Please be patient as the page loads
|
KAD7_MOUSE
|
||||||
| SwissProt Accessions | Q9D2H2, Q8BVH3 | Gene names | Ak7 | |||
|
Domain Architecture |

|
|||||
| Description | Putative adenylate kinase 7 (EC 2.7.4.3). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
KAD7_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.982719 (rank : 2) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q96M32, Q8IYP6 | Gene names | AK7 | |||
|
Domain Architecture |

|
|||||
| Description | Putative adenylate kinase 7 (EC 2.7.4.3). | |||||
|
KAD7_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9D2H2, Q8BVH3 | Gene names | Ak7 | |||
|
Domain Architecture |

|
|||||
| Description | Putative adenylate kinase 7 (EC 2.7.4.3). | |||||
|
DPY30_MOUSE
|
||||||
| θ value | 1.80886e-08 (rank : 3) | NC score | 0.524249 (rank : 3) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q99LT0 | Gene names | ||||
|
Domain Architecture |
|
|||||
| Description | Dpy-30-like protein. | |||||
|
DPY30_HUMAN
|
||||||
| θ value | 2.36244e-08 (rank : 4) | NC score | 0.521092 (rank : 4) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9C005 | Gene names | ||||
|
Domain Architecture |
|
|||||
| Description | Dpy-30-like protein. | |||||
|
NDK5_HUMAN
|
||||||
| θ value | 3.41135e-07 (rank : 5) | NC score | 0.273337 (rank : 7) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P56597 | Gene names | NME5 | |||
|
Domain Architecture |
|
|||||
| Description | Nucleoside diphosphate kinase homolog 5 (NDK-H 5) (NDP kinase homolog 5) (nm23-H5) (Testis-specific nm23 homolog) (Inhibitor of p53-induced apoptosis-beta) (IPIA-beta). | |||||
|
KAD2_MOUSE
|
||||||
| θ value | 7.59969e-07 (rank : 6) | NC score | 0.294311 (rank : 5) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9WTP6, Q3TKI6, Q9CY37 | Gene names | Ak2, Ak-2 | |||
|
Domain Architecture |
|
|||||
| Description | Adenylate kinase isoenzyme 2, mitochondrial (EC 2.7.4.3) (ATP-AMP transphosphorylase). | |||||
|
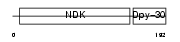
NDK5_MOUSE
|
||||||
| θ value | 7.59969e-07 (rank : 7) | NC score | 0.272616 (rank : 8) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q99MH5, Q810R1 | Gene names | Nme5 | |||
|
Domain Architecture |
|
|||||
| Description | Nucleoside diphosphate kinase homolog 5 (NDK-H 5) (NDP kinase homolog 5) (nm23-M5). | |||||
|
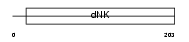
KAD2_HUMAN
|
||||||
| θ value | 1.69304e-06 (rank : 8) | NC score | 0.288040 (rank : 6) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P54819, Q16856, Q5TIF7, Q8TCY2, Q8TCY3 | Gene names | AK2, ADK2 | |||
|
Domain Architecture |
|
|||||
| Description | Adenylate kinase isoenzyme 2, mitochondrial (EC 2.7.4.3) (ATP-AMP transphosphorylase). | |||||
|
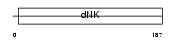
KAD4_HUMAN
|
||||||
| θ value | 0.00134147 (rank : 9) | NC score | 0.240214 (rank : 9) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P27144 | Gene names | AK3L1, AK3, AK4 | |||
|
Domain Architecture |
|
|||||
| Description | Adenylate kinase isoenzyme 4, mitochondrial (EC 2.7.4.3) (Adenylate kinase 3-like 1) (ATP-AMP transphosphorylase). | |||||
|
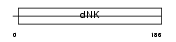
KAD4_MOUSE
|
||||||
| θ value | 0.00509761 (rank : 10) | NC score | 0.227803 (rank : 10) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9WUR9, Q9R1X7 | Gene names | Ak3l1, Ak-4, Ak3b, Ak4 | |||
|
Domain Architecture |
|
|||||
| Description | Adenylate kinase isoenzyme 4, mitochondrial (EC 2.7.4.3) (Adenylate kinase 3-like 1) (ATP-AMP transphosphorylase). | |||||
|
DYDC1_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 11) | NC score | 0.220150 (rank : 11) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9D9T0, Q810Q6 | Gene names | Dydc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DPY30 domain-containing protein 1. | |||||
|
KAD3_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 12) | NC score | 0.195000 (rank : 12) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9WTP7, Q3UDN7, Q8BGX5, Q9D7Z1, Q9DB57, Q9DBM5 | Gene names | Ak3, Ak3l, Ak3l1, Akl3l | |||
|
Domain Architecture |
|
|||||
| Description | GTP:AMP phosphotransferase mitochondrial (EC 2.7.4.10) (Adenylate kinase 3) (AK3) (Adenylate kinase 3 alpha-like 1). | |||||
|
DYDC1_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 13) | NC score | 0.183669 (rank : 13) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8WWB3, Q5QP03, Q5QP04, Q6WNP4, Q6ZU87 | Gene names | DYDC1, DPY30D1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DPY30 domain-containing protein 1 (RSD-9). | |||||
|
STMN1_HUMAN
|
||||||
| θ value | 0.0961366 (rank : 14) | NC score | 0.086304 (rank : 22) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 554 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P16949 | Gene names | STMN1, LAP18, OP18 | |||
|
Domain Architecture |

|
|||||
| Description | Stathmin (Phosphoprotein p19) (pp19) (Oncoprotein 18) (Op18) (Leukemia-associated phosphoprotein p18) (pp17) (Prosolin) (Metablastin) (Protein Pr22). | |||||
|
STMN1_MOUSE
|
||||||
| θ value | 0.125558 (rank : 15) | NC score | 0.084919 (rank : 23) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 560 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P54227 | Gene names | Stmn1, Lag, Lap18, Pr22 | |||
|
Domain Architecture |

