Please be patient as the page loads
|
DCHS_MOUSE
|
||||||
| SwissProt Accessions | P23738 | Gene names | Hdc | |||
|
Domain Architecture |

|
|||||
| Description | Histidine decarboxylase (EC 4.1.1.22) (HDC). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
DCHS_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.997270 (rank : 2) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P19113 | Gene names | HDC | |||
|
Domain Architecture |

|
|||||
| Description | Histidine decarboxylase (EC 4.1.1.22) (HDC). | |||||
|
DCHS_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P23738 | Gene names | Hdc | |||
|
Domain Architecture |

|
|||||
| Description | Histidine decarboxylase (EC 4.1.1.22) (HDC). | |||||
|
DDC_HUMAN
|
||||||
| θ value | 6.59259e-152 (rank : 3) | NC score | 0.979186 (rank : 4) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P20711, Q16723 | Gene names | DDC, AADC | |||
|
Domain Architecture |

|
|||||
| Description | Aromatic-L-amino-acid decarboxylase (EC 4.1.1.28) (AADC) (DOPA decarboxylase) (DDC). | |||||
|
DDC_MOUSE
|
||||||
| θ value | 1.12453e-151 (rank : 4) | NC score | 0.980071 (rank : 3) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O88533 | Gene names | Ddc | |||
|
Domain Architecture |

|
|||||
| Description | Aromatic-L-amino-acid decarboxylase (EC 4.1.1.28) (AADC) (DOPA decarboxylase) (DDC). | |||||
|
DCE2_MOUSE
|
||||||
| θ value | 5.97985e-28 (rank : 5) | NC score | 0.733864 (rank : 6) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P48320, O35519 | Gene names | Gad2, Gad65 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate decarboxylase 2 (EC 4.1.1.15) (Glutamate decarboxylase 65 kDa isoform) (GAD-65) (65 kDa glutamic acid decarboxylase). | |||||
|
DCE2_HUMAN
|
||||||
| θ value | 7.80994e-28 (rank : 6) | NC score | 0.733996 (rank : 5) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q05329, Q9UD87 | Gene names | GAD2, GAD65 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate decarboxylase 2 (EC 4.1.1.15) (Glutamate decarboxylase 65 kDa isoform) (GAD-65) (65 kDa glutamic acid decarboxylase). | |||||
|
DCE1_HUMAN
|
||||||
| θ value | 7.30988e-26 (rank : 7) | NC score | 0.730461 (rank : 7) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q99259, Q9BU91, Q9UHH4 | Gene names | GAD1, GAD, GAD67 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate decarboxylase 1 (EC 4.1.1.15) (Glutamate decarboxylase 67 kDa isoform) (GAD-67) (67 kDa glutamic acid decarboxylase). | |||||
|
CSAD_HUMAN
|
||||||
| θ value | 4.73814e-25 (rank : 8) | NC score | 0.728723 (rank : 8) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Y600, Q9UNJ5, Q9Y601 | Gene names | CSAD, CSD | |||
|
Domain Architecture |

|
|||||
| Description | Cysteine sulfinic acid decarboxylase (EC 4.1.1.29) (Sulfinoalanine decarboxylase) (Cysteine-sulfinate decarboxylase). | |||||
|
DCE1_MOUSE
|
||||||
| θ value | 4.73814e-25 (rank : 9) | NC score | 0.725667 (rank : 9) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P48318, O08685 | Gene names | Gad1, Gad67 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate decarboxylase 1 (EC 4.1.1.15) (Glutamate decarboxylase 67 kDa isoform) (GAD-67) (67 kDa glutamic acid decarboxylase). | |||||
|
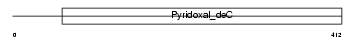
CSAD_MOUSE
|
||||||
| θ value | 5.23862e-24 (rank : 10) | NC score | 0.725506 (rank : 10) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9DBE0 | Gene names | Csad | |||
|
Domain Architecture |
|
|||||
| Description | Cysteine sulfinic acid decarboxylase (EC 4.1.1.29) (Sulfinoalanine decarboxylase) (Cysteine-sulfinate decarboxylase). | |||||
|
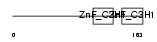
TTP_HUMAN
|
||||||
| θ value | 0.365318 (rank : 11) | NC score | 0.038858 (rank : 13) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P26651 | Gene names | ZFP36, G0S24, TIS11A, TTP | |||
|
Domain Architecture |
|
|||||
| Description | Tristetraproline (TTP) (Zinc finger protein 36 homolog) (Zfp-36) (TIS11A protein) (TIS11) (Growth factor-inducible nuclear protein NUP475) (G0/G1 switch regulatory protein 24). | |||||
|
TM16B_HUMAN
|
||||||
| θ value | 1.38821 (rank : 12) | NC score | 0.018137 (rank : 18) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NQ90 | Gene names | TMEM16B, C12orf3 | |||
|
Domain Architecture |

