Please be patient as the page loads
|
CDA7L_HUMAN
|
||||||
| SwissProt Accessions | Q96GN5, Q6PIL4, Q86YT0, Q8IXN5, Q96C70, Q9H9A2, Q9NPV2 | Gene names | CDCA7L, HR1, JPO2, R1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle-associated 7-like protein (Protein JPO2) (Transcription factor RAM2). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
CDA7L_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q96GN5, Q6PIL4, Q86YT0, Q8IXN5, Q96C70, Q9H9A2, Q9NPV2 | Gene names | CDCA7L, HR1, JPO2, R1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle-associated 7-like protein (Protein JPO2) (Transcription factor RAM2). | |||||
|
CDA7L_MOUSE
|
||||||
| θ value | 6.32649e-179 (rank : 2) | NC score | 0.979264 (rank : 2) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q922M5 | Gene names | Cdca7l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle-associated 7-like protein (Transcription factor RAM2). | |||||
|
CDCA7_HUMAN
|
||||||
| θ value | 3.70236e-70 (rank : 3) | NC score | 0.923504 (rank : 3) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9BWT1, Q53EW5, Q580W9, Q658K4, Q658N4, Q8NBY9, Q96BV8, Q96SP5 | Gene names | CDCA7, JPO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle-associated protein 7 (Protein JPO1). | |||||
|
CDCA7_MOUSE
|
||||||
| θ value | 5.17823e-64 (rank : 4) | NC score | 0.921354 (rank : 4) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9D0M2, Q3TIS1, Q3U511, Q6NZE5, Q8C1A0 | Gene names | Cdca7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle-associated protein 7. | |||||
|
RFC1_HUMAN
|
||||||
| θ value | 0.00869519 (rank : 5) | NC score | 0.073713 (rank : 5) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P35251, Q5XKF5, Q6PKU0, Q86V41, Q86V46 | Gene names | RFC1, RFC140 | |||
|
Domain Architecture |

|
|||||
| Description | Replication factor C subunit 1 (Replication factor C large subunit) (RF-C 140 kDa subunit) (Activator 1 140 kDa subunit) (Activator 1 large subunit) (A1 140 kDa subunit) (DNA-binding protein PO-GA). | |||||
|
RN168_MOUSE
|
||||||
| θ value | 0.163984 (rank : 6) | NC score | 0.043045 (rank : 9) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 440 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q80XJ2 | Gene names | Rnf168 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 168. | |||||
|
NEST_HUMAN
|
||||||
| θ value | 0.21417 (rank : 7) | NC score | 0.024614 (rank : 15) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 1305 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P48681, O00552 | Gene names | NES | |||
|
Domain Architecture |

|
|||||
| Description | Nestin. | |||||
|
ATRN_HUMAN
|
||||||
| θ value | 0.279714 (rank : 8) | NC score | 0.046333 (rank : 7) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 525 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O75882, O60295, O95414, Q3MIT3, Q5TDA2, Q5TDA4, Q5VYW3, Q9NTQ3, Q9NTQ4, Q9NU01, Q9NZ57, Q9NZ58 | Gene names | ATRN, KIAA0548, MGCA | |||
|
Domain Architecture |

|
|||||
| Description | Attractin precursor (Mahogany homolog) (DPPT-L). | |||||
|
IWS1_MOUSE
|
||||||
| θ value | 0.365318 (rank : 9) | NC score | 0.045589 (rank : 8) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
STAB1_MOUSE
|
||||||
| θ value | 0.62314 (rank : 10) | NC score | 0.023798 (rank : 16) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 566 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8R4Y4, Q8K0K6, Q8VC09 | Gene names | Stab1, Feel1 | |||
|
Domain Architecture |

|
|||||
| Description | Stabilin-1 precursor (FEEL-1 protein). | |||||
|
CCD39_HUMAN
|
||||||
| θ value | 0.813845 (rank : 11) | NC score | 0.015061 (rank : 25) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 868 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UFE4 | Gene names | CCDC39 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 39. | |||||
|
NUF2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 12) | NC score | 0.020486 (rank : 20) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 661 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9BZD4, Q8WU69, Q96HJ4, Q96Q78 | Gene names | CDCA1, NUF2, NUF2R | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinetochore protein Nuf2 (hNuf2) (hNuf2R) (hsNuf2) (Cell division cycle-associated protein 1). | |||||
|
ATRN_MOUSE
|
||||||
| θ value | 1.06291 (rank : 13) | NC score | 0.040409 (rank : 10) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 519 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9WU60, Q9R263, Q9WU77 | Gene names | Atrn, mg, Mgca | |||
|
Domain Architecture |

