Please be patient as the page loads
|
CCNF_HUMAN
|
||||||
| SwissProt Accessions | P41002, Q96EG9 | Gene names | CCNF | |||
|
Domain Architecture |

|
|||||
| Description | G2/mitotic-specific cyclin-F. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
CCNF_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P41002, Q96EG9 | Gene names | CCNF | |||
|
Domain Architecture |

|
|||||
| Description | G2/mitotic-specific cyclin-F. | |||||
|
CCNF_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.992573 (rank : 2) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P51944, Q60797, Q60799 | Gene names | Ccnf | |||
|
Domain Architecture |

|
|||||
| Description | G2/mitotic-specific cyclin-F. | |||||
|
CCNA2_MOUSE
|
||||||
| θ value | 3.07116e-24 (rank : 3) | NC score | 0.733728 (rank : 3) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P51943, Q61459 | Gene names | Ccna2, Ccna, Cyca | |||
|
Domain Architecture |
|
|||||
| Description | Cyclin-A2 (Cyclin-A). | |||||
|
CCNA2_HUMAN
|
||||||
| θ value | 3.39556e-23 (rank : 4) | NC score | 0.729441 (rank : 5) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P20248 | Gene names | CCNA2, CCN1, CCNA | |||
|
Domain Architecture |
|
|||||
| Description | Cyclin-A2 (Cyclin-A). | |||||
|
CCNA1_MOUSE
|
||||||
| θ value | 5.79196e-23 (rank : 5) | NC score | 0.730380 (rank : 4) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q61456 | Gene names | Ccna1 | |||
|
Domain Architecture |
|
|||||
| Description | Cyclin-A1. | |||||
|
CCNA1_HUMAN
|
||||||
| θ value | 4.58923e-20 (rank : 6) | NC score | 0.721688 (rank : 6) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P78396 | Gene names | CCNA1 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin-A1. | |||||
|
CCNB1_HUMAN
|
||||||
| θ value | 1.24977e-17 (rank : 7) | NC score | 0.705529 (rank : 7) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P14635 | Gene names | CCNB1, CCNB | |||
|
Domain Architecture |
|
|||||
| Description | G2/mitotic-specific cyclin-B1. | |||||
|
CCNB1_MOUSE
|
||||||
| θ value | 2.7842e-17 (rank : 8) | NC score | 0.705307 (rank : 8) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P24860 | Gene names | Ccnb1, Ccn-2, Ccnb1-rs13, Cycb, Cycb1 | |||
|
Domain Architecture |
|
|||||
| Description | G2/mitotic-specific cyclin-B1. | |||||
|
CCNB3_MOUSE
|
||||||
| θ value | 1.058e-16 (rank : 9) | NC score | 0.658690 (rank : 11) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q810T2, Q810T3, Q8VDC8 | Gene names | Ccnb3, Cycb3 | |||
|
Domain Architecture |

|
|||||
| Description | G2/mitotic-specific cyclin-B3. | |||||
|
CCNB2_MOUSE
|
||||||
| θ value | 5.25075e-16 (rank : 10) | NC score | 0.695675 (rank : 9) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P30276 | Gene names | Ccnb2, Cycb2 | |||
|
Domain Architecture |
|
|||||
| Description | G2/mitotic-specific cyclin-B2. | |||||
|
CCNB2_HUMAN
|
||||||
| θ value | 1.52774e-15 (rank : 11) | NC score | 0.692818 (rank : 10) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O95067 | Gene names | CCNB2 | |||
|
Domain Architecture |
|
|||||
| Description | G2/mitotic-specific cyclin-B2. | |||||
|
CCNB3_HUMAN
|
||||||
| θ value | 7.09661e-13 (rank : 12) | NC score | 0.562769 (rank : 12) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8WWL7, Q96SB5, Q96SB6, Q96SB7, Q9NT38 | Gene names | CCNB3, CYCB3 | |||
|
Domain Architecture |

