Please be patient as the page loads
|
DGKE_HUMAN
|
||||||
| SwissProt Accessions | P52429, Q9UKQ3 | Gene names | DGKE, DAGK5 | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase epsilon (EC 2.7.1.107) (Diglyceride kinase epsilon) (DGK-epsilon) (DAG kinase epsilon). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
DGKE_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P52429, Q9UKQ3 | Gene names | DGKE, DAGK5 | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase epsilon (EC 2.7.1.107) (Diglyceride kinase epsilon) (DGK-epsilon) (DAG kinase epsilon). | |||||
|
DGKE_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.995351 (rank : 2) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9R1C6 | Gene names | Dgke | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase epsilon (EC 2.7.1.107) (Diglyceride kinase epsilon) (DGK-epsilon) (DAG kinase epsilon). | |||||
|
DGKB_HUMAN
|
||||||
| θ value | 1.5519e-76 (rank : 3) | NC score | 0.838304 (rank : 8) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9Y6T7, O75241, Q75MF9, Q75MU7, Q86UI5, Q86UM9, Q9UQ29 | Gene names | DGKB, DAGK2, KIAA0718 | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase beta (EC 2.7.1.107) (Diglyceride kinase beta) (DGK-beta) (DAG kinase beta) (90 kDa diacylglycerol kinase). | |||||
|
DGKG_MOUSE
|
||||||
| θ value | 9.41505e-74 (rank : 4) | NC score | 0.866351 (rank : 6) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q91WG7 | Gene names | Dgkg, Dagk3 | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase gamma (EC 2.7.1.107) (Diglyceride kinase gamma) (DGK-gamma) (DAG kinase gamma) (88 kDa diacylglycerol kinase). | |||||
|
DGKG_HUMAN
|
||||||
| θ value | 1.60597e-73 (rank : 5) | NC score | 0.874037 (rank : 5) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P49619 | Gene names | DGKG, DAGK3 | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase gamma (EC 2.7.1.107) (Diglyceride kinase gamma) (DGK-gamma) (DAG kinase gamma). | |||||
|
DGKA_HUMAN
|
||||||
| θ value | 4.67263e-73 (rank : 6) | NC score | 0.893616 (rank : 3) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P23743, O75481, O75482, O75483 | Gene names | DGKA, DAGK, DAGK1 | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase alpha (EC 2.7.1.107) (Diglyceride kinase alpha) (DGK-alpha) (DAG kinase alpha) (80 kDa diacylglycerol kinase). | |||||
|
DGKA_MOUSE
|
||||||
| θ value | 1.66191e-70 (rank : 7) | NC score | 0.888069 (rank : 4) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O88673, Q922X2 | Gene names | Dgka, Dagk1 | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase alpha (EC 2.7.1.107) (Diglyceride kinase alpha) (DGK-alpha) (DAG kinase alpha) (80 kDa diacylglycerol kinase). | |||||
|
DGKQ_HUMAN
|
||||||
| θ value | 1.90147e-66 (rank : 8) | NC score | 0.864511 (rank : 7) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P52824 | Gene names | DGKQ, DAGK4 | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase theta (EC 2.7.1.107) (Diglyceride kinase theta) (DGK-theta) (DAG kinase theta). | |||||
|
DGKI_HUMAN
|
||||||
| θ value | 5.92465e-60 (rank : 9) | NC score | 0.765278 (rank : 14) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 609 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O75912, Q9NZ49 | Gene names | DGKI | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase iota (EC 2.7.1.107) (Diglyceride kinase iota) (DGK-iota) (DAG kinase iota). | |||||
|
DGKZ_MOUSE
|
||||||
| θ value | 3.59435e-57 (rank : 10) | NC score | 0.778158 (rank : 11) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 542 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q80UP3 | Gene names | Dgkz | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase zeta (EC 2.7.1.107) (Diglyceride kinase zeta) (DGK-zeta) (DAG kinase zeta). | |||||
|
DGKZ_HUMAN
|
||||||
| θ value | 5.19021e-56 (rank : 11) | NC score | 0.773016 (rank : 12) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 551 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q13574, O00542 | Gene names | DGKZ, DAGK6 | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase zeta (EC 2.7.1.107) (Diglyceride kinase zeta) (DGK-zeta) (DAG kinase zeta). | |||||
|
DGKD_HUMAN
|
||||||
| θ value | 5.20228e-48 (rank : 12) | NC score | 0.783386 (rank : 9) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q16760, Q14158, Q6PK55, Q8NG53 | Gene names | DGKD, KIAA0145 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Diacylglycerol kinase delta (EC 2.7.1.107) (Diglyceride kinase delta) (DGK-delta) (DAG kinase delta) (130 kDa diacylglycerol kinase). | |||||
|
DGKH_HUMAN
|
||||||
| θ value | 1.28137e-46 (rank : 13) | NC score | 0.778213 (rank : 10) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q86XP1, Q5VZW0, Q6PI56, Q86XP2, Q8N3N0, Q8N7J9 | Gene names | DGKH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Diacylglycerol kinase eta (EC 2.7.1.107) (Diglyceride kinase eta) (DGK-eta) (DAG kinase eta). | |||||
|
DGKK_HUMAN
|
||||||
| θ value | 4.55743e-44 (rank : 14) | NC score | 0.767798 (rank : 13) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 378 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q5KSL6 | Gene names | DGKK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Diacylglycerol kinase kappa (EC 2.7.1.107) (Diglyceride kinase kappa) (DGK-kappa) (DAG kinase kappa) (142 kDa diacylglycerol kinase). | |||||
|
KPCL_HUMAN
|
||||||
| θ value | 7.1131e-05 (rank : 15) | NC score | 0.062238 (rank : 32) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 887 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P24723, Q16246 | Gene names | PRKCH, PKCL | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C eta type (EC 2.7.11.13) (nPKC-eta) (PKC-L). | |||||
|
KPCL_MOUSE
|
||||||
| θ value | 0.000121331 (rank : 16) | NC score | 0.060059 (rank : 38) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 891 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P23298 | Gene names | Prkch, Pkch | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C eta type (EC 2.7.11.13) (nPKC-eta) (PKC-L). | |||||
|
USH2A_HUMAN
|
||||||
| θ value | 0.21417 (rank : 17) | NC score | 0.012556 (rank : 68) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 600 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O75445, Q5VVM9, Q6S362, Q9NS27 | Gene names | USH2A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Usherin precursor (Usher syndrome type-2A protein) (Usher syndrome type IIa protein). | |||||
|
KPCT_HUMAN
|
||||||
| θ value | 0.279714 (rank : 18) | NC score | 0.057170 (rank : 49) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 897 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q04759, Q3MJF1, Q64FY5, Q9H508, Q9H549 | Gene names | PRKCQ, PRKCT | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C theta type (EC 2.7.11.13) (nPKC-theta). | |||||
|
KPCT_MOUSE
|
||||||
| θ value | 0.279714 (rank : 19) | NC score | 0.057402 (rank : 48) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 895 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q02111 | Gene names | Prkcq, Pkcq | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C theta type (EC 2.7.11.13) (nPKC-theta). | |||||
|
ZN277_HUMAN
|
||||||
| θ value | 0.47712 (rank : 20) | NC score | 0.037589 (rank : 60) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NRM2 | Gene names | ZNF277 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 277. | |||||
|
NTNG1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 21) | NC score | 0.017258 (rank : 63) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y2I2, Q5VU86, Q5VU87, Q5VU89, Q5VU90, Q5VU91, Q7Z2Y3, Q8N633 | Gene names | NTNG1, KIAA0976, LMNT1 | |||
|
Domain Architecture |
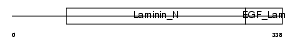
|
|||||
| Description | Netrin G1 precursor (Laminet-1). | |||||
|
NTNG1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 22) | NC score | 0.016887 (rank : 64) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8R4G0, Q68FE5, Q69ZU3, Q8R4F3, Q8R4F4, Q8R4F5, Q8R4F6, Q8R4F7, Q8R4F8, Q8R4F9, Q9ESR3, Q9ESR4, Q9ESR5, Q9ESR6, Q9ESR7, Q9ESR8 | Gene names | Ntng1, Kiaa0976, Lmnt1 | |||
|
Domain Architecture |
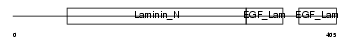
|
|||||
| Description | Netrin G1 precursor (Laminet-1). | |||||
|
KPCD_MOUSE
|
||||||
| θ value | 1.06291 (rank : 23) | NC score | 0.059715 (rank : 39) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 888 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P28867, Q91V85, Q9Z333 | Gene names | Prkcd, Pkcd | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C delta type (EC 2.7.11.13) (nPKC-delta). | |||||
|
KPCD_HUMAN
|
||||||
| θ value | 1.38821 (rank : 24) | NC score | 0.059298 (rank : 41) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 900 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q05655, Q15144 | Gene names | PRKCD | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C delta type (EC 2.7.11.13) (nPKC-delta). | |||||
|
KPCG_HUMAN
|
||||||
| θ value | 1.81305 (rank : 25) | NC score | 0.053829 (rank : 53) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 1126 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P05129 | Gene names | PRKCG, PKCG | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C gamma type (EC 2.7.11.13) (PKC-gamma). | |||||
|
KPCG_MOUSE
|
||||||
| θ value | 1.81305 (rank : 26) | NC score | 0.053454 (rank : 56) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 1123 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P63318, P05697 | Gene names | Prkcg, Pkcc, Pkcg, Prkcc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein kinase C gamma type (EC 2.7.11.13) (PKC-gamma). | |||||
|
BRPF3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 27) | NC score | 0.010500 (rank : 69) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9ULD4, Q5R3K8 | Gene names | BRPF3, KIAA1286 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain and PHD finger-containing protein 3. | |||||
|
PHF14_HUMAN
|
||||||
| θ value | 6.88961 (rank : 28) | NC score | 0.014746 (rank : 66) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O94880 | Gene names | PHF14, KIAA0783 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 14. | |||||
|
PHF14_MOUSE
|
||||||
| θ value | 6.88961 (rank : 29) | NC score | 0.015307 (rank : 65) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9D4H9, Q810Z6, Q8C284, Q8CGF8, Q9D1Q8 | Gene names | Phf14 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 14. | |||||
|
KSR1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 30) | NC score | 0.013124 (rank : 67) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 870 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q61097, Q61648, Q78DX8 | Gene names | Ksr1, Ksr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinase suppressor of ras-1 (Kinase suppressor of ras) (mKSR1) (Hb protein). | |||||
|
RAF1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 31) | NC score | 0.021583 (rank : 62) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 886 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P04049 | Gene names | RAF1, RAF | |||
|
Domain Architecture |

