Please be patient as the page loads
|
POLK_MOUSE
|
||||||
| SwissProt Accessions | Q9QUG2, Q7TPY7 | Gene names | Polk, Dinb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA polymerase kappa (EC 2.7.7.7) (DINB protein) (DINP). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
POLK_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.985975 (rank : 2) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UBT6, Q86VJ8, Q8IZY0, Q8IZY1, Q8NB30, Q96L01, Q96Q86, Q96Q87, Q9UHC5 | Gene names | POLK, DINB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA polymerase kappa (EC 2.7.7.7) (DINB protein) (DINP). | |||||
|
POLK_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q9QUG2, Q7TPY7 | Gene names | Polk, Dinb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA polymerase kappa (EC 2.7.7.7) (DINB protein) (DINP). | |||||
|
POLI_MOUSE
|
||||||
| θ value | 1.79631e-32 (rank : 3) | NC score | 0.742718 (rank : 3) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q6R3M4, Q641P1, Q6R3M3, Q9R1A6 | Gene names | Poli, Rad30b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA polymerase iota (EC 2.7.7.7) (Rad30 homolog B). | |||||
|
POLI_HUMAN
|
||||||
| θ value | 9.8567e-31 (rank : 4) | NC score | 0.737770 (rank : 4) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UNA4, Q8N590, Q9H0S1, Q9NYH6 | Gene names | POLI, RAD30B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA polymerase iota (EC 2.7.7.7) (RAD30 homolog B) (Eta2). | |||||
|
POLH_MOUSE
|
||||||
| θ value | 2.77775e-25 (rank : 5) | NC score | 0.690122 (rank : 5) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9JJN0, Q9JJJ2 | Gene names | Polh, Rad30a, Xpv | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA polymerase eta (EC 2.7.7.7) (RAD30 homolog A) (Xeroderma pigmentosum variant type protein homolog). | |||||
|
POLH_HUMAN
|
||||||
| θ value | 5.79196e-23 (rank : 6) | NC score | 0.681310 (rank : 6) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Y253, Q7L8E3, Q96BC4, Q9BX13 | Gene names | POLH, RAD30, RAD30A, XPV | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA polymerase eta (EC 2.7.7.7) (RAD30 homolog A) (Xeroderma pigmentosum variant type protein). | |||||
|
REV1_HUMAN
|
||||||
| θ value | 2.43343e-21 (rank : 7) | NC score | 0.600833 (rank : 7) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UBZ9, O95941, Q9C0J4, Q9NUP2 | Gene names | REV1L, REV1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein REV1 (EC 2.7.7.-) (Rev1-like terminal deoxycytidyl transferase) (Alpha integrin-binding protein 80) (AIBP80). | |||||
|
REV1_MOUSE
|
||||||
| θ value | 1.5773e-20 (rank : 8) | NC score | 0.579750 (rank : 8) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q920Q2, Q9QXV2 | Gene names | Rev1l, Rev1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein REV1 (EC 2.7.7.-) (Rev1-like terminal deoxycytidyl transferase). | |||||
|
DMP1_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 9) | NC score | 0.054980 (rank : 9) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O55188 | Gene names | Dmp1, Dmp | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1) (AG1). | |||||
|
HIP1_HUMAN
|
||||||
| θ value | 0.163984 (rank : 10) | NC score | 0.035859 (rank : 13) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 1016 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O00291, O00328, Q8TDL4 | Gene names | HIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin-interacting protein 1 (HIP-I). | |||||
|
HRX_HUMAN
|
||||||
| θ value | 0.279714 (rank : 11) | NC score | 0.030011 (rank : 21) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
NIN_HUMAN
|
||||||
| θ value | 0.279714 (rank : 12) | NC score | 0.034324 (rank : 16) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 1416 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8N4C6, Q6P0P6, Q9BWU6, Q9C012, Q9C013, Q9C014, Q9H5I6, Q9HAT7, Q9HBY5, Q9HCK7, Q9UH61 | Gene names | NIN, KIAA1565 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ninein (hNinein) (Glycogen synthase kinase 3 beta-interacting protein) (GSK3B-interacting protein). | |||||
|
RAD50_HUMAN
|
||||||
| θ value | 0.62314 (rank : 13) | NC score | 0.036914 (rank : 12) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 1317 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q92878, O43254, Q6GMT7, Q6P5X3, Q9UP86 | Gene names | RAD50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein RAD50 (EC 3.6.-.-) (hRAD50). | |||||
|
CTRO_HUMAN
|
||||||
| θ value | 0.813845 (rank : 14) | NC score | 0.012845 (rank : 55) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 2092 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O14578, Q6XUH8, Q86UQ9, Q9UPZ7 | Gene names | CIT, KIAA0949, STK21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Citron Rho-interacting kinase (EC 2.7.11.1) (CRIK) (Rho-interacting, serine/threonine-protein kinase 21). | |||||
|
MRCKB_HUMAN
|
||||||
| θ value | 0.813845 (rank : 15) | NC score | 0.014490 (rank : 53) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 2239 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9Y5S2, Q2L7A5, Q86TJ1, Q9ULU5 | Gene names | CDC42BPB, KIAA1124 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MRCK beta (EC 2.7.11.1) (CDC42-binding protein kinase beta) (Myotonic dystrophy kinase-related CDC42-binding kinase beta) (Myotonic dystrophy protein kinase-like beta) (MRCK beta) (DMPK-like beta). | |||||
|
PLEC1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 16) | NC score | 0.022715 (rank : 34) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 1326 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q15149, Q15148, Q16640 | Gene names | PLEC1 | |||
|
Domain Architecture |