|
|||||
| Description | G2/mitotic-specific cyclin-B3. | |||||
|
CCND2_HUMAN
|
||||||
| θ value | 4.0297e-08 (rank : 13) | NC score | 0.512096 (rank : 19) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P30279, Q13955 | Gene names | CCND2 | |||
|
Domain Architecture |
|
|||||
| Description | G1/S-specific cyclin-D2. | |||||
|
CCND1_HUMAN
|
||||||
| θ value | 1.99992e-07 (rank : 14) | NC score | 0.533628 (rank : 16) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P24385, Q6LEF0 | Gene names | CCND1, BCL1, PRAD1 | |||
|
Domain Architecture |
|
|||||
| Description | G1/S-specific cyclin-D1 (PRAD1 oncogene) (BCL-1 oncogene). | |||||
|
CCND1_MOUSE
|
||||||
| θ value | 3.41135e-07 (rank : 15) | NC score | 0.535376 (rank : 15) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P25322 | Gene names | Ccnd1, Cyl-1 | |||
|
Domain Architecture |
|
|||||
| Description | G1/S-specific cyclin-D1. | |||||
|
CCNE2_HUMAN
|
||||||
| θ value | 1.69304e-06 (rank : 16) | NC score | 0.541131 (rank : 14) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O96020, O95439 | Gene names | CCNE2 | |||
|
Domain Architecture |
|
|||||
| Description | G1/S-specific cyclin-E2. | |||||
|
CCND2_MOUSE
|
||||||
| θ value | 2.21117e-06 (rank : 17) | NC score | 0.497316 (rank : 20) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P30280 | Gene names | Ccnd2, Cyl-2 | |||
|
Domain Architecture |
|
|||||
| Description | G1/S-specific cyclin-D2. | |||||
|
CCNE2_MOUSE
|
||||||
| θ value | 2.21117e-06 (rank : 18) | NC score | 0.543966 (rank : 13) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9Z238, Q9QWU5 | Gene names | Ccne2 | |||
|
Domain Architecture |
|
|||||
| Description | G1/S-specific cyclin-E2. | |||||
|
CCNE1_HUMAN
|
||||||
| θ value | 0.000121331 (rank : 19) | NC score | 0.528171 (rank : 17) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P24864, Q14091, Q8NFG1, Q92501 | Gene names | CCNE1, CCNE | |||
|
Domain Architecture |
|
|||||
| Description | G1/S-specific cyclin-E1. | |||||
|
CCNE1_MOUSE
|
||||||
| θ value | 0.000158464 (rank : 20) | NC score | 0.520961 (rank : 18) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q61457 | Gene names | Ccne1, Ccne | |||
|
Domain Architecture |
|
|||||
| Description | G1/S-specific cyclin-E1. | |||||
|
U2AF2_HUMAN
|
||||||
| θ value | 0.0113563 (rank : 21) | NC score | 0.072084 (rank : 31) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 192 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P26368 | Gene names | U2AF2, U2AF65 | |||
|
Domain Architecture |
|
|||||
| Description | Splicing factor U2AF 65 kDa subunit (U2 auxiliary factor 65 kDa subunit) (U2 snRNP auxiliary factor large subunit) (hU2AF(65)). | |||||
|
CCND3_HUMAN
|
||||||
| θ value | 0.0193708 (rank : 22) | NC score | 0.476354 (rank : 21) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P30281, Q5T8J0, Q96F49 | Gene names | CCND3 | |||
|
Domain Architecture |
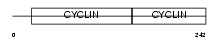
|
|||||
| Description | G1/S-specific cyclin-D3. | |||||
|
GIDRP_HUMAN
|
||||||
| θ value | 0.125558 (rank : 23) | NC score | 0.067853 (rank : 32) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7Z5L2, O60598, Q5W0B4, Q5W0B5, Q86VT9, Q8N9H0 | Gene names | GIDRP88, C10orf28 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Growth inhibition and differentiation-related protein 88 (Putative mitochondrial space protein 32.1). | |||||
|
U2AF2_MOUSE
|
||||||
| θ value | 0.163984 (rank : 24) | NC score | 0.052719 (rank : 34) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 192 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P26369 | Gene names | U2af2, U2af65 | |||
|
Domain Architecture |
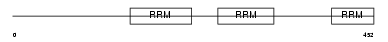
|
|||||
| Description | Splicing factor U2AF 65 kDa subunit (U2 auxiliary factor 65 kDa subunit) (U2 snRNP auxiliary factor large subunit). | |||||
|
EGR4_HUMAN
|
||||||
| θ value | 0.47712 (rank : 25) | NC score | 0.003532 (rank : 52) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 939 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q05215 | Gene names | EGR4 | |||
|
Domain Architecture |

|
|||||
| Description | Early growth response protein 4 (EGR-4) (AT133). | |||||
|
GP152_HUMAN
|
||||||
| θ value | 0.47712 (rank : 26) | NC score | 0.015838 (rank : 43) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 636 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8TDT2, Q86SM0 | Gene names | GPR152, PGR5 | |||
|
Domain Architecture |