|
|||||
| Description | RAF proto-oncogene serine/threonine-protein kinase (EC 2.7.11.1) (Raf- 1) (C-RAF) (cRaf). | |||||
|
RAF1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 32) | NC score | 0.021785 (rank : 61) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 886 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q99N57, Q91WH1 | Gene names | Raf1, Craf | |||
|
Domain Architecture |

|
|||||
| Description | RAF proto-oncogene serine/threonine-protein kinase (EC 2.7.11.1) (Raf- 1) (C-RAF) (cRaf). | |||||
|
CANB1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 33) | NC score | 0.055156 (rank : 52) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P63098, P06705, P15117, Q08044 | Gene names | PPP3R1, CNA2, CNB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcineurin subunit B isoform 1 (Protein phosphatase 2B regulatory subunit 1) (Protein phosphatase 3 regulatory subunit B alpha isoform 1). | |||||
|
CANB1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 34) | NC score | 0.055161 (rank : 51) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q63810 | Gene names | Ppp3r1, Cnb | |||
|
Domain Architecture |
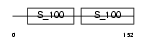
|
|||||
| Description | Calcineurin subunit B isoform 1 (Protein phosphatase 2B regulatory subunit 1) (Protein phosphatase 3 regulatory subunit B alpha isoform 1). | |||||
|
CANB2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 35) | NC score | 0.053516 (rank : 55) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q96LZ3, Q7Z4V8, Q8WYJ4 | Gene names | PPP3R2, CBLP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcineurin subunit B isoform 2 (Protein phosphatase 2B regulatory subunit 2) (Protein phosphatase 3 regulatory subunit B beta isoform) (Calcineurin B-like protein) (CBLP) (CNBII). | |||||
|
CANB2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 36) | NC score | 0.059135 (rank : 42) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q63811 | Gene names | Ppp3r2 | |||
|
Domain Architecture |
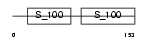
|
|||||
| Description | Calcineurin subunit B isoform 2 (Protein phosphatase 2B regulatory subunit 2) (Protein phosphatase 3 regulatory subunit B beta isoform). | |||||
|
CHIN_HUMAN
|
||||||
| θ value | θ > 10 (rank : 37) | NC score | 0.059422 (rank : 40) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P15882, Q53SD6, Q53SH5, Q96FB0 | Gene names | CHN1, ARHGAP2, CHN | |||
|
Domain Architecture |
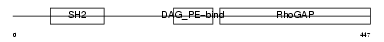
|
|||||
| Description | N-chimaerin (NC) (N-chimerin) (Alpha chimerin) (A-chimaerin) (Rho- GTPase-activating protein 2). | |||||
|
CHIN_MOUSE
|
||||||
| θ value | θ > 10 (rank : 38) | NC score | 0.055474 (rank : 50) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 229 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q91V57, Q3UGY3, Q7TQE5, Q8BWU6, Q9D9B3 | Gene names | Chn1, Arhgap2 | |||
|
Domain Architecture |
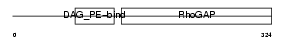
|
|||||
| Description | N-chimaerin (NC) (N-chimerin) (Alpha chimerin) (A-chimaerin) (Rho- GTPase-activating protein 2). | |||||
|
CHIO_HUMAN
|
||||||
| θ value | θ > 10 (rank : 39) | NC score | 0.063407 (rank : 30) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 220 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P52757, Q75MM2 | Gene names | CHN2, ARHGAP3, BCH | |||
|
Domain Architecture |
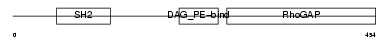
|
|||||
| Description | Beta-chimaerin (Beta-chimerin) (Rho-GTPase-activating protein 3). | |||||
|
CHIO_MOUSE
|
||||||
| θ value | θ > 10 (rank : 40) | NC score | 0.067266 (rank : 26) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q80XD1, Q9D9W2, Q9ER57 | Gene names | Chn2, Arhgap3, Bch | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-chimaerin (Beta-chimerin) (Rho-GTPase-activating protein 3). | |||||
|
CHP2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 41) | NC score | 0.058555 (rank : 45) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O43745 | Gene names | CHP2, HCA520 | |||
|
Domain Architecture |
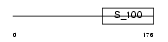
|
|||||
| Description | Calcineurin B homologous protein 2 (Hepatocellular carcinoma- associated antigen 520). | |||||
|
CHP2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 42) | NC score | 0.066114 (rank : 27) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9D869 | Gene names | Chp2, Hca520 | |||
|
Domain Architecture |
|
|||||
| Description | Calcineurin B homologous protein 2 (Hepatocellular carcinoma- associated antigen 520 homolog). | |||||
|
EFHD1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 43) | NC score | 0.052466 (rank : 59) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9BUP0, Q9BTF8, Q9H8I2, Q9HBQ0 | Gene names | EFHD1, SWS2 | |||
|
Domain Architecture |
|
|||||
| Description | EF-hand domain-containing protein 1 (Swiprosin-2). | |||||
|
GRP3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 44) | NC score | 0.104950 (rank : 17) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8IV61, O94931 | Gene names | RASGRP3, GRP3, KIAA0846 | |||
|
Domain Architecture |

