_MOUSE
|
||||||
| NC score | 0.738001 (rank : 11) | θ value | 4.37348e-71 (rank : 9) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 542 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q80UP3 | Gene names | Dgkz | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase zeta (EC 2.7.1.107) (Diglyceride kinase zeta) (DGK-zeta) (DAG kinase zeta). | |||||
|
DGKZ_HUMAN
|
||||||
| NC score | 0.733732 (rank : 12) | θ value | 1.15091e-71 (rank : 7) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 551 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q13574, O00542 | Gene names | DGKZ, DAGK6 | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase zeta (EC 2.7.1.107) (Diglyceride kinase zeta) (DGK-zeta) (DAG kinase zeta). | |||||
|
DGKK_HUMAN
|
||||||
| NC score | 0.726448 (rank : 13) | θ value | 3.15612e-45 (rank : 14) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 378 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q5KSL6 | Gene names | DGKK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Diacylglycerol kinase kappa (EC 2.7.1.107) (Diglyceride kinase kappa) (DGK-kappa) (DAG kinase kappa) (142 kDa diacylglycerol kinase). | |||||
|
DGKI_HUMAN
|
||||||
| NC score | 0.725792 (rank : 14) | θ value | 5.1662e-72 (rank : 6) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 609 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O75912, Q9NZ49 | Gene names | DGKI | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase iota (EC 2.7.1.107) (Diglyceride kinase iota) (DGK-iota) (DAG kinase iota). | |||||
|
KIP1_MOUSE
|
||||||
| NC score | 0.330343 (rank : 15) | θ value | 0.00175202 (rank : 46) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q9Z0F4, Q3TN80 | Gene names | Cib1, Cib, Kip, Prkdcip | |||
|
Domain Architecture |

|
|||||
| Description | Calcium and integrin-binding protein 1 (Calmyrin) (DNA-PKcs- interacting protein) (Kinase-interacting protein) (KIP) (CIB). | |||||
|
GUC1A_MOUSE
|
||||||
| NC score | 0.325436 (rank : 16) | θ value | 0.00020696 (rank : 32) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | P43081 | Gene names | Guca1a, Gcap, Gcap1, Guca1 | |||
|
Domain Architecture |

|
|||||
| Description | Guanylyl cyclase-activating protein 1 (GCAP 1) (Guanylate cyclase activator 1A). | |||||
|
GUC1A_HUMAN
|
||||||
| NC score | 0.324543 (rank : 17) | θ value | 0.000461057 (rank : 38) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | P43080, Q9NU14 | Gene names | GUCA1A, GCAP, GCAP1, GUCA1 | |||
|
Domain Architecture |

|
|||||
| Description | Guanylyl cyclase-activating protein 1 (GCAP 1) (Guanylate cyclase activator 1A). | |||||
|
NCALD_HUMAN
|
||||||
| NC score | 0.322661 (rank : 18) | θ value | 1.09739e-05 (rank : 19) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 77 | |
| SwissProt Accessions | P61601, P29554, Q8IYC3, Q9H0W2 | Gene names | NCALD | |||
|
Domain Architecture |

|
|||||
| Description | Neurocalcin delta. | |||||
|
NCALD_MOUSE
|
||||||
| NC score | 0.322661 (rank : 19) | θ value | 1.09739e-05 (rank : 20) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 77 | |
| SwissProt Accessions | Q91X97, Q3TJS9, Q8BZN9 | Gene names | Ncald, D15Ertd412e | |||
|
Domain Architecture |

|
|||||
| Description | Neurocalcin delta. | |||||
|
HPCL1_MOUSE
|
||||||
| NC score | 0.322392 (rank : 20) | θ value | 3.77169e-06 (rank : 17) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | P62748, P35333 | Gene names | Hpcal1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hippocalcin-like protein 1 (Visinin-like protein 3) (VILIP-3) (Neural visinin-like protein 3) (NVL-3) (NVP-3). | |||||
|
HPCA_HUMAN
|
||||||
| NC score | 0.320899 (rank : 21) | θ value | 3.77169e-06 (rank : 15) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | P84074, P32076, P41211, P70510 | Gene names | HPCA, BDR2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuron-specific calcium-binding protein hippocalcin (Calcium-binding protein BDR-2). | |||||
|
HPCA_MOUSE
|
||||||
| NC score | 0.320899 (rank : 22) | θ value | 3.77169e-06 (rank : 16) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | P84075, P32076, P41211, P70510 | Gene names | Hpca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuron-specific calcium-binding protein hippocalcin. | |||||
|
HPCL1_HUMAN
|
||||||
| NC score | 0.320732 (rank : 23) | θ value | 8.40245e-06 (rank : 18) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | P37235, Q969S5 | Gene names | HPCAL1, BDR1 | |||
|
Domain Architecture |

|
|||||
| Description | Hippocalcin-like protein 1 (Visinin-like protein 3) (VILIP-3) (Calcium-binding protein BDR-1) (HLP2). | |||||
|
KIP1_HUMAN
|
||||||
| NC score | 0.318805 (rank : 24) | θ value | 0.00509761 (rank : 56) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q99828, O00693, O00735, Q6IB49, Q96J54, Q99971 | Gene names | CIB1, CIB, KIP, PRKDCIP | |||
|
Domain Architecture |

|
|||||
| Description | Calcium and integrin-binding protein 1 (Calmyrin) (DNA-PKcs- interacting protein) (Kinase-interacting protein) (KIP) (CIB) (SNK- interacting protein 2-28) (SIP2-28). | |||||
|
GUC1B_MOUSE
|
||||||
| NC score | 0.316686 (rank : 25) | θ value | 2.44474e-05 (rank : 24) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q8VBV8 | Gene names | Guca1b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Guanylyl cyclase-activating protein 2 (GCAP 2) (Guanylate cyclase activator 1B). | |||||
|
GUC1B_HUMAN
|
||||||
| NC score | 0.313813 (rank : 26) | θ value | 0.000158464 (rank : 29) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q9UMX6, Q9NU15 | Gene names | GUCA1B, GCAP2 | |||
|
Domain Architecture |

|
|||||
| Description | Guanylyl cyclase-activating protein 2 (GCAP 2) (Guanylate cyclase activator 1B). | |||||
|
CANB2_MOUSE
|
||||||
| NC score | 0.304451 (rank : 27) | θ value | 1.87187e-05 (rank : 21) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | Q63811 | Gene names | Ppp3r2 | |||
|
Domain Architecture |

|
|||||
| Description | Calcineurin subunit B isoform 2 (Protein phosphatase 2B regulatory subunit 2) (Protein phosphatase 3 regulatory subunit B beta isoform). | |||||
|
GUC1C_HUMAN
|
||||||
| NC score | 0.302940 (rank : 28) | θ value | 0.0193708 (rank : 65) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | O95843, O95844, Q9UNM0 | Gene names | GUCA1C, GCAP3 | |||
|
Domain Architecture |

|
|||||
| Description | Guanylyl cyclase-activating protein 3 (GCAP 3) (Guanylate cyclase activator 1C). | |||||
|
CANB2_HUMAN
|
||||||
| NC score | 0.298086 (rank : 29) | θ value | 2.44474e-05 (rank : 23) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 75 | |
| SwissProt Accessions | Q96LZ3, Q7Z4V8, Q8WYJ4 | Gene names | PPP3R2, CBLP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcineurin subunit B isoform 2 (Protein phosphatase 2B regulatory subunit 2) (Protein phosphatase 3 regulatory subunit B beta isoform) (Calcineurin B-like protein) (CBLP) (CNBII). | |||||
|
HPCL4_MOUSE
|
||||||
| NC score | 0.297194 (rank : 30) | θ value | 5.44631e-05 (rank : 26) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q8BGZ1 | Gene names | Hpcal4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hippocalcin-like protein 4 (Neural visinin-like protein 2) (NVP-2). | |||||
|
HPCL4_HUMAN
|
||||||
| NC score | 0.297002 (rank : 31) | θ value | 7.1131e-05 (rank : 28) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q9UM19, Q5TG97, Q8N611 | Gene names | HPCAL4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hippocalcin-like protein 4 (HLP4). | |||||
|
CANB1_HUMAN
|
||||||
| NC score | 0.296401 (rank : 32) | θ value | 0.00020696 (rank : 30) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | P63098, P06705, P15117, Q08044 | Gene names | PPP3R1, CNA2, CNB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcineurin subunit B isoform 1 (Protein phosphatase 2B regulatory subunit 1) (Protein phosphatase 3 regulatory subunit B alpha isoform 1). | |||||
|
CANB1_MOUSE
|
||||||
| NC score | 0.296347 (rank : 33) | θ value | 0.00020696 (rank : 31) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 76 | |
| SwissProt Accessions | Q63810 | Gene names | Ppp3r1, Cnb | |||
|
Domain Architecture |