|
|||||
| Description | Attractin precursor (Mahogany protein). | |||||
|
CD016_HUMAN
|
||||||
| θ value | 1.38821 (rank : 14) | NC score | 0.056258 (rank : 6) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q63HQ0, Q96GG6, Q9H0V0, Q9P1L4 | Gene names | C4orf16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C4orf16. | |||||
|
ESCO2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 15) | NC score | 0.027031 (rank : 12) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q56NI9 | Gene names | ESCO2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyltransferase ESCO2 (EC 2.3.1.-) (Establishment of cohesion 1 homolog 2) (ECO1 homolog 2). | |||||
|
IWS1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 16) | NC score | 0.036792 (rank : 11) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
MAON_MOUSE
|
||||||
| θ value | 2.36792 (rank : 17) | NC score | 0.026290 (rank : 13) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BMF3 | Gene names | Me3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NADP-dependent malic enzyme, mitochondrial precursor (EC 1.1.1.40) (NADP-ME) (Malic enzyme 3). | |||||
|
NPBL_MOUSE
|
||||||
| θ value | 2.36792 (rank : 18) | NC score | 0.020906 (rank : 19) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
MAON_HUMAN
|
||||||
| θ value | 3.0926 (rank : 19) | NC score | 0.025397 (rank : 14) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q16798 | Gene names | ME3 | |||
|
Domain Architecture |

|
|||||
| Description | NADP-dependent malic enzyme, mitochondrial precursor (EC 1.1.1.40) (NADP-ME) (Malic enzyme 3). | |||||
|
SESN2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 20) | NC score | 0.016958 (rank : 22) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P58004, Q96SI5 | Gene names | SESN2, SEST2 | |||
|
Domain Architecture |
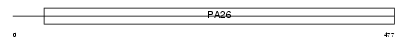
|
|||||
| Description | Sestrin-2 (Hi95). | |||||
|
ANXA1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 21) | NC score | 0.009228 (rank : 31) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P04083 | Gene names | ANXA1, ANX1, LPC1 | |||
|
Domain Architecture |
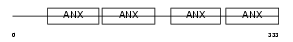
|
|||||
| Description | Annexin A1 (Annexin I) (Lipocortin I) (Calpactin II) (Chromobindin-9) (p35) (Phospholipase A2 inhibitory protein). | |||||
|
CCNB3_MOUSE
|
||||||
| θ value | 4.03905 (rank : 22) | NC score | 0.017391 (rank : 21) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q810T2, Q810T3, Q8VDC8 | Gene names | Ccnb3, Cycb3 | |||
|
Domain Architecture |

|
|||||
| Description | G2/mitotic-specific cyclin-B3. | |||||
|
MILK1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 23) | NC score | 0.011462 (rank : 27) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 788 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8BGT6, Q8BJ60 | Gene names | Mirab13, D15Mit260, Kiaa1668 | |||
|
Domain Architecture |

|
|||||
| Description | Molecule interacting with Rab13 (MIRab13) (MICAL-like protein 1). | |||||
|
MYH9_MOUSE
|
||||||
| θ value | 4.03905 (rank : 24) | NC score | 0.006093 (rank : 33) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 1829 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8VDD5 | Gene names | Myh9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
STAB1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 25) | NC score | 0.021517 (rank : 18) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 546 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NY15, Q8IUH0, Q8IUH1, Q93072 | Gene names | STAB1, FEEL1, KIAA0246 | |||
|
Domain Architecture |

