Please be patient as the page loads
|
TOPB1_HUMAN
|
||||||
| SwissProt Accessions | Q92547, Q7LGC1, Q9UEB9 | Gene names | TOPBP1, KIAA0259 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA topoisomerase 2-binding protein 1 (DNA topoisomerase II-binding protein 1) (DNA topoisomerase IIbeta-binding protein 1) (TopBP1). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
TOPB1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | Q92547, Q7LGC1, Q9UEB9 | Gene names | TOPBP1, KIAA0259 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA topoisomerase 2-binding protein 1 (DNA topoisomerase II-binding protein 1) (DNA topoisomerase IIbeta-binding protein 1) (TopBP1). | |||||
|
TOPB1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.973910 (rank : 2) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q6ZQF0, Q6P6P0, Q80Y33, Q8BUI0, Q8BUK1, Q8R348, Q91VX3, Q922X8 | Gene names | Topbp1, Kiaa0259 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA topoisomerase II-binding protein 1 (DNA topoisomerase IIbeta- binding protein 1) (TopBP1). | |||||
|
ECT2_HUMAN
|
||||||
| θ value | 5.40856e-29 (rank : 3) | NC score | 0.363965 (rank : 3) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H8V3, Q9NSV8, Q9NVW9 | Gene names | ECT2 | |||
|
Domain Architecture |

|
|||||
| Description | ECT2 protein (Epithelial cell-transforming sequence 2 oncogene). | |||||
|
ECT2_MOUSE
|
||||||
| θ value | 3.88503e-19 (rank : 4) | NC score | 0.322011 (rank : 4) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q07139 | Gene names | Ect2 | |||
|
Domain Architecture |

|
|||||
| Description | ECT2 protein (Epithelial cell-transforming sequence 2 oncogene). | |||||
|
MCPH1_HUMAN
|
||||||
| θ value | 2.44474e-05 (rank : 5) | NC score | 0.197331 (rank : 5) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8NEM0, Q66GU1, Q9H9C7 | Gene names | MCPH1 | |||
|
Domain Architecture |

|
|||||
| Description | Microcephalin. | |||||
|
XRCC1_MOUSE
|
||||||
| θ value | 0.000602161 (rank : 6) | NC score | 0.180190 (rank : 6) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q60596, Q3THC5, Q5U435, Q7TNQ5 | Gene names | Xrcc1, Xrcc-1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-repair protein XRCC1 (X-ray repair cross-complementing protein 1). | |||||
|
REV1_MOUSE
|
||||||
| θ value | 0.00175202 (rank : 7) | NC score | 0.130748 (rank : 9) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q920Q2, Q9QXV2 | Gene names | Rev1l, Rev1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein REV1 (EC 2.7.7.-) (Rev1-like terminal deoxycytidyl transferase). | |||||
|
REV1_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 8) | NC score | 0.109051 (rank : 10) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UBZ9, O95941, Q9C0J4, Q9NUP2 | Gene names | REV1L, REV1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein REV1 (EC 2.7.7.-) (Rev1-like terminal deoxycytidyl transferase) (Alpha integrin-binding protein 80) (AIBP80). | |||||
|
BARD1_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 9) | NC score | 0.041484 (rank : 22) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 411 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O70445 | Gene names | Bard1 | |||
|
Domain Architecture |

|
|||||
| Description | BRCA1-associated RING domain protein 1 (BARD-1). | |||||
|
MCPH1_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 10) | NC score | 0.149967 (rank : 7) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q7TT79, Q8BNI9, Q8C9I7 | Gene names | Mcph1 | |||
|
Domain Architecture |

|
|||||
| Description | Microcephalin. | |||||
|
XRCC1_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 11) | NC score | 0.147604 (rank : 8) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P18887, Q9HCB1 | Gene names | XRCC1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-repair protein XRCC1 (X-ray repair cross-complementing protein 1). | |||||
|
MLL2_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 12) | NC score | 0.036554 (rank : 25) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
PCLO_MOUSE
|
||||||
| θ value | 0.163984 (rank : 13) | NC score | 0.031415 (rank : 29) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
CE152_HUMAN
|
||||||
| θ value | 0.21417 (rank : 14) | NC score | 0.026676 (rank : 35) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 976 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O94986, Q6NTA0 | Gene names | CEP152, KIAA0912 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 152 kDa (Cep152 protein). | |||||
|
VCIP1_HUMAN
|
||||||
| θ value | 0.21417 (rank : 15) | NC score | 0.056330 (rank : 13) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96JH7, Q504T4, Q86T93, Q86W01, Q8N3A9, Q9H5R8 | Gene names | VCPIP1, KIAA1850, VCIP135 | |||
|
Domain Architecture |

|
|||||
| Description | Deubiquitinating protein VCIP135 (EC 3.4.22.-) (Valosin-containing protein p97/p47 complex-interacting protein p135) (Valosin-containing protein p97/p47 complex-interacting protein 1). | |||||
|
LAMB2_MOUSE
|
||||||
| θ value | 0.279714 (rank : 16) | NC score | 0.014741 (rank : 59) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 722 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q61292, Q62182 | Gene names | Lamb2, Lams | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-2 chain precursor (S-laminin) (S-LAM). | |||||
|
SMBT1_HUMAN
|
||||||
| θ value | 0.279714 (rank : 17) | NC score | 0.042997 (rank : 19) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UHJ3, Q96C73, Q9Y4Q9 | Gene names | SFMBT1, RU1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Scm-like with four MBT domains protein 1 (Renal ubiquitous protein 1). | |||||
|
SMBT1_MOUSE
|
||||||
| θ value | 0.279714 (rank : 18) | NC score | 0.043064 (rank : 18) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9JMD1, Q6NZD3, Q8CFS1 | Gene names | Sfmbt1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Scm-like with four MBT domains protein 1. | |||||
|
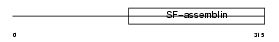
DMN_HUMAN
|
||||||
| θ value | 0.365318 (rank : 19) | NC score | 0.026560 (rank : 36) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O15061 | Gene names | DMN, KIAA0353 | |||
|
Domain Architecture |
|
|||||
| Description | Desmuslin. | |||||
|
PARP1_MOUSE
|
||||||
| θ value | 0.47712 (rank : 20) | NC score | 0.043679 (rank : 17) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P11103, Q9JLX4, Q9QVQ3 | Gene names | Parp1, Adprp, Adprt, Adprt1 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 1 (EC 2.4.2.30) (PARP-1) (ADPRT) (NAD(+) ADP-ribosyltransferase 1) (Poly[ADP-ribose] synthetase 1) (msPARP). | |||||
|
CDKL2_MOUSE
|
||||||
| θ value | 0.62314 (rank : 21) | NC score | 0.003394 (rank : 81) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 854 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9QUK0, Q3UL60, Q3V3X7, Q9QYI1, Q9QYI2 | Gene names | Cdkl2, Kkiamre, Kkm | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-dependent kinase-like 2 (EC 2.7.11.22) (Serine/threonine- protein kinase KKIAMRE). | |||||
|
MINT_HUMAN
|
||||||
| θ value | 0.62314 (rank : 22) | NC score | 0.025649 (rank : 37) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1291 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q96T58, Q9H9A8, Q9NWH5, Q9UQ01, Q9Y556 | Gene names | SPEN, KIAA0929, MINT, SHARP | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
MYC_HUMAN
|
||||||
| θ value | 0.62314 (rank : 23) | NC score | 0.031265 (rank : 30) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P01106, P01107, Q14026 | Gene names | MYC | |||
|
Domain Architecture |

