Please be patient as the page loads
|
SGOL1_MOUSE
|
||||||
| SwissProt Accessions | Q9CXH7, Q3U4K4, Q588H1, Q8BKW2 | Gene names | Sgol1, Sgo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Shugoshin-like 1. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
SGOL1_MOUSE
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9CXH7, Q3U4K4, Q588H1, Q8BKW2 | Gene names | Sgol1, Sgo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Shugoshin-like 1. | |||||
|
SGOL1_HUMAN
|
||||||
| θ value | 1.37783e-141 (rank : 2) | NC score | 0.926816 (rank : 2) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q5FBB7, Q588H5, Q5FBB4, Q5FBB5, Q5FBB6, Q5FBB8, Q8N579, Q8WVL0, Q9BVA8, Q9H275 | Gene names | SGOL1, SGO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Shugoshin-like 1 (hSgo1) (Serologically defined breast cancer antigen NY-BR-85). | |||||
|
FLIP1_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 3) | NC score | 0.059791 (rank : 8) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1405 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q7Z7B0, Q5VUL6, Q86TC3, Q8N8B9, Q96SK6, Q9NVI8, Q9ULE5 | Gene names | FILIP1, KIAA1275 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filamin-A-interacting protein 1 (FILIP). | |||||
|
FLIP1_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 4) | NC score | 0.064667 (rank : 4) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1176 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9CS72 | Gene names | Filip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filamin-A-interacting protein 1 (FILIP). | |||||
|
SRRM1_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 5) | NC score | 0.070768 (rank : 3) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
SRRM2_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 6) | NC score | 0.064531 (rank : 5) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
PSIP1_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 7) | NC score | 0.062896 (rank : 7) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O75475, O00256, O95368, Q6P391, Q86YB9, Q9NZI3, Q9UER6 | Gene names | PSIP1, DFS70, LEDGF, PSIP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PC4 and SFRS1-interacting protein (Lens epithelium-derived growth factor) (Transcriptional coactivator p75/p52) (Dense fine speckles 70 kDa protein) (DFS 70) (CLL-associated antigen KW-7). | |||||
|
ERC2_HUMAN
|
||||||
| θ value | 0.279714 (rank : 8) | NC score | 0.048419 (rank : 11) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1156 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O15083, Q86TK4 | Gene names | ERC2, KIAA0378 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ERC protein 2. | |||||
|
ERC2_MOUSE
|
||||||
| θ value | 0.279714 (rank : 9) | NC score | 0.048160 (rank : 13) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1166 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q6PH08, Q80U20, Q8CCP1 | Gene names | Erc2, Cast1, D14Ertd171e, Kiaa0378 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ERC protein 2 (CAZ-associated structural protein 1) (CAST1). | |||||
|
SRRM1_MOUSE
|
||||||
| θ value | 0.279714 (rank : 10) | NC score | 0.064500 (rank : 6) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
ZF106_HUMAN
|
||||||
| θ value | 0.279714 (rank : 11) | NC score | 0.041791 (rank : 17) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 470 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9H2Y7, Q6NSD9, Q6PEK1, Q86T43, Q86T45, Q86T50, Q86T58, Q86TA9, Q96M37, Q9H7B8 | Gene names | ZFP106, ZNF474 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 106 homolog (Zfp-106) (Zinc finger protein 474). | |||||
|
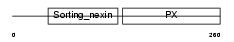
SNX2_HUMAN
|
||||||
| θ value | 0.365318 (rank : 12) | NC score | 0.044528 (rank : 15) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O60749, O43650, P82862, Q53XK8, Q597H6, Q9BTS8 | Gene names | SNX2, TRG9 | |||
|
Domain Architecture |
|
|||||
| Description | Sorting nexin-2 (Transformation-related gene 9 protein) (TRG-9). | |||||
|
SRGP2_HUMAN
|
||||||
| θ value | 0.47712 (rank : 13) | NC score | 0.040708 (rank : 18) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 331 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O43295, Q8IX13, Q8IZV8 | Gene names | SRGAP3, ARHGAP14, KIAA0411, MEGAP, SRGAP2 | |||
|
Domain Architecture |

|
|||||
| Description | SLIT-ROBO Rho GTPase-activating protein 3 (srGAP3) (srGAP2) (WAVE- associated Rac GTPase-activating protein) (WRP) (Mental disorder- associated GAP) (Rho-GTPase-activating protein 14). | |||||
|
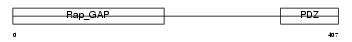
SIPA1_MOUSE
|
||||||
| θ value | 0.62314 (rank : 14) | NC score | 0.039454 (rank : 24) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P46062, P70204 | Gene names | Sipa1, Spa-1, Spa1 | |||
|
Domain Architecture |
|
|||||
| Description | Signal-induced proliferation-associated protein 1 (Sipa-1) (GTPase- activating protein Spa-1). | |||||
|
SRGP2_MOUSE
|
||||||
| θ value | 0.62314 (rank : 15) | NC score | 0.040169 (rank : 21) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 324 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q812A2, Q80U09, Q8BKP4, Q8BLD0, Q925I2 | Gene names | Srgap3, Arhgap14, Kiaa0411, Srgap2 | |||
|
Domain Architecture |

|
|||||
| Description | SLIT-ROBO Rho GTPase-activating protein 3 (srGAP3) (srGAP2) (WAVE- associated Rac GTPase-activating protein) (WRP) (Rho-GTPase-activating protein 14). | |||||
|
MAP1B_MOUSE
|
||||||
| θ value | 0.813845 (rank : 16) | NC score | 0.043498 (rank : 16) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 977 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P14873 | Gene names | Map1b, Mtap1b, Mtap5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) (MAP1.2) (MAP1(X)) [Contains: MAP1 light chain LC1]. | |||||
|
NEST_HUMAN
|
||||||
| θ value | 0.813845 (rank : 17) | NC score | 0.039380 (rank : 25) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1305 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P48681, O00552 | Gene names | NES | |||
|
Domain Architecture |