|
|||||
| Description | Stabilin-1 precursor (FEEL-1 protein) (MS-1 antigen). | |||||
|
LAMA1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 26) | NC score | 0.012603 (rank : 26) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 804 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P25391 | Gene names | LAMA1, LAMA | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-1 chain precursor (Laminin A chain). | |||||
|
CAC1C_HUMAN
|
||||||
| θ value | 6.88961 (rank : 27) | NC score | 0.006077 (rank : 34) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13936, Q13917, Q13918, Q13919, Q13920, Q13921, Q13922, Q13923, Q13924, Q13925, Q13926, Q13927, Q13928, Q13929, Q13930, Q13932, Q13933, Q14743, Q14744, Q15877, Q99025, Q99241, Q99875 | Gene names | CACNA1C, CACH2, CACN2, CACNL1A1, CCHL1A1 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1C (Voltage- gated calcium channel subunit alpha Cav1.2) (Calcium channel, L type, alpha-1 polypeptide, isoform 1, cardiac muscle). | |||||
|
MSPD2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 28) | NC score | 0.015808 (rank : 24) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8NHP6, Q8N3H2, Q8NA83 | Gene names | MOSPD2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Motile sperm domain-containing protein 2. | |||||
|
AT10A_HUMAN
|
||||||
| θ value | 8.99809 (rank : 29) | NC score | 0.008941 (rank : 32) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O60312, Q969I4 | Gene names | ATP10A, ATP10C, ATPVC, KIAA0566 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase VA (EC 3.6.3.1) (ATPVA) (Aminophospholipid translocase VA). | |||||
|
BRCA1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 30) | NC score | 0.016017 (rank : 23) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 436 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P48754, Q60957, Q60983 | Gene names | Brca1 | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer type 1 susceptibility protein homolog. | |||||
|
HTSF1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 31) | NC score | 0.021588 (rank : 17) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8BGC0, Q1WWK0, Q9CT41, Q9DAU3 | Gene names | Htatsf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HIV Tat-specific factor 1 homolog. | |||||
|
LAMA5_MOUSE
|
||||||
| θ value | 8.99809 (rank : 32) | NC score | 0.010513 (rank : 29) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q61001, Q9JHQ6 | Gene names | Lama5 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-5 chain precursor. | |||||
|
MY18A_MOUSE
|
||||||
| θ value | 8.99809 (rank : 33) | NC score | 0.005835 (rank : 35) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 1989 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9JMH9, Q811D7 | Gene names | Myo18a, Myspdz | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18A (Myosin XVIIIa) (Myosin containing PDZ domain) (Molecule associated with JAK3 N-terminus) (MAJN). | |||||
|
POGZ_MOUSE
|
||||||
| θ value | 8.99809 (rank : 34) | NC score | 0.001787 (rank : 36) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 765 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BZH4, Q4VA94, Q80TZ8, Q8C0K1, Q8K294 | Gene names | Pogz, Kiaa0461 | |||
|
Domain Architecture |

|
|||||
| Description | Pogo transposable element with ZNF domain. | |||||
|
SOAT1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 35) | NC score | 0.010045 (rank : 30) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P35610, Q8N1E4 | Gene names | SOAT1, ACACT, ACACT1, ACAT, ACAT1, SOAT, STAT | |||
|
Domain Architecture |

|
|||||
| Description | Sterol O-acyltransferase 1 (EC 2.3.1.26) (Cholesterol acyltransferase 1) (Acyl coenzyme A:cholesterol acyltransferase 1) (ACAT-1). | |||||
|
TNR16_MOUSE
|
||||||
| θ value | 8.99809 (rank : 36) | NC score | 0.010751 (rank : 28) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Z0W1 | Gene names | Ngfr, Tnfrsf16 | |||
|
Domain Architecture |
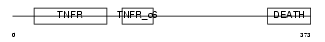
|
|||||
| Description | Tumor necrosis factor receptor superfamily member 16 precursor (Low- affinity nerve growth factor receptor) (NGF receptor) (Low affinity neurotrophin receptor p75NTR). | |||||
|
CDA7L_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 36 | |
| SwissProt Accessions | Q96GN5, Q6PIL4, Q86YT0, Q8IXN5, Q96C70, Q9H9A2, Q9NPV2 | Gene names | CDCA7L, HR1, JPO2, R1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle-associated 7-like protein (Protein JPO2) (Transcription factor RAM2). | |||||
|
CDA7L_MOUSE
|
||||||
| NC score | 0.979264 (rank : 2) | θ value | 6.32649e-179 (rank : 2) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q922M5 | Gene names | Cdca7l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle-associated 7-like protein (Transcription factor RAM2). | |||||
|
CDCA7_HUMAN
|
||||||
| NC score | 0.923504 (rank : 3) | θ value | 3.70236e-70 (rank : 3) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9BWT1, Q53EW5, Q580W9, Q658K4, Q658N4, Q8NBY9, Q96BV8, Q96SP5 | Gene names | CDCA7, JPO1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle-associated protein 7 (Protein JPO1). | |||||
|
CDCA7_MOUSE
|
||||||
| NC score | 0.921354 (rank : 4) | θ value | 5.17823e-64 (rank : 4) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9D0M2, Q3TIS1, Q3U511, Q6NZE5, Q8C1A0 | Gene names | Cdca7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cell division cycle-associated protein 7. | |||||
|
RFC1_HUMAN
|
||||||
| NC score | 0.073713 (rank : 5) | θ value | 0.00869519 (rank : 5) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 352 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P35251, Q5XKF5, Q6PKU0, Q86V41, Q86V46 | Gene names | RFC1, RFC140 | |||
|
Domain Architecture |