|
|||||
| Description | Myc proto-oncogene protein (c-Myc) (Transcription factor p64). | |||||
|
FBX38_MOUSE
|
||||||
| θ value | 0.813845 (rank : 24) | NC score | 0.035143 (rank : 27) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BMI0, Q8BTQ4, Q8C0H4 | Gene names | Fbxo38 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 38 (Modulator of KLF7 activity) (MoKA). | |||||
|
THOC2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 25) | NC score | 0.044571 (rank : 15) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8NI27, Q9H8I6 | Gene names | THOC2, CXorf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | THO complex subunit 2 (Tho2). | |||||
|
ATP5S_MOUSE
|
||||||
| θ value | 1.06291 (rank : 26) | NC score | 0.074643 (rank : 11) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9CRA7, Q8R2M3 | Gene names | Atp5s, Atpw | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP synthase subunit s, mitochondrial precursor (ATP synthase coupling factor B) (Mitochondrial ATP synthase regulatory component factor B). | |||||
|
BRD3_MOUSE
|
||||||
| θ value | 1.06291 (rank : 27) | NC score | 0.021424 (rank : 43) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 402 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8K2F0, Q8C665, Q8CAX7, Q9JI25 | Gene names | Brd3, Fsrg2 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 3 (Bromodomain-containing FSH-like protein FSRG2). | |||||
|
ELK4_MOUSE
|
||||||
| θ value | 1.06291 (rank : 28) | NC score | 0.018023 (rank : 50) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P41158 | Gene names | Elk4, Sap1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-4 (Serum response factor accessory protein 1) (SAP-1). | |||||
|
PCLO_HUMAN
|
||||||
| θ value | 1.06291 (rank : 29) | NC score | 0.030011 (rank : 33) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
RAD9B_HUMAN
|
||||||
| θ value | 1.06291 (rank : 30) | NC score | 0.044128 (rank : 16) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 5 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6WBX8, Q5U5K0, Q6NVJ1, Q6ZVT7, Q8N7T9, Q96LI8 | Gene names | RAD9B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell cycle checkpoint control protein RAD9B homolog (RAD9 homolog B) (hRAD9B). | |||||
|
BARD1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 31) | NC score | 0.042053 (rank : 21) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q99728, O43574 | Gene names | BARD1 | |||
|
Domain Architecture |

|
|||||
| Description | BRCA1-associated RING domain protein 1 (BARD-1). | |||||
|
PARP1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 32) | NC score | 0.039959 (rank : 23) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P09874, Q8IUZ9 | Gene names | PARP1, ADPRT, PPOL | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 1 (EC 2.4.2.30) (PARP-1) (ADPRT) (NAD(+) ADP-ribosyltransferase 1) (Poly[ADP-ribose] synthetase 1). | |||||
|
CCNB3_MOUSE
|
||||||
| θ value | 1.81305 (rank : 33) | NC score | 0.020368 (rank : 45) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q810T2, Q810T3, Q8VDC8 | Gene names | Ccnb3, Cycb3 | |||
|
Domain Architecture |

|
|||||
| Description | G2/mitotic-specific cyclin-B3. | |||||
|
CG1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 34) | NC score | 0.038062 (rank : 24) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q13495 | Gene names | CXorf6, CG1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CG1 protein (F18). | |||||
|
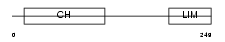
MILK2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 35) | NC score | 0.021421 (rank : 44) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 457 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8IY33, Q7RTP4, Q7Z655, Q8TEQ4, Q9H5F9 | Gene names | ||||
|
Domain Architecture |
|
|||||
| Description | MICAL-like protein 2. | |||||
|
PARG_MOUSE
|
||||||
| θ value | 1.81305 (rank : 36) | NC score | 0.030429 (rank : 31) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O88622, Q80YQ6, Q8CB72 | Gene names | Parg | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly(ADP-ribose) glycohydrolase (EC 3.2.1.143). | |||||
|
TAF3_HUMAN
|
||||||
| θ value | 1.81305 (rank : 37) | NC score | 0.030097 (rank : 32) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 568 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q5VWG9, Q6GMS5, Q6P6B5, Q86VY6, Q9BQS9, Q9UFI8 | Gene names | TAF3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID subunit 3 (TBP-associated factor 3) (Transcription initiation factor TFIID 140 kDa subunit) (140 kDa TATA box-binding protein-associated factor) (TAF140) (TAFII140). | |||||
|
ZFP59_MOUSE
|
||||||
| θ value | 1.81305 (rank : 38) | NC score | 0.000708 (rank : 84) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 768 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P16373, Q61849 | Gene names | Zfp59, Mfg2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 59 (Zfp-59) (Zinc finger protein Mfg-2). | |||||
|
ATX2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 39) | NC score | 0.022045 (rank : 41) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 375 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q99700, Q6ZQZ7, Q99493 | Gene names | ATXN2, ATX2, SCA2, TNRC13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-2 (Spinocerebellar ataxia type 2 protein) (Trinucleotide repeat-containing gene 13 protein). | |||||
|
BRCA1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 40) | NC score | 0.029296 (rank : 34) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 381 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P38398 | Gene names | BRCA1, RNF53 | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer type 1 susceptibility protein (RING finger protein 53). | |||||
|
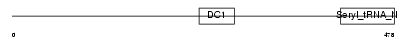
DTNA_MOUSE
|
||||||
| θ value | 2.36792 (rank : 41) | NC score | 0.024301 (rank : 39) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 264 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9D2N4, P97319, Q61498, Q61499, Q9QZZ5, Q9WUL9, Q9WUM0 | Gene names | Dtna, Dtn | |||
|
Domain Architecture |
|
|||||
| Description | Dystrobrevin alpha (Alpha-dystrobrevin). | |||||
|
OFD1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 42) | NC score | 0.021740 (rank : 42) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 753 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O75665, O75666 | Gene names | OFD1, CXorf5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Oral-facial-digital syndrome 1 protein (Protein 71-7A). | |||||
|
SMC2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 43) | NC score | 0.022803 (rank : 40) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1405 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O95347, Q9P1P2 | Gene names | SMC2L1, CAPE, SMC2 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 2-like 1 protein (Chromosome- associated protein E) (hCAP-E) (XCAP-E homolog). | |||||
|
FYCO1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 44) | NC score | 0.015078 (rank : 58) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1059 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8VDC1, Q3V385, Q7TMT0, Q8BJN2 | Gene names | Fyco1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE and coiled-coil domain-containing protein 1. | |||||
|
GP158_MOUSE
|
||||||
| θ value | 3.0926 (rank : 45) | NC score | 0.019406 (rank : 47) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8C419, Q8CHB0 | Gene names | Gpr158, Kiaa1136 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 158 precursor. | |||||
|
ITAD_MOUSE
|
||||||
| θ value | 3.0926 (rank : 46) | NC score | 0.008544 (rank : 73) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q3V0T4 | Gene names | Itgad | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Integrin alpha-D precursor (CD11d antigen). | |||||
|
SPTA1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 47) | NC score | 0.012287 (rank : 65) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 675 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P08032, P97502 | Gene names | Spta1, Spna1, Spta | |||
|
Domain Architecture |