|
|||||
| Description | Probable G-protein coupled receptor 152 (G-protein coupled receptor PGR5). | |||||
|
CCNG1_MOUSE
|
||||||
| θ value | 0.813845 (rank : 27) | NC score | 0.285339 (rank : 23) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P51945, O54779 | Gene names | Ccng1, Ccng | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin-G1 (Cyclin-G). | |||||
|
FMN1B_MOUSE
|
||||||
| θ value | 1.38821 (rank : 28) | NC score | 0.029069 (rank : 37) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q05859 | Gene names | Fmn, Ld | |||
|
Domain Architecture |

|
|||||
| Description | Formin-1 isoform IV (Limb deformity protein). | |||||
|
CDSN_MOUSE
|
||||||
| θ value | 1.81305 (rank : 29) | NC score | 0.029627 (rank : 36) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7TPC1 | Gene names | Cdsn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Corneodesmosin precursor. | |||||
|
MAGE1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 30) | NC score | 0.007522 (rank : 50) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1389 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9HCI5, Q86TG0, Q8TD92, Q9H216 | Gene names | MAGEE1, HCA1, KIAA1587 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma-associated antigen E1 (MAGE-E1 antigen) (Hepatocellular carcinoma-associated protein 1). | |||||
|
VHL_HUMAN
|
||||||
| θ value | 2.36792 (rank : 31) | NC score | 0.032366 (rank : 35) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P40337, Q13599 | Gene names | VHL | |||
|
Domain Architecture |
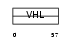
|
|||||
| Description | Von Hippel-Lindau disease tumor suppressor (pVHL) (G7 protein). | |||||
|
CD2L1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 32) | NC score | 0.000058 (rank : 53) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1463 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P24788, Q3UI03, Q61399, Q7TST4, Q8BP53 | Gene names | Cdc2l1 | |||
|
Domain Architecture |

|
|||||
| Description | PITSLRE serine/threonine-protein kinase CDC2L1 (EC 2.7.11.22) (Galactosyltransferase-associated protein kinase p58/GTA) (Cell division cycle 2-like protein kinase 1). | |||||
|
NACAM_MOUSE
|
||||||
| θ value | 3.0926 (rank : 33) | NC score | 0.008533 (rank : 49) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
SYNJ1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 34) | NC score | 0.013601 (rank : 44) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 516 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8CHC4 | Gene names | Synj1, Kiaa0910 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptojanin-1 (EC 3.1.3.36) (Synaptic inositol-1,4,5-trisphosphate 5- phosphatase 1). | |||||
|
CCDC9_MOUSE
|
||||||
| θ value | 4.03905 (rank : 35) | NC score | 0.020573 (rank : 42) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8VC31 | Gene names | Ccdc9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 9. | |||||
|
CCNG1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 36) | NC score | 0.272717 (rank : 24) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P51959, Q15757, Q96L32 | Gene names | CCNG1, CCNG, CYCG1 | |||
|
Domain Architecture |
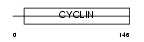
|
|||||
| Description | Cyclin-G1 (Cyclin-G). | |||||
|
BAT2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 37) | NC score | 0.011713 (rank : 46) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P48634, O95875, Q5SQ29, Q5SQ30, Q5ST84, Q5STX6, Q5STX7, Q68DW9, Q6P9P7, Q6PIN1, Q96QC6 | Gene names | BAT2, G2 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
CENPI_HUMAN
|
||||||
| θ value | 6.88961 (rank : 38) | NC score | 0.021253 (rank : 41) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q92674, Q5JWZ9, Q96ED0 | Gene names | CENPI, FSHPRH1, ICEN19, LRPR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein I (CENP-I) (Interphase centromere complex protein 19) (Follicle-stimulating hormone primary response protein) (FSH primary response protein 1) (Leucine-rich primary response protein 1). | |||||
|
CENPI_MOUSE
|
||||||
| θ value | 6.88961 (rank : 39) | NC score | 0.021281 (rank : 40) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8K1K4, Q8BXU9 | Gene names | Cenpi, Fshprh1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein I (CENP-I) (Follicle-stimulating hormone primary response protein) (FSH primary response protein 1). | |||||
|
IRF7_HUMAN
|
||||||
| θ value | 6.88961 (rank : 40) | NC score | 0.010318 (rank : 48) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q92985, O00331, O00332, O00333, O75924 | Gene names | IRF7 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon regulatory factor 7 (IRF-7). | |||||
|
OGFR_HUMAN
|
||||||
| θ value | 6.88961 (rank : 41) | NC score | 0.005704 (rank : 51) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
SEL1L_HUMAN
|
||||||
| θ value | 6.88961 (rank : 42) | NC score | 0.023733 (rank : 38) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UBV2, Q6UWT6, Q9P1T9, Q9UHK7 | Gene names | SEL1L, TSA305 | |||
|
Domain Architecture |

