Please be patient as the page loads
|
SARDH_MOUSE
|
||||||
| SwissProt Accessions | Q99LB7 | Gene names | Sardh | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcosine dehydrogenase, mitochondrial precursor (EC 1.5.99.1) (SarDH). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
SARDH_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.996500 (rank : 2) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9UL12, Q9Y280, Q9Y2Y3 | Gene names | SARDH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcosine dehydrogenase, mitochondrial precursor (EC 1.5.99.1) (SarDH) (BPR-2). | |||||
|
SARDH_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q99LB7 | Gene names | Sardh | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcosine dehydrogenase, mitochondrial precursor (EC 1.5.99.1) (SarDH). | |||||
|
M2GD_HUMAN
|
||||||
| θ value | 2.79562e-118 (rank : 3) | NC score | 0.916142 (rank : 3) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9UI17 | Gene names | DMGDH | |||
|
Domain Architecture |

|
|||||
| Description | Dimethylglycine dehydrogenase, mitochondrial precursor (EC 1.5.99.2) (ME2GLYDH). | |||||
|
M2GD_MOUSE
|
||||||
| θ value | 8.99321e-117 (rank : 4) | NC score | 0.914358 (rank : 4) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9DBT9, Q8R1S7 | Gene names | Dmgdh | |||
|
Domain Architecture |

|
|||||
| Description | Dimethylglycine dehydrogenase, mitochondrial precursor (EC 1.5.99.2) (ME2GLYDH). | |||||
|
GCST_HUMAN
|
||||||
| θ value | 1.02475e-11 (rank : 5) | NC score | 0.527112 (rank : 5) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P48728 | Gene names | AMT, GCST | |||
|
Domain Architecture |
|
|||||
| Description | Aminomethyltransferase, mitochondrial precursor (EC 2.1.2.10) (Glycine cleavage system T protein) (GCVT). | |||||
|
GCST_MOUSE
|
||||||
| θ value | 2.13673e-09 (rank : 6) | NC score | 0.501524 (rank : 6) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CFA2 | Gene names | Amt | |||
|
Domain Architecture |
|
|||||
| Description | Aminomethyltransferase, mitochondrial precursor (EC 2.1.2.10) (Glycine cleavage system T protein) (GCVT). | |||||
|
SOX_HUMAN
|
||||||
| θ value | 0.00228821 (rank : 7) | NC score | 0.281893 (rank : 7) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9P0Z9, Q96H28, Q9C070 | Gene names | PIPOX, LPIPOX, PSO | |||
|
Domain Architecture |
|
|||||
| Description | Peroxisomal sarcosine oxidase (EC 1.5.3.1) (PSO) (L-pipecolate oxidase) (EC 1.5.3.7) (L-pipecolic acid oxidase). | |||||
|
SOX_MOUSE
|
||||||
| θ value | 0.0193708 (rank : 8) | NC score | 0.217448 (rank : 8) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9D826, O55223 | Gene names | Pipox, Pso | |||
|
Domain Architecture |
|
|||||
| Description | Peroxisomal sarcosine oxidase (EC 1.5.3.1) (PSO) (L-pipecolate oxidase) (EC 1.5.3.7) (L-pipecolic acid oxidase). | |||||
|
L2HDH_HUMAN
|
||||||
| θ value | 0.125558 (rank : 9) | NC score | 0.153025 (rank : 10) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9H9P8, Q9BRR1 | Gene names | L2HGDH, C14orf160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | L-2-hydroxyglutarate dehydrogenase, mitochondrial precursor (EC 1.1.99.2) (Duranin). | |||||
|
L2HDH_MOUSE
|
||||||
| θ value | 0.163984 (rank : 10) | NC score | 0.123747 (rank : 11) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q91YP0, Q3TH61, Q3U7Z0, Q3ULY6 | Gene names | L2hgdh | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | L-2-hydroxyglutarate dehydrogenase, mitochondrial precursor (EC 1.1.99.2) (Duranin). | |||||
|
DLDH_MOUSE
|
||||||
| θ value | 0.279714 (rank : 11) | NC score | 0.049925 (rank : 13) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O08749 | Gene names | Dld | |||
|
Domain Architecture |