|
|||||
| Description | Transmembrane protein 16B. | |||||
|
USBP1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 13) | NC score | 0.025233 (rank : 15) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8N6Y0, Q8NBX7, Q96KH3, Q9BYI8 | Gene names | USHBP1, AIEBP, MCC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | USH1C-binding protein 1 (Usher syndrome type-1C protein-binding protein 1) (MCC-2) (AIE-75 binding protein). | |||||
|
NEB2_MOUSE
|
||||||
| θ value | 1.81305 (rank : 14) | NC score | 0.012919 (rank : 20) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 889 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6R891, Q8K0X7 | Gene names | Ppp1r9b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neurabin-2 (Neurabin-II) (Spinophilin) (Protein phosphatase 1 regulatory subunit 9B). | |||||
|
TGON1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 15) | NC score | 0.020428 (rank : 16) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q62313 | Gene names | Tgoln1, Ttgn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trans-Golgi network integral membrane protein 1 precursor (TGN38A). | |||||
|
KLHL6_HUMAN
|
||||||
| θ value | 3.0926 (rank : 16) | NC score | 0.004850 (rank : 28) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 192 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8WZ60, Q8N5I1, Q8N892 | Gene names | KLHL6 | |||
|
Domain Architecture |

|
|||||
| Description | Kelch-like protein 6. | |||||
|
RIN2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 17) | NC score | 0.018978 (rank : 17) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9D684, Q99K06 | Gene names | Rin2 | |||
|
Domain Architecture |

|
|||||
| Description | Ras and Rab interactor 2 (Ras interaction/interference protein 2). | |||||
|
SIGL5_MOUSE
|
||||||
| θ value | 4.03905 (rank : 18) | NC score | 0.007064 (rank : 27) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q920G3 | Gene names | Siglec5, Siglecf | |||
|
Domain Architecture |

|
|||||
| Description | Sialic acid-binding Ig-like lectin 5 precursor (Siglec-5) (Sialic acid-binding Ig-like lectin-F) (mSiglec-F). | |||||
|
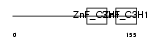
TTP_MOUSE
|
||||||
| θ value | 4.03905 (rank : 19) | NC score | 0.028112 (rank : 14) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P22893, P11520 | Gene names | Zfp36, Tis11, Tis11a | |||
|
Domain Architecture |
|
|||||
| Description | Tristetraproline (TTP) (Zinc finger protein 36) (Zfp-36) (TIS11A protein) (TIS11) (Growth factor-inducible nuclear protein NUP475) (TPA-induced sequence 11). | |||||
|
NEB2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 20) | NC score | 0.010967 (rank : 23) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 883 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q96SB3, Q8TCR9 | Gene names | PPP1R9B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neurabin-2 (Neurabin-II) (Spinophilin) (Protein phosphatase 1 regulatory subunit 9B). | |||||
|
THSD1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 21) | NC score | 0.010997 (rank : 22) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NS62, Q6P3U1, Q6UXZ2 | Gene names | THSD1, TMTSP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thrombospondin type-1 domain-containing protein 1 precursor (Transmembrane molecule with thrombospondin module). | |||||
|
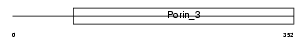
TOM40_HUMAN
|
||||||
| θ value | 5.27518 (rank : 22) | NC score | 0.012793 (rank : 21) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O96008, Q86VW4, Q8WY09, Q8WY10, Q8WY11, Q9BR95 | Gene names | TOMM40, PEREC1, TOM40 | |||
|
Domain Architecture |
|
|||||
| Description | Probable mitochondrial import receptor subunit TOM40 homolog (Translocase of outer membrane 40 kDa subunit homolog) (Haymaker protein) (p38.5). | |||||
|
FA47B_HUMAN
|
||||||
| θ value | 6.88961 (rank : 23) | NC score | 0.008939 (rank : 25) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 431 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8NA70, Q5JQN5, Q6PIG3 | Gene names | FAM47B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM47B. | |||||
|
KSR2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 24) | NC score | -0.000049 (rank : 29) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 920 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6VAB6, Q8N775 | Gene names | KSR2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinase suppressor of ras-2 (hKSR2). | |||||
|
RIN2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 25) | NC score | 0.016642 (rank : 19) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8WYP3, Q00425, Q5TFT8, Q9BQL3, Q9H071 | Gene names | RIN2, RASSF4 | |||
|
Domain Architecture |

