Please be patient as the page loads
|
BSCL2_HUMAN
|
||||||
| SwissProt Accessions | Q96G97, Q96SV1, Q9BSQ0 | Gene names | BSCL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Seipin (Bernardinelli-Seip congenital lipodystrophy type 2 protein). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
BSCL2_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 148 | |
| SwissProt Accessions | Q96G97, Q96SV1, Q9BSQ0 | Gene names | BSCL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Seipin (Bernardinelli-Seip congenital lipodystrophy type 2 protein). | |||||
|
BSCL2_MOUSE
|
||||||
| θ value | 5.9214e-177 (rank : 2) | NC score | 0.942970 (rank : 2) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9Z2E9, Q810B0, Q9JJC2, Q9JMF1 | Gene names | Bscl2, Gng3lg | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Seipin (Bernardinelli-Seip congenital lipodystrophy type 2 protein homolog). | |||||
|
HUWE1_HUMAN
|
||||||
| θ value | 0.00665767 (rank : 3) | NC score | 0.101893 (rank : 17) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 443 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q7Z6Z7, O15029, Q4G2Z2, Q5H961, Q6P4D0, Q8NG67, Q9BUI0, Q9HCJ4, Q9NSL6, Q9P0A9 | Gene names | HUWE1, KIAA0312, KIAA1578, UREB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT, UBA and WWE domain-containing protein 1 (EC 6.3.2.-) (E3 ubiquitin protein ligase URE-B1) (Mcl-1 ubiquitin ligase E3) (Mule) (ARF-binding protein 1) (ARF-BP1). | |||||
|
HUWE1_MOUSE
|
||||||
| θ value | 0.00665767 (rank : 4) | NC score | 0.103027 (rank : 16) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q7TMY8, Q4G2Z1, Q5BMM7, Q6NS61, Q8BNJ7, Q8CFH2, Q8VD14, Q921M5, Q9R0P2 | Gene names | Huwe1, Kiaa0312, Ureb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT, UBA and WWE domain-containing protein 1 (EC 6.3.2.-) (E3 ubiquitin protein ligase URE-B1) (E3Histone). | |||||
|
T2FA_HUMAN
|
||||||
| θ value | 0.00665767 (rank : 5) | NC score | 0.136214 (rank : 6) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P35269, Q9BWN0 | Gene names | GTF2F1, RAP74 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor IIF alpha subunit (EC 2.7.11.1) (TFIIF-alpha) (Transcription initiation factor RAP74) (General transcription factor IIF polypeptide 1 74 kDa subunit protein). | |||||
|
FOG1_MOUSE
|
||||||
| θ value | 0.0330416 (rank : 6) | NC score | 0.045122 (rank : 87) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1237 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O35615 | Gene names | Zfpm1, Fog, Fog1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein ZFPM1 (Zinc finger protein multitype 1) (Friend of GATA protein 1) (Friend of GATA-1) (FOG-1). | |||||
|
CCD96_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 7) | NC score | 0.057107 (rank : 61) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9CR92 | Gene names | Ccdc96 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 96. | |||||
|
EDD1_HUMAN
|
||||||
| θ value | 0.0431538 (rank : 8) | NC score | 0.110273 (rank : 12) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O95071, O94970, Q9NPL3 | Gene names | EDD1, EDD, HYD, KIAA0896 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-protein ligase EDD1 (EC 6.3.2.-) (Hyperplastic discs protein homolog) (hHYD) (Progestin-induced protein). | |||||
|
EDD1_MOUSE
|
||||||
| θ value | 0.0563607 (rank : 9) | NC score | 0.109119 (rank : 13) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q80TP3, Q698K9, Q6PEQ8, Q6PFQ9, Q80VL4, Q810V6, Q9CXE9 | Gene names | Edd1, Edd, Kiaa0896 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase EDD1 (EC 6.3.2.-) (Hyperplastic discs protein homolog). | |||||
|
LIPS_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 10) | NC score | 0.075517 (rank : 31) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 314 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q05469, Q3LRT2, Q6NSL7 | Gene names | LIPE | |||
|
Domain Architecture |
|
|||||
| Description | Hormone-sensitive lipase (EC 3.1.1.79) (HSL). | |||||
|
RP1L1_HUMAN
|
||||||
| θ value | 0.125558 (rank : 11) | NC score | 0.077989 (rank : 26) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1072 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q8IWN7, Q86SQ1, Q8IWN8, Q8IWN9, Q8IWP0, Q8IWP1, Q8IWP2 | Gene names | RP1L1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinitis pigmentosa 1-like 1 protein. | |||||
|
FGD5_HUMAN
|
||||||
| θ value | 0.163984 (rank : 12) | NC score | 0.035825 (rank : 100) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6ZNL6, Q6MZY1, Q7Z303, Q8IYP3, Q8N861, Q8N8G4 | Gene names | FGD5, ZFYVE23 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 5 (Zinc finger FYVE domain-containing protein 23). | |||||
|
AKA12_HUMAN
|
||||||
| θ value | 0.21417 (rank : 13) | NC score | 0.126972 (rank : 9) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | Q02952, O00310, O00498, Q99970 | Gene names | AKAP12, AKAP250 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 12 (A-kinase anchor protein 250 kDa) (AKAP 250) (Myasthenia gravis autoantigen gravin). | |||||
|
UBP35_HUMAN
|
||||||
| θ value | 0.21417 (rank : 14) | NC score | 0.062495 (rank : 49) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9P2H5 | Gene names | USP35, KIAA1372, USP34 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 35 (EC 3.1.2.15) (Ubiquitin thioesterase 35) (Ubiquitin-specific-processing protease 35) (Deubiquitinating enzyme 35). | |||||
|
CAH9_HUMAN
|
||||||
| θ value | 0.279714 (rank : 15) | NC score | 0.033835 (rank : 102) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q16790 | Gene names | CA9, G250, MN | |||
|
Domain Architecture |
|
|||||
| Description | Carbonic anhydrase 9 precursor (EC 4.2.1.1) (Carbonic anhydrase IX) (Carbonate dehydratase IX) (CA-IX) (CAIX) (Membrane antigen MN) (P54/58N) (Renal cell carcinoma-associated antigen G250) (RCC- associated antigen G250) (pMW1). | |||||
|
MYO9B_HUMAN
|
||||||
| θ value | 0.279714 (rank : 16) | NC score | 0.024767 (rank : 120) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 641 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q13459, O75314, Q9NUJ2, Q9UHN0 | Gene names | MYO9B, MYR5 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9B (Myosin IXb) (Unconventional myosin-9b). | |||||
|
NSBP1_MOUSE
|
||||||
| θ value | 0.279714 (rank : 17) | NC score | 0.158327 (rank : 5) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 96 | |
| SwissProt Accessions | Q9JL35, O88832, Q8VC71, Q9CUW1 | Gene names | Nsbp1, Garp45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1 (Nucleosome-binding protein 45) (NBP-45) (GARP45 protein). | |||||
|
RGS3_HUMAN
|
||||||
| θ value | 0.279714 (rank : 18) | NC score | 0.040742 (rank : 95) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
SH3BG_MOUSE
|
||||||
| θ value | 0.279714 (rank : 19) | NC score | 0.087104 (rank : 20) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9WUZ7 | Gene names | Sh3bgr | |||
|
Domain Architecture |