|
|||||
| Description | Dihydrolipoyl dehydrogenase, mitochondrial precursor (EC 1.8.1.4) (Dihydrolipoamide dehydrogenase). | |||||
|
GPDM_MOUSE
|
||||||
| θ value | 0.279714 (rank : 12) | NC score | 0.169524 (rank : 9) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q64521, Q61507 | Gene names | Gpd2, Gdm1 | |||
|
Domain Architecture |

|
|||||
| Description | Glycerol-3-phosphate dehydrogenase, mitochondrial precursor (EC 1.1.99.5) (GPD-M) (GPDH-M). | |||||
|
GPDM_HUMAN
|
||||||
| θ value | 0.813845 (rank : 13) | NC score | 0.113460 (rank : 12) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P43304, Q59FR1, Q9HAP9 | Gene names | GPD2 | |||
|
Domain Architecture |

|
|||||
| Description | Glycerol-3-phosphate dehydrogenase, mitochondrial precursor (EC 1.1.99.5) (GPD-M) (GPDH-M) (mtGPD). | |||||
|
PTN4_HUMAN
|
||||||
| θ value | 1.81305 (rank : 14) | NC score | 0.007090 (rank : 24) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P29074 | Gene names | PTPN4 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 4 (EC 3.1.3.48) (Protein-tyrosine phosphatase MEG1) (PTPase-MEG1) (MEG). | |||||
|
FREM1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 15) | NC score | 0.008084 (rank : 22) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q5H8C1, Q5VV00, Q5VV01, Q6MZI4, Q8NEG9, Q96LI3 | Gene names | FREM1, C9orf154 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FRAS1-related extracellular matrix protein 1 precursor (Protein QBRICK). | |||||
|
MSLN_HUMAN
|
||||||
| θ value | 5.27518 (rank : 16) | NC score | 0.029664 (rank : 14) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q13421, Q14859, Q4VQD5, Q96GR6, Q96KJ5, Q9BR17, Q9BTR2, Q9UK57 | Gene names | MSLN, MPF | |||
|
Domain Architecture |

|
|||||
| Description | Mesothelin precursor (Pre-pro-megakaryocyte potentiating factor) (CAK1 antigen) [Contains: Megakaryocyte-potentiating factor (MPF); Mesothelin, cleaved form]. | |||||
|
DLDH_HUMAN
|
||||||
| θ value | 6.88961 (rank : 17) | NC score | 0.026859 (rank : 15) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P09622, Q14131, Q14167 | Gene names | DLD, GCSL, LAD, PHE3 | |||
|
Domain Architecture |

|
|||||
| Description | Dihydrolipoyl dehydrogenase, mitochondrial precursor (EC 1.8.1.4) (Dihydrolipoamide dehydrogenase) (Glycine cleavage system L protein). | |||||
|
SIA7D_HUMAN
|
||||||
| θ value | 6.88961 (rank : 18) | NC score | 0.014379 (rank : 16) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H4F1, Q9NWU6, Q9UKU1, Q9ULB9, Q9Y3G3, Q9Y3G4 | Gene names | ST6GALNAC4, SIAT3C, SIAT7D | |||
|
Domain Architecture |
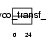
|
|||||
| Description | Alpha-N-acetyl-neuraminyl-2,3-beta-galactosyl-1,3-N-acetyl- galactosaminide alpha-2,6-sialyltransferase (EC 2.4.99.7) (NeuAc- alpha-2,3-Gal-beta-1,3-GalNAc-alpha-2,6-sialyltransferase) (ST6GalNAc IV) (Sialyltransferase 7D) (Sialyltransferase 3C). | |||||
|
CLCA_HUMAN
|
||||||
| θ value | 8.99809 (rank : 19) | NC score | 0.010460 (rank : 21) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P09496 | Gene names | CLTA | |||
|
Domain Architecture |
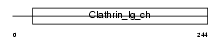
|
|||||
| Description | Clathrin light chain A (Lca). | |||||
|
MYST3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 20) | NC score | 0.012501 (rank : 19) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q92794 | Gene names | MYST3, MOZ, RUNXBP2, ZNF220 | |||
|
Domain Architecture |

