Please be patient as the page loads
|
FBSH_HUMAN
|
||||||
| SwissProt Accessions | Q9HAH7, Q96CI9, Q9H9X4 | Gene names | FBS1, FBS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable fibrosin-1 long transcript protein. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
FBSH_HUMAN
|
||||||
| θ value | 8.9516e-133 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9HAH7, Q96CI9, Q9H9X4 | Gene names | FBS1, FBS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable fibrosin-1 long transcript protein. | |||||
|
FBSH_MOUSE
|
||||||
| θ value | 1.74701e-128 (rank : 2) | NC score | 0.985639 (rank : 2) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8R089 | Gene names | Fbs1, Fbs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable fibrosin-1 long transcript protein. | |||||
|
AUTS2_HUMAN
|
||||||
| θ value | 4.14116e-29 (rank : 3) | NC score | 0.635765 (rank : 3) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8WXX7, Q9Y4F2 | Gene names | AUTS2, KIAA0442 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Autism susceptibility gene 2 protein. | |||||
|
NFH_MOUSE
|
||||||
| θ value | 0.000158464 (rank : 4) | NC score | 0.062459 (rank : 21) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
MUC1_HUMAN
|
||||||
| θ value | 0.00102713 (rank : 5) | NC score | 0.081943 (rank : 6) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
CEBPE_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 6) | NC score | 0.077675 (rank : 7) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q15744, Q15745, Q8IYI2, Q99803 | Gene names | CEBPE | |||
|
Domain Architecture |

|
|||||
| Description | CCAAT/enhancer-binding protein epsilon (C/EBP epsilon). | |||||
|
LPP_HUMAN
|
||||||
| θ value | 0.125558 (rank : 7) | NC score | 0.042633 (rank : 55) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 775 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q93052, Q8NFX5 | Gene names | LPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lipoma-preferred partner (LIM domain-containing preferred translocation partner in lipoma). | |||||
|
PCLO_MOUSE
|
||||||
| θ value | 0.21417 (rank : 8) | NC score | 0.074380 (rank : 10) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
CBP_MOUSE
|
||||||
| θ value | 0.279714 (rank : 9) | NC score | 0.072472 (rank : 12) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1051 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P45481 | Gene names | Crebbp, Cbp | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
HNF1A_HUMAN
|
||||||
| θ value | 1.06291 (rank : 10) | NC score | 0.051123 (rank : 42) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P20823, Q99861 | Gene names | TCF1, HNF1A | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte nuclear factor 1-alpha (HNF-1A) (Liver-specific transcription factor LF-B1) (LFB1) (Transcription factor 1) (TCF-1). | |||||
|
MISS_MOUSE
|
||||||
| θ value | 1.06291 (rank : 11) | NC score | 0.070283 (rank : 14) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 444 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9D7G9, Q7M749 | Gene names | Mapk1ip1, Miss | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MAPK-interacting and spindle-stabilizing protein (Mitogen-activated protein kinase 1-interacting protein 1). | |||||
|
ZN687_HUMAN
|
||||||
| θ value | 1.06291 (rank : 12) | NC score | 0.026482 (rank : 65) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1244 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8N1G0, Q68DQ8, Q9H937, Q9P2A7 | Gene names | ZNF687, KIAA1441 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 687. | |||||
|
INSM1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 13) | NC score | 0.031482 (rank : 62) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 678 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q63ZV0, Q9Z113 | Gene names | Insm1, Ia1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Insulinoma-associated protein 1 (Zinc finger protein IA-1). | |||||
|
NRX2A_HUMAN
|
||||||
| θ value | 1.38821 (rank : 14) | NC score | 0.040854 (rank : 56) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 614 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9P2S2, Q9Y2D6 | Gene names | NRXN2, KIAA0921 | |||
|
Domain Architecture |