|
|||||
| Description | SH3 domain-binding glutamic acid-rich protein (SH3BGR protein). | |||||
|
IWS1_MOUSE
|
||||||
| θ value | 0.365318 (rank : 20) | NC score | 0.129535 (rank : 8) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 109 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
TCAL3_MOUSE
|
||||||
| θ value | 0.365318 (rank : 21) | NC score | 0.076799 (rank : 30) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8R0A5, Q9CYK5, Q9D1S4 | Gene names | Tceal3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 3 (TCEA-like protein 3) (Transcription elongation factor S-II protein-like 3). | |||||
|
TRI44_HUMAN
|
||||||
| θ value | 0.365318 (rank : 22) | NC score | 0.079147 (rank : 25) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 393 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q96DX7, Q96QY2, Q9UGK0 | Gene names | TRIM44, DIPB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 44 (Protein DIPB). | |||||
|
CD2L1_MOUSE
|
||||||
| θ value | 0.47712 (rank : 23) | NC score | 0.017105 (rank : 138) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1463 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P24788, Q3UI03, Q61399, Q7TST4, Q8BP53 | Gene names | Cdc2l1 | |||
|
Domain Architecture |

|
|||||
| Description | PITSLRE serine/threonine-protein kinase CDC2L1 (EC 2.7.11.22) (Galactosyltransferase-associated protein kinase p58/GTA) (Cell division cycle 2-like protein kinase 1). | |||||
|
CSTN1_HUMAN
|
||||||
| θ value | 0.47712 (rank : 24) | NC score | 0.051336 (rank : 77) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O94985 | Gene names | CLSTN1, CS1, KIAA0911 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calsyntenin-1 precursor. | |||||
|
MYST4_HUMAN
|
||||||
| θ value | 0.47712 (rank : 25) | NC score | 0.053064 (rank : 71) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8WYB5, O15087, Q86Y05, Q8WU81, Q9UKW2, Q9UKW3, Q9UKX0 | Gene names | MYST4, KIAA0383, MORF, MOZ2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST4 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 4) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 4) (Histone acetyltransferase MOZ2) (Monocytic leukemia zinc finger protein- related factor) (Histone acetyltransferase MORF). | |||||
|
MYT1_MOUSE
|
||||||
| θ value | 0.47712 (rank : 26) | NC score | 0.073863 (rank : 32) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8CFC2, O08995, Q8CFH1 | Gene names | Myt1, Kiaa0835, Nzf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myelin transcription factor 1 (MyT1) (Neural zinc finger factor 2) (NZF-2). | |||||
|
TCAL5_HUMAN
|
||||||
| θ value | 0.47712 (rank : 27) | NC score | 0.134984 (rank : 7) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 806 | Shared Neighborhood Hits | 96 | |
| SwissProt Accessions | Q5H9L2 | Gene names | TCEAL5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 5 (TCEA-like protein 5) (Transcription elongation factor S-II protein-like 5). | |||||
|
BASP_MOUSE
|
||||||
| θ value | 0.62314 (rank : 28) | NC score | 0.077272 (rank : 28) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q91XV3 | Gene names | Basp1, Nap22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
FOG1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 29) | NC score | 0.034965 (rank : 101) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 981 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8IX07 | Gene names | ZFPM1, FOG1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein ZFPM1 (Zinc finger protein multitype 1) (Friend of GATA protein 1) (Friend of GATA-1) (FOG-1). | |||||
|
LSD1_HUMAN
|
||||||
| θ value | 0.62314 (rank : 30) | NC score | 0.056143 (rank : 63) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O60341, Q5TH94, Q5TH95, Q86VT7, Q8IXK4, Q8NDP6, Q8TAZ3, Q96AW4 | Gene names | AOF2, KIAA0601, LSD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lysine-specific histone demethylase 1 (EC 1.-.-.-) (Flavin-containing amine oxidase domain-containing protein 2) (BRAF35-HDAC complex protein BHC110). | |||||
|
LSD1_MOUSE
|
||||||
| θ value | 0.62314 (rank : 31) | NC score | 0.057537 (rank : 59) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 224 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q6ZQ88, Q6PB53, Q8VEA1 | Gene names | Aof2, Kiaa0601, Lsd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lysine-specific histone demethylase 1 (EC 1.-.-.-) (Flavin-containing amine oxidase domain-containing protein 2) (BRAF35-HDAC complex protein BHC110). | |||||
|
NOL3_MOUSE
|
||||||
| θ value | 0.62314 (rank : 32) | NC score | 0.080638 (rank : 24) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9D1X0 | Gene names | Nol3 | |||
|
Domain Architecture |
|
|||||
| Description | Nucleolar protein 3. | |||||
|
RPGR1_MOUSE
|
||||||
| θ value | 0.62314 (rank : 33) | NC score | 0.065630 (rank : 43) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 713 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9EPQ2, Q8CAC2, Q8CDJ9, Q91WE0, Q9CUK6, Q9D5Q1 | Gene names | Rpgrip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator-interacting protein 1 (RPGR-interacting protein 1). | |||||
|
SCN5A_HUMAN
|
||||||
| θ value | 0.62314 (rank : 34) | NC score | 0.024247 (rank : 124) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q14524 | Gene names | SCN5A | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 5 subunit alpha (Sodium channel protein type V subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.5) (Sodium channel protein, cardiac muscle alpha-subunit) (HH1). | |||||
|
TCF8_HUMAN
|
||||||
| θ value | 0.62314 (rank : 35) | NC score | 0.026977 (rank : 113) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1057 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P37275, Q12924, Q13800 | Gene names | TCF8, AREB6 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 8 (NIL-2-A zinc finger protein) (Negative regulator of IL2). | |||||
|
TXD13_HUMAN
|
||||||
| θ value | 0.62314 (rank : 36) | NC score | 0.069244 (rank : 38) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9H1E5, Q8N4P7, Q8NCC1, Q9UJA1, Q9ULQ8 | Gene names | TXNDC13, KIAA1162 | |||
|
Domain Architecture |
|
|||||
| Description | Thioredoxin domain-containing protein 13 precursor. | |||||
|
CASC3_HUMAN
|
||||||
| θ value | 0.813845 (rank : 37) | NC score | 0.068221 (rank : 39) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O15234 | Gene names | CASC3, MLN51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein CASC3 (Cancer susceptibility candidate gene 3 protein) (Metastatic lymph node protein 51) (Protein MLN 51) (Protein barentsz) (Btz). | |||||
|
CD2L1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 38) | NC score | 0.016543 (rank : 141) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1467 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P21127, O95265, Q12817, Q12818, Q12819, Q12820, Q12822, Q8N530, Q9NZS5, Q9UBJ0, Q9UBQ1, Q9UBR0, Q9UNY2, Q9UP57, Q9UP58, Q9UP59 | Gene names | CDC2L1, PITSLREA, PK58 | |||
|
Domain Architecture |

|
|||||
| Description | PITSLRE serine/threonine-protein kinase CDC2L1 (EC 2.7.11.22) (Galactosyltransferase-associated protein kinase p58/GTA) (Cell division cycle 2-like protein kinase 1) (CLK-1) (CDK11) (p58 CLK-1). | |||||
|
JHD3B_HUMAN
|
||||||
| θ value | 0.813845 (rank : 39) | NC score | 0.031474 (rank : 107) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O94953, O75274, Q6P3R5, Q9P1V1, Q9UF40 | Gene names | JMJD2B, JHDM3B, KIAA0876 | |||
|
Domain Architecture |

|
|||||
| Description | JmjC domain-containing histone demethylation protein 3B (EC 1.14.11.-) (Jumonji domain-containing protein 2B). | |||||
|
AP3B2_HUMAN
|
||||||
| θ value | 1.06291 (rank : 40) | NC score | 0.047461 (rank : 84) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q13367, O14808 | Gene names | AP3B2 | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit beta-2 (Adapter-related protein complex 3 beta-2 subunit) (Beta3B-adaptin) (Adaptor protein complex AP-3 beta-2 subunit) (AP-3 complex beta-2 subunit) (Clathrin assembly protein complex 3 beta-2 large chain) (Neuron-specific vesicle coat protein beta-NAP). | |||||
|
CENPB_HUMAN
|
||||||
| θ value | 1.06291 (rank : 41) | NC score | 0.081104 (rank : 23) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P07199 | Gene names | CENPB | |||
|
Domain Architecture |

