Please be patient as the page loads
|
MPRI_MOUSE
|
||||||
| SwissProt Accessions | Q07113, Q61822, Q6LED1 | Gene names | Igf2r | |||
|
Domain Architecture |

|
|||||
| Description | Cation-independent mannose-6-phosphate receptor precursor (CI Man-6-P receptor) (CI-MPR) (M6PR) (Insulin-like growth factor 2 receptor) (Insulin-like growth factor II receptor) (IGF-II receptor) (M6P/IGF2 receptor) (M6P/IGF2R) (300 kDa mannose 6-phosphate receptor) (MPR 300) (MPR300). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
MPRI_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.992617 (rank : 2) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P11717, Q7Z7G9, Q96PT5 | Gene names | IGF2R, MPRI | |||
|
Domain Architecture |

|
|||||
| Description | Cation-independent mannose-6-phosphate receptor precursor (CI Man-6-P receptor) (CI-MPR) (M6PR) (Insulin-like growth factor 2 receptor) (Insulin-like growth factor II receptor) (IGF-II receptor) (M6P/IGF2 receptor) (M6P/IGF2R) (300 kDa mannose 6-phosphate receptor) (MPR 300) (MPR300) (CD222 antigen). | |||||
|
MPRI_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q07113, Q61822, Q6LED1 | Gene names | Igf2r | |||
|
Domain Architecture |

|
|||||
| Description | Cation-independent mannose-6-phosphate receptor precursor (CI Man-6-P receptor) (CI-MPR) (M6PR) (Insulin-like growth factor 2 receptor) (Insulin-like growth factor II receptor) (IGF-II receptor) (M6P/IGF2 receptor) (M6P/IGF2R) (300 kDa mannose 6-phosphate receptor) (MPR 300) (MPR300). | |||||
|
MMP2_HUMAN
|
||||||
| θ value | 1.47974e-10 (rank : 3) | NC score | 0.267652 (rank : 7) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P08253 | Gene names | MMP2, CLG4A | |||
|
Domain Architecture |

|
|||||
| Description | 72 kDa type IV collagenase precursor (EC 3.4.24.24) (72 kDa gelatinase) (Matrix metalloproteinase-2) (MMP-2) (Gelatinase A) (TBE- 1). | |||||
|
MMP9_HUMAN
|
||||||
| θ value | 4.30538e-10 (rank : 4) | NC score | 0.290011 (rank : 6) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P14780, Q3LR70, Q8N725, Q9H4Z1 | Gene names | MMP9, CLG4B | |||
|
Domain Architecture |

|
|||||
| Description | Matrix metalloproteinase-9 precursor (EC 3.4.24.35) (MMP-9) (92 kDa type IV collagenase) (92 kDa gelatinase) (Gelatinase B) (GELB) [Contains: 67 kDa matrix metalloproteinase-9; 82 kDa matrix metalloproteinase-9]. | |||||
|
MMP9_MOUSE
|
||||||
| θ value | 4.30538e-10 (rank : 5) | NC score | 0.290683 (rank : 5) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P41245, Q06788, Q9DC02 | Gene names | Mmp9, Clg4b | |||
|
Domain Architecture |

|
|||||
| Description | Matrix metalloproteinase-9 precursor (EC 3.4.24.35) (MMP-9) (92 kDa type IV collagenase) (92 kDa gelatinase) (Gelatinase B) (GELB). | |||||
|
MMP2_MOUSE
|
||||||
| θ value | 7.34386e-10 (rank : 6) | NC score | 0.264661 (rank : 8) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P33434 | Gene names | Mmp2 | |||
|
Domain Architecture |

|
|||||
| Description | 72 kDa type IV collagenase precursor (EC 3.4.24.24) (72 kDa gelatinase) (Matrix metalloproteinase-2) (MMP-2) (Gelatinase A). | |||||
|
SEL1L_HUMAN
|
||||||
| θ value | 9.59137e-10 (rank : 7) | NC score | 0.445356 (rank : 3) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9UBV2, Q6UWT6, Q9P1T9, Q9UHK7 | Gene names | SEL1L, TSA305 | |||
|
Domain Architecture |

|
|||||
| Description | Sel-1 homolog precursor (Suppressor of lin-12-like protein) (Sel-1L). | |||||
|
SEL1L_MOUSE
|
||||||
| θ value | 9.59137e-10 (rank : 8) | NC score | 0.442330 (rank : 4) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9Z2G6, Q9DBD8 | Gene names | Sel1l, Sel1h | |||
|
Domain Architecture |

|
|||||
| Description | Sel-1 homolog precursor (Suppressor of lin-12-like protein) (Sel-1L). | |||||
|
HGFA_HUMAN
|
||||||
| θ value | 2.79066e-09 (rank : 9) | NC score | 0.125975 (rank : 19) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 436 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q04756, Q14726 | Gene names | HGFAC | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte growth factor activator precursor (EC 3.4.21.-) (HGF activator) (HGFA) [Contains: Hepatocyte growth factor activator short chain; Hepatocyte growth factor activator long chain]. | |||||
|
HGFA_MOUSE
|
||||||
| θ value | 4.76016e-09 (rank : 10) | NC score | 0.125099 (rank : 20) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9R098, Q9JKV4 | Gene names | Hgfac | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte growth factor activator precursor (EC 3.4.21.-) (HGF activator) (HGFA) [Contains: Hepatocyte growth factor activator short chain; Hepatocyte growth factor activator long chain]. | |||||
|
FINC_HUMAN
|
||||||
| θ value | 8.11959e-09 (rank : 11) | NC score | 0.225829 (rank : 16) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 302 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P02751, O95609, O95610, Q14312, Q14325, Q14326, Q6LDP6, Q86T27, Q8IVI8, Q96KP7, Q96KP8, Q96KP9, Q9H1B8, Q9HAP3, Q9UMK2 | Gene names | FN1, FN | |||
|
Domain Architecture |

|
|||||
| Description | Fibronectin precursor (FN) (Cold-insoluble globulin) (CIG). | |||||
|
FINC_MOUSE
|
||||||
| θ value | 8.11959e-09 (rank : 12) | NC score | 0.225370 (rank : 17) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 309 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P11276, Q61567, Q61568, Q61569, Q64233, Q80UI4 | Gene names | Fn1 | |||
|
Domain Architecture |

|
|||||
| Description | Fibronectin precursor (FN). | |||||
|
FA12_HUMAN
|
||||||
| θ value | 4.45536e-07 (rank : 13) | NC score | 0.120787 (rank : 21) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P00748, P78339 | Gene names | F12 | |||
|
Domain Architecture |