|
|||||
| Description | Calcineurin subunit B isoform 1 (Protein phosphatase 2B regulatory subunit 1) (Protein phosphatase 3 regulatory subunit B alpha isoform 1). | |||||
|
KCIP1_MOUSE
|
||||||
| NC score | 0.291875 (rank : 34) | θ value | 0.00020696 (rank : 33) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9JJ57, Q5SSA3, Q6DTJ1, Q8BGJ4, Q8C4K4, Q8CGL1, Q8K1U1, Q8K3M2 | Gene names | Kcnip1, Kchip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kv channel-interacting protein 1 (KChIP1) (A-type potassium channel modulatory protein 1) (Potassium channel-interacting protein 1). | |||||
|
KCIP1_HUMAN
|
||||||
| NC score | 0.290896 (rank : 35) | θ value | 0.000270298 (rank : 35) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9NZI2, Q5U822 | Gene names | KCNIP1, KCHIP1, VABP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kv channel-interacting protein 1 (KChIP1) (A-type potassium channel modulatory protein 1) (Potassium channel-interacting protein 1) (Vesicle APC-binding protein). | |||||
|
VISL1_HUMAN
|
||||||
| NC score | 0.289097 (rank : 36) | θ value | 0.00035302 (rank : 36) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | P62760, P28677, P29103, P42323, Q9UM20 | Gene names | VSNL1, VISL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Visinin-like protein 1 (VILIP) (Hippocalcin-like protein 3) (HLP3). | |||||
|
VISL1_MOUSE
|
||||||
| NC score | 0.289097 (rank : 37) | θ value | 0.00035302 (rank : 37) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | P62761, P28677, P29103, P42323, Q9UM20 | Gene names | Vsnl1, Visl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Visinin-like protein 1 (VILIP) (Neural visinin-like protein 1) (NVL-1) (NVP-1). | |||||
|
KCIP2_HUMAN
|
||||||
| NC score | 0.288557 (rank : 38) | θ value | 0.000602161 (rank : 40) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q9NS61, Q7Z6F1, Q96K86, Q96T41, Q96T42, Q96T43, Q96T44, Q9H0N4, Q9HD10, Q9HD11, Q9NS60, Q9NY10, Q9NZI1 | Gene names | KCNIP2, KCHIP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kv channel-interacting protein 2 (KChIP2) (A-type potassium channel modulatory protein 2) (Potassium channel-interacting protein 2) (Cardiac voltage-gated potassium channel modulatory subunit). | |||||
|
RECO_MOUSE
|
||||||
| NC score | 0.288516 (rank : 39) | θ value | 0.0148317 (rank : 64) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | P34057 | Gene names | Rcvrn, Rcv1 | |||
|
Domain Architecture |

|
|||||
| Description | Recoverin (Cancer-associated retinopathy protein) (Protein CAR) (23 kDa photoreceptor cell-specific protein). | |||||
|
KCIP2_MOUSE
|
||||||
| NC score | 0.286310 (rank : 40) | θ value | 0.000786445 (rank : 42) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q9JJ69, Q6DTJ2, Q8K1T8, Q8K1T9, Q8K1U0, Q8VHN4, Q8VHN5, Q8VHN6, Q9JJ68 | Gene names | Kcnip2, Kchip2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kv channel-interacting protein 2 (KChIP2) (A-type potassium channel modulatory protein 2) (Potassium channel-interacting protein 2). | |||||
|
NCS1_HUMAN
|
||||||
| NC score | 0.285782 (rank : 41) | θ value | 0.0148317 (rank : 62) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | P62166, P36610, Q9UK26 | Gene names | FREQ, FLUP, NCS1 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal calcium sensor 1 (NCS-1) (Frequenin homolog) (Frequenin-like protein) (Frequenin-like ubiquitous protein). | |||||
|
NCS1_MOUSE
|
||||||
| NC score | 0.285782 (rank : 42) | θ value | 0.0148317 (rank : 63) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q8BNY6 | Gene names | Freq, Ncs1 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal calcium sensor 1 (NCS-1) (Frequenin homolog). | |||||
|
CHP2_MOUSE
|
||||||
| NC score | 0.285621 (rank : 43) | θ value | 7.1131e-05 (rank : 27) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q9D869 | Gene names | Chp2, Hca520 | |||
|
Domain Architecture |

|
|||||
| Description | Calcineurin B homologous protein 2 (Hepatocellular carcinoma- associated antigen 520 homolog). | |||||
|
KCIP4_MOUSE
|
||||||
| NC score | 0.285347 (rank : 44) | θ value | 0.00869519 (rank : 61) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q6PHZ8, Q6DTJ3, Q8CAD0, Q8R4I2, Q9EQ01 | Gene names | Kcnip4, Calp, Kchip4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kv channel-interacting protein 4 (KChIP4) (A-type potassium channel modulatory protein 4) (Potassium channel-interacting protein 4) (Calsenilin-like protein). | |||||
|
KCIP4_HUMAN
|
||||||
| NC score | 0.285220 (rank : 45) | θ value | 0.00869519 (rank : 60) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q6PIL6, Q4W5G8, Q8NEU0, Q9BWT2, Q9H294, Q9H2A4 | Gene names | KCNIP4, CALP, KCHIP4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kv channel-interacting protein 4 (KChIP4) (A-type potassium channel modulatory protein 4) (Potassium channel-interacting protein 4) (Calsenilin-like protein). | |||||
|
RECO_HUMAN
|
||||||
| NC score | 0.277401 (rank : 46) | θ value | 0.0961366 (rank : 76) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | P35243 | Gene names | RCVRN, RCV1 | |||
|
Domain Architecture |

|
|||||
| Description | Recoverin (Cancer-associated retinopathy protein) (Protein CAR). | |||||
|
CHP1_MOUSE
|
||||||
| NC score | 0.272636 (rank : 47) | θ value | 0.000602161 (rank : 39) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | P61022, Q62877, Q6ZWQ8 | Gene names | Chp, Sid470 | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-binding protein p22 (Calcium-binding protein CHP) (Calcineurin homologous protein) (Sid 470). | |||||
|
CHP2_HUMAN
|
||||||
| NC score | 0.272567 (rank : 48) | θ value | 0.000270298 (rank : 34) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | O43745 | Gene names | CHP2, HCA520 | |||
|
Domain Architecture |

|
|||||
| Description | Calcineurin B homologous protein 2 (Hepatocellular carcinoma- associated antigen 520). | |||||
|
CSEN_HUMAN
|
||||||
| NC score | 0.271324 (rank : 49) | θ value | 0.365318 (rank : 93) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q9Y2W7, Q53TJ5, Q96T40, Q9UJ84, Q9UJ85 | Gene names | KCNIP3, CSEN, DREAM, KCHIP3 | |||
|
Domain Architecture |

|
|||||
| Description | Calsenilin (DRE-antagonist modulator) (DREAM) (Kv channel-interacting protein 3) (KChIP3) (A-type potassium channel modulatory protein 3). | |||||
|
CHP1_HUMAN
|
||||||
| NC score | 0.269611 (rank : 50) | θ value | 0.000786445 (rank : 41) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 114 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q99653 | Gene names | CHP | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-binding protein p22 (Calcium-binding protein CHP) (Calcineurin homologous protein) (Calcineurin B homolog). | |||||
|
EFCB1_HUMAN
|
||||||
| NC score | 0.268985 (rank : 51) | θ value | 3.0926 (rank : 120) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q9HAE3 | Gene names | EFCAB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EF-hand calcium-binding domain-containing protein 1. | |||||
|
CSEN_MOUSE
|
||||||
| NC score | 0.268007 (rank : 52) | θ value | 0.365318 (rank : 94) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 117 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q9QXT8, Q924L0, Q99PH9, Q99PI0, Q99PI2, Q99PI3, Q9JHZ5 | Gene names | Kcnip3, Csen, Dream, Kchip3 | |||
|
Domain Architecture |