|
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Runt-related transcription factor-binding protein 2) (Monocytic leukemia zinc finger protein) (Zinc finger protein 220). | |||||
|
MYST3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 21) | NC score | 0.011851 (rank : 20) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 688 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BZ21, Q8BW52, Q8BYH1, Q8C1F3 | Gene names | Myst3, Moz | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Monocytic leukemia zinc finger protein) (Monocytic leukemia zinc finger homolog). | |||||
|
SNTG1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 22) | NC score | 0.006781 (rank : 25) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NSN8, Q9NY98 | Gene names | SNTG1 | |||
|
Domain Architecture |

|
|||||
| Description | Gamma-1-syntrophin (G1SYN) (Syntrophin 4) (SYN4). | |||||
|
STXB3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 23) | NC score | 0.007197 (rank : 23) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q60770 | Gene names | Stxbp3, Stxbp3a, Unc18c | |||
|
Domain Architecture |

|
|||||
| Description | Syntaxin-binding protein 3 (Unc-18 homolog 3) (Unc-18C) (MUNC-18-3). | |||||
|
UBA3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 24) | NC score | 0.013754 (rank : 17) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8TBC4, O76088, Q9NTU3 | Gene names | UBE1C, UBA3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NEDD8-activating enzyme E1 catalytic subunit (EC 6.3.2.-) (Ubiquitin- activating enzyme 3) (NEDD8-activating enzyme E1C) (Ubiquitin- activating enzyme E1C). | |||||
|
UBA3_MOUSE
|
||||||
| θ value | 8.99809 (rank : 25) | NC score | 0.013745 (rank : 18) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8C878, O88598, Q9D6B7 | Gene names | Ube1c, Uba3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NEDD8-activating enzyme E1 catalytic subunit (EC 6.3.2.-) (Ubiquitin- activating enzyme 3) (NEDD8-activating enzyme E1C) (Ubiquitin- activating enzyme E1C). | |||||
|
ZHX3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 26) | NC score | 0.006524 (rank : 26) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 331 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9H4I2, O43145 | Gene names | ZHX3, KIAA0395, TIX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and homeodomain protein 3 (Triple homeobox protein 1). | |||||
|
SARDH_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q99LB7 | Gene names | Sardh | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcosine dehydrogenase, mitochondrial precursor (EC 1.5.99.1) (SarDH). | |||||
|
SARDH_HUMAN
|
||||||
| NC score | 0.996500 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9UL12, Q9Y280, Q9Y2Y3 | Gene names | SARDH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcosine dehydrogenase, mitochondrial precursor (EC 1.5.99.1) (SarDH) (BPR-2). | |||||
|
M2GD_HUMAN
|
||||||
| NC score | 0.916142 (rank : 3) | θ value | 2.79562e-118 (rank : 3) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9UI17 | Gene names | DMGDH | |||
|
Domain Architecture |

|
|||||
| Description | Dimethylglycine dehydrogenase, mitochondrial precursor (EC 1.5.99.2) (ME2GLYDH). | |||||
|
M2GD_MOUSE
|
||||||
| NC score | 0.914358 (rank : 4) | θ value | 8.99321e-117 (rank : 4) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9DBT9, Q8R1S7 | Gene names | Dmgdh | |||
|
Domain Architecture |

|
|||||
| Description | Dimethylglycine dehydrogenase, mitochondrial precursor (EC 1.5.99.2) (ME2GLYDH). | |||||
|
GCST_HUMAN
|
||||||
| NC score | 0.527112 (rank : 5) | θ value | 1.02475e-11 (rank : 5) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P48728 | Gene names | AMT, GCST | |||
|
Domain Architecture |

|
|||||
| Description | Aminomethyltransferase, mitochondrial precursor (EC 2.1.2.10) (Glycine cleavage system T protein) (GCVT). | |||||
|
GCST_MOUSE
|
||||||
| NC score | 0.501524 (rank : 6) | θ value | 2.13673e-09 (rank : 6) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CFA2 | Gene names | Amt | |||
|
Domain Architecture |