|
|||||
| Description | Spectrin alpha chain, erythrocyte (Erythroid alpha-spectrin). | |||||
|
TXLNB_HUMAN
|
||||||
| θ value | 3.0926 (rank : 48) | NC score | 0.016457 (rank : 54) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 912 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8N3L3, Q76L25, Q86T52, Q8N3S2 | Gene names | TXLNB, C6orf198, MDP77 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-taxilin (Muscle-derived protein 77) (hMDP77). | |||||
|
CO027_HUMAN
|
||||||
| θ value | 4.03905 (rank : 49) | NC score | 0.025482 (rank : 38) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q2M3C6, Q8N993, Q96LL5 | Gene names | C15orf27 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C15orf27. | |||||
|
DSPP_MOUSE
|
||||||
| θ value | 4.03905 (rank : 50) | NC score | 0.018468 (rank : 48) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P97399, O70567 | Gene names | Dspp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor (Dentin matrix protein 3) (DMP-3) [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
KINH_HUMAN
|
||||||
| θ value | 4.03905 (rank : 51) | NC score | 0.012495 (rank : 63) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1231 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P33176, Q5VZ85 | Gene names | KIF5B, KNS, KNS1 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin heavy chain (Ubiquitous kinesin heavy chain) (UKHC). | |||||
|
KINH_MOUSE
|
||||||
| θ value | 4.03905 (rank : 52) | NC score | 0.012436 (rank : 64) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1082 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q61768, O08711, Q61580 | Gene names | Kif5b, Khcs, Kns1 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin heavy chain (Ubiquitous kinesin heavy chain) (UKHC). | |||||
|
PTPRZ_HUMAN
|
||||||
| θ value | 4.03905 (rank : 53) | NC score | 0.006220 (rank : 78) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P23471 | Gene names | PTPRZ1, PTPRZ, PTPZ | |||
|
Domain Architecture |

|
|||||
| Description | Receptor-type tyrosine-protein phosphatase zeta precursor (EC 3.1.3.48) (R-PTP-zeta). | |||||
|
RAD9A_HUMAN
|
||||||
| θ value | 4.03905 (rank : 54) | NC score | 0.035898 (rank : 26) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99638, Q6FI29, Q96C41 | Gene names | RAD9A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell cycle checkpoint control protein RAD9A (EC 3.1.11.2) (DNA repair exonuclease rad9 homolog A) (hRAD9). | |||||
|
VCIP1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 55) | NC score | 0.042906 (rank : 20) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8CDG3, Q7TMU9, Q8BP90 | Gene names | Vcpip1, Vcip135 | |||
|
Domain Architecture |

|
|||||
| Description | Deubiquitinating protein VCIP135 (EC 3.4.22.-) (Valosin-containing protein p97/p47 complex-interacting protein p135) (Valosin-containing protein p97/p47 complex-interacting protein 1). | |||||
|
DMD_HUMAN
|
||||||
| θ value | 5.27518 (rank : 56) | NC score | 0.017582 (rank : 51) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P11532, Q02295, Q14169, Q14170, Q5JYU0, Q7KZ48 | Gene names | DMD | |||
|
Domain Architecture |

|
|||||
| Description | Dystrophin. | |||||
|
DMD_MOUSE
|
||||||
| θ value | 5.27518 (rank : 57) | NC score | 0.017382 (rank : 52) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1111 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P11531, O35653, Q60703 | Gene names | Dmd | |||
|
Domain Architecture |

|
|||||
| Description | Dystrophin. | |||||
|
RAD9A_MOUSE
|
||||||
| θ value | 5.27518 (rank : 58) | NC score | 0.034365 (rank : 28) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Z0F6, Q8QZZ3 | Gene names | Rad9a, Rad9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell cycle checkpoint control protein RAD9A (EC 3.1.11.2) (DNA repair exonuclease rad9 homolog A) (Rad9-like protein) (mRAD9). | |||||
|
TAF1L_HUMAN
|
||||||
| θ value | 5.27518 (rank : 59) | NC score | 0.015734 (rank : 55) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8IZX4 | Gene names | TAF1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID 210 kDa subunit (TBP-associated factor 210 kDa) (TAF(II)210) (TBP-associated factor 1-like). | |||||
|
TARA_MOUSE
|
||||||
| θ value | 5.27518 (rank : 60) | NC score | 0.017098 (rank : 53) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1133 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q99KW3, Q2PZW9, Q2Q402, Q2Q403, Q2Q404, Q6ZPK4, Q8C6T3, Q8CG90 | Gene names | Triobp, Kiaa1662, Tara | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
ZBT20_HUMAN
|
||||||
| θ value | 5.27518 (rank : 61) | NC score | 0.001096 (rank : 83) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 835 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9HC78 | Gene names | ZBTB20, DPZF, ZNF288 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and BTB domain-containing protein 20 (Zinc finger protein 288) (Dendritic-derived BTB/POZ zinc finger protein). | |||||
|
ZN281_HUMAN
|
||||||
| θ value | 5.27518 (rank : 62) | NC score | 0.001625 (rank : 82) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 745 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Y2X9, Q9NY92 | Gene names | ZNF281, GZP1, ZBP99 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 281 (Zinc finger DNA-binding protein 99) (Transcription factor ZBP-99) (GC-box-binding zinc finger protein 1). | |||||
|
BPA1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 63) | NC score | 0.019781 (rank : 46) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1477 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q91ZU6, Q91ZU7 | Gene names | Dst, Bpag1 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoforms 1/2/3/4 (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
CTDP1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 64) | NC score | 0.018355 (rank : 49) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Y5B0, Q7Z644, Q96BZ1, Q9Y6F5 | Gene names | CTDP1, FCP1 | |||
|
Domain Architecture |