|
|||||
| Description | Calsenilin (DRE-antagonist modulator) (DREAM) (Kv channel-interacting protein 3) (A-type potassium channel modulatory protein 3) (KChIP3). | |||||
|
EFCB1_MOUSE
|
||||||
| NC score | 0.250127 (rank : 53) | θ value | θ > 10 (rank : 164) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | Q9D3N2, Q8BSW3, Q8C982, Q8CAI0, Q9D527, Q9D5E7 | Gene names | Efcab1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EF-hand calcium-binding domain-containing protein 1. | |||||
|
KIP2_HUMAN
|
||||||
| NC score | 0.199152 (rank : 54) | θ value | 0.813845 (rank : 106) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | O75838 | Gene names | CIB2, KIP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcium and integrin-binding protein 2 (Kinase-interacting protein 2) (KIP 2). | |||||
|
KIP2_MOUSE
|
||||||
| NC score | 0.190864 (rank : 55) | θ value | 6.88961 (rank : 137) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q9Z309 | Gene names | Cib2, Kip2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcium and integrin-binding protein 2 (Kinase-interacting protein 2) (KIP 2). | |||||
|
KIP3_HUMAN
|
||||||
| NC score | 0.177703 (rank : 56) | θ value | 2.36792 (rank : 116) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | Q96Q77 | Gene names | CIB3, KIP3 | |||
|
Domain Architecture |

|
|||||
| Description | Calcium and integrin-binding protein 3 (Kinase-interacting protein 3) (KIP 3). | |||||
|
GRP3_HUMAN
|
||||||
| NC score | 0.168572 (rank : 57) | θ value | θ > 10 (rank : 168) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 58 | |
| SwissProt Accessions | Q8IV61, O94931 | Gene names | RASGRP3, GRP3, KIAA0846 | |||
|
Domain Architecture |

|
|||||
| Description | RAS guanyl-releasing protein 3 (Calcium and DAG-regulated guanine nucleotide exchange factor III) (Guanine nucleotide exchange factor for Rap1). | |||||
|
CABP7_HUMAN
|
||||||
| NC score | 0.166235 (rank : 58) | θ value | 0.0736092 (rank : 71) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q86V35 | Gene names | CABP7 | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-binding protein 7 (CaBP7). | |||||
|
CABP7_MOUSE
|
||||||
| NC score | 0.166235 (rank : 59) | θ value | 0.0736092 (rank : 72) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q91ZM8 | Gene names | Cabp7 | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-binding protein 7 (CaBP7). | |||||
|
DUOX1_HUMAN
|
||||||
| NC score | 0.162433 (rank : 60) | θ value | 0.21417 (rank : 86) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 172 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | Q9NRD9, Q6ZMB3, Q6ZR09, Q9NZC1 | Gene names | DUOX1, DUOX, LNOX1, THOX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dual oxidase 1 precursor (EC 1.6.3.1) (EC 1.11.1.-) (NADPH thyroid oxidase 1) (Thyroid oxidase 1) (Large NOX 1) (Long NOX 1). | |||||
|
DUOX2_HUMAN
|
||||||
| NC score | 0.161806 (rank : 61) | θ value | 0.163984 (rank : 82) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q9NRD8, Q9NR02, Q9UHF9 | Gene names | DUOX2, LNOX2, THOX2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dual oxidase 2 precursor (EC 1.6.3.1) (EC 1.11.1.-) (NADPH oxidase/peroxidase DUOX2) (NADPH thyroid oxidase 2) (Thyroid oxidase 2) (NADH/NADPH thyroid oxidase p138-tox) (p138 thyroid oxidase) (Large NOX 2) (Long NOX 2). | |||||
|
CALM_HUMAN
|
||||||
| NC score | 0.161431 (rank : 62) | θ value | 0.0431538 (rank : 68) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | P62158, P02593, P70667, P99014, Q13942, Q53S29, Q61379, Q61380, Q96HK3 | Gene names | CALM1, CALM, CAM, CAM1 | |||
|
Domain Architecture |

|
|||||
| Description | Calmodulin (CaM). | |||||
|
CALM_MOUSE
|
||||||
| NC score | 0.161431 (rank : 63) | θ value | 0.0431538 (rank : 69) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | P62204, P02593, P70667, P99014, Q61379, Q61380 | Gene names | Calm1, Calm, Cam, Cam1 | |||
|
Domain Architecture |

|
|||||
| Description | Calmodulin (CaM). | |||||
|
CALL3_HUMAN
|
||||||
| NC score | 0.160292 (rank : 64) | θ value | 0.00869519 (rank : 59) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 74 | |
| SwissProt Accessions | P27482 | Gene names | CALML3 | |||
|
Domain Architecture |

|
|||||
| Description | Calmodulin-like protein 3 (Calmodulin-related protein NB-1) (CaM-like protein) (CLP). | |||||
|
RASF1_MOUSE
|
||||||
| NC score | 0.159524 (rank : 65) | θ value | 5.27518 (rank : 135) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q99MK9, Q9WUF5 | Gene names | Rassf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras association domain-containing protein 1 (Protein 123F2). | |||||
|
STAC_MOUSE
|
||||||
| NC score | 0.157756 (rank : 66) | θ value | 1.06291 (rank : 112) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 268 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P97306 | Gene names | Stac | |||
|
Domain Architecture |

|
|||||
| Description | SH3 and cysteine-rich domain-containing protein (SRC homology 3 and cysteine-rich domain protein). | |||||
|
TNNC1_HUMAN
|
||||||
| NC score | 0.155501 (rank : 67) | θ value | 0.62314 (rank : 102) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | P63316, O14800, P02590, P04463 | Gene names | TNNC1, TNNC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Troponin C, slow skeletal and cardiac muscles (TN-C). | |||||
|
TNNC1_MOUSE
|
||||||
| NC score | 0.154896 (rank : 68) | θ value | 1.38821 (rank : 115) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | P19123 | Gene names | Tnnc1, Tncc | |||
|
Domain Architecture |

|
|||||
| Description | Troponin C, slow skeletal and cardiac muscles (TN-C). | |||||
|
NOX5_HUMAN
|
||||||
| NC score | 0.151028 (rank : 69) | θ value | 0.125558 (rank : 79) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q96PH1, Q8TEQ1, Q8TER4, Q96PH2, Q96PJ8, Q96PJ9, Q9H6E0, Q9HAM8 | Gene names | NOX5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NADPH oxidase 5 (EC 1.6.3.-). | |||||
|
STAC_HUMAN
|
||||||
| NC score | 0.150431 (rank : 70) | θ value | 4.03905 (rank : 129) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 276 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q99469 | Gene names | STAC | |||
|
Domain Architecture |

|
|||||
| Description | SH3 and cysteine-rich domain-containing protein (SRC homology 3 and cysteine-rich domain protein). | |||||
|
STAC3_HUMAN
|
||||||
| NC score | 0.149430 (rank : 71) | θ value | 0.163984 (rank : 84) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q96MF2, Q96HU5 | Gene names | STAC3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH3 and cysteine-rich domain-containing protein 3. | |||||
|
STAC3_MOUSE
|
||||||
| NC score | 0.149253 (rank : 72) | θ value | 0.163984 (rank : 85) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q8BZ71 | Gene names | Stac3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH3 and cysteine-rich domain-containing protein 3. | |||||
|
CALM4_MOUSE
|
||||||
| NC score | 0.149247 (rank : 73) | θ value | θ > 10 (rank : 158) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 229 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q9JM83, Q9CR31, Q9D1E9 | Gene names | Calm4, Dd112 | |||
|
Domain Architecture |

|
|||||
| Description | Calmodulin-4 (Calcium-binding protein Dd112). | |||||
|
TNNC2_MOUSE
|
||||||
| NC score | 0.147941 (rank : 74) | θ value | θ > 10 (rank : 194) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | P20801 | Gene names | Tnnc2, Tncs | |||
|
Domain Architecture |

|
|||||
| Description | Troponin C, skeletal muscle (STNC). | |||||
|
CALL5_HUMAN
|
||||||
| NC score | 0.147255 (rank : 75) | θ value | 5.27518 (rank : 132) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q9NZT1, Q8IXU8 | Gene names | CALML5, CLSP | |||
|
Domain Architecture |

|
|||||
| Description | Calmodulin-like protein 5 (Calmodulin-like skin protein). | |||||
|
TESC_HUMAN
|
||||||
| NC score | 0.145627 (rank : 76) | θ value | θ > 10 (rank : 191) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q96BS2 | Gene names | TESC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tescalcin (TSC). | |||||
|
TNNC2_HUMAN
|
||||||
| NC score | 0.145598 (rank : 77) | θ value | θ > 10 (rank : 193) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 234 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | P02585 | Gene names | TNNC2 | |||
|
Domain Architecture |