|
|||||
| Description | Replication factor C subunit 1 (Replication factor C large subunit) (RF-C 140 kDa subunit) (Activator 1 140 kDa subunit) (Activator 1 large subunit) (A1 140 kDa subunit) (DNA-binding protein PO-GA). | |||||
|
CD016_HUMAN
|
||||||
| NC score | 0.056258 (rank : 6) | θ value | 1.38821 (rank : 14) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q63HQ0, Q96GG6, Q9H0V0, Q9P1L4 | Gene names | C4orf16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C4orf16. | |||||
|
ATRN_HUMAN
|
||||||
| NC score | 0.046333 (rank : 7) | θ value | 0.279714 (rank : 8) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 525 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O75882, O60295, O95414, Q3MIT3, Q5TDA2, Q5TDA4, Q5VYW3, Q9NTQ3, Q9NTQ4, Q9NU01, Q9NZ57, Q9NZ58 | Gene names | ATRN, KIAA0548, MGCA | |||
|
Domain Architecture |

|
|||||
| Description | Attractin precursor (Mahogany homolog) (DPPT-L). | |||||
|
IWS1_MOUSE
|
||||||
| NC score | 0.045589 (rank : 8) | θ value | 0.365318 (rank : 9) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
RN168_MOUSE
|
||||||
| NC score | 0.043045 (rank : 9) | θ value | 0.163984 (rank : 6) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 440 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q80XJ2 | Gene names | Rnf168 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING finger protein 168. | |||||
|
ATRN_MOUSE
|
||||||
| NC score | 0.040409 (rank : 10) | θ value | 1.06291 (rank : 13) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 519 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9WU60, Q9R263, Q9WU77 | Gene names | Atrn, mg, Mgca | |||
|
Domain Architecture |

|
|||||
| Description | Attractin precursor (Mahogany protein). | |||||
|
IWS1_HUMAN
|
||||||
| NC score | 0.036792 (rank : 11) | θ value | 1.81305 (rank : 16) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
ESCO2_HUMAN
|
||||||
| NC score | 0.027031 (rank : 12) | θ value | 1.81305 (rank : 15) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q56NI9 | Gene names | ESCO2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyltransferase ESCO2 (EC 2.3.1.-) (Establishment of cohesion 1 homolog 2) (ECO1 homolog 2). | |||||
|
MAON_MOUSE
|
||||||
| NC score | 0.026290 (rank : 13) | θ value | 2.36792 (rank : 17) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8BMF3 | Gene names | Me3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NADP-dependent malic enzyme, mitochondrial precursor (EC 1.1.1.40) (NADP-ME) (Malic enzyme 3). | |||||
|
MAON_HUMAN
|
||||||
| NC score | 0.025397 (rank : 14) | θ value | 3.0926 (rank : 19) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q16798 | Gene names | ME3 | |||
|
Domain Architecture |

|
|||||
| Description | NADP-dependent malic enzyme, mitochondrial precursor (EC 1.1.1.40) (NADP-ME) (Malic enzyme 3). | |||||
|
NEST_HUMAN
|
||||||
| NC score | 0.024614 (rank : 15) | θ value | 0.21417 (rank : 7) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 1305 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P48681, O00552 | Gene names | NES | |||
|
Domain Architecture |