|
|||||
| Description | Sel-1 homolog precursor (Suppressor of lin-12-like protein) (Sel-1L). | |||||
|
SEL1L_MOUSE
|
||||||
| θ value | 6.88961 (rank : 43) | NC score | 0.022367 (rank : 39) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Z2G6, Q9DBD8 | Gene names | Sel1l, Sel1h | |||
|
Domain Architecture |

|
|||||
| Description | Sel-1 homolog precursor (Suppressor of lin-12-like protein) (Sel-1L). | |||||
|
SNPH_HUMAN
|
||||||
| θ value | 6.88961 (rank : 44) | NC score | 0.012166 (rank : 45) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O15079 | Gene names | SNPH, KIAA0374 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Syntaphilin. | |||||
|
CD2L2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 45) | NC score | -0.000780 (rank : 54) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1439 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UQ88, O95227, O95228, O96012, Q12821, Q12853, Q12854, Q5QPR0, Q5QPR1, Q5QPR2, Q9UBC4, Q9UBI3, Q9UEI1, Q9UEI2, Q9UP53, Q9UP54, Q9UP55, Q9UP56, Q9UQ86, Q9UQ87, Q9UQ89 | Gene names | CDC2L2, PITSLREB | |||
|
Domain Architecture |

|
|||||
| Description | PITSLRE serine/threonine-protein kinase CDC2L2 (EC 2.7.11.22) (Galactosyltransferase-associated protein kinase p58/GTA) (Cell division cycle 2-like protein kinase 2) (CDK11). | |||||
|
ODO1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 46) | NC score | 0.010802 (rank : 47) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q60597, Q80Y57, Q8K2Z3 | Gene names | Ogdh | |||
|
Domain Architecture |

|
|||||
| Description | 2-oxoglutarate dehydrogenase E1 component, mitochondrial precursor (EC 1.2.4.2) (Alpha-ketoglutarate dehydrogenase). | |||||
|
CCND3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 47) | NC score | 0.448339 (rank : 22) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P30282, Q62391, Q99KP2 | Gene names | Ccnd3, Cyl-3 | |||
|
Domain Architecture |
|
|||||
| Description | G1/S-specific cyclin-D3. | |||||
|
CCNG2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 48) | NC score | 0.223719 (rank : 28) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q16589 | Gene names | CCNG2 | |||
|
Domain Architecture |
|
|||||
| Description | Cyclin-G2. | |||||
|
CCNG2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 49) | NC score | 0.224441 (rank : 27) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O08918, O35612 | Gene names | Ccng2 | |||
|
Domain Architecture |
|
|||||
| Description | Cyclin-G2. | |||||
|
CCNI_HUMAN
|
||||||
| θ value | θ > 10 (rank : 50) | NC score | 0.254340 (rank : 25) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q14094 | Gene names | CCNI | |||
|
Domain Architecture |
|
|||||
| Description | Cyclin-I. | |||||
|
CCNI_MOUSE
|
||||||
| θ value | θ > 10 (rank : 51) | NC score | 0.235392 (rank : 26) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9Z2V9 | Gene names | Ccni | |||
|
Domain Architecture |
|
|||||
| Description | Cyclin-I. | |||||
|
CCNK_HUMAN
|
||||||
| θ value | θ > 10 (rank : 52) | NC score | 0.072124 (rank : 30) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O75909, Q96B63, Q9NNY9 | Gene names | CCNK, CPR4 | |||
|
Domain Architecture |
|
|||||
| Description | Cyclin-K. | |||||
|
CCNK_MOUSE
|
||||||
| θ value | θ > 10 (rank : 53) | NC score | 0.061577 (rank : 33) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O88874, Q8R068 | Gene names | Ccnk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-K. | |||||
|
UNG2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 54) | NC score | 0.200228 (rank : 29) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P22674 | Gene names | UNG2 | |||
|
Domain Architecture |
|
|||||
| Description | Uracil-DNA glycosylase 2 (EC 3.2.2.-) (UDG 2). | |||||
|
CCNF_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 46 | |
| SwissProt Accessions | P41002, Q96EG9 | Gene names | CCNF | |||
|
Domain Architecture |

