Please be patient as the page loads
|
CIKS_HUMAN
|
||||||
| SwissProt Accessions | O43734, Q5R3A3, Q9H5W2, Q9H6Y3, Q9NS14, Q9UG72 | Gene names | TRAF3IP2, C6orf4, C6orf5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Adapter protein CIKS (Connection to IKK and SAPK/JNK) (TRAF3- interacting protein 2) (Nuclear factor NF-kappa-B activator 1) (ACT1). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
CIKS_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | O43734, Q5R3A3, Q9H5W2, Q9H6Y3, Q9NS14, Q9UG72 | Gene names | TRAF3IP2, C6orf4, C6orf5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Adapter protein CIKS (Connection to IKK and SAPK/JNK) (TRAF3- interacting protein 2) (Nuclear factor NF-kappa-B activator 1) (ACT1). | |||||
|
RGS3_MOUSE
|
||||||
| θ value | 0.00298849 (rank : 2) | NC score | 0.077284 (rank : 10) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
PHC2_HUMAN
|
||||||
| θ value | 0.00665767 (rank : 3) | NC score | 0.097016 (rank : 5) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8IXK0, Q5T0C1, Q6NUJ6, Q6ZQR1, Q8N306, Q8TAG8, Q96BL4, Q9Y4Y7 | Gene names | PHC2, EDR2, PH2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polyhomeotic-like protein 2 (hPH2) (Early development regulatory protein 2). | |||||
|
ADA12_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 4) | NC score | 0.039409 (rank : 29) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 474 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q61824 | Gene names | Adam12, Mltna | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 12 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 12) (Meltrin alpha). | |||||
|
CASL_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 5) | NC score | 0.111155 (rank : 4) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q14511 | Gene names | NEDD9, CASL | |||
|
Domain Architecture |
|
|||||
| Description | Enhancer of filamentation 1 (HEF1) (CRK-associated substrate-related protein) (CAS-L) (CasL) (p105) (Protein NEDD9) (NY-REN-12 antigen). | |||||
|
MLL4_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 6) | NC score | 0.089833 (rank : 8) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 722 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9UMN6, O15022, O95836, Q96GP2, Q96IP3, Q9UK25, Q9Y668, Q9Y669 | Gene names | MLL4, HRX2, KIAA0304, MLL2, TRX2 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 4 (Trithorax homolog 2). | |||||
|
ARS2_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 7) | NC score | 0.143152 (rank : 2) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 186 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9BXP5, O95808, Q9BWP6, Q9BXP4 | Gene names | ARS2, ASR2 | |||
|
Domain Architecture |
|
|||||
| Description | Arsenite-resistance protein 2. | |||||
|
ARS2_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 8) | NC score | 0.141987 (rank : 3) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q99MR6, Q99MR4, Q99MR5, Q99MR7 | Gene names | Ars2, Asr2 | |||
|
Domain Architecture |

|
|||||
| Description | Arsenite-resistance protein 2. | |||||
|
CASL_MOUSE
|
||||||
| θ value | 0.125558 (rank : 9) | NC score | 0.096943 (rank : 6) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O35177, Q8BJL8, Q8BK90, Q8BL52, Q8BM94, Q8BMI9, Q99KE7 | Gene names | Nedd9, Casl | |||
|
Domain Architecture |
|
|||||
| Description | Enhancer of filamentation 1 (MEF1) (CRK-associated substrate-related protein) (CAS-L) (p105) (Protein NEDD9) (Neural precursor cell expressed developmentally down-regulated protein 9). | |||||
|
PAR3L_HUMAN
|
||||||
| θ value | 0.279714 (rank : 10) | NC score | 0.049587 (rank : 19) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 297 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8TEW8, Q8IUC7, Q8IUC9, Q96DK9, Q96N09, Q96NX6, Q96NX7, Q96Q29 | Gene names | PARD3B, ALS2CR19, PAR3B, PAR3L | |||
|
Domain Architecture |

|
|||||
| Description | Partitioning-defective 3 homolog B (PAR3-beta) (Partitioning-defective 3-like protein) (PAR3-L protein) (Amyotrophic lateral sclerosis 2 chromosome region candidate gene 19 protein). | |||||
|
LRBA_HUMAN
|
||||||
| θ value | 0.365318 (rank : 11) | NC score | 0.045888 (rank : 22) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P50851, Q9H2U3, Q9H2U4 | Gene names | LRBA, BGL, CDC4L, LBA | |||
|
Domain Architecture |