|
|||||
| Description | Ras and Rab interactor 2 (Ras interaction/interference protein 2) (Ras inhibitor JC265) (Ras association domain family 4). | |||||
|
SYNJ2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 26) | NC score | 0.007377 (rank : 26) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O15056, Q5TA13, Q5TA16, Q5TA19, Q86XK0, Q8IZA8, Q9H226 | Gene names | SYNJ2, KIAA0348 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptojanin-2 (EC 3.1.3.36) (Synaptic inositol-1,4,5-trisphosphate 5- phosphatase 2). | |||||
|
UGTAP_MOUSE
|
||||||
| θ value | 8.99809 (rank : 27) | NC score | 0.010862 (rank : 24) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 4 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6P6M7, Q8BZS6 | Gene names | D5Ertd135e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UGA suppressor tRNA-associated protein. | |||||
|
SGPL1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 28) | NC score | 0.057840 (rank : 11) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O95470, Q7Z732, Q9ULG8, Q9UN89 | Gene names | SGPL1, KIAA1252 | |||
|
Domain Architecture |
|
|||||
| Description | Sphingosine-1-phosphate lyase 1 (EC 4.1.2.27) (SP-lyase) (hSPL) (Sphingosine-1-phosphate aldolase). | |||||
|
SGPL1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 29) | NC score | 0.056589 (rank : 12) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8R0X7, O54955, Q8C942 | Gene names | Sgpl1 | |||
|
Domain Architecture |

|
|||||
| Description | Sphingosine-1-phosphate lyase 1 (EC 4.1.2.27) (SP-lyase) (mSPL) (Sphingosine-1-phosphate aldolase). | |||||
|
DCHS_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P23738 | Gene names | Hdc | |||
|
Domain Architecture |

|
|||||
| Description | Histidine decarboxylase (EC 4.1.1.22) (HDC). | |||||
|
DCHS_HUMAN
|
||||||
| NC score | 0.997270 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P19113 | Gene names | HDC | |||
|
Domain Architecture |

|
|||||
| Description | Histidine decarboxylase (EC 4.1.1.22) (HDC). | |||||
|
DDC_MOUSE
|
||||||
| NC score | 0.980071 (rank : 3) | θ value | 1.12453e-151 (rank : 4) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O88533 | Gene names | Ddc | |||
|
Domain Architecture |

|
|||||
| Description | Aromatic-L-amino-acid decarboxylase (EC 4.1.1.28) (AADC) (DOPA decarboxylase) (DDC). | |||||
|
DDC_HUMAN
|
||||||
| NC score | 0.979186 (rank : 4) | θ value | 6.59259e-152 (rank : 3) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P20711, Q16723 | Gene names | DDC, AADC | |||
|
Domain Architecture |

|
|||||
| Description | Aromatic-L-amino-acid decarboxylase (EC 4.1.1.28) (AADC) (DOPA decarboxylase) (DDC). | |||||
|
DCE2_HUMAN
|
||||||
| NC score | 0.733996 (rank : 5) | θ value | 7.80994e-28 (rank : 6) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q05329, Q9UD87 | Gene names | GAD2, GAD65 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate decarboxylase 2 (EC 4.1.1.15) (Glutamate decarboxylase 65 kDa isoform) (GAD-65) (65 kDa glutamic acid decarboxylase). | |||||
|
DCE2_MOUSE
|
||||||
| NC score | 0.733864 (rank : 6) | θ value | 5.97985e-28 (rank : 5) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P48320, O35519 | Gene names | Gad2, Gad65 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate decarboxylase 2 (EC 4.1.1.15) (Glutamate decarboxylase 65 kDa isoform) (GAD-65) (65 kDa glutamic acid decarboxylase). | |||||
|
DCE1_HUMAN
|
||||||
| NC score | 0.730461 (rank : 7) | θ value | 7.30988e-26 (rank : 7) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q99259, Q9BU91, Q9UHH4 | Gene names | GAD1, GAD, GAD67 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate decarboxylase 1 (EC 4.1.1.15) (Glutamate decarboxylase 67 kDa isoform) (GAD-67) (67 kDa glutamic acid decarboxylase). | |||||
|
CSAD_HUMAN
|
||||||
| NC score | 0.728723 (rank : 8) | θ value | 4.73814e-25 (rank : 8) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Y600, Q9UNJ5, Q9Y601 | Gene names | CSAD, CSD | |||
|
Domain Architecture |