|
|||||
| Description | G2/mitotic-specific cyclin-F. | |||||
|
CCNF_MOUSE
|
||||||
| NC score | 0.992573 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 45 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P51944, Q60797, Q60799 | Gene names | Ccnf | |||
|
Domain Architecture |

|
|||||
| Description | G2/mitotic-specific cyclin-F. | |||||
|
CCNA2_MOUSE
|
||||||
| NC score | 0.733728 (rank : 3) | θ value | 3.07116e-24 (rank : 3) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P51943, Q61459 | Gene names | Ccna2, Ccna, Cyca | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin-A2 (Cyclin-A). | |||||
|
CCNA1_MOUSE
|
||||||
| NC score | 0.730380 (rank : 4) | θ value | 5.79196e-23 (rank : 5) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q61456 | Gene names | Ccna1 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin-A1. | |||||
|
CCNA2_HUMAN
|
||||||
| NC score | 0.729441 (rank : 5) | θ value | 3.39556e-23 (rank : 4) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P20248 | Gene names | CCNA2, CCN1, CCNA | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin-A2 (Cyclin-A). | |||||
|
CCNA1_HUMAN
|
||||||
| NC score | 0.721688 (rank : 6) | θ value | 4.58923e-20 (rank : 6) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P78396 | Gene names | CCNA1 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin-A1. | |||||
|
CCNB1_HUMAN
|
||||||
| NC score | 0.705529 (rank : 7) | θ value | 1.24977e-17 (rank : 7) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P14635 | Gene names | CCNB1, CCNB | |||
|
Domain Architecture |

|
|||||
| Description | G2/mitotic-specific cyclin-B1. | |||||
|
CCNB1_MOUSE
|
||||||
| NC score | 0.705307 (rank : 8) | θ value | 2.7842e-17 (rank : 8) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P24860 | Gene names | Ccnb1, Ccn-2, Ccnb1-rs13, Cycb, Cycb1 | |||
|
Domain Architecture |

|
|||||
| Description | G2/mitotic-specific cyclin-B1. | |||||
|
CCNB2_MOUSE
|
||||||
| NC score | 0.695675 (rank : 9) | θ value | 5.25075e-16 (rank : 10) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P30276 | Gene names | Ccnb2, Cycb2 | |||
|
Domain Architecture |

|
|||||
| Description | G2/mitotic-specific cyclin-B2. | |||||
|
CCNB2_HUMAN
|
||||||
| NC score | 0.692818 (rank : 10) | θ value | 1.52774e-15 (rank : 11) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O95067 | Gene names | CCNB2 | |||
|
Domain Architecture |

|
|||||
| Description | G2/mitotic-specific cyclin-B2. | |||||
|
CCNB3_MOUSE
|
||||||
| NC score | 0.658690 (rank : 11) | θ value | 1.058e-16 (rank : 9) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q810T2, Q810T3, Q8VDC8 | Gene names | Ccnb3, Cycb3 | |||
|
Domain Architecture |

|
|||||
| Description | G2/mitotic-specific cyclin-B3. | |||||
|
CCNB3_HUMAN
|
||||||
| NC score | 0.562769 (rank : 12) | θ value | 7.09661e-13 (rank : 12) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 368 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8WWL7, Q96SB5, Q96SB6, Q96SB7, Q9NT38 | Gene names | CCNB3, CYCB3 | |||
|
Domain Architecture |

|
|||||
| Description | G2/mitotic-specific cyclin-B3. | |||||
|
CCNE2_MOUSE
|
||||||
| NC score | 0.543966 (rank : 13) | θ value | 2.21117e-06 (rank : 18) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q9Z238, Q9QWU5 | Gene names | Ccne2 | |||
|
Domain Architecture |

|
|||||
| Description | G1/S-specific cyclin-E2. | |||||
|
CCNE2_HUMAN
|
||||||
| NC score | 0.541131 (rank : 14) | θ value | 1.69304e-06 (rank : 16) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O96020, O95439 | Gene names | CCNE2 | |||
|
Domain Architecture |

|
|||||
| Description | G1/S-specific cyclin-E2. | |||||
|
CCND1_MOUSE
|
||||||
| NC score | 0.535376 (rank : 15) | θ value | 3.41135e-07 (rank : 15) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P25322 | Gene names | Ccnd1, Cyl-1 | |||
|
Domain Architecture |