|
|||||
| Description | Lipopolysaccharide-responsive and beige-like anchor protein (CDC4-like protein) (Beige-like protein). | |||||
|
WNK2_HUMAN
|
||||||
| θ value | 0.365318 (rank : 12) | NC score | 0.023791 (rank : 47) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1407 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9Y3S1, Q8IY36, Q9C0A3, Q9H3P4 | Gene names | WNK2, KIAA1760, PRKWNK2 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK2 (EC 2.7.11.1) (Protein kinase with no lysine 2) (Protein kinase, lysine-deficient 2). | |||||
|
FOXB2_MOUSE
|
||||||
| θ value | 0.47712 (rank : 13) | NC score | 0.025868 (rank : 45) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q64733 | Gene names | Foxb2, Fkh4 | |||
|
Domain Architecture |
|
|||||
| Description | Forkhead box protein B2 (Transcription factor FKH-4). | |||||
|
PHLA1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 14) | NC score | 0.095013 (rank : 7) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8WV24, Q15184, Q2TAN2, Q9NZ17 | Gene names | PHLDA1, PHRIP, TDAG51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology-like domain family A member 1 (T-cell death- associated gene 51 protein) (Apoptosis-associated nuclear protein) (Proline- and histidine-rich protein) (Proline- and glutamine-rich protein) (PQ-rich protein). | |||||
|
JHD2C_HUMAN
|
||||||
| θ value | 0.62314 (rank : 15) | NC score | 0.038503 (rank : 30) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q15652, Q5SQZ8, Q5SQZ9, Q5SR00, Q7Z3E7, Q8N3U0, Q96KB9, Q9P2G7 | Gene names | JMJD1C, JHDM2C, KIAA1380, TRIP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable JmjC domain-containing histone demethylation protein 2C (EC 1.14.11.-) (Jumonji domain-containing protein 1C) (Thyroid receptor-interacting protein 8) (TRIP-8). | |||||
|
SELPL_MOUSE
|
||||||
| θ value | 0.62314 (rank : 16) | NC score | 0.062395 (rank : 12) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 588 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q62170 | Gene names | Selplg, Selp1, Selpl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | P-selectin glycoprotein ligand 1 precursor (PSGL-1) (Selectin P ligand). | |||||
|
MLL3_HUMAN
|
||||||
| θ value | 0.813845 (rank : 17) | NC score | 0.055596 (rank : 16) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
AMELX_HUMAN
|
||||||
| θ value | 1.38821 (rank : 18) | NC score | 0.062000 (rank : 13) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q99217, Q96NW6, Q9UCA7 | Gene names | AMELX, AMG, AMGX | |||
|
Domain Architecture |

|
|||||
| Description | Amelogenin, X isoform precursor. | |||||
|
PCLO_MOUSE
|
||||||
| θ value | 1.38821 (rank : 19) | NC score | 0.047345 (rank : 21) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
PHLA1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 20) | NC score | 0.080697 (rank : 9) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q62392, Q3TY48, Q3UG87 | Gene names | Phlda1, Tdag51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology-like domain family A member 1 (T-cell death- associated gene 51 protein) (Proline- and glutamine-rich protein) (PQR protein). | |||||
|
PLEC1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 21) | NC score | 0.013084 (rank : 57) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1326 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q15149, Q15148, Q16640 | Gene names | PLEC1 | |||
|
Domain Architecture |

|
|||||
| Description | Plectin-1 (PLTN) (PCN) (Hemidesmosomal protein 1) (HD1). | |||||
|
TMC8_HUMAN
|
||||||
| θ value | 1.38821 (rank : 22) | NC score | 0.042591 (rank : 25) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8IU68, Q8IWU7, Q8N358, Q8NF04 | Gene names | TMC8, EVER2, EVIN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane channel-like protein 8 (Epidermodysplasia verruciformis protein 2). | |||||
|
AKAP2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 23) | NC score | 0.037988 (rank : 31) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 507 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Y2D5, Q9UG26 | Gene names | AKAP2, KIAA0920 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 2 (Protein kinase A-anchoring protein 2) (PRKA2) (AKAP-2). | |||||
|
MAST4_HUMAN
|
||||||
| θ value | 1.81305 (rank : 24) | NC score | 0.011069 (rank : 58) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1804 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O15021, Q6ZN07, Q96LY3 | Gene names | MAST4, KIAA0303 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated serine/threonine-protein kinase 4 (EC 2.7.11.1). | |||||
|
SYNM_HUMAN
|
||||||
| θ value | 1.81305 (rank : 25) | NC score | 0.040561 (rank : 27) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96I59 | Gene names | NARS2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable asparaginyl-tRNA synthetase, mitochondrial precursor (EC 6.1.1.22) (Asparagine--tRNA ligase) (AsnRS). | |||||
|
TMBI1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 26) | NC score | 0.041951 (rank : 26) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BJZ3, Q8CH91, Q99KB6 | Gene names | Tmbim1, Recs1 | |||
|
Domain Architecture |
|
|||||
| Description | Transmembrane BAX inhibitor motif-containing protein 1 (RECS1 protein) (Responsive to centrifugal force and shear stress gene 1 protein). | |||||
|
KLF4_MOUSE
|
||||||
| θ value | 2.36792 (rank : 27) | NC score | 0.005794 (rank : 62) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 694 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q60793, P70421 | Gene names | Klf4, Ezf, Gklf, Zie | |||
|
Domain Architecture |

|
|||||
| Description | Krueppel-like factor 4 (Gut-enriched krueppel-like factor) (Epithelial zinc-finger protein EZF). | |||||
|
NPAS2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 28) | NC score | 0.025726 (rank : 46) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P97460 | Gene names | Npas2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 2 (Neuronal PAS2). | |||||
|
PO6F2_MOUSE
|
||||||
| θ value | 2.36792 (rank : 29) | NC score | 0.036765 (rank : 34) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 639 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8BJI4 | Gene names | Pou6f2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | POU domain, class 6, transcription factor 2. | |||||
|
PTN23_MOUSE
|
||||||
| θ value | 2.36792 (rank : 30) | NC score | 0.032820 (rank : 37) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 726 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q6PB44, Q69ZJ0, Q8R1Z5, Q923E6 | Gene names | Ptpn23, Kiaa1471 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 23 (EC 3.1.3.48). | |||||
|
RGS3_HUMAN
|
||||||
| θ value | 2.36792 (rank : 31) | NC score | 0.061348 (rank : 15) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
IF35_MOUSE
|
||||||
| θ value | 3.0926 (rank : 32) | NC score | 0.044302 (rank : 24) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9DCH4 | Gene names | Eif3s5 | |||
|
Domain Architecture |
|
|||||
| Description | Eukaryotic translation initiation factor 3 subunit 5 (eIF-3 epsilon) (eIF3 p47 subunit) (eIF3f). | |||||
|
K1024_HUMAN
|
||||||
| θ value | 3.0926 (rank : 33) | NC score | 0.035506 (rank : 35) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UPX6 | Gene names | KIAA1024 | |||
|
Domain Architecture |