|
|||||
| Description | Major centromere autoantigen B (Centromere protein B) (CENP-B). | |||||
|
CIR_HUMAN
|
||||||
| θ value | 1.06291 (rank : 42) | NC score | 0.049765 (rank : 81) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q86X95, O95367, Q12804, Q4G1B9, Q6PJI4, Q8IWI2 | Gene names | CIR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CBF1-interacting corepressor (Recepin). | |||||
|
JHD2C_HUMAN
|
||||||
| θ value | 1.06291 (rank : 43) | NC score | 0.030811 (rank : 108) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q15652, Q5SQZ8, Q5SQZ9, Q5SR00, Q7Z3E7, Q8N3U0, Q96KB9, Q9P2G7 | Gene names | JMJD1C, JHDM2C, KIAA1380, TRIP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable JmjC domain-containing histone demethylation protein 2C (EC 1.14.11.-) (Jumonji domain-containing protein 1C) (Thyroid receptor-interacting protein 8) (TRIP-8). | |||||
|
MATR3_HUMAN
|
||||||
| θ value | 1.06291 (rank : 44) | NC score | 0.072733 (rank : 33) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P43243, Q9UHW0, Q9UQ27 | Gene names | MATR3, KIAA0723 | |||
|
Domain Architecture |

|
|||||
| Description | Matrin-3. | |||||
|
RGP1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 45) | NC score | 0.065302 (rank : 44) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P46061, Q60801 | Gene names | Rangap1, Fug1 | |||
|
Domain Architecture |
|
|||||
| Description | Ran GTPase-activating protein 1. | |||||
|
SYTL1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 46) | NC score | 0.033626 (rank : 104) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8IYJ3, Q96BB6, Q96GU6, Q96S89, Q96SI0 | Gene names | SYTL1, SLP1 | |||
|
Domain Architecture |
|
|||||
| Description | Synaptotagmin-like protein 1 (Exophilin-7) (Protein JFC1) (SB146). | |||||
|
NFM_MOUSE
|
||||||
| θ value | 1.38821 (rank : 47) | NC score | 0.022640 (rank : 127) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1159 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P08553, Q61961 | Gene names | Nef3, Nefm, Nfm | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet M protein (160 kDa neurofilament protein) (Neurofilament medium polypeptide) (NF-M). | |||||
|
NGEF_MOUSE
|
||||||
| θ value | 1.38821 (rank : 48) | NC score | 0.021933 (rank : 131) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8CHT1, Q8R204, Q923H2, Q9JHT9 | Gene names | Ngef | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ephexin-1 (Eph-interacting exchange protein) (Neuronal guanine nucleotide exchange factor). | |||||
|
NOLC1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 49) | NC score | 0.059669 (rank : 55) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q14978, Q15030 | Gene names | NOLC1, KIAA0035 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar phosphoprotein p130 (Nucleolar 130 kDa protein) (140 kDa nucleolar phosphoprotein) (Nopp140) (Nucleolar and coiled-body phosphoprotein 1). | |||||
|
SPEG_HUMAN
|
||||||
| θ value | 1.38821 (rank : 50) | NC score | 0.011582 (rank : 158) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q15772, Q27J74, Q695L1, Q6FGA6, Q6ZQW1, Q6ZTL8, Q9P2P9 | Gene names | SPEG, APEG1, KIAA1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle preferentially expressed protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
TMAP1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 51) | NC score | 0.072072 (rank : 36) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P84157 | Gene names | TMAP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane anchor protein 1. | |||||
|
UBF1_HUMAN
|
||||||
| θ value | 1.38821 (rank : 52) | NC score | 0.055665 (rank : 67) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 415 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P17480 | Gene names | UBTF, UBF, UBF1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar transcription factor 1 (Upstream-binding factor 1) (UBF-1) (Autoantigen NOR-90). | |||||
|
AN32C_HUMAN
|
||||||
| θ value | 1.81305 (rank : 53) | NC score | 0.060886 (rank : 52) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O43423 | Gene names | ANP32C, PP32R1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acidic leucine-rich nuclear phosphoprotein 32 family member C (Phosphoprotein 32-related protein 1) (Tumorigenic protein pp32r1). | |||||
|
CAC1C_MOUSE
|
||||||
| θ value | 1.81305 (rank : 54) | NC score | 0.024753 (rank : 121) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q01815, Q04476, Q61242, Q99242 | Gene names | Cacna1c, Cach2, Cacn2, Cacnl1a1, Cchl1a1 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1C (Voltage- gated calcium channel subunit alpha Cav1.2) (Calcium channel, L type, alpha-1 polypeptide, isoform 1, cardiac muscle) (Mouse brain class C) (MBC) (MELC-CC). | |||||
|
NCOR1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 55) | NC score | 0.050276 (rank : 79) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 521 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q60974, Q60812 | Gene names | Ncor1, Rxrip13 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 1 (N-CoR1) (N-CoR) (Retinoid X receptor- interacting protein 13) (RIP13). | |||||
|
OGFR_HUMAN
|
||||||
| θ value | 1.81305 (rank : 56) | NC score | 0.042332 (rank : 91) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
PTMS_HUMAN
|
||||||
| θ value | 1.81305 (rank : 57) | NC score | 0.097394 (rank : 18) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P20962 | Gene names | PTMS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Parathymosin. | |||||
|
SPTN2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 58) | NC score | 0.011741 (rank : 157) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 535 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O15020, O14872, O14873 | Gene names | SPTBN2, KIAA0302 | |||
|
Domain Architecture |

|
|||||
| Description | Spectrin beta chain, brain 2 (Spectrin, non-erythroid beta chain 2) (Beta-III spectrin). | |||||
|
TCF8_MOUSE
|
||||||
| θ value | 1.81305 (rank : 59) | NC score | 0.024453 (rank : 123) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1067 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q64318, Q62519 | Gene names | Tcf8, Zfhx1a, Zfx1a, Zfx1ha | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 8 (Zinc finger homeobox protein 1a) (MEB1) (Delta EF1). | |||||
|
TRI44_MOUSE
|
||||||
| θ value | 1.81305 (rank : 60) | NC score | 0.065146 (rank : 45) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 327 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9QXA7, Q80SV9, Q9JHR8 | Gene names | Trim44, Dipb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 44 (Protein DIPB) (Protein Mc7). | |||||
|
YTDC1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 61) | NC score | 0.077160 (rank : 29) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q96MU7, Q7Z622, Q8TF35 | Gene names | YTHDC1, KIAA1966, YT521 | |||
|
Domain Architecture |