|
|||||
| Description | Cysteine sulfinic acid decarboxylase (EC 4.1.1.29) (Sulfinoalanine decarboxylase) (Cysteine-sulfinate decarboxylase). | |||||
|
DCE1_MOUSE
|
||||||
| NC score | 0.725667 (rank : 9) | θ value | 4.73814e-25 (rank : 9) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P48318, O08685 | Gene names | Gad1, Gad67 | |||
|
Domain Architecture |

|
|||||
| Description | Glutamate decarboxylase 1 (EC 4.1.1.15) (Glutamate decarboxylase 67 kDa isoform) (GAD-67) (67 kDa glutamic acid decarboxylase). | |||||
|
CSAD_MOUSE
|
||||||
| NC score | 0.725506 (rank : 10) | θ value | 5.23862e-24 (rank : 10) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9DBE0 | Gene names | Csad | |||
|
Domain Architecture |

|
|||||
| Description | Cysteine sulfinic acid decarboxylase (EC 4.1.1.29) (Sulfinoalanine decarboxylase) (Cysteine-sulfinate decarboxylase). | |||||
|
SGPL1_HUMAN
|
||||||
| NC score | 0.057840 (rank : 11) | θ value | θ > 10 (rank : 28) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O95470, Q7Z732, Q9ULG8, Q9UN89 | Gene names | SGPL1, KIAA1252 | |||
|
Domain Architecture |

|
|||||
| Description | Sphingosine-1-phosphate lyase 1 (EC 4.1.2.27) (SP-lyase) (hSPL) (Sphingosine-1-phosphate aldolase). | |||||
|
SGPL1_MOUSE
|
||||||
| NC score | 0.056589 (rank : 12) | θ value | θ > 10 (rank : 29) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8R0X7, O54955, Q8C942 | Gene names | Sgpl1 | |||
|
Domain Architecture |

|
|||||
| Description | Sphingosine-1-phosphate lyase 1 (EC 4.1.2.27) (SP-lyase) (mSPL) (Sphingosine-1-phosphate aldolase). | |||||
|
TTP_HUMAN
|
||||||
| NC score | 0.038858 (rank : 13) | θ value | 0.365318 (rank : 11) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 86 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P26651 | Gene names | ZFP36, G0S24, TIS11A, TTP | |||
|
Domain Architecture |

|
|||||
| Description | Tristetraproline (TTP) (Zinc finger protein 36 homolog) (Zfp-36) (TIS11A protein) (TIS11) (Growth factor-inducible nuclear protein NUP475) (G0/G1 switch regulatory protein 24). | |||||
|
TTP_MOUSE
|
||||||
| NC score | 0.028112 (rank : 14) | θ value | 4.03905 (rank : 19) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P22893, P11520 | Gene names | Zfp36, Tis11, Tis11a | |||
|
Domain Architecture |

|
|||||
| Description | Tristetraproline (TTP) (Zinc finger protein 36) (Zfp-36) (TIS11A protein) (TIS11) (Growth factor-inducible nuclear protein NUP475) (TPA-induced sequence 11). | |||||
|
USBP1_HUMAN
|
||||||
| NC score | 0.025233 (rank : 15) | θ value | 1.38821 (rank : 13) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8N6Y0, Q8NBX7, Q96KH3, Q9BYI8 | Gene names | USHBP1, AIEBP, MCC2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | USH1C-binding protein 1 (Usher syndrome type-1C protein-binding protein 1) (MCC-2) (AIE-75 binding protein). | |||||
|
TGON1_MOUSE
|
||||||
| NC score | 0.020428 (rank : 16) | θ value | 1.81305 (rank : 15) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q62313 | Gene names | Tgoln1, Ttgn1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trans-Golgi network integral membrane protein 1 precursor (TGN38A). | |||||
|
RIN2_MOUSE
|
||||||
| NC score | 0.018978 (rank : 17) | θ value | 3.0926 (rank : 17) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9D684, Q99K06 | Gene names | Rin2 | |||
|
Domain Architecture |

