Please be patient as the page loads
|
CRYL1_MOUSE
|
||||||
| SwissProt Accessions | Q99KP3, Q542R9, Q8R4W7 | Gene names | Cryl1, Cry | |||
|
Domain Architecture |
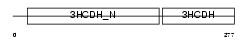
|
|||||
| Description | Lambda-crystallin homolog. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
CRYL1_HUMAN
|
||||||
| θ value | 5.77539e-148 (rank : 1) | NC score | 0.985402 (rank : 2) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Y2S2, Q7Z4Z9 | Gene names | CRYL1, CRY | |||
|
Domain Architecture |
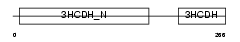
|
|||||
| Description | Lambda-crystallin homolog. | |||||
|
CRYL1_MOUSE
|
||||||
| θ value | 1.57279e-145 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q99KP3, Q542R9, Q8R4W7 | Gene names | Cryl1, Cry | |||
|
Domain Architecture |

|
|||||
| Description | Lambda-crystallin homolog. | |||||
|
HCDH_MOUSE
|
||||||
| θ value | 3.88503e-19 (rank : 3) | NC score | 0.674546 (rank : 3) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q61425, Q3TF75, Q3THK8, Q3UFI0, Q8K149, Q925U9 | Gene names | Hadhsc, Hadh, Mschad, Schad | |||
|
Domain Architecture |
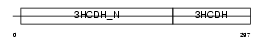
|
|||||
| Description | Short chain 3-hydroxyacyl-CoA dehydrogenase, mitochondrial precursor (EC 1.1.1.35) (HCDH) (Medium and short chain L-3-hydroxyacyl-coenzyme A dehydrogenase). | |||||
|
HCDH_HUMAN
|
||||||
| θ value | 5.07402e-19 (rank : 4) | NC score | 0.671567 (rank : 4) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q16836, O00324, O00397, O00753 | Gene names | HADHSC, HAD, SCHAD | |||
|
Domain Architecture |
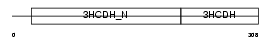
|
|||||
| Description | Short chain 3-hydroxyacyl-CoA dehydrogenase, mitochondrial precursor (EC 1.1.1.35) (HCDH) (Medium and short chain L-3-hydroxyacyl-coenzyme A dehydrogenase). | |||||
|
ECHP_MOUSE
|
||||||
| θ value | 2.43908e-13 (rank : 5) | NC score | 0.469491 (rank : 5) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9DBM2 | Gene names | Ehhadh | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisomal bifunctional enzyme (PBE) (PBFE) [Includes: Enoyl-CoA hydratase (EC 4.2.1.17); 3,2-trans-enoyl-CoA isomerase (EC 5.3.3.8); 3-hydroxyacyl-CoA dehydrogenase (EC 1.1.1.35)]. | |||||
|
ECHP_HUMAN
|
||||||
| θ value | 1.69304e-06 (rank : 6) | NC score | 0.414117 (rank : 6) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q08426 | Gene names | EHHADH, ECHD | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisomal bifunctional enzyme (PBE) (PBFE) [Includes: Enoyl-CoA hydratase (EC 4.2.1.17); 3,2-trans-enoyl-CoA isomerase (EC 5.3.3.8); 3-hydroxyacyl-CoA dehydrogenase (EC 1.1.1.35)]. | |||||
|
EVPL_MOUSE
|
||||||
| θ value | 0.279714 (rank : 7) | NC score | 0.021530 (rank : 18) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 952 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9D952 | Gene names | Evpl | |||
|
Domain Architecture |

|
|||||
| Description | Envoplakin (p210) (210 kDa cornified envelope precursor protein). | |||||
|
COA2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 8) | NC score | 0.017827 (rank : 19) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O00763, Q16852 | Gene names | ACACB, ACC2, ACCB | |||
|
Domain Architecture |

|
|||||
| Description | Acetyl-CoA carboxylase 2 (EC 6.4.1.2) (ACC-beta) [Includes: Biotin carboxylase (EC 6.3.4.14)]. | |||||
|
KIF4A_MOUSE
|
||||||
| θ value | 5.27518 (rank : 9) | NC score | 0.006071 (rank : 20) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 1033 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P33174 | Gene names | Kif4a, Kif4, Kns4 | |||
|
Domain Architecture |