|
|||||
| Description | Coagulation factor XII precursor (EC 3.4.21.38) (Hageman factor) (HAF) [Contains: Coagulation factor XIIa heavy chain; Beta-factor XIIa part 1; Beta-factor XIIa part 2; Coagulation factor XIIa light chain]. | |||||
|
LY75_HUMAN
|
||||||
| θ value | 6.43352e-06 (rank : 14) | NC score | 0.257914 (rank : 9) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O60449, O75913, Q7Z575, Q7Z577 | Gene names | LY75, CD205 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphocyte antigen 75 precursor (DEC-205) (gp200-MR6) (CD205 antigen). | |||||
|
MRC2_HUMAN
|
||||||
| θ value | 1.43324e-05 (rank : 15) | NC score | 0.249512 (rank : 12) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9UBG0, Q7LGE7, Q9Y5P9 | Gene names | MRC2, ENDO180, KIAA0709, UPARAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Macrophage mannose receptor 2 precursor (Urokinase receptor-associated protein) (Endocytic receptor 180) (CD280 antigen). | |||||
|
MRC2_MOUSE
|
||||||
| θ value | 1.43324e-05 (rank : 16) | NC score | 0.250426 (rank : 11) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q64449, Q6ZQ64, Q8C6P0 | Gene names | Mrc2, Kiaa0709 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Macrophage mannose receptor 2 precursor (Lectin lambda) (CD280 antigen). | |||||
|
MRC1_MOUSE
|
||||||
| θ value | 1.87187e-05 (rank : 17) | NC score | 0.257779 (rank : 10) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q61830, Q8C502 | Gene names | Mrc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Macrophage mannose receptor 1 precursor (MMR) (CD206 antigen). | |||||
|
MRC1_HUMAN
|
||||||
| θ value | 7.1131e-05 (rank : 18) | NC score | 0.248968 (rank : 13) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P22897 | Gene names | MRC1 | |||
|
Domain Architecture |

|
|||||
| Description | Macrophage mannose receptor 1 precursor (MMR) (CD206 antigen). | |||||
|
LY75_MOUSE
|
||||||
| θ value | 0.000461057 (rank : 19) | NC score | 0.247984 (rank : 14) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q60767, Q8C7T3, Q91XL8, Q9QUZ6 | Gene names | Ly75, Cd205 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphocyte antigen 75 precursor (DEC-205) (CD205 antigen). | |||||
|
MPRD_MOUSE
|
||||||
| θ value | 0.000786445 (rank : 20) | NC score | 0.228771 (rank : 15) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 4 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P24668 | Gene names | M6pr, 46mpr | |||
|
Domain Architecture |
|
|||||
| Description | Cation-dependent mannose-6-phosphate receptor precursor (CD Man-6-P receptor) (CD-MPR) (46 kDa mannose 6-phosphate receptor) (MPR 46). | |||||
|
MPRD_HUMAN
|
||||||
| θ value | 0.00665767 (rank : 21) | NC score | 0.217054 (rank : 18) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P20645 | Gene names | M6PR, MPR46, MPRD | |||
|
Domain Architecture |
|
|||||
| Description | Cation-dependent mannose-6-phosphate receptor precursor (CD Man-6-P receptor) (CD-MPR) (46 kDa mannose 6-phosphate receptor) (MPR 46). | |||||
|
NOTC4_HUMAN
|
||||||
| θ value | 0.163984 (rank : 22) | NC score | 0.023943 (rank : 49) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 905 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q99466, O00306, Q99458, Q99940, Q9H3S8, Q9UII9, Q9UIJ0 | Gene names | NOTCH4 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 4 precursor (Notch 4) (hNotch4) [Contains: Notch 4 extracellular truncation; Notch 4 intracellular domain]. | |||||
|
GLU2B_HUMAN
|
||||||
| θ value | 0.365318 (rank : 23) | NC score | 0.059118 (rank : 26) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 377 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P14314, Q96BU9, Q96D06, Q9P0W9 | Gene names | PRKCSH, G19P1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glucosidase 2 subunit beta precursor (Glucosidase II subunit beta) (Protein kinase C substrate, 60.1 kDa protein, heavy chain) (PKCSH) (80K-H protein). | |||||
|
LRBA_HUMAN
|
||||||
| θ value | 0.365318 (rank : 24) | NC score | 0.027677 (rank : 46) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P50851, Q9H2U3, Q9H2U4 | Gene names | LRBA, BGL, CDC4L, LBA | |||
|
Domain Architecture |

|
|||||
| Description | Lipopolysaccharide-responsive and beige-like anchor protein (CDC4-like protein) (Beige-like protein). | |||||
|
GLU2B_MOUSE
|
||||||
| θ value | 0.62314 (rank : 25) | NC score | 0.055677 (rank : 27) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O08795, Q921X2 | Gene names | Prkcsh | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glucosidase 2 subunit beta precursor (Glucosidase II subunit beta) (Protein kinase C substrate, 60.1 kDa protein, heavy chain) (PKCSH) (80K-H protein). | |||||
|
CAD23_MOUSE
|
||||||
| θ value | 0.813845 (rank : 26) | NC score | 0.009320 (rank : 66) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99PF4, Q99NH1, Q9D4N9 | Gene names | Cdh23 | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin-23 precursor (Otocadherin). | |||||
|
MUG1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 27) | NC score | 0.018621 (rank : 52) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P28665 | Gene names | Mug1, Mug-1 | |||
|
Domain Architecture |

|
|||||
| Description | Murinoglobulin-1 precursor (MuG1). | |||||
|
BTNL8_HUMAN
|
||||||
| θ value | 1.06291 (rank : 28) | NC score | 0.012499 (rank : 63) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6UX41, Q9H730 | Gene names | BTNL8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Butyrophilin-like protein 8 precursor. | |||||
|
RBM19_MOUSE
|
||||||
| θ value | 1.38821 (rank : 29) | NC score | 0.014203 (rank : 58) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8R3C6, Q8BHR0, Q9CW63 | Gene names | Rbm19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable RNA-binding protein 19 (RNA-binding motif protein 19). | |||||
|
DLG7_MOUSE
|
||||||
| θ value | 1.81305 (rank : 30) | NC score | 0.025229 (rank : 48) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8K4R9, Q6ZQL3, Q80VS7, Q8BUC8, Q8BZ79 | Gene names | Dlg7, Kiaa0008 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Discs large homolog 7 (Hepatoma up-regulated protein homolog) (HURP). | |||||
|
RBFAL_HUMAN
|
||||||
| θ value | 1.81305 (rank : 31) | NC score | 0.034110 (rank : 45) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8N0V3, Q6PF07, Q8WZ65, Q9H776 | Gene names | C18orf22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative ribosome-binding factor A, mitochondrial precursor. | |||||
|
ATS20_HUMAN
|
||||||
| θ value | 2.36792 (rank : 32) | NC score | 0.008420 (rank : 68) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 192 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P59510 | Gene names | ADAMTS20 | |||
|
Domain Architecture |

|
|||||
| Description | ADAMTS-20 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase with thrombospondin motifs 20) (ADAM-TS 20) (ADAM-TS20). | |||||
|
GRP78_HUMAN
|
||||||
| θ value | 2.36792 (rank : 33) | NC score | 0.016595 (rank : 54) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P11021, Q2EF78, Q9NPF1, Q9UK02 | Gene names | HSPA5, GRP78 | |||
|
Domain Architecture |

|
|||||
| Description | 78 kDa glucose-regulated protein precursor (GRP 78) (Heat shock 70 kDa protein 5) (Immunoglobulin heavy chain-binding protein) (BiP) (Endoplasmic reticulum lumenal Ca(2+)-binding protein grp78). | |||||
|
GRP78_MOUSE
|
||||||
| θ value | 2.36792 (rank : 34) | NC score | 0.016601 (rank : 53) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P20029, O35642, Q3UFF2, Q61630 | Gene names | Hspa5, Grp78 | |||
|
Domain Architecture |