|
|||||
| Description | UPF0258 protein KIAA1024. | |||||
|
MILK1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 34) | NC score | 0.037101 (rank : 32) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8N3F8, Q5TI16, Q7RTP5, Q8N3N8, Q9BVL9, Q9BY92, Q9UH43, Q9UH44, Q9UH45 | Gene names | MIRAB13, KIAA1668 | |||
|
Domain Architecture |
|
|||||
| Description | Molecule interacting with Rab13 (MIRab13) (MICAL-like protein 1). | |||||
|
SRBP1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 35) | NC score | 0.049400 (rank : 20) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P36956, Q16062 | Gene names | SREBF1, SREBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
4ET_HUMAN
|
||||||
| θ value | 4.03905 (rank : 36) | NC score | 0.036780 (rank : 33) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9NRA8, Q8NCF2, Q9H708 | Gene names | EIF4ENIF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 4E transporter (eIF4E transporter) (4E-T) (Eukaryotic translation initiation factor 4E nuclear import factor 1). | |||||
|
CQ039_HUMAN
|
||||||
| θ value | 4.03905 (rank : 37) | NC score | 0.040366 (rank : 28) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8IVV7, Q8TEB5, Q9BW50 | Gene names | C17orf39 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C17orf39. | |||||
|
GP179_HUMAN
|
||||||
| θ value | 4.03905 (rank : 38) | NC score | 0.029115 (rank : 41) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 372 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6PRD1 | Gene names | GPR179, GPR158L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 179 precursor (Probable G-protein coupled receptor 158-like 1). | |||||
|
NFAC1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 39) | NC score | 0.030982 (rank : 40) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O95644, Q12865, Q15793 | Gene names | NFATC1, NFAT2, NFATC | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor of activated T-cells, cytoplasmic 1 (NFAT transcription complex cytosolic component) (NF-ATc1) (NF-ATc). | |||||
|
SOX7_HUMAN
|
||||||
| θ value | 4.03905 (rank : 40) | NC score | 0.016499 (rank : 54) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9BT81 | Gene names | SOX7 | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor SOX-7. | |||||
|
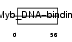
TERF2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 41) | NC score | 0.032116 (rank : 39) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q15554 | Gene names | TERF2, TRF2 | |||
|
Domain Architecture |
|
|||||
| Description | Telomeric repeat-binding factor 2 (TTAGGG repeat-binding factor 2) (Telomeric DNA-binding protein). | |||||
|
DNER_HUMAN
|
||||||
| θ value | 5.27518 (rank : 42) | NC score | 0.008269 (rank : 60) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 460 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8NFT8, Q53R88, Q53TP7, Q53TQ5, Q8IYT0, Q8TB42, Q9NTF1, Q9UDM2 | Gene names | DNER, BET | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Delta and Notch-like epidermal growth factor-related receptor precursor. | |||||
|
MAP1B_HUMAN
|
||||||
| θ value | 5.27518 (rank : 43) | NC score | 0.022415 (rank : 48) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 903 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P46821 | Gene names | MAP1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) [Contains: MAP1 light chain LC1]. | |||||
|
PCLO_HUMAN
|
||||||
| θ value | 5.27518 (rank : 44) | NC score | 0.045358 (rank : 23) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
TNR6A_HUMAN
|
||||||
| θ value | 5.27518 (rank : 45) | NC score | 0.028542 (rank : 42) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8NDV7, O15408, Q658L5, Q6NVB5, Q8NEZ0, Q8TBT8, Q8TCR0, Q9NV59, Q9P268 | Gene names | TNRC6A, CAGH26, KIAA1460, TNRC6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trinucleotide repeat-containing gene 6A protein (CAG repeat protein 26) (Glycine-tryptophan protein of 182 kDa) (GW182 autoantigen) (Protein GW1) (EMSY interactor protein). | |||||
|
ZN645_HUMAN
|
||||||
| θ value | 5.27518 (rank : 46) | NC score | 0.051732 (rank : 17) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 265 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8N7E2, Q6DJY9 | Gene names | ZNF645 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 645. | |||||
|
KCNK4_MOUSE
|
||||||
| θ value | 6.88961 (rank : 47) | NC score | 0.017230 (rank : 52) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O88454 | Gene names | Kcnk4, Traak | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 4 (TWIK-related arachidonic acid- stimulated potassium channel protein) (TRAAK). | |||||
|
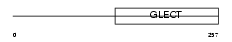
LEG3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 48) | NC score | 0.019661 (rank : 50) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P16110 | Gene names | Lgals3 | |||
|
Domain Architecture |
|
|||||
| Description | Galectin-3 (Galactose-specific lectin 3) (Mac-2 antigen) (IgE-binding protein) (35 kDa lectin) (Carbohydrate-binding protein 35) (CBP 35) (Laminin-binding protein) (Lectin L-29) (L-34 galactoside-binding lectin). | |||||
|
RC3H1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 49) | NC score | 0.021660 (rank : 49) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 433 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q5TC82, Q5W180, Q5W181, Q8IVE6, Q8N9V1 | Gene names | RC3H1, KIAA2025 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Roquin (RING finger and C3H zinc finger protein 1). | |||||
|
RNF38_HUMAN
|
||||||
| θ value | 6.88961 (rank : 50) | NC score | 0.019184 (rank : 51) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9H0F5, Q8N0Y0 | Gene names | RNF38 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 38. | |||||
|
SRBP2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 51) | NC score | 0.033999 (rank : 36) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q12772, Q9UH04 | Gene names | SREBF2, SREBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 2 (SREBP-2) (Sterol regulatory element-binding transcription factor 2). | |||||
|
BNC1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 52) | NC score | 0.013962 (rank : 56) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 460 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q01954, Q15840 | Gene names | BNC1, BNC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein basonuclin-1. | |||||
|
CBPD_MOUSE
|
||||||
| θ value | 8.99809 (rank : 53) | NC score | 0.010243 (rank : 59) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O89001 | Gene names | Cpd | |||
|
Domain Architecture |