|
|||||
| Description | Neurexin-2-alpha precursor (Neurexin II-alpha). | |||||
|
NRX2B_HUMAN
|
||||||
| θ value | 1.38821 (rank : 15) | NC score | 0.057320 (rank : 25) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 481 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P58401 | Gene names | NRXN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neurexin-2-beta precursor (Neurexin II-beta). | |||||
|
BHLH3_HUMAN
|
||||||
| θ value | 1.81305 (rank : 16) | NC score | 0.033766 (rank : 61) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9C0J9 | Gene names | BHLHB3, DEC2, SHARP1 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 3 (bHLHB3) (Differentially expressed in chondrocytes protein 2) (hDEC2) (Enhancer-of-split and hairy-related protein 1) (SHARP-1). | |||||
|
KCNH6_HUMAN
|
||||||
| θ value | 1.81305 (rank : 17) | NC score | 0.019124 (rank : 68) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9H252, Q9BRD7 | Gene names | KCNH6, ERG2, HERG2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 6 (Voltage-gated potassium channel subunit Kv11.2) (Ether-a-go-go-related gene potassium channel 2) (Ether-a-go-go-related protein 2) (Eag-related protein 2). | |||||
|
NFH_HUMAN
|
||||||
| θ value | 1.81305 (rank : 18) | NC score | 0.038016 (rank : 58) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1242 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P12036, Q9UJS7, Q9UQ14 | Gene names | NEFH, KIAA0845, NFH | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
SON_HUMAN
|
||||||
| θ value | 1.81305 (rank : 19) | NC score | 0.043527 (rank : 51) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
SPEG_MOUSE
|
||||||
| θ value | 1.81305 (rank : 20) | NC score | 0.016487 (rank : 71) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1959 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q62407, Q3TPH8, Q6P5V1, Q80TF7, Q80ZN0, Q8BZF4, Q9EQJ5 | Gene names | Speg, Apeg1, Kiaa1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle-specific serine/threonine protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
BTBD7_MOUSE
|
||||||
| θ value | 2.36792 (rank : 21) | NC score | 0.035690 (rank : 60) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CFE5, Q80TC3, Q8BZY9 | Gene names | Btbd7, Kiaa1525 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein 7. | |||||
|
CAC1A_MOUSE
|
||||||
| θ value | 2.36792 (rank : 22) | NC score | 0.014467 (rank : 72) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P97445 | Gene names | Cacna1a, Cach4, Cacn3, Cacnl1a4, Ccha1a | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent P/Q-type calcium channel subunit alpha-1A (Voltage- gated calcium channel subunit alpha Cav2.1) (Calcium channel, L type, alpha-1 polypeptide isoform 4) (Brain calcium channel I) (BI). | |||||
|
SHAN1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 23) | NC score | 0.035982 (rank : 59) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9Y566, Q9NYW9 | Gene names | SHANK1 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 and multiple ankyrin repeat domains protein 1 (Shank1) (Somatostatin receptor-interacting protein) (SSTR-interacting protein) (SSTRIP). | |||||
|
BSN_HUMAN
|
||||||
| θ value | 3.0926 (rank : 24) | NC score | 0.089929 (rank : 4) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
BSN_MOUSE
|
||||||
| θ value | 3.0926 (rank : 25) | NC score | 0.076693 (rank : 8) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
K1802_MOUSE
|
||||||
| θ value | 3.0926 (rank : 26) | NC score | 0.042660 (rank : 54) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1596 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8K327, Q3UZ85, Q6ZPI1 | Gene names | Kiaa1802, D8Ertd457e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
MLL2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 27) | NC score | 0.043276 (rank : 52) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
SEPT9_HUMAN
|
||||||
| θ value | 4.03905 (rank : 28) | NC score | 0.029150 (rank : 63) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 706 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9UHD8, Q96QF3, Q96QF4, Q96QF5, Q9HA04, Q9UG40, Q9Y5W4 | Gene names | SEPT9, KIAA0991, MSF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Septin-9 (MLL septin-like fusion protein) (MLL septin-like fusion protein MSF-A) (Ovarian/Breast septin) (Ov/Br septin) (Septin D1). | |||||
|
HAX1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 29) | NC score | 0.026812 (rank : 64) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O35387 | Gene names | Hax1, Hs1bp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HS1-associating protein X-1 (HAX-1) (HS1-binding protein). | |||||
|
HCFC1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 30) | NC score | 0.044944 (rank : 50) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 331 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P51610, Q6P4G5 | Gene names | HCFC1, HCF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Host cell factor (HCF) (HCF-1) (C1 factor) (VP16 accessory protein) (VCAF) (CFF) [Contains: HCF N-terminal chain 1; HCF N-terminal chain 2; HCF N-terminal chain 3; HCF N-terminal chain 4; HCF N-terminal chain 5; HCF N-terminal chain 6; HCF C-terminal chain 1; HCF C- terminal chain 2; HCF C-terminal chain 3; HCF C-terminal chain 4; HCF C-terminal chain 5; HCF C-terminal chain 6]. | |||||
|
HCN3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 31) | NC score | 0.013950 (rank : 73) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O88705 | Gene names | Hcn3, Hac3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 3 (Hyperpolarization-activated cation channel 3) (HAC-3). | |||||
|
LNK_HUMAN
|
||||||
| θ value | 5.27518 (rank : 32) | NC score | 0.017191 (rank : 70) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UQQ2, O95184 | Gene names | LNK | |||
|
Domain Architecture |
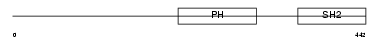
|
|||||
| Description | Lymphocyte-specific adapter protein Lnk (Signal transduction protein Lnk) (Lymphocyte adapter protein). | |||||
|
SC24C_HUMAN
|
||||||
| θ value | 5.27518 (rank : 33) | NC score | 0.049425 (rank : 47) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 662 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P53992, Q8WV25 | Gene names | SEC24C, KIAA0079 | |||
|
Domain Architecture |

|
|||||
| Description | Protein transport protein Sec24C (SEC24-related protein C). | |||||
|
HNF1A_MOUSE
|
||||||
| θ value | 6.88961 (rank : 34) | NC score | 0.042691 (rank : 53) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P22361 | Gene names | Tcf1, Hnf-1, Hnf-1a, Hnf1a | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte nuclear factor 1-alpha (HNF-1A) (Liver-specific transcription factor LF-B1) (LFB1). | |||||
|
IRF8_HUMAN
|
||||||
| θ value | 6.88961 (rank : 35) | NC score | 0.011055 (rank : 74) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q02556 | Gene names | IRF8, ICSBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon regulatory factor 8 (IRF-8) (Interferon consensus sequence- binding protein) (ICSBP). | |||||
|
K0174_MOUSE
|
||||||
| θ value | 6.88961 (rank : 36) | NC score | 0.023042 (rank : 66) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9CX00, Q80U68 | Gene names | Kiaa0174 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0174. | |||||
|
NDF1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 37) | NC score | 0.018943 (rank : 69) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 186 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q13562, O00343, Q13340, Q5U095, Q96TH0, Q99455, Q9UEC8 | Gene names | NEUROD1, NEUROD | |||
|
Domain Architecture |
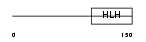
|
|||||
| Description | Neurogenic differentiation factor 1 (NeuroD1) (NeuroD). | |||||
|
OGFR_MOUSE
|
||||||
| θ value | 6.88961 (rank : 38) | NC score | 0.040807 (rank : 57) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 584 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q99PG2, Q99L33 | Gene names | Ogfr | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor). | |||||
|
RBM16_HUMAN
|
||||||
| θ value | 6.88961 (rank : 39) | NC score | 0.051362 (rank : 39) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 300 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9UPN6, Q5TBU6, Q9BQN8, Q9BX43 | Gene names | RBM16, KIAA1116 | |||
|
Domain Architecture |