|
|||||
| Description | G1/S-specific cyclin-D1. | |||||
|
CCND1_HUMAN
|
||||||
| NC score | 0.533628 (rank : 16) | θ value | 1.99992e-07 (rank : 14) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P24385, Q6LEF0 | Gene names | CCND1, BCL1, PRAD1 | |||
|
Domain Architecture |

|
|||||
| Description | G1/S-specific cyclin-D1 (PRAD1 oncogene) (BCL-1 oncogene). | |||||
|
CCNE1_HUMAN
|
||||||
| NC score | 0.528171 (rank : 17) | θ value | 0.000121331 (rank : 19) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P24864, Q14091, Q8NFG1, Q92501 | Gene names | CCNE1, CCNE | |||
|
Domain Architecture |

|
|||||
| Description | G1/S-specific cyclin-E1. | |||||
|
CCNE1_MOUSE
|
||||||
| NC score | 0.520961 (rank : 18) | θ value | 0.000158464 (rank : 20) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q61457 | Gene names | Ccne1, Ccne | |||
|
Domain Architecture |

|
|||||
| Description | G1/S-specific cyclin-E1. | |||||
|
CCND2_HUMAN
|
||||||
| NC score | 0.512096 (rank : 19) | θ value | 4.0297e-08 (rank : 13) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P30279, Q13955 | Gene names | CCND2 | |||
|
Domain Architecture |

|
|||||
| Description | G1/S-specific cyclin-D2. | |||||
|
CCND2_MOUSE
|
||||||
| NC score | 0.497316 (rank : 20) | θ value | 2.21117e-06 (rank : 17) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P30280 | Gene names | Ccnd2, Cyl-2 | |||
|
Domain Architecture |

|
|||||
| Description | G1/S-specific cyclin-D2. | |||||
|
CCND3_HUMAN
|
||||||
| NC score | 0.476354 (rank : 21) | θ value | 0.0193708 (rank : 22) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P30281, Q5T8J0, Q96F49 | Gene names | CCND3 | |||
|
Domain Architecture |

|
|||||
| Description | G1/S-specific cyclin-D3. | |||||
|
CCND3_MOUSE
|
||||||
| NC score | 0.448339 (rank : 22) | θ value | θ > 10 (rank : 47) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P30282, Q62391, Q99KP2 | Gene names | Ccnd3, Cyl-3 | |||
|
Domain Architecture |

|
|||||
| Description | G1/S-specific cyclin-D3. | |||||
|
CCNG1_MOUSE
|
||||||
| NC score | 0.285339 (rank : 23) | θ value | 0.813845 (rank : 27) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P51945, O54779 | Gene names | Ccng1, Ccng | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin-G1 (Cyclin-G). | |||||
|
CCNG1_HUMAN
|
||||||
| NC score | 0.272717 (rank : 24) | θ value | 4.03905 (rank : 36) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P51959, Q15757, Q96L32 | Gene names | CCNG1, CCNG, CYCG1 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin-G1 (Cyclin-G). | |||||
|
CCNI_HUMAN
|
||||||
| NC score | 0.254340 (rank : 25) | θ value | θ > 10 (rank : 50) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q14094 | Gene names | CCNI | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin-I. | |||||
|
CCNI_MOUSE
|
||||||
| NC score | 0.235392 (rank : 26) | θ value | θ > 10 (rank : 51) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9Z2V9 | Gene names | Ccni | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin-I. | |||||
|
CCNG2_MOUSE
|
||||||
| NC score | 0.224441 (rank : 27) | θ value | θ > 10 (rank : 49) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | O08918, O35612 | Gene names | Ccng2 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin-G2. | |||||
|
CCNG2_HUMAN
|
||||||
| NC score | 0.223719 (rank : 28) | θ value | θ > 10 (rank : 48) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q16589 | Gene names | CCNG2 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin-G2. | |||||
|
UNG2_HUMAN
|
||||||
| NC score | 0.200228 (rank : 29) | θ value | θ > 10 (rank : 54) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P22674 | Gene names | UNG2 | |||
|
Domain Architecture |