|
|||||
| Description | RAS guanyl-releasing protein 3 (Calcium and DAG-regulated guanine nucleotide exchange factor III) (Guanine nucleotide exchange factor for Rap1). | |||||
|
GUC1A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 45) | NC score | 0.064076 (rank : 29) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P43080, Q9NU14 | Gene names | GUCA1A, GCAP, GCAP1, GUCA1 | |||
|
Domain Architecture |
|
|||||
| Description | Guanylyl cyclase-activating protein 1 (GCAP 1) (Guanylate cyclase activator 1A). | |||||
|
GUC1A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 46) | NC score | 0.064756 (rank : 28) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P43081 | Gene names | Guca1a, Gcap, Gcap1, Guca1 | |||
|
Domain Architecture |
|
|||||
| Description | Guanylyl cyclase-activating protein 1 (GCAP 1) (Guanylate cyclase activator 1A). | |||||
|
GUC1B_HUMAN
|
||||||
| θ value | θ > 10 (rank : 47) | NC score | 0.057456 (rank : 47) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UMX6, Q9NU15 | Gene names | GUCA1B, GCAP2 | |||
|
Domain Architecture |
|
|||||
| Description | Guanylyl cyclase-activating protein 2 (GCAP 2) (Guanylate cyclase activator 1B). | |||||
|
GUC1B_MOUSE
|
||||||
| θ value | θ > 10 (rank : 48) | NC score | 0.058779 (rank : 43) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8VBV8 | Gene names | Guca1b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Guanylyl cyclase-activating protein 2 (GCAP 2) (Guanylate cyclase activator 1B). | |||||
|
GUC1C_HUMAN
|
||||||
| θ value | θ > 10 (rank : 49) | NC score | 0.053141 (rank : 58) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O95843, O95844, Q9UNM0 | Gene names | GUCA1C, GCAP3 | |||
|
Domain Architecture |
|
|||||
| Description | Guanylyl cyclase-activating protein 3 (GCAP 3) (Guanylate cyclase activator 1C). | |||||
|
HPCA_HUMAN
|
||||||
| θ value | θ > 10 (rank : 50) | NC score | 0.060087 (rank : 36) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P84074, P32076, P41211, P70510 | Gene names | HPCA, BDR2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuron-specific calcium-binding protein hippocalcin (Calcium-binding protein BDR-2). | |||||
|
HPCA_MOUSE
|
||||||
| θ value | θ > 10 (rank : 51) | NC score | 0.060087 (rank : 37) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P84075, P32076, P41211, P70510 | Gene names | Hpca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuron-specific calcium-binding protein hippocalcin. | |||||
|
HPCL1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 52) | NC score | 0.058632 (rank : 44) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P37235, Q969S5 | Gene names | HPCAL1, BDR1 | |||
|
Domain Architecture |
|
|||||
| Description | Hippocalcin-like protein 1 (Visinin-like protein 3) (VILIP-3) (Calcium-binding protein BDR-1) (HLP2). | |||||
|
HPCL1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 53) | NC score | 0.060278 (rank : 35) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P62748, P35333 | Gene names | Hpcal1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hippocalcin-like protein 1 (Visinin-like protein 3) (VILIP-3) (Neural visinin-like protein 3) (NVL-3) (NVP-3). | |||||
|
KIP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 54) | NC score | 0.096463 (rank : 20) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q99828, O00693, O00735, Q6IB49, Q96J54, Q99971 | Gene names | CIB1, CIB, KIP, PRKDCIP | |||
|
Domain Architecture |
|
|||||
| Description | Calcium and integrin-binding protein 1 (Calmyrin) (DNA-PKcs- interacting protein) (Kinase-interacting protein) (KIP) (CIB) (SNK- interacting protein 2-28) (SIP2-28). | |||||
|
KIP1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 55) | NC score | 0.105704 (rank : 15) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Z0F4, Q3TN80 | Gene names | Cib1, Cib, Kip, Prkdcip | |||
|
Domain Architecture |
|
|||||
| Description | Calcium and integrin-binding protein 1 (Calmyrin) (DNA-PKcs- interacting protein) (Kinase-interacting protein) (KIP) (CIB). | |||||
|
KPCE_HUMAN
|
||||||
| θ value | θ > 10 (rank : 56) | NC score | 0.053158 (rank : 57) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q02156, Q53SL4, Q53SM5, Q9UE81 | Gene names | PRKCE, PKCE | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C epsilon type (EC 2.7.11.13) (nPKC-epsilon). | |||||
|
KPCE_MOUSE
|
||||||
| θ value | θ > 10 (rank : 57) | NC score | 0.053684 (rank : 54) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P16054 | Gene names | Prkce, Pkce, Pkcea | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C epsilon type (EC 2.7.11.13) (nPKC-epsilon). | |||||
|
NCALD_HUMAN
|
||||||
| θ value | θ > 10 (rank : 58) | NC score | 0.061187 (rank : 33) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P61601, P29554, Q8IYC3, Q9H0W2 | Gene names | NCALD | |||
|
Domain Architecture |
|
|||||
| Description | Neurocalcin delta. | |||||
|
NCALD_MOUSE
|
||||||
| θ value | θ > 10 (rank : 59) | NC score | 0.061187 (rank : 34) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q91X97, Q3TJS9, Q8BZN9 | Gene names | Ncald, D15Ertd412e | |||
|
Domain Architecture |
|
|||||
| Description | Neurocalcin delta. | |||||
|
PDZD8_HUMAN
|
||||||
| θ value | θ > 10 (rank : 60) | NC score | 0.068499 (rank : 24) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 185 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8NEN9, Q86WE0, Q86WE5, Q9UFF1 | Gene names | PDZD8, PDZK8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing protein 8 (Sarcoma antigen NY-SAR-84/NY-SAR- 104). | |||||
|
RASF1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 61) | NC score | 0.057856 (rank : 46) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9NS23, O14571, O60539, O60710, Q9HB04, Q9HB18, Q9NS22, Q9UND4, Q9UND5 | Gene names | RASSF1, RDA32 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras association domain-containing protein 1. | |||||
|
RASF1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 62) | NC score | 0.104957 (rank : 16) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q99MK9, Q9WUF5 | Gene names | Rassf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras association domain-containing protein 1 (Protein 123F2). | |||||
|
STAC3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 63) | NC score | 0.085171 (rank : 21) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q96MF2, Q96HU5 | Gene names | STAC3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH3 and cysteine-rich domain-containing protein 3. | |||||
|
STAC3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 64) | NC score | 0.084169 (rank : 22) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8BZ71 | Gene names | Stac3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH3 and cysteine-rich domain-containing protein 3. | |||||