|
|||||
| Description | Chromosome-associated kinesin KIF4A (Chromokinesin). | |||||
|
INSI1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 10) | NC score | 0.026454 (rank : 16) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O15503, Q9BUV5 | Gene names | INSIG1 | |||
|
Domain Architecture |
|
|||||
| Description | Insulin-induced gene 1 protein (INSIG-1). | |||||
|
INSI1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 11) | NC score | 0.026116 (rank : 17) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BGI3, Q3UAX9 | Gene names | Insig1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Insulin-induced gene 1 protein (INSIG-1). | |||||
|
TRIPB_HUMAN
|
||||||
| θ value | 8.99809 (rank : 12) | NC score | 0.004882 (rank : 21) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 1587 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q15643, O14689, O15154, O95949 | Gene names | TRIP11, CEV14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid receptor-interacting protein 11 (TRIP-11) (Golgi-associated microtubule-binding protein 210) (GMAP-210) (Trip230) (Clonal evolution-related gene on chromosome 14). | |||||
|
AUHM_MOUSE
|
||||||
| θ value | θ > 10 (rank : 13) | NC score | 0.063457 (rank : 11) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9JLZ3, Q80YD7, Q8QZS0, Q9CY78, Q9D155 | Gene names | Auh | |||
|
Domain Architecture |
|
|||||
| Description | Methylglutaconyl-CoA hydratase, mitochondrial precursor (EC 4.2.1.18) (AU-specific RNA-binding enoyl-CoA hydratase) (AU-binding enoyl-CoA hydratase) (muAUH). | |||||
|
AUMH_HUMAN
|
||||||
| θ value | θ > 10 (rank : 14) | NC score | 0.061993 (rank : 12) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q13825, Q8WUE4 | Gene names | AUH | |||
|
Domain Architecture |

|
|||||
| Description | Methylglutaconyl-CoA hydratase, mitochondrial precursor (EC 4.2.1.18) (AU-specific RNA-binding enoyl-CoA hydratase) (AU-binding protein/enoyl-CoA hydratase). | |||||
|
D3D2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 15) | NC score | 0.059664 (rank : 13) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P42126, Q13290, Q7Z2L6, Q9BUB8, Q9BW05 | Gene names | DCI | |||
|
Domain Architecture |
|
|||||
| Description | 3,2-trans-enoyl-CoA isomerase, mitochondrial precursor (EC 5.3.3.8) (Dodecenoyl-CoA isomerase) (Delta(3),delta(2)-enoyl-CoA isomerase) (D3,D2-enoyl-CoA isomerase). | |||||
|
D3D2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 16) | NC score | 0.067006 (rank : 10) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P42125 | Gene names | Dci | |||
|
Domain Architecture |
|
|||||
| Description | 3,2-trans-enoyl-CoA isomerase, mitochondrial precursor (EC 5.3.3.8) (Dodecenoyl-CoA isomerase) (Delta(3),delta(2)-enoyl-CoA isomerase) (D3,D2-enoyl-CoA isomerase). | |||||
|
ECH1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 17) | NC score | 0.057155 (rank : 14) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q13011, Q8WVX0, Q96EZ9 | Gene names | ECH1 | |||
|
Domain Architecture |
|
|||||
| Description | Delta3,5-delta2,4-dienoyl-CoA isomerase, mitochondrial precursor (EC 5.3.3.-). | |||||
|
ECH1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 18) | NC score | 0.051020 (rank : 15) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O35459, Q5M8P6 | Gene names | Ech1 | |||
|
Domain Architecture |
|
|||||
| Description | Delta3,5-delta2,4-dienoyl-CoA isomerase, mitochondrial precursor (EC 5.3.3.-). | |||||
|
ECHA_HUMAN
|
||||||
| θ value | θ > 10 (rank : 19) | NC score | 0.313764 (rank : 7) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P40939, Q16679, Q96GT7 | Gene names | HADHA, HADH | |||
|
Domain Architecture |

|
|||||
| Description | Trifunctional enzyme subunit alpha, mitochondrial precursor (TP-alpha) (78 kDa gastrin-binding protein) [Includes: Long-chain enoyl-CoA hydratase (EC 4.2.1.17); Long chain 3-hydroxyacyl-CoA dehydrogenase (EC 1.1.1.211)]. | |||||
|
ECHM_HUMAN
|
||||||
| θ value | θ > 10 (rank : 20) | NC score | 0.073627 (rank : 9) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P30084, O00739, Q5VWY1, Q96H54 | Gene names | ECHS1 | |||
|
Domain Architecture |
|
|||||
| Description | Enoyl-CoA hydratase, mitochondrial precursor (EC 4.2.1.17) (Short chain enoyl-CoA hydratase) (SCEH) (Enoyl-CoA hydratase 1). | |||||
|
ECHM_MOUSE
|
||||||
| θ value | θ > 10 (rank : 21) | NC score | 0.075837 (rank : 8) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BH95, Q3TJK2, Q6GQS2, Q6PEN1, Q99LX7 | Gene names | Echs1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Enoyl-CoA hydratase, mitochondrial precursor (EC 4.2.1.17) (Short chain enoyl-CoA hydratase) (SCEH) (Enoyl-CoA hydratase 1). | |||||
|
CRYL1_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 1.57279e-145 (rank : 2) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q99KP3, Q542R9, Q8R4W7 | Gene names | Cryl1, Cry | |||
|
Domain Architecture |