|
|||||
| Description | Plectin-1 (PLTN) (PCN) (Hemidesmosomal protein 1) (HD1). | |||||
|
BRCA2_HUMAN
|
||||||
| θ value | 1.38821 (rank : 17) | NC score | 0.029134 (rank : 23) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P51587, O00183, O15008, Q13879 | Gene names | BRCA2, FANCD1 | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer type 2 susceptibility protein (Fanconi anemia group D1 protein). | |||||
|
TRIPC_HUMAN
|
||||||
| θ value | 1.38821 (rank : 18) | NC score | 0.018861 (rank : 42) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 319 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q14669, Q15644 | Gene names | TRIP12, KIAA0045 | |||
|
Domain Architecture |

|
|||||
| Description | Thyroid receptor-interacting protein 12 (TRIP12). | |||||
|
CTNL1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 19) | NC score | 0.023383 (rank : 31) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O88327, Q810J3 | Gene names | Ctnnal1, Catnal1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Alpha-catulin (Catenin alpha-like protein 1) (Alpha-catenin-related protein) (ACRP). | |||||
|
IL12A_HUMAN
|
||||||
| θ value | 1.81305 (rank : 20) | NC score | 0.039416 (rank : 10) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 3 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P29459, Q96QZ1 | Gene names | IL12A, NKSF1 | |||
|
Domain Architecture |
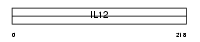
|
|||||
| Description | Interleukin-12 alpha chain precursor (IL-12A) (Cytotoxic lymphocyte maturation factor 35 kDa subunit) (CLMF p35) (NK cell stimulatory factor chain 1) (NKSF1). | |||||
|
SACS_MOUSE
|
||||||
| θ value | 1.81305 (rank : 21) | NC score | 0.035840 (rank : 14) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9JLC8 | Gene names | Sacs | |||
|
Domain Architecture |

|
|||||
| Description | Sacsin. | |||||
|
CLMN_MOUSE
|
||||||
| θ value | 2.36792 (rank : 22) | NC score | 0.017573 (rank : 47) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8C5W0, Q91V71, Q91XT7, Q91XT8, Q91XU9 | Gene names | Clmn | |||
|
Domain Architecture |
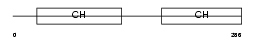
|
|||||
| Description | Calmin. | |||||
|
DSPP_HUMAN
|
||||||
| θ value | 2.36792 (rank : 23) | NC score | 0.022648 (rank : 35) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9NZW4, O95815 | Gene names | DSPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
DSPP_MOUSE
|
||||||
| θ value | 2.36792 (rank : 24) | NC score | 0.026359 (rank : 28) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P97399, O70567 | Gene names | Dspp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor (Dentin matrix protein 3) (DMP-3) [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
FYCO1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 25) | NC score | 0.023354 (rank : 32) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 1149 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9BQS8, Q3MJE6, Q86T41, Q86TB1, Q8TEF9, Q96IV5, Q9H8P9 | Gene names | FYCO1, ZFYVE7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE and coiled-coil domain-containing protein 1 (Zinc finger FYVE domain-containing protein 7). | |||||
|
GCC2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 26) | NC score | 0.027522 (rank : 26) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 1558 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8IWJ2, O15045, Q8TDH3, Q9H2G8 | Gene names | GCC2, KIAA0336 | |||
|
Domain Architecture |

|
|||||
| Description | GRIP and coiled-coil domain-containing protein 2 (Golgi coiled coil protein GCC185) (CTCL tumor antigen se1-1) (CLL-associated antigen KW- 11) (NY-REN-53 antigen). | |||||
|
RBP24_HUMAN
|
||||||
| θ value | 2.36792 (rank : 27) | NC score | 0.027583 (rank : 25) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q4ZG46 | Gene names | RANBP2L4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ran-binding protein 2-like 4 (RanBP2L4). | |||||
|
SACS_HUMAN
|
||||||
| θ value | 2.36792 (rank : 28) | NC score | 0.034844 (rank : 15) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NZJ4, O94835 | Gene names | SACS, KIAA0730 | |||
|
Domain Architecture |

|
|||||
| Description | Sacsin. | |||||
|
TS13_MOUSE
|
||||||
| θ value | 2.36792 (rank : 29) | NC score | 0.033779 (rank : 17) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q01755 | Gene names | Tcp11 | |||
|
Domain Architecture |

|
|||||
| Description | Testis-specific protein PBS13. | |||||
|
FLIP1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 30) | NC score | 0.029674 (rank : 22) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 1176 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9CS72 | Gene names | Filip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filamin-A-interacting protein 1 (FILIP). | |||||
|
GOGA4_HUMAN
|
||||||
| θ value | 3.0926 (rank : 31) | NC score | 0.020914 (rank : 37) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 2034 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q13439, Q13270, Q13654, Q14436 | Gene names | GOLGA4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (Trans-Golgi p230) (256 kDa golgin) (Golgin-245) (Protein 72.1). | |||||
|
ROCK2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 32) | NC score | 0.011876 (rank : 57) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 2073 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O75116, Q9UQN5 | Gene names | ROCK2, KIAA0619 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 2 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 2) (p164 ROCK-2) (Rho kinase 2). | |||||
|
ADDG_MOUSE
|
||||||
| θ value | 4.03905 (rank : 33) | NC score | 0.019241 (rank : 40) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9QYB5, Q9D2M5, Q9QYB6 | Gene names | Add3, Addl | |||
|
Domain Architecture |

|
|||||
| Description | Gamma-adducin (Adducin-like protein 70). | |||||
|
ERCC5_MOUSE
|
||||||
| θ value | 4.03905 (rank : 34) | NC score | 0.038009 (rank : 11) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P35689, Q61528, Q64248 | Gene names | Ercc5, Ercc-5, Xpg | |||
|
Domain Architecture |