|
|||||
| Description | Aminomethyltransferase, mitochondrial precursor (EC 2.1.2.10) (Glycine cleavage system T protein) (GCVT). | |||||
|
SOX_HUMAN
|
||||||
| NC score | 0.281893 (rank : 7) | θ value | 0.00228821 (rank : 7) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9P0Z9, Q96H28, Q9C070 | Gene names | PIPOX, LPIPOX, PSO | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisomal sarcosine oxidase (EC 1.5.3.1) (PSO) (L-pipecolate oxidase) (EC 1.5.3.7) (L-pipecolic acid oxidase). | |||||
|
SOX_MOUSE
|
||||||
| NC score | 0.217448 (rank : 8) | θ value | 0.0193708 (rank : 8) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9D826, O55223 | Gene names | Pipox, Pso | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisomal sarcosine oxidase (EC 1.5.3.1) (PSO) (L-pipecolate oxidase) (EC 1.5.3.7) (L-pipecolic acid oxidase). | |||||
|
GPDM_MOUSE
|
||||||
| NC score | 0.169524 (rank : 9) | θ value | 0.279714 (rank : 12) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q64521, Q61507 | Gene names | Gpd2, Gdm1 | |||
|
Domain Architecture |

|
|||||
| Description | Glycerol-3-phosphate dehydrogenase, mitochondrial precursor (EC 1.1.99.5) (GPD-M) (GPDH-M). | |||||
|
L2HDH_HUMAN
|
||||||
| NC score | 0.153025 (rank : 10) | θ value | 0.125558 (rank : 9) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9H9P8, Q9BRR1 | Gene names | L2HGDH, C14orf160 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | L-2-hydroxyglutarate dehydrogenase, mitochondrial precursor (EC 1.1.99.2) (Duranin). | |||||
|
L2HDH_MOUSE
|
||||||
| NC score | 0.123747 (rank : 11) | θ value | 0.163984 (rank : 10) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q91YP0, Q3TH61, Q3U7Z0, Q3ULY6 | Gene names | L2hgdh | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | L-2-hydroxyglutarate dehydrogenase, mitochondrial precursor (EC 1.1.99.2) (Duranin). | |||||
|
GPDM_HUMAN
|
||||||
| NC score | 0.113460 (rank : 12) | θ value | 0.813845 (rank : 13) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P43304, Q59FR1, Q9HAP9 | Gene names | GPD2 | |||
|
Domain Architecture |

|
|||||
| Description | Glycerol-3-phosphate dehydrogenase, mitochondrial precursor (EC 1.1.99.5) (GPD-M) (GPDH-M) (mtGPD). | |||||
|
DLDH_MOUSE
|
||||||
| NC score | 0.049925 (rank : 13) | θ value | 0.279714 (rank : 11) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O08749 | Gene names | Dld | |||
|
Domain Architecture |

|
|||||
| Description | Dihydrolipoyl dehydrogenase, mitochondrial precursor (EC 1.8.1.4) (Dihydrolipoamide dehydrogenase). | |||||
|
MSLN_HUMAN
|
||||||
| NC score | 0.029664 (rank : 14) | θ value | 5.27518 (rank : 16) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q13421, Q14859, Q4VQD5, Q96GR6, Q96KJ5, Q9BR17, Q9BTR2, Q9UK57 | Gene names | MSLN, MPF | |||
|
Domain Architecture |

|
|||||
| Description | Mesothelin precursor (Pre-pro-megakaryocyte potentiating factor) (CAK1 antigen) [Contains: Megakaryocyte-potentiating factor (MPF); Mesothelin, cleaved form]. | |||||
|
DLDH_HUMAN
|
||||||
| NC score | 0.026859 (rank : 15) | θ value | 6.88961 (rank : 17) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P09622, Q14131, Q14167 | Gene names | DLD, GCSL, LAD, PHE3 | |||
|
Domain Architecture |