|
|||||
| Description | Stathmin (Phosphoprotein p19) (pp19) (Oncoprotein 18) (Op18) (Leukemia-associated phosphoprotein p18) (pp17) (Prosolin) (Metablastin) (Pr22 protein) (Leukemia-associated gene protein). | |||||
|
BPA1_MOUSE
|
||||||
| θ value | 0.21417 (rank : 16) | NC score | 0.030885 (rank : 42) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1477 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q91ZU6, Q91ZU7 | Gene names | Dst, Bpag1 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoforms 1/2/3/4 (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
DYDC2_HUMAN
|
||||||
| θ value | 0.21417 (rank : 17) | NC score | 0.162553 (rank : 14) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96IM9, Q5QP07, Q5QP11 | Gene names | DYDC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DPY30 domain-containing protein 2. | |||||
|
PEPL_HUMAN
|
||||||
| θ value | 0.21417 (rank : 18) | NC score | 0.043370 (rank : 35) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1141 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O60437, O60314, O60454 | Gene names | PPL, KIAA0568 | |||
|
Domain Architecture |
|
|||||
| Description | Periplakin (195 kDa cornified envelope precursor protein) (190 kDa paraneoplastic pemphigus antigen). | |||||
|
KI13B_HUMAN
|
||||||
| θ value | 0.47712 (rank : 19) | NC score | 0.020502 (rank : 53) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 492 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9NQT8, O75134, Q9BYJ6 | Gene names | KIF13B, GAKIN, KIAA0639 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF13B (Kinesin-like protein GAKIN). | |||||
|
PLVAP_HUMAN
|
||||||
| θ value | 0.62314 (rank : 20) | NC score | 0.040021 (rank : 36) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9BX97, Q86VP0, Q8N8Y0, Q8ND68, Q8TER8, Q9BZD5 | Gene names | PLVAP, FELS, PV1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plasmalemma vesicle-associated protein (Plasmalemma vesicle protein 1) (PV-1) (Fenestrated endothelial-linked structure protein). | |||||
|
AP1B1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 21) | NC score | 0.033254 (rank : 39) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q10567, P78436 | Gene names | AP1B1, ADTB1, BAM22 | |||
|
Domain Architecture |

|
|||||
| Description | AP-1 complex subunit beta-1 (Adapter-related protein complex 1 beta-1 subunit) (Beta-adaptin 1) (Adaptor protein complex AP-1 beta-1 subunit) (Golgi adaptor HA1/AP1 adaptin beta subunit) (Clathrin assembly protein complex 1 beta large chain). | |||||
|
AP1B1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 22) | NC score | 0.033150 (rank : 40) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O35643, Q922E2 | Gene names | Ap1b1, Adtb1 | |||
|
Domain Architecture |

|
|||||
| Description | AP-1 complex subunit beta-1 (Adapter-related protein complex 1 beta-1 subunit) (Beta-adaptin 1) (Adaptor protein complex AP-1 beta-1 subunit) (Golgi adaptor HA1/AP1 adaptin beta subunit) (Clathrin assembly protein complex 1 beta large chain). | |||||
|
KAD5_MOUSE
|
||||||
| θ value | 1.06291 (rank : 23) | NC score | 0.132610 (rank : 16) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q920P5 | Gene names | Ak5 | |||
|
Domain Architecture |
|
|||||
| Description | Adenylate kinase isoenzyme 5 (EC 2.7.4.3) (ATP-AMP transphosphorylase). | |||||
|
LONM_HUMAN
|
||||||
| θ value | 1.06291 (rank : 24) | NC score | 0.058545 (rank : 26) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P36776, P36777, Q9UQ95 | Gene names | PRSS15 | |||
|
Domain Architecture |

|
|||||
| Description | Lon protease homolog, mitochondrial precursor (EC 3.4.21.-) (Lon protease-like protein) (LONP) (LONHs). | |||||
|
DDX56_MOUSE
|
||||||
| θ value | 1.38821 (rank : 25) | NC score | 0.015929 (rank : 60) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9D0R4 | Gene names | Ddx56, D11Ertd619e, Noh61 | |||
|
Domain Architecture |

|
|||||
| Description | Probable ATP-dependent RNA helicase DDX56 (EC 3.6.1.-) (DEAD box protein 56) (ATP-dependent 61 kDa nucleolar RNA helicase). | |||||
|
IAG2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 26) | NC score | 0.038572 (rank : 37) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 5 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H0U3, Q53G00, Q8NBN6 | Gene names | IAG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Implantation-associated protein precursor (IAP) (Magnesium transporter protein 1) (MagT1). | |||||
|
AR13B_HUMAN
|
||||||
| θ value | 3.0926 (rank : 27) | NC score | 0.018597 (rank : 54) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 528 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q3SXY8, Q504W8 | Gene names | ARL13B, ARL2L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADP-ribosylation factor-like protein 13B (ADP-ribosylation factor-like protein 2-like 1) (ARL2-like protein 1). | |||||
|
BPAEA_HUMAN
|
||||||
| θ value | 3.0926 (rank : 28) | NC score | 0.029445 (rank : 44) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1369 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O94833, Q5TBT0, Q8N1T8, Q8N8J3, Q8WXK9, Q96AK9, Q96DQ5, Q9H555 | Gene names | DST, BPAG1, DMH, DT, KIAA0728 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoforms 6/9/10 (Trabeculin-beta) (Bullous pemphigoid antigen) (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
CDC37_HUMAN
|
||||||
| θ value | 3.0926 (rank : 29) | NC score | 0.044843 (rank : 34) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q16543 | Gene names | CDC37 | |||
|
Domain Architecture |
|
|||||
| Description | Hsp90 co-chaperone Cdc37 (Hsp90 chaperone protein kinase-targeting subunit) (p50Cdc37). | |||||
|
LRRF2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 30) | NC score | 0.021460 (rank : 51) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 556 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q91WK0, Q8BVD1, Q9CU89 | Gene names | Lrrfip2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat flightless-interacting protein 2 (LRR FLII- interacting protein 2). | |||||
|
TRM6_MOUSE
|
||||||
| θ value | 3.0926 (rank : 31) | NC score | 0.028583 (rank : 47) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CE96, Q3TJZ8, Q3TME7, Q6ZPW8, Q80Y59, Q8CEU0 | Gene names | Trm6, Kiaa1153 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | tRNA (adenine-N(1)-)-methyltransferase non-catalytic subunit TRM6 (tRNA(m1A58)-methyltransferase subunit TRM6) (tRNA(m1A58)MTase subunit TRM6). | |||||
|
CE290_HUMAN
|
||||||
| θ value | 4.03905 (rank : 32) | NC score | 0.018325 (rank : 55) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1553 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O15078, Q1PSK5, Q66GS8, Q9H2G6, Q9H6Q7, Q9H8I0 | Gene names | CEP290, KIAA0373, NPHP6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein Cep290 (Nephrocystin-6) (Tumor antigen se2-2). | |||||
|
CT077_HUMAN
|
||||||
| θ value | 4.03905 (rank : 33) | NC score | 0.037486 (rank : 38) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NQG5 | Gene names | C20orf77 | |||
|
Domain Architecture |