|
|||||
| Description | RNA polymerase II subunit A C-terminal domain phosphatase (EC 3.1.3.16) (TFIIF-associating CTD phosphatase). | |||||
|
DDX51_HUMAN
|
||||||
| θ value | 6.88961 (rank : 65) | NC score | 0.005913 (rank : 79) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8N8A6, Q5CZ71, Q8IXK5, Q96ED1 | Gene names | DDX51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent RNA helicase DDX51 (EC 3.6.1.-) (DEAD box protein 51). | |||||
|
FIGN_HUMAN
|
||||||
| θ value | 6.88961 (rank : 66) | NC score | 0.007725 (rank : 77) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5HY92, Q9H6M5, Q9NVZ9 | Gene names | FIGN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fidgetin. | |||||
|
K0319_HUMAN
|
||||||
| θ value | 6.88961 (rank : 67) | NC score | 0.015595 (rank : 56) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q5VV43, Q9UJC8, Q9Y4G7 | Gene names | KIAA0319 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0319 precursor. | |||||
|
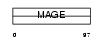
MAGB6_HUMAN
|
||||||
| θ value | 6.88961 (rank : 68) | NC score | 0.008123 (rank : 75) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8N7X4, Q6GS19, Q9H219 | Gene names | MAGEB6 | |||
|
Domain Architecture |
|
|||||
| Description | Melanoma-associated antigen B6 (MAGE-B6 antigen). | |||||
|
RAI14_HUMAN
|
||||||
| θ value | 6.88961 (rank : 69) | NC score | 0.009781 (rank : 71) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1367 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9P0K7, Q6V1W9, Q7Z5I4, Q7Z733, Q9P2L2, Q9Y3T5 | Gene names | RAI14, KIAA1334, NORPEG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankycorbin (Ankyrin repeat and coiled-coil structure-containing protein) (Retinoic acid-induced protein 14) (Novel retinal pigment epithelial cell protein). | |||||
|
ARMX5_HUMAN
|
||||||
| θ value | 8.99809 (rank : 70) | NC score | 0.009328 (rank : 72) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6P1M9, Q68DB4, Q9BVZ3, Q9H969 | Gene names | ARMCX5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing X-linked protein 5. | |||||
|
CLMN_HUMAN
|
||||||
| θ value | 8.99809 (rank : 71) | NC score | 0.013862 (rank : 60) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q96JQ2, Q9H713, Q9HA23, Q9HA57, Q9UFP4, Q9ULN2 | Gene names | CLMN, KIAA1188 | |||
|
Domain Architecture |
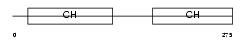
|
|||||
| Description | Calmin. | |||||
|
ERC2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 72) | NC score | 0.011044 (rank : 69) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1166 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q6PH08, Q80U20, Q8CCP1 | Gene names | Erc2, Cast1, D14Ertd171e, Kiaa0378 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ERC protein 2 (CAZ-associated structural protein 1) (CAST1). | |||||
|
FIGLA_MOUSE
|
||||||
| θ value | 8.99809 (rank : 73) | NC score | 0.011799 (rank : 67) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O55208 | Gene names | Figla | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Factor in the germline alpha (Transcription factor FIGa) (FIGalpha). | |||||
|
GON4L_MOUSE
|
||||||
| θ value | 8.99809 (rank : 74) | NC score | 0.013320 (rank : 61) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9DB00, Q80TB4, Q91YI9 | Gene names | Gon4l, Gon4, KIAA1606 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GON-4-like protein (GON-4 homolog). | |||||
|
KI18A_MOUSE
|
||||||
| θ value | 8.99809 (rank : 75) | NC score | 0.007833 (rank : 76) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q91WD7, Q3TP77, Q8BLL1 | Gene names | Kif18a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin family member 18A. | |||||
|
LRP12_HUMAN
|
||||||
| θ value | 8.99809 (rank : 76) | NC score | 0.004641 (rank : 80) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y561 | Gene names | LRP12, ST7 | |||
|
Domain Architecture |
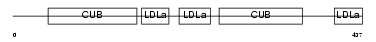
|
|||||
| Description | Low-density lipoprotein receptor-related protein 12 precursor (Suppressor of tumorigenicity protein 7). | |||||
|
NFH_HUMAN
|
||||||
| θ value | 8.99809 (rank : 77) | NC score | 0.008474 (rank : 74) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1242 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P12036, Q9UJS7, Q9UQ14 | Gene names | NEFH, KIAA0845, NFH | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
RRBP1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 78) | NC score | 0.011896 (rank : 66) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1650 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q99PL5, Q99PK5, Q99PK6, Q99PK7, Q99PK8, Q99PK9, Q99PL0, Q99PL1, Q99PL2, Q99PL3, Q99PL4, Q9CS20 | Gene names | Rrbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (mRRp). | |||||
|
SCG2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 79) | NC score | 0.012654 (rank : 62) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q03517, Q80Y79, Q9CW80 | Gene names | Scg2, Chgc, Scg-2 | |||
|
Domain Architecture |

|
|||||
| Description | Secretogranin-2 precursor (Secretogranin II) (SgII) (Chromogranin C) [Contains: Secretoneurin (SN)]. | |||||
|
SETX_HUMAN
|
||||||
| θ value | 8.99809 (rank : 80) | NC score | 0.011262 (rank : 68) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q7Z333, O75120, Q3KQX4, Q68DW5, Q6AZD7, Q7Z3J6, Q8WX33, Q9H9D1, Q9NVP9 | Gene names | SETX, ALS4, KIAA0625, SCAR1 | |||
|
Domain Architecture |

|
|||||
| Description | Probable helicase senataxin (EC 3.6.1.-) (SEN1 homolog). | |||||
|
TDT_HUMAN
|
||||||
| θ value | 8.99809 (rank : 81) | NC score | 0.009863 (rank : 70) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P04053, Q96E50 | Gene names | DNTT, TDT | |||
|
Domain Architecture |

|
|||||
| Description | DNA nucleotidylexotransferase (EC 2.7.7.31) (Terminal addition enzyme) (Terminal deoxynucleotidyltransferase) (Terminal transferase). | |||||
|
ZC11A_MOUSE
|
||||||
| θ value | 8.99809 (rank : 82) | NC score | 0.015120 (rank : 57) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6NZF1, Q6NXW9, Q6PDR6, Q80TU7, Q8C5L5, Q99JN6 | Gene names | Zc3h11a, Kiaa0663 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 11A. | |||||
|
ARHGA_HUMAN
|
||||||
| θ value | θ > 10 (rank : 83) | NC score | 0.055562 (rank : 14) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O15013, O14665, Q68D55, Q8IWD9, Q8IY77 | Gene names | ARHGEF10, KIAA0294 | |||
|
Domain Architecture |
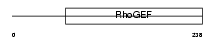
|
|||||
| Description | Rho guanine nucleotide exchange factor 10. | |||||
|
ARHGA_MOUSE
|
||||||
| θ value | θ > 10 (rank : 84) | NC score | 0.057992 (rank : 12) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8C033, Q5DU38, Q80VH8, Q8BW76, Q922S7 | Gene names | Arhgef10, Kiaa0294 | |||
|
Domain Architecture |
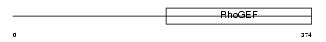
|
|||||
| Description | Rho guanine nucleotide exchange factor 10. | |||||
|
TOPB1_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | Q92547, Q7LGC1, Q9UEB9 | Gene names | TOPBP1, KIAA0259 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA topoisomerase 2-binding protein 1 (DNA topoisomerase II-binding protein 1) (DNA topoisomerase IIbeta-binding protein 1) (TopBP1). | |||||
|
TOPB1_MOUSE
|
||||||
| NC score | 0.973910 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q6ZQF0, Q6P6P0, Q80Y33, Q8BUI0, Q8BUK1, Q8R348, Q91VX3, Q922X8 | Gene names | Topbp1, Kiaa0259 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA topoisomerase II-binding protein 1 (DNA topoisomerase IIbeta- binding protein 1) (TopBP1). | |||||
|
ECT2_HUMAN
|
||||||
| NC score | 0.363965 (rank : 3) | θ value | 5.40856e-29 (rank : 3) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9H8V3, Q9NSV8, Q9NVW9 | Gene names | ECT2 | |||
|
Domain Architecture |