|
|||||
| Description | 78 kDa glucose-regulated protein precursor (GRP 78) (Heat shock 70 kDa protein 5) (Immunoglobulin heavy chain-binding protein) (BiP). | |||||
|
TNR8_MOUSE
|
||||||
| θ value | 2.36792 (rank : 35) | NC score | 0.023378 (rank : 50) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q60846 | Gene names | Tnfrsf8 | |||
|
Domain Architecture |
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 8 precursor (CD30L receptor) (Lymphocyte activation antigen CD30). | |||||
|
RUSC2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 36) | NC score | 0.025956 (rank : 47) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8N2Y8, O15080, Q6P1W7 | Gene names | RUSC2, KIAA0375 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RUN and SH3 domain-containing protein 2. | |||||
|
CUBN_MOUSE
|
||||||
| θ value | 4.03905 (rank : 37) | NC score | 0.019034 (rank : 51) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 477 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9JLB4 | Gene names | Cubn, Ifcr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cubilin precursor (Intrinsic factor-cobalamin receptor). | |||||
|
ATS12_MOUSE
|
||||||
| θ value | 5.27518 (rank : 38) | NC score | 0.006770 (rank : 69) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 220 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q811B3, Q8BK92, Q8BKY1 | Gene names | Adamts12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADAMTS-12 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase with thrombospondin motifs 12) (ADAM-TS 12) (ADAM-TS12). | |||||
|
P52K_MOUSE
|
||||||
| θ value | 5.27518 (rank : 39) | NC score | 0.015298 (rank : 56) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9CUX1, Q80Y58 | Gene names | Prkrir, Thap0 | |||
|
Domain Architecture |

|
|||||
| Description | 52 kDa repressor of the inhibitor of the protein kinase (p58IPK- interacting protein) (58 kDa interferon-induced protein kinase- interacting protein) (P52rIPK) (THAP domain-containing protein 0). | |||||
|
ZADH1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 40) | NC score | 0.015908 (rank : 55) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8N8N7, Q6MZH8 | Gene names | ZADH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc-binding alcohol dehydrogenase domain-containing protein 1 (EC 1.-.-.-). | |||||
|
CAF1A_HUMAN
|
||||||
| θ value | 6.88961 (rank : 41) | NC score | 0.009239 (rank : 67) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 750 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q13111, Q7Z7K3, Q9UJY8 | Gene names | CHAF1A, CAF, CAF1P150 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromatin assembly factor 1 subunit A (CAF-1 subunit A) (Chromatin assembly factor I p150 subunit) (CAF-I 150 kDa subunit) (CAF-Ip150). | |||||
|
LRP2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 42) | NC score | 0.011456 (rank : 64) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 573 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P98164, O00711, Q16215 | Gene names | LRP2 | |||
|
Domain Architecture |

|
|||||
| Description | Low-density lipoprotein receptor-related protein 2 precursor (Megalin) (Glycoprotein 330) (gp330). | |||||
|
NAR3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 43) | NC score | 0.013333 (rank : 60) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8R2G4, Q8R2F9, Q8R2G0, Q8R2G1, Q8R2G2, Q8R2G3 | Gene names | Art3 | |||
|
Domain Architecture |
|
|||||
| Description | Ecto-ADP-ribosyltransferase 3 precursor (EC 2.4.2.31) (NAD(P)(+)-- arginine ADP-ribosyltransferase 3) (Mono(ADP-ribosyl)transferase 3). | |||||
|
NELL1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 44) | NC score | 0.013817 (rank : 59) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 451 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q92832, Q9Y472 | Gene names | NELL1, NRP1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C-binding protein NELL1 precursor (NEL-like protein 1) (Nel-related protein 1). | |||||
|
SMYD1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 45) | NC score | 0.011080 (rank : 65) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8NB12 | Gene names | SMYD1 | |||
|
Domain Architecture |

|
|||||
| Description | SET and MYND domain-containing protein 1. | |||||
|
TF3C1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 46) | NC score | 0.012771 (rank : 61) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q12789, Q12838, Q9Y4W9 | Gene names | GTF3C1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | General transcription factor 3C polypeptide 1 (Transcription factor IIIC-subunit alpha) (TF3C-alpha) (TFIIIC 220 kDa subunit) (TFIIIC220) (TFIIIC box B-binding subunit). | |||||
|
CABYR_HUMAN
|
||||||
| θ value | 8.99809 (rank : 47) | NC score | 0.012672 (rank : 62) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O75952, Q8WXW5, Q9HAY3, Q9HAY4, Q9HAY5, Q9HCY9 | Gene names | CABYR, CBP86, FSP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcium-binding tyrosine phosphorylation-regulated protein (Calcium- binding protein 86) (Testis-specific calcium-binding protein CBP86) (Fibrousheathin-2) (FSP-2). | |||||
|
CUBN_HUMAN
|
||||||
| θ value | 8.99809 (rank : 48) | NC score | 0.015090 (rank : 57) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 482 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O60494, Q5VTA6, Q96RU9 | Gene names | CUBN, IFCR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cubilin precursor (Intrinsic factor-cobalamin receptor) (Intrinsic factor-vitamin B12 receptor) (460 kDa receptor) (Intestinal intrinsic factor receptor). | |||||
|
RIMB1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 49) | NC score | 0.006390 (rank : 70) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 1376 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q7TNF8, Q5NCP7, Q80TV9, Q8BIH5 | Gene names | Bzrap1, Kiaa0612, Rbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Peripheral-type benzodiazepine receptor-associated protein 1 (PRAX-1) (Peripheral benzodiazepine receptor-interacting protein) (PBR-IP) (RIM-binding protein 1) (RIM-BP1). | |||||
|
SEM4F_MOUSE
|
||||||
| θ value | 8.99809 (rank : 50) | NC score | 0.003601 (rank : 71) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Z123, Q9R1Y1 | Gene names | Sema4f, Semaw | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4F precursor (Semaphorin W) (Sema W). | |||||
|
SLIK1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 51) | NC score | 0.002372 (rank : 72) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q810C1, Q9CXL0 | Gene names | Slitrk1, Sltk1 | |||
|
Domain Architecture |

|
|||||
| Description | SLIT and NTRK-like protein 1 precursor. | |||||
|
CD302_HUMAN
|
||||||
| θ value | θ > 10 (rank : 52) | NC score | 0.068926 (rank : 22) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8IX05, Q15009 | Gene names | CD302, CLEC13A, DCL1, KIAA0022 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CD302 antigen precursor (C-type lectin domain family 13 member A) (C- type lectin BIMLEC) (Type I transmembrane C-type lectin receptor DCL- 1). | |||||
|
CD302_MOUSE
|
||||||
| θ value | θ > 10 (rank : 53) | NC score | 0.068272 (rank : 23) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9DCG2, Q78KD8, Q9D0X7 | Gene names | Cd302, Clec13a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CD302 antigen precursor (C-type lectin domain family 13 member A) (Type I transmembrane C-type lectin receptor DCL-1). | |||||
|
CHODL_HUMAN
|
||||||
| θ value | θ > 10 (rank : 54) | NC score | 0.052624 (rank : 33) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9H9P2, Q7Z798, Q7Z799, Q7Z7A0, Q9HCY3 | Gene names | CHODL, C21orf68 | |||
|
Domain Architecture |
|
|||||
| Description | Chondrolectin precursor (Transmembrane protein MT75). | |||||
|
CHODL_MOUSE
|
||||||
| θ value | θ > 10 (rank : 55) | NC score | 0.052704 (rank : 32) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9CXM0, Q8VI31 | Gene names | Chodl | |||
|
Domain Architecture |
|
|||||
| Description | Chondrolectin precursor (Transmembrane protein MT75). | |||||
|
MMP10_HUMAN
|
||||||
| θ value | θ > 10 (rank : 56) | NC score | 0.051750 (rank : 36) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P09238 | Gene names | MMP10, STMY2 | |||
|
Domain Architecture |