|
|||||
| Description | Lambda-crystallin homolog. | |||||
|
CRYL1_HUMAN
|
||||||
| NC score | 0.985402 (rank : 2) | θ value | 5.77539e-148 (rank : 1) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Y2S2, Q7Z4Z9 | Gene names | CRYL1, CRY | |||
|
Domain Architecture |

|
|||||
| Description | Lambda-crystallin homolog. | |||||
|
HCDH_MOUSE
|
||||||
| NC score | 0.674546 (rank : 3) | θ value | 3.88503e-19 (rank : 3) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q61425, Q3TF75, Q3THK8, Q3UFI0, Q8K149, Q925U9 | Gene names | Hadhsc, Hadh, Mschad, Schad | |||
|
Domain Architecture |

|
|||||
| Description | Short chain 3-hydroxyacyl-CoA dehydrogenase, mitochondrial precursor (EC 1.1.1.35) (HCDH) (Medium and short chain L-3-hydroxyacyl-coenzyme A dehydrogenase). | |||||
|
HCDH_HUMAN
|
||||||
| NC score | 0.671567 (rank : 4) | θ value | 5.07402e-19 (rank : 4) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q16836, O00324, O00397, O00753 | Gene names | HADHSC, HAD, SCHAD | |||
|
Domain Architecture |

|
|||||
| Description | Short chain 3-hydroxyacyl-CoA dehydrogenase, mitochondrial precursor (EC 1.1.1.35) (HCDH) (Medium and short chain L-3-hydroxyacyl-coenzyme A dehydrogenase). | |||||
|
ECHP_MOUSE
|
||||||
| NC score | 0.469491 (rank : 5) | θ value | 2.43908e-13 (rank : 5) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9DBM2 | Gene names | Ehhadh | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisomal bifunctional enzyme (PBE) (PBFE) [Includes: Enoyl-CoA hydratase (EC 4.2.1.17); 3,2-trans-enoyl-CoA isomerase (EC 5.3.3.8); 3-hydroxyacyl-CoA dehydrogenase (EC 1.1.1.35)]. | |||||
|
ECHP_HUMAN
|
||||||
| NC score | 0.414117 (rank : 6) | θ value | 1.69304e-06 (rank : 6) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q08426 | Gene names | EHHADH, ECHD | |||
|
Domain Architecture |

|
|||||
| Description | Peroxisomal bifunctional enzyme (PBE) (PBFE) [Includes: Enoyl-CoA hydratase (EC 4.2.1.17); 3,2-trans-enoyl-CoA isomerase (EC 5.3.3.8); 3-hydroxyacyl-CoA dehydrogenase (EC 1.1.1.35)]. | |||||
|
ECHA_HUMAN
|
||||||
| NC score | 0.313764 (rank : 7) | θ value | θ > 10 (rank : 19) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P40939, Q16679, Q96GT7 | Gene names | HADHA, HADH | |||
|
Domain Architecture |

|
|||||
| Description | Trifunctional enzyme subunit alpha, mitochondrial precursor (TP-alpha) (78 kDa gastrin-binding protein) [Includes: Long-chain enoyl-CoA hydratase (EC 4.2.1.17); Long chain 3-hydroxyacyl-CoA dehydrogenase (EC 1.1.1.211)]. | |||||
|
ECHM_MOUSE
|
||||||
| NC score | 0.075837 (rank : 8) | θ value | θ > 10 (rank : 21) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BH95, Q3TJK2, Q6GQS2, Q6PEN1, Q99LX7 | Gene names | Echs1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Enoyl-CoA hydratase, mitochondrial precursor (EC 4.2.1.17) (Short chain enoyl-CoA hydratase) (SCEH) (Enoyl-CoA hydratase 1). | |||||
|
ECHM_HUMAN
|
||||||
| NC score | 0.073627 (rank : 9) | θ value | θ > 10 (rank : 20) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P30084, O00739, Q5VWY1, Q96H54 | Gene names | ECHS1 | |||
|
Domain Architecture |

|
|||||
| Description | Enoyl-CoA hydratase, mitochondrial precursor (EC 4.2.1.17) (Short chain enoyl-CoA hydratase) (SCEH) (Enoyl-CoA hydratase 1). | |||||
|
D3D2_MOUSE
|
||||||
| NC score | 0.067006 (rank : 10) | θ value | θ > 10 (rank : 16) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P42125 | Gene names | Dci | |||
|
Domain Architecture |