|
|||||
| Description | Putative RNA-binding protein 16 (RNA-binding motif protein 16). | |||||
|
ATN1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 40) | NC score | 0.073710 (rank : 11) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P54259, Q99495, Q99621, Q9UEK7 | Gene names | ATN1, DRPLA | |||
|
Domain Architecture |

|
|||||
| Description | Atrophin-1 (Dentatorubral-pallidoluysian atrophy protein). | |||||
|
CSPG2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 41) | NC score | 0.009448 (rank : 75) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 711 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q62059, Q62058, Q9CUU0 | Gene names | Cspg2 | |||
|
Domain Architecture |

|
|||||
| Description | Versican core protein precursor (Large fibroblast proteoglycan) (Chondroitin sulfate proteoglycan core protein 2) (PG-M). | |||||
|
MAVS_HUMAN
|
||||||
| θ value | 8.99809 (rank : 42) | NC score | 0.047317 (rank : 48) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q7Z434, Q3I0Y2, Q5T7I6, Q86VY7, Q9H1H3, Q9H4Y1, Q9H8D3, Q9ULE9 | Gene names | MAVS, IPS1, KIAA1271, VISA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitochondrial antiviral signaling protein (Interferon-beta promoter stimulator protein 1) (IPS-1) (Virus-induced signaling adapter) (CARD adapter inducing interferon-beta) (Cardif) (Putative NF-kappa-B- activating protein 031N). | |||||
|
MEGF8_HUMAN
|
||||||
| θ value | 8.99809 (rank : 43) | NC score | 0.006622 (rank : 76) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 528 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q7Z7M0, O75097 | Gene names | MEGF8, EGFL4, KIAA0817 | |||
|
Domain Architecture |
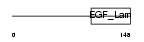
|
|||||
| Description | Multiple epidermal growth factor-like domains 8 (EGF-like domain- containing protein 4) (Multiple EGF-like domain protein 4). | |||||
|
NACAM_MOUSE
|
||||||
| θ value | 8.99809 (rank : 44) | NC score | 0.055997 (rank : 29) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
NUP98_HUMAN
|
||||||
| θ value | 8.99809 (rank : 45) | NC score | 0.021042 (rank : 67) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P52948, Q8IUT2, Q8WYB0, Q96E54, Q9H3Q4, Q9NT02, Q9UF57, Q9UHX0, Q9Y6J4, Q9Y6J5 | Gene names | NUP98 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup98-Nup96 precursor [Contains: Nuclear pore complex protein Nup98 (Nucleoporin Nup98) (98 kDa nucleoporin); Nuclear pore complex protein Nup96 (Nucleoporin Nup96) (96 kDa nucleoporin)]. | |||||
|
OGFR_HUMAN
|
||||||
| θ value | 8.99809 (rank : 46) | NC score | 0.047220 (rank : 49) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
BASP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 47) | NC score | 0.051483 (rank : 37) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P80723, O43596, Q5U0S0 | Gene names | BASP1, NAP22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
BASP_MOUSE
|
||||||
| θ value | θ > 10 (rank : 48) | NC score | 0.051348 (rank : 40) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q91XV3 | Gene names | Basp1, Nap22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
CBP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 49) | NC score | 0.050667 (rank : 45) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 941 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q92793, O00147, Q16376 | Gene names | CREBBP, CBP | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
CDN1C_HUMAN
|
||||||
| θ value | θ > 10 (rank : 50) | NC score | 0.062260 (rank : 22) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P49918 | Gene names | CDKN1C, KIP2 | |||
|
Domain Architecture |
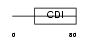
|
|||||
| Description | Cyclin-dependent kinase inhibitor 1C (Cyclin-dependent kinase inhibitor p57) (p57KIP2). | |||||
|
CN032_HUMAN
|
||||||
| θ value | θ > 10 (rank : 51) | NC score | 0.066340 (rank : 16) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8NDC0, Q96BG5 | Gene names | C14orf32 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf32. | |||||
|
CN032_MOUSE
|
||||||
| θ value | θ > 10 (rank : 52) | NC score | 0.086638 (rank : 5) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8BH93, Q3TF12, Q3TGL0, Q8CC90 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf32 homolog. | |||||
|
CN155_HUMAN
|
||||||
| θ value | θ > 10 (rank : 53) | NC score | 0.052491 (rank : 34) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q5H9T9, Q5H9U7, Q86YI2, Q9H0J3 | Gene names | C14orf155 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf155. | |||||
|
EP300_HUMAN
|
||||||
| θ value | θ > 10 (rank : 54) | NC score | 0.050674 (rank : 44) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q09472 | Gene names | EP300, P300 | |||
|
Domain Architecture |