|
STAC_HUMAN
|
||||||
| θ value | θ > 10 (rank : 65) | NC score | 0.096466 (rank : 19) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 276 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q99469 | Gene names | STAC | |||
|
Domain Architecture |
|
|||||
| Description | SH3 and cysteine-rich domain-containing protein (SRC homology 3 and cysteine-rich domain protein). | |||||
|
STAC_MOUSE
|
||||||
| θ value | θ > 10 (rank : 66) | NC score | 0.098941 (rank : 18) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 268 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P97306 | Gene names | Stac | |||
|
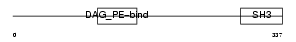
Domain Architecture |
|
|||||
| Description | SH3 and cysteine-rich domain-containing protein (SRC homology 3 and cysteine-rich domain protein). | |||||
|
UN13A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 67) | NC score | 0.067440 (rank : 25) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9UPW8 | Gene names | UNC13A, KIAA1032 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Unc-13 homolog A (Munc13-1). | |||||
|
UN13B_HUMAN
|
||||||
| θ value | θ > 10 (rank : 68) | NC score | 0.068875 (rank : 23) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O14795 | Gene names | UNC13B, UNC13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Unc-13 homolog B (Munc13-2) (munc13). | |||||
|
UN13B_MOUSE
|
||||||
| θ value | θ > 10 (rank : 69) | NC score | 0.062246 (rank : 31) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Z1N9 | Gene names | Unc13b, Unc13a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Unc-13 homolog B (Munc13-2) (munc13). | |||||
|
DGKE_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | P52429, Q9UKQ3 | Gene names | DGKE, DAGK5 | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase epsilon (EC 2.7.1.107) (Diglyceride kinase epsilon) (DGK-epsilon) (DAG kinase epsilon). | |||||
|
DGKE_MOUSE
|
||||||
| NC score | 0.995351 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9R1C6 | Gene names | Dgke | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase epsilon (EC 2.7.1.107) (Diglyceride kinase epsilon) (DGK-epsilon) (DAG kinase epsilon). | |||||
|
DGKA_HUMAN
|
||||||
| NC score | 0.893616 (rank : 3) | θ value | 4.67263e-73 (rank : 6) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P23743, O75481, O75482, O75483 | Gene names | DGKA, DAGK, DAGK1 | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase alpha (EC 2.7.1.107) (Diglyceride kinase alpha) (DGK-alpha) (DAG kinase alpha) (80 kDa diacylglycerol kinase). | |||||
|
DGKA_MOUSE
|
||||||
| NC score | 0.888069 (rank : 4) | θ value | 1.66191e-70 (rank : 7) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O88673, Q922X2 | Gene names | Dgka, Dagk1 | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase alpha (EC 2.7.1.107) (Diglyceride kinase alpha) (DGK-alpha) (DAG kinase alpha) (80 kDa diacylglycerol kinase). | |||||
|
DGKG_HUMAN
|
||||||
| NC score | 0.874037 (rank : 5) | θ value | 1.60597e-73 (rank : 5) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P49619 | Gene names | DGKG, DAGK3 | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase gamma (EC 2.7.1.107) (Diglyceride kinase gamma) (DGK-gamma) (DAG kinase gamma). | |||||
|
DGKG_MOUSE
|
||||||
| NC score | 0.866351 (rank : 6) | θ value | 9.41505e-74 (rank : 4) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q91WG7 | Gene names | Dgkg, Dagk3 | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase gamma (EC 2.7.1.107) (Diglyceride kinase gamma) (DGK-gamma) (DAG kinase gamma) (88 kDa diacylglycerol kinase). | |||||
|
DGKQ_HUMAN
|
||||||
| NC score | 0.864511 (rank : 7) | θ value | 1.90147e-66 (rank : 8) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P52824 | Gene names | DGKQ, DAGK4 | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase theta (EC 2.7.1.107) (Diglyceride kinase theta) (DGK-theta) (DAG kinase theta). | |||||
|
DGKB_HUMAN
|
||||||
| NC score | 0.838304 (rank : 8) | θ value | 1.5519e-76 (rank : 3) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 238 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9Y6T7, O75241, Q75MF9, Q75MU7, Q86UI5, Q86UM9, Q9UQ29 | Gene names | DGKB, DAGK2, KIAA0718 | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase beta (EC 2.7.1.107) (Diglyceride kinase beta) (DGK-beta) (DAG kinase beta) (90 kDa diacylglycerol kinase). | |||||
|
DGKD_HUMAN
|
||||||
| NC score | 0.783386 (rank : 9) | θ value | 5.20228e-48 (rank : 12) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q16760, Q14158, Q6PK55, Q8NG53 | Gene names | DGKD, KIAA0145 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Diacylglycerol kinase delta (EC 2.7.1.107) (Diglyceride kinase delta) (DGK-delta) (DAG kinase delta) (130 kDa diacylglycerol kinase). | |||||
|
DGKH_HUMAN
|
||||||
| NC score | 0.778213 (rank : 10) | θ value | 1.28137e-46 (rank : 13) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q86XP1, Q5VZW0, Q6PI56, Q86XP2, Q8N3N0, Q8N7J9 | Gene names | DGKH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Diacylglycerol kinase eta (EC 2.7.1.107) (Diglyceride kinase eta) (DGK-eta) (DAG kinase eta). | |||||
|
DGKZ_MOUSE
|
||||||
| NC score | 0.778158 (rank : 11) | θ value | 3.59435e-57 (rank : 10) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 542 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q80UP3 | Gene names | Dgkz | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase zeta (EC 2.7.1.107) (Diglyceride kinase zeta) (DGK-zeta) (DAG kinase zeta). | |||||
|
DGKZ_HUMAN
|
||||||
| NC score | 0.773016 (rank : 12) | θ value | 5.19021e-56 (rank : 11) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 551 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q13574, O00542 | Gene names | DGKZ, DAGK6 | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase zeta (EC 2.7.1.107) (Diglyceride kinase zeta) (DGK-zeta) (DAG kinase zeta). | |||||
|
DGKK_HUMAN
|
||||||
| NC score | 0.767798 (rank : 13) | θ value | 4.55743e-44 (rank : 14) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 378 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q5KSL6 | Gene names | DGKK | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Diacylglycerol kinase kappa (EC 2.7.1.107) (Diglyceride kinase kappa) (DGK-kappa) (DAG kinase kappa) (142 kDa diacylglycerol kinase). | |||||
|
DGKI_HUMAN
|
||||||
| NC score | 0.765278 (rank : 14) | θ value | 5.92465e-60 (rank : 9) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 609 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O75912, Q9NZ49 | Gene names | DGKI | |||
|
Domain Architecture |