|
|||||
| Description | ECT2 protein (Epithelial cell-transforming sequence 2 oncogene). | |||||
|
ECT2_MOUSE
|
||||||
| NC score | 0.322011 (rank : 4) | θ value | 3.88503e-19 (rank : 4) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q07139 | Gene names | Ect2 | |||
|
Domain Architecture |

|
|||||
| Description | ECT2 protein (Epithelial cell-transforming sequence 2 oncogene). | |||||
|
MCPH1_HUMAN
|
||||||
| NC score | 0.197331 (rank : 5) | θ value | 2.44474e-05 (rank : 5) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8NEM0, Q66GU1, Q9H9C7 | Gene names | MCPH1 | |||
|
Domain Architecture |

|
|||||
| Description | Microcephalin. | |||||
|
XRCC1_MOUSE
|
||||||
| NC score | 0.180190 (rank : 6) | θ value | 0.000602161 (rank : 6) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q60596, Q3THC5, Q5U435, Q7TNQ5 | Gene names | Xrcc1, Xrcc-1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-repair protein XRCC1 (X-ray repair cross-complementing protein 1). | |||||
|
MCPH1_MOUSE
|
||||||
| NC score | 0.149967 (rank : 7) | θ value | 0.0431538 (rank : 10) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q7TT79, Q8BNI9, Q8C9I7 | Gene names | Mcph1 | |||
|
Domain Architecture |

|
|||||
| Description | Microcephalin. | |||||
|
XRCC1_HUMAN
|
||||||
| NC score | 0.147604 (rank : 8) | θ value | 0.0431538 (rank : 11) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P18887, Q9HCB1 | Gene names | XRCC1 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-repair protein XRCC1 (X-ray repair cross-complementing protein 1). | |||||
|
REV1_MOUSE
|
||||||
| NC score | 0.130748 (rank : 9) | θ value | 0.00175202 (rank : 7) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q920Q2, Q9QXV2 | Gene names | Rev1l, Rev1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein REV1 (EC 2.7.7.-) (Rev1-like terminal deoxycytidyl transferase). | |||||
|
REV1_HUMAN
|
||||||
| NC score | 0.109051 (rank : 10) | θ value | 0.0330416 (rank : 8) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UBZ9, O95941, Q9C0J4, Q9NUP2 | Gene names | REV1L, REV1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein REV1 (EC 2.7.7.-) (Rev1-like terminal deoxycytidyl transferase) (Alpha integrin-binding protein 80) (AIBP80). | |||||
|
ATP5S_MOUSE
|
||||||
| NC score | 0.074643 (rank : 11) | θ value | 1.06291 (rank : 26) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9CRA7, Q8R2M3 | Gene names | Atp5s, Atpw | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP synthase subunit s, mitochondrial precursor (ATP synthase coupling factor B) (Mitochondrial ATP synthase regulatory component factor B). | |||||
|
ARHGA_MOUSE
|
||||||
| NC score | 0.057992 (rank : 12) | θ value | θ > 10 (rank : 84) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8C033, Q5DU38, Q80VH8, Q8BW76, Q922S7 | Gene names | Arhgef10, Kiaa0294 | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 10. | |||||
|
VCIP1_HUMAN
|
||||||
| NC score | 0.056330 (rank : 13) | θ value | 0.21417 (rank : 15) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96JH7, Q504T4, Q86T93, Q86W01, Q8N3A9, Q9H5R8 | Gene names | VCPIP1, KIAA1850, VCIP135 | |||
|
Domain Architecture |

|
|||||
| Description | Deubiquitinating protein VCIP135 (EC 3.4.22.-) (Valosin-containing protein p97/p47 complex-interacting protein p135) (Valosin-containing protein p97/p47 complex-interacting protein 1). | |||||
|
ARHGA_HUMAN
|
||||||
| NC score | 0.055562 (rank : 14) | θ value | θ > 10 (rank : 83) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O15013, O14665, Q68D55, Q8IWD9, Q8IY77 | Gene names | ARHGEF10, KIAA0294 | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 10. | |||||
|
THOC2_HUMAN
|
||||||
| NC score | 0.044571 (rank : 15) | θ value | 0.813845 (rank : 25) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8NI27, Q9H8I6 | Gene names | THOC2, CXorf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | THO complex subunit 2 (Tho2). | |||||
|
RAD9B_HUMAN
|
||||||
| NC score | 0.044128 (rank : 16) | θ value | 1.06291 (rank : 30) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 5 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6WBX8, Q5U5K0, Q6NVJ1, Q6ZVT7, Q8N7T9, Q96LI8 | Gene names | RAD9B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell cycle checkpoint control protein RAD9B homolog (RAD9 homolog B) (hRAD9B). | |||||
|
PARP1_MOUSE
|
||||||
| NC score | 0.043679 (rank : 17) | θ value | 0.47712 (rank : 20) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P11103, Q9JLX4, Q9QVQ3 | Gene names | Parp1, Adprp, Adprt, Adprt1 | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 1 (EC 2.4.2.30) (PARP-1) (ADPRT) (NAD(+) ADP-ribosyltransferase 1) (Poly[ADP-ribose] synthetase 1) (msPARP). | |||||
|
SMBT1_MOUSE
|
||||||
| NC score | 0.043064 (rank : 18) | θ value | 0.279714 (rank : 18) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9JMD1, Q6NZD3, Q8CFS1 | Gene names | Sfmbt1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Scm-like with four MBT domains protein 1. | |||||
|
SMBT1_HUMAN
|
||||||
| NC score | 0.042997 (rank : 19) | θ value | 0.279714 (rank : 17) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UHJ3, Q96C73, Q9Y4Q9 | Gene names | SFMBT1, RU1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Scm-like with four MBT domains protein 1 (Renal ubiquitous protein 1). | |||||
|
VCIP1_MOUSE
|
||||||
| NC score | 0.042906 (rank : 20) | θ value | 4.03905 (rank : 55) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8CDG3, Q7TMU9, Q8BP90 | Gene names | Vcpip1, Vcip135 | |||
|
Domain Architecture |

|
|||||
| Description | Deubiquitinating protein VCIP135 (EC 3.4.22.-) (Valosin-containing protein p97/p47 complex-interacting protein p135) (Valosin-containing protein p97/p47 complex-interacting protein 1). | |||||
|
BARD1_HUMAN
|
||||||
| NC score | 0.042053 (rank : 21) | θ value | 1.38821 (rank : 31) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q99728, O43574 | Gene names | BARD1 | |||
|
Domain Architecture |