|
|||||
| Description | Uncharacterized protein C20orf77. | |||||
|
GEMI_MOUSE
|
||||||
| θ value | 4.03905 (rank : 34) | NC score | 0.059410 (rank : 24) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O88513 | Gene names | Gmnn | |||
|
Domain Architecture |

|
|||||
| Description | Geminin. | |||||
|
KAD6_HUMAN
|
||||||
| θ value | 4.03905 (rank : 35) | NC score | 0.046334 (rank : 32) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y3D8 | Gene names | AK6 | |||
|
Domain Architecture |
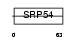
|
|||||
| Description | Adenylate kinase isoenzyme 6 (EC 2.7.4.3) (ATP-AMP transphosphorylase 6). | |||||
|
PEPL_MOUSE
|
||||||
| θ value | 4.03905 (rank : 36) | NC score | 0.032077 (rank : 41) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1105 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9R269, O70231, Q9CUT1, Q9JLZ7 | Gene names | Ppl | |||
|
Domain Architecture |

|
|||||
| Description | Periplakin. | |||||
|
RBCC1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 37) | NC score | 0.017259 (rank : 57) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1048 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9ESK9, Q3TXX2, Q61384, Q8BRY9, Q9JK14 | Gene names | Rb1cc1, Cc1, Kiaa0203 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RB1-inducible coiled-coil protein 1 (Coiled-coil-forming protein 1) (LaXp180 protein). | |||||
|
ZN232_HUMAN
|
||||||
| θ value | 4.03905 (rank : 38) | NC score | -0.000245 (rank : 65) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 754 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UNY5 | Gene names | ZNF232, ZSCAN11 | |||
|
Domain Architecture |
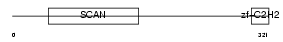
|
|||||
| Description | Zinc finger protein 232 (Zinc finger and SCAN domain-containing protein 11). | |||||
|
ZN638_HUMAN
|
||||||
| θ value | 4.03905 (rank : 39) | NC score | 0.020816 (rank : 52) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q14966, Q53R34, Q5XJ05, Q68DP3, Q6P2H2, Q7Z3T7, Q8NF92, Q8TCA1, Q9H2G1, Q9NP37 | Gene names | ZNF638, NP220, ZFML | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 638 (Nuclear protein 220) (Zinc-finger matrin-like protein) (Cutaneous T-cell lymphoma-associated antigen se33-1) (CTCL tumor antigen se33-1). | |||||
|
GEMI_HUMAN
|
||||||
| θ value | 5.27518 (rank : 40) | NC score | 0.059143 (rank : 25) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 192 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O75496, Q9H1Z1 | Gene names | GMNN | |||
|
Domain Architecture |

|
|||||
| Description | Geminin. | |||||
|
SM1L2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 41) | NC score | 0.023530 (rank : 49) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1019 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8NDV3, Q5TIC3, Q9Y3G5 | Gene names | SMC1L2 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 1-like 2 protein (SMC1beta protein). | |||||
|
CLH1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 42) | NC score | 0.028597 (rank : 45) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q00610, Q6N0A0, Q86TF2 | Gene names | CLTC, CLH17, KIAA0034 | |||
|
Domain Architecture |