|
|||||
| Description | YTH domain-containing protein 1 (Putative splicing factor YT521). | |||||
|
AN32A_HUMAN
|
||||||
| θ value | 2.36792 (rank : 62) | NC score | 0.058789 (rank : 56) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 234 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P39687 | Gene names | ANP32A, C15orf1, LANP, MAPM, PHAP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acidic leucine-rich nuclear phosphoprotein 32 family member A (Potent heat-stable protein phosphatase 2A inhibitor I1PP2A) (Acidic nuclear phosphoprotein pp32) (Leucine-rich acidic nuclear protein) (Lanp) (Putative HLA-DR-associated protein I) (PHAPI) (Mapmodulin). | |||||
|
AN32B_MOUSE
|
||||||
| θ value | 2.36792 (rank : 63) | NC score | 0.061271 (rank : 50) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9EST5 | Gene names | Anp32b, Pal31 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acidic leucine-rich nuclear phosphoprotein 32 family member B (Proliferation-related acidic leucine-rich protein PAL31). | |||||
|
AN32E_HUMAN
|
||||||
| θ value | 2.36792 (rank : 64) | NC score | 0.062908 (rank : 48) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 291 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9BTT0, Q8N1S4, Q8WWW9 | Gene names | ANP32E | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acidic leucine-rich nuclear phosphoprotein 32 family member E (LANP- like protein) (LANP-L). | |||||
|
CAC1C_HUMAN
|
||||||
| θ value | 2.36792 (rank : 65) | NC score | 0.024496 (rank : 122) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q13936, Q13917, Q13918, Q13919, Q13920, Q13921, Q13922, Q13923, Q13924, Q13925, Q13926, Q13927, Q13928, Q13929, Q13930, Q13932, Q13933, Q14743, Q14744, Q15877, Q99025, Q99241, Q99875 | Gene names | CACNA1C, CACH2, CACN2, CACNL1A1, CCHL1A1 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1C (Voltage- gated calcium channel subunit alpha Cav1.2) (Calcium channel, L type, alpha-1 polypeptide, isoform 1, cardiac muscle). | |||||
|
DDX51_HUMAN
|
||||||
| θ value | 2.36792 (rank : 66) | NC score | 0.022544 (rank : 129) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8N8A6, Q5CZ71, Q8IXK5, Q96ED1 | Gene names | DDX51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent RNA helicase DDX51 (EC 3.6.1.-) (DEAD box protein 51). | |||||
|
DMP1_MOUSE
|
||||||
| θ value | 2.36792 (rank : 67) | NC score | 0.085439 (rank : 21) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | O55188 | Gene names | Dmp1, Dmp | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1) (AG1). | |||||
|
FBX34_MOUSE
|
||||||
| θ value | 2.36792 (rank : 68) | NC score | 0.037257 (rank : 97) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q80XI1, Q9D290 | Gene names | Fbxo34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 34. | |||||
|
HTSF1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 69) | NC score | 0.052678 (rank : 72) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O43719, Q59G06, Q99730 | Gene names | HTATSF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HIV Tat-specific factor 1 (Tat-SF1). | |||||
|
MARCS_MOUSE
|
||||||
| θ value | 2.36792 (rank : 70) | NC score | 0.055495 (rank : 68) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P26645 | Gene names | Marcks, Macs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myristoylated alanine-rich C-kinase substrate (MARCKS). | |||||
|
NPAS3_MOUSE
|
||||||
| θ value | 2.36792 (rank : 71) | NC score | 0.026834 (rank : 114) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9QZQ0, Q9EQP4 | Gene names | Npas3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 3 (Neuronal PAS3) (Member of PAS protein 6) (MOP6). | |||||
|
PCNT_MOUSE
|
||||||
| θ value | 2.36792 (rank : 72) | NC score | 0.019032 (rank : 136) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1207 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P48725 | Gene names | Pcnt, Pcnt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pericentrin. | |||||
|
PTMA_MOUSE
|
||||||
| θ value | 2.36792 (rank : 73) | NC score | 0.164462 (rank : 3) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P26350, Q3UQV6 | Gene names | Ptma | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prothymosin alpha [Contains: Thymosin alpha]. | |||||
|
SRCA_HUMAN
|
||||||
| θ value | 2.36792 (rank : 74) | NC score | 0.050208 (rank : 80) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q86TD4 | Gene names | SRL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcalumenin precursor. | |||||
|
ZKSC1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 75) | NC score | 0.009606 (rank : 163) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 949 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | P17029, P52745, Q8TBW5, Q8TEK7 | Gene names | ZKSCAN1, KOX18, ZNF139, ZNF36 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger with KRAB and SCAN domain-containing protein 1 (Zinc finger protein 36) (Zinc finger protein KOX18). | |||||
|
ZN295_HUMAN
|
||||||
| θ value | 2.36792 (rank : 76) | NC score | 0.011168 (rank : 159) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 847 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9ULJ3, Q6P4R0 | Gene names | ZNF295, KIAA1227, ZBTB21 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger protein 295 (Zinc finger and BTB domain-containing protein 21). | |||||
|
B4GN4_MOUSE
|
||||||
| θ value | 3.0926 (rank : 77) | NC score | 0.020865 (rank : 134) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q766D5, Q8CHV2 | Gene names | B4galnt4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyl-beta-glucosaminyl-glycoprotein 4-beta-N- acetylgalactosaminyltransferase 1 (EC 2.4.1.244) (NGalNAc-T1) (Beta- 1,4-N-acetylgalactosaminyltransferase IV) (Beta4GalNAc-T4) (Beta4GalNAcT4). | |||||
|
FOG2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 78) | NC score | 0.033479 (rank : 105) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 714 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8CCH7, Q9Z0F2 | Gene names | Zfpm2, Fog2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein ZFPM2 (Zinc finger protein multitype 2) (Friend of GATA protein 2) (Friend of GATA-2) (FOG-2) (mFOG-2). | |||||
|
GP124_MOUSE
|
||||||
| θ value | 3.0926 (rank : 79) | NC score | 0.011010 (rank : 160) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 423 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q91ZV8 | Gene names | Gpr124, Tem5 | |||
|
Domain Architecture |

|
|||||
| Description | Probable G-protein coupled receptor 124 precursor (Tumor endothelial marker 5). | |||||
|
KIRR3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 80) | NC score | 0.012799 (rank : 153) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 302 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8IZU9, Q6UWJ9, Q6UWL5, Q96JG0 | Gene names | KIRREL3, KIAA1867, NEPH2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kin of IRRE-like protein 3 precursor (Kin of irregular chiasm-like protein 3) (Nephrin-like 2). | |||||
|
MYT1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 81) | NC score | 0.067301 (rank : 41) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 475 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q01538, O94922, Q9UPV2 | Gene names | MYT1, KIAA0835, KIAA1050, MTF1, MYTI, PLPB1 | |||
|
Domain Architecture |

|
|||||
| Description | Myelin transcription factor 1 (MyT1) (MyTI) (Proteolipid protein- binding protein) (PLPB1). | |||||
|
NCKX1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 82) | NC score | 0.055856 (rank : 65) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O60721, O43485, O75184 | Gene names | SLC24A1, KIAA0702, NCKX1 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium/calcium exchanger 1 (Na(+)/K(+)/Ca(2+)-exchange protein 1) (Retinal rod Na-Ca+K exchanger). | |||||
|
PTMA_HUMAN
|
||||||
| θ value | 3.0926 (rank : 83) | NC score | 0.161061 (rank : 4) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P06454, Q15249, Q15592 | Gene names | PTMA, TMSA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prothymosin alpha [Contains: Thymosin alpha-1]. | |||||
|
PTPRG_HUMAN
|
||||||
| θ value | 3.0926 (rank : 84) | NC score | 0.014411 (rank : 147) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P23470, Q15623 | Gene names | PTPRG | |||
|
Domain Architecture |