|
|||||
| Description | BRCA1-associated RING domain protein 1 (BARD-1). | |||||
|
BARD1_MOUSE
|
||||||
| NC score | 0.041484 (rank : 22) | θ value | 0.0431538 (rank : 9) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 411 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O70445 | Gene names | Bard1 | |||
|
Domain Architecture |

|
|||||
| Description | BRCA1-associated RING domain protein 1 (BARD-1). | |||||
|
PARP1_HUMAN
|
||||||
| NC score | 0.039959 (rank : 23) | θ value | 1.38821 (rank : 32) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P09874, Q8IUZ9 | Gene names | PARP1, ADPRT, PPOL | |||
|
Domain Architecture |

|
|||||
| Description | Poly [ADP-ribose] polymerase 1 (EC 2.4.2.30) (PARP-1) (ADPRT) (NAD(+) ADP-ribosyltransferase 1) (Poly[ADP-ribose] synthetase 1). | |||||
|
CG1_HUMAN
|
||||||
| NC score | 0.038062 (rank : 24) | θ value | 1.81305 (rank : 34) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q13495 | Gene names | CXorf6, CG1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CG1 protein (F18). | |||||
|
MLL2_HUMAN
|
||||||
| NC score | 0.036554 (rank : 25) | θ value | 0.0736092 (rank : 12) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
RAD9A_HUMAN
|
||||||
| NC score | 0.035898 (rank : 26) | θ value | 4.03905 (rank : 54) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99638, Q6FI29, Q96C41 | Gene names | RAD9A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell cycle checkpoint control protein RAD9A (EC 3.1.11.2) (DNA repair exonuclease rad9 homolog A) (hRAD9). | |||||
|
FBX38_MOUSE
|
||||||
| NC score | 0.035143 (rank : 27) | θ value | 0.813845 (rank : 24) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BMI0, Q8BTQ4, Q8C0H4 | Gene names | Fbxo38 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 38 (Modulator of KLF7 activity) (MoKA). | |||||
|
RAD9A_MOUSE
|
||||||
| NC score | 0.034365 (rank : 28) | θ value | 5.27518 (rank : 58) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Z0F6, Q8QZZ3 | Gene names | Rad9a, Rad9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell cycle checkpoint control protein RAD9A (EC 3.1.11.2) (DNA repair exonuclease rad9 homolog A) (Rad9-like protein) (mRAD9). | |||||
|
PCLO_MOUSE
|
||||||
| NC score | 0.031415 (rank : 29) | θ value | 0.163984 (rank : 13) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
MYC_HUMAN
|
||||||
| NC score | 0.031265 (rank : 30) | θ value | 0.62314 (rank : 23) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 146 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P01106, P01107, Q14026 | Gene names | MYC | |||
|
Domain Architecture |

|
|||||
| Description | Myc proto-oncogene protein (c-Myc) (Transcription factor p64). | |||||
|
PARG_MOUSE
|
||||||
| NC score | 0.030429 (rank : 31) | θ value | 1.81305 (rank : 36) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O88622, Q80YQ6, Q8CB72 | Gene names | Parg | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Poly(ADP-ribose) glycohydrolase (EC 3.2.1.143). | |||||
|
TAF3_HUMAN
|
||||||
| NC score | 0.030097 (rank : 32) | θ value | 1.81305 (rank : 37) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 568 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q5VWG9, Q6GMS5, Q6P6B5, Q86VY6, Q9BQS9, Q9UFI8 | Gene names | TAF3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID subunit 3 (TBP-associated factor 3) (Transcription initiation factor TFIID 140 kDa subunit) (140 kDa TATA box-binding protein-associated factor) (TAF140) (TAFII140). | |||||
|
PCLO_HUMAN
|
||||||
| NC score | 0.030011 (rank : 33) | θ value | 1.06291 (rank : 29) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
BRCA1_HUMAN
|
||||||
| NC score | 0.029296 (rank : 34) | θ value | 2.36792 (rank : 40) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 381 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P38398 | Gene names | BRCA1, RNF53 | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer type 1 susceptibility protein (RING finger protein 53). | |||||
|
CE152_HUMAN
|
||||||
| NC score | 0.026676 (rank : 35) | θ value | 0.21417 (rank : 14) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 976 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O94986, Q6NTA0 | Gene names | CEP152, KIAA0912 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 152 kDa (Cep152 protein). | |||||
|
DMN_HUMAN
|
||||||
| NC score | 0.026560 (rank : 36) | θ value | 0.365318 (rank : 19) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | O15061 | Gene names | DMN, KIAA0353 | |||
|
Domain Architecture |

|
|||||
| Description | Desmuslin. | |||||
|
MINT_HUMAN
|
||||||
| NC score | 0.025649 (rank : 37) | θ value | 0.62314 (rank : 22) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1291 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q96T58, Q9H9A8, Q9NWH5, Q9UQ01, Q9Y556 | Gene names | SPEN, KIAA0929, MINT, SHARP | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
CO027_HUMAN
|
||||||
| NC score | 0.025482 (rank : 38) | θ value | 4.03905 (rank : 49) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q2M3C6, Q8N993, Q96LL5 | Gene names | C15orf27 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C15orf27. | |||||
|
DTNA_MOUSE
|
||||||
| NC score | 0.024301 (rank : 39) | θ value | 2.36792 (rank : 41) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 264 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9D2N4, P97319, Q61498, Q61499, Q9QZZ5, Q9WUL9, Q9WUM0 | Gene names | Dtna, Dtn | |||
|
Domain Architecture |

|
|||||
| Description | Dystrobrevin alpha (Alpha-dystrobrevin). | |||||
|
SMC2_HUMAN
|
||||||
| NC score | 0.022803 (rank : 40) | θ value | 2.36792 (rank : 43) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1405 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O95347, Q9P1P2 | Gene names | SMC2L1, CAPE, SMC2 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 2-like 1 protein (Chromosome- associated protein E) (hCAP-E) (XCAP-E homolog). | |||||
|
ATX2_HUMAN
|
||||||
| NC score | 0.022045 (rank : 41) | θ value | 2.36792 (rank : 39) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 375 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q99700, Q6ZQZ7, Q99493 | Gene names | ATXN2, ATX2, SCA2, TNRC13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-2 (Spinocerebellar ataxia type 2 protein) (Trinucleotide repeat-containing gene 13 protein). | |||||
|
OFD1_HUMAN
|
||||||
| NC score | 0.021740 (rank : 42) | θ value | 2.36792 (rank : 42) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 753 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | O75665, O75666 | Gene names | OFD1, CXorf5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Oral-facial-digital syndrome 1 protein (Protein 71-7A). | |||||
|
BRD3_MOUSE
|
||||||
| NC score | 0.021424 (rank : 43) | θ value | 1.06291 (rank : 27) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 402 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8K2F0, Q8C665, Q8CAX7, Q9JI25 | Gene names | Brd3, Fsrg2 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 3 (Bromodomain-containing FSH-like protein FSRG2). | |||||
|
MILK2_HUMAN
|
||||||
| NC score | 0.021421 (rank : 44) | θ value | 1.81305 (rank : 35) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 457 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8IY33, Q7RTP4, Q7Z655, Q8TEQ4, Q9H5F9 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | MICAL-like protein 2. | |||||
|
CCNB3_MOUSE
|
||||||
| NC score | 0.020368 (rank : 45) | θ value | 1.81305 (rank : 33) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q810T2, Q810T3, Q8VDC8 | Gene names | Ccnb3, Cycb3 | |||
|
Domain Architecture |