|
|||||
| Description | Carboxypeptidase D precursor (EC 3.4.17.22) (Metallocarboxypeptidase D) (gp180). | |||||
|
GLIS3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 54) | NC score | 0.002892 (rank : 63) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 758 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8NEA6 | Gene names | GLIS3, ZNF515 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein GLIS3 (GLI-similar 3) (Zinc finger protein 515). | |||||
|
HXD10_MOUSE
|
||||||
| θ value | 8.99809 (rank : 55) | NC score | 0.006550 (rank : 61) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P28359, Q91WU9 | Gene names | Hoxd10, Hox-4.5, Hoxd-10 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-D10 (Hox-4.5) (Hox-5.3). | |||||
|
PO6F2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 56) | NC score | 0.032351 (rank : 38) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 618 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P78424, P78425, Q75ME8, Q86UM6, Q9UDS7 | Gene names | POU6F2, RPF1 | |||
|
Domain Architecture |

|
|||||
| Description | POU domain, class 6, transcription factor 2 (Retina-derived POU-domain factor 1) (RPF-1). | |||||
|
PTN23_HUMAN
|
||||||
| θ value | 8.99809 (rank : 57) | NC score | 0.027547 (rank : 44) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 646 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9H3S7, Q7KZF8, Q8N6Z5, Q9BSR5, Q9P257, Q9UG03, Q9UMZ4 | Gene names | PTPN23, KIAA1471 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 23 (EC 3.1.3.48) (His- domain-containing protein tyrosine phosphatase) (HD-PTP) (Protein tyrosine phosphatase TD14) (PTP-TD14). | |||||
|
TIF1G_MOUSE
|
||||||
| θ value | 8.99809 (rank : 58) | NC score | 0.014973 (rank : 55) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q99PP7, Q6SI71, Q6ZPX5 | Gene names | Trim33, Kiaa1113 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription intermediary factor 1-gamma (TIF1-gamma) (Tripartite motif-containing protein 33). | |||||
|
ZF106_MOUSE
|
||||||
| θ value | 8.99809 (rank : 59) | NC score | 0.016569 (rank : 53) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 472 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O88466, O55185, O88465, O88467, Q792M1, Q792P4, Q8CDZ8, Q8R3I4, Q9ESU3 | Gene names | Zfp106, H3a, Sh3bp3, Sirm, Znf474 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 106 (Zfp-106) (Zinc finger protein 474) (H3a minor histocompatibility antigen) (Son of insulin receptor mutant). | |||||
|
ZN750_MOUSE
|
||||||
| θ value | 8.99809 (rank : 60) | NC score | 0.027646 (rank : 43) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 346 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8BH05, Q66JP3, Q8C0L1 | Gene names | Znf750 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein ZNF750. | |||||
|
BCAR1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 61) | NC score | 0.062759 (rank : 11) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 554 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P56945 | Gene names | BCAR1, CAS, CRKAS | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer anti-estrogen resistance protein 1 (CRK-associated substrate) (p130cas). | |||||
|
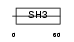
BCAR1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 62) | NC score | 0.061460 (rank : 14) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 438 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q61140, Q60869 | Gene names | Bcar1, Cas, Crkas | |||
|
Domain Architecture |
|
|||||
| Description | Breast cancer anti-estrogen resistance protein 1 (CRK-associated substrate) (p130cas). | |||||
|
PHC2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 63) | NC score | 0.050700 (rank : 18) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9QWH1, O88463, Q8K5D9 | Gene names | Phc2, Edr2, Ph2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polyhomeotic-like protein 2 (mPH2) (Early development regulatory protein 2) (p36). | |||||
|
CIKS_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | O43734, Q5R3A3, Q9H5W2, Q9H6Y3, Q9NS14, Q9UG72 | Gene names | TRAF3IP2, C6orf4, C6orf5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Adapter protein CIKS (Connection to IKK and SAPK/JNK) (TRAF3- interacting protein 2) (Nuclear factor NF-kappa-B activator 1) (ACT1). | |||||
|
ARS2_HUMAN
|
||||||
| NC score | 0.143152 (rank : 2) | θ value | 0.0563607 (rank : 7) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 186 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9BXP5, O95808, Q9BWP6, Q9BXP4 | Gene names | ARS2, ASR2 | |||
|
Domain Architecture |

|
|||||
| Description | Arsenite-resistance protein 2. | |||||
|
ARS2_MOUSE
|
||||||
| NC score | 0.141987 (rank : 3) | θ value | 0.0563607 (rank : 8) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q99MR6, Q99MR4, Q99MR5, Q99MR7 | Gene names | Ars2, Asr2 | |||
|
Domain Architecture |

|
|||||
| Description | Arsenite-resistance protein 2. | |||||
|
CASL_HUMAN
|
||||||
| NC score | 0.111155 (rank : 4) | θ value | 0.0148317 (rank : 5) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 259 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q14511 | Gene names | NEDD9, CASL | |||
|
Domain Architecture |