|
|||||
| Description | Nestin. | |||||
|
STAB1_MOUSE
|
||||||
| NC score | 0.023798 (rank : 16) | θ value | 0.62314 (rank : 10) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 566 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8R4Y4, Q8K0K6, Q8VC09 | Gene names | Stab1, Feel1 | |||
|
Domain Architecture |

|
|||||
| Description | Stabilin-1 precursor (FEEL-1 protein). | |||||
|
HTSF1_MOUSE
|
||||||
| NC score | 0.021588 (rank : 17) | θ value | 8.99809 (rank : 31) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 425 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q8BGC0, Q1WWK0, Q9CT41, Q9DAU3 | Gene names | Htatsf1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HIV Tat-specific factor 1 homolog. | |||||
|
STAB1_HUMAN
|
||||||
| NC score | 0.021517 (rank : 18) | θ value | 4.03905 (rank : 25) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 546 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NY15, Q8IUH0, Q8IUH1, Q93072 | Gene names | STAB1, FEEL1, KIAA0246 | |||
|
Domain Architecture |

|
|||||
| Description | Stabilin-1 precursor (FEEL-1 protein) (MS-1 antigen). | |||||
|
NPBL_MOUSE
|
||||||
| NC score | 0.020906 (rank : 19) | θ value | 2.36792 (rank : 18) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 1041 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6KCD5, Q6KC78, Q7TNS4, Q8BKV4, Q8CES9, Q9CUC6 | Gene names | Nipbl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin homolog) (SCC2 homolog). | |||||
|
NUF2_HUMAN
|
||||||
| NC score | 0.020486 (rank : 20) | θ value | 0.813845 (rank : 12) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 661 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9BZD4, Q8WU69, Q96HJ4, Q96Q78 | Gene names | CDCA1, NUF2, NUF2R | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kinetochore protein Nuf2 (hNuf2) (hNuf2R) (hsNuf2) (Cell division cycle-associated protein 1). | |||||
|
CCNB3_MOUSE
|
||||||
| NC score | 0.017391 (rank : 21) | θ value | 4.03905 (rank : 22) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q810T2, Q810T3, Q8VDC8 | Gene names | Ccnb3, Cycb3 | |||
|
Domain Architecture |

|
|||||
| Description | G2/mitotic-specific cyclin-B3. | |||||
|
SESN2_HUMAN
|
||||||
| NC score | 0.016958 (rank : 22) | θ value | 3.0926 (rank : 20) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P58004, Q96SI5 | Gene names | SESN2, SEST2 | |||
|
Domain Architecture |

|
|||||
| Description | Sestrin-2 (Hi95). | |||||
|
BRCA1_MOUSE
|
||||||
| NC score | 0.016017 (rank : 23) | θ value | 8.99809 (rank : 30) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 436 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P48754, Q60957, Q60983 | Gene names | Brca1 | |||
|
Domain Architecture |

|
|||||
| Description | Breast cancer type 1 susceptibility protein homolog. | |||||
|
MSPD2_HUMAN
|
||||||
| NC score | 0.015808 (rank : 24) | θ value | 6.88961 (rank : 28) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 52 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8NHP6, Q8N3H2, Q8NA83 | Gene names | MOSPD2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Motile sperm domain-containing protein 2. | |||||
|
CCD39_HUMAN
|
||||||
| NC score | 0.015061 (rank : 25) | θ value | 0.813845 (rank : 11) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 868 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9UFE4 | Gene names | CCDC39 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 39. | |||||
|
LAMA1_HUMAN
|
||||||
| NC score | 0.012603 (rank : 26) | θ value | 5.27518 (rank : 26) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 804 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | P25391 | Gene names | LAMA1, LAMA | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-1 chain precursor (Laminin A chain). | |||||
|
MILK1_MOUSE
|
||||||
| NC score | 0.011462 (rank : 27) | θ value | 4.03905 (rank : 23) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 788 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8BGT6, Q8BJ60 | Gene names | Mirab13, D15Mit260, Kiaa1668 | |||
|
Domain Architecture |