|
|||||
| Description | Receptor-type tyrosine-protein phosphatase gamma precursor (EC 3.1.3.48) (Protein-tyrosine phosphatase gamma) (R-PTP-gamma). | |||||
|
TXLNA_HUMAN
|
||||||
| θ value | 3.0926 (rank : 85) | NC score | 0.024841 (rank : 119) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 982 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P40222, Q66K62, Q86T54, Q86T85, Q86T86, Q86Y86, Q86YW3, Q8N2Y3 | Gene names | TXLNA, TXLN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Alpha-taxilin. | |||||
|
UBF1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 86) | NC score | 0.052571 (rank : 73) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P25976 | Gene names | Ubtf, Tcfubf, Ubf-1, Ubf1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar transcription factor 1 (Upstream-binding factor 1) (UBF-1). | |||||
|
ZFHX2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 87) | NC score | 0.015279 (rank : 144) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1178 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9C0A1 | Gene names | ZFHX2, KIAA1762 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger homeobox protein 2. | |||||
|
ABTAP_MOUSE
|
||||||
| θ value | 4.03905 (rank : 88) | NC score | 0.043362 (rank : 89) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 381 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q3V1V3 | Gene names | Abtap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ABT1-associated protein. | |||||
|
CSTN1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 89) | NC score | 0.035873 (rank : 99) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9EPL2 | Gene names | Clstn1, Cs1, Cstn1 | |||
|
Domain Architecture |
|
|||||
| Description | Calsyntenin-1 precursor. | |||||
|
LRRF1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 90) | NC score | 0.054146 (rank : 69) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 625 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q32MZ4, O75766, O75799, Q32MZ5, Q53T49, Q6PKG2 | Gene names | LRRFIP1, GCF2, TRIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat flightless-interacting protein 1 (LRR FLII- interacting protein 1) (TAR RNA-interacting protein) (GC-binding factor 2). | |||||
|
MLL2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 91) | NC score | 0.029203 (rank : 111) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
MYT1L_HUMAN
|
||||||
| θ value | 4.03905 (rank : 92) | NC score | 0.048960 (rank : 82) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 471 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9UL68, Q6IQ17, Q9UPP6 | Gene names | MYT1L, KIAA1106 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myelin transcription factor 1-like protein (MyT1L protein) (MyT1-L). | |||||
|
NPAS3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 93) | NC score | 0.023881 (rank : 125) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8IXF0, Q86US6, Q86US7, Q8IXF2, Q9BY81, Q9H323, Q9Y4L8 | Gene names | NPAS3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 3 (Neuronal PAS3) (Member of PAS protein 6) (MOP6). | |||||
|
NSD1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 94) | NC score | 0.025669 (rank : 117) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 472 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O88491, Q8C480, Q9CT70 | Gene names | Nsd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein). | |||||
|
PELP1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 95) | NC score | 0.043901 (rank : 88) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9DBD5, Q5F2E2, Q6PEM0, Q91YM9 | Gene names | Pelp1, Mnar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-, glutamic acid- and leucine-rich protein 1 (Modulator of nongenomic activity of estrogen receptor). | |||||
|
SALL2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 96) | NC score | 0.014138 (rank : 149) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 836 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9QX96 | Gene names | Sall2, Sal2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sal-like protein 2 (Spalt-like protein 2) (MSal-2). | |||||
|
SMRC2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 97) | NC score | 0.059955 (rank : 54) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 499 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q8TAQ2, Q92923, Q96E12, Q96GY4 | Gene names | SMARCC2, BAF170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 2 (SWI/SNF complex 170 kDa subunit) (BRG1-associated factor 170). | |||||
|
SPEG_MOUSE
|
||||||
| θ value | 4.03905 (rank : 98) | NC score | 0.010816 (rank : 161) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1959 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q62407, Q3TPH8, Q6P5V1, Q80TF7, Q80ZN0, Q8BZF4, Q9EQJ5 | Gene names | Speg, Apeg1, Kiaa1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle-specific serine/threonine protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
TM16H_HUMAN
|
||||||
| θ value | 4.03905 (rank : 99) | NC score | 0.033726 (rank : 103) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9HCE9 | Gene names | TMEM16H, KIAA1623 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein 16H. | |||||
|
TXND2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 100) | NC score | 0.062925 (rank : 47) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 793 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q86VQ3, Q8N7U4, Q96RX3, Q9H0L8 | Gene names | TXNDC2, SPTRX, SPTRX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1). | |||||
|
UBP4_HUMAN
|
||||||
| θ value | 4.03905 (rank : 101) | NC score | 0.015680 (rank : 143) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q13107, O43452, O43453 | Gene names | USP4, UNP, UNPH | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 4 (EC 3.1.2.15) (Ubiquitin thioesterase 4) (Ubiquitin-specific-processing protease 4) (Deubiquitinating enzyme 4) (Ubiquitous nuclear protein homolog). | |||||
|
ZN174_HUMAN
|
||||||
| θ value | 4.03905 (rank : 102) | NC score | 0.006711 (rank : 167) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q15697, Q9BQ34 | Gene names | ZNF174, ZSCAN8 | |||
|
Domain Architecture |
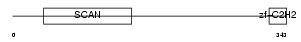
|
|||||
| Description | Zinc finger protein 174 (AW-1) (Zinc finger and SCAN domain-containing protein 8). | |||||
|
AP3B1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 103) | NC score | 0.040931 (rank : 94) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O00203, O00580, Q7Z393, Q9HD66 | Gene names | AP3B1, ADTB3A | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit beta-1 (Adapter-related protein complex 3 beta-1 subunit) (Beta3A-adaptin) (Adaptor protein complex AP-3 beta-1 subunit) (Clathrin assembly protein complex 3 beta-1 large chain). | |||||
|
ARI3A_HUMAN
|
||||||
| θ value | 5.27518 (rank : 104) | NC score | 0.046614 (rank : 85) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q99856, Q8IZA7 | Gene names | ARID3A, DRIL1, DRX, E2FBP1 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 3A (ARID domain- containing protein 3A) (B-cell regulator of IgH transcription) (Bright) (E2F-binding protein 1). | |||||
|
LEO1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 105) | NC score | 0.114528 (rank : 10) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 93 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
MYST3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 106) | NC score | 0.039338 (rank : 96) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q92794 | Gene names | MYST3, MOZ, RUNXBP2, ZNF220 | |||
|
Domain Architecture |

|
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Runt-related transcription factor-binding protein 2) (Monocytic leukemia zinc finger protein) (Zinc finger protein 220). | |||||
|
RGP1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 107) | NC score | 0.048197 (rank : 83) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P46060 | Gene names | RANGAP1 | |||
|
Domain Architecture |
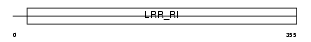
|
|||||
| Description | Ran GTPase-activating protein 1. | |||||
|
SSRP1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 108) | NC score | 0.023556 (rank : 126) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q08945, Q5BJG8 | Gene names | SSRP1, FACT80 | |||
|
Domain Architecture |

|
|||||
| Description | FACT complex subunit SSRP1 (Facilitates chromatin transcription complex subunit SSRP1) (FACT 80 kDa subunit) (FACTp80) (Chromatin- specific transcription elongation factor 80 kDa subunit) (Structure- specific recognition protein 1) (hSSRP1) (Recombination signal sequence recognition protein 1) (T160). | |||||
|
AP3B2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 109) | NC score | 0.041610 (rank : 93) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9JME5, Q6QR53, Q8R1E5 | Gene names | Ap3b2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AP-3 complex subunit beta-2 (Adapter-related protein complex 3 beta-2 subunit) (Beta3B-adaptin) (Adaptor protein complex AP-3 beta-2 subunit) (Clathrin assembly protein complex 3 beta-2 large chain). | |||||
|
BFSP1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 110) | NC score | 0.014786 (rank : 146) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 508 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q12934, O43595, O76034, Q9HBX4 | Gene names | BFSP1 | |||
|
Domain Architecture |
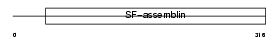
|
|||||
| Description | Filensin (Beaded filament structural protein 1) (Lens fiber cell beaded-filament structural protein CP 115) (CP115) (Lens intermediate filament-like heavy) (LIFL-H). | |||||
|
CD2L2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 111) | NC score | 0.013898 (rank : 150) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1439 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9UQ88, O95227, O95228, O96012, Q12821, Q12853, Q12854, Q5QPR0, Q5QPR1, Q5QPR2, Q9UBC4, Q9UBI3, Q9UEI1, Q9UEI2, Q9UP53, Q9UP54, Q9UP55, Q9UP56, Q9UQ86, Q9UQ87, Q9UQ89 | Gene names | CDC2L2, PITSLREB | |||
|
Domain Architecture |