|
|||||
| Description | Enhancer of filamentation 1 (HEF1) (CRK-associated substrate-related protein) (CAS-L) (CasL) (p105) (Protein NEDD9) (NY-REN-12 antigen). | |||||
|
PHC2_HUMAN
|
||||||
| NC score | 0.097016 (rank : 5) | θ value | 0.00665767 (rank : 3) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 388 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8IXK0, Q5T0C1, Q6NUJ6, Q6ZQR1, Q8N306, Q8TAG8, Q96BL4, Q9Y4Y7 | Gene names | PHC2, EDR2, PH2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polyhomeotic-like protein 2 (hPH2) (Early development regulatory protein 2). | |||||
|
CASL_MOUSE
|
||||||
| NC score | 0.096943 (rank : 6) | θ value | 0.125558 (rank : 9) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O35177, Q8BJL8, Q8BK90, Q8BL52, Q8BM94, Q8BMI9, Q99KE7 | Gene names | Nedd9, Casl | |||
|
Domain Architecture |

|
|||||
| Description | Enhancer of filamentation 1 (MEF1) (CRK-associated substrate-related protein) (CAS-L) (p105) (Protein NEDD9) (Neural precursor cell expressed developmentally down-regulated protein 9). | |||||
|
PHLA1_HUMAN
|
||||||
| NC score | 0.095013 (rank : 7) | θ value | 0.47712 (rank : 14) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8WV24, Q15184, Q2TAN2, Q9NZ17 | Gene names | PHLDA1, PHRIP, TDAG51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology-like domain family A member 1 (T-cell death- associated gene 51 protein) (Apoptosis-associated nuclear protein) (Proline- and histidine-rich protein) (Proline- and glutamine-rich protein) (PQ-rich protein). | |||||
|
MLL4_HUMAN
|
||||||
| NC score | 0.089833 (rank : 8) | θ value | 0.0148317 (rank : 6) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 722 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9UMN6, O15022, O95836, Q96GP2, Q96IP3, Q9UK25, Q9Y668, Q9Y669 | Gene names | MLL4, HRX2, KIAA0304, MLL2, TRX2 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 4 (Trithorax homolog 2). | |||||
|
PHLA1_MOUSE
|
||||||
| NC score | 0.080697 (rank : 9) | θ value | 1.38821 (rank : 20) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q62392, Q3TY48, Q3UG87 | Gene names | Phlda1, Tdag51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology-like domain family A member 1 (T-cell death- associated gene 51 protein) (Proline- and glutamine-rich protein) (PQR protein). | |||||
|
RGS3_MOUSE
|
||||||
| NC score | 0.077284 (rank : 10) | θ value | 0.00298849 (rank : 2) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1621 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9DC04, Q5SRB1, Q5SRB4, Q5SRB8, Q8CEE3, Q8VI25, Q925G9, Q9JL22, Q9JL23, Q9QXA2 | Gene names | Rgs3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (C2PA). | |||||
|
BCAR1_HUMAN
|
||||||
| NC score | 0.062759 (rank : 11) | θ value | θ > 10 (rank : 61) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 554 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | P56945 | Gene names | BCAR1, CAS, CRKAS | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer anti-estrogen resistance protein 1 (CRK-associated substrate) (p130cas). | |||||
|
SELPL_MOUSE
|
||||||
| NC score | 0.062395 (rank : 12) | θ value | 0.62314 (rank : 16) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 588 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q62170 | Gene names | Selplg, Selp1, Selpl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | P-selectin glycoprotein ligand 1 precursor (PSGL-1) (Selectin P ligand). | |||||
|
AMELX_HUMAN
|
||||||
| NC score | 0.062000 (rank : 13) | θ value | 1.38821 (rank : 18) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 95 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q99217, Q96NW6, Q9UCA7 | Gene names | AMELX, AMG, AMGX | |||
|
Domain Architecture |

|
|||||
| Description | Amelogenin, X isoform precursor. | |||||
|
BCAR1_MOUSE
|
||||||
| NC score | 0.061460 (rank : 14) | θ value | θ > 10 (rank : 62) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 438 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q61140, Q60869 | Gene names | Bcar1, Cas, Crkas | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer anti-estrogen resistance protein 1 (CRK-associated substrate) (p130cas). | |||||
|
RGS3_HUMAN
|
||||||
| NC score | 0.061348 (rank : 15) | θ value | 2.36792 (rank : 31) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
MLL3_HUMAN
|
||||||
| NC score | 0.055596 (rank : 16) | θ value | 0.813845 (rank : 17) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
ZN645_HUMAN
|
||||||
| NC score | 0.051732 (rank : 17) | θ value | 5.27518 (rank : 46) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 265 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8N7E2, Q6DJY9 | Gene names | ZNF645 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 645. | |||||
|
PHC2_MOUSE
|
||||||
| NC score | 0.050700 (rank : 18) | θ value | θ > 10 (rank : 63) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 156 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9QWH1, O88463, Q8K5D9 | Gene names | Phc2, Edr2, Ph2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Polyhomeotic-like protein 2 (mPH2) (Early development regulatory protein 2) (p36). | |||||
|
PAR3L_HUMAN
|
||||||
| NC score | 0.049587 (rank : 19) | θ value | 0.279714 (rank : 10) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 297 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8TEW8, Q8IUC7, Q8IUC9, Q96DK9, Q96N09, Q96NX6, Q96NX7, Q96Q29 | Gene names | PARD3B, ALS2CR19, PAR3B, PAR3L | |||
|
Domain Architecture |