|
|||||
| Description | G2/mitotic-specific cyclin-B3. | |||||
|
BPA1_MOUSE
|
||||||
| NC score | 0.019781 (rank : 46) | θ value | 6.88961 (rank : 63) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1477 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q91ZU6, Q91ZU7 | Gene names | Dst, Bpag1 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoforms 1/2/3/4 (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
GP158_MOUSE
|
||||||
| NC score | 0.019406 (rank : 47) | θ value | 3.0926 (rank : 45) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8C419, Q8CHB0 | Gene names | Gpr158, Kiaa1136 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 158 precursor. | |||||
|
DSPP_MOUSE
|
||||||
| NC score | 0.018468 (rank : 48) | θ value | 4.03905 (rank : 50) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P97399, O70567 | Gene names | Dspp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor (Dentin matrix protein 3) (DMP-3) [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
CTDP1_HUMAN
|
||||||
| NC score | 0.018355 (rank : 49) | θ value | 6.88961 (rank : 64) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 416 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Y5B0, Q7Z644, Q96BZ1, Q9Y6F5 | Gene names | CTDP1, FCP1 | |||
|
Domain Architecture |

|
|||||
| Description | RNA polymerase II subunit A C-terminal domain phosphatase (EC 3.1.3.16) (TFIIF-associating CTD phosphatase). | |||||
|
ELK4_MOUSE
|
||||||
| NC score | 0.018023 (rank : 50) | θ value | 1.06291 (rank : 28) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 107 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P41158 | Gene names | Elk4, Sap1 | |||
|
Domain Architecture |

|
|||||
| Description | ETS domain-containing protein Elk-4 (Serum response factor accessory protein 1) (SAP-1). | |||||
|
DMD_HUMAN
|
||||||
| NC score | 0.017582 (rank : 51) | θ value | 5.27518 (rank : 56) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1135 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P11532, Q02295, Q14169, Q14170, Q5JYU0, Q7KZ48 | Gene names | DMD | |||
|
Domain Architecture |

|
|||||
| Description | Dystrophin. | |||||
|
DMD_MOUSE
|
||||||
| NC score | 0.017382 (rank : 52) | θ value | 5.27518 (rank : 57) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1111 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P11531, O35653, Q60703 | Gene names | Dmd | |||
|
Domain Architecture |

|
|||||
| Description | Dystrophin. | |||||
|
TARA_MOUSE
|
||||||
| NC score | 0.017098 (rank : 53) | θ value | 5.27518 (rank : 60) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1133 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q99KW3, Q2PZW9, Q2Q402, Q2Q403, Q2Q404, Q6ZPK4, Q8C6T3, Q8CG90 | Gene names | Triobp, Kiaa1662, Tara | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TRIO and F-actin-binding protein (Protein Tara) (Trio-associated repeat on actin). | |||||
|
TXLNB_HUMAN
|
||||||
| NC score | 0.016457 (rank : 54) | θ value | 3.0926 (rank : 48) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 912 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8N3L3, Q76L25, Q86T52, Q8N3S2 | Gene names | TXLNB, C6orf198, MDP77 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-taxilin (Muscle-derived protein 77) (hMDP77). | |||||
|
TAF1L_HUMAN
|
||||||
| NC score | 0.015734 (rank : 55) | θ value | 5.27518 (rank : 59) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8IZX4 | Gene names | TAF1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID 210 kDa subunit (TBP-associated factor 210 kDa) (TAF(II)210) (TBP-associated factor 1-like). | |||||
|
K0319_HUMAN
|
||||||
| NC score | 0.015595 (rank : 56) | θ value | 6.88961 (rank : 67) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q5VV43, Q9UJC8, Q9Y4G7 | Gene names | KIAA0319 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0319 precursor. | |||||
|
ZC11A_MOUSE
|
||||||
| NC score | 0.015120 (rank : 57) | θ value | 8.99809 (rank : 82) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6NZF1, Q6NXW9, Q6PDR6, Q80TU7, Q8C5L5, Q99JN6 | Gene names | Zc3h11a, Kiaa0663 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CCCH domain-containing protein 11A. | |||||
|
FYCO1_MOUSE
|
||||||
| NC score | 0.015078 (rank : 58) | θ value | 3.0926 (rank : 44) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1059 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8VDC1, Q3V385, Q7TMT0, Q8BJN2 | Gene names | Fyco1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE and coiled-coil domain-containing protein 1. | |||||
|
LAMB2_MOUSE
|
||||||
| NC score | 0.014741 (rank : 59) | θ value | 0.279714 (rank : 16) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 722 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q61292, Q62182 | Gene names | Lamb2, Lams | |||
|
Domain Architecture |

|
|||||
| Description | Laminin beta-2 chain precursor (S-laminin) (S-LAM). | |||||
|
CLMN_HUMAN
|
||||||
| NC score | 0.013862 (rank : 60) | θ value | 8.99809 (rank : 71) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q96JQ2, Q9H713, Q9HA23, Q9HA57, Q9UFP4, Q9ULN2 | Gene names | CLMN, KIAA1188 | |||
|
Domain Architecture |

|
|||||
| Description | Calmin. | |||||
|
GON4L_MOUSE
|
||||||
| NC score | 0.013320 (rank : 61) | θ value | 8.99809 (rank : 74) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9DB00, Q80TB4, Q91YI9 | Gene names | Gon4l, Gon4, KIAA1606 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GON-4-like protein (GON-4 homolog). | |||||
|
SCG2_MOUSE
|
||||||
| NC score | 0.012654 (rank : 62) | θ value | 8.99809 (rank : 79) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q03517, Q80Y79, Q9CW80 | Gene names | Scg2, Chgc, Scg-2 | |||
|
Domain Architecture |

|
|||||
| Description | Secretogranin-2 precursor (Secretogranin II) (SgII) (Chromogranin C) [Contains: Secretoneurin (SN)]. | |||||
|
KINH_HUMAN
|
||||||
| NC score | 0.012495 (rank : 63) | θ value | 4.03905 (rank : 51) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1231 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P33176, Q5VZ85 | Gene names | KIF5B, KNS, KNS1 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin heavy chain (Ubiquitous kinesin heavy chain) (UKHC). | |||||
|
KINH_MOUSE
|
||||||
| NC score | 0.012436 (rank : 64) | θ value | 4.03905 (rank : 52) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1082 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q61768, O08711, Q61580 | Gene names | Kif5b, Khcs, Kns1 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin heavy chain (Ubiquitous kinesin heavy chain) (UKHC). | |||||
|
SPTA1_MOUSE
|
||||||
| NC score | 0.012287 (rank : 65) | θ value | 3.0926 (rank : 47) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 675 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P08032, P97502 | Gene names | Spta1, Spna1, Spta | |||
|
Domain Architecture |