|
|||||
| Description | Troponin C, skeletal muscle. | |||||
|
AYTL1_HUMAN
|
||||||
| NC score | 0.141986 (rank : 78) | θ value | 5.27518 (rank : 131) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 70 | |
| SwissProt Accessions | Q7L5N7, Q6MZJ6, Q9NX23 | Gene names | AYTL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acyltransferase-like 1 (EC 2.3.1.-). | |||||
|
TESC_MOUSE
|
||||||
| NC score | 0.140677 (rank : 79) | θ value | θ > 10 (rank : 192) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q9JKL5 | Gene names | Tesc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tescalcin (TSC). | |||||
|
CETN2_HUMAN
|
||||||
| NC score | 0.139698 (rank : 80) | θ value | 0.279714 (rank : 89) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 213 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | P41208, Q53XW1 | Gene names | CETN2, CALT, CEN2 | |||
|
Domain Architecture |

|
|||||
| Description | Centrin-2 (Caltractin isoform 1). | |||||
|
CALB2_MOUSE
|
||||||
| NC score | 0.138072 (rank : 81) | θ value | 0.163984 (rank : 81) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q08331, Q60964, Q9JM81 | Gene names | Calb2 | |||
|
Domain Architecture |

|
|||||
| Description | Calretinin (CR). | |||||
|
CETN2_MOUSE
|
||||||
| NC score | 0.137968 (rank : 82) | θ value | 0.62314 (rank : 98) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | Q9R1K9, Q3UBB4, Q9CWM0 | Gene names | Cetn2, Calt | |||
|
Domain Architecture |

|
|||||
| Description | Centrin-2 (Caltractin isoform 1). | |||||
|
CALB2_HUMAN
|
||||||
| NC score | 0.135661 (rank : 83) | θ value | 0.163984 (rank : 80) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | P22676, Q53HD2, Q96BK4 | Gene names | CALB2, CAB29 | |||
|
Domain Architecture |

|
|||||
| Description | Calretinin (CR) (29 kDa calbindin). | |||||
|
EFHD1_HUMAN
|
||||||
| NC score | 0.135157 (rank : 84) | θ value | 0.21417 (rank : 87) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9BUP0, Q9BTF8, Q9H8I2, Q9HBQ0 | Gene names | EFHD1, SWS2 | |||
|
Domain Architecture |

|
|||||
| Description | EF-hand domain-containing protein 1 (Swiprosin-2). | |||||
|
CABP4_HUMAN
|
||||||
| NC score | 0.134905 (rank : 85) | θ value | 0.0736092 (rank : 70) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 250 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | P57796, Q8N4Z2, Q8WWY5 | Gene names | CABP4 | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-binding protein 4 (CaBP4). | |||||
|
CETN1_HUMAN
|
||||||
| NC score | 0.134078 (rank : 86) | θ value | 1.06291 (rank : 108) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q12798 | Gene names | CETN1, CEN1, CETN | |||
|
Domain Architecture |

|
|||||
| Description | Centrin-1 (Caltractin isoform 2). | |||||
|
UBP32_HUMAN
|
||||||
| NC score | 0.132935 (rank : 87) | θ value | 0.0736092 (rank : 75) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q8NFA0, Q9BX85, Q9Y591 | Gene names | USP32, USP10 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 32 (EC 3.1.2.15) (Ubiquitin thioesterase 32) (Ubiquitin-specific-processing protease 32) (Deubiquitinating enzyme 32) (NY-REN-60 antigen). | |||||
|
CETN3_MOUSE
|
||||||
| NC score | 0.132638 (rank : 88) | θ value | 0.47712 (rank : 96) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | O35648 | Gene names | Cetn3, Cen3 | |||
|
Domain Architecture |

|
|||||
| Description | Centrin-3. | |||||
|
CETN3_HUMAN
|
||||||
| NC score | 0.132249 (rank : 89) | θ value | 0.47712 (rank : 95) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 71 | |
| SwissProt Accessions | O15182, Q9BS23 | Gene names | CETN3, CEN3 | |||
|
Domain Architecture |

|
|||||
| Description | Centrin-3. | |||||
|
AYT1B_MOUSE
|
||||||
| NC score | 0.131842 (rank : 90) | θ value | θ > 10 (rank : 148) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q9D5U0 | Gene names | Aytl1b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acyltransferase-like 1-B (EC 2.3.1.-). | |||||
|
AYT1A_MOUSE
|
||||||
| NC score | 0.131187 (rank : 91) | θ value | θ > 10 (rank : 147) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 68 | |
| SwissProt Accessions | Q8BYI6 | Gene names | Aytl1a, Aytl1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acyltransferase-like 1-A (EC 2.3.1.-). | |||||
|
CETN1_MOUSE
|
||||||
| NC score | 0.130710 (rank : 92) | θ value | 1.38821 (rank : 113) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 72 | |
| SwissProt Accessions | P41209, Q3V119, Q9D9G9, Q9DAL6 | Gene names | Cetn1, Calt | |||
|
Domain Architecture |

|
|||||
| Description | Centrin-1 (Caltractin). | |||||
|
CABP4_MOUSE
|
||||||
| NC score | 0.129137 (rank : 93) | θ value | 0.279714 (rank : 88) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 69 | |
| SwissProt Accessions | Q8VHC5 | Gene names | Cabp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcium-binding protein 4 (CaBP4). | |||||
|
CABP3_HUMAN
|
||||||
| NC score | 0.121082 (rank : 94) | θ value | θ > 10 (rank : 153) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9NZU6, Q9NYY0, Q9NZT9 | Gene names | CABP3 | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-binding protein 3 (CaBP3). | |||||
|
CALN1_HUMAN
|
||||||
| NC score | 0.119417 (rank : 95) | θ value | 0.813845 (rank : 103) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q9BXU9 | Gene names | CALN1, CABP8 | |||
|
Domain Architecture |

|
|||||
| Description | Calneuron-1 (Calcium-binding protein CaBP8). | |||||
|
CALN1_MOUSE
|
||||||
| NC score | 0.119189 (rank : 96) | θ value | 0.813845 (rank : 104) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q9JJG7 | Gene names | Caln1 | |||
|
Domain Architecture |

|
|||||
| Description | Calneuron-1. | |||||
|
PDZD8_HUMAN
|
||||||
| NC score | 0.118866 (rank : 97) | θ value | 3.0926 (rank : 121) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 185 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q8NEN9, Q86WE0, Q86WE5, Q9UFF1 | Gene names | PDZD8, PDZK8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing protein 8 (Sarcoma antigen NY-SAR-84/NY-SAR- 104). | |||||
|
EFHD1_MOUSE
|
||||||
| NC score | 0.118319 (rank : 98) | θ value | 1.06291 (rank : 109) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9D4J1, Q543U4 | Gene names | Efhd1, Sws2 | |||
|
Domain Architecture |

|
|||||
| Description | EF-hand domain-containing protein 1 (Swiprosin-2). | |||||
|
CALB1_HUMAN
|
||||||
| NC score | 0.114323 (rank : 99) | θ value | θ > 10 (rank : 156) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | P05937 | Gene names | CALB1, CAB27 | |||
|
Domain Architecture |

|
|||||
| Description | Calbindin (Vitamin D-dependent calcium-binding protein, avian-type) (Calbindin D28) (D-28K). | |||||
|
CALB1_MOUSE
|
||||||
| NC score | 0.113412 (rank : 100) | θ value | θ > 10 (rank : 157) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | P12658, Q545N6 | Gene names | Calb1 | |||
|
Domain Architecture |

|
|||||
| Description | Calbindin (Vitamin D-dependent calcium-binding protein, avian-type) (Calbindin D28) (D-28K) (Spot 35 protein) (PCD-29). | |||||
|
CAYP1_HUMAN
|
||||||
| NC score | 0.112860 (rank : 101) | θ value | θ > 10 (rank : 159) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q13938, Q3MJA7, Q8WUZ3 | Gene names | CAPS | |||
|
Domain Architecture |