|
|||||
| Description | Nestin. | |||||
|
CCD45_HUMAN
|
||||||
| θ value | 1.06291 (rank : 18) | NC score | 0.048363 (rank : 12) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 525 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q96GE4, Q96M81 | Gene names | CCDC45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 45. | |||||
|
CENPC_MOUSE
|
||||||
| θ value | 1.81305 (rank : 19) | NC score | 0.039690 (rank : 23) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P49452 | Gene names | Cenpc1, Cenpc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein C 1 (CENP-C) (Centromere autoantigen C). | |||||
|
EEA1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 20) | NC score | 0.034163 (rank : 29) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1637 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8BL66, Q6DIC2 | Gene names | Eea1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Early endosome antigen 1. | |||||
|
GOGA4_MOUSE
|
||||||
| θ value | 1.81305 (rank : 21) | NC score | 0.034812 (rank : 27) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1939 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q91VW5, O70365, Q8CGH6 | Gene names | Golga4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (tGolgin-1). | |||||
|
KTN1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 22) | NC score | 0.039959 (rank : 22) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1324 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q61595, Q8BG49, Q8BHF4, Q8BHM8, Q8C9Y5, Q8CG51, Q8CG52, Q8CG53, Q8CG54, Q8CG55, Q8CG56, Q8CG57, Q8CG58, Q8CG59, Q8CG60, Q8CG61, Q8CG62, Q8CG63 | Gene names | Ktn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinectin. | |||||
|
RNPS1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 23) | NC score | 0.040323 (rank : 19) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q15287, O75308, Q8WY42, Q9NYG3 | Gene names | RNPS1, LDC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein with serine-rich domain 1 (SR-related protein LDC2). | |||||
|
RNPS1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 24) | NC score | 0.040323 (rank : 20) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q99M28, Q3TMJ1, Q922H8 | Gene names | Rnps1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein with serine-rich domain 1. | |||||
|
ST18_HUMAN
|
||||||
| θ value | 1.81305 (rank : 25) | NC score | 0.029407 (rank : 38) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 252 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O60284 | Gene names | ST18, KIAA0535, ZNF387 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppression of tumorigenicity protein 18 (Zinc finger protein 387). | |||||
|
ROCK2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 26) | NC score | 0.017675 (rank : 54) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 2072 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P70336, Q8BR64, Q8CBR0 | Gene names | Rock2 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 2 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 2) (p164 ROCK-2). | |||||
|
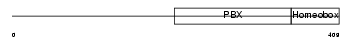
TFE2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 27) | NC score | 0.022767 (rank : 48) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P15923, P15883, Q14208, Q14635, Q14636, Q9UPI9 | Gene names | TCF3, E2A, ITF1 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor E2-alpha (Immunoglobulin enhancer-binding factor E12/E47) (Transcription factor 3) (TCF-3) (Immunoglobulin transcription factor 1) (Transcription factor ITF-1) (Kappa-E2-binding factor). | |||||
|
USP9X_MOUSE
|
||||||
| θ value | 2.36792 (rank : 28) | NC score | 0.025687 (rank : 45) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P70398, Q62497 | Gene names | Usp9x, Fafl, Fam | |||
|
Domain Architecture |

|
|||||
| Description | Probable ubiquitin carboxyl-terminal hydrolase FAF-X (EC 3.1.2.15) (Ubiquitin thioesterase FAF-X) (Ubiquitin-specific-processing protease FAF-X) (Deubiquitinating enzyme FAF-X) (Fat facets protein-related, X- linked) (Ubiquitin-specific protease 9, X chromosome) (Ubiquitin carboxyl-terminal hydrolase FAM) (Fat facets homolog). | |||||
|
AP3B1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 29) | NC score | 0.020048 (rank : 51) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O00203, O00580, Q7Z393, Q9HD66 | Gene names | AP3B1, ADTB3A | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit beta-1 (Adapter-related protein complex 3 beta-1 subunit) (Beta3A-adaptin) (Adaptor protein complex AP-3 beta-1 subunit) (Clathrin assembly protein complex 3 beta-1 large chain). | |||||
|
CLSPN_MOUSE
|
||||||
| θ value | 3.0926 (rank : 30) | NC score | 0.031200 (rank : 35) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q80YR7, Q69GM2 | Gene names | Clspn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Claspin. | |||||
|
CP135_MOUSE
|
||||||
| θ value | 3.0926 (rank : 31) | NC score | 0.034126 (rank : 30) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1278 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q6P5D4, Q6A030 | Gene names | Cep135, Cep4, Kiaa0635 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 135 kDa (Cep135 protein) (Centrosomal protein 4). | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 32) | NC score | 0.046617 (rank : 14) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
T53G5_HUMAN
|
||||||
| θ value | 3.0926 (rank : 33) | NC score | 0.049678 (rank : 10) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y2B4 | Gene names | TP53TG5, C20orf10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TP53-target gene 5 protein (TP53-inducible gene 5 protein). | |||||
|
TCF25_MOUSE
|
||||||
| θ value | 3.0926 (rank : 34) | NC score | 0.030979 (rank : 36) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8R3L2, Q3THR8, Q3UBI7, Q3UD64, Q3UG75, Q80ZX3, Q8BR80, Q8C200, Q8C2M3, Q8C6B4, Q9CUW0, Q9ER19 | Gene names | Tcf25, D8Ertd325e, Nulp1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 25 (Nuclear localized protein 1). | |||||
|
PREX1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 35) | NC score | 0.013586 (rank : 58) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8TCU6, Q5JS95, Q5JS96, Q69YL2, Q7Z2L9, Q9BQH0, Q9BX55, Q9H4Q6, Q9P2D2, Q9UGQ4 | Gene names | PREX1, KIAA1415 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 3,4,5-trisphosphate-dependent Rac exchanger 1 protein (P-Rex1 protein). | |||||
|
RBL2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 36) | NC score | 0.029058 (rank : 39) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q08999, Q15073, Q16084, Q92812 | Gene names | RBL2, RB2 | |||
|
Domain Architecture |