|
|||||
| Description | E1A-associated protein p300 (EC 2.3.1.48). | |||||
|
GGN_HUMAN
|
||||||
| θ value | θ > 10 (rank : 55) | NC score | 0.055132 (rank : 31) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q86UU5, Q7RTU6, Q86UU4, Q8NAA1 | Gene names | GGN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gametogenetin. | |||||
|
GGN_MOUSE
|
||||||
| θ value | θ > 10 (rank : 56) | NC score | 0.059515 (rank : 24) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q80WJ1, Q5EBP4, Q80WI9, Q80WJ0 | Gene names | Ggn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gametogenetin. | |||||
|
GSCR1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 57) | NC score | 0.064465 (rank : 17) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 674 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9NZM4 | Gene names | GLTSCR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glioma tumor suppressor candidate region gene 1 protein. | |||||
|
MARCS_HUMAN
|
||||||
| θ value | θ > 10 (rank : 58) | NC score | 0.055955 (rank : 30) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P29966, Q2LA83, Q5TDB7 | Gene names | MARCKS, MACS, PRKCSL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myristoylated alanine-rich C-kinase substrate (MARCKS) (Protein kinase C substrate, 80 kDa protein, light chain) (PKCSL) (80K-L protein). | |||||
|
MARCS_MOUSE
|
||||||
| θ value | θ > 10 (rank : 59) | NC score | 0.051157 (rank : 41) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P26645 | Gene names | Marcks, Macs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myristoylated alanine-rich C-kinase substrate (MARCKS). | |||||
|
MBD6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 60) | NC score | 0.062759 (rank : 20) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q96DN6, Q8N3M0, Q8NA81, Q96Q00 | Gene names | MBD6, KIAA1887 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 6 (Methyl-CpG-binding protein MBD6). | |||||
|
MUC1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 61) | NC score | 0.074507 (rank : 9) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q02496 | Gene names | Muc1, Muc-1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (Polymorphic epithelial mucin) (PEMT) (Episialin). | |||||
|
MUC7_HUMAN
|
||||||
| θ value | θ > 10 (rank : 62) | NC score | 0.056270 (rank : 28) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 538 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8TAX7 | Gene names | MUC7, MG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucin-7 precursor (MUC-7) (Salivary mucin-7) (Apo-MG2). | |||||
|
PCLO_HUMAN
|
||||||
| θ value | θ > 10 (rank : 63) | NC score | 0.063935 (rank : 18) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
PRG4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 64) | NC score | 0.056329 (rank : 27) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q92954, Q6DNC4, Q6DNC5, Q6ZMZ5, Q9BX49 | Gene names | PRG4, MSF, SZP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
PRP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 65) | NC score | 0.067665 (rank : 15) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P04280, Q08805, Q15186, Q15187, Q15214, Q15215, Q16038 | Gene names | PRB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 1 precursor (Salivary proline-rich protein) [Contains: Basic peptide IB-6; Peptide P-H]. | |||||
|
PRP2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 66) | NC score | 0.070651 (rank : 13) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P05142 | Gene names | Prh1, Prp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein MP-2 precursor. | |||||
|
PRR12_HUMAN
|
||||||
| θ value | θ > 10 (rank : 67) | NC score | 0.061522 (rank : 23) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 460 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9ULL5, Q8N4J6 | Gene names | PRR12, KIAA1205 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 12. | |||||
|
SELV_HUMAN
|
||||||
| θ value | θ > 10 (rank : 68) | NC score | 0.056609 (rank : 26) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P59797 | Gene names | SELV | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Selenoprotein V. | |||||
|
SF3A2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 69) | NC score | 0.052599 (rank : 33) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 263 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q15428, O75245 | Gene names | SF3A2, SAP62 | |||
|
Domain Architecture |
|
|||||
| Description | Splicing factor 3A subunit 2 (Spliceosome-associated protein 62) (SAP 62) (SF3a66). | |||||
|
SF3A2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 70) | NC score | 0.051943 (rank : 35) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q62203 | Gene names | Sf3a2, Sap62 | |||
|
Domain Architecture |
|
|||||
| Description | Splicing factor 3A subunit 2 (Spliceosome-associated protein 62) (SAP 62) (SF3a66). | |||||
|
SMR3A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 71) | NC score | 0.050832 (rank : 43) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q99954 | Gene names | SMR3A, PBI, PROL5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Submaxillary gland androgen-regulated protein 3 homolog A precursor (Proline-rich protein 5) (Proline-rich protein PBI). | |||||
|
SRRM1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 72) | NC score | 0.050066 (rank : 46) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 73) | NC score | 0.051435 (rank : 38) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
T22D2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 74) | NC score | 0.053626 (rank : 32) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 250 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O75157, Q6PI50, Q9H2Z6, Q9H2Z7, Q9H2Z8 | Gene names | TSC22D2, KIAA0669, TILZ4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TSC22 domain family protein 2 (TSC22-related-inducible leucine zipper protein 4). | |||||
|
TAF4_HUMAN
|
||||||
| θ value | θ > 10 (rank : 75) | NC score | 0.063028 (rank : 19) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 687 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O00268, Q5TBP6, Q99721, Q9BR40, Q9BX42 | Gene names | TAF4, TAF2C, TAF2C1, TAF4A, TAFII130, TAFII135 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 4 (TBP-associated factor 4) (Transcription initiation factor TFIID 135 kDa subunit) (TAF(II)135) (TAFII-135) (TAFII135) (TAFII-130) (TAFII130). | |||||
|
WASIP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 76) | NC score | 0.051924 (rank : 36) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O43516, Q15220, Q6MZU9, Q9BU37, Q9UNP1 | Gene names | WASPIP, WIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein) (PRPL-2 protein). | |||||
|
FBSH_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 8.9516e-133 (rank : 1) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | Q9HAH7, Q96CI9, Q9H9X4 | Gene names | FBS1, FBS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable fibrosin-1 long transcript protein. | |||||
|
FBSH_MOUSE
|
||||||
| NC score | 0.985639 (rank : 2) | θ value | 1.74701e-128 (rank : 2) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q8R089 | Gene names | Fbs1, Fbs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable fibrosin-1 long transcript protein. | |||||
|
AUTS2_HUMAN
|
||||||
| NC score | 0.635765 (rank : 3) | θ value | 4.14116e-29 (rank : 3) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 342 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8WXX7, Q9Y4F2 | Gene names | AUTS2, KIAA0442 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Autism susceptibility gene 2 protein. | |||||
|
BSN_HUMAN
|
||||||
| NC score | 0.089929 (rank : 4) | θ value | 3.0926 (rank : 24) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1537 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9UPA5, O43161, Q7LGH3 | Gene names | BSN, KIAA0434, ZNF231 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon (Zinc finger protein 231). | |||||
|
CN032_MOUSE
|
||||||
| NC score | 0.086638 (rank : 5) | θ value | θ > 10 (rank : 52) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 233 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8BH93, Q3TF12, Q3TGL0, Q8CC90 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf32 homolog. | |||||
|
MUC1_HUMAN
|
||||||
| NC score | 0.081943 (rank : 6) | θ value | 0.00102713 (rank : 5) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 975 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P15941, P13931, P15942, P17626, Q14128, Q14876, Q16437, Q16442, Q16615, Q9BXA4, Q9UE75, Q9UE76, Q9UQL1, Q9Y4J2 | Gene names | MUC1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (MUC-1) (Polymorphic epithelial mucin) (PEM) (PEMT) (Episialin) (Tumor-associated mucin) (Carcinoma-associated mucin) (Tumor-associated epithelial membrane antigen) (EMA) (H23AG) (Peanut- reactive urinary mucin) (PUM) (Breast carcinoma-associated antigen DF3) (CD227 antigen). | |||||
|
CEBPE_HUMAN
|
||||||
| NC score | 0.077675 (rank : 7) | θ value | 0.0736092 (rank : 6) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q15744, Q15745, Q8IYI2, Q99803 | Gene names | CEBPE | |||
|
Domain Architecture |