|
|||||
| Description | Diacylglycerol kinase iota (EC 2.7.1.107) (Diglyceride kinase iota) (DGK-iota) (DAG kinase iota). | |||||
|
KIP1_MOUSE
|
||||||
| NC score | 0.105704 (rank : 15) | θ value | θ > 10 (rank : 55) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Z0F4, Q3TN80 | Gene names | Cib1, Cib, Kip, Prkdcip | |||
|
Domain Architecture |

|
|||||
| Description | Calcium and integrin-binding protein 1 (Calmyrin) (DNA-PKcs- interacting protein) (Kinase-interacting protein) (KIP) (CIB). | |||||
|
RASF1_MOUSE
|
||||||
| NC score | 0.104957 (rank : 16) | θ value | θ > 10 (rank : 62) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q99MK9, Q9WUF5 | Gene names | Rassf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras association domain-containing protein 1 (Protein 123F2). | |||||
|
GRP3_HUMAN
|
||||||
| NC score | 0.104950 (rank : 17) | θ value | θ > 10 (rank : 44) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8IV61, O94931 | Gene names | RASGRP3, GRP3, KIAA0846 | |||
|
Domain Architecture |

|
|||||
| Description | RAS guanyl-releasing protein 3 (Calcium and DAG-regulated guanine nucleotide exchange factor III) (Guanine nucleotide exchange factor for Rap1). | |||||
|
STAC_MOUSE
|
||||||
| NC score | 0.098941 (rank : 18) | θ value | θ > 10 (rank : 66) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 268 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P97306 | Gene names | Stac | |||
|
Domain Architecture |