|
|||||
| Description | DNA-repair protein complementing XP-G cells homolog (Xeroderma pigmentosum group G-complementing protein homolog) (DNA excision repair protein ERCC-5). | |||||
|
KIF5C_HUMAN
|
||||||
| θ value | 4.03905 (rank : 35) | NC score | 0.014532 (rank : 52) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 974 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O60282, O95079 | Gene names | KIF5C, KIAA0531, NKHC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin heavy chain isoform 5C (Kinesin heavy chain neuron-specific 2). | |||||
|
NEUL_HUMAN
|
||||||
| θ value | 4.03905 (rank : 36) | NC score | 0.020539 (rank : 39) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9BYT8, Q9ULJ4 | Gene names | NLN, KIAA1226 | |||
|
Domain Architecture |

|
|||||
| Description | Neurolysin, mitochondrial precursor (EC 3.4.24.16) (Neurotensin endopeptidase) (Mitochondrial oligopeptidase M) (Microsomal endopeptidase) (MEP). | |||||
|
ADDG_HUMAN
|
||||||
| θ value | 5.27518 (rank : 37) | NC score | 0.017893 (rank : 46) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UEY8, O43243, Q92773, Q9UEY7 | Gene names | ADD3, ADDL | |||
|
Domain Architecture |

|
|||||
| Description | Gamma-adducin (Adducin-like protein 70). | |||||
|
DAAM2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 38) | NC score | 0.012146 (rank : 56) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q86T65, Q5T4U0, Q9NQI5, Q9Y4G0 | Gene names | DAAM2, KIAA0381 | |||
|
Domain Architecture |

|
|||||
| Description | Disheveled-associated activator of morphogenesis 2. | |||||
|
GP158_HUMAN
|
||||||
| θ value | 5.27518 (rank : 39) | NC score | 0.018384 (rank : 45) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q5T848, Q6QR81, Q9ULT3 | Gene names | GPR158, KIAA1136 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 158 precursor. | |||||
|
MAFB_HUMAN
|
||||||
| θ value | 5.27518 (rank : 40) | NC score | 0.018663 (rank : 44) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Y5Q3, Q9H1F1 | Gene names | MAFB, KRML | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor MafB (V-maf musculoaponeurotic fibrosarcoma oncogene homolog B). | |||||
|
MAFB_MOUSE
|
||||||
| θ value | 5.27518 (rank : 41) | NC score | 0.018672 (rank : 43) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P54841 | Gene names | Mafb, Krml, Maf1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor MafB (V-maf musculoaponeurotic fibrosarcoma oncogene homolog B) (Transcription factor MAF1) (Segmentation protein KR) (Kreisler). | |||||
|
MDC1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 42) | NC score | 0.009974 (rank : 60) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
REST_HUMAN
|
||||||
| θ value | 5.27518 (rank : 43) | NC score | 0.018981 (rank : 41) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 1605 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P30622 | Gene names | RSN, CYLN1 | |||
|
Domain Architecture |

|
|||||
| Description | Restin (Cytoplasmic linker protein 170 alpha-2) (CLIP-170) (Reed- Sternberg intermediate filament-associated protein) (Cytoplasmic linker protein 1). | |||||
|
RUSC2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 44) | NC score | 0.017123 (rank : 49) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q80U22, Q8R1D6 | Gene names | Rusc2, Kiaa0375 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RUN and SH3 domain-containing protein 2. | |||||
|
AKAP2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 45) | NC score | 0.028820 (rank : 24) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O54931, O54932, O54933 | Gene names | Akap2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 2 (Protein kinase A-anchoring protein 2) (PRKA2) (AKAP-2) (AKAP expressed in kidney and lung) (AKAP-KL). | |||||
|
AMPH_HUMAN
|
||||||
| θ value | 6.88961 (rank : 46) | NC score | 0.015992 (rank : 50) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P49418, O43538, Q75MJ8, Q75MK5, Q75MM3, Q8N4G0 | Gene names | AMPH, AMPH1 | |||
|
Domain Architecture |

|
|||||
| Description | Amphiphysin. | |||||
|
FV1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 47) | NC score | 0.024471 (rank : 29) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P70213, P70214 | Gene names | Fv1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Friend virus susceptibility protein 1. | |||||
|
GOGA3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 48) | NC score | 0.023313 (rank : 33) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 1437 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P55937, Q80VF5, Q8CCK4, Q9QYT2, Q9QYT3 | Gene names | Golga3, Mea2 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 3 (Golgin-160) (Male-enhanced antigen 2) (MEA-2). | |||||
|
KMHN1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 49) | NC score | 0.027185 (rank : 27) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 950 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8TBZ0, Q86YI9, Q8N7W0 | Gene names | KMHN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cancer/testis antigen KM-HN-1. | |||||
|
L1CAM_MOUSE
|
||||||
| θ value | 6.88961 (rank : 50) | NC score | 0.005644 (rank : 63) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 457 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P11627 | Gene names | L1cam, Caml1 | |||
|
Domain Architecture |

|
|||||
| Description | Neural cell adhesion molecule L1 precursor (N-CAM L1) (CD171 antigen). | |||||
|
PRDM1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 51) | NC score | -0.000976 (rank : 66) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 734 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O75626 | Gene names | PRDM1, BLIMP1 | |||
|
Domain Architecture |
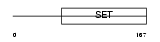
|
|||||
| Description | PR domain zinc finger protein 1 (PR domain-containing protein 1) (Beta-interferon gene positive-regulatory domain I-binding factor) (BLIMP-1) (Positive regulatory domain I-binding factor 1) (PRDI- binding factor 1) (PRDI-BF1). | |||||
|
RPGF6_HUMAN
|
||||||
| θ value | 6.88961 (rank : 52) | NC score | 0.008609 (rank : 61) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8TEU7, Q8NI21, Q8TEU6, Q96PC1 | Gene names | RAPGEF6, PDZGEF2 | |||
|
Domain Architecture |