|
|||||
| Description | CCAAT/enhancer-binding protein epsilon (C/EBP epsilon). | |||||
|
BSN_MOUSE
|
||||||
| NC score | 0.076693 (rank : 8) | θ value | 3.0926 (rank : 25) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 2057 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O88737, Q6ZQB5 | Gene names | Bsn, Kiaa0434 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein bassoon. | |||||
|
MUC1_MOUSE
|
||||||
| NC score | 0.074507 (rank : 9) | θ value | θ > 10 (rank : 61) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q02496 | Gene names | Muc1, Muc-1 | |||
|
Domain Architecture |

|
|||||
| Description | Mucin-1 precursor (Polymorphic epithelial mucin) (PEMT) (Episialin). | |||||
|
PCLO_MOUSE
|
||||||
| NC score | 0.074380 (rank : 10) | θ value | 0.21417 (rank : 8) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
ATN1_HUMAN
|
||||||
| NC score | 0.073710 (rank : 11) | θ value | 8.99809 (rank : 40) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 676 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P54259, Q99495, Q99621, Q9UEK7 | Gene names | ATN1, DRPLA | |||
|
Domain Architecture |

|
|||||
| Description | Atrophin-1 (Dentatorubral-pallidoluysian atrophy protein). | |||||
|
CBP_MOUSE
|
||||||
| NC score | 0.072472 (rank : 12) | θ value | 0.279714 (rank : 9) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1051 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P45481 | Gene names | Crebbp, Cbp | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
PRP2_MOUSE
|
||||||
| NC score | 0.070651 (rank : 13) | θ value | θ > 10 (rank : 66) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 347 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P05142 | Gene names | Prh1, Prp | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein MP-2 precursor. | |||||
|
MISS_MOUSE
|
||||||
| NC score | 0.070283 (rank : 14) | θ value | 1.06291 (rank : 11) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 444 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9D7G9, Q7M749 | Gene names | Mapk1ip1, Miss | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MAPK-interacting and spindle-stabilizing protein (Mitogen-activated protein kinase 1-interacting protein 1). | |||||
|
PRP1_HUMAN
|
||||||
| NC score | 0.067665 (rank : 15) | θ value | θ > 10 (rank : 65) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P04280, Q08805, Q15186, Q15187, Q15214, Q15215, Q16038 | Gene names | PRB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Basic salivary proline-rich protein 1 precursor (Salivary proline-rich protein) [Contains: Basic peptide IB-6; Peptide P-H]. | |||||
|
CN032_HUMAN
|
||||||
| NC score | 0.066340 (rank : 16) | θ value | θ > 10 (rank : 51) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8NDC0, Q96BG5 | Gene names | C14orf32 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf32. | |||||
|
GSCR1_HUMAN
|
||||||
| NC score | 0.064465 (rank : 17) | θ value | θ > 10 (rank : 57) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 674 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9NZM4 | Gene names | GLTSCR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glioma tumor suppressor candidate region gene 1 protein. | |||||
|
PCLO_HUMAN
|
||||||
| NC score | 0.063935 (rank : 18) | θ value | θ > 10 (rank : 63) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
TAF4_HUMAN
|
||||||
| NC score | 0.063028 (rank : 19) | θ value | θ > 10 (rank : 75) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 687 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O00268, Q5TBP6, Q99721, Q9BR40, Q9BX42 | Gene names | TAF4, TAF2C, TAF2C1, TAF4A, TAFII130, TAFII135 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor TFIID subunit 4 (TBP-associated factor 4) (Transcription initiation factor TFIID 135 kDa subunit) (TAF(II)135) (TAFII-135) (TAFII135) (TAFII-130) (TAFII130). | |||||
|
MBD6_HUMAN
|
||||||
| NC score | 0.062759 (rank : 20) | θ value | θ > 10 (rank : 60) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 349 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q96DN6, Q8N3M0, Q8NA81, Q96Q00 | Gene names | MBD6, KIAA1887 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 6 (Methyl-CpG-binding protein MBD6). | |||||
|
NFH_MOUSE
|
||||||
| NC score | 0.062459 (rank : 21) | θ value | 0.000158464 (rank : 4) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 2247 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P19246, Q5SVF6, Q61959 | Gene names | Nefh, Nfh | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
CDN1C_HUMAN
|
||||||
| NC score | 0.062260 (rank : 22) | θ value | θ > 10 (rank : 50) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P49918 | Gene names | CDKN1C, KIP2 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin-dependent kinase inhibitor 1C (Cyclin-dependent kinase inhibitor p57) (p57KIP2). | |||||
|
PRR12_HUMAN
|
||||||
| NC score | 0.061522 (rank : 23) | θ value | θ > 10 (rank : 67) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 460 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9ULL5, Q8N4J6 | Gene names | PRR12, KIAA1205 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-rich protein 12. | |||||
|
GGN_MOUSE
|
||||||
| NC score | 0.059515 (rank : 24) | θ value | θ > 10 (rank : 56) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 311 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q80WJ1, Q5EBP4, Q80WI9, Q80WJ0 | Gene names | Ggn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gametogenetin. | |||||
|
NRX2B_HUMAN
|
||||||
| NC score | 0.057320 (rank : 25) | θ value | 1.38821 (rank : 15) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 481 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P58401 | Gene names | NRXN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neurexin-2-beta precursor (Neurexin II-beta). | |||||
|
SELV_HUMAN
|
||||||