|
|||||
| Description | Stromelysin-2 precursor (EC 3.4.24.22) (Matrix metalloproteinase-10) (MMP-10) (Transin-2) (SL-2). | |||||
|
MMP10_MOUSE
|
||||||
| θ value | θ > 10 (rank : 57) | NC score | 0.052435 (rank : 35) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O55123 | Gene names | Mmp10 | |||
|
Domain Architecture |
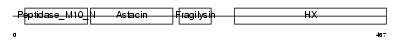
|
|||||
| Description | Stromelysin-2 precursor (EC 3.4.24.22) (Matrix metalloproteinase-10) (MMP-10) (Transin-2) (SL-2). | |||||
|
MMP11_HUMAN
|
||||||
| θ value | θ > 10 (rank : 58) | NC score | 0.052821 (rank : 31) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P24347, Q5FX24, Q6PEZ6, Q9UC26 | Gene names | MMP11, STMY3 | |||
|
Domain Architecture |

|
|||||
| Description | Stromelysin-3 precursor (EC 3.4.24.-) (ST3) (SL-3) (Matrix metalloproteinase-11) (MMP-11). | |||||
|
MMP11_MOUSE
|
||||||
| θ value | θ > 10 (rank : 59) | NC score | 0.053021 (rank : 30) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q02853 | Gene names | Mmp11 | |||
|
Domain Architecture |

|
|||||
| Description | Stromelysin-3 precursor (EC 3.4.24.-) (ST3) (SL-3) (Matrix metalloproteinase-11) (MMP-11). | |||||
|
MMP12_MOUSE
|
||||||
| θ value | θ > 10 (rank : 60) | NC score | 0.050724 (rank : 41) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P34960, Q8BJ92, Q8BJB3, Q8BJB6, Q8BJB8, Q8BJB9, Q8VED6 | Gene names | Mmp12, Mme, Mmel | |||
|
Domain Architecture |

|
|||||
| Description | Macrophage metalloelastase precursor (EC 3.4.24.65) (MME) (Matrix metalloproteinase-12) (MMP-12). | |||||
|
MMP13_HUMAN
|
||||||
| θ value | θ > 10 (rank : 61) | NC score | 0.053140 (rank : 29) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P45452 | Gene names | MMP13 | |||
|
Domain Architecture |

|
|||||
| Description | Collagenase 3 precursor (EC 3.4.24.-) (Matrix metalloproteinase-13) (MMP-13). | |||||
|
MMP13_MOUSE
|
||||||
| θ value | θ > 10 (rank : 62) | NC score | 0.052614 (rank : 34) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P33435 | Gene names | Mmp13 | |||
|
Domain Architecture |

|
|||||
| Description | Collagenase 3 precursor (EC 3.4.24.-) (Matrix metalloproteinase-13) (MMP-13). | |||||
|
MMP1A_MOUSE
|
||||||
| θ value | θ > 10 (rank : 63) | NC score | 0.051111 (rank : 40) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9EPL5 | Gene names | Mmp1a, McolA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interstitial collagenase A precursor (EC 3.4.24.7) (Matrix metalloproteinase-1a) (MMP-1a) (Mcol-A). | |||||
|
MMP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 64) | NC score | 0.051683 (rank : 37) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P03956, P08156 | Gene names | MMP1, CLG | |||
|
Domain Architecture |

|
|||||
| Description | Interstitial collagenase precursor (EC 3.4.24.7) (Matrix metalloproteinase-1) (MMP-1) (Fibroblast collagenase) [Contains: 22 kDa interstitial collagenase; 27 kDa interstitial collagenase]. | |||||
|
MMP20_HUMAN
|
||||||
| θ value | θ > 10 (rank : 65) | NC score | 0.050082 (rank : 43) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O60882, Q6DKT9 | Gene names | MMP20 | |||
|
Domain Architecture |

|
|||||
| Description | Matrix metalloproteinase-20 precursor (EC 3.4.24.-) (MMP-20) (Enamel metalloproteinase) (Enamelysin). | |||||
|
MMP20_MOUSE
|
||||||
| θ value | θ > 10 (rank : 66) | NC score | 0.050138 (rank : 42) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P57748 | Gene names | Mmp20 | |||
|
Domain Architecture |

|
|||||
| Description | Matrix metalloproteinase-20 precursor (EC 3.4.24.-) (MMP-20) (Enamel metalloproteinase) (Enamelysin). | |||||
|
MMP26_HUMAN
|
||||||
| θ value | θ > 10 (rank : 67) | NC score | 0.055188 (rank : 28) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NRE1, Q9GZS2, Q9NR87 | Gene names | MMP26 | |||
|
Domain Architecture |
|
|||||
| Description | Matrix metalloproteinase-26 precursor (EC 3.4.24.-) (MMP-26) (Matrilysin-2) (Endometase). | |||||
|
MMP3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 68) | NC score | 0.051487 (rank : 38) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P08254, Q3B7S0 | Gene names | MMP3, STMY1 | |||
|
Domain Architecture |

|
|||||
| Description | Stromelysin-1 precursor (EC 3.4.24.17) (Matrix metalloproteinase-3) (MMP-3) (Transin-1) (SL-1). | |||||
|
MMP3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 69) | NC score | 0.051440 (rank : 39) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P28862 | Gene names | Mmp3 | |||
|
Domain Architecture |

|
|||||
| Description | Stromelysin-1 precursor (EC 3.4.24.17) (Matrix metalloproteinase-3) (MMP-3) (Transin-1) (SL-1) (EMS-2). | |||||
|
MMP7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 70) | NC score | 0.059618 (rank : 25) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P09237, Q9BTK9 | Gene names | MMP7, MPSL1, PUMP1 | |||
|
Domain Architecture |
|
|||||
| Description | Matrilysin precursor (EC 3.4.24.23) (Pump-1 protease) (Uterine metalloproteinase) (Matrix metalloproteinase-7) (MMP-7) (Matrin). | |||||
|
MMP7_MOUSE
|
||||||
| θ value | θ > 10 (rank : 71) | NC score | 0.059718 (rank : 24) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q10738 | Gene names | Mmp7 | |||
|
Domain Architecture |
|
|||||
| Description | Matrilysin precursor (EC 3.4.24.23) (Pump-1 protease) (Uterine metalloproteinase) (Matrix metalloproteinase-7) (MMP-7) (Matrin). | |||||
|
MMP8_MOUSE
|
||||||
| θ value | θ > 10 (rank : 72) | NC score | 0.050018 (rank : 44) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O70138, O88733 | Gene names | Mmp8 | |||
|
Domain Architecture |

|
|||||
| Description | Neutrophil collagenase precursor (EC 3.4.24.34) (Matrix metalloproteinase-8) (MMP-8) (Collagenase 2). | |||||
|
MPRI_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 51 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | Q07113, Q61822, Q6LED1 | Gene names | Igf2r | |||
|
Domain Architecture |