|
|||||
| Description | Calcyphosin (Calcyphosine). | |||||
|
UN13B_HUMAN
|
||||||
| NC score | 0.112218 (rank : 102) | θ value | θ > 10 (rank : 196) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | O14795 | Gene names | UNC13B, UNC13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Unc-13 homolog B (Munc13-2) (munc13). | |||||
|
CABP2_MOUSE
|
||||||
| NC score | 0.112215 (rank : 103) | θ value | θ > 10 (rank : 152) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q9JLK4, Q9JLK5 | Gene names | Cabp2 | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-binding protein 2 (CaBP2). | |||||
|
UN13A_HUMAN
|
||||||
| NC score | 0.111491 (rank : 104) | θ value | θ > 10 (rank : 195) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9UPW8 | Gene names | UNC13A, KIAA1032 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Unc-13 homolog A (Munc13-1). | |||||
|
CABP2_HUMAN
|
||||||
| NC score | 0.110334 (rank : 105) | θ value | θ > 10 (rank : 151) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9NPB3 | Gene names | CABP2 | |||
|
Domain Architecture |
|
|||||
| Description | Calcium-binding protein 2 (CaBP2). | |||||
|
CABP5_MOUSE
|
||||||
| NC score | 0.107886 (rank : 106) | θ value | θ > 10 (rank : 155) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 207 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q9JLK3 | Gene names | Cabp5 | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-binding protein 5 (CaBP5). | |||||
|
CHIO_MOUSE
|
||||||
| NC score | 0.107371 (rank : 107) | θ value | θ > 10 (rank : 163) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q80XD1, Q9D9W2, Q9ER57 | Gene names | Chn2, Arhgap3, Bch | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-chimaerin (Beta-chimerin) (Rho-GTPase-activating protein 3). | |||||
|
CABP5_HUMAN
|
||||||
| NC score | 0.106896 (rank : 108) | θ value | θ > 10 (rank : 154) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q9NP86 | Gene names | CABP5 | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-binding protein 5 (CaBP5). | |||||
|
CABP1_MOUSE
|
||||||
| NC score | 0.104138 (rank : 109) | θ value | θ > 10 (rank : 150) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9JLK7, Q9JLK6 | Gene names | Cabp1 | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-binding protein 1 (CaBP1). | |||||
|
CABP1_HUMAN
|
||||||
| NC score | 0.104120 (rank : 110) | θ value | θ > 10 (rank : 149) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 197 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | Q9NZU7, O95663, Q9NZU8 | Gene names | CABP1 | |||
|
Domain Architecture |

|
|||||
| Description | Calcium-binding protein 1 (CaBP1) (Calbrain). | |||||
|
UN13B_MOUSE
|
||||||
| NC score | 0.102754 (rank : 111) | θ value | θ > 10 (rank : 197) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q9Z1N9 | Gene names | Unc13b, Unc13a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Unc-13 homolog B (Munc13-2) (munc13). | |||||
|
CHIO_HUMAN
|
||||||
| NC score | 0.101560 (rank : 112) | θ value | θ > 10 (rank : 162) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 220 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P52757, Q75MM2 | Gene names | CHN2, ARHGAP3, BCH | |||
|
Domain Architecture |
|
|||||
| Description | Beta-chimaerin (Beta-chimerin) (Rho-GTPase-activating protein 3). | |||||
|
KPCL_HUMAN
|
||||||
| NC score | 0.100366 (rank : 113) | θ value | 0.00175202 (rank : 47) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 887 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P24723, Q16246 | Gene names | PRKCH, PKCL | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C eta type (EC 2.7.11.13) (nPKC-eta) (PKC-L). | |||||
|
CHIN_HUMAN
|
||||||
| NC score | 0.099946 (rank : 114) | θ value | θ > 10 (rank : 160) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P15882, Q53SD6, Q53SH5, Q96FB0 | Gene names | CHN1, ARHGAP2, CHN | |||
|
Domain Architecture |

|
|||||
| Description | N-chimaerin (NC) (N-chimerin) (Alpha chimerin) (A-chimaerin) (Rho- GTPase-activating protein 2). | |||||
|
KPCD_MOUSE
|
||||||
| NC score | 0.098417 (rank : 115) | θ value | 0.00298849 (rank : 53) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 888 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | P28867, Q91V85, Q9Z333 | Gene names | Prkcd, Pkcd | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C delta type (EC 2.7.11.13) (nPKC-delta). | |||||
|
KPCD_HUMAN
|
||||||
| NC score | 0.098261 (rank : 116) | θ value | 0.00298849 (rank : 52) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 900 | Shared Neighborhood Hits | 49 | |
| SwissProt Accessions | Q05655, Q15144 | Gene names | PRKCD | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C delta type (EC 2.7.11.13) (nPKC-delta). | |||||
|
KPCG_HUMAN
|
||||||
| NC score | 0.097843 (rank : 117) | θ value | 0.00102713 (rank : 43) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1126 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P05129 | Gene names | PRKCG, PKCG | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C gamma type (EC 2.7.11.13) (PKC-gamma). | |||||
|
KPCL_MOUSE
|
||||||
| NC score | 0.097542 (rank : 118) | θ value | 0.00390308 (rank : 55) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 891 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P23298 | Gene names | Prkch, Pkch | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C eta type (EC 2.7.11.13) (nPKC-eta) (PKC-L). | |||||
|
KPCG_MOUSE
|
||||||
| NC score | 0.097507 (rank : 119) | θ value | 0.00102713 (rank : 44) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1123 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P63318, P05697 | Gene names | Prkcg, Pkcc, Pkcg, Prkcc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein kinase C gamma type (EC 2.7.11.13) (PKC-gamma). | |||||
|
RASF1_HUMAN
|
||||||
| NC score | 0.096671 (rank : 120) | θ value | θ > 10 (rank : 187) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q9NS23, O14571, O60539, O60710, Q9HB04, Q9HB18, Q9NS22, Q9UND4, Q9UND5 | Gene names | RASSF1, RDA32 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras association domain-containing protein 1. | |||||
|
KPCE_MOUSE
|
||||||
| NC score | 0.096303 (rank : 121) | θ value | 0.00228821 (rank : 49) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P16054 | Gene names | Prkce, Pkce, Pkcea | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C epsilon type (EC 2.7.11.13) (nPKC-epsilon). | |||||
|
MCFD2_HUMAN
|
||||||
| NC score | 0.095896 (rank : 122) | θ value | θ > 10 (rank : 169) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | Q8NI22, Q8N3M5 | Gene names | MCFD2, SDNSF | |||
|
Domain Architecture |

|
|||||
| Description | Multiple coagulation factor deficiency protein 2 precursor (Neural stem cell-derived neuronal survival protein). | |||||
|
KPCE_HUMAN
|
||||||
| NC score | 0.095882 (rank : 123) | θ value | 0.00390308 (rank : 54) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q02156, Q53SL4, Q53SM5, Q9UE81 | Gene names | PRKCE, PKCE | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C epsilon type (EC 2.7.11.13) (nPKC-epsilon). | |||||
|
CALL6_HUMAN
|
||||||
| NC score | 0.095026 (rank : 124) | θ value | 4.03905 (rank : 124) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q8TD86, Q6Q2C4 | Gene names | CALML6, CAGLP, CALGP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calmodulin-like protein 6 (Calglandulin-like protein). | |||||
|
CHIN_MOUSE
|
||||||
| NC score | 0.094972 (rank : 125) | θ value | θ > 10 (rank : 161) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 229 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q91V57, Q3UGY3, Q7TQE5, Q8BWU6, Q9D9B3 | Gene names | Chn1, Arhgap2 | |||
|
Domain Architecture |