|
|||||
| Description | Retinoblastoma-like protein 2 (130 kDa retinoblastoma-associated protein) (PRB2) (P130) (RBR-2). | |||||
|
RBL2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 37) | NC score | 0.029576 (rank : 37) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q64700 | Gene names | Rbl2 | |||
|
Domain Architecture |

|
|||||
| Description | Retinoblastoma-like protein 2 (130 kDa retinoblastoma-associated protein) (PRB2) (P130) (RBR-2). | |||||
|
SIPA1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 38) | NC score | 0.035670 (rank : 26) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 250 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q96FS4, O14518, O60484, O60618 | Gene names | SIPA1, SPA1 | |||
|
Domain Architecture |

|
|||||
| Description | Signal-induced proliferation-associated protein 1 (Sipa-1) (GTPase- activating protein Spa-1) (p130 SPA-1). | |||||
|
USP9X_HUMAN
|
||||||
| θ value | 4.03905 (rank : 39) | NC score | 0.024045 (rank : 46) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q93008, O75550, Q8WWT3, Q8WX12 | Gene names | USP9X, DFFRX, USP9 | |||
|
Domain Architecture |

|
|||||
| Description | Probable ubiquitin carboxyl-terminal hydrolase FAF-X (EC 3.1.2.15) (Ubiquitin thioesterase FAF-X) (Ubiquitin-specific-processing protease FAF-X) (Deubiquitinating enzyme FAF-X) (Fat facets protein-related, X- linked) (Ubiquitin-specific protease 9, X chromosome). | |||||
|
ANR26_MOUSE
|
||||||
| θ value | 5.27518 (rank : 40) | NC score | 0.026104 (rank : 43) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1554 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q811D2, Q69ZS2, Q9CS61 | Gene names | Ankrd26, Kiaa1074 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 26. | |||||
|
BSN_MOUSE
|
||||||
| θ value | 5.27518 (rank : 41) | NC score | 0.026359 (rank : 42) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
DDFL1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 42) | NC score | 0.011534 (rank : 60) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 353 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8TDY4, Q6P9F4, Q86UY1, Q9NXK2 | Gene names | DDEFL1, UPLC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Development and differentiation-enhancing factor-like 1 (Protein up- regulated in liver cancer 1). | |||||
|
EVPL_MOUSE
|
||||||
| θ value | 5.27518 (rank : 43) | NC score | 0.031715 (rank : 34) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 952 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9D952 | Gene names | Evpl | |||
|
Domain Architecture |

|
|||||
| Description | Envoplakin (p210) (210 kDa cornified envelope precursor protein). | |||||
|
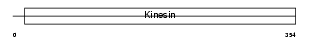
KIF11_HUMAN
|
||||||
| θ value | 5.27518 (rank : 44) | NC score | 0.019279 (rank : 52) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 583 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P52732, Q15716, Q5VWX0 | Gene names | KIF11, EG5, KNSL1 | |||
|
Domain Architecture |
|
|||||
| Description | Kinesin-like protein KIF11 (Kinesin-related motor protein Eg5) (Kinesin-like spindle protein HKSP) (Thyroid receptor-interacting protein 5) (TRIP5) (Kinesin-like protein 1). | |||||
|
MILK2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 45) | NC score | 0.016202 (rank : 55) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 457 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8IY33, Q7RTP4, Q7Z655, Q8TEQ4, Q9H5F9 | Gene names | ||||
|
Domain Architecture |
|
|||||
| Description | MICAL-like protein 2. | |||||
|
MY18A_HUMAN
|
||||||
| θ value | 5.27518 (rank : 46) | NC score | 0.033608 (rank : 31) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1929 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q92614, Q8IXP8 | Gene names | MYO18A, KIAA0216, MYSPDZ | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18A (Myosin XVIIIa) (Myosin containing PDZ domain) (Molecule associated with JAK3 N-terminus) (MAJN). | |||||
|
MYCB2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 47) | NC score | 0.027093 (rank : 41) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 390 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O75592, Q5JSX8, Q5VZN6, Q6PIB6, Q9UQ11, Q9Y6E4 | Gene names | MYCBP2, KIAA0916, PAM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ubiquitin ligase protein MYCBP2 (EC 6.3.2.-) (Myc-binding protein 2) (Protein associated with Myc) (Pam/highwire/rpm-1 protein). | |||||
|
R51A1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 48) | NC score | 0.031747 (rank : 33) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 164 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8C551, O55219, Q8BP36, Q99L94, Q9D0J0 | Gene names | Rad51ap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RAD51-associated protein 1 (RAD51-interacting protein) (RAB22). | |||||
|
ZAN_HUMAN
|
||||||
| θ value | 5.27518 (rank : 49) | NC score | 0.015751 (rank : 56) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 679 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y493, O00218, Q96L85, Q96L86, Q96L87, Q96L88, Q96L89, Q96L90, Q9BXN9, Q9BZ83, Q9BZ84, Q9BZ85, Q9BZ86, Q9BZ87, Q9BZ88 | Gene names | ZAN | |||
|
Domain Architecture |