|
|||||
| Description | Partitioning-defective 3 homolog B (PAR3-beta) (Partitioning-defective 3-like protein) (PAR3-L protein) (Amyotrophic lateral sclerosis 2 chromosome region candidate gene 19 protein). | |||||
|
SRBP1_HUMAN
|
||||||
| NC score | 0.049400 (rank : 20) | θ value | 3.0926 (rank : 35) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P36956, Q16062 | Gene names | SREBF1, SREBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 1 (SREBP-1) (Sterol regulatory element-binding transcription factor 1). | |||||
|
PCLO_MOUSE
|
||||||
| NC score | 0.047345 (rank : 21) | θ value | 1.38821 (rank : 19) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 2275 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q9QYX7, Q8CF91, Q8CF92, Q9QYX6, Q9QZJ0 | Gene names | Pclo, Acz | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin) (Multidomain presynaptic cytomatrix protein) (Brain-derived HLMN protein). | |||||
|
LRBA_HUMAN
|
||||||
| NC score | 0.045888 (rank : 22) | θ value | 0.365318 (rank : 11) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P50851, Q9H2U3, Q9H2U4 | Gene names | LRBA, BGL, CDC4L, LBA | |||
|
Domain Architecture |

|
|||||
| Description | Lipopolysaccharide-responsive and beige-like anchor protein (CDC4-like protein) (Beige-like protein). | |||||
|
PCLO_HUMAN
|
||||||
| NC score | 0.045358 (rank : 23) | θ value | 5.27518 (rank : 44) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 2046 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9Y6V0, O43373, O60305, Q9BVC8, Q9UIV2, Q9Y6U9 | Gene names | PCLO, ACZ, KIAA0559 | |||
|
Domain Architecture |

|
|||||
| Description | Protein piccolo (Aczonin). | |||||
|
IF35_MOUSE
|
||||||
| NC score | 0.044302 (rank : 24) | θ value | 3.0926 (rank : 32) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 210 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9DCH4 | Gene names | Eif3s5 | |||
|
Domain Architecture |

|
|||||
| Description | Eukaryotic translation initiation factor 3 subunit 5 (eIF-3 epsilon) (eIF3 p47 subunit) (eIF3f). | |||||
|
TMC8_HUMAN
|
||||||
| NC score | 0.042591 (rank : 25) | θ value | 1.38821 (rank : 22) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8IU68, Q8IWU7, Q8N358, Q8NF04 | Gene names | TMC8, EVER2, EVIN2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane channel-like protein 8 (Epidermodysplasia verruciformis protein 2). | |||||
|
TMBI1_MOUSE
|
||||||
| NC score | 0.041951 (rank : 26) | θ value | 1.81305 (rank : 26) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BJZ3, Q8CH91, Q99KB6 | Gene names | Tmbim1, Recs1 | |||
|
Domain Architecture |

|
|||||
| Description | Transmembrane BAX inhibitor motif-containing protein 1 (RECS1 protein) (Responsive to centrifugal force and shear stress gene 1 protein). | |||||
|
SYNM_HUMAN
|
||||||
| NC score | 0.040561 (rank : 27) | θ value | 1.81305 (rank : 25) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96I59 | Gene names | NARS2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable asparaginyl-tRNA synthetase, mitochondrial precursor (EC 6.1.1.22) (Asparagine--tRNA ligase) (AsnRS). | |||||
|
CQ039_HUMAN
|
||||||
| NC score | 0.040366 (rank : 28) | θ value | 4.03905 (rank : 37) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8IVV7, Q8TEB5, Q9BW50 | Gene names | C17orf39 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C17orf39. | |||||
|
ADA12_MOUSE
|
||||||
| NC score | 0.039409 (rank : 29) | θ value | 0.0113563 (rank : 4) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 474 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q61824 | Gene names | Adam12, Mltna | |||
|
Domain Architecture |

|
|||||
| Description | ADAM 12 precursor (EC 3.4.24.-) (A disintegrin and metalloproteinase domain 12) (Meltrin alpha). | |||||
|
JHD2C_HUMAN
|
||||||
| NC score | 0.038503 (rank : 30) | θ value | 0.62314 (rank : 15) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q15652, Q5SQZ8, Q5SQZ9, Q5SR00, Q7Z3E7, Q8N3U0, Q96KB9, Q9P2G7 | Gene names | JMJD1C, JHDM2C, KIAA1380, TRIP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable JmjC domain-containing histone demethylation protein 2C (EC 1.14.11.-) (Jumonji domain-containing protein 1C) (Thyroid receptor-interacting protein 8) (TRIP-8). | |||||
|
AKAP2_HUMAN
|
||||||
| NC score | 0.037988 (rank : 31) | θ value | 1.81305 (rank : 23) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 507 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Y2D5, Q9UG26 | Gene names | AKAP2, KIAA0920 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 2 (Protein kinase A-anchoring protein 2) (PRKA2) (AKAP-2). | |||||
|
MILK1_HUMAN
|
||||||
| NC score | 0.037101 (rank : 32) | θ value | 3.0926 (rank : 34) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 427 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8N3F8, Q5TI16, Q7RTP5, Q8N3N8, Q9BVL9, Q9BY92, Q9UH43, Q9UH44, Q9UH45 | Gene names | MIRAB13, KIAA1668 | |||
|
Domain Architecture |