|
|||||
| Description | Cation-independent mannose-6-phosphate receptor precursor (CI Man-6-P receptor) (CI-MPR) (M6PR) (Insulin-like growth factor 2 receptor) (Insulin-like growth factor II receptor) (IGF-II receptor) (M6P/IGF2 receptor) (M6P/IGF2R) (300 kDa mannose 6-phosphate receptor) (MPR 300) (MPR300). | |||||
|
MPRI_HUMAN
|
||||||
| NC score | 0.992617 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P11717, Q7Z7G9, Q96PT5 | Gene names | IGF2R, MPRI | |||
|
Domain Architecture |

|
|||||
| Description | Cation-independent mannose-6-phosphate receptor precursor (CI Man-6-P receptor) (CI-MPR) (M6PR) (Insulin-like growth factor 2 receptor) (Insulin-like growth factor II receptor) (IGF-II receptor) (M6P/IGF2 receptor) (M6P/IGF2R) (300 kDa mannose 6-phosphate receptor) (MPR 300) (MPR300) (CD222 antigen). | |||||
|
SEL1L_HUMAN
|
||||||
| NC score | 0.445356 (rank : 3) | θ value | 9.59137e-10 (rank : 7) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9UBV2, Q6UWT6, Q9P1T9, Q9UHK7 | Gene names | SEL1L, TSA305 | |||
|
Domain Architecture |

|
|||||
| Description | Sel-1 homolog precursor (Suppressor of lin-12-like protein) (Sel-1L). | |||||
|
SEL1L_MOUSE
|
||||||
| NC score | 0.442330 (rank : 4) | θ value | 9.59137e-10 (rank : 8) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9Z2G6, Q9DBD8 | Gene names | Sel1l, Sel1h | |||
|
Domain Architecture |

|
|||||
| Description | Sel-1 homolog precursor (Suppressor of lin-12-like protein) (Sel-1L). | |||||
|
MMP9_MOUSE
|
||||||
| NC score | 0.290683 (rank : 5) | θ value | 4.30538e-10 (rank : 5) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P41245, Q06788, Q9DC02 | Gene names | Mmp9, Clg4b | |||
|
Domain Architecture |

|
|||||
| Description | Matrix metalloproteinase-9 precursor (EC 3.4.24.35) (MMP-9) (92 kDa type IV collagenase) (92 kDa gelatinase) (Gelatinase B) (GELB). | |||||
|
MMP9_HUMAN
|
||||||
| NC score | 0.290011 (rank : 6) | θ value | 4.30538e-10 (rank : 4) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 125 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P14780, Q3LR70, Q8N725, Q9H4Z1 | Gene names | MMP9, CLG4B | |||
|
Domain Architecture |

|
|||||
| Description | Matrix metalloproteinase-9 precursor (EC 3.4.24.35) (MMP-9) (92 kDa type IV collagenase) (92 kDa gelatinase) (Gelatinase B) (GELB) [Contains: 67 kDa matrix metalloproteinase-9; 82 kDa matrix metalloproteinase-9]. | |||||
|
MMP2_HUMAN
|
||||||
| NC score | 0.267652 (rank : 7) | θ value | 1.47974e-10 (rank : 3) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 79 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P08253 | Gene names | MMP2, CLG4A | |||
|
Domain Architecture |

|
|||||
| Description | 72 kDa type IV collagenase precursor (EC 3.4.24.24) (72 kDa gelatinase) (Matrix metalloproteinase-2) (MMP-2) (Gelatinase A) (TBE- 1). | |||||
|
MMP2_MOUSE
|
||||||
| NC score | 0.264661 (rank : 8) | θ value | 7.34386e-10 (rank : 6) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P33434 | Gene names | Mmp2 | |||
|
Domain Architecture |

|
|||||
| Description | 72 kDa type IV collagenase precursor (EC 3.4.24.24) (72 kDa gelatinase) (Matrix metalloproteinase-2) (MMP-2) (Gelatinase A). | |||||
|
LY75_HUMAN
|
||||||
| NC score | 0.257914 (rank : 9) | θ value | 6.43352e-06 (rank : 14) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 169 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O60449, O75913, Q7Z575, Q7Z577 | Gene names | LY75, CD205 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphocyte antigen 75 precursor (DEC-205) (gp200-MR6) (CD205 antigen). | |||||
|
MRC1_MOUSE
|
||||||
| NC score | 0.257779 (rank : 10) | θ value | 1.87187e-05 (rank : 17) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q61830, Q8C502 | Gene names | Mrc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Macrophage mannose receptor 1 precursor (MMR) (CD206 antigen). | |||||
|
MRC2_MOUSE
|
||||||
| NC score | 0.250426 (rank : 11) | θ value | 1.43324e-05 (rank : 16) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 159 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q64449, Q6ZQ64, Q8C6P0 | Gene names | Mrc2, Kiaa0709 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Macrophage mannose receptor 2 precursor (Lectin lambda) (CD280 antigen). | |||||
|
MRC2_HUMAN
|
||||||
| NC score | 0.249512 (rank : 12) | θ value | 1.43324e-05 (rank : 15) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 162 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9UBG0, Q7LGE7, Q9Y5P9 | Gene names | MRC2, ENDO180, KIAA0709, UPARAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Macrophage mannose receptor 2 precursor (Urokinase receptor-associated protein) (Endocytic receptor 180) (CD280 antigen). | |||||
|
MRC1_HUMAN
|
||||||
| NC score | 0.248968 (rank : 13) | θ value | 7.1131e-05 (rank : 18) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 157 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P22897 | Gene names | MRC1 | |||
|
Domain Architecture |

|
|||||
| Description | Macrophage mannose receptor 1 precursor (MMR) (CD206 antigen). | |||||
|
LY75_MOUSE
|
||||||
| NC score | 0.247984 (rank : 14) | θ value | 0.000461057 (rank : 19) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q60767, Q8C7T3, Q91XL8, Q9QUZ6 | Gene names | Ly75, Cd205 | |||
|
Domain Architecture |

|
|||||
| Description | Lymphocyte antigen 75 precursor (DEC-205) (CD205 antigen). | |||||
|
MPRD_MOUSE
|
||||||
| NC score | 0.228771 (rank : 15) | θ value | 0.000786445 (rank : 20) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 4 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P24668 | Gene names | M6pr, 46mpr | |||
|
Domain Architecture |
|
|||||
| Description | Cation-dependent mannose-6-phosphate receptor precursor (CD Man-6-P receptor) (CD-MPR) (46 kDa mannose 6-phosphate receptor) (MPR 46). | |||||
|
FINC_HUMAN
|
||||||
| NC score | 0.225829 (rank : 16) | θ value | 8.11959e-09 (rank : 11) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 302 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P02751, O95609, O95610, Q14312, Q14325, Q14326, Q6LDP6, Q86T27, Q8IVI8, Q96KP7, Q96KP8, Q96KP9, Q9H1B8, Q9HAP3, Q9UMK2 | Gene names | FN1, FN | |||
|
Domain Architecture |

|
|||||
| Description | Fibronectin precursor (FN) (Cold-insoluble globulin) (CIG). | |||||
|
FINC_MOUSE
|
||||||
| NC score | 0.225370 (rank : 17) | θ value | 8.11959e-09 (rank : 12) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 309 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P11276, Q61567, Q61568, Q61569, Q64233, Q80UI4 | Gene names | Fn1 | |||
|
Domain Architecture |