|
|||||
| Description | Rap guanine nucleotide exchange factor 6 (PDZ domain-containing guanine nucleotide exchange factor 2) (PDZ-GEF2) (RA-GEF-2). | |||||
|
SCG1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 53) | NC score | 0.017524 (rank : 48) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P05060, Q9BQV6, Q9UJA6 | Gene names | CHGB, SCG1 | |||
|
Domain Architecture |

|
|||||
| Description | Secretogranin-1 precursor (Secretogranin I) (SgI) (Chromogranin B) (CgB) [Contains: GAWK peptide; CCB peptide]. | |||||
|
SPY4_HUMAN
|
||||||
| θ value | 6.88961 (rank : 54) | NC score | 0.011215 (rank : 58) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9C004, Q6QIX2, Q9C003 | Gene names | SPRY4 | |||
|
Domain Architecture |
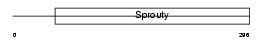
|
|||||
| Description | Sprouty homolog 4 (Spry-4). | |||||
|
SRPK2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 55) | NC score | 0.006684 (rank : 62) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 765 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O54781, Q8VCD9 | Gene names | Srpk2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SRPK2 (EC 2.7.11.1) (Serine/arginine- rich protein-specific kinase 2) (SR-protein-specific kinase 2) (SFRS protein kinase 2). | |||||
|
TPM1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 56) | NC score | 0.031440 (rank : 19) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 716 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P09493, P09494, P10469, Q86W64, Q96IK2, Q9UCY9 | Gene names | TPM1, TMSA | |||
|
Domain Architecture |
|
|||||
| Description | Tropomyosin 1 alpha chain (Alpha-tropomyosin). | |||||
|
TPM1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 57) | NC score | 0.031363 (rank : 20) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 729 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P58771, P02558, P19354, P46902, P99034 | Gene names | Tpm1, Tpm-1, Tpma | |||
|
Domain Architecture |
|
|||||
| Description | Tropomyosin 1 alpha chain (Alpha-tropomyosin). | |||||
|
EME1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 58) | NC score | 0.013177 (rank : 54) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96AY2, Q96N62 | Gene names | EME1, MMS4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Crossover junction endonuclease EME1 (EC 3.1.22.-) (hMMS4). | |||||
|
FLIP1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 59) | NC score | 0.023867 (rank : 30) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 1405 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q7Z7B0, Q5VUL6, Q86TC3, Q8N8B9, Q96SK6, Q9NVI8, Q9ULE5 | Gene names | FILIP1, KIAA1275 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filamin-A-interacting protein 1 (FILIP). | |||||
|
IRK8_HUMAN
|
||||||
| θ value | 8.99809 (rank : 60) | NC score | 0.003977 (rank : 65) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q15842, O00657 | Gene names | KCNJ8 | |||
|
Domain Architecture |
|
|||||
| Description | ATP-sensitive inward rectifier potassium channel 8 (Potassium channel, inwardly rectifying subfamily J member 8) (Inwardly rectifier K(+) channel Kir6.1) (uKATP-1). | |||||
|
MACF1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 61) | NC score | 0.020835 (rank : 38) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 1313 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9QXZ0, P97394, P97395, P97396 | Gene names | Macf1, Acf7, Aclp7, Macf | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-actin cross-linking factor 1 (Actin cross-linking family 7). | |||||
|
MDM4_HUMAN
|
||||||
| θ value | 8.99809 (rank : 62) | NC score | 0.010831 (rank : 59) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O15151, Q6GS18, Q8IV83 | Gene names | MDM4, MDMX | |||
|
Domain Architecture |
|
|||||
| Description | Mdm4 protein (p53-binding protein Mdm4) (Mdm2-like p53-binding protein) (Mdmx protein) (Double minute 4 protein). | |||||
|
RAD50_MOUSE
|
||||||
| θ value | 8.99809 (rank : 63) | NC score | 0.031689 (rank : 18) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 1366 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P70388, Q6PEB0, Q8C2T7, Q9CU59 | Gene names | Rad50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein RAD50 (EC 3.6.-.-) (mRad50). | |||||
|
T23O_HUMAN
|
||||||
| θ value | 8.99809 (rank : 64) | NC score | 0.021986 (rank : 36) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P48775 | Gene names | TDO2, TDO | |||
|
Domain Architecture |