|
|||||
| Description | SH3 and cysteine-rich domain-containing protein (SRC homology 3 and cysteine-rich domain protein). | |||||
|
STAC_HUMAN
|
||||||
| NC score | 0.096466 (rank : 19) | θ value | θ > 10 (rank : 65) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 276 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q99469 | Gene names | STAC | |||
|
Domain Architecture |

|
|||||
| Description | SH3 and cysteine-rich domain-containing protein (SRC homology 3 and cysteine-rich domain protein). | |||||
|
KIP1_HUMAN
|
||||||
| NC score | 0.096463 (rank : 20) | θ value | θ > 10 (rank : 54) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q99828, O00693, O00735, Q6IB49, Q96J54, Q99971 | Gene names | CIB1, CIB, KIP, PRKDCIP | |||
|
Domain Architecture |

|
|||||
| Description | Calcium and integrin-binding protein 1 (Calmyrin) (DNA-PKcs- interacting protein) (Kinase-interacting protein) (KIP) (CIB) (SNK- interacting protein 2-28) (SIP2-28). | |||||
|
STAC3_HUMAN
|
||||||
| NC score | 0.085171 (rank : 21) | θ value | θ > 10 (rank : 63) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q96MF2, Q96HU5 | Gene names | STAC3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH3 and cysteine-rich domain-containing protein 3. | |||||
|
STAC3_MOUSE
|
||||||
| NC score | 0.084169 (rank : 22) | θ value | θ > 10 (rank : 64) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8BZ71 | Gene names | Stac3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH3 and cysteine-rich domain-containing protein 3. | |||||
|
UN13B_HUMAN
|
||||||
| NC score | 0.068875 (rank : 23) | θ value | θ > 10 (rank : 68) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O14795 | Gene names | UNC13B, UNC13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Unc-13 homolog B (Munc13-2) (munc13). | |||||
|
PDZD8_HUMAN
|
||||||
| NC score | 0.068499 (rank : 24) | θ value | θ > 10 (rank : 60) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 185 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8NEN9, Q86WE0, Q86WE5, Q9UFF1 | Gene names | PDZD8, PDZK8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PDZ domain-containing protein 8 (Sarcoma antigen NY-SAR-84/NY-SAR- 104). | |||||
|
UN13A_HUMAN
|
||||||
| NC score | 0.067440 (rank : 25) | θ value | θ > 10 (rank : 67) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9UPW8 | Gene names | UNC13A, KIAA1032 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Unc-13 homolog A (Munc13-1). | |||||
|
CHIO_MOUSE
|
||||||
| NC score | 0.067266 (rank : 26) | θ value | θ > 10 (rank : 40) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q80XD1, Q9D9W2, Q9ER57 | Gene names | Chn2, Arhgap3, Bch | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-chimaerin (Beta-chimerin) (Rho-GTPase-activating protein 3). | |||||
|
CHP2_MOUSE
|
||||||
| NC score | 0.066114 (rank : 27) | θ value | θ > 10 (rank : 42) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9D869 | Gene names | Chp2, Hca520 | |||
|
Domain Architecture |

|
|||||
| Description | Calcineurin B homologous protein 2 (Hepatocellular carcinoma- associated antigen 520 homolog). | |||||
|
GUC1A_MOUSE
|
||||||
| NC score | 0.064756 (rank : 28) | θ value | θ > 10 (rank : 46) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 152 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P43081 | Gene names | Guca1a, Gcap, Gcap1, Guca1 | |||
|
Domain Architecture |

|
|||||
| Description | Guanylyl cyclase-activating protein 1 (GCAP 1) (Guanylate cyclase activator 1A). | |||||
|
GUC1A_HUMAN
|
||||||
| NC score | 0.064076 (rank : 29) | θ value | θ > 10 (rank : 45) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 147 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P43080, Q9NU14 | Gene names | GUCA1A, GCAP, GCAP1, GUCA1 | |||
|
Domain Architecture |

|
|||||
| Description | Guanylyl cyclase-activating protein 1 (GCAP 1) (Guanylate cyclase activator 1A). | |||||
|
CHIO_HUMAN
|
||||||
| NC score | 0.063407 (rank : 30) | θ value | θ > 10 (rank : 39) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 220 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P52757, Q75MM2 | Gene names | CHN2, ARHGAP3, BCH | |||
|
Domain Architecture |
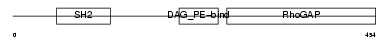
|
|||||
| Description | Beta-chimaerin (Beta-chimerin) (Rho-GTPase-activating protein 3). | |||||
|
UN13B_MOUSE
|
||||||
| NC score | 0.062246 (rank : 31) | θ value | θ > 10 (rank : 69) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9Z1N9 | Gene names | Unc13b, Unc13a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Unc-13 homolog B (Munc13-2) (munc13). | |||||
|
KPCL_HUMAN
|
||||||
| NC score | 0.062238 (rank : 32) | θ value | 7.1131e-05 (rank : 15) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 887 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P24723, Q16246 | Gene names | PRKCH, PKCL | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C eta type (EC 2.7.11.13) (nPKC-eta) (PKC-L). | |||||
|
NCALD_HUMAN
|
||||||
| NC score | 0.061187 (rank : 33) | θ value | θ > 10 (rank : 58) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P61601, P29554, Q8IYC3, Q9H0W2 | Gene names | NCALD | |||
|
Domain Architecture |

|
|||||
| Description | Neurocalcin delta. | |||||
|
NCALD_MOUSE
|
||||||
| NC score | 0.061187 (rank : 34) | θ value | θ > 10 (rank : 59) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 153 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q91X97, Q3TJS9, Q8BZN9 | Gene names | Ncald, D15Ertd412e | |||
|
Domain Architecture |