|
|||||
| Description | Molecule interacting with Rab13 (MIRab13) (MICAL-like protein 1). | |||||
|
TNR16_MOUSE
|
||||||
| NC score | 0.010751 (rank : 28) | θ value | 8.99809 (rank : 36) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 178 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Z0W1 | Gene names | Ngfr, Tnfrsf16 | |||
|
Domain Architecture |

|
|||||
| Description | Tumor necrosis factor receptor superfamily member 16 precursor (Low- affinity nerve growth factor receptor) (NGF receptor) (Low affinity neurotrophin receptor p75NTR). | |||||
|
LAMA5_MOUSE
|
||||||
| NC score | 0.010513 (rank : 29) | θ value | 8.99809 (rank : 32) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q61001, Q9JHQ6 | Gene names | Lama5 | |||
|
Domain Architecture |

|
|||||
| Description | Laminin alpha-5 chain precursor. | |||||
|
SOAT1_HUMAN
|
||||||
| NC score | 0.010045 (rank : 30) | θ value | 8.99809 (rank : 35) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P35610, Q8N1E4 | Gene names | SOAT1, ACACT, ACACT1, ACAT, ACAT1, SOAT, STAT | |||
|
Domain Architecture |

|
|||||
| Description | Sterol O-acyltransferase 1 (EC 2.3.1.26) (Cholesterol acyltransferase 1) (Acyl coenzyme A:cholesterol acyltransferase 1) (ACAT-1). | |||||
|
ANXA1_HUMAN
|
||||||
| NC score | 0.009228 (rank : 31) | θ value | 4.03905 (rank : 21) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P04083 | Gene names | ANXA1, ANX1, LPC1 | |||
|
Domain Architecture |

|
|||||
| Description | Annexin A1 (Annexin I) (Lipocortin I) (Calpactin II) (Chromobindin-9) (p35) (Phospholipase A2 inhibitory protein). | |||||
|
AT10A_HUMAN
|
||||||
| NC score | 0.008941 (rank : 32) | θ value | 8.99809 (rank : 29) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 98 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O60312, Q969I4 | Gene names | ATP10A, ATP10C, ATPVC, KIAA0566 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable phospholipid-transporting ATPase VA (EC 3.6.3.1) (ATPVA) (Aminophospholipid translocase VA). | |||||
|
MYH9_MOUSE
|
||||||
| NC score | 0.006093 (rank : 33) | θ value | 4.03905 (rank : 24) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 1829 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8VDD5 | Gene names | Myh9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myosin-9 (Myosin heavy chain, nonmuscle IIa) (Nonmuscle myosin heavy chain IIa) (NMMHC II-a) (NMMHC-IIA) (Cellular myosin heavy chain, type A) (Nonmuscle myosin heavy chain-A) (NMMHC-A). | |||||
|
CAC1C_HUMAN
|
||||||
| NC score | 0.006077 (rank : 34) | θ value | 6.88961 (rank : 27) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q13936, Q13917, Q13918, Q13919, Q13920, Q13921, Q13922, Q13923, Q13924, Q13925, Q13926, Q13927, Q13928, Q13929, Q13930, Q13932, Q13933, Q14743, Q14744, Q15877, Q99025, Q99241, Q99875 | Gene names | CACNA1C, CACH2, CACN2, CACNL1A1, CCHL1A1 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1C (Voltage- gated calcium channel subunit alpha Cav1.2) (Calcium channel, L type, alpha-1 polypeptide, isoform 1, cardiac muscle). | |||||
|
MY18A_MOUSE
|
||||||
| NC score | 0.005835 (rank : 35) | θ value | 8.99809 (rank : 33) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 1989 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q9JMH9, Q811D7 | Gene names | Myo18a, Myspdz | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-18A (Myosin XVIIIa) (Myosin containing PDZ domain) (Molecule associated with JAK3 N-terminus) (MAJN). | |||||
|
POGZ_MOUSE
|
||||||
| NC score | 0.001787 (rank : 36) | θ value | 8.99809 (rank : 34) | |||
| Query Neighborhood Hits | 36 | Target Neighborhood Hits | 765 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BZH4, Q4VA94, Q80TZ8, Q8C0K1, Q8K294 | Gene names | Pogz, Kiaa0461 | |||
|
Domain Architecture |

|
|||||
| Description | Pogo transposable element with ZNF domain. | |||||