|
|||||
| Description | Clathrin heavy chain 1 (CLH-17). | |||||
|
CLH_MOUSE
|
||||||
| θ value | 6.88961 (rank : 43) | NC score | 0.028593 (rank : 46) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q68FD5 | Gene names | Cltc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Clathrin heavy chain. | |||||
|
IAG2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 44) | NC score | 0.030505 (rank : 43) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9CQY5, Q3UW45, Q9CWX5, Q9CZT3 | Gene names | Iag2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Implantation-associated protein precursor (IAP) (Magnesium transporter protein 1) (MagT1). | |||||
|
KAD1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 45) | NC score | 0.109149 (rank : 17) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P00568, Q9BVK9, Q9UQC7 | Gene names | AK1 | |||
|
Domain Architecture |
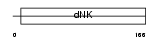
|
|||||
| Description | Adenylate kinase isoenzyme 1 (EC 2.7.4.3) (ATP-AMP transphosphorylase) (AK1) (Myokinase). | |||||
|
NDUA5_MOUSE
|
||||||
| θ value | 6.88961 (rank : 46) | NC score | 0.055151 (rank : 29) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9CPP6, Q9CY90, Q9D2P2, Q9D703, Q9D739 | Gene names | Ndufa5 | |||
|
Domain Architecture |
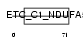
|
|||||
| Description | NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 5 (EC 1.6.5.3) (EC 1.6.99.3) (NADH-ubiquinone oxidoreductase 13 kDa-B subunit) (Complex I-13kD-B) (CI-13kD-B) (Complex I subunit B13). | |||||
|
NPAS2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 47) | NC score | 0.008944 (rank : 62) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P97460 | Gene names | Npas2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 2 (Neuronal PAS2). | |||||
|
NUF2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 48) | NC score | 0.017524 (rank : 56) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 585 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q99P69, Q8BTK7, Q8VE05, Q9CST5 | Gene names | Cdca1, Nuf2r | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinetochore protein Nuf2 (Cell division cycle-associated protein 1). | |||||
|
S4A7_HUMAN
|
||||||
| θ value | 6.88961 (rank : 49) | NC score | 0.007821 (rank : 63) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y6M7, O60350, Q6AHZ9, Q9HC88, Q9UIB9 | Gene names | SLC4A7, BT, NBC2, NBC2B, NBC3, SBC2, SLC4A6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium bicarbonate cotransporter 3 (Sodium bicarbonate cotransporter 2) (Sodium bicarbonate cotransporter 2b) (Bicarbonate transporter) (Solute carrier family 4 member 7). | |||||
|
SMC1A_HUMAN
|
||||||
| θ value | 6.88961 (rank : 50) | NC score | 0.023705 (rank : 48) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1346 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q14683, O14995, Q16351 | Gene names | SMC1L1, DXS423E, KIAA0178, SB1.8, SMC1 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 1-like 1 protein (SMC1alpha protein) (Sb1.8). | |||||
|
SMC1A_MOUSE
|
||||||
| θ value | 6.88961 (rank : 51) | NC score | 0.023023 (rank : 50) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1384 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9CU62, Q3V480, Q9CUX9, Q9D959, Q9WTU1 | Gene names | Smc1l1, Sb1.8, Smc1, Smcb | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 1-like 1 protein (SMC1alpha protein) (Chromosome segregation protein SmcB) (Sb1.8). | |||||
|
ABCB9_MOUSE
|
||||||
| θ value | 8.99809 (rank : 52) | NC score | 0.004481 (rank : 64) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9JJ59, Q8CHA1, Q9D212, Q9JIN1 | Gene names | Abcb9, Kiaa1520 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-binding cassette sub-family B member 9 precursor (ATP-binding cassette transporter 9) (ABC transporter 9 protein) (TAP-like protein) (TAPL) (mABCB9). | |||||
|
KIF5A_HUMAN
|
||||||
| θ value | 8.99809 (rank : 53) | NC score | 0.013691 (rank : 61) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 843 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q12840, Q4LE26 | Gene names | KIF5A, NKHC1 | |||
|
Domain Architecture |
|
|||||
| Description | Kinesin heavy chain isoform 5A (Neuronal kinesin heavy chain) (NKHC) (Kinesin heavy chain neuron-specific 1). | |||||
|
NDUA5_HUMAN
|
||||||
| θ value | 8.99809 (rank : 54) | NC score | 0.045732 (rank : 33) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q16718, Q6IRX7 | Gene names | NDUFA5 | |||
|
Domain Architecture |
|
|||||
| Description | NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 5 (EC 1.6.5.3) (EC 1.6.99.3) (NADH-ubiquinone oxidoreductase 13 kDa-B subunit) (Complex I-13kD-B) (CI-13kD-B) (Complex I subunit B13). | |||||
|
PICAL_HUMAN
|
||||||
| θ value | 8.99809 (rank : 55) | NC score | 0.016604 (rank : 58) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q13492, O60700, Q86XZ9 | Gene names | PICALM, CALM | |||
|
Domain Architecture |
|
|||||
| Description | Phosphatidylinositol-binding clathrin assembly protein (Clathrin assembly lymphoid myeloid leukemia protein). | |||||
|
PICA_MOUSE
|
||||||
| θ value | 8.99809 (rank : 56) | NC score | 0.016573 (rank : 59) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7M6Y3, Q3TS04, Q811P1, Q8BUF6, Q8CIH8, Q8R0A9, Q8R3E1, Q8VDN5, Q921L0 | Gene names | Picalm, Calm, Fit1 | |||
|
Domain Architecture |
|
|||||
| Description | Phosphatidylinositol-binding clathrin assembly protein (Clathrin assembly lymphoid myeloid leukemia) (CALM). | |||||
|
KAD1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 57) | NC score | 0.105116 (rank : 18) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9R0Y5 | Gene names | Ak1 | |||
|
Domain Architecture |
|
|||||
| Description | Adenylate kinase isoenzyme 1 (EC 2.7.4.3) (ATP-AMP transphosphorylase) (AK1) (Myokinase). | |||||
|
KAD3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 58) | NC score | 0.158280 (rank : 15) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UIJ7, Q5VYW6, Q9H576, Q9NPB4 | Gene names | AK3, AK3L1, AKL3L | |||
|
Domain Architecture |
|
|||||
| Description | GTP:AMP phosphotransferase mitochondrial (EC 2.7.4.10) (Adenylate kinase 3) (AK3) (Adenylate kinase 3 alpha-like 1). | |||||
|
KAD5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 59) | NC score | 0.097796 (rank : 19) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Y6K8, Q96EC9 | Gene names | AK5 | |||
|
Domain Architecture |
|
|||||
| Description | Adenylate kinase isoenzyme 5 (EC 2.7.4.3) (ATP-AMP transphosphorylase). | |||||
|
KCY_HUMAN
|
||||||
| θ value | θ > 10 (rank : 60) | NC score | 0.095303 (rank : 21) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P30085, Q53GB7, Q5SVZ0, Q96C07, Q9UBQ8, Q9UIA2 | Gene names | CMPK, CMK, UCK, UMK, UMPK | |||
|
Domain Architecture |
|
|||||
| Description | UMP-CMP kinase (EC 2.7.4.14) (Cytidylate kinase) (Deoxycytidylate kinase) (Cytidine monophosphate kinase) (Uridine monophosphate/cytidine monophosphate kinase) (UMP/CMP kinase) (UMP/CMPK) (Uridine monophosphate kinase). | |||||
|
KCY_MOUSE
|
||||||
| θ value | θ > 10 (rank : 61) | NC score | 0.095579 (rank : 20) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9DBP5 | Gene names | Cmpk, Cmk, Uck, Umk, Umpk | |||
|
Domain Architecture |
|
|||||
| Description | UMP-CMP kinase (EC 2.7.4.14) (Cytidylate kinase) (Deoxycytidylate kinase) (Cytidine monophosphate kinase) (Uridine monophosphate/cytidine monophosphate kinase) (UMP/CMP kinase) (UMP/CMPK) (Uridine monophosphate kinase). | |||||
|
NDK6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 62) | NC score | 0.056654 (rank : 27) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O75414, Q96E73, Q9BQ63 | Gene names | NME6 | |||
|
Domain Architecture |
|
|||||
| Description | Nucleoside diphosphate kinase 6 (EC 2.7.4.6) (NDK 6) (NDP kinase 6) (nm23-H6) (Inhibitor of p53-induced apoptosis-alpha) (IPIA-alpha). | |||||
|
NDK6_MOUSE
|
||||||
| θ value | θ > 10 (rank : 63) | NC score | 0.056645 (rank : 28) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O88425, Q99M53 | Gene names | Nme6 | |||
|
Domain Architecture |
|
|||||
| Description | Nucleoside diphosphate kinase 6 (EC 2.7.4.6) (NDK 6) (NDP kinase 6) (nm23-M6). | |||||
|
NDK7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 64) | NC score | 0.053523 (rank : 30) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y5B8, Q5TGZ4 | Gene names | NME7 | |||
|
Domain Architecture |
|
|||||
| Description | Nucleoside diphosphate kinase 7 (EC 2.7.4.6) (NDK 7) (NDP kinase 7) (nm23-H7). | |||||
|
NDK7_MOUSE
|
||||||
| θ value | θ > 10 (rank : 65) | NC score | 0.053022 (rank : 31) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9QXL8 | Gene names | Nme7 | |||
|
Domain Architecture |
|
|||||
| Description | Nucleoside diphosphate kinase 7 (EC 2.7.4.6) (NDK 7) (NDP kinase 7) (nm23-M7). | |||||
|
KAD7_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 56 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9D2H2, Q8BVH3 | Gene names | Ak7 | |||
|
Domain Architecture |