|
|||||
| Description | PITSLRE serine/threonine-protein kinase CDC2L2 (EC 2.7.11.22) (Galactosyltransferase-associated protein kinase p58/GTA) (Cell division cycle 2-like protein kinase 2) (CDK11). | |||||
|
CENPB_MOUSE
|
||||||
| θ value | 6.88961 (rank : 112) | NC score | 0.060237 (rank : 53) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P27790 | Gene names | Cenpb, Cenp-b | |||
|
Domain Architecture |

|
|||||
| Description | Major centromere autoantigen B (Centromere protein B) (CENP-B). | |||||
|
CLIC6_HUMAN
|
||||||
| θ value | 6.88961 (rank : 113) | NC score | 0.029674 (rank : 109) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q96NY7, Q8IX31 | Gene names | CLIC6, CLIC1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chloride intracellular channel 6. | |||||
|
DCDC2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 114) | NC score | 0.069303 (rank : 37) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q5DU00, Q5SZU0, Q80Y99 | Gene names | Dcdc2, Kiaa1154 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Doublecortin domain-containing protein 2. | |||||
|
DPOD3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 115) | NC score | 0.027078 (rank : 112) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q15054 | Gene names | POLD3, KIAA0039 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA polymerase subunit delta 3 (DNA polymerase subunit delta p66). | |||||
|
K0256_HUMAN
|
||||||
| θ value | 6.88961 (rank : 116) | NC score | 0.016927 (rank : 139) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q93073, Q8N767 | Gene names | KIAA0256 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0256. | |||||
|
MDN1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 117) | NC score | 0.072293 (rank : 35) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 662 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q9NU22, O15019 | Gene names | MDN1, KIAA0301 | |||
|
Domain Architecture |

|
|||||
| Description | Midasin (MIDAS-containing protein). | |||||
|
MINT_MOUSE
|
||||||
| θ value | 6.88961 (rank : 118) | NC score | 0.033082 (rank : 106) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
NFM_HUMAN
|
||||||
| θ value | 6.88961 (rank : 119) | NC score | 0.021628 (rank : 132) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1328 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P07197 | Gene names | NEF3, NEFM, NFM | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet M protein (160 kDa neurofilament protein) (Neurofilament medium polypeptide) (NF-M) (Neurofilament 3). | |||||
|
PEG3_MOUSE
|
||||||
| θ value | 6.88961 (rank : 120) | NC score | 0.021369 (rank : 133) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1101 | Shared Neighborhood Hits | 84 | |
| SwissProt Accessions | Q3URU2, O54978, Q3TQ69, Q5EBP7, Q61138, Q6GQS0, Q80U47, Q8R5N0, Q9QX53 | Gene names | Peg3, Kiaa0287, Pw1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Paternally expressed gene 3 protein (ASF-1). | |||||
|
PTN13_MOUSE
|
||||||
| θ value | 6.88961 (rank : 121) | NC score | 0.007957 (rank : 166) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 426 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q64512, Q61494, Q62135, Q64499 | Gene names | Ptpn13, Ptp14 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 13 (EC 3.1.3.48) (Protein tyrosine phosphatase PTP-BL) (Protein-tyrosine phosphatase RIP) (protein tyrosine phosphatase DPZPTP) (PTP36). | |||||
|
REM1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 122) | NC score | 0.003339 (rank : 168) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O75628, Q9NP57 | Gene names | REM1, GES, REM | |||
|
Domain Architecture |

|
|||||
| Description | GTP-binding protein REM 1 (Rad and Gem-like GTP-binding protein 1) (GTPase-regulating endothelial cell sprouting). | |||||
|
SMCA2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 123) | NC score | 0.013396 (rank : 152) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 503 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P51531 | Gene names | SMARCA2, BRM, SNF2A, SNF2L2 | |||
|
Domain Architecture |

|
|||||
| Description | Probable global transcription activator SNF2L2 (EC 3.6.1.-) (ATP- dependent helicase SMARCA2) (SNF2-alpha) (SWI/SNF-related matrix- associated actin-dependent regulator of chromatin subfamily A member 2) (hBRM). | |||||
|
SMRC1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 124) | NC score | 0.043080 (rank : 90) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 459 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | P97496, Q7TS80, Q7TT29 | Gene names | Smarcc1, Baf155, Srg3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 1 (SWI/SNF complex 155 kDa subunit) (BRG1-associated factor 155) (SWI3-related protein). | |||||
|
SMRCD_MOUSE
|
||||||
| θ value | 6.88961 (rank : 125) | NC score | 0.009202 (rank : 164) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q04692 | Gene names | Smarcad1, Etl1, Kiaa1122 | |||
|
Domain Architecture |

|
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A containing DEAD/H box 1 (EC 3.6.1.-) (ATP- dependent helicase SMARCAD1) (Enhancer trap locus homolog 1) (Etl-1). | |||||
|
SYTL1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 126) | NC score | 0.041620 (rank : 92) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 314 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q99N80, Q99J26 | Gene names | Sytl1, Slp1 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-like protein 1 (Exophilin-7). | |||||
|
TGON2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 127) | NC score | 0.077400 (rank : 27) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1037 | Shared Neighborhood Hits | 80 | |
| SwissProt Accessions | O43493, O15282, O43492, O43499, O43500, O43501, Q92760 | Gene names | TGOLN2, TGN46, TGN51 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (Trans-Golgi network protein TGN51) (TGN46) (TGN48) (TGN38 homolog). | |||||
|
TRI56_HUMAN
|
||||||
| θ value | 6.88961 (rank : 128) | NC score | 0.016822 (rank : 140) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 339 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9BRZ2, Q86VT6, Q8N2H8, Q8NAC0, Q9H031 | Gene names | TRIM56, RNF109 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 56 (RING finger protein 109). | |||||
|
ZBTB5_HUMAN
|
||||||
| θ value | 6.88961 (rank : 129) | NC score | 0.012048 (rank : 154) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 811 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O15062 | Gene names | ZBTB5, KIAA0354 | |||
|
Domain Architecture |
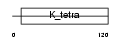
|
|||||
| Description | Zinc finger and BTB domain-containing protein 5. | |||||
|
ZBTB5_MOUSE
|
||||||
| θ value | 6.88961 (rank : 130) | NC score | 0.010797 (rank : 162) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 801 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q7TQG0, Q3TVR7, Q6GV09, Q8BIB5 | Gene names | Zbtb5, Kiaa0354 | |||
|
Domain Architecture |
|
|||||
| Description | Zinc finger and BTB domain-containing protein 5 (Transcription factor ZNF-POZ). | |||||
|
ZEP2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 131) | NC score | 0.015132 (rank : 145) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 937 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P31629, Q02646, Q5THT5, Q9NS05 | Gene names | HIVEP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Human immunodeficiency virus type I enhancer-binding protein 2 (HIV- EP2) (MHC-binding protein 2) (MBP-2). | |||||
|
ARS2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 132) | NC score | 0.046519 (rank : 86) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q99MR6, Q99MR4, Q99MR5, Q99MR7 | Gene names | Ars2, Asr2 | |||
|
Domain Architecture |

|
|||||
| Description | Arsenite-resistance protein 2. | |||||
|
BRD2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 133) | NC score | 0.014182 (rank : 148) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P25440, O00699, O00700, Q15310, Q5STC9, Q969U4 | Gene names | BRD2, KIAA9001, RING3 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 2 (Protein RING3) (O27.1.1). | |||||
|
DAXX_MOUSE
|
||||||
| θ value | 8.99809 (rank : 134) | NC score | 0.036023 (rank : 98) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O35613, Q9QWT8, Q9QWV3 | Gene names | Daxx | |||
|
Domain Architecture |