| NC score | 0.056609 (rank : 26) | θ value | θ > 10 (rank : 68) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P59797 | Gene names | SELV | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Selenoprotein V. | |||||
|
PRG4_HUMAN
|
||||||
| NC score | 0.056329 (rank : 27) | θ value | θ > 10 (rank : 64) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 764 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q92954, Q6DNC4, Q6DNC5, Q6ZMZ5, Q9BX49 | Gene names | PRG4, MSF, SZP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proteoglycan-4 precursor (Lubricin) (Megakaryocyte-stimulating factor) (Superficial zone proteoglycan) [Contains: Proteoglycan-4 C-terminal part]. | |||||
|
MUC7_HUMAN
|
||||||
| NC score | 0.056270 (rank : 28) | θ value | θ > 10 (rank : 62) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 538 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8TAX7 | Gene names | MUC7, MG2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucin-7 precursor (MUC-7) (Salivary mucin-7) (Apo-MG2). | |||||
|
NACAM_MOUSE
|
||||||
| NC score | 0.055997 (rank : 29) | θ value | 8.99809 (rank : 44) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
MARCS_HUMAN
|
||||||
| NC score | 0.055955 (rank : 30) | θ value | θ > 10 (rank : 58) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P29966, Q2LA83, Q5TDB7 | Gene names | MARCKS, MACS, PRKCSL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myristoylated alanine-rich C-kinase substrate (MARCKS) (Protein kinase C substrate, 80 kDa protein, light chain) (PKCSL) (80K-L protein). | |||||
|
GGN_HUMAN
|
||||||
| NC score | 0.055132 (rank : 31) | θ value | θ > 10 (rank : 55) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 269 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q86UU5, Q7RTU6, Q86UU4, Q8NAA1 | Gene names | GGN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Gametogenetin. | |||||
|
T22D2_HUMAN
|
||||||
| NC score | 0.053626 (rank : 32) | θ value | θ > 10 (rank : 74) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 250 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O75157, Q6PI50, Q9H2Z6, Q9H2Z7, Q9H2Z8 | Gene names | TSC22D2, KIAA0669, TILZ4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TSC22 domain family protein 2 (TSC22-related-inducible leucine zipper protein 4). | |||||
|
SF3A2_HUMAN
|
||||||
| NC score | 0.052599 (rank : 33) | θ value | θ > 10 (rank : 69) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 263 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q15428, O75245 | Gene names | SF3A2, SAP62 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 3A subunit 2 (Spliceosome-associated protein 62) (SAP 62) (SF3a66). | |||||
|
CN155_HUMAN
|
||||||
| NC score | 0.052491 (rank : 34) | θ value | θ > 10 (rank : 53) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q5H9T9, Q5H9U7, Q86YI2, Q9H0J3 | Gene names | C14orf155 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf155. | |||||
|
SF3A2_MOUSE
|
||||||
| NC score | 0.051943 (rank : 35) | θ value | θ > 10 (rank : 70) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q62203 | Gene names | Sf3a2, Sap62 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor 3A subunit 2 (Spliceosome-associated protein 62) (SAP 62) (SF3a66). | |||||
|
WASIP_HUMAN
|
||||||
| NC score | 0.051924 (rank : 36) | θ value | θ > 10 (rank : 76) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O43516, Q15220, Q6MZU9, Q9BU37, Q9UNP1 | Gene names | WASPIP, WIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wiskott-Aldrich syndrome protein-interacting protein (WASP-interacting protein) (PRPL-2 protein). | |||||
|
BASP_HUMAN
|
||||||
| NC score | 0.051483 (rank : 37) | θ value | θ > 10 (rank : 47) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P80723, O43596, Q5U0S0 | Gene names | BASP1, NAP22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.051435 (rank : 38) | θ value | θ > 10 (rank : 73) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
RBM16_HUMAN
|
||||||
| NC score | 0.051362 (rank : 39) | θ value | 6.88961 (rank : 39) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 300 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9UPN6, Q5TBU6, Q9BQN8, Q9BX43 | Gene names | RBM16, KIAA1116 | |||
|
Domain Architecture |

|
|||||
| Description | Putative RNA-binding protein 16 (RNA-binding motif protein 16). | |||||
|
BASP_MOUSE
|
||||||
| NC score | 0.051348 (rank : 40) | θ value | θ > 10 (rank : 48) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q91XV3 | Gene names | Basp1, Nap22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
MARCS_MOUSE
|
||||||
| NC score | 0.051157 (rank : 41) | θ value | θ > 10 (rank : 59) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P26645 | Gene names | Marcks, Macs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myristoylated alanine-rich C-kinase substrate (MARCKS). | |||||
|
HNF1A_HUMAN
|
||||||
| NC score | 0.051123 (rank : 42) | θ value | 1.06291 (rank : 10) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P20823, Q99861 | Gene names | TCF1, HNF1A | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte nuclear factor 1-alpha (HNF-1A) (Liver-specific transcription factor LF-B1) (LFB1) (Transcription factor 1) (TCF-1). | |||||
|
SMR3A_HUMAN
|
||||||
| NC score | 0.050832 (rank : 43) | θ value | θ > 10 (rank : 71) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 93 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q99954 | Gene names | SMR3A, PBI, PROL5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Submaxillary gland androgen-regulated protein 3 homolog A precursor (Proline-rich protein 5) (Proline-rich protein PBI). | |||||
|
EP300_HUMAN
|
||||||
| NC score | 0.050674 (rank : 44) | θ value | θ > 10 (rank : 54) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 821 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q09472 | Gene names | EP300, P300 | |||
|
Domain Architecture |