|
|||||
| Description | Dihydrolipoyl dehydrogenase, mitochondrial precursor (EC 1.8.1.4) (Dihydrolipoamide dehydrogenase) (Glycine cleavage system L protein). | |||||
|
SIA7D_HUMAN
|
||||||
| NC score | 0.014379 (rank : 16) | θ value | 6.88961 (rank : 18) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H4F1, Q9NWU6, Q9UKU1, Q9ULB9, Q9Y3G3, Q9Y3G4 | Gene names | ST6GALNAC4, SIAT3C, SIAT7D | |||
|
Domain Architecture |
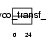
|
|||||
| Description | Alpha-N-acetyl-neuraminyl-2,3-beta-galactosyl-1,3-N-acetyl- galactosaminide alpha-2,6-sialyltransferase (EC 2.4.99.7) (NeuAc- alpha-2,3-Gal-beta-1,3-GalNAc-alpha-2,6-sialyltransferase) (ST6GalNAc IV) (Sialyltransferase 7D) (Sialyltransferase 3C). | |||||
|
UBA3_HUMAN
|
||||||
| NC score | 0.013754 (rank : 17) | θ value | 8.99809 (rank : 24) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8TBC4, O76088, Q9NTU3 | Gene names | UBE1C, UBA3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NEDD8-activating enzyme E1 catalytic subunit (EC 6.3.2.-) (Ubiquitin- activating enzyme 3) (NEDD8-activating enzyme E1C) (Ubiquitin- activating enzyme E1C). | |||||
|
UBA3_MOUSE
|
||||||
| NC score | 0.013745 (rank : 18) | θ value | 8.99809 (rank : 25) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8C878, O88598, Q9D6B7 | Gene names | Ube1c, Uba3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NEDD8-activating enzyme E1 catalytic subunit (EC 6.3.2.-) (Ubiquitin- activating enzyme 3) (NEDD8-activating enzyme E1C) (Ubiquitin- activating enzyme E1C). | |||||
|
MYST3_HUMAN
|
||||||
| NC score | 0.012501 (rank : 19) | θ value | 8.99809 (rank : 20) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q92794 | Gene names | MYST3, MOZ, RUNXBP2, ZNF220 | |||
|
Domain Architecture |

|
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Runt-related transcription factor-binding protein 2) (Monocytic leukemia zinc finger protein) (Zinc finger protein 220). | |||||
|
MYST3_MOUSE
|
||||||
| NC score | 0.011851 (rank : 20) | θ value | 8.99809 (rank : 21) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 688 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BZ21, Q8BW52, Q8BYH1, Q8C1F3 | Gene names | Myst3, Moz | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Monocytic leukemia zinc finger protein) (Monocytic leukemia zinc finger homolog). | |||||
|
CLCA_HUMAN
|
||||||
| NC score | 0.010460 (rank : 21) | θ value | 8.99809 (rank : 19) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 90 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P09496 | Gene names | CLTA | |||
|
Domain Architecture |

|
|||||
| Description | Clathrin light chain A (Lca). | |||||
|
FREM1_HUMAN
|
||||||
| NC score | 0.008084 (rank : 22) | θ value | 5.27518 (rank : 15) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q5H8C1, Q5VV00, Q5VV01, Q6MZI4, Q8NEG9, Q96LI3 | Gene names | FREM1, C9orf154 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FRAS1-related extracellular matrix protein 1 precursor (Protein QBRICK). | |||||
|
STXB3_MOUSE
|
||||||
| NC score | 0.007197 (rank : 23) | θ value | 8.99809 (rank : 23) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q60770 | Gene names | Stxbp3, Stxbp3a, Unc18c | |||
|
Domain Architecture |

|
|||||
| Description | Syntaxin-binding protein 3 (Unc-18 homolog 3) (Unc-18C) (MUNC-18-3). | |||||
|
PTN4_HUMAN
|
||||||
| NC score | 0.007090 (rank : 24) | θ value | 1.81305 (rank : 14) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 303 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P29074 | Gene names | PTPN4 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 4 (EC 3.1.3.48) (Protein-tyrosine phosphatase MEG1) (PTPase-MEG1) (MEG). | |||||
|
SNTG1_HUMAN
|
||||||
| NC score | 0.006781 (rank : 25) | θ value | 8.99809 (rank : 22) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 181 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NSN8, Q9NY98 | Gene names | SNTG1 | |||
|
Domain Architecture |

|
|||||
| Description | Gamma-1-syntrophin (G1SYN) (Syntrophin 4) (SYN4). | |||||
|
ZHX3_HUMAN
|
||||||
| NC score | 0.006524 (rank : 26) | θ value | 8.99809 (rank : 26) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 331 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9H4I2, O43145 | Gene names | ZHX3, KIAA0395, TIX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and homeodomain protein 3 (Triple homeobox protein 1). | |||||