|
|||||
| Description | Uracil-DNA glycosylase 2 (EC 3.2.2.-) (UDG 2). | |||||
|
CCNK_HUMAN
|
||||||
| NC score | 0.072124 (rank : 30) | θ value | θ > 10 (rank : 52) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | O75909, Q96B63, Q9NNY9 | Gene names | CCNK, CPR4 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin-K. | |||||
|
U2AF2_HUMAN
|
||||||
| NC score | 0.072084 (rank : 31) | θ value | 0.0113563 (rank : 21) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 192 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P26368 | Gene names | U2AF2, U2AF65 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor U2AF 65 kDa subunit (U2 auxiliary factor 65 kDa subunit) (U2 snRNP auxiliary factor large subunit) (hU2AF(65)). | |||||
|
GIDRP_HUMAN
|
||||||
| NC score | 0.067853 (rank : 32) | θ value | 0.125558 (rank : 23) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7Z5L2, O60598, Q5W0B4, Q5W0B5, Q86VT9, Q8N9H0 | Gene names | GIDRP88, C10orf28 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Growth inhibition and differentiation-related protein 88 (Putative mitochondrial space protein 32.1). | |||||
|
CCNK_MOUSE
|
||||||
| NC score | 0.061577 (rank : 33) | θ value | θ > 10 (rank : 53) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O88874, Q8R068 | Gene names | Ccnk | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclin-K. | |||||
|
U2AF2_MOUSE
|
||||||
| NC score | 0.052719 (rank : 34) | θ value | 0.163984 (rank : 24) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 192 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P26369 | Gene names | U2af2, U2af65 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor U2AF 65 kDa subunit (U2 auxiliary factor 65 kDa subunit) (U2 snRNP auxiliary factor large subunit). | |||||
|
VHL_HUMAN
|
||||||
| NC score | 0.032366 (rank : 35) | θ value | 2.36792 (rank : 31) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P40337, Q13599 | Gene names | VHL | |||
|
Domain Architecture |

|
|||||
| Description | Von Hippel-Lindau disease tumor suppressor (pVHL) (G7 protein). | |||||
|
CDSN_MOUSE
|
||||||
| NC score | 0.029627 (rank : 36) | θ value | 1.81305 (rank : 29) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 131 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7TPC1 | Gene names | Cdsn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Corneodesmosin precursor. | |||||
|
FMN1B_MOUSE
|
||||||
| NC score | 0.029069 (rank : 37) | θ value | 1.38821 (rank : 28) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 447 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q05859 | Gene names | Fmn, Ld | |||
|
Domain Architecture |

|
|||||
| Description | Formin-1 isoform IV (Limb deformity protein). | |||||
|
SEL1L_HUMAN
|
||||||
| NC score | 0.023733 (rank : 38) | θ value | 6.88961 (rank : 42) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UBV2, Q6UWT6, Q9P1T9, Q9UHK7 | Gene names | SEL1L, TSA305 | |||
|
Domain Architecture |

|
|||||
| Description | Sel-1 homolog precursor (Suppressor of lin-12-like protein) (Sel-1L). | |||||
|
SEL1L_MOUSE
|
||||||
| NC score | 0.022367 (rank : 39) | θ value | 6.88961 (rank : 43) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Z2G6, Q9DBD8 | Gene names | Sel1l, Sel1h | |||
|
Domain Architecture |

|
|||||
| Description | Sel-1 homolog precursor (Suppressor of lin-12-like protein) (Sel-1L). | |||||
|
CENPI_MOUSE
|
||||||
| NC score | 0.021281 (rank : 40) | θ value | 6.88961 (rank : 39) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8K1K4, Q8BXU9 | Gene names | Cenpi, Fshprh1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein I (CENP-I) (Follicle-stimulating hormone primary response protein) (FSH primary response protein 1). | |||||
|
CENPI_HUMAN
|
||||||
| NC score | 0.021253 (rank : 41) | θ value | 6.88961 (rank : 38) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q92674, Q5JWZ9, Q96ED0 | Gene names | CENPI, FSHPRH1, ICEN19, LRPR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centromere protein I (CENP-I) (Interphase centromere complex protein 19) (Follicle-stimulating hormone primary response protein) (FSH primary response protein 1) (Leucine-rich primary response protein 1). | |||||
|
CCDC9_MOUSE
|
||||||
| NC score | 0.020573 (rank : 42) | θ value | 4.03905 (rank : 35) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 196 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8VC31 | Gene names | Ccdc9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 9. | |||||
|
GP152_HUMAN
|
||||||
| NC score | 0.015838 (rank : 43) | θ value | 0.47712 (rank : 26) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 636 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8TDT2, Q86SM0 | Gene names | GPR152, PGR5 | |||
|
Domain Architecture |