|
|||||
| Description | E1A-associated protein p300 (EC 2.3.1.48). | |||||
|
CBP_HUMAN
|
||||||
| NC score | 0.050667 (rank : 45) | θ value | θ > 10 (rank : 49) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 941 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q92793, O00147, Q16376 | Gene names | CREBBP, CBP | |||
|
Domain Architecture |

|
|||||
| Description | CREB-binding protein (EC 2.3.1.48). | |||||
|
SRRM1_MOUSE
|
||||||
| NC score | 0.050066 (rank : 46) | θ value | θ > 10 (rank : 72) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 966 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q52KI8, O70495, Q9CVG5 | Gene names | Srrm1, Pop101 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 1 (Plenty-of-prolines 101). | |||||
|
SC24C_HUMAN
|
||||||
| NC score | 0.049425 (rank : 47) | θ value | 5.27518 (rank : 33) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 662 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P53992, Q8WV25 | Gene names | SEC24C, KIAA0079 | |||
|
Domain Architecture |

|
|||||
| Description | Protein transport protein Sec24C (SEC24-related protein C). | |||||
|
MAVS_HUMAN
|
||||||
| NC score | 0.047317 (rank : 48) | θ value | 8.99809 (rank : 42) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q7Z434, Q3I0Y2, Q5T7I6, Q86VY7, Q9H1H3, Q9H4Y1, Q9H8D3, Q9ULE9 | Gene names | MAVS, IPS1, KIAA1271, VISA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitochondrial antiviral signaling protein (Interferon-beta promoter stimulator protein 1) (IPS-1) (Virus-induced signaling adapter) (CARD adapter inducing interferon-beta) (Cardif) (Putative NF-kappa-B- activating protein 031N). | |||||
|
OGFR_HUMAN
|
||||||
| NC score | 0.047220 (rank : 49) | θ value | 8.99809 (rank : 46) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
HCFC1_HUMAN
|
||||||
| NC score | 0.044944 (rank : 50) | θ value | 5.27518 (rank : 30) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 331 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P51610, Q6P4G5 | Gene names | HCFC1, HCF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Host cell factor (HCF) (HCF-1) (C1 factor) (VP16 accessory protein) (VCAF) (CFF) [Contains: HCF N-terminal chain 1; HCF N-terminal chain 2; HCF N-terminal chain 3; HCF N-terminal chain 4; HCF N-terminal chain 5; HCF N-terminal chain 6; HCF C-terminal chain 1; HCF C- terminal chain 2; HCF C-terminal chain 3; HCF C-terminal chain 4; HCF C-terminal chain 5; HCF C-terminal chain 6]. | |||||
|
SON_HUMAN
|
||||||
| NC score | 0.043527 (rank : 51) | θ value | 1.81305 (rank : 19) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
MLL2_HUMAN
|
||||||
| NC score | 0.043276 (rank : 52) | θ value | 4.03905 (rank : 27) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
HNF1A_MOUSE
|
||||||
| NC score | 0.042691 (rank : 53) | θ value | 6.88961 (rank : 34) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P22361 | Gene names | Tcf1, Hnf-1, Hnf-1a, Hnf1a | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte nuclear factor 1-alpha (HNF-1A) (Liver-specific transcription factor LF-B1) (LFB1). | |||||
|
K1802_MOUSE
|
||||||
| NC score | 0.042660 (rank : 54) | θ value | 3.0926 (rank : 26) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1596 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q8K327, Q3UZ85, Q6ZPI1 | Gene names | Kiaa1802, D8Ertd457e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
LPP_HUMAN
|
||||||
| NC score | 0.042633 (rank : 55) | θ value | 0.125558 (rank : 7) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 775 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q93052, Q8NFX5 | Gene names | LPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lipoma-preferred partner (LIM domain-containing preferred translocation partner in lipoma). | |||||
|
NRX2A_HUMAN
|
||||||
| NC score | 0.040854 (rank : 56) | θ value | 1.38821 (rank : 14) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 614 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9P2S2, Q9Y2D6 | Gene names | NRXN2, KIAA0921 | |||
|
Domain Architecture |

|
|||||
| Description | Neurexin-2-alpha precursor (Neurexin II-alpha). | |||||
|
OGFR_MOUSE
|
||||||
| NC score | 0.040807 (rank : 57) | θ value | 6.88961 (rank : 38) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 584 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q99PG2, Q99L33 | Gene names | Ogfr | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor). | |||||
|
NFH_HUMAN
|
||||||
| NC score | 0.038016 (rank : 58) | θ value | 1.81305 (rank : 18) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1242 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P12036, Q9UJS7, Q9UQ14 | Gene names | NEFH, KIAA0845, NFH | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
SHAN1_HUMAN
|
||||||
| NC score | 0.035982 (rank : 59) | θ value | 2.36792 (rank : 23) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9Y566, Q9NYW9 | Gene names | SHANK1 | |||
|
Domain Architecture |