|
|||||
| Description | Neurocalcin delta. | |||||
|
HPCL1_MOUSE
|
||||||
| NC score | 0.060278 (rank : 35) | θ value | θ > 10 (rank : 53) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 149 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P62748, P35333 | Gene names | Hpcal1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hippocalcin-like protein 1 (Visinin-like protein 3) (VILIP-3) (Neural visinin-like protein 3) (NVL-3) (NVP-3). | |||||
|
HPCA_HUMAN
|
||||||
| NC score | 0.060087 (rank : 36) | θ value | θ > 10 (rank : 50) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P84074, P32076, P41211, P70510 | Gene names | HPCA, BDR2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuron-specific calcium-binding protein hippocalcin (Calcium-binding protein BDR-2). | |||||
|
HPCA_MOUSE
|
||||||
| NC score | 0.060087 (rank : 37) | θ value | θ > 10 (rank : 51) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P84075, P32076, P41211, P70510 | Gene names | Hpca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neuron-specific calcium-binding protein hippocalcin. | |||||
|
KPCL_MOUSE
|
||||||
| NC score | 0.060059 (rank : 38) | θ value | 0.000121331 (rank : 16) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 891 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P23298 | Gene names | Prkch, Pkch | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C eta type (EC 2.7.11.13) (nPKC-eta) (PKC-L). | |||||
|
KPCD_MOUSE
|
||||||
| NC score | 0.059715 (rank : 39) | θ value | 1.06291 (rank : 23) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 888 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P28867, Q91V85, Q9Z333 | Gene names | Prkcd, Pkcd | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C delta type (EC 2.7.11.13) (nPKC-delta). | |||||
|
CHIN_HUMAN
|
||||||
| NC score | 0.059422 (rank : 40) | θ value | θ > 10 (rank : 37) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P15882, Q53SD6, Q53SH5, Q96FB0 | Gene names | CHN1, ARHGAP2, CHN | |||
|
Domain Architecture |

|
|||||
| Description | N-chimaerin (NC) (N-chimerin) (Alpha chimerin) (A-chimaerin) (Rho- GTPase-activating protein 2). | |||||
|
KPCD_HUMAN
|
||||||
| NC score | 0.059298 (rank : 41) | θ value | 1.38821 (rank : 24) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 900 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q05655, Q15144 | Gene names | PRKCD | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C delta type (EC 2.7.11.13) (nPKC-delta). | |||||
|
CANB2_MOUSE
|
||||||
| NC score | 0.059135 (rank : 42) | θ value | θ > 10 (rank : 36) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q63811 | Gene names | Ppp3r2 | |||
|
Domain Architecture |

|
|||||
| Description | Calcineurin subunit B isoform 2 (Protein phosphatase 2B regulatory subunit 2) (Protein phosphatase 3 regulatory subunit B beta isoform). | |||||
|
GUC1B_MOUSE
|
||||||
| NC score | 0.058779 (rank : 43) | θ value | θ > 10 (rank : 48) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8VBV8 | Gene names | Guca1b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Guanylyl cyclase-activating protein 2 (GCAP 2) (Guanylate cyclase activator 1B). | |||||
|
HPCL1_HUMAN
|
||||||
| NC score | 0.058632 (rank : 44) | θ value | θ > 10 (rank : 52) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 150 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P37235, Q969S5 | Gene names | HPCAL1, BDR1 | |||
|
Domain Architecture |

|
|||||
| Description | Hippocalcin-like protein 1 (Visinin-like protein 3) (VILIP-3) (Calcium-binding protein BDR-1) (HLP2). | |||||
|
CHP2_HUMAN
|
||||||
| NC score | 0.058555 (rank : 45) | θ value | θ > 10 (rank : 41) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O43745 | Gene names | CHP2, HCA520 | |||
|
Domain Architecture |

|
|||||
| Description | Calcineurin B homologous protein 2 (Hepatocellular carcinoma- associated antigen 520). | |||||
|
RASF1_HUMAN
|
||||||
| NC score | 0.057856 (rank : 46) | θ value | θ > 10 (rank : 61) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9NS23, O14571, O60539, O60710, Q9HB04, Q9HB18, Q9NS22, Q9UND4, Q9UND5 | Gene names | RASSF1, RDA32 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ras association domain-containing protein 1. | |||||
|
GUC1B_HUMAN
|
||||||
| NC score | 0.057456 (rank : 47) | θ value | θ > 10 (rank : 47) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UMX6, Q9NU15 | Gene names | GUCA1B, GCAP2 | |||
|
Domain Architecture |

|
|||||
| Description | Guanylyl cyclase-activating protein 2 (GCAP 2) (Guanylate cyclase activator 1B). | |||||
|
KPCT_MOUSE
|
||||||
| NC score | 0.057402 (rank : 48) | θ value | 0.279714 (rank : 19) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 895 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q02111 | Gene names | Prkcq, Pkcq | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C theta type (EC 2.7.11.13) (nPKC-theta). | |||||
|
KPCT_HUMAN
|
||||||
| NC score | 0.057170 (rank : 49) | θ value | 0.279714 (rank : 18) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 897 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q04759, Q3MJF1, Q64FY5, Q9H508, Q9H549 | Gene names | PRKCQ, PRKCT | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C theta type (EC 2.7.11.13) (nPKC-theta). | |||||
|
CHIN_MOUSE
|
||||||
| NC score | 0.055474 (rank : 50) | θ value | θ > 10 (rank : 38) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 229 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q91V57, Q3UGY3, Q7TQE5, Q8BWU6, Q9D9B3 | Gene names | Chn1, Arhgap2 | |||
|
Domain Architecture |

|
|||||
| Description | N-chimaerin (NC) (N-chimerin) (Alpha chimerin) (A-chimaerin) (Rho- GTPase-activating protein 2). | |||||
|
CANB1_MOUSE
|
||||||
| NC score | 0.055161 (rank : 51) | θ value | θ > 10 (rank : 34) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q63810 | Gene names | Ppp3r1, Cnb | |||
|
Domain Architecture |