|
|||||
| Description | Tryptophan 2,3-dioxygenase (EC 1.13.11.11) (TO) (Tryptophan pyrrolase) (Tryptophanase) (Tryptophan oxygenase) (Tryptamin 2,3-dioxygenase) (TRPO). | |||||
|
TEX2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 65) | NC score | 0.014583 (rank : 51) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8IWB9, Q6AHZ5, Q8N3L0, Q9C0C5 | Gene names | TEX2, KIAA1738 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Testis-expressed sequence 2 protein. | |||||
|
TRI12_MOUSE
|
||||||
| θ value | 8.99809 (rank : 66) | NC score | 0.004095 (rank : 64) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q99PQ1, Q9D704 | Gene names | Trim12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 12. | |||||
|
POLK_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q9QUG2, Q7TPY7 | Gene names | Polk, Dinb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA polymerase kappa (EC 2.7.7.7) (DINB protein) (DINP). | |||||
|
POLK_HUMAN
|
||||||
| NC score | 0.985975 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q9UBT6, Q86VJ8, Q8IZY0, Q8IZY1, Q8NB30, Q96L01, Q96Q86, Q96Q87, Q9UHC5 | Gene names | POLK, DINB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA polymerase kappa (EC 2.7.7.7) (DINB protein) (DINP). | |||||
|
POLI_MOUSE
|
||||||
| NC score | 0.742718 (rank : 3) | θ value | 1.79631e-32 (rank : 3) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q6R3M4, Q641P1, Q6R3M3, Q9R1A6 | Gene names | Poli, Rad30b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA polymerase iota (EC 2.7.7.7) (Rad30 homolog B). | |||||
|
POLI_HUMAN
|
||||||
| NC score | 0.737770 (rank : 4) | θ value | 9.8567e-31 (rank : 4) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UNA4, Q8N590, Q9H0S1, Q9NYH6 | Gene names | POLI, RAD30B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA polymerase iota (EC 2.7.7.7) (RAD30 homolog B) (Eta2). | |||||
|
POLH_MOUSE
|
||||||
| NC score | 0.690122 (rank : 5) | θ value | 2.77775e-25 (rank : 5) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9JJN0, Q9JJJ2 | Gene names | Polh, Rad30a, Xpv | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA polymerase eta (EC 2.7.7.7) (RAD30 homolog A) (Xeroderma pigmentosum variant type protein homolog). | |||||
|
POLH_HUMAN
|
||||||
| NC score | 0.681310 (rank : 6) | θ value | 5.79196e-23 (rank : 6) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Y253, Q7L8E3, Q96BC4, Q9BX13 | Gene names | POLH, RAD30, RAD30A, XPV | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA polymerase eta (EC 2.7.7.7) (RAD30 homolog A) (Xeroderma pigmentosum variant type protein). | |||||
|
REV1_HUMAN
|
||||||
| NC score | 0.600833 (rank : 7) | θ value | 2.43343e-21 (rank : 7) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UBZ9, O95941, Q9C0J4, Q9NUP2 | Gene names | REV1L, REV1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein REV1 (EC 2.7.7.-) (Rev1-like terminal deoxycytidyl transferase) (Alpha integrin-binding protein 80) (AIBP80). | |||||
|
REV1_MOUSE
|
||||||
| NC score | 0.579750 (rank : 8) | θ value | 1.5773e-20 (rank : 8) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q920Q2, Q9QXV2 | Gene names | Rev1l, Rev1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein REV1 (EC 2.7.7.-) (Rev1-like terminal deoxycytidyl transferase). | |||||
|
DMP1_MOUSE
|
||||||
| NC score | 0.054980 (rank : 9) | θ value | 0.0252991 (rank : 9) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O55188 | Gene names | Dmp1, Dmp | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1) (AG1). | |||||
|
IL12A_HUMAN
|
||||||
| NC score | 0.039416 (rank : 10) | θ value | 1.81305 (rank : 20) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 3 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P29459, Q96QZ1 | Gene names | IL12A, NKSF1 | |||
|
Domain Architecture |

|
|||||
| Description | Interleukin-12 alpha chain precursor (IL-12A) (Cytotoxic lymphocyte maturation factor 35 kDa subunit) (CLMF p35) (NK cell stimulatory factor chain 1) (NKSF1). | |||||
|
ERCC5_MOUSE
|
||||||
| NC score | 0.038009 (rank : 11) | θ value | 4.03905 (rank : 34) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 206 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P35689, Q61528, Q64248 | Gene names | Ercc5, Ercc-5, Xpg | |||
|
Domain Architecture |

|
|||||
| Description | DNA-repair protein complementing XP-G cells homolog (Xeroderma pigmentosum group G-complementing protein homolog) (DNA excision repair protein ERCC-5). | |||||
|
RAD50_HUMAN
|
||||||
| NC score | 0.036914 (rank : 12) | θ value | 0.62314 (rank : 13) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 1317 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | Q92878, O43254, Q6GMT7, Q6P5X3, Q9UP86 | Gene names | RAD50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein RAD50 (EC 3.6.-.-) (hRAD50). | |||||
|
HIP1_HUMAN
|
||||||
| NC score | 0.035859 (rank : 13) | θ value | 0.163984 (rank : 10) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 1016 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O00291, O00328, Q8TDL4 | Gene names | HIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin-interacting protein 1 (HIP-I). | |||||
|
SACS_MOUSE
|
||||||
| NC score | 0.035840 (rank : 14) | θ value | 1.81305 (rank : 21) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9JLC8 | Gene names | Sacs | |||
|
Domain Architecture |

|
|||||
| Description | Sacsin. | |||||
|
SACS_HUMAN
|
||||||
| NC score | 0.034844 (rank : 15) | θ value | 2.36792 (rank : 28) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NZJ4, O94835 | Gene names | SACS, KIAA0730 | |||
|
Domain Architecture |

|
|||||
| Description | Sacsin. | |||||
|
NIN_HUMAN
|
||||||
| NC score | 0.034324 (rank : 16) | θ value | 0.279714 (rank : 12) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 1416 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8N4C6, Q6P0P6, Q9BWU6, Q9C012, Q9C013, Q9C014, Q9H5I6, Q9HAT7, Q9HBY5, Q9HCK7, Q9UH61 | Gene names | NIN, KIAA1565 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ninein (hNinein) (Glycogen synthase kinase 3 beta-interacting protein) (GSK3B-interacting protein). | |||||
|
TS13_MOUSE
|
||||||
| NC score | 0.033779 (rank : 17) | θ value | 2.36792 (rank : 29) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 191 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q01755 | Gene names | Tcp11 | |||
|
Domain Architecture |

|
|||||
| Description | Testis-specific protein PBS13. | |||||
|
RAD50_MOUSE
|
||||||
| NC score | 0.031689 (rank : 18) | θ value | 8.99809 (rank : 63) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 1366 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P70388, Q6PEB0, Q8C2T7, Q9CU59 | Gene names | Rad50 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA repair protein RAD50 (EC 3.6.-.-) (mRad50). | |||||
|
TPM1_HUMAN
|
||||||
| NC score | 0.031440 (rank : 19) | θ value | 6.88961 (rank : 56) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 716 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P09493, P09494, P10469, Q86W64, Q96IK2, Q9UCY9 | Gene names | TPM1, TMSA | |||
|
Domain Architecture |