|
|||||
| Description | Molecule interacting with Rab13 (MIRab13) (MICAL-like protein 1). | |||||
|
4ET_HUMAN
|
||||||
| NC score | 0.036780 (rank : 33) | θ value | 4.03905 (rank : 36) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9NRA8, Q8NCF2, Q9H708 | Gene names | EIF4ENIF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Eukaryotic translation initiation factor 4E transporter (eIF4E transporter) (4E-T) (Eukaryotic translation initiation factor 4E nuclear import factor 1). | |||||
|
PO6F2_MOUSE
|
||||||
| NC score | 0.036765 (rank : 34) | θ value | 2.36792 (rank : 29) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 639 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8BJI4 | Gene names | Pou6f2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | POU domain, class 6, transcription factor 2. | |||||
|
K1024_HUMAN
|
||||||
| NC score | 0.035506 (rank : 35) | θ value | 3.0926 (rank : 33) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9UPX6 | Gene names | KIAA1024 | |||
|
Domain Architecture |

|
|||||
| Description | UPF0258 protein KIAA1024. | |||||
|
SRBP2_HUMAN
|
||||||
| NC score | 0.033999 (rank : 36) | θ value | 6.88961 (rank : 51) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q12772, Q9UH04 | Gene names | SREBF2, SREBP2 | |||
|
Domain Architecture |

|
|||||
| Description | Sterol regulatory element-binding protein 2 (SREBP-2) (Sterol regulatory element-binding transcription factor 2). | |||||
|
PTN23_MOUSE
|
||||||
| NC score | 0.032820 (rank : 37) | θ value | 2.36792 (rank : 30) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 726 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q6PB44, Q69ZJ0, Q8R1Z5, Q923E6 | Gene names | Ptpn23, Kiaa1471 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 23 (EC 3.1.3.48). | |||||
|
PO6F2_HUMAN
|
||||||
| NC score | 0.032351 (rank : 38) | θ value | 8.99809 (rank : 56) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 618 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P78424, P78425, Q75ME8, Q86UM6, Q9UDS7 | Gene names | POU6F2, RPF1 | |||
|
Domain Architecture |

|
|||||
| Description | POU domain, class 6, transcription factor 2 (Retina-derived POU-domain factor 1) (RPF-1). | |||||
|
TERF2_HUMAN
|
||||||
| NC score | 0.032116 (rank : 39) | θ value | 4.03905 (rank : 41) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q15554 | Gene names | TERF2, TRF2 | |||
|
Domain Architecture |

|
|||||
| Description | Telomeric repeat-binding factor 2 (TTAGGG repeat-binding factor 2) (Telomeric DNA-binding protein). | |||||
|
NFAC1_HUMAN
|
||||||
| NC score | 0.030982 (rank : 40) | θ value | 4.03905 (rank : 39) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O95644, Q12865, Q15793 | Gene names | NFATC1, NFAT2, NFATC | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor of activated T-cells, cytoplasmic 1 (NFAT transcription complex cytosolic component) (NF-ATc1) (NF-ATc). | |||||
|
GP179_HUMAN
|
||||||
| NC score | 0.029115 (rank : 41) | θ value | 4.03905 (rank : 38) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 372 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6PRD1 | Gene names | GPR179, GPR158L1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable G-protein coupled receptor 179 precursor (Probable G-protein coupled receptor 158-like 1). | |||||
|
TNR6A_HUMAN
|
||||||
| NC score | 0.028542 (rank : 42) | θ value | 5.27518 (rank : 45) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8NDV7, O15408, Q658L5, Q6NVB5, Q8NEZ0, Q8TBT8, Q8TCR0, Q9NV59, Q9P268 | Gene names | TNRC6A, CAGH26, KIAA1460, TNRC6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trinucleotide repeat-containing gene 6A protein (CAG repeat protein 26) (Glycine-tryptophan protein of 182 kDa) (GW182 autoantigen) (Protein GW1) (EMSY interactor protein). | |||||
|
ZN750_MOUSE
|
||||||
| NC score | 0.027646 (rank : 43) | θ value | 8.99809 (rank : 60) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 346 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8BH05, Q66JP3, Q8C0L1 | Gene names | Znf750 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein ZNF750. | |||||
|
PTN23_HUMAN
|
||||||
| NC score | 0.027547 (rank : 44) | θ value | 8.99809 (rank : 57) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 646 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9H3S7, Q7KZF8, Q8N6Z5, Q9BSR5, Q9P257, Q9UG03, Q9UMZ4 | Gene names | PTPN23, KIAA1471 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 23 (EC 3.1.3.48) (His- domain-containing protein tyrosine phosphatase) (HD-PTP) (Protein tyrosine phosphatase TD14) (PTP-TD14). | |||||
|
FOXB2_MOUSE
|
||||||
| NC score | 0.025868 (rank : 45) | θ value | 0.47712 (rank : 13) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q64733 | Gene names | Foxb2, Fkh4 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein B2 (Transcription factor FKH-4). | |||||
|
NPAS2_MOUSE
|
||||||
| NC score | 0.025726 (rank : 46) | θ value | 2.36792 (rank : 28) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P97460 | Gene names | Npas2 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 2 (Neuronal PAS2). | |||||
|
WNK2_HUMAN
|
||||||
| NC score | 0.023791 (rank : 47) | θ value | 0.365318 (rank : 12) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1407 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9Y3S1, Q8IY36, Q9C0A3, Q9H3P4 | Gene names | WNK2, KIAA1760, PRKWNK2 | |||
|
Domain Architecture |