|
|||||
| Description | 3,2-trans-enoyl-CoA isomerase, mitochondrial precursor (EC 5.3.3.8) (Dodecenoyl-CoA isomerase) (Delta(3),delta(2)-enoyl-CoA isomerase) (D3,D2-enoyl-CoA isomerase). | |||||
|
AUHM_MOUSE
|
||||||
| NC score | 0.063457 (rank : 11) | θ value | θ > 10 (rank : 13) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9JLZ3, Q80YD7, Q8QZS0, Q9CY78, Q9D155 | Gene names | Auh | |||
|
Domain Architecture |

|
|||||
| Description | Methylglutaconyl-CoA hydratase, mitochondrial precursor (EC 4.2.1.18) (AU-specific RNA-binding enoyl-CoA hydratase) (AU-binding enoyl-CoA hydratase) (muAUH). | |||||
|
AUMH_HUMAN
|
||||||
| NC score | 0.061993 (rank : 12) | θ value | θ > 10 (rank : 14) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q13825, Q8WUE4 | Gene names | AUH | |||
|
Domain Architecture |

|
|||||
| Description | Methylglutaconyl-CoA hydratase, mitochondrial precursor (EC 4.2.1.18) (AU-specific RNA-binding enoyl-CoA hydratase) (AU-binding protein/enoyl-CoA hydratase). | |||||
|
D3D2_HUMAN
|
||||||
| NC score | 0.059664 (rank : 13) | θ value | θ > 10 (rank : 15) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P42126, Q13290, Q7Z2L6, Q9BUB8, Q9BW05 | Gene names | DCI | |||
|
Domain Architecture |

|
|||||
| Description | 3,2-trans-enoyl-CoA isomerase, mitochondrial precursor (EC 5.3.3.8) (Dodecenoyl-CoA isomerase) (Delta(3),delta(2)-enoyl-CoA isomerase) (D3,D2-enoyl-CoA isomerase). | |||||
|
ECH1_HUMAN
|
||||||
| NC score | 0.057155 (rank : 14) | θ value | θ > 10 (rank : 17) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q13011, Q8WVX0, Q96EZ9 | Gene names | ECH1 | |||
|
Domain Architecture |

|
|||||
| Description | Delta3,5-delta2,4-dienoyl-CoA isomerase, mitochondrial precursor (EC 5.3.3.-). | |||||
|
ECH1_MOUSE
|
||||||
| NC score | 0.051020 (rank : 15) | θ value | θ > 10 (rank : 18) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O35459, Q5M8P6 | Gene names | Ech1 | |||
|
Domain Architecture |

|
|||||
| Description | Delta3,5-delta2,4-dienoyl-CoA isomerase, mitochondrial precursor (EC 5.3.3.-). | |||||
|
INSI1_HUMAN
|
||||||
| NC score | 0.026454 (rank : 16) | θ value | 8.99809 (rank : 10) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O15503, Q9BUV5 | Gene names | INSIG1 | |||
|
Domain Architecture |

|
|||||
| Description | Insulin-induced gene 1 protein (INSIG-1). | |||||
|
INSI1_MOUSE
|
||||||
| NC score | 0.026116 (rank : 17) | θ value | 8.99809 (rank : 11) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BGI3, Q3UAX9 | Gene names | Insig1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Insulin-induced gene 1 protein (INSIG-1). | |||||
|
EVPL_MOUSE
|
||||||
| NC score | 0.021530 (rank : 18) | θ value | 0.279714 (rank : 7) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 952 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9D952 | Gene names | Evpl | |||
|
Domain Architecture |

|
|||||
| Description | Envoplakin (p210) (210 kDa cornified envelope precursor protein). | |||||
|
COA2_HUMAN
|
||||||
| NC score | 0.017827 (rank : 19) | θ value | 4.03905 (rank : 8) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 184 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O00763, Q16852 | Gene names | ACACB, ACC2, ACCB | |||
|
Domain Architecture |

|
|||||
| Description | Acetyl-CoA carboxylase 2 (EC 6.4.1.2) (ACC-beta) [Includes: Biotin carboxylase (EC 6.3.4.14)]. | |||||
|
KIF4A_MOUSE
|
||||||
| NC score | 0.006071 (rank : 20) | θ value | 5.27518 (rank : 9) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 1033 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P33174 | Gene names | Kif4a, Kif4, Kns4 | |||
|
Domain Architecture |

|
|||||
| Description | Chromosome-associated kinesin KIF4A (Chromokinesin). | |||||
|
TRIPB_HUMAN
|
||||||
| NC score | 0.004882 (rank : 21) | θ value | 8.99809 (rank : 12) | |||
| Query Neighborhood Hits | 12 | Target Neighborhood Hits | 1587 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q15643, O14689, O15154, O95949 | Gene names | TRIP11, CEV14 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyroid receptor-interacting protein 11 (TRIP-11) (Golgi-associated microtubule-binding protein 210) (GMAP-210) (Trip230) (Clonal evolution-related gene on chromosome 14). | |||||