|
|||||
| Description | Tropomyosin 1 alpha chain (Alpha-tropomyosin). | |||||
|
TPM1_MOUSE
|
||||||
| NC score | 0.031363 (rank : 20) | θ value | 6.88961 (rank : 57) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 729 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P58771, P02558, P19354, P46902, P99034 | Gene names | Tpm1, Tpm-1, Tpma | |||
|
Domain Architecture |

|
|||||
| Description | Tropomyosin 1 alpha chain (Alpha-tropomyosin). | |||||
|
HRX_HUMAN
|
||||||
| NC score | 0.030011 (rank : 21) | θ value | 0.279714 (rank : 11) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q03164, Q13743, Q13744, Q14845, Q16364, Q6UBD1, Q9UMA3 | Gene names | MLL, ALL1, HRX, HTRX, TRX1 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein HRX (ALL-1) (Trithorax-like protein). | |||||
|
FLIP1_MOUSE
|
||||||
| NC score | 0.029674 (rank : 22) | θ value | 3.0926 (rank : 30) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 1176 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9CS72 | Gene names | Filip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filamin-A-interacting protein 1 (FILIP). | |||||
|
BRCA2_HUMAN
|
||||||
| NC score | 0.029134 (rank : 23) | θ value | 1.38821 (rank : 17) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 202 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P51587, O00183, O15008, Q13879 | Gene names | BRCA2, FANCD1 | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer type 2 susceptibility protein (Fanconi anemia group D1 protein). | |||||
|
AKAP2_MOUSE
|
||||||
| NC score | 0.028820 (rank : 24) | θ value | 6.88961 (rank : 45) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O54931, O54932, O54933 | Gene names | Akap2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 2 (Protein kinase A-anchoring protein 2) (PRKA2) (AKAP-2) (AKAP expressed in kidney and lung) (AKAP-KL). | |||||
|
RBP24_HUMAN
|
||||||
| NC score | 0.027583 (rank : 25) | θ value | 2.36792 (rank : 27) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 1559 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q4ZG46 | Gene names | RANBP2L4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ran-binding protein 2-like 4 (RanBP2L4). | |||||
|
GCC2_HUMAN
|
||||||
| NC score | 0.027522 (rank : 26) | θ value | 2.36792 (rank : 26) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 1558 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8IWJ2, O15045, Q8TDH3, Q9H2G8 | Gene names | GCC2, KIAA0336 | |||
|
Domain Architecture |

|
|||||
| Description | GRIP and coiled-coil domain-containing protein 2 (Golgi coiled coil protein GCC185) (CTCL tumor antigen se1-1) (CLL-associated antigen KW- 11) (NY-REN-53 antigen). | |||||
|
KMHN1_HUMAN
|
||||||
| NC score | 0.027185 (rank : 27) | θ value | 6.88961 (rank : 49) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 950 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8TBZ0, Q86YI9, Q8N7W0 | Gene names | KMHN1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cancer/testis antigen KM-HN-1. | |||||
|
DSPP_MOUSE
|
||||||
| NC score | 0.026359 (rank : 28) | θ value | 2.36792 (rank : 24) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P97399, O70567 | Gene names | Dspp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor (Dentin matrix protein 3) (DMP-3) [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
FV1_MOUSE
|
||||||
| NC score | 0.024471 (rank : 29) | θ value | 6.88961 (rank : 47) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P70213, P70214 | Gene names | Fv1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Friend virus susceptibility protein 1. | |||||
|
FLIP1_HUMAN
|
||||||
| NC score | 0.023867 (rank : 30) | θ value | 8.99809 (rank : 59) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 1405 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q7Z7B0, Q5VUL6, Q86TC3, Q8N8B9, Q96SK6, Q9NVI8, Q9ULE5 | Gene names | FILIP1, KIAA1275 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filamin-A-interacting protein 1 (FILIP). | |||||
|
CTNL1_MOUSE
|
||||||
| NC score | 0.023383 (rank : 31) | θ value | 1.81305 (rank : 19) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O88327, Q810J3 | Gene names | Ctnnal1, Catnal1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Alpha-catulin (Catenin alpha-like protein 1) (Alpha-catenin-related protein) (ACRP). | |||||
|
FYCO1_HUMAN
|
||||||
| NC score | 0.023354 (rank : 32) | θ value | 2.36792 (rank : 25) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 1149 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9BQS8, Q3MJE6, Q86T41, Q86TB1, Q8TEF9, Q96IV5, Q9H8P9 | Gene names | FYCO1, ZFYVE7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE and coiled-coil domain-containing protein 1 (Zinc finger FYVE domain-containing protein 7). | |||||
|
GOGA3_MOUSE
|
||||||
| NC score | 0.023313 (rank : 33) | θ value | 6.88961 (rank : 48) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 1437 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P55937, Q80VF5, Q8CCK4, Q9QYT2, Q9QYT3 | Gene names | Golga3, Mea2 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 3 (Golgin-160) (Male-enhanced antigen 2) (MEA-2). | |||||
|
PLEC1_HUMAN
|
||||||
| NC score | 0.022715 (rank : 34) | θ value | 1.06291 (rank : 16) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 1326 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q15149, Q15148, Q16640 | Gene names | PLEC1 | |||
|
Domain Architecture |

|
|||||
| Description | Plectin-1 (PLTN) (PCN) (Hemidesmosomal protein 1) (HD1). | |||||
|
DSPP_HUMAN
|
||||||
| NC score | 0.022648 (rank : 35) | θ value | 2.36792 (rank : 23) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 645 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9NZW4, O95815 | Gene names | DSPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dentin sialophosphoprotein precursor [Contains: Dentin phosphoprotein (Dentin phosphophoryn) (DPP); Dentin sialoprotein (DSP)]. | |||||
|
T23O_HUMAN
|
||||||
| NC score | 0.021986 (rank : 36) | θ value | 8.99809 (rank : 64) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P48775 | Gene names | TDO2, TDO | |||
|
Domain Architecture |