|
|||||
| Description | Zonadhesin precursor. | |||||
|
DSPP_HUMAN
|
||||||
| θ value | 6.88961 (rank : 50) | NC score | 0.020926 (rank : 49) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9NZW4, O95815 | Gene names | DSPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
HSP7C_MOUSE
|
||||||
| θ value | 6.88961 (rank : 51) | NC score | 0.008734 (rank : 61) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P63017, P08109, P12225, Q62373, Q62374, Q62375 | Gene names | Hspa8, Hsc70, Hsc73 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heat shock cognate 71 kDa protein (Heat shock 70 kDa protein 8). | |||||
|
MYH7_HUMAN
|
||||||
| θ value | 6.88961 (rank : 52) | NC score | 0.020919 (rank : 50) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P12883, Q14836, Q14837, Q14904, Q16579, Q92679, Q9H1D5, Q9UMM8 | Gene names | MYH7, MYHCB | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle beta isoform (MyHC-beta). | |||||
|
NCKX3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 53) | NC score | 0.012408 (rank : 59) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99PD7, Q99JR2, Q99PD8 | Gene names | Slc24a3, Nckx3 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium/calcium exchanger 3 precursor (Na(+)/K(+)/Ca(2+)- exchange protein 3). | |||||
|
SALL2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 54) | NC score | 0.003596 (rank : 65) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 835 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Y467, Q9Y4G1 | Gene names | SALL2, KIAA0360, SAL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sal-like protein 2 (Zinc finger protein SALL2) (HSal2). | |||||
|
SNX2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 55) | NC score | 0.032355 (rank : 32) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9CWK8, Q3UKW3, Q9CQV0 | Gene names | Snx2 | |||
|
Domain Architecture |
|
|||||
| Description | Sorting nexin-2. | |||||
|
TITIN_HUMAN
|
||||||
| θ value | 6.88961 (rank : 56) | NC score | 0.006584 (rank : 62) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 2923 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8WZ42, Q10465, Q10466, Q15598, Q2XUS3, Q32Q60, Q4U1Z6, Q6NSG0, Q6PDB1, Q6PJP0, Q7KYM2, Q7KYN4, Q7KYN5, Q7LDM3, Q7Z2X3, Q8TCG8, Q8WZ51, Q8WZ52, Q8WZ53, Q8WZB3, Q92761, Q92762, Q9UD97, Q9UP84, Q9Y6L9 | Gene names | TTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Titin (EC 2.7.11.1) (Connectin) (Rhabdomyosarcoma antigen MU-RMS- 40.14). | |||||
|
WTAP_MOUSE
|
||||||
| θ value | 6.88961 (rank : 57) | NC score | 0.027568 (rank : 40) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 250 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9ER69 | Gene names | Wtap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wilms' tumor 1-associating protein (WT1-associated protein) (Putative pre-mRNA-splicing regulator female-lethal(2D) homolog). | |||||
|
ZKSC1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 58) | NC score | -0.000310 (rank : 66) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 755 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BGS3, Q7TS88, Q8BJ55, Q9CRN6 | Gene names | Zkscan1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger with KRAB and SCAN domain-containing protein 1. | |||||
|
CENPE_HUMAN
|
||||||
| θ value | 8.99809 (rank : 59) | NC score | 0.025873 (rank : 44) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 2060 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q02224 | Gene names | CENPE | |||
|
Domain Architecture |

|
|||||
| Description | Centromeric protein E (CENP-E). | |||||
|
IQGA1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 60) | NC score | 0.015479 (rank : 57) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 250 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P46940 | Gene names | IQGAP1, KIAA0051 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP1 (p195). | |||||
|
IWS1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 61) | NC score | 0.023895 (rank : 47) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
PRGC2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 62) | NC score | 0.019060 (rank : 53) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8VHJ7, Q8C1C0 | Gene names | Ppargc1b, Errl1, Pgc1, Pgc1b, Ppargc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor gamma coactivator 1-beta (PPAR gamma coactivator-1beta protein) (PGC-1beta) (ERR ligand 1) (PPARGC-1-beta) (PGC-1-beta). | |||||
|
SRGP1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 63) | NC score | 0.034517 (rank : 28) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 390 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q7Z6B7, Q9H8A3, Q9P2P2 | Gene names | SRGAP1, ARHGAP13, KIAA1304 | |||
|
Domain Architecture |