|
|||||
| Description | Death domain-associated protein 6 (Daxx). | |||||
|
DDIT3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 135) | NC score | 0.022395 (rank : 130) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P35638 | Gene names | DDIT3, CHOP, GADD153 | |||
|
Domain Architecture |
|
|||||
| Description | DNA damage-inducible transcript 3 (DDIT-3) (Growth arrest and DNA- damage-inducible protein GADD153) (C/EBP-homologous protein) (CHOP). | |||||
|
DMP1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 136) | NC score | 0.072689 (rank : 34) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q13316, O43265 | Gene names | DMP1 | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1). | |||||
|
FIP1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 137) | NC score | 0.017831 (rank : 137) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q6UN15, Q499Y4, Q49AU3, Q7Z608, Q8WVN3, Q96F80, Q9H077 | Gene names | FIP1L1, FIP1, RHE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pre-mRNA 3'-end-processing factor FIP1 (FIP1-like 1) (Factor interacting with PAP) (hFip1) (Rearranged in hypereosinophilia). | |||||
|
G3BP_HUMAN
|
||||||
| θ value | 8.99809 (rank : 138) | NC score | 0.011818 (rank : 155) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q13283 | Gene names | G3BP, G3BP1 | |||
|
Domain Architecture |

|
|||||
| Description | Ras-GTPase-activating protein-binding protein 1 (EC 3.6.1.-) (ATP- dependent DNA helicase VIII) (GAP SH3-domain-binding protein 1) (G3BP- 1) (HDH-VIII). | |||||
|
HLX1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 139) | NC score | 0.013531 (rank : 151) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 362 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q61670 | Gene names | Hlx1, Hlx | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein HLX1. | |||||
|
MADCA_HUMAN
|
||||||
| θ value | 8.99809 (rank : 140) | NC score | 0.026804 (rank : 115) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q13477, O60222, O75867 | Gene names | MADCAM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucosal addressin cell adhesion molecule 1 precursor (MAdCAM-1) (hMAdCAM-1). | |||||
|
MAP2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 141) | NC score | 0.026055 (rank : 116) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 348 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P11137, Q99975, Q99976 | Gene names | MAP2 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 2 (MAP 2) (MAP-2). | |||||
|
NFH_HUMAN
|
||||||
| θ value | 8.99809 (rank : 142) | NC score | 0.029432 (rank : 110) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1242 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P12036, Q9UJS7, Q9UQ14 | Gene names | NEFH, KIAA0845, NFH | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
SAS10_HUMAN
|
||||||
| θ value | 8.99809 (rank : 143) | NC score | 0.022618 (rank : 128) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NQZ2, Q6FI82 | Gene names | CRLZ1, SAS10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Something about silencing protein 10 (Disrupter of silencing SAS10) (Charged amino acid-rich leucine zipper 1). | |||||
|
SPT5H_HUMAN
|
||||||
| θ value | 8.99809 (rank : 144) | NC score | 0.025459 (rank : 118) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 436 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O00267, O43279, Q59G52, Q99639 | Gene names | SUPT5H, SPT5, SPT5H | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor SPT5 (hSPT5) (DRB sensitivity-inducing factor large subunit) (DSIF large subunit) (DSIF p160) (Tat- cotransactivator 1 protein) (Tat-CT1 protein). | |||||
|
TAF1L_HUMAN
|
||||||
| θ value | 8.99809 (rank : 145) | NC score | 0.015773 (rank : 142) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8IZX4 | Gene names | TAF1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID 210 kDa subunit (TBP-associated factor 210 kDa) (TAF(II)210) (TBP-associated factor 1-like). | |||||
|
TBC16_HUMAN
|
||||||
| θ value | 8.99809 (rank : 146) | NC score | 0.011743 (rank : 156) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8TBP0 | Gene names | TBC1D16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TBC1 domain family member 16. | |||||
|
UBIP1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 147) | NC score | 0.008900 (rank : 165) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NZI7, Q68CT0, Q86Y57, Q9H8V0, Q9UD76, Q9UD78 | Gene names | UBP1, LBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Upstream-binding protein 1 (LBP-1). | |||||
|
UBP8_MOUSE
|
||||||
| θ value | 8.99809 (rank : 148) | NC score | 0.019993 (rank : 135) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 362 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q80U87, Q80YP2, Q8R0D3, Q9EQU1, Q9WVP5 | Gene names | Usp8, Kiaa0055, Ubpy | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 8 (EC 3.1.2.15) (Ubiquitin thioesterase 8) (Ubiquitin-specific-processing protease 8) (Deubiquitinating enzyme 8) (mUBPy). | |||||
|
BASP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 149) | NC score | 0.051767 (rank : 76) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P80723, O43596, Q5U0S0 | Gene names | BASP1, NAP22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
CC47_MOUSE
|
||||||
| θ value | θ > 10 (rank : 150) | NC score | 0.051336 (rank : 78) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9D024, Q3TSB5, Q3V1S7, Q8C5D9, Q920S6 | Gene names | Ccdc47, Asp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 47 precursor (Adipocyte-specific protein 4). | |||||
|
ERIC1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 151) | NC score | 0.111014 (rank : 11) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q86X53, Q9P063 | Gene names | ERICH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate-rich protein 1. | |||||
|
HSP3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 152) | NC score | 0.053399 (rank : 70) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q62100 | Gene names | Prm3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm protamine P3. | |||||
|
IWS1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 153) | NC score | 0.104191 (rank : 15) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 104 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
LEO1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 154) | NC score | 0.107603 (rank : 14) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 95 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
MATR3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 155) | NC score | 0.061209 (rank : 51) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q8K310 | Gene names | Matr3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Matrin-3. | |||||
|
NASP_HUMAN
|
||||||
| θ value | θ > 10 (rank : 156) | NC score | 0.082982 (rank : 22) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 81 | |
| SwissProt Accessions | P49321, Q96A69, Q9BTW2 | Gene names | NASP | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear autoantigenic sperm protein (NASP). | |||||
|
NIPBL_HUMAN
|
||||||
| θ value | θ > 10 (rank : 157) | NC score | 0.057179 (rank : 60) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 86 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
NSBP1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 158) | NC score | 0.066785 (rank : 42) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P82970 | Gene names | NSBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1. | |||||
|
OGFR_MOUSE
|
||||||
| θ value | θ > 10 (rank : 159) | NC score | 0.058077 (rank : 58) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 584 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q99PG2, Q99L33 | Gene names | Ogfr | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor). | |||||
|
SEC63_HUMAN
|
||||||
| θ value | θ > 10 (rank : 160) | NC score | 0.055975 (rank : 64) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 586 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q9UGP8, O95380, Q86VS9, Q8IWL0, Q9NTE0 | Gene names | SEC63, SEC63L | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
SEC63_MOUSE
|
||||||
| θ value | θ > 10 (rank : 161) | NC score | 0.056338 (rank : 62) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 601 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q8VHE0, Q8VEB9 | Gene names | Sec63, Sec63l | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
SPRL1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 162) | NC score | 0.067799 (rank : 40) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | P70663, P97810, Q99L82 | Gene names | Sparcl1, Ecm2, Sc1 | |||
|
Domain Architecture |