|
|||||
| Description | Tryptophan 2,3-dioxygenase (EC 1.13.11.11) (TO) (Tryptophan pyrrolase) (Tryptophanase) (Tryptophan oxygenase) (Tryptamin 2,3-dioxygenase) (TRPO). | |||||
|
GOGA4_HUMAN
|
||||||
| NC score | 0.020914 (rank : 37) | θ value | 3.0926 (rank : 31) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 2034 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q13439, Q13270, Q13654, Q14436 | Gene names | GOLGA4 | |||
|
Domain Architecture |

|
|||||
| Description | Golgin subfamily A member 4 (Trans-Golgi p230) (256 kDa golgin) (Golgin-245) (Protein 72.1). | |||||
|
MACF1_MOUSE
|
||||||
| NC score | 0.020835 (rank : 38) | θ value | 8.99809 (rank : 61) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 1313 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9QXZ0, P97394, P97395, P97396 | Gene names | Macf1, Acf7, Aclp7, Macf | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-actin cross-linking factor 1 (Actin cross-linking family 7). | |||||
|
NEUL_HUMAN
|
||||||
| NC score | 0.020539 (rank : 39) | θ value | 4.03905 (rank : 36) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9BYT8, Q9ULJ4 | Gene names | NLN, KIAA1226 | |||
|
Domain Architecture |

|
|||||
| Description | Neurolysin, mitochondrial precursor (EC 3.4.24.16) (Neurotensin endopeptidase) (Mitochondrial oligopeptidase M) (Microsomal endopeptidase) (MEP). | |||||
|
ADDG_MOUSE
|
||||||
| NC score | 0.019241 (rank : 40) | θ value | 4.03905 (rank : 33) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 43 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9QYB5, Q9D2M5, Q9QYB6 | Gene names | Add3, Addl | |||
|
Domain Architecture |

|
|||||
| Description | Gamma-adducin (Adducin-like protein 70). | |||||
|
REST_HUMAN
|
||||||
| NC score | 0.018981 (rank : 41) | θ value | 5.27518 (rank : 43) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 1605 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P30622 | Gene names | RSN, CYLN1 | |||
|
Domain Architecture |

|
|||||
| Description | Restin (Cytoplasmic linker protein 170 alpha-2) (CLIP-170) (Reed- Sternberg intermediate filament-associated protein) (Cytoplasmic linker protein 1). | |||||
|
TRIPC_HUMAN
|
||||||
| NC score | 0.018861 (rank : 42) | θ value | 1.38821 (rank : 18) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 319 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q14669, Q15644 | Gene names | TRIP12, KIAA0045 | |||
|
Domain Architecture |

|
|||||
| Description | Thyroid receptor-interacting protein 12 (TRIP12). | |||||
|
MAFB_MOUSE
|
||||||
| NC score | 0.018672 (rank : 43) | θ value | 5.27518 (rank : 41) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P54841 | Gene names | Mafb, Krml, Maf1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor MafB (V-maf musculoaponeurotic fibrosarcoma oncogene homolog B) (Transcription factor MAF1) (Segmentation protein KR) (Kreisler). | |||||
|
MAFB_HUMAN
|
||||||
| NC score | 0.018663 (rank : 44) | θ value | 5.27518 (rank : 40) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9Y5Q3, Q9H1F1 | Gene names | MAFB, KRML | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor MafB (V-maf musculoaponeurotic fibrosarcoma oncogene homolog B). | |||||
|
GP158_HUMAN
|
||||||
| NC score | 0.018384 (rank : 45) | θ value | 5.27518 (rank : 39) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q5T848, Q6QR81, Q9ULT3 | Gene names | GPR158, KIAA1136 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 158 precursor. | |||||
|
ADDG_HUMAN
|
||||||
| NC score | 0.017893 (rank : 46) | θ value | 5.27518 (rank : 37) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UEY8, O43243, Q92773, Q9UEY7 | Gene names | ADD3, ADDL | |||
|
Domain Architecture |

|
|||||
| Description | Gamma-adducin (Adducin-like protein 70). | |||||
|
CLMN_MOUSE
|
||||||
| NC score | 0.017573 (rank : 47) | θ value | 2.36792 (rank : 22) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 180 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8C5W0, Q91V71, Q91XT7, Q91XT8, Q91XU9 | Gene names | Clmn | |||
|
Domain Architecture |

|
|||||
| Description | Calmin. | |||||
|
SCG1_HUMAN
|
||||||
| NC score | 0.017524 (rank : 48) | θ value | 6.88961 (rank : 53) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P05060, Q9BQV6, Q9UJA6 | Gene names | CHGB, SCG1 | |||
|
Domain Architecture |

|
|||||
| Description | Secretogranin-1 precursor (Secretogranin I) (SgI) (Chromogranin B) (CgB) [Contains: GAWK peptide; CCB peptide]. | |||||
|
RUSC2_MOUSE
|
||||||
| NC score | 0.017123 (rank : 49) | θ value | 5.27518 (rank : 44) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 305 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q80U22, Q8R1D6 | Gene names | Rusc2, Kiaa0375 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RUN and SH3 domain-containing protein 2. | |||||
|
AMPH_HUMAN
|
||||||
| NC score | 0.015992 (rank : 50) | θ value | 6.88961 (rank : 46) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P49418, O43538, Q75MJ8, Q75MK5, Q75MM3, Q8N4G0 | Gene names | AMPH, AMPH1 | |||
|
Domain Architecture |