|
|||||
| Description | Putative adenylate kinase 7 (EC 2.7.4.3). | |||||
|
KAD7_HUMAN
|
||||||
| NC score | 0.982719 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q96M32, Q8IYP6 | Gene names | AK7 | |||
|
Domain Architecture |

|
|||||
| Description | Putative adenylate kinase 7 (EC 2.7.4.3). | |||||
|
DPY30_MOUSE
|
||||||
| NC score | 0.524249 (rank : 3) | θ value | 1.80886e-08 (rank : 3) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q99LT0 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | Dpy-30-like protein. | |||||
|
DPY30_HUMAN
|
||||||
| NC score | 0.521092 (rank : 4) | θ value | 2.36244e-08 (rank : 4) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9C005 | Gene names | ||||
|
Domain Architecture |

|
|||||
| Description | Dpy-30-like protein. | |||||
|
KAD2_MOUSE
|
||||||
| NC score | 0.294311 (rank : 5) | θ value | 7.59969e-07 (rank : 6) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9WTP6, Q3TKI6, Q9CY37 | Gene names | Ak2, Ak-2 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylate kinase isoenzyme 2, mitochondrial (EC 2.7.4.3) (ATP-AMP transphosphorylase). | |||||
|
KAD2_HUMAN
|
||||||
| NC score | 0.288040 (rank : 6) | θ value | 1.69304e-06 (rank : 8) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P54819, Q16856, Q5TIF7, Q8TCY2, Q8TCY3 | Gene names | AK2, ADK2 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylate kinase isoenzyme 2, mitochondrial (EC 2.7.4.3) (ATP-AMP transphosphorylase). | |||||
|
NDK5_HUMAN
|
||||||
| NC score | 0.273337 (rank : 7) | θ value | 3.41135e-07 (rank : 5) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P56597 | Gene names | NME5 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoside diphosphate kinase homolog 5 (NDK-H 5) (NDP kinase homolog 5) (nm23-H5) (Testis-specific nm23 homolog) (Inhibitor of p53-induced apoptosis-beta) (IPIA-beta). | |||||
|
NDK5_MOUSE
|
||||||
| NC score | 0.272616 (rank : 8) | θ value | 7.59969e-07 (rank : 7) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q99MH5, Q810R1 | Gene names | Nme5 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoside diphosphate kinase homolog 5 (NDK-H 5) (NDP kinase homolog 5) (nm23-M5). | |||||
|
KAD4_HUMAN
|
||||||
| NC score | 0.240214 (rank : 9) | θ value | 0.00134147 (rank : 9) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P27144 | Gene names | AK3L1, AK3, AK4 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylate kinase isoenzyme 4, mitochondrial (EC 2.7.4.3) (Adenylate kinase 3-like 1) (ATP-AMP transphosphorylase). | |||||
|
KAD4_MOUSE
|
||||||
| NC score | 0.227803 (rank : 10) | θ value | 0.00509761 (rank : 10) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9WUR9, Q9R1X7 | Gene names | Ak3l1, Ak-4, Ak3b, Ak4 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylate kinase isoenzyme 4, mitochondrial (EC 2.7.4.3) (Adenylate kinase 3-like 1) (ATP-AMP transphosphorylase). | |||||
|
DYDC1_MOUSE
|
||||||
| NC score | 0.220150 (rank : 11) | θ value | 0.0330416 (rank : 11) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9D9T0, Q810Q6 | Gene names | Dydc1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DPY30 domain-containing protein 1. | |||||
|
KAD3_MOUSE
|
||||||
| NC score | 0.195000 (rank : 12) | θ value | 0.0736092 (rank : 12) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9WTP7, Q3UDN7, Q8BGX5, Q9D7Z1, Q9DB57, Q9DBM5 | Gene names | Ak3, Ak3l, Ak3l1, Akl3l | |||
|
Domain Architecture |

|
|||||
| Description | GTP:AMP phosphotransferase mitochondrial (EC 2.7.4.10) (Adenylate kinase 3) (AK3) (Adenylate kinase 3 alpha-like 1). | |||||
|
DYDC1_HUMAN
|
||||||
| NC score | 0.183669 (rank : 13) | θ value | 0.0961366 (rank : 13) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8WWB3, Q5QP03, Q5QP04, Q6WNP4, Q6ZU87 | Gene names | DYDC1, DPY30D1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DPY30 domain-containing protein 1 (RSD-9). | |||||
|
DYDC2_HUMAN
|
||||||
| NC score | 0.162553 (rank : 14) | θ value | 0.21417 (rank : 17) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q96IM9, Q5QP07, Q5QP11 | Gene names | DYDC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DPY30 domain-containing protein 2. | |||||
|
KAD3_HUMAN
|
||||||
| NC score | 0.158280 (rank : 15) | θ value | θ > 10 (rank : 58) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UIJ7, Q5VYW6, Q9H576, Q9NPB4 | Gene names | AK3, AK3L1, AKL3L | |||
|
Domain Architecture |

|
|||||
| Description | GTP:AMP phosphotransferase mitochondrial (EC 2.7.4.10) (Adenylate kinase 3) (AK3) (Adenylate kinase 3 alpha-like 1). | |||||
|
KAD5_MOUSE
|
||||||
| NC score | 0.132610 (rank : 16) | θ value | 1.06291 (rank : 23) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q920P5 | Gene names | Ak5 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylate kinase isoenzyme 5 (EC 2.7.4.3) (ATP-AMP transphosphorylase). | |||||
|
KAD1_HUMAN
|
||||||
| NC score | 0.109149 (rank : 17) | θ value | 6.88961 (rank : 45) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P00568, Q9BVK9, Q9UQC7 | Gene names | AK1 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylate kinase isoenzyme 1 (EC 2.7.4.3) (ATP-AMP transphosphorylase) (AK1) (Myokinase). | |||||
|
KAD1_MOUSE
|
||||||
| NC score | 0.105116 (rank : 18) | θ value | θ > 10 (rank : 57) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9R0Y5 | Gene names | Ak1 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylate kinase isoenzyme 1 (EC 2.7.4.3) (ATP-AMP transphosphorylase) (AK1) (Myokinase). | |||||
|
KAD5_HUMAN
|
||||||
| NC score | 0.097796 (rank : 19) | θ value | θ > 10 (rank : 59) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9Y6K8, Q96EC9 | Gene names | AK5 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylate kinase isoenzyme 5 (EC 2.7.4.3) (ATP-AMP transphosphorylase). | |||||
|
KCY_MOUSE
|
||||||
| NC score | 0.095579 (rank : 20) | θ value | θ > 10 (rank : 61) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9DBP5 | Gene names | Cmpk, Cmk, Uck, Umk, Umpk | |||
|
Domain Architecture |