|
|||||
| Description | Calcineurin subunit B isoform 1 (Protein phosphatase 2B regulatory subunit 1) (Protein phosphatase 3 regulatory subunit B alpha isoform 1). | |||||
|
CANB1_HUMAN
|
||||||
| NC score | 0.055156 (rank : 52) | θ value | θ > 10 (rank : 33) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P63098, P06705, P15117, Q08044 | Gene names | PPP3R1, CNA2, CNB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcineurin subunit B isoform 1 (Protein phosphatase 2B regulatory subunit 1) (Protein phosphatase 3 regulatory subunit B alpha isoform 1). | |||||
|
KPCG_HUMAN
|
||||||
| NC score | 0.053829 (rank : 53) | θ value | 1.81305 (rank : 25) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 1126 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P05129 | Gene names | PRKCG, PKCG | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C gamma type (EC 2.7.11.13) (PKC-gamma). | |||||
|
KPCE_MOUSE
|
||||||
| NC score | 0.053684 (rank : 54) | θ value | θ > 10 (rank : 57) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P16054 | Gene names | Prkce, Pkce, Pkcea | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C epsilon type (EC 2.7.11.13) (nPKC-epsilon). | |||||
|
CANB2_HUMAN
|
||||||
| NC score | 0.053516 (rank : 55) | θ value | θ > 10 (rank : 35) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q96LZ3, Q7Z4V8, Q8WYJ4 | Gene names | PPP3R2, CBLP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcineurin subunit B isoform 2 (Protein phosphatase 2B regulatory subunit 2) (Protein phosphatase 3 regulatory subunit B beta isoform) (Calcineurin B-like protein) (CBLP) (CNBII). | |||||
|
KPCG_MOUSE
|
||||||
| NC score | 0.053454 (rank : 56) | θ value | 1.81305 (rank : 26) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 1123 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P63318, P05697 | Gene names | Prkcg, Pkcc, Pkcg, Prkcc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein kinase C gamma type (EC 2.7.11.13) (PKC-gamma). | |||||
|
KPCE_HUMAN
|
||||||
| NC score | 0.053158 (rank : 57) | θ value | θ > 10 (rank : 56) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 906 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q02156, Q53SL4, Q53SM5, Q9UE81 | Gene names | PRKCE, PKCE | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C epsilon type (EC 2.7.11.13) (nPKC-epsilon). | |||||
|
GUC1C_HUMAN
|
||||||
| NC score | 0.053141 (rank : 58) | θ value | θ > 10 (rank : 49) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O95843, O95844, Q9UNM0 | Gene names | GUCA1C, GCAP3 | |||
|
Domain Architecture |

|
|||||
| Description | Guanylyl cyclase-activating protein 3 (GCAP 3) (Guanylate cyclase activator 1C). | |||||
|
EFHD1_HUMAN
|
||||||
| NC score | 0.052466 (rank : 59) | θ value | θ > 10 (rank : 43) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9BUP0, Q9BTF8, Q9H8I2, Q9HBQ0 | Gene names | EFHD1, SWS2 | |||
|
Domain Architecture |

|
|||||
| Description | EF-hand domain-containing protein 1 (Swiprosin-2). | |||||
|
ZN277_HUMAN
|
||||||
| NC score | 0.037589 (rank : 60) | θ value | 0.47712 (rank : 20) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NRM2 | Gene names | ZNF277 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 277. | |||||
|
RAF1_MOUSE
|
||||||
| NC score | 0.021785 (rank : 61) | θ value | 8.99809 (rank : 32) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 886 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q99N57, Q91WH1 | Gene names | Raf1, Craf | |||
|
Domain Architecture |

|
|||||
| Description | RAF proto-oncogene serine/threonine-protein kinase (EC 2.7.11.1) (Raf- 1) (C-RAF) (cRaf). | |||||
|
RAF1_HUMAN
|
||||||
| NC score | 0.021583 (rank : 62) | θ value | 8.99809 (rank : 31) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 886 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P04049 | Gene names | RAF1, RAF | |||
|
Domain Architecture |

|
|||||
| Description | RAF proto-oncogene serine/threonine-protein kinase (EC 2.7.11.1) (Raf- 1) (C-RAF) (cRaf). | |||||
|
NTNG1_HUMAN
|
||||||
| NC score | 0.017258 (rank : 63) | θ value | 0.62314 (rank : 21) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Y2I2, Q5VU86, Q5VU87, Q5VU89, Q5VU90, Q5VU91, Q7Z2Y3, Q8N633 | Gene names | NTNG1, KIAA0976, LMNT1 | |||
|
Domain Architecture |

|
|||||
| Description | Netrin G1 precursor (Laminet-1). | |||||
|
NTNG1_MOUSE
|
||||||
| NC score | 0.016887 (rank : 64) | θ value | 0.813845 (rank : 22) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8R4G0, Q68FE5, Q69ZU3, Q8R4F3, Q8R4F4, Q8R4F5, Q8R4F6, Q8R4F7, Q8R4F8, Q8R4F9, Q9ESR3, Q9ESR4, Q9ESR5, Q9ESR6, Q9ESR7, Q9ESR8 | Gene names | Ntng1, Kiaa0976, Lmnt1 | |||
|
Domain Architecture |

|
|||||
| Description | Netrin G1 precursor (Laminet-1). | |||||
|
PHF14_MOUSE
|
||||||
| NC score | 0.015307 (rank : 65) | θ value | 6.88961 (rank : 29) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9D4H9, Q810Z6, Q8C284, Q8CGF8, Q9D1Q8 | Gene names | Phf14 | |||
|
Domain Architecture |

|
|||||
| Description | PHD finger protein 14. | |||||
|
PHF14_HUMAN
|
||||||
| NC score | 0.014746 (rank : 66) | θ value | 6.88961 (rank : 28) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O94880 | Gene names | PHF14, KIAA0783 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PHD finger protein 14. | |||||
|
KSR1_MOUSE
|
||||||
| NC score | 0.013124 (rank : 67) | θ value | 8.99809 (rank : 30) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 870 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q61097, Q61648, Q78DX8 | Gene names | Ksr1, Ksr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinase suppressor of ras-1 (Kinase suppressor of ras) (mKSR1) (Hb protein). | |||||
|
USH2A_HUMAN
|
||||||
| NC score | 0.012556 (rank : 68) | θ value | 0.21417 (rank : 17) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 600 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O75445, Q5VVM9, Q6S362, Q9NS27 | Gene names | USH2A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Usherin precursor (Usher syndrome type-2A protein) (Usher syndrome type IIa protein). | |||||
|
BRPF3_HUMAN
|
||||||
| NC score | 0.010500 (rank : 69) | θ value | 4.03905 (rank : 27) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9ULD4, Q5R3K8 | Gene names | BRPF3, KIAA1286 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain and PHD finger-containing protein 3. | |||||