|
|||||
| Description | Probable G-protein coupled receptor 152 (G-protein coupled receptor PGR5). | |||||
|
SYNJ1_MOUSE
|
||||||
| NC score | 0.013601 (rank : 44) | θ value | 3.0926 (rank : 34) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 516 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8CHC4 | Gene names | Synj1, Kiaa0910 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptojanin-1 (EC 3.1.3.36) (Synaptic inositol-1,4,5-trisphosphate 5- phosphatase 1). | |||||
|
SNPH_HUMAN
|
||||||
| NC score | 0.012166 (rank : 45) | θ value | 6.88961 (rank : 44) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O15079 | Gene names | SNPH, KIAA0374 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Syntaphilin. | |||||
|
BAT2_HUMAN
|
||||||
| NC score | 0.011713 (rank : 46) | θ value | 6.88961 (rank : 37) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 931 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P48634, O95875, Q5SQ29, Q5SQ30, Q5ST84, Q5STX6, Q5STX7, Q68DW9, Q6P9P7, Q6PIN1, Q96QC6 | Gene names | BAT2, G2 | |||
|
Domain Architecture |

|
|||||
| Description | Large proline-rich protein BAT2 (HLA-B-associated transcript 2). | |||||
|
ODO1_MOUSE
|
||||||
| NC score | 0.010802 (rank : 47) | θ value | 8.99809 (rank : 46) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q60597, Q80Y57, Q8K2Z3 | Gene names | Ogdh | |||
|
Domain Architecture |

|
|||||
| Description | 2-oxoglutarate dehydrogenase E1 component, mitochondrial precursor (EC 1.2.4.2) (Alpha-ketoglutarate dehydrogenase). | |||||
|
IRF7_HUMAN
|
||||||
| NC score | 0.010318 (rank : 48) | θ value | 6.88961 (rank : 40) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q92985, O00331, O00332, O00333, O75924 | Gene names | IRF7 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon regulatory factor 7 (IRF-7). | |||||
|
NACAM_MOUSE
|
||||||
| NC score | 0.008533 (rank : 49) | θ value | 3.0926 (rank : 33) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
MAGE1_HUMAN
|
||||||
| NC score | 0.007522 (rank : 50) | θ value | 2.36792 (rank : 30) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1389 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9HCI5, Q86TG0, Q8TD92, Q9H216 | Gene names | MAGEE1, HCA1, KIAA1587 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Melanoma-associated antigen E1 (MAGE-E1 antigen) (Hepatocellular carcinoma-associated protein 1). | |||||
|
OGFR_HUMAN
|
||||||
| NC score | 0.005704 (rank : 51) | θ value | 6.88961 (rank : 41) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
EGR4_HUMAN
|
||||||
| NC score | 0.003532 (rank : 52) | θ value | 0.47712 (rank : 25) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 939 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q05215 | Gene names | EGR4 | |||
|
Domain Architecture |

|
|||||
| Description | Early growth response protein 4 (EGR-4) (AT133). | |||||
|
CD2L1_MOUSE
|
||||||
| NC score | 0.000058 (rank : 53) | θ value | 3.0926 (rank : 32) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1463 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P24788, Q3UI03, Q61399, Q7TST4, Q8BP53 | Gene names | Cdc2l1 | |||
|
Domain Architecture |

|
|||||
| Description | PITSLRE serine/threonine-protein kinase CDC2L1 (EC 2.7.11.22) (Galactosyltransferase-associated protein kinase p58/GTA) (Cell division cycle 2-like protein kinase 1). | |||||
|
CD2L2_HUMAN
|
||||||
| NC score | -0.000780 (rank : 54) | θ value | 8.99809 (rank : 45) | |||
| Query Neighborhood Hits | 46 | Target Neighborhood Hits | 1439 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UQ88, O95227, O95228, O96012, Q12821, Q12853, Q12854, Q5QPR0, Q5QPR1, Q5QPR2, Q9UBC4, Q9UBI3, Q9UEI1, Q9UEI2, Q9UP53, Q9UP54, Q9UP55, Q9UP56, Q9UQ86, Q9UQ87, Q9UQ89 | Gene names | CDC2L2, PITSLREB | |||
|
Domain Architecture |

|
|||||
| Description | PITSLRE serine/threonine-protein kinase CDC2L2 (EC 2.7.11.22) (Galactosyltransferase-associated protein kinase p58/GTA) (Cell division cycle 2-like protein kinase 2) (CDK11). | |||||