|
|||||
| Description | UMP-CMP kinase (EC 2.7.4.14) (Cytidylate kinase) (Deoxycytidylate kinase) (Cytidine monophosphate kinase) (Uridine monophosphate/cytidine monophosphate kinase) (UMP/CMP kinase) (UMP/CMPK) (Uridine monophosphate kinase). | |||||
|
KCY_HUMAN
|
||||||
| NC score | 0.095303 (rank : 21) | θ value | θ > 10 (rank : 60) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P30085, Q53GB7, Q5SVZ0, Q96C07, Q9UBQ8, Q9UIA2 | Gene names | CMPK, CMK, UCK, UMK, UMPK | |||
|
Domain Architecture |

|
|||||
| Description | UMP-CMP kinase (EC 2.7.4.14) (Cytidylate kinase) (Deoxycytidylate kinase) (Cytidine monophosphate kinase) (Uridine monophosphate/cytidine monophosphate kinase) (UMP/CMP kinase) (UMP/CMPK) (Uridine monophosphate kinase). | |||||
|
STMN1_HUMAN
|
||||||
| NC score | 0.086304 (rank : 22) | θ value | 0.0961366 (rank : 14) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 554 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P16949 | Gene names | STMN1, LAP18, OP18 | |||
|
Domain Architecture |

|
|||||
| Description | Stathmin (Phosphoprotein p19) (pp19) (Oncoprotein 18) (Op18) (Leukemia-associated phosphoprotein p18) (pp17) (Prosolin) (Metablastin) (Protein Pr22). | |||||
|
STMN1_MOUSE
|
||||||
| NC score | 0.084919 (rank : 23) | θ value | 0.125558 (rank : 15) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 560 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P54227 | Gene names | Stmn1, Lag, Lap18, Pr22 | |||
|
Domain Architecture |

|
|||||
| Description | Stathmin (Phosphoprotein p19) (pp19) (Oncoprotein 18) (Op18) (Leukemia-associated phosphoprotein p18) (pp17) (Prosolin) (Metablastin) (Pr22 protein) (Leukemia-associated gene protein). | |||||
|
GEMI_MOUSE
|
||||||
| NC score | 0.059410 (rank : 24) | θ value | 4.03905 (rank : 34) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O88513 | Gene names | Gmnn | |||
|
Domain Architecture |

|
|||||
| Description | Geminin. | |||||
|
GEMI_HUMAN
|
||||||
| NC score | 0.059143 (rank : 25) | θ value | 5.27518 (rank : 40) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 192 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O75496, Q9H1Z1 | Gene names | GMNN | |||
|
Domain Architecture |

|
|||||
| Description | Geminin. | |||||
|
LONM_HUMAN
|
||||||
| NC score | 0.058545 (rank : 26) | θ value | 1.06291 (rank : 24) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P36776, P36777, Q9UQ95 | Gene names | PRSS15 | |||
|
Domain Architecture |

|
|||||
| Description | Lon protease homolog, mitochondrial precursor (EC 3.4.21.-) (Lon protease-like protein) (LONP) (LONHs). | |||||
|
NDK6_HUMAN
|
||||||
| NC score | 0.056654 (rank : 27) | θ value | θ > 10 (rank : 62) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O75414, Q96E73, Q9BQ63 | Gene names | NME6 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoside diphosphate kinase 6 (EC 2.7.4.6) (NDK 6) (NDP kinase 6) (nm23-H6) (Inhibitor of p53-induced apoptosis-alpha) (IPIA-alpha). | |||||
|
NDK6_MOUSE
|
||||||
| NC score | 0.056645 (rank : 28) | θ value | θ > 10 (rank : 63) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O88425, Q99M53 | Gene names | Nme6 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoside diphosphate kinase 6 (EC 2.7.4.6) (NDK 6) (NDP kinase 6) (nm23-M6). | |||||
|
NDUA5_MOUSE
|
||||||
| NC score | 0.055151 (rank : 29) | θ value | 6.88961 (rank : 46) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9CPP6, Q9CY90, Q9D2P2, Q9D703, Q9D739 | Gene names | Ndufa5 | |||
|
Domain Architecture |

|
|||||
| Description | NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 5 (EC 1.6.5.3) (EC 1.6.99.3) (NADH-ubiquinone oxidoreductase 13 kDa-B subunit) (Complex I-13kD-B) (CI-13kD-B) (Complex I subunit B13). | |||||
|
NDK7_HUMAN
|
||||||
| NC score | 0.053523 (rank : 30) | θ value | θ > 10 (rank : 64) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y5B8, Q5TGZ4 | Gene names | NME7 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoside diphosphate kinase 7 (EC 2.7.4.6) (NDK 7) (NDP kinase 7) (nm23-H7). | |||||
|
NDK7_MOUSE
|
||||||
| NC score | 0.053022 (rank : 31) | θ value | θ > 10 (rank : 65) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9QXL8 | Gene names | Nme7 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleoside diphosphate kinase 7 (EC 2.7.4.6) (NDK 7) (NDP kinase 7) (nm23-M7). | |||||
|
KAD6_HUMAN
|
||||||
| NC score | 0.046334 (rank : 32) | θ value | 4.03905 (rank : 35) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y3D8 | Gene names | AK6 | |||
|
Domain Architecture |

|
|||||
| Description | Adenylate kinase isoenzyme 6 (EC 2.7.4.3) (ATP-AMP transphosphorylase 6). | |||||
|
NDUA5_HUMAN
|
||||||
| NC score | 0.045732 (rank : 33) | θ value | 8.99809 (rank : 54) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q16718, Q6IRX7 | Gene names | NDUFA5 | |||
|
Domain Architecture |

|
|||||
| Description | NADH dehydrogenase [ubiquinone] 1 alpha subcomplex subunit 5 (EC 1.6.5.3) (EC 1.6.99.3) (NADH-ubiquinone oxidoreductase 13 kDa-B subunit) (Complex I-13kD-B) (CI-13kD-B) (Complex I subunit B13). | |||||
|
CDC37_HUMAN
|
||||||
| NC score | 0.044843 (rank : 34) | θ value | 3.0926 (rank : 29) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q16543 | Gene names | CDC37 | |||
|
Domain Architecture |