|
|||||
| Description | SLIT-ROBO Rho GTPase-activating protein 1 (srGAP1) (Rho-GTPase- activating protein 13). | |||||
|
TLK1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 64) | NC score | 0.005171 (rank : 63) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 967 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9UKI8, Q14150, Q8N591, Q9NYH2, Q9Y4F6 | Gene names | TLK1, KIAA0137 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase tousled-like 1 (EC 2.7.11.1) (Tousled- like kinase 1) (PKU-beta). | |||||
|
TLK1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 65) | NC score | 0.004889 (rank : 64) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8C0V0 | Gene names | Tlk1 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase tousled-like 1 (EC 2.7.11.1) (Tousled- like kinase 1). | |||||
|
TXND2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 66) | NC score | 0.050359 (rank : 9) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
SGOL1_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 65 | |
| SwissProt Accessions | Q9CXH7, Q3U4K4, Q588H1, Q8BKW2 | Gene names | Sgol1, Sgo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Shugoshin-like 1. | |||||
|
SGOL1_HUMAN
|
||||||
| NC score | 0.926816 (rank : 2) | θ value | 1.37783e-141 (rank : 2) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q5FBB7, Q588H5, Q5FBB4, Q5FBB5, Q5FBB6, Q5FBB8, Q8N579, Q8WVL0, Q9BVA8, Q9H275 | Gene names | SGOL1, SGO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Shugoshin-like 1 (hSgo1) (Serologically defined breast cancer antigen NY-BR-85). | |||||
|
SRRM1_HUMAN
|
||||||
| NC score | 0.070768 (rank : 3) | θ value | 0.0563607 (rank : 5) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 690 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8IYB3, O60585, Q5VVN4 | Gene names | SRRM1, SRM160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Ser/Arg-related nuclear matrix protein) (SR-related nuclear matrix protein of 160 kDa) (SRm160). | |||||
|
FLIP1_MOUSE
|
||||||
| NC score | 0.064667 (rank : 4) | θ value | 0.0330416 (rank : 4) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1176 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9CS72 | Gene names | Filip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filamin-A-interacting protein 1 (FILIP). | |||||
|
SRRM2_HUMAN
|
||||||
| NC score | 0.064531 (rank : 5) | θ value | 0.0563607 (rank : 6) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
SRRM1_MOUSE
|
||||||
| NC score | 0.064500 (rank : 6) | θ value | 0.279714 (rank : 10) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
PSIP1_HUMAN
|
||||||
| NC score | 0.062896 (rank : 7) | θ value | 0.0736092 (rank : 7) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O75475, O00256, O95368, Q6P391, Q86YB9, Q9NZI3, Q9UER6 | Gene names | PSIP1, DFS70, LEDGF, PSIP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | PC4 and SFRS1-interacting protein (Lens epithelium-derived growth factor) (Transcriptional coactivator p75/p52) (Dense fine speckles 70 kDa protein) (DFS 70) (CLL-associated antigen KW-7). | |||||
|
FLIP1_HUMAN
|
||||||
| NC score | 0.059791 (rank : 8) | θ value | 0.0193708 (rank : 3) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1405 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q7Z7B0, Q5VUL6, Q86TC3, Q8N8B9, Q96SK6, Q9NVI8, Q9ULE5 | Gene names | FILIP1, KIAA1275 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filamin-A-interacting protein 1 (FILIP). | |||||
|
TXND2_MOUSE
|
||||||
| NC score | 0.050359 (rank : 9) | θ value | θ > 10 (rank : 66) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
T53G5_HUMAN
|
||||||
| NC score | 0.049678 (rank : 10) | θ value | 3.0926 (rank : 33) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y2B4 | Gene names | TP53TG5, C20orf10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TP53-target gene 5 protein (TP53-inducible gene 5 protein). | |||||
|
ERC2_HUMAN
|
||||||
| NC score | 0.048419 (rank : 11) | θ value | 0.279714 (rank : 8) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1156 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O15083, Q86TK4 | Gene names | ERC2, KIAA0378 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ERC protein 2. | |||||
|
CCD45_HUMAN
|
||||||
| NC score | 0.048363 (rank : 12) | θ value | 1.06291 (rank : 18) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 525 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q96GE4, Q96M81 | Gene names | CCDC45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 45. | |||||
|
ERC2_MOUSE
|
||||||
| NC score | 0.048160 (rank : 13) | θ value | 0.279714 (rank : 9) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1166 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q6PH08, Q80U20, Q8CCP1 | Gene names | Erc2, Cast1, D14Ertd171e, Kiaa0378 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ERC protein 2 (CAZ-associated structural protein 1) (CAST1). | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.046617 (rank : 14) | θ value | 3.0926 (rank : 32) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
SNX2_HUMAN
|
||||||
| NC score | 0.044528 (rank : 15) | θ value | 0.365318 (rank : 12) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O60749, O43650, P82862, Q53XK8, Q597H6, Q9BTS8 | Gene names | SNX2, TRG9 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-2 (Transformation-related gene 9 protein) (TRG-9). | |||||
|
MAP1B_MOUSE
|
||||||
| NC score | 0.043498 (rank : 16) | θ value | 0.813845 (rank : 16) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 977 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P14873 | Gene names | Map1b, Mtap1b, Mtap5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) (MAP1.2) (MAP1(X)) [Contains: MAP1 light chain LC1]. | |||||
|
ZF106_HUMAN
|
||||||
| NC score | 0.041791 (rank : 17) | θ value | 0.279714 (rank : 11) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 470 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9H2Y7, Q6NSD9, Q6PEK1, Q86T43, Q86T45, Q86T50, Q86T58, Q86TA9, Q96M37, Q9H7B8 | Gene names | ZFP106, ZNF474 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 106 homolog (Zfp-106) (Zinc finger protein 474). | |||||
|
SRGP2_HUMAN
|
||||||
| NC score | 0.040708 (rank : 18) | θ value | 0.47712 (rank : 13) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 331 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O43295, Q8IX13, Q8IZV8 | Gene names | SRGAP3, ARHGAP14, KIAA0411, MEGAP, SRGAP2 | |||
|
Domain Architecture |

|
|||||
| Description | SLIT-ROBO Rho GTPase-activating protein 3 (srGAP3) (srGAP2) (WAVE- associated Rac GTPase-activating protein) (WRP) (Mental disorder- associated GAP) (Rho-GTPase-activating protein 14). | |||||
|
RNPS1_HUMAN
|
||||||
| NC score | 0.040323 (rank : 19) | θ value | 1.81305 (rank : 23) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q15287, O75308, Q8WY42, Q9NYG3 | Gene names | RNPS1, LDC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein with serine-rich domain 1 (SR-related protein LDC2). | |||||
|
RNPS1_MOUSE
|
||||||
| NC score | 0.040323 (rank : 20) | θ value | 1.81305 (rank : 24) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 285 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q99M28, Q3TMJ1, Q922H8 | Gene names | Rnps1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein with serine-rich domain 1. | |||||
|
SRGP2_MOUSE
|
||||||
| NC score | 0.040169 (rank : 21) | θ value | 0.62314 (rank : 15) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 324 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q812A2, Q80U09, Q8BKP4, Q8BLD0, Q925I2 | Gene names | Srgap3, Arhgap14, Kiaa0411, Srgap2 | |||
|
Domain Architecture |