|
|||||
| Description | Serine/threonine-protein kinase WNK2 (EC 2.7.11.1) (Protein kinase with no lysine 2) (Protein kinase, lysine-deficient 2). | |||||
|
MAP1B_HUMAN
|
||||||
| NC score | 0.022415 (rank : 48) | θ value | 5.27518 (rank : 43) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 903 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P46821 | Gene names | MAP1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) [Contains: MAP1 light chain LC1]. | |||||
|
RC3H1_HUMAN
|
||||||
| NC score | 0.021660 (rank : 49) | θ value | 6.88961 (rank : 49) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 433 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q5TC82, Q5W180, Q5W181, Q8IVE6, Q8N9V1 | Gene names | RC3H1, KIAA2025 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Roquin (RING finger and C3H zinc finger protein 1). | |||||
|
LEG3_MOUSE
|
||||||
| NC score | 0.019661 (rank : 50) | θ value | 6.88961 (rank : 48) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 113 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P16110 | Gene names | Lgals3 | |||
|
Domain Architecture |

|
|||||
| Description | Galectin-3 (Galactose-specific lectin 3) (Mac-2 antigen) (IgE-binding protein) (35 kDa lectin) (Carbohydrate-binding protein 35) (CBP 35) (Laminin-binding protein) (Lectin L-29) (L-34 galactoside-binding lectin). | |||||
|
RNF38_HUMAN
|
||||||
| NC score | 0.019184 (rank : 51) | θ value | 6.88961 (rank : 50) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 155 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9H0F5, Q8N0Y0 | Gene names | RNF38 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 38. | |||||
|
KCNK4_MOUSE
|
||||||
| NC score | 0.017230 (rank : 52) | θ value | 6.88961 (rank : 47) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O88454 | Gene names | Kcnk4, Traak | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Potassium channel subfamily K member 4 (TWIK-related arachidonic acid- stimulated potassium channel protein) (TRAAK). | |||||
|
ZF106_MOUSE
|
||||||
| NC score | 0.016569 (rank : 53) | θ value | 8.99809 (rank : 59) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 472 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O88466, O55185, O88465, O88467, Q792M1, Q792P4, Q8CDZ8, Q8R3I4, Q9ESU3 | Gene names | Zfp106, H3a, Sh3bp3, Sirm, Znf474 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 106 (Zfp-106) (Zinc finger protein 474) (H3a minor histocompatibility antigen) (Son of insulin receptor mutant). | |||||
|
SOX7_HUMAN
|
||||||
| NC score | 0.016499 (rank : 54) | θ value | 4.03905 (rank : 40) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9BT81 | Gene names | SOX7 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor SOX-7. | |||||
|
TIF1G_MOUSE
|
||||||
| NC score | 0.014973 (rank : 55) | θ value | 8.99809 (rank : 58) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q99PP7, Q6SI71, Q6ZPX5 | Gene names | Trim33, Kiaa1113 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription intermediary factor 1-gamma (TIF1-gamma) (Tripartite motif-containing protein 33). | |||||
|
BNC1_HUMAN
|
||||||
| NC score | 0.013962 (rank : 56) | θ value | 8.99809 (rank : 52) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 460 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q01954, Q15840 | Gene names | BNC1, BNC | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein basonuclin-1. | |||||
|
PLEC1_HUMAN
|
||||||
| NC score | 0.013084 (rank : 57) | θ value | 1.38821 (rank : 21) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1326 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q15149, Q15148, Q16640 | Gene names | PLEC1 | |||
|
Domain Architecture |

|
|||||
| Description | Plectin-1 (PLTN) (PCN) (Hemidesmosomal protein 1) (HD1). | |||||
|
MAST4_HUMAN
|
||||||
| NC score | 0.011069 (rank : 58) | θ value | 1.81305 (rank : 24) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 1804 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O15021, Q6ZN07, Q96LY3 | Gene names | MAST4, KIAA0303 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated serine/threonine-protein kinase 4 (EC 2.7.11.1). | |||||
|
CBPD_MOUSE
|
||||||
| NC score | 0.010243 (rank : 59) | θ value | 8.99809 (rank : 53) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O89001 | Gene names | Cpd | |||
|
Domain Architecture |

|
|||||
| Description | Carboxypeptidase D precursor (EC 3.4.17.22) (Metallocarboxypeptidase D) (gp180). | |||||
|
DNER_HUMAN
|
||||||
| NC score | 0.008269 (rank : 60) | θ value | 5.27518 (rank : 42) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 460 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8NFT8, Q53R88, Q53TP7, Q53TQ5, Q8IYT0, Q8TB42, Q9NTF1, Q9UDM2 | Gene names | DNER, BET | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Delta and Notch-like epidermal growth factor-related receptor precursor. | |||||
|
HXD10_MOUSE
|
||||||
| NC score | 0.006550 (rank : 61) | θ value | 8.99809 (rank : 55) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 392 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P28359, Q91WU9 | Gene names | Hoxd10, Hox-4.5, Hoxd-10 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein Hox-D10 (Hox-4.5) (Hox-5.3). | |||||
|
KLF4_MOUSE
|
||||||
| NC score | 0.005794 (rank : 62) | θ value | 2.36792 (rank : 27) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 694 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q60793, P70421 | Gene names | Klf4, Ezf, Gklf, Zie | |||
|
Domain Architecture |

|
|||||
| Description | Krueppel-like factor 4 (Gut-enriched krueppel-like factor) (Epithelial zinc-finger protein EZF). | |||||
|
GLIS3_HUMAN
|
||||||
| NC score | 0.002892 (rank : 63) | θ value | 8.99809 (rank : 54) | |||
| Query Neighborhood Hits | 60 | Target Neighborhood Hits | 758 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8NEA6 | Gene names | GLIS3, ZNF515 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein GLIS3 (GLI-similar 3) (Zinc finger protein 515). | |||||