|
|||||
| Description | Ras and Rab interactor 2 (Ras interaction/interference protein 2). | |||||
|
TM16B_HUMAN
|
||||||
| NC score | 0.018137 (rank : 18) | θ value | 1.38821 (rank : 12) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NQ90 | Gene names | TMEM16B, C12orf3 | |||
|
Domain Architecture |

|
|||||
| Description | Transmembrane protein 16B. | |||||
|
RIN2_HUMAN
|
||||||
| NC score | 0.016642 (rank : 19) | θ value | 6.88961 (rank : 25) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8WYP3, Q00425, Q5TFT8, Q9BQL3, Q9H071 | Gene names | RIN2, RASSF4 | |||
|
Domain Architecture |

|
|||||
| Description | Ras and Rab interactor 2 (Ras interaction/interference protein 2) (Ras inhibitor JC265) (Ras association domain family 4). | |||||
|
NEB2_MOUSE
|
||||||
| NC score | 0.012919 (rank : 20) | θ value | 1.81305 (rank : 14) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 889 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6R891, Q8K0X7 | Gene names | Ppp1r9b | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neurabin-2 (Neurabin-II) (Spinophilin) (Protein phosphatase 1 regulatory subunit 9B). | |||||
|
TOM40_HUMAN
|
||||||
| NC score | 0.012793 (rank : 21) | θ value | 5.27518 (rank : 22) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O96008, Q86VW4, Q8WY09, Q8WY10, Q8WY11, Q9BR95 | Gene names | TOMM40, PEREC1, TOM40 | |||
|
Domain Architecture |

|
|||||
| Description | Probable mitochondrial import receptor subunit TOM40 homolog (Translocase of outer membrane 40 kDa subunit homolog) (Haymaker protein) (p38.5). | |||||
|
THSD1_HUMAN
|
||||||
| NC score | 0.010997 (rank : 22) | θ value | 5.27518 (rank : 21) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NS62, Q6P3U1, Q6UXZ2 | Gene names | THSD1, TMTSP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thrombospondin type-1 domain-containing protein 1 precursor (Transmembrane molecule with thrombospondin module). | |||||
|
NEB2_HUMAN
|
||||||
| NC score | 0.010967 (rank : 23) | θ value | 5.27518 (rank : 20) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 883 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q96SB3, Q8TCR9 | Gene names | PPP1R9B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Neurabin-2 (Neurabin-II) (Spinophilin) (Protein phosphatase 1 regulatory subunit 9B). | |||||
|
UGTAP_MOUSE
|
||||||
| NC score | 0.010862 (rank : 24) | θ value | 8.99809 (rank : 27) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 4 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6P6M7, Q8BZS6 | Gene names | D5Ertd135e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UGA suppressor tRNA-associated protein. | |||||
|
FA47B_HUMAN
|
||||||
| NC score | 0.008939 (rank : 25) | θ value | 6.88961 (rank : 23) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 431 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8NA70, Q5JQN5, Q6PIG3 | Gene names | FAM47B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein FAM47B. | |||||
|
SYNJ2_HUMAN
|
||||||
| NC score | 0.007377 (rank : 26) | θ value | 6.88961 (rank : 26) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O15056, Q5TA13, Q5TA16, Q5TA19, Q86XK0, Q8IZA8, Q9H226 | Gene names | SYNJ2, KIAA0348 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptojanin-2 (EC 3.1.3.36) (Synaptic inositol-1,4,5-trisphosphate 5- phosphatase 2). | |||||
|
SIGL5_MOUSE
|
||||||
| NC score | 0.007064 (rank : 27) | θ value | 4.03905 (rank : 18) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 271 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q920G3 | Gene names | Siglec5, Siglecf | |||
|
Domain Architecture |

|
|||||
| Description | Sialic acid-binding Ig-like lectin 5 precursor (Siglec-5) (Sialic acid-binding Ig-like lectin-F) (mSiglec-F). | |||||
|
KLHL6_HUMAN
|
||||||
| NC score | 0.004850 (rank : 28) | θ value | 3.0926 (rank : 16) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 192 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8WZ60, Q8N5I1, Q8N892 | Gene names | KLHL6 | |||
|
Domain Architecture |

|
|||||
| Description | Kelch-like protein 6. | |||||
|
KSR2_HUMAN
|
||||||
| NC score | -0.000049 (rank : 29) | θ value | 6.88961 (rank : 24) | |||
| Query Neighborhood Hits | 27 | Target Neighborhood Hits | 920 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6VAB6, Q8N775 | Gene names | KSR2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinase suppressor of ras-2 (hKSR2). | |||||