|
|||||
| Description | N-chimaerin (NC) (N-chimerin) (Alpha chimerin) (A-chimaerin) (Rho- GTPase-activating protein 2). | |||||
|
KPCT_MOUSE
|
||||||
| NC score | 0.094035 (rank : 126) | θ value | 0.279714 (rank : 92) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 895 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q02111 | Gene names | Prkcq, Pkcq | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C theta type (EC 2.7.11.13) (nPKC-theta). | |||||
|
PCAT1_HUMAN
|
||||||
| NC score | 0.093792 (rank : 127) | θ value | θ > 10 (rank : 178) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q8NF37, Q1HAQ1, Q7Z4G6, Q8N3U7, Q8WUL8, Q9GZW6 | Gene names | AYTL2, LPCAT1, PFAAP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 1-acylglycerophosphocholine O-acyltransferase 1 (EC 2.3.1.23) (Lung- type acyl-coa:lysophosphatidylcholine acyltransferase 1) (Acyltransferase-like 2) (Phosphonoformate immuno-associated protein 3). | |||||
|
KPCT_HUMAN
|
||||||
| NC score | 0.093711 (rank : 128) | θ value | 0.279714 (rank : 91) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 897 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q04759, Q3MJF1, Q64FY5, Q9H508, Q9H549 | Gene names | PRKCQ, PRKCT | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C theta type (EC 2.7.11.13) (nPKC-theta). | |||||
|
KPCB_MOUSE
|
||||||
| NC score | 0.092602 (rank : 129) | θ value | 0.00509761 (rank : 58) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1117 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P68404, P04410, P04411 | Gene names | Prkcb1, Pkcb, Prkcb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein kinase C beta type (EC 2.7.11.13) (PKC-beta) (PKC-B). | |||||
|
KPCB_HUMAN
|
||||||
| NC score | 0.092428 (rank : 130) | θ value | 0.00509761 (rank : 57) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1121 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P05771, O43744, P05127, Q15138, Q93060, Q9UE49, Q9UE50, Q9UEH8, Q9UJ30, Q9UJ33 | Gene names | PRKCB1, PKCB, PRKCB | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C beta type (EC 2.7.11.13) (PKC-beta) (PKC-B). | |||||
|
MCFD2_MOUSE
|
||||||
| NC score | 0.091981 (rank : 131) | θ value | θ > 10 (rank : 170) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | Q8K5B2, Q3U9G5 | Gene names | Mcfd2, Sdnsf | |||
|
Domain Architecture |

|
|||||
| Description | Multiple coagulation factor deficiency protein 2 homolog precursor (Neural stem cell-derived neuronal survival protein). | |||||
|
PDCD6_MOUSE
|
||||||
| NC score | 0.089841 (rank : 132) | θ value | θ > 10 (rank : 181) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | P12815, Q545I0, Q61145 | Gene names | Pdcd6, Alg2 | |||
|
Domain Architecture |

|
|||||
| Description | Programmed cell death protein 6 (Probable calcium-binding protein ALG- 2) (PMP41) (ALG-257). | |||||
|
PDCD6_HUMAN
|
||||||
| NC score | 0.089516 (rank : 133) | θ value | θ > 10 (rank : 180) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | O75340 | Gene names | PDCD6, ALG2 | |||
|
Domain Architecture |

|
|||||
| Description | Programmed cell death protein 6 (Probable calcium-binding protein ALG- 2). | |||||
|
KPCA_HUMAN
|
||||||
| NC score | 0.088381 (rank : 134) | θ value | 0.0330416 (rank : 67) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1070 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P17252, Q15137, Q96RE4 | Gene names | PRKCA, PKCA, PRKACA | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C alpha type (EC 2.7.11.13) (PKC-alpha) (PKC-A). | |||||
|
KPCA_MOUSE
|
||||||
| NC score | 0.088161 (rank : 135) | θ value | 0.0252991 (rank : 66) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1067 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P20444 | Gene names | Prkca, Pkca | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C alpha type (EC 2.7.11.13) (PKC-alpha) (PKC-A). | |||||
|
PCAT1_MOUSE
|
||||||
| NC score | 0.087740 (rank : 136) | θ value | θ > 10 (rank : 179) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q3TFD2, Q3TAX4, Q6NXZ6, Q8BG23, Q8BUX7, Q99JU6 | Gene names | Aytl2, Lpcat1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 1-acylglycerophosphocholine O-acyltransferase 1 (EC 2.3.1.23) (Lung- type acyl-coa:lysophosphatidylcholine acyltransferase 1) (mLPCAT1) (Acyltransferase-like 2). | |||||
|
KPCZ_MOUSE
|
||||||
| NC score | 0.084032 (rank : 137) | θ value | 0.00175202 (rank : 48) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 882 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q02956 | Gene names | Prkcz, Pkcz | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C zeta type (EC 2.7.11.13) (nPKC-zeta). | |||||
|
SEGN_HUMAN
|
||||||
| NC score | 0.083472 (rank : 138) | θ value | θ > 10 (rank : 189) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | O76038, Q5VV44, Q96QV7, Q9UJF6 | Gene names | SCGN, SECRET | |||
|
Domain Architecture |

|
|||||
| Description | Secretagogin. | |||||
|
KPCZ_HUMAN
|
||||||
| NC score | 0.082568 (rank : 139) | θ value | 0.00102713 (rank : 45) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 884 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q05513, Q15207, Q5SYT5, Q969S4 | Gene names | PRKCZ, PKC2 | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C zeta type (EC 2.7.11.13) (nPKC-zeta). | |||||
|
ONCO_MOUSE
|
||||||
| NC score | 0.082269 (rank : 140) | θ value | θ > 10 (rank : 177) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P51879, Q62004 | Gene names | Ocm | |||
|
Domain Architecture |

|
|||||
| Description | Oncomodulin (OM) (Parvalbumin beta). | |||||
|
ONCO_HUMAN
|
||||||
| NC score | 0.081221 (rank : 141) | θ value | θ > 10 (rank : 176) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P32930 | Gene names | OCM, OCMN | |||
|
Domain Architecture |

|
|||||
| Description | Oncomodulin (OM) (Parvalbumin beta). | |||||
|
EFHD2_MOUSE
|
||||||
| NC score | 0.080758 (rank : 142) | θ value | θ > 10 (rank : 167) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9D8Y0, Q9D0P4 | Gene names | Efhd2, Sws1 | |||
|
Domain Architecture |

|
|||||
| Description | EF-hand domain-containing protein 2 (Swiprosin-1). | |||||
|
KPCI_MOUSE
|
||||||
| NC score | 0.077711 (rank : 143) | θ value | 0.00228821 (rank : 51) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 888 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q62074 | Gene names | Prkci, Pkcl | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C iota type (EC 2.7.11.13) (nPKC-iota) (Atypical protein kinase C-lambda/iota) (aPKC-lambda/iota). | |||||
|
KPCI_HUMAN
|
||||||
| NC score | 0.077699 (rank : 144) | θ value | 0.00228821 (rank : 50) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 885 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P41743, Q8WW06 | Gene names | PRKCI, DXS1179E | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C iota type (EC 2.7.11.13) (nPKC-iota) (Atypical protein kinase C-lambda/iota) (aPKC-lambda/iota) (PRKC-lambda/iota). | |||||
|
MRCKG_MOUSE
|
||||||
| NC score | 0.077439 (rank : 145) | θ value | 1.87187e-05 (rank : 22) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1466 | Shared Neighborhood Hits | 42 | |
| SwissProt Accessions | Q80UW5 | Gene names | Cdc42bpg | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MRCK gamma (EC 2.7.11.1) (CDC42- binding protein kinase gamma) (Myotonic dystrophy kinase-related CDC42-binding kinase gamma) (Myotonic dystrophy protein kinase-like alpha) (MRCK gamma) (DMPK-like gamma). | |||||
|
PRVA_MOUSE
|
||||||
| NC score | 0.075435 (rank : 146) | θ value | θ > 10 (rank : 186) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P32848 | Gene names | Pvalb, Pva | |||
|
Domain Architecture |

|
|||||
| Description | Parvalbumin alpha. | |||||
|
MRCKG_HUMAN
|
||||||
| NC score | 0.074945 (rank : 147) | θ value | 3.19293e-05 (rank : 25) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1519 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q6DT37, O00565 | Gene names | CDC42BPG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MRCK gamma (EC 2.7.11.1) (CDC42- binding protein kinase gamma) (Myotonic dystrophy kinase-related CDC42-binding kinase gamma) (Myotonic dystrophy protein kinase-like alpha) (MRCK gamma) (DMPK-like gamma). | |||||
|
SEGN_MOUSE
|
||||||
| NC score | 0.074575 (rank : 148) | θ value | θ > 10 (rank : 190) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | Q91WD9 | Gene names | Scgn | |||
|
Domain Architecture |

|
|||||
| Description | Secretagogin. | |||||
|
EFHD2_HUMAN
|
||||||
| NC score | 0.073814 (rank : 149) | θ value | θ > 10 (rank : 166) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | Q96C19 | Gene names | EFHD2, SWS1 | |||
|
Domain Architecture |

|
|||||
| Description | EF-hand domain-containing protein 2 (Swiprosin-1). | |||||
|
PLSI_HUMAN
|
||||||
| NC score | 0.072025 (rank : 150) | θ value | θ > 10 (rank : 182) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 209 | Shared Neighborhood Hits | 64 | |
| SwissProt Accessions | Q14651 | Gene names | PLS1 | |||
|
Domain Architecture |