|
|||||
| Description | Fibronectin precursor (FN). | |||||
|
MPRD_HUMAN
|
||||||
| NC score | 0.217054 (rank : 18) | θ value | 0.00665767 (rank : 21) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P20645 | Gene names | M6PR, MPR46, MPRD | |||
|
Domain Architecture |
|
|||||
| Description | Cation-dependent mannose-6-phosphate receptor precursor (CD Man-6-P receptor) (CD-MPR) (46 kDa mannose 6-phosphate receptor) (MPR 46). | |||||
|
HGFA_HUMAN
|
||||||
| NC score | 0.125975 (rank : 19) | θ value | 2.79066e-09 (rank : 9) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 436 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q04756, Q14726 | Gene names | HGFAC | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte growth factor activator precursor (EC 3.4.21.-) (HGF activator) (HGFA) [Contains: Hepatocyte growth factor activator short chain; Hepatocyte growth factor activator long chain]. | |||||
|
HGFA_MOUSE
|
||||||
| NC score | 0.125099 (rank : 20) | θ value | 4.76016e-09 (rank : 10) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9R098, Q9JKV4 | Gene names | Hgfac | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte growth factor activator precursor (EC 3.4.21.-) (HGF activator) (HGFA) [Contains: Hepatocyte growth factor activator short chain; Hepatocyte growth factor activator long chain]. | |||||
|
FA12_HUMAN
|
||||||
| NC score | 0.120787 (rank : 21) | θ value | 4.45536e-07 (rank : 13) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P00748, P78339 | Gene names | F12 | |||
|
Domain Architecture |

|
|||||
| Description | Coagulation factor XII precursor (EC 3.4.21.38) (Hageman factor) (HAF) [Contains: Coagulation factor XIIa heavy chain; Beta-factor XIIa part 1; Beta-factor XIIa part 2; Coagulation factor XIIa light chain]. | |||||
|
CD302_HUMAN
|
||||||
| NC score | 0.068926 (rank : 22) | θ value | θ > 10 (rank : 52) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8IX05, Q15009 | Gene names | CD302, CLEC13A, DCL1, KIAA0022 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CD302 antigen precursor (C-type lectin domain family 13 member A) (C- type lectin BIMLEC) (Type I transmembrane C-type lectin receptor DCL- 1). | |||||
|
CD302_MOUSE
|
||||||
| NC score | 0.068272 (rank : 23) | θ value | θ > 10 (rank : 53) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9DCG2, Q78KD8, Q9D0X7 | Gene names | Cd302, Clec13a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CD302 antigen precursor (C-type lectin domain family 13 member A) (Type I transmembrane C-type lectin receptor DCL-1). | |||||
|
MMP7_MOUSE
|
||||||
| NC score | 0.059718 (rank : 24) | θ value | θ > 10 (rank : 71) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q10738 | Gene names | Mmp7 | |||
|
Domain Architecture |

|
|||||
| Description | Matrilysin precursor (EC 3.4.24.23) (Pump-1 protease) (Uterine metalloproteinase) (Matrix metalloproteinase-7) (MMP-7) (Matrin). | |||||
|
MMP7_HUMAN
|
||||||
| NC score | 0.059618 (rank : 25) | θ value | θ > 10 (rank : 70) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P09237, Q9BTK9 | Gene names | MMP7, MPSL1, PUMP1 | |||
|
Domain Architecture |

|
|||||
| Description | Matrilysin precursor (EC 3.4.24.23) (Pump-1 protease) (Uterine metalloproteinase) (Matrix metalloproteinase-7) (MMP-7) (Matrin). | |||||
|
GLU2B_HUMAN
|
||||||
| NC score | 0.059118 (rank : 26) | θ value | 0.365318 (rank : 23) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 377 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P14314, Q96BU9, Q96D06, Q9P0W9 | Gene names | PRKCSH, G19P1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glucosidase 2 subunit beta precursor (Glucosidase II subunit beta) (Protein kinase C substrate, 60.1 kDa protein, heavy chain) (PKCSH) (80K-H protein). | |||||
|
GLU2B_MOUSE
|
||||||
| NC score | 0.055677 (rank : 27) | θ value | 0.62314 (rank : 25) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 357 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O08795, Q921X2 | Gene names | Prkcsh | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glucosidase 2 subunit beta precursor (Glucosidase II subunit beta) (Protein kinase C substrate, 60.1 kDa protein, heavy chain) (PKCSH) (80K-H protein). | |||||
|
MMP26_HUMAN
|
||||||
| NC score | 0.055188 (rank : 28) | θ value | θ > 10 (rank : 67) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NRE1, Q9GZS2, Q9NR87 | Gene names | MMP26 | |||
|
Domain Architecture |

|
|||||
| Description | Matrix metalloproteinase-26 precursor (EC 3.4.24.-) (MMP-26) (Matrilysin-2) (Endometase). | |||||
|
MMP13_HUMAN
|
||||||
| NC score | 0.053140 (rank : 29) | θ value | θ > 10 (rank : 61) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P45452 | Gene names | MMP13 | |||
|
Domain Architecture |

|
|||||
| Description | Collagenase 3 precursor (EC 3.4.24.-) (Matrix metalloproteinase-13) (MMP-13). | |||||
|
MMP11_MOUSE
|
||||||
| NC score | 0.053021 (rank : 30) | θ value | θ > 10 (rank : 59) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q02853 | Gene names | Mmp11 | |||
|
Domain Architecture |

|
|||||
| Description | Stromelysin-3 precursor (EC 3.4.24.-) (ST3) (SL-3) (Matrix metalloproteinase-11) (MMP-11). | |||||
|
MMP11_HUMAN
|
||||||
| NC score | 0.052821 (rank : 31) | θ value | θ > 10 (rank : 58) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P24347, Q5FX24, Q6PEZ6, Q9UC26 | Gene names | MMP11, STMY3 | |||
|
Domain Architecture |

|
|||||
| Description | Stromelysin-3 precursor (EC 3.4.24.-) (ST3) (SL-3) (Matrix metalloproteinase-11) (MMP-11). | |||||
|
CHODL_MOUSE
|
||||||
| NC score | 0.052704 (rank : 32) | θ value | θ > 10 (rank : 55) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9CXM0, Q8VI31 | Gene names | Chodl | |||
|
Domain Architecture |

|
|||||
| Description | Chondrolectin precursor (Transmembrane protein MT75). | |||||
|
CHODL_HUMAN
|
||||||
| NC score | 0.052624 (rank : 33) | θ value | θ > 10 (rank : 54) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9H9P2, Q7Z798, Q7Z799, Q7Z7A0, Q9HCY3 | Gene names | CHODL, C21orf68 | |||
|
Domain Architecture |

|
|||||
| Description | Chondrolectin precursor (Transmembrane protein MT75). | |||||
|
MMP13_MOUSE
|
||||||
| NC score | 0.052614 (rank : 34) | θ value | θ > 10 (rank : 62) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P33435 | Gene names | Mmp13 | |||
|
Domain Architecture |

|
|||||
| Description | Collagenase 3 precursor (EC 3.4.24.-) (Matrix metalloproteinase-13) (MMP-13). | |||||
|
MMP10_MOUSE
|
||||||
| NC score | 0.052435 (rank : 35) | θ value | θ > 10 (rank : 57) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O55123 | Gene names | Mmp10 | |||
|
Domain Architecture |

|
|||||
| Description | Stromelysin-2 precursor (EC 3.4.24.22) (Matrix metalloproteinase-10) (MMP-10) (Transin-2) (SL-2). | |||||
|
MMP10_HUMAN
|
||||||
| NC score | 0.051750 (rank : 36) | θ value | θ > 10 (rank : 56) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P09238 | Gene names | MMP10, STMY2 | |||
|
Domain Architecture |