|
|||||
| Description | Hsp90 co-chaperone Cdc37 (Hsp90 chaperone protein kinase-targeting subunit) (p50Cdc37). | |||||
|
PEPL_HUMAN
|
||||||
| NC score | 0.043370 (rank : 35) | θ value | 0.21417 (rank : 18) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1141 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O60437, O60314, O60454 | Gene names | PPL, KIAA0568 | |||
|
Domain Architecture |

|
|||||
| Description | Periplakin (195 kDa cornified envelope precursor protein) (190 kDa paraneoplastic pemphigus antigen). | |||||
|
PLVAP_HUMAN
|
||||||
| NC score | 0.040021 (rank : 36) | θ value | 0.62314 (rank : 20) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9BX97, Q86VP0, Q8N8Y0, Q8ND68, Q8TER8, Q9BZD5 | Gene names | PLVAP, FELS, PV1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Plasmalemma vesicle-associated protein (Plasmalemma vesicle protein 1) (PV-1) (Fenestrated endothelial-linked structure protein). | |||||
|
IAG2_HUMAN
|
||||||
| NC score | 0.038572 (rank : 37) | θ value | 1.81305 (rank : 26) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 5 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H0U3, Q53G00, Q8NBN6 | Gene names | IAG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Implantation-associated protein precursor (IAP) (Magnesium transporter protein 1) (MagT1). | |||||
|
CT077_HUMAN
|
||||||
| NC score | 0.037486 (rank : 38) | θ value | 4.03905 (rank : 33) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9NQG5 | Gene names | C20orf77 | |||
|
Domain Architecture |

|
|||||
| Description | Uncharacterized protein C20orf77. | |||||
|
AP1B1_HUMAN
|
||||||
| NC score | 0.033254 (rank : 39) | θ value | 0.813845 (rank : 21) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q10567, P78436 | Gene names | AP1B1, ADTB1, BAM22 | |||
|
Domain Architecture |

|
|||||
| Description | AP-1 complex subunit beta-1 (Adapter-related protein complex 1 beta-1 subunit) (Beta-adaptin 1) (Adaptor protein complex AP-1 beta-1 subunit) (Golgi adaptor HA1/AP1 adaptin beta subunit) (Clathrin assembly protein complex 1 beta large chain). | |||||
|
AP1B1_MOUSE
|
||||||
| NC score | 0.033150 (rank : 40) | θ value | 0.813845 (rank : 22) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 59 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O35643, Q922E2 | Gene names | Ap1b1, Adtb1 | |||
|
Domain Architecture |

|
|||||
| Description | AP-1 complex subunit beta-1 (Adapter-related protein complex 1 beta-1 subunit) (Beta-adaptin 1) (Adaptor protein complex AP-1 beta-1 subunit) (Golgi adaptor HA1/AP1 adaptin beta subunit) (Clathrin assembly protein complex 1 beta large chain). | |||||
|
PEPL_MOUSE
|
||||||
| NC score | 0.032077 (rank : 41) | θ value | 4.03905 (rank : 36) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1105 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9R269, O70231, Q9CUT1, Q9JLZ7 | Gene names | Ppl | |||
|
Domain Architecture |

|
|||||
| Description | Periplakin. | |||||
|
BPA1_MOUSE
|
||||||
| NC score | 0.030885 (rank : 42) | θ value | 0.21417 (rank : 16) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1477 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q91ZU6, Q91ZU7 | Gene names | Dst, Bpag1 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoforms 1/2/3/4 (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
IAG2_MOUSE
|
||||||
| NC score | 0.030505 (rank : 43) | θ value | 6.88961 (rank : 44) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9CQY5, Q3UW45, Q9CWX5, Q9CZT3 | Gene names | Iag2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Implantation-associated protein precursor (IAP) (Magnesium transporter protein 1) (MagT1). | |||||
|
BPAEA_HUMAN
|
||||||
| NC score | 0.029445 (rank : 44) | θ value | 3.0926 (rank : 28) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1369 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O94833, Q5TBT0, Q8N1T8, Q8N8J3, Q8WXK9, Q96AK9, Q96DQ5, Q9H555 | Gene names | DST, BPAG1, DMH, DT, KIAA0728 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoforms 6/9/10 (Trabeculin-beta) (Bullous pemphigoid antigen) (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
CLH1_HUMAN
|
||||||
| NC score | 0.028597 (rank : 45) | θ value | 6.88961 (rank : 42) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q00610, Q6N0A0, Q86TF2 | Gene names | CLTC, CLH17, KIAA0034 | |||
|
Domain Architecture |

|
|||||
| Description | Clathrin heavy chain 1 (CLH-17). | |||||
|
CLH_MOUSE
|
||||||
| NC score | 0.028593 (rank : 46) | θ value | 6.88961 (rank : 43) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q68FD5 | Gene names | Cltc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Clathrin heavy chain. | |||||
|
TRM6_MOUSE
|
||||||
| NC score | 0.028583 (rank : 47) | θ value | 3.0926 (rank : 31) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CE96, Q3TJZ8, Q3TME7, Q6ZPW8, Q80Y59, Q8CEU0 | Gene names | Trm6, Kiaa1153 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | tRNA (adenine-N(1)-)-methyltransferase non-catalytic subunit TRM6 (tRNA(m1A58)-methyltransferase subunit TRM6) (tRNA(m1A58)MTase subunit TRM6). | |||||
|
SMC1A_HUMAN
|
||||||
| NC score | 0.023705 (rank : 48) | θ value | 6.88961 (rank : 50) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1346 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q14683, O14995, Q16351 | Gene names | SMC1L1, DXS423E, KIAA0178, SB1.8, SMC1 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 1-like 1 protein (SMC1alpha protein) (Sb1.8). | |||||
|
SM1L2_HUMAN
|
||||||
| NC score | 0.023530 (rank : 49) | θ value | 5.27518 (rank : 41) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1019 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8NDV3, Q5TIC3, Q9Y3G5 | Gene names | SMC1L2 | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 1-like 2 protein (SMC1beta protein). | |||||
|
SMC1A_MOUSE
|
||||||
| NC score | 0.023023 (rank : 50) | θ value | 6.88961 (rank : 51) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1384 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9CU62, Q3V480, Q9CUX9, Q9D959, Q9WTU1 | Gene names | Smc1l1, Sb1.8, Smc1, Smcb | |||
|
Domain Architecture |