|
|||||
| Description | SH3 and multiple ankyrin repeat domains protein 1 (Shank1) (Somatostatin receptor-interacting protein) (SSTR-interacting protein) (SSTRIP). | |||||
|
BTBD7_MOUSE
|
||||||
| NC score | 0.035690 (rank : 60) | θ value | 2.36792 (rank : 21) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8CFE5, Q80TC3, Q8BZY9 | Gene names | Btbd7, Kiaa1525 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | BTB/POZ domain-containing protein 7. | |||||
|
BHLH3_HUMAN
|
||||||
| NC score | 0.033766 (rank : 61) | θ value | 1.81305 (rank : 16) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9C0J9 | Gene names | BHLHB3, DEC2, SHARP1 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 3 (bHLHB3) (Differentially expressed in chondrocytes protein 2) (hDEC2) (Enhancer-of-split and hairy-related protein 1) (SHARP-1). | |||||
|
INSM1_MOUSE
|
||||||
| NC score | 0.031482 (rank : 62) | θ value | 1.38821 (rank : 13) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 678 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q63ZV0, Q9Z113 | Gene names | Insm1, Ia1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Insulinoma-associated protein 1 (Zinc finger protein IA-1). | |||||
|
SEPT9_HUMAN
|
||||||
| NC score | 0.029150 (rank : 63) | θ value | 4.03905 (rank : 28) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 706 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9UHD8, Q96QF3, Q96QF4, Q96QF5, Q9HA04, Q9UG40, Q9Y5W4 | Gene names | SEPT9, KIAA0991, MSF | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Septin-9 (MLL septin-like fusion protein) (MLL septin-like fusion protein MSF-A) (Ovarian/Breast septin) (Ov/Br septin) (Septin D1). | |||||
|
HAX1_MOUSE
|
||||||
| NC score | 0.026812 (rank : 64) | θ value | 5.27518 (rank : 29) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O35387 | Gene names | Hax1, Hs1bp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HS1-associating protein X-1 (HAX-1) (HS1-binding protein). | |||||
|
ZN687_HUMAN
|
||||||
| NC score | 0.026482 (rank : 65) | θ value | 1.06291 (rank : 12) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1244 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8N1G0, Q68DQ8, Q9H937, Q9P2A7 | Gene names | ZNF687, KIAA1441 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 687. | |||||
|
K0174_MOUSE
|
||||||
| NC score | 0.023042 (rank : 66) | θ value | 6.88961 (rank : 36) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9CX00, Q80U68 | Gene names | Kiaa0174 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0174. | |||||
|
NUP98_HUMAN
|
||||||
| NC score | 0.021042 (rank : 67) | θ value | 8.99809 (rank : 45) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P52948, Q8IUT2, Q8WYB0, Q96E54, Q9H3Q4, Q9NT02, Q9UF57, Q9UHX0, Q9Y6J4, Q9Y6J5 | Gene names | NUP98 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup98-Nup96 precursor [Contains: Nuclear pore complex protein Nup98 (Nucleoporin Nup98) (98 kDa nucleoporin); Nuclear pore complex protein Nup96 (Nucleoporin Nup96) (96 kDa nucleoporin)]. | |||||
|
KCNH6_HUMAN
|
||||||
| NC score | 0.019124 (rank : 68) | θ value | 1.81305 (rank : 17) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9H252, Q9BRD7 | Gene names | KCNH6, ERG2, HERG2 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium voltage-gated channel subfamily H member 6 (Voltage-gated potassium channel subunit Kv11.2) (Ether-a-go-go-related gene potassium channel 2) (Ether-a-go-go-related protein 2) (Eag-related protein 2). | |||||
|
NDF1_HUMAN
|
||||||
| NC score | 0.018943 (rank : 69) | θ value | 6.88961 (rank : 37) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 186 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q13562, O00343, Q13340, Q5U095, Q96TH0, Q99455, Q9UEC8 | Gene names | NEUROD1, NEUROD | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic differentiation factor 1 (NeuroD1) (NeuroD). | |||||
|
LNK_HUMAN
|
||||||
| NC score | 0.017191 (rank : 70) | θ value | 5.27518 (rank : 32) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UQQ2, O95184 | Gene names | LNK | |||
|
Domain Architecture |

|
|||||
| Description | Lymphocyte-specific adapter protein Lnk (Signal transduction protein Lnk) (Lymphocyte adapter protein). | |||||
|
SPEG_MOUSE
|
||||||
| NC score | 0.016487 (rank : 71) | θ value | 1.81305 (rank : 20) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1959 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q62407, Q3TPH8, Q6P5V1, Q80TF7, Q80ZN0, Q8BZF4, Q9EQJ5 | Gene names | Speg, Apeg1, Kiaa1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle-specific serine/threonine protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
CAC1A_MOUSE
|
||||||
| NC score | 0.014467 (rank : 72) | θ value | 2.36792 (rank : 22) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 267 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P97445 | Gene names | Cacna1a, Cach4, Cacn3, Cacnl1a4, Ccha1a | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent P/Q-type calcium channel subunit alpha-1A (Voltage- gated calcium channel subunit alpha Cav2.1) (Calcium channel, L type, alpha-1 polypeptide isoform 4) (Brain calcium channel I) (BI). | |||||
|
HCN3_MOUSE
|
||||||
| NC score | 0.013950 (rank : 73) | θ value | 5.27518 (rank : 31) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 112 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O88705 | Gene names | Hcn3, Hac3 | |||
|
Domain Architecture |

|
|||||
| Description | Potassium/sodium hyperpolarization-activated cyclic nucleotide-gated channel 3 (Hyperpolarization-activated cation channel 3) (HAC-3). | |||||
|
IRF8_HUMAN
|
||||||
| NC score | 0.011055 (rank : 74) | θ value | 6.88961 (rank : 35) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q02556 | Gene names | IRF8, ICSBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon regulatory factor 8 (IRF-8) (Interferon consensus sequence- binding protein) (ICSBP). | |||||
|
CSPG2_MOUSE
|
||||||
| NC score | 0.009448 (rank : 75) | θ value | 8.99809 (rank : 41) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 711 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q62059, Q62058, Q9CUU0 | Gene names | Cspg2 | |||
|
Domain Architecture |

|
|||||
| Description | Versican core protein precursor (Large fibroblast proteoglycan) (Chondroitin sulfate proteoglycan core protein 2) (PG-M). | |||||
|
MEGF8_HUMAN
|
||||||
| NC score | 0.006622 (rank : 76) | θ value | 8.99809 (rank : 43) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 528 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q7Z7M0, O75097 | Gene names | MEGF8, EGFL4, KIAA0817 | |||
|
Domain Architecture |

|
|||||
| Description | Multiple epidermal growth factor-like domains 8 (EGF-like domain- containing protein 4) (Multiple EGF-like domain protein 4). | |||||