|
|||||
| Description | SLIT-ROBO Rho GTPase-activating protein 3 (srGAP3) (srGAP2) (WAVE- associated Rac GTPase-activating protein) (WRP) (Rho-GTPase-activating protein 14). | |||||
|
KTN1_MOUSE
|
||||||
| NC score | 0.039959 (rank : 22) | θ value | 1.81305 (rank : 22) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1324 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q61595, Q8BG49, Q8BHF4, Q8BHM8, Q8C9Y5, Q8CG51, Q8CG52, Q8CG53, Q8CG54, Q8CG55, Q8CG56, Q8CG57, Q8CG58, Q8CG59, Q8CG60, Q8CG61, Q8CG62, Q8CG63 | Gene names | Ktn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinectin. | |||||
|
CENPC_MOUSE
|
||||||
| NC score | 0.039690 (rank : 23) | θ value | 1.81305 (rank : 19) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P49452 | Gene names | Cenpc1, Cenpc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein C 1 (CENP-C) (Centromere autoantigen C). | |||||
|
SIPA1_MOUSE
|
||||||
| NC score | 0.039454 (rank : 24) | θ value | 0.62314 (rank : 14) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P46062, P70204 | Gene names | Sipa1, Spa-1, Spa1 | |||
|
Domain Architecture |

|
|||||
| Description | Signal-induced proliferation-associated protein 1 (Sipa-1) (GTPase- activating protein Spa-1). | |||||
|
NEST_HUMAN
|
||||||
| NC score | 0.039380 (rank : 25) | θ value | 0.813845 (rank : 17) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1305 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P48681, O00552 | Gene names | NES | |||
|
Domain Architecture |

|
|||||
| Description | Nestin. | |||||
|
SIPA1_HUMAN
|
||||||
| NC score | 0.035670 (rank : 26) | θ value | 4.03905 (rank : 38) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 250 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q96FS4, O14518, O60484, O60618 | Gene names | SIPA1, SPA1 | |||
|
Domain Architecture |

|
|||||
| Description | Signal-induced proliferation-associated protein 1 (Sipa-1) (GTPase- activating protein Spa-1) (p130 SPA-1). | |||||
|
GOGA4_MOUSE
|
||||||
| NC score | 0.034812 (rank : 27) | θ value | 1.81305 (rank : 21) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1939 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q91VW5, O70365, Q8CGH6 | Gene names | Golga4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (tGolgin-1). | |||||
|
SRGP1_HUMAN
|
||||||
| NC score | 0.034517 (rank : 28) | θ value | 8.99809 (rank : 63) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 390 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q7Z6B7, Q9H8A3, Q9P2P2 | Gene names | SRGAP1, ARHGAP13, KIAA1304 | |||
|
Domain Architecture |

|
|||||
| Description | SLIT-ROBO Rho GTPase-activating protein 1 (srGAP1) (Rho-GTPase- activating protein 13). | |||||
|
EEA1_MOUSE
|
||||||
| NC score | 0.034163 (rank : 29) | θ value | 1.81305 (rank : 20) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1637 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q8BL66, Q6DIC2 | Gene names | Eea1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Early endosome antigen 1. | |||||
|
CP135_MOUSE
|
||||||
| NC score | 0.034126 (rank : 30) | θ value | 3.0926 (rank : 31) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1278 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q6P5D4, Q6A030 | Gene names | Cep135, Cep4, Kiaa0635 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein of 135 kDa (Cep135 protein) (Centrosomal protein 4). | |||||
|
MY18A_HUMAN
|
||||||
| NC score | 0.033608 (rank : 31) | θ value | 5.27518 (rank : 46) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1929 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q92614, Q8IXP8 | Gene names | MYO18A, KIAA0216, MYSPDZ | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18A (Myosin XVIIIa) (Myosin containing PDZ domain) (Molecule associated with JAK3 N-terminus) (MAJN). | |||||
|
SNX2_MOUSE
|
||||||
| NC score | 0.032355 (rank : 32) | θ value | 6.88961 (rank : 55) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9CWK8, Q3UKW3, Q9CQV0 | Gene names | Snx2 | |||
|
Domain Architecture |

|
|||||
| Description | Sorting nexin-2. | |||||
|
R51A1_MOUSE
|
||||||
| NC score | 0.031747 (rank : 33) | θ value | 5.27518 (rank : 48) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 164 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8C551, O55219, Q8BP36, Q99L94, Q9D0J0 | Gene names | Rad51ap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RAD51-associated protein 1 (RAD51-interacting protein) (RAB22). | |||||
|
EVPL_MOUSE
|
||||||
| NC score | 0.031715 (rank : 34) | θ value | 5.27518 (rank : 43) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 952 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9D952 | Gene names | Evpl | |||
|
Domain Architecture |

|
|||||
| Description | Envoplakin (p210) (210 kDa cornified envelope precursor protein). | |||||
|
CLSPN_MOUSE
|
||||||
| NC score | 0.031200 (rank : 35) | θ value | 3.0926 (rank : 30) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 292 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q80YR7, Q69GM2 | Gene names | Clspn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Claspin. | |||||
|
TCF25_MOUSE
|
||||||
| NC score | 0.030979 (rank : 36) | θ value | 3.0926 (rank : 34) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8R3L2, Q3THR8, Q3UBI7, Q3UD64, Q3UG75, Q80ZX3, Q8BR80, Q8C200, Q8C2M3, Q8C6B4, Q9CUW0, Q9ER19 | Gene names | Tcf25, D8Ertd325e, Nulp1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 25 (Nuclear localized protein 1). | |||||
|
RBL2_MOUSE
|
||||||
| NC score | 0.029576 (rank : 37) | θ value | 4.03905 (rank : 37) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q64700 | Gene names | Rbl2 | |||
|
Domain Architecture |