|
|||||
| Description | Plastin-1 (I-plastin) (Intestine-specific plastin). | |||||
|
KPCD3_MOUSE
|
||||||
| NC score | 0.068769 (rank : 151) | θ value | 0.125558 (rank : 78) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 901 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q8K1Y2 | Gene names | Prkd3, Prkcn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase D3 (EC 2.7.11.13) (Protein kinase C nu type) (nPKC-nu). | |||||
|
KPCD3_HUMAN
|
||||||
| NC score | 0.068632 (rank : 152) | θ value | 0.279714 (rank : 90) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 895 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | O94806 | Gene names | PRKD3, EPK2, PRKCN | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase D3 (EC 2.7.11.13) (Protein kinase C nu type) (nPKC-nu) (Protein kinase EPK2). | |||||
|
NUCB2_MOUSE
|
||||||
| NC score | 0.065301 (rank : 153) | θ value | 1.06291 (rank : 111) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 459 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | P81117 | Gene names | Nucb2, Nefa | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleobindin-2 precursor (DNA-binding protein NEFA). | |||||
|
KPCD1_MOUSE
|
||||||
| NC score | 0.065283 (rank : 154) | θ value | 0.813845 (rank : 107) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 891 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q62101 | Gene names | Prkd1, Pkcm, Pkd, Prkcm | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase D1 (EC 2.7.11.13) (nPKC-D1) (Protein kinase D) (Protein kinase C mu type) (nPKC-mu). | |||||
|
PRVA_HUMAN
|
||||||
| NC score | 0.065177 (rank : 155) | θ value | θ > 10 (rank : 185) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 38 | |
| SwissProt Accessions | P20472, P78378, Q4VB78, Q5R3Q9 | Gene names | PVALB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Parvalbumin alpha. | |||||
|
KPCD2_HUMAN
|
||||||
| NC score | 0.064952 (rank : 156) | θ value | 0.62314 (rank : 100) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 910 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q9BZL6, Q8TB08, Q9P0T6, Q9Y3X8 | Gene names | PRKD2, PKD2 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase D2 (EC 2.7.11.13) (nPKC-D2). | |||||
|
KPCD1_HUMAN
|
||||||
| NC score | 0.064482 (rank : 157) | θ value | 1.06291 (rank : 110) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 901 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q15139 | Gene names | PRKD1, PKD, PKD1, PRKCM | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase D1 (EC 2.7.11.13) (nPKC-D1) (Protein kinase D) (Protein kinase C mu type) (nPKC-mu). | |||||
|
NUCB2_HUMAN
|
||||||
| NC score | 0.064000 (rank : 158) | θ value | 4.03905 (rank : 125) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | P80303, Q8NFT5 | Gene names | NUCB2, NEFA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleobindin-2 precursor (DNA-binding protein NEFA) (Gastric cancer antigen Zg4). | |||||
|
MRCKA_HUMAN
|
||||||
| NC score | 0.063348 (rank : 159) | θ value | 6.88961 (rank : 139) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1999 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | Q5VT25, Q59GZ1, Q5H9N9, Q5T797, Q5VT26, Q5VT27, Q86XX2, Q86XX3, Q99646 | Gene names | CDC42BPA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MRCK alpha (EC 2.7.11.1) (CDC42- binding protein kinase alpha) (Myotonic dystrophy kinase-related CDC42-binding kinase alpha) (Myotonic dystrophy protein kinase-like alpha) (MRCK alpha) (DMPK-like alpha). | |||||
|
PLSL_MOUSE
|
||||||
| NC score | 0.060309 (rank : 160) | θ value | θ > 10 (rank : 184) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q61233, Q3UE24, Q8R1X3, Q9CV77 | Gene names | Lcp1, Pls2 | |||
|
Domain Architecture |

|
|||||
| Description | Plastin-2 (L-plastin) (Lymphocyte cytosolic protein 1) (LCP-1) (65 kDa macrophage protein) (pp65). | |||||
|
MRCKB_MOUSE
|
||||||
| NC score | 0.060127 (rank : 161) | θ value | θ > 10 (rank : 175) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 2180 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q7TT50, Q69ZR5, Q80W33 | Gene names | Cdc42bpb, Kiaa1124 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MRCK beta (EC 2.7.11.1) (CDC42-binding protein kinase beta) (Myotonic dystrophy kinase-related CDC42-binding kinase beta) (Myotonic dystrophy protein kinase-like beta) (MRCK beta) (DMPK-like beta). | |||||
|
PLSL_HUMAN
|
||||||
| NC score | 0.059304 (rank : 162) | θ value | θ > 10 (rank : 183) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | P13796 | Gene names | LCP1, PLS2 | |||
|
Domain Architecture |

|
|||||
| Description | Plastin-2 (L-plastin) (Lymphocyte cytosolic protein 1) (LCP-1) (LC64P). | |||||
|
MRCKB_HUMAN
|
||||||
| NC score | 0.058524 (rank : 163) | θ value | θ > 10 (rank : 174) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 2239 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q9Y5S2, Q2L7A5, Q86TJ1, Q9ULU5 | Gene names | CDC42BPB, KIAA1124 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MRCK beta (EC 2.7.11.1) (CDC42-binding protein kinase beta) (Myotonic dystrophy kinase-related CDC42-binding kinase beta) (Myotonic dystrophy protein kinase-like beta) (MRCK beta) (DMPK-like beta). | |||||
|
MLRN_HUMAN
|
||||||
| NC score | 0.056684 (rank : 164) | θ value | θ > 10 (rank : 172) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P24844, Q9H136 | Gene names | MYL9, MLC2, MRLC1, MYRL2 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin regulatory light chain 2, smooth muscle isoform (Myosin RLC) (Myosin regulatory light chain 9) (LC20). | |||||
|
EFHB_HUMAN
|
||||||
| NC score | 0.056296 (rank : 165) | θ value | θ > 10 (rank : 165) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q8N7U6, Q6ZWK9, Q8IV58, Q96LQ6 | Gene names | EFHB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | EF-hand domain-containing family member B. | |||||
|
MLRN_MOUSE
|
||||||
| NC score | 0.055620 (rank : 166) | θ value | θ > 10 (rank : 173) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q9CQ19, Q80X77 | Gene names | Myl9, Myrl2 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin regulatory light chain 2, smooth muscle isoform (Myosin RLC) (Myosin regulatory light chain 9). | |||||
|
MLRM_HUMAN
|
||||||
| NC score | 0.054493 (rank : 167) | θ value | θ > 10 (rank : 171) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 41 | |
| SwissProt Accessions | P19105 | Gene names | MRLC2, MLCB, RLC | |||
|
Domain Architecture |

|
|||||
| Description | Myosin regulatory light chain 2, nonsarcomeric (Myosin RLC). | |||||
|
RCN2_MOUSE
|
||||||
| NC score | 0.054037 (rank : 168) | θ value | θ > 10 (rank : 188) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q8BP92, O70341 | Gene names | Rcn2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Reticulocalbin-2 precursor (Taipoxin-associated calcium-binding protein 49) (TCBP-49). | |||||
|
RGNEF_MOUSE
|
||||||
| NC score | 0.052077 (rank : 169) | θ value | 0.0961366 (rank : 77) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 641 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P97433 | Gene names | Rgnef, Rhoip2 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-guanine nucleotide exchange factor (Rho-interacting protein 2) (RhoGEF) (RIP2). | |||||
|
PLST_HUMAN
|
||||||
| NC score | 0.051741 (rank : 170) | θ value | 1.38821 (rank : 114) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P13797, Q86YI6 | Gene names | PLS3 | |||
|
Domain Architecture |

|
|||||
| Description | Plastin-3 (T-plastin). | |||||
|
RAF1_MOUSE
|
||||||
| NC score | 0.045023 (rank : 171) | θ value | 0.0736092 (rank : 74) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 886 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q99N57, Q91WH1 | Gene names | Raf1, Craf | |||
|
Domain Architecture |

|
|||||
| Description | RAF proto-oncogene serine/threonine-protein kinase (EC 2.7.11.1) (Raf- 1) (C-RAF) (cRaf). | |||||
|
RAF1_HUMAN
|
||||||
| NC score | 0.044621 (rank : 172) | θ value | 0.0736092 (rank : 73) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 886 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | P04049 | Gene names | RAF1, RAF | |||
|
Domain Architecture |