|
|||||
| Description | Amphiphysin. | |||||
|
TEX2_HUMAN
|
||||||
| NC score | 0.014583 (rank : 51) | θ value | 8.99809 (rank : 65) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8IWB9, Q6AHZ5, Q8N3L0, Q9C0C5 | Gene names | TEX2, KIAA1738 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Testis-expressed sequence 2 protein. | |||||
|
KIF5C_HUMAN
|
||||||
| NC score | 0.014532 (rank : 52) | θ value | 4.03905 (rank : 35) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 974 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O60282, O95079 | Gene names | KIF5C, KIAA0531, NKHC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinesin heavy chain isoform 5C (Kinesin heavy chain neuron-specific 2). | |||||
|
MRCKB_HUMAN
|
||||||
| NC score | 0.014490 (rank : 53) | θ value | 0.813845 (rank : 15) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 2239 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9Y5S2, Q2L7A5, Q86TJ1, Q9ULU5 | Gene names | CDC42BPB, KIAA1124 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase MRCK beta (EC 2.7.11.1) (CDC42-binding protein kinase beta) (Myotonic dystrophy kinase-related CDC42-binding kinase beta) (Myotonic dystrophy protein kinase-like beta) (MRCK beta) (DMPK-like beta). | |||||
|
EME1_HUMAN
|
||||||
| NC score | 0.013177 (rank : 54) | θ value | 8.99809 (rank : 58) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96AY2, Q96N62 | Gene names | EME1, MMS4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Crossover junction endonuclease EME1 (EC 3.1.22.-) (hMMS4). | |||||
|
CTRO_HUMAN
|
||||||
| NC score | 0.012845 (rank : 55) | θ value | 0.813845 (rank : 14) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 2092 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O14578, Q6XUH8, Q86UQ9, Q9UPZ7 | Gene names | CIT, KIAA0949, STK21 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Citron Rho-interacting kinase (EC 2.7.11.1) (CRIK) (Rho-interacting, serine/threonine-protein kinase 21). | |||||
|
DAAM2_HUMAN
|
||||||
| NC score | 0.012146 (rank : 56) | θ value | 5.27518 (rank : 38) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q86T65, Q5T4U0, Q9NQI5, Q9Y4G0 | Gene names | DAAM2, KIAA0381 | |||
|
Domain Architecture |

|
|||||
| Description | Disheveled-associated activator of morphogenesis 2. | |||||
|
ROCK2_HUMAN
|
||||||
| NC score | 0.011876 (rank : 57) | θ value | 3.0926 (rank : 32) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 2073 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O75116, Q9UQN5 | Gene names | ROCK2, KIAA0619 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-associated protein kinase 2 (EC 2.7.11.1) (Rho-associated, coiled- coil-containing protein kinase 2) (p164 ROCK-2) (Rho kinase 2). | |||||
|
SPY4_HUMAN
|
||||||
| NC score | 0.011215 (rank : 58) | θ value | 6.88961 (rank : 54) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9C004, Q6QIX2, Q9C003 | Gene names | SPRY4 | |||
|
Domain Architecture |

|
|||||
| Description | Sprouty homolog 4 (Spry-4). | |||||
|
MDM4_HUMAN
|
||||||
| NC score | 0.010831 (rank : 59) | θ value | 8.99809 (rank : 62) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O15151, Q6GS18, Q8IV83 | Gene names | MDM4, MDMX | |||
|
Domain Architecture |

|
|||||
| Description | Mdm4 protein (p53-binding protein Mdm4) (Mdm2-like p53-binding protein) (Mdmx protein) (Double minute 4 protein). | |||||
|
MDC1_HUMAN
|
||||||
| NC score | 0.009974 (rank : 60) | θ value | 5.27518 (rank : 42) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
RPGF6_HUMAN
|
||||||
| NC score | 0.008609 (rank : 61) | θ value | 6.88961 (rank : 52) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8TEU7, Q8NI21, Q8TEU6, Q96PC1 | Gene names | RAPGEF6, PDZGEF2 | |||
|
Domain Architecture |

|
|||||
| Description | Rap guanine nucleotide exchange factor 6 (PDZ domain-containing guanine nucleotide exchange factor 2) (PDZ-GEF2) (RA-GEF-2). | |||||
|
SRPK2_MOUSE
|
||||||
| NC score | 0.006684 (rank : 62) | θ value | 6.88961 (rank : 55) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 765 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O54781, Q8VCD9 | Gene names | Srpk2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase SRPK2 (EC 2.7.11.1) (Serine/arginine- rich protein-specific kinase 2) (SR-protein-specific kinase 2) (SFRS protein kinase 2). | |||||
|
L1CAM_MOUSE
|
||||||
| NC score | 0.005644 (rank : 63) | θ value | 6.88961 (rank : 50) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 457 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P11627 | Gene names | L1cam, Caml1 | |||
|
Domain Architecture |

|
|||||
| Description | Neural cell adhesion molecule L1 precursor (N-CAM L1) (CD171 antigen). | |||||
|
TRI12_MOUSE
|
||||||
| NC score | 0.004095 (rank : 64) | θ value | 8.99809 (rank : 66) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q99PQ1, Q9D704 | Gene names | Trim12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 12. | |||||
|
IRK8_HUMAN
|
||||||
| NC score | 0.003977 (rank : 65) | θ value | 8.99809 (rank : 60) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q15842, O00657 | Gene names | KCNJ8 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-sensitive inward rectifier potassium channel 8 (Potassium channel, inwardly rectifying subfamily J member 8) (Inwardly rectifier K(+) channel Kir6.1) (uKATP-1). | |||||
|
PRDM1_HUMAN
|
||||||
| NC score | -0.000976 (rank : 66) | θ value | 6.88961 (rank : 51) | |||
| Query Neighborhood Hits | 66 | Target Neighborhood Hits | 734 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O75626 | Gene names | PRDM1, BLIMP1 | |||
|
Domain Architecture |

|
|||||
| Description | PR domain zinc finger protein 1 (PR domain-containing protein 1) (Beta-interferon gene positive-regulatory domain I-binding factor) (BLIMP-1) (Positive regulatory domain I-binding factor 1) (PRDI- binding factor 1) (PRDI-BF1). | |||||