|
|||||
| Description | Spectrin alpha chain, erythrocyte (Erythroid alpha-spectrin). | |||||
|
RRBP1_MOUSE
|
||||||
| NC score | 0.011896 (rank : 66) | θ value | 8.99809 (rank : 78) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1650 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q99PL5, Q99PK5, Q99PK6, Q99PK7, Q99PK8, Q99PK9, Q99PL0, Q99PL1, Q99PL2, Q99PL3, Q99PL4, Q9CS20 | Gene names | Rrbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (mRRp). | |||||
|
FIGLA_MOUSE
|
||||||
| NC score | 0.011799 (rank : 67) | θ value | 8.99809 (rank : 73) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O55208 | Gene names | Figla | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Factor in the germline alpha (Transcription factor FIGa) (FIGalpha). | |||||
|
SETX_HUMAN
|
||||||
| NC score | 0.011262 (rank : 68) | θ value | 8.99809 (rank : 80) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q7Z333, O75120, Q3KQX4, Q68DW5, Q6AZD7, Q7Z3J6, Q8WX33, Q9H9D1, Q9NVP9 | Gene names | SETX, ALS4, KIAA0625, SCAR1 | |||
|
Domain Architecture |

|
|||||
| Description | Probable helicase senataxin (EC 3.6.1.-) (SEN1 homolog). | |||||
|
ERC2_MOUSE
|
||||||
| NC score | 0.011044 (rank : 69) | θ value | 8.99809 (rank : 72) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1166 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q6PH08, Q80U20, Q8CCP1 | Gene names | Erc2, Cast1, D14Ertd171e, Kiaa0378 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ERC protein 2 (CAZ-associated structural protein 1) (CAST1). | |||||
|
TDT_HUMAN
|
||||||
| NC score | 0.009863 (rank : 70) | θ value | 8.99809 (rank : 81) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P04053, Q96E50 | Gene names | DNTT, TDT | |||
|
Domain Architecture |

|
|||||
| Description | DNA nucleotidylexotransferase (EC 2.7.7.31) (Terminal addition enzyme) (Terminal deoxynucleotidyltransferase) (Terminal transferase). | |||||
|
RAI14_HUMAN
|
||||||
| NC score | 0.009781 (rank : 71) | θ value | 6.88961 (rank : 69) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1367 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9P0K7, Q6V1W9, Q7Z5I4, Q7Z733, Q9P2L2, Q9Y3T5 | Gene names | RAI14, KIAA1334, NORPEG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankycorbin (Ankyrin repeat and coiled-coil structure-containing protein) (Retinoic acid-induced protein 14) (Novel retinal pigment epithelial cell protein). | |||||
|
ARMX5_HUMAN
|
||||||
| NC score | 0.009328 (rank : 72) | θ value | 8.99809 (rank : 70) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6P1M9, Q68DB4, Q9BVZ3, Q9H969 | Gene names | ARMCX5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Armadillo repeat-containing X-linked protein 5. | |||||
|
ITAD_MOUSE
|
||||||
| NC score | 0.008544 (rank : 73) | θ value | 3.0926 (rank : 46) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 85 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q3V0T4 | Gene names | Itgad | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Integrin alpha-D precursor (CD11d antigen). | |||||
|
NFH_HUMAN
|
||||||
| NC score | 0.008474 (rank : 74) | θ value | 8.99809 (rank : 77) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 1242 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | P12036, Q9UJS7, Q9UQ14 | Gene names | NEFH, KIAA0845, NFH | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
MAGB6_HUMAN
|
||||||
| NC score | 0.008123 (rank : 75) | θ value | 6.88961 (rank : 68) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8N7X4, Q6GS19, Q9H219 | Gene names | MAGEB6 | |||
|
Domain Architecture |

|
|||||
| Description | Melanoma-associated antigen B6 (MAGE-B6 antigen). | |||||
|
KI18A_MOUSE
|
||||||
| NC score | 0.007833 (rank : 76) | θ value | 8.99809 (rank : 75) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q91WD7, Q3TP77, Q8BLL1 | Gene names | Kif18a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin family member 18A. | |||||
|
FIGN_HUMAN
|
||||||
| NC score | 0.007725 (rank : 77) | θ value | 6.88961 (rank : 66) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5HY92, Q9H6M5, Q9NVZ9 | Gene names | FIGN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fidgetin. | |||||
|
PTPRZ_HUMAN
|
||||||
| NC score | 0.006220 (rank : 78) | θ value | 4.03905 (rank : 53) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P23471 | Gene names | PTPRZ1, PTPRZ, PTPZ | |||
|
Domain Architecture |

|
|||||
| Description | Receptor-type tyrosine-protein phosphatase zeta precursor (EC 3.1.3.48) (R-PTP-zeta). | |||||
|
DDX51_HUMAN
|
||||||
| NC score | 0.005913 (rank : 79) | θ value | 6.88961 (rank : 65) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8N8A6, Q5CZ71, Q8IXK5, Q96ED1 | Gene names | DDX51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent RNA helicase DDX51 (EC 3.6.1.-) (DEAD box protein 51). | |||||
|
LRP12_HUMAN
|
||||||
| NC score | 0.004641 (rank : 80) | θ value | 8.99809 (rank : 76) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y561 | Gene names | LRP12, ST7 | |||
|
Domain Architecture |

|
|||||
| Description | Low-density lipoprotein receptor-related protein 12 precursor (Suppressor of tumorigenicity protein 7). | |||||
|
CDKL2_MOUSE
|
||||||
| NC score | 0.003394 (rank : 81) | θ value | 0.62314 (rank : 21) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 854 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9QUK0, Q3UL60, Q3V3X7, Q9QYI1, Q9QYI2 | Gene names | Cdkl2, Kkiamre, Kkm | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-dependent kinase-like 2 (EC 2.7.11.22) (Serine/threonine- protein kinase KKIAMRE). | |||||
|
ZN281_HUMAN
|
||||||
| NC score | 0.001625 (rank : 82) | θ value | 5.27518 (rank : 62) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 745 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Y2X9, Q9NY92 | Gene names | ZNF281, GZP1, ZBP99 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 281 (Zinc finger DNA-binding protein 99) (Transcription factor ZBP-99) (GC-box-binding zinc finger protein 1). | |||||
|
ZBT20_HUMAN
|
||||||
| NC score | 0.001096 (rank : 83) | θ value | 5.27518 (rank : 61) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 835 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9HC78 | Gene names | ZBTB20, DPZF, ZNF288 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and BTB domain-containing protein 20 (Zinc finger protein 288) (Dendritic-derived BTB/POZ zinc finger protein). | |||||
|
ZFP59_MOUSE
|
||||||
| NC score | 0.000708 (rank : 84) | θ value | 1.81305 (rank : 38) | |||
| Query Neighborhood Hits | 82 | Target Neighborhood Hits | 768 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P16373, Q61849 | Gene names | Zfp59, Mfg2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 59 (Zfp-59) (Zinc finger protein Mfg-2). | |||||