|
|||||
| Description | Retinoblastoma-like protein 2 (130 kDa retinoblastoma-associated protein) (PRB2) (P130) (RBR-2). | |||||
|
ST18_HUMAN
|
||||||
| NC score | 0.029407 (rank : 38) | θ value | 1.81305 (rank : 25) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 252 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O60284 | Gene names | ST18, KIAA0535, ZNF387 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Suppression of tumorigenicity protein 18 (Zinc finger protein 387). | |||||
|
RBL2_HUMAN
|
||||||
| NC score | 0.029058 (rank : 39) | θ value | 4.03905 (rank : 36) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q08999, Q15073, Q16084, Q92812 | Gene names | RBL2, RB2 | |||
|
Domain Architecture |

|
|||||
| Description | Retinoblastoma-like protein 2 (130 kDa retinoblastoma-associated protein) (PRB2) (P130) (RBR-2). | |||||
|
WTAP_MOUSE
|
||||||
| NC score | 0.027568 (rank : 40) | θ value | 6.88961 (rank : 57) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 250 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9ER69 | Gene names | Wtap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wilms' tumor 1-associating protein (WT1-associated protein) (Putative pre-mRNA-splicing regulator female-lethal(2D) homolog). | |||||
|
MYCB2_HUMAN
|
||||||
| NC score | 0.027093 (rank : 41) | θ value | 5.27518 (rank : 47) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 390 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O75592, Q5JSX8, Q5VZN6, Q6PIB6, Q9UQ11, Q9Y6E4 | Gene names | MYCBP2, KIAA0916, PAM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ubiquitin ligase protein MYCBP2 (EC 6.3.2.-) (Myc-binding protein 2) (Protein associated with Myc) (Pam/highwire/rpm-1 protein). | |||||
|
BSN_MOUSE
|
||||||
| NC score | 0.026359 (rank : 42) | θ value | 5.27518 (rank : 41) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
ANR26_MOUSE
|
||||||
| NC score | 0.026104 (rank : 43) | θ value | 5.27518 (rank : 40) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1554 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q811D2, Q69ZS2, Q9CS61 | Gene names | Ankrd26, Kiaa1074 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 26. | |||||
|
CENPE_HUMAN
|
||||||
| NC score | 0.025873 (rank : 44) | θ value | 8.99809 (rank : 59) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 2060 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q02224 | Gene names | CENPE | |||
|
Domain Architecture |

|
|||||
| Description | Centromeric protein E (CENP-E). | |||||
|
USP9X_MOUSE
|
||||||
| NC score | 0.025687 (rank : 45) | θ value | 2.36792 (rank : 28) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P70398, Q62497 | Gene names | Usp9x, Fafl, Fam | |||
|
Domain Architecture |

|
|||||
| Description | Probable ubiquitin carboxyl-terminal hydrolase FAF-X (EC 3.1.2.15) (Ubiquitin thioesterase FAF-X) (Ubiquitin-specific-processing protease FAF-X) (Deubiquitinating enzyme FAF-X) (Fat facets protein-related, X- linked) (Ubiquitin-specific protease 9, X chromosome) (Ubiquitin carboxyl-terminal hydrolase FAM) (Fat facets homolog). | |||||
|
USP9X_HUMAN
|
||||||
| NC score | 0.024045 (rank : 46) | θ value | 4.03905 (rank : 39) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q93008, O75550, Q8WWT3, Q8WX12 | Gene names | USP9X, DFFRX, USP9 | |||
|
Domain Architecture |

|
|||||
| Description | Probable ubiquitin carboxyl-terminal hydrolase FAF-X (EC 3.1.2.15) (Ubiquitin thioesterase FAF-X) (Ubiquitin-specific-processing protease FAF-X) (Deubiquitinating enzyme FAF-X) (Fat facets protein-related, X- linked) (Ubiquitin-specific protease 9, X chromosome). | |||||
|
IWS1_MOUSE
|
||||||
| NC score | 0.023895 (rank : 47) | θ value | 8.99809 (rank : 61) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
TFE2_HUMAN
|
||||||
| NC score | 0.022767 (rank : 48) | θ value | 2.36792 (rank : 27) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P15923, P15883, Q14208, Q14635, Q14636, Q9UPI9 | Gene names | TCF3, E2A, ITF1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor E2-alpha (Immunoglobulin enhancer-binding factor E12/E47) (Transcription factor 3) (TCF-3) (Immunoglobulin transcription factor 1) (Transcription factor ITF-1) (Kappa-E2-binding factor). | |||||
|
DSPP_HUMAN
|
||||||
| NC score | 0.020926 (rank : 49) | θ value | 6.88961 (rank : 50) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9NZW4, O95815 | Gene names | DSPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
MYH7_HUMAN
|
||||||
| NC score | 0.020919 (rank : 50) | θ value | 6.88961 (rank : 52) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P12883, Q14836, Q14837, Q14904, Q16579, Q92679, Q9H1D5, Q9UMM8 | Gene names | MYH7, MYHCB | |||
|
Domain Architecture |