|
|||||
| Description | SPARC-like protein 1 precursor (Matrix glycoprotein Sc1) (Extracellular matrix protein 2). | |||||
|
SRCH_HUMAN
|
||||||
| θ value | θ > 10 (rank : 163) | NC score | 0.058599 (rank : 57) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P23327, Q504Y6 | Gene names | HRC, HCP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcoplasmic reticulum histidine-rich calcium-binding protein precursor. | |||||
|
TCAL3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 164) | NC score | 0.089710 (rank : 19) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q969E4 | Gene names | TCEAL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 3 (TCEA-like protein 3) (Transcription elongation factor S-II protein-like 3). | |||||
|
TCAL6_HUMAN
|
||||||
| θ value | θ > 10 (rank : 165) | NC score | 0.052511 (rank : 74) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q6IPX3, Q5H9J8 | Gene names | TCEAL6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 6 (TCEA-like protein 6) (Transcription elongation factor S-II protein-like 6). | |||||
|
TXND2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 166) | NC score | 0.063781 (rank : 46) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
VPS72_HUMAN
|
||||||
| θ value | θ > 10 (rank : 167) | NC score | 0.052281 (rank : 75) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q15906, Q53GJ2, Q5U0R4 | Gene names | VPS72, TCFL1, YL1 | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar protein sorting-associated protein 72 homolog (Transcription factor-like 1) (Protein YL-1). | |||||
|
VPS72_MOUSE
|
||||||
| θ value | θ > 10 (rank : 168) | NC score | 0.055746 (rank : 66) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q62481, Q3U2N4, Q810A9, Q99K81 | Gene names | Vps72, Tcfl1, Yl1 | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar protein sorting-associated protein 72 homolog (Transcription factor-like 1) (Protein YL-1). | |||||
|
BSCL2_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 148 | Shared Neighborhood Hits | 148 | |
| SwissProt Accessions | Q96G97, Q96SV1, Q9BSQ0 | Gene names | BSCL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Seipin (Bernardinelli-Seip congenital lipodystrophy type 2 protein). | |||||
|
BSCL2_MOUSE
|
||||||
| NC score | 0.942970 (rank : 2) | θ value | 5.9214e-177 (rank : 2) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9Z2E9, Q810B0, Q9JJC2, Q9JMF1 | Gene names | Bscl2, Gng3lg | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Seipin (Bernardinelli-Seip congenital lipodystrophy type 2 protein homolog). | |||||
|
PTMA_MOUSE
|
||||||
| NC score | 0.164462 (rank : 3) | θ value | 2.36792 (rank : 73) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P26350, Q3UQV6 | Gene names | Ptma | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prothymosin alpha [Contains: Thymosin alpha]. | |||||
|
PTMA_HUMAN
|
||||||
| NC score | 0.161061 (rank : 4) | θ value | 3.0926 (rank : 83) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P06454, Q15249, Q15592 | Gene names | PTMA, TMSA | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prothymosin alpha [Contains: Thymosin alpha-1]. | |||||
|
NSBP1_MOUSE
|
||||||
| NC score | 0.158327 (rank : 5) | θ value | 0.279714 (rank : 17) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 96 | |
| SwissProt Accessions | Q9JL35, O88832, Q8VC71, Q9CUW1 | Gene names | Nsbp1, Garp45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1 (Nucleosome-binding protein 45) (NBP-45) (GARP45 protein). | |||||
|
T2FA_HUMAN
|
||||||
| NC score | 0.136214 (rank : 6) | θ value | 0.00665767 (rank : 5) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P35269, Q9BWN0 | Gene names | GTF2F1, RAP74 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription initiation factor IIF alpha subunit (EC 2.7.11.1) (TFIIF-alpha) (Transcription initiation factor RAP74) (General transcription factor IIF polypeptide 1 74 kDa subunit protein). | |||||
|
TCAL5_HUMAN
|
||||||
| NC score | 0.134984 (rank : 7) | θ value | 0.47712 (rank : 27) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 806 | Shared Neighborhood Hits | 96 | |
| SwissProt Accessions | Q5H9L2 | Gene names | TCEAL5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 5 (TCEA-like protein 5) (Transcription elongation factor S-II protein-like 5). | |||||
|
IWS1_MOUSE
|
||||||
| NC score | 0.129535 (rank : 8) | θ value | 0.365318 (rank : 20) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 109 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
AKA12_HUMAN
|
||||||
| NC score | 0.126972 (rank : 9) | θ value | 0.21417 (rank : 13) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 825 | Shared Neighborhood Hits | 82 | |
| SwissProt Accessions | Q02952, O00310, O00498, Q99970 | Gene names | AKAP12, AKAP250 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | A-kinase anchor protein 12 (A-kinase anchor protein 250 kDa) (AKAP 250) (Myasthenia gravis autoantigen gravin). | |||||
|
LEO1_HUMAN
|
||||||
| NC score | 0.114528 (rank : 10) | θ value | 5.27518 (rank : 105) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 958 | Shared Neighborhood Hits | 93 | |
| SwissProt Accessions | Q8WVC0, Q96N99 | Gene names | LEO1, RDL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1 (Replicative senescence down- regulated leo1-like protein). | |||||
|
ERIC1_HUMAN
|
||||||
| NC score | 0.111014 (rank : 11) | θ value | θ > 10 (rank : 151) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 364 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q86X53, Q9P063 | Gene names | ERICH1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glutamate-rich protein 1. | |||||
|
EDD1_HUMAN
|
||||||
| NC score | 0.110273 (rank : 12) | θ value | 0.0431538 (rank : 8) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 170 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O95071, O94970, Q9NPL3 | Gene names | EDD1, EDD, HYD, KIAA0896 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin-protein ligase EDD1 (EC 6.3.2.-) (Hyperplastic discs protein homolog) (hHYD) (Progestin-induced protein). | |||||
|
EDD1_MOUSE
|
||||||
| NC score | 0.109119 (rank : 13) | θ value | 0.0563607 (rank : 9) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q80TP3, Q698K9, Q6PEQ8, Q6PFQ9, Q80VL4, Q810V6, Q9CXE9 | Gene names | Edd1, Edd, Kiaa0896 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ubiquitin-protein ligase EDD1 (EC 6.3.2.-) (Hyperplastic discs protein homolog). | |||||
|
LEO1_MOUSE
|
||||||
| NC score | 0.107603 (rank : 14) | θ value | θ > 10 (rank : 154) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 946 | Shared Neighborhood Hits | 95 | |
| SwissProt Accessions | Q5XJE5, Q640R1 | Gene names | Leo1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA polymerase-associated protein LEO1. | |||||
|
IWS1_HUMAN
|
||||||
| NC score | 0.104191 (rank : 15) | θ value | θ > 10 (rank : 153) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1241 | Shared Neighborhood Hits | 104 | |
| SwissProt Accessions | Q96ST2, Q6P157, Q8N3E8, Q96MI7, Q9NV97, Q9NWH8 | Gene names | IWS1, IWS1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
HUWE1_MOUSE
|
||||||
| NC score | 0.103027 (rank : 16) | θ value | 0.00665767 (rank : 4) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q7TMY8, Q4G2Z1, Q5BMM7, Q6NS61, Q8BNJ7, Q8CFH2, Q8VD14, Q921M5, Q9R0P2 | Gene names | Huwe1, Kiaa0312, Ureb1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT, UBA and WWE domain-containing protein 1 (EC 6.3.2.-) (E3 ubiquitin protein ligase URE-B1) (E3Histone). | |||||
|
HUWE1_HUMAN
|
||||||
| NC score | 0.101893 (rank : 17) | θ value | 0.00665767 (rank : 3) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 443 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q7Z6Z7, O15029, Q4G2Z2, Q5H961, Q6P4D0, Q8NG67, Q9BUI0, Q9HCJ4, Q9NSL6, Q9P0A9 | Gene names | HUWE1, KIAA0312, KIAA1578, UREB1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HECT, UBA and WWE domain-containing protein 1 (EC 6.3.2.-) (E3 ubiquitin protein ligase URE-B1) (Mcl-1 ubiquitin ligase E3) (Mule) (ARF-binding protein 1) (ARF-BP1). | |||||
|
PTMS_HUMAN
|
||||||
| NC score | 0.097394 (rank : 18) | θ value | 1.81305 (rank : 57) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 111 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P20962 | Gene names | PTMS | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Parathymosin. | |||||
|
TCAL3_HUMAN
|
||||||
| NC score | 0.089710 (rank : 