|
|||||
| Description | Stromelysin-2 precursor (EC 3.4.24.22) (Matrix metalloproteinase-10) (MMP-10) (Transin-2) (SL-2). | |||||
|
MMP1_HUMAN
|
||||||
| NC score | 0.051683 (rank : 37) | θ value | θ > 10 (rank : 64) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P03956, P08156 | Gene names | MMP1, CLG | |||
|
Domain Architecture |

|
|||||
| Description | Interstitial collagenase precursor (EC 3.4.24.7) (Matrix metalloproteinase-1) (MMP-1) (Fibroblast collagenase) [Contains: 22 kDa interstitial collagenase; 27 kDa interstitial collagenase]. | |||||
|
MMP3_HUMAN
|
||||||
| NC score | 0.051487 (rank : 38) | θ value | θ > 10 (rank : 68) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P08254, Q3B7S0 | Gene names | MMP3, STMY1 | |||
|
Domain Architecture |

|
|||||
| Description | Stromelysin-1 precursor (EC 3.4.24.17) (Matrix metalloproteinase-3) (MMP-3) (Transin-1) (SL-1). | |||||
|
MMP3_MOUSE
|
||||||
| NC score | 0.051440 (rank : 39) | θ value | θ > 10 (rank : 69) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P28862 | Gene names | Mmp3 | |||
|
Domain Architecture |

|
|||||
| Description | Stromelysin-1 precursor (EC 3.4.24.17) (Matrix metalloproteinase-3) (MMP-3) (Transin-1) (SL-1) (EMS-2). | |||||
|
MMP1A_MOUSE
|
||||||
| NC score | 0.051111 (rank : 40) | θ value | θ > 10 (rank : 63) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9EPL5 | Gene names | Mmp1a, McolA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Interstitial collagenase A precursor (EC 3.4.24.7) (Matrix metalloproteinase-1a) (MMP-1a) (Mcol-A). | |||||
|
MMP12_MOUSE
|
||||||
| NC score | 0.050724 (rank : 41) | θ value | θ > 10 (rank : 60) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P34960, Q8BJ92, Q8BJB3, Q8BJB6, Q8BJB8, Q8BJB9, Q8VED6 | Gene names | Mmp12, Mme, Mmel | |||
|
Domain Architecture |

|
|||||
| Description | Macrophage metalloelastase precursor (EC 3.4.24.65) (MME) (Matrix metalloproteinase-12) (MMP-12). | |||||
|
MMP20_MOUSE
|
||||||
| NC score | 0.050138 (rank : 42) | θ value | θ > 10 (rank : 66) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P57748 | Gene names | Mmp20 | |||
|
Domain Architecture |

|
|||||
| Description | Matrix metalloproteinase-20 precursor (EC 3.4.24.-) (MMP-20) (Enamel metalloproteinase) (Enamelysin). | |||||
|
MMP20_HUMAN
|
||||||
| NC score | 0.050082 (rank : 43) | θ value | θ > 10 (rank : 65) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O60882, Q6DKT9 | Gene names | MMP20 | |||
|
Domain Architecture |

|
|||||
| Description | Matrix metalloproteinase-20 precursor (EC 3.4.24.-) (MMP-20) (Enamel metalloproteinase) (Enamelysin). | |||||
|
MMP8_MOUSE
|
||||||
| NC score | 0.050018 (rank : 44) | θ value | θ > 10 (rank : 72) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O70138, O88733 | Gene names | Mmp8 | |||
|
Domain Architecture |

|
|||||
| Description | Neutrophil collagenase precursor (EC 3.4.24.34) (Matrix metalloproteinase-8) (MMP-8) (Collagenase 2). | |||||
|
RBFAL_HUMAN
|
||||||
| NC score | 0.034110 (rank : 45) | θ value | 1.81305 (rank : 31) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8N0V3, Q6PF07, Q8WZ65, Q9H776 | Gene names | C18orf22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative ribosome-binding factor A, mitochondrial precursor. | |||||
|
LRBA_HUMAN
|
||||||
| NC score | 0.027677 (rank : 46) | θ value | 0.365318 (rank : 24) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P50851, Q9H2U3, Q9H2U4 | Gene names | LRBA, BGL, CDC4L, LBA | |||
|
Domain Architecture |

|
|||||
| Description | Lipopolysaccharide-responsive and beige-like anchor protein (CDC4-like protein) (Beige-like protein). | |||||
|
RUSC2_HUMAN
|
||||||
| NC score | 0.025956 (rank : 47) | θ value | 3.0926 (rank : 36) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 205 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8N2Y8, O15080, Q6P1W7 | Gene names | RUSC2, KIAA0375 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RUN and SH3 domain-containing protein 2. | |||||
|
DLG7_MOUSE
|
||||||
| NC score | 0.025229 (rank : 48) | θ value | 1.81305 (rank : 30) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 84 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8K4R9, Q6ZQL3, Q80VS7, Q8BUC8, Q8BZ79 | Gene names | Dlg7, Kiaa0008 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Discs large homolog 7 (Hepatoma up-regulated protein homolog) (HURP). | |||||
|
NOTC4_HUMAN
|
||||||
| NC score | 0.023943 (rank : 49) | θ value | 0.163984 (rank : 22) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 905 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q99466, O00306, Q99458, Q99940, Q9H3S8, Q9UII9, Q9UIJ0 | Gene names | NOTCH4 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 4 precursor (Notch 4) (hNotch4) [Contains: Notch 4 extracellular truncation; Notch 4 intracellular domain]. | |||||
|
TNR8_MOUSE
|
||||||
| NC score | 0.023378 (rank : 50) | θ value | 2.36792 (rank : 35) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q60846 | Gene names | Tnfrsf8 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 8 precursor (CD30L receptor) (Lymphocyte activation antigen CD30). | |||||
|
CUBN_MOUSE
|
||||||
| NC score | 0.019034 (rank : 51) | θ value | 4.03905 (rank : 37) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 477 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9JLB4 | Gene names | Cubn, Ifcr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cubilin precursor (Intrinsic factor-cobalamin receptor). | |||||
|
MUG1_MOUSE
|
||||||
| NC score | 0.018621 (rank : 52) | θ value | 0.813845 (rank : 27) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P28665 | Gene names | Mug1, Mug-1 | |||
|
Domain Architecture |

|
|||||
| Description | Murinoglobulin-1 precursor (MuG1). | |||||
|
GRP78_MOUSE
|
||||||
| NC score | 0.016601 (rank : 53) | θ value | 2.36792 (rank : 34) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P20029, O35642, Q3UFF2, Q61630 | Gene names | Hspa5, Grp78 | |||
|
Domain Architecture |

|
|||||
| Description | 78 kDa glucose-regulated protein precursor (GRP 78) (Heat shock 70 kDa protein 5) (Immunoglobulin heavy chain-binding protein) (BiP). | |||||
|
GRP78_HUMAN
|
||||||
| NC score | 0.016595 (rank : 54) | θ value | 2.36792 (rank : 33) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 218 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P11021, Q2EF78, Q9NPF1, Q9UK02 | Gene names | HSPA5, GRP78 | |||
|
Domain Architecture |