|
|||||
| Description | Myosin heavy chain, cardiac muscle beta isoform (MyHC-beta). | |||||
|
AP3B1_HUMAN
|
||||||
| NC score | 0.020048 (rank : 51) | θ value | 3.0926 (rank : 29) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O00203, O00580, Q7Z393, Q9HD66 | Gene names | AP3B1, ADTB3A | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit beta-1 (Adapter-related protein complex 3 beta-1 subunit) (Beta3A-adaptin) (Adaptor protein complex AP-3 beta-1 subunit) (Clathrin assembly protein complex 3 beta-1 large chain). | |||||
|
KIF11_HUMAN
|
||||||
| NC score | 0.019279 (rank : 52) | θ value | 5.27518 (rank : 44) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 583 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P52732, Q15716, Q5VWX0 | Gene names | KIF11, EG5, KNSL1 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF11 (Kinesin-related motor protein Eg5) (Kinesin-like spindle protein HKSP) (Thyroid receptor-interacting protein 5) (TRIP5) (Kinesin-like protein 1). | |||||
|
PRGC2_MOUSE
|
||||||
| NC score | 0.019060 (rank : 53) | θ value | 8.99809 (rank : 62) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8VHJ7, Q8C1C0 | Gene names | Ppargc1b, Errl1, Pgc1, Pgc1b, Ppargc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Peroxisome proliferator-activated receptor gamma coactivator 1-beta (PPAR gamma coactivator-1beta protein) (PGC-1beta) (ERR ligand 1) (PPARGC-1-beta) (PGC-1-beta). | |||||
|
ROCK2_MOUSE
|
||||||
| NC score | 0.017675 (rank : 54) | θ value | 2.36792 (rank : 26) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 2072 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P70336, Q8BR64, Q8CBR0 | Gene names | Rock2 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 2 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 2) (p164 ROCK-2). | |||||
|
MILK2_HUMAN
|
||||||
| NC score | 0.016202 (rank : 55) | θ value | 5.27518 (rank : 45) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 457 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8IY33, Q7RTP4, Q7Z655, Q8TEQ4, Q9H5F9 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | MICAL-like protein 2. | |||||
|
ZAN_HUMAN
|
||||||
| NC score | 0.015751 (rank : 56) | θ value | 5.27518 (rank : 49) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 679 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y493, O00218, Q96L85, Q96L86, Q96L87, Q96L88, Q96L89, Q96L90, Q9BXN9, Q9BZ83, Q9BZ84, Q9BZ85, Q9BZ86, Q9BZ87, Q9BZ88 | Gene names | ZAN | |||
|
Domain Architecture |

|
|||||
| Description | Zonadhesin precursor. | |||||
|
IQGA1_HUMAN
|
||||||
| NC score | 0.015479 (rank : 57) | θ value | 8.99809 (rank : 60) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 250 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P46940 | Gene names | IQGAP1, KIAA0051 | |||
|
Domain Architecture |

|
|||||
| Description | Ras GTPase-activating-like protein IQGAP1 (p195). | |||||
|
PREX1_HUMAN
|
||||||
| NC score | 0.013586 (rank : 58) | θ value | 4.03905 (rank : 35) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8TCU6, Q5JS95, Q5JS96, Q69YL2, Q7Z2L9, Q9BQH0, Q9BX55, Q9H4Q6, Q9P2D2, Q9UGQ4 | Gene names | PREX1, KIAA1415 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol 3,4,5-trisphosphate-dependent Rac exchanger 1 protein (P-Rex1 protein). | |||||
|
NCKX3_MOUSE
|
||||||
| NC score | 0.012408 (rank : 59) | θ value | 6.88961 (rank : 53) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99PD7, Q99JR2, Q99PD8 | Gene names | Slc24a3, Nckx3 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium/calcium exchanger 3 precursor (Na(+)/K(+)/Ca(2+)- exchange protein 3). | |||||
|
DDFL1_HUMAN
|
||||||
| NC score | 0.011534 (rank : 60) | θ value | 5.27518 (rank : 42) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 353 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8TDY4, Q6P9F4, Q86UY1, Q9NXK2 | Gene names | DDEFL1, UPLC1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Development and differentiation-enhancing factor-like 1 (Protein up- regulated in liver cancer 1). | |||||
|
HSP7C_MOUSE
|
||||||
| NC score | 0.008734 (rank : 61) | θ value | 6.88961 (rank : 51) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 166 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P63017, P08109, P12225, Q62373, Q62374, Q62375 | Gene names | Hspa8, Hsc70, Hsc73 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heat shock cognate 71 kDa protein (Heat shock 70 kDa protein 8). | |||||
|
TITIN_HUMAN
|
||||||
| NC score | 0.006584 (rank : 62) | θ value | 6.88961 (rank : 56) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 2923 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8WZ42, Q10465, Q10466, Q15598, Q2XUS3, Q32Q60, Q4U1Z6, Q6NSG0, Q6PDB1, Q6PJP0, Q7KYM2, Q7KYN4, Q7KYN5, Q7LDM3, Q7Z2X3, Q8TCG8, Q8WZ51, Q8WZ52, Q8WZ53, Q8WZB3, Q92761, Q92762, Q9UD97, Q9UP84, Q9Y6L9 | Gene names | TTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Titin (EC 2.7.11.1) (Connectin) (Rhabdomyosarcoma antigen MU-RMS- 40.14). | |||||
|
TLK1_HUMAN
|
||||||
| NC score | 0.005171 (rank : 63) | θ value | 8.99809 (rank : 64) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 967 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9UKI8, Q14150, Q8N591, Q9NYH2, Q9Y4F6 | Gene names | TLK1, KIAA0137 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase tousled-like 1 (EC 2.7.11.1) (Tousled- like kinase 1) (PKU-beta). | |||||
|
TLK1_MOUSE
|
||||||
| NC score | 0.004889 (rank : 64) | θ value | 8.99809 (rank : 65) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 947 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8C0V0 | Gene names | Tlk1 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase tousled-like 1 (EC 2.7.11.1) (Tousled- like kinase 1). | |||||
|
SALL2_HUMAN
|
||||||
| NC score | 0.003596 (rank : 65) | θ value | 6.88961 (rank : 54) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 835 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Y467, Q9Y4G1 | Gene names | SALL2, KIAA0360, SAL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sal-like protein 2 (Zinc finger protein SALL2) (HSal2). | |||||
|
ZKSC1_MOUSE
|
||||||
| NC score | -0.000310 (rank : 66) | θ value | 6.88961 (rank : 58) | |||
| Query Neighborhood Hits | 65 | Target Neighborhood Hits | 755 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BGS3, Q7TS88, Q8BJ55, Q9CRN6 | Gene names | Zkscan1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger with KRAB and SCAN domain-containing protein 1. | |||||