|
|||||
| Description | Structural maintenance of chromosome 1-like 1 protein (SMC1alpha protein) (Chromosome segregation protein SmcB) (Sb1.8). | |||||
|
LRRF2_MOUSE
|
||||||
| NC score | 0.021460 (rank : 51) | θ value | 3.0926 (rank : 30) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 556 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q91WK0, Q8BVD1, Q9CU89 | Gene names | Lrrfip2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat flightless-interacting protein 2 (LRR FLII- interacting protein 2). | |||||
|
ZN638_HUMAN
|
||||||
| NC score | 0.020816 (rank : 52) | θ value | 4.03905 (rank : 39) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q14966, Q53R34, Q5XJ05, Q68DP3, Q6P2H2, Q7Z3T7, Q8NF92, Q8TCA1, Q9H2G1, Q9NP37 | Gene names | ZNF638, NP220, ZFML | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 638 (Nuclear protein 220) (Zinc-finger matrin-like protein) (Cutaneous T-cell lymphoma-associated antigen se33-1) (CTCL tumor antigen se33-1). | |||||
|
KI13B_HUMAN
|
||||||
| NC score | 0.020502 (rank : 53) | θ value | 0.47712 (rank : 19) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 492 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9NQT8, O75134, Q9BYJ6 | Gene names | KIF13B, GAKIN, KIAA0639 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin-like protein KIF13B (Kinesin-like protein GAKIN). | |||||
|
AR13B_HUMAN
|
||||||
| NC score | 0.018597 (rank : 54) | θ value | 3.0926 (rank : 27) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 528 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q3SXY8, Q504W8 | Gene names | ARL13B, ARL2L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ADP-ribosylation factor-like protein 13B (ADP-ribosylation factor-like protein 2-like 1) (ARL2-like protein 1). | |||||
|
CE290_HUMAN
|
||||||
| NC score | 0.018325 (rank : 55) | θ value | 4.03905 (rank : 32) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1553 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | O15078, Q1PSK5, Q66GS8, Q9H2G6, Q9H6Q7, Q9H8I0 | Gene names | CEP290, KIAA0373, NPHP6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein Cep290 (Nephrocystin-6) (Tumor antigen se2-2). | |||||
|
NUF2_MOUSE
|
||||||
| NC score | 0.017524 (rank : 56) | θ value | 6.88961 (rank : 48) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 585 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q99P69, Q8BTK7, Q8VE05, Q9CST5 | Gene names | Cdca1, Nuf2r | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinetochore protein Nuf2 (Cell division cycle-associated protein 1). | |||||
|
RBCC1_MOUSE
|
||||||
| NC score | 0.017259 (rank : 57) | θ value | 4.03905 (rank : 37) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 1048 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9ESK9, Q3TXX2, Q61384, Q8BRY9, Q9JK14 | Gene names | Rb1cc1, Cc1, Kiaa0203 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RB1-inducible coiled-coil protein 1 (Coiled-coil-forming protein 1) (LaXp180 protein). | |||||
|
PICAL_HUMAN
|
||||||
| NC score | 0.016604 (rank : 58) | θ value | 8.99809 (rank : 55) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q13492, O60700, Q86XZ9 | Gene names | PICALM, CALM | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-binding clathrin assembly protein (Clathrin assembly lymphoid myeloid leukemia protein). | |||||
|
PICA_MOUSE
|
||||||
| NC score | 0.016573 (rank : 59) | θ value | 8.99809 (rank : 56) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 63 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7M6Y3, Q3TS04, Q811P1, Q8BUF6, Q8CIH8, Q8R0A9, Q8R3E1, Q8VDN5, Q921L0 | Gene names | Picalm, Calm, Fit1 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-binding clathrin assembly protein (Clathrin assembly lymphoid myeloid leukemia) (CALM). | |||||
|
DDX56_MOUSE
|
||||||
| NC score | 0.015929 (rank : 60) | θ value | 1.38821 (rank : 25) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 87 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9D0R4 | Gene names | Ddx56, D11Ertd619e, Noh61 | |||
|
Domain Architecture |

|
|||||
| Description | Probable ATP-dependent RNA helicase DDX56 (EC 3.6.1.-) (DEAD box protein 56) (ATP-dependent 61 kDa nucleolar RNA helicase). | |||||
|
KIF5A_HUMAN
|
||||||
| NC score | 0.013691 (rank : 61) | θ value | 8.99809 (rank : 53) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 843 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q12840, Q4LE26 | Gene names | KIF5A, NKHC1 | |||
|
Domain Architecture |

|
|||||
| Description | Kinesin heavy chain isoform 5A (Neuronal kinesin heavy chain) (NKHC) (Kinesin heavy chain neuron-specific 1). | |||||
|
NPAS2_MOUSE
|
||||||
| NC score | 0.008944 (rank : 62) | θ value | 6.88961 (rank : 47) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P97460 | Gene names | Npas2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 2 (Neuronal PAS2). | |||||
|
S4A7_HUMAN
|
||||||
| NC score | 0.007821 (rank : 63) | θ value | 6.88961 (rank : 49) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y6M7, O60350, Q6AHZ9, Q9HC88, Q9UIB9 | Gene names | SLC4A7, BT, NBC2, NBC2B, NBC3, SBC2, SLC4A6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sodium bicarbonate cotransporter 3 (Sodium bicarbonate cotransporter 2) (Sodium bicarbonate cotransporter 2b) (Bicarbonate transporter) (Solute carrier family 4 member 7). | |||||
|
ABCB9_MOUSE
|
||||||
| NC score | 0.004481 (rank : 64) | θ value | 8.99809 (rank : 52) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9JJ59, Q8CHA1, Q9D212, Q9JIN1 | Gene names | Abcb9, Kiaa1520 | |||
|
Domain Architecture |

|
|||||
| Description | ATP-binding cassette sub-family B member 9 precursor (ATP-binding cassette transporter 9) (ABC transporter 9 protein) (TAP-like protein) (TAPL) (mABCB9). | |||||
|
ZN232_HUMAN
|
||||||
| NC score | -0.000245 (rank : 65) | θ value | 4.03905 (rank : 38) | |||
| Query Neighborhood Hits | 56 | Target Neighborhood Hits | 754 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UNY5 | Gene names | ZNF232, ZSCAN11 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 232 (Zinc finger and SCAN domain-containing protein 11). | |||||