|
|||||
| Description | RAF proto-oncogene serine/threonine-protein kinase (EC 2.7.11.1) (Raf- 1) (C-RAF) (cRaf). | |||||
|
SPHK2_HUMAN
|
||||||
| NC score | 0.043896 (rank : 173) | θ value | 6.88961 (rank : 141) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9NRA0, Q9BRN1, Q9H0Q2, Q9NWU7 | Gene names | SPHK2 | |||
|
Domain Architecture |

|
|||||
| Description | Sphingosine kinase 2 (EC 2.7.1.-) (SK 2) (SPK 2). | |||||
|
ARAF_MOUSE
|
||||||
| NC score | 0.038013 (rank : 174) | θ value | 4.03905 (rank : 123) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 874 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P04627, Q99J44, Q9CTT5, Q9D6R6, Q9DBU7 | Gene names | Araf, A-raf, Araf1 | |||
|
Domain Architecture |

|
|||||
| Description | A-Raf proto-oncogene serine/threonine-protein kinase (EC 2.7.11.1). | |||||
|
ARAF_HUMAN
|
||||||
| NC score | 0.037079 (rank : 175) | θ value | 4.03905 (rank : 122) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 924 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | P10398, P07557, Q5H9B3 | Gene names | ARAF, ARAF1, PKS, PKS2 | |||
|
Domain Architecture |

|
|||||
| Description | A-Raf proto-oncogene serine/threonine-protein kinase (EC 2.7.11.1) (A- raf-1) (Proto-oncogene Pks). | |||||
|
FSTL5_MOUSE
|
||||||
| NC score | 0.036357 (rank : 176) | θ value | 0.62314 (rank : 99) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 355 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8BFR2, Q80TG3, Q8C4T3 | Gene names | Fstl5, Kiaa1263 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Follistatin-related protein 5 precursor (Follistatin-like 5) (m- D/Bsp120I 1-2). | |||||
|
KSR2_HUMAN
|
||||||
| NC score | 0.023249 (rank : 177) | θ value | 8.99809 (rank : 144) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 920 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q6VAB6, Q8N775 | Gene names | KSR2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinase suppressor of ras-2 (hKSR2). | |||||
|
FKBP7_HUMAN
|
||||||
| NC score | 0.018994 (rank : 178) | θ value | 5.27518 (rank : 133) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Y680, Q86U65, Q96DA4, Q9Y6B0 | Gene names | FKBP7, FKBP23 | |||
|
Domain Architecture |

|
|||||
| Description | FK506-binding protein 7 precursor (EC 5.2.1.8) (Peptidyl-prolyl cis- trans isomerase) (PPIase) (Rotamase) (FKBP-23). | |||||
|
MLL3_MOUSE
|
||||||
| NC score | 0.015731 (rank : 179) | θ value | 0.62314 (rank : 101) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1399 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8BRH4, Q5YLV9, Q8BK12, Q8C6M3, Q923H5, Q923H6 | Gene names | Mll3 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3). | |||||
|
ITB6_MOUSE
|
||||||
| NC score | 0.013142 (rank : 180) | θ value | 0.813845 (rank : 105) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9Z0T9 | Gene names | Itgb6 | |||
|
Domain Architecture |

|
|||||
| Description | Integrin beta-6 precursor. | |||||
|
LAMC3_HUMAN
|
||||||
| NC score | 0.012760 (rank : 181) | θ value | 0.163984 (rank : 83) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 588 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9Y6N6 | Gene names | LAMC3 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin gamma-3 chain precursor (Laminin 12 gamma 3 subunit). | |||||
|
LAMB2_MOUSE
|
||||||
| NC score | 0.012735 (rank : 182) | θ value | 2.36792 (rank : 117) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 722 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q61292, Q62182 | Gene names | Lamb2, Lams | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-2 chain precursor (S-laminin) (S-LAM). | |||||
|
LAMB2_HUMAN
|
||||||
| NC score | 0.011989 (rank : 183) | θ value | 5.27518 (rank : 134) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 692 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P55268, Q16321 | Gene names | LAMB2, LAMS | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-2 chain precursor (S-laminin) (Laminin B1s chain). | |||||
|
MEGF9_MOUSE
|
||||||
| NC score | 0.011833 (rank : 184) | θ value | 8.99809 (rank : 146) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8BH27, Q8BWI4 | Gene names | Megf9, Egfl5, Kiaa0818 | |||
|
Domain Architecture |

|
|||||
| Description | Multiple epidermal growth factor-like domains 9 precursor (EGF-like domain-containing protein 5) (Multiple EGF-like domain protein 5). | |||||
|
RN183_HUMAN
|
||||||
| NC score | 0.010556 (rank : 185) | θ value | 4.03905 (rank : 128) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 126 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96D59 | Gene names | RNF183 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 183. | |||||
|
LAMB1_HUMAN
|
||||||
| NC score | 0.009723 (rank : 186) | θ value | 6.88961 (rank : 138) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1018 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P07942 | Gene names | LAMB1 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-1 chain precursor (Laminin B1 chain). | |||||
|
LAMA2_MOUSE
|
||||||
| NC score | 0.008312 (rank : 187) | θ value | 8.99809 (rank : 145) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1198 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q60675, Q05003, Q64061 | Gene names | Lama2 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-2 chain precursor (Laminin M chain) (Merosin heavy chain). | |||||
|
SSPO_MOUSE
|
||||||
| NC score | 0.006695 (rank : 188) | θ value | 0.47712 (rank : 97) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 800 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8CG65 | Gene names | Sspo | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SCO-spondin precursor. | |||||
|
ADA29_HUMAN
|
||||||
| NC score | 0.003596 (rank : 189) | θ value | 8.99809 (rank : 143) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9UKF5, Q9UHP1, Q9UKF3, Q9UKF4 | Gene names | ADAM29 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 29 precursor (A disintegrin and metalloproteinase domain 29). | |||||
|
ADA20_HUMAN
|
||||||
| NC score | 0.002607 (rank : 190) | θ value | 8.99809 (rank : 142) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 143 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O43506, Q9UKJ9 | Gene names | ADAM20 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 20 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 20). | |||||
|
TCF8_MOUSE
|
||||||
| NC score | 0.001875 (rank : 191) | θ value | 2.36792 (rank : 118) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 1067 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q64318, Q62519 | Gene names | Tcf8, Zfhx1a, Zfx1a, Zfx1ha | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 8 (Zinc finger homeobox protein 1a) (MEB1) (Delta EF1). | |||||
|
ADAM8_MOUSE
|
||||||
| NC score | 0.001766 (rank : 192) | θ value | 6.88961 (rank : 136) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 326 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q05910 | Gene names | Adam8, Ms2 | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 8 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 8) (Cell surface antigen MS2) (Macrophage cysteine-rich glycoprotein) (CD156 antigen). | |||||
|
PRD15_HUMAN
|
||||||
| NC score | -0.000314 (rank : 193) | θ value | 6.88961 (rank : 140) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 805 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P57071, Q8N0X3, Q8NEX0, Q9NQV3 | Gene names | PRDM15, C21orf83, ZNF298 | |||
|
Domain Architecture |

|
|||||
| Description | PR domain zinc finger protein 15 (PR domain-containing protein 15) (Zinc finger protein 298). | |||||
|
ZN545_HUMAN
|
||||||
| NC score | -0.001966 (rank : 194) | θ value | 2.36792 (rank : 119) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8N141, Q8NC63, Q8TF53 | Gene names | ZNF545, KIAA1948 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 545. | |||||
|
PRDM4_MOUSE
|
||||||
| NC score | -0.002279 (rank : 195) | θ value | 4.03905 (rank : 127) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q80V63 | Gene names | Prdm4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PR domain zinc finger protein 4 (PR domain-containing protein 4). | |||||
|
PRDM4_HUMAN
|
||||||
| NC score | -0.002290 (rank : 196) | θ value | 4.03905 (rank : 126) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 749 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9UKN5, Q9UFA6 | Gene names | PRDM4, PFM1 | |||
|
Domain Architecture |

|
|||||
| Description | PR domain zinc finger protein 4 (PR domain-containing protein 4). | |||||
|
ZN545_MOUSE
|
||||||
| NC score | -0.002774 (rank : 197) | θ value | 4.03905 (rank : 130) | |||
| Query Neighborhood Hits | 146 | Target Neighborhood Hits | 739 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q6P9Y7, Q6ZPG0 | Gene names | Znf545, Kiaa1948, Zfp82 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 545 (Zinc finger protein 82). | |||||