|
|||||
| Description | 78 kDa glucose-regulated protein precursor (GRP 78) (Heat shock 70 kDa protein 5) (Immunoglobulin heavy chain-binding protein) (BiP) (Endoplasmic reticulum lumenal Ca(2+)-binding protein grp78). | |||||
|
ZADH1_HUMAN
|
||||||
| NC score | 0.015908 (rank : 55) | θ value | 5.27518 (rank : 40) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8N8N7, Q6MZH8 | Gene names | ZADH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc-binding alcohol dehydrogenase domain-containing protein 1 (EC 1.-.-.-). | |||||
|
P52K_MOUSE
|
||||||
| NC score | 0.015298 (rank : 56) | θ value | 5.27518 (rank : 39) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9CUX1, Q80Y58 | Gene names | Prkrir, Thap0 | |||
|
Domain Architecture |

|
|||||
| Description | 52 kDa repressor of the inhibitor of the protein kinase (p58IPK- interacting protein) (58 kDa interferon-induced protein kinase- interacting protein) (P52rIPK) (THAP domain-containing protein 0). | |||||
|
CUBN_HUMAN
|
||||||
| NC score | 0.015090 (rank : 57) | θ value | 8.99809 (rank : 48) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 482 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O60494, Q5VTA6, Q96RU9 | Gene names | CUBN, IFCR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cubilin precursor (Intrinsic factor-cobalamin receptor) (Intrinsic factor-vitamin B12 receptor) (460 kDa receptor) (Intestinal intrinsic factor receptor). | |||||
|
RBM19_MOUSE
|
||||||
| NC score | 0.014203 (rank : 58) | θ value | 1.38821 (rank : 29) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 296 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8R3C6, Q8BHR0, Q9CW63 | Gene names | Rbm19 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable RNA-binding protein 19 (RNA-binding motif protein 19). | |||||
|
NELL1_HUMAN
|
||||||
| NC score | 0.013817 (rank : 59) | θ value | 6.88961 (rank : 44) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 451 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q92832, Q9Y472 | Gene names | NELL1, NRP1 | |||
|
Domain Architecture |

|
|||||
| Description | Protein kinase C-binding protein NELL1 precursor (NEL-like protein 1) (Nel-related protein 1). | |||||
|
NAR3_MOUSE
|
||||||
| NC score | 0.013333 (rank : 60) | θ value | 6.88961 (rank : 43) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8R2G4, Q8R2F9, Q8R2G0, Q8R2G1, Q8R2G2, Q8R2G3 | Gene names | Art3 | |||
|
Domain Architecture |

|
|||||
| Description | Ecto-ADP-ribosyltransferase 3 precursor (EC 2.4.2.31) (NAD(P)(+)-- arginine ADP-ribosyltransferase 3) (Mono(ADP-ribosyl)transferase 3). | |||||
|
TF3C1_HUMAN
|
||||||
| NC score | 0.012771 (rank : 61) | θ value | 6.88961 (rank : 46) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q12789, Q12838, Q9Y4W9 | Gene names | GTF3C1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | General transcription factor 3C polypeptide 1 (Transcription factor IIIC-subunit alpha) (TF3C-alpha) (TFIIIC 220 kDa subunit) (TFIIIC220) (TFIIIC box B-binding subunit). | |||||
|
CABYR_HUMAN
|
||||||
| NC score | 0.012672 (rank : 62) | θ value | 8.99809 (rank : 47) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O75952, Q8WXW5, Q9HAY3, Q9HAY4, Q9HAY5, Q9HCY9 | Gene names | CABYR, CBP86, FSP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calcium-binding tyrosine phosphorylation-regulated protein (Calcium- binding protein 86) (Testis-specific calcium-binding protein CBP86) (Fibrousheathin-2) (FSP-2). | |||||
|
BTNL8_HUMAN
|
||||||
| NC score | 0.012499 (rank : 63) | θ value | 1.06291 (rank : 28) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 177 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6UX41, Q9H730 | Gene names | BTNL8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Butyrophilin-like protein 8 precursor. | |||||
|
LRP2_HUMAN
|
||||||
| NC score | 0.011456 (rank : 64) | θ value | 6.88961 (rank : 42) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 573 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P98164, O00711, Q16215 | Gene names | LRP2 | |||
|
Domain Architecture |

|
|||||
| Description | Low-density lipoprotein receptor-related protein 2 precursor (Megalin) (Glycoprotein 330) (gp330). | |||||
|
SMYD1_HUMAN
|
||||||
| NC score | 0.011080 (rank : 65) | θ value | 6.88961 (rank : 45) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8NB12 | Gene names | SMYD1 | |||
|
Domain Architecture |

|
|||||
| Description | SET and MYND domain-containing protein 1. | |||||
|
CAD23_MOUSE
|
||||||
| NC score | 0.009320 (rank : 66) | θ value | 0.813845 (rank : 26) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 208 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q99PF4, Q99NH1, Q9D4N9 | Gene names | Cdh23 | |||
|
Domain Architecture |

|
|||||
| Description | Cadherin-23 precursor (Otocadherin). | |||||
|
CAF1A_HUMAN
|
||||||
| NC score | 0.009239 (rank : 67) | θ value | 6.88961 (rank : 41) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 750 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q13111, Q7Z7K3, Q9UJY8 | Gene names | CHAF1A, CAF, CAF1P150 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromatin assembly factor 1 subunit A (CAF-1 subunit A) (Chromatin assembly factor I p150 subunit) (CAF-I 150 kDa subunit) (CAF-Ip150). | |||||
|
ATS20_HUMAN
|
||||||
| NC score | 0.008420 (rank : 68) | θ value | 2.36792 (rank : 32) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 192 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P59510 | Gene names | ADAMTS20 | |||
|
Domain Architecture |

|
|||||
| Description | ADAMTS-20 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase with thrombospondin motifs 20) (ADAM-TS 20) (ADAM-TS20). | |||||
|
ATS12_MOUSE
|
||||||
| NC score | 0.006770 (rank : 69) | θ value | 5.27518 (rank : 38) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 220 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q811B3, Q8BK92, Q8BKY1 | Gene names | Adamts12 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADAMTS-12 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase with thrombospondin motifs 12) (ADAM-TS 12) (ADAM-TS12). | |||||
|
RIMB1_MOUSE
|
||||||
| NC score | 0.006390 (rank : 70) | θ value | 8.99809 (rank : 49) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 1376 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q7TNF8, Q5NCP7, Q80TV9, Q8BIH5 | Gene names | Bzrap1, Kiaa0612, Rbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Peripheral-type benzodiazepine receptor-associated protein 1 (PRAX-1) (Peripheral benzodiazepine receptor-interacting protein) (PBR-IP) (RIM-binding protein 1) (RIM-BP1). | |||||
|
SEM4F_MOUSE
|
||||||
| NC score | 0.003601 (rank : 71) | θ value | 8.99809 (rank : 50) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Z123, Q9R1Y1 | Gene names | Sema4f, Semaw | |||
|
Domain Architecture |

|
|||||
| Description | Semaphorin-4F precursor (Semaphorin W) (Sema W). | |||||
|
SLIK1_MOUSE
|
||||||
| NC score | 0.002372 (rank : 72) | θ value | 8.99809 (rank : 51) | |||
| Query Neighborhood Hits | 51 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q810C1, Q9CXL0 | Gene names | Slitrk1, Sltk1 | |||
|
Domain Architecture |

|
|||||
| Description | SLIT and NTRK-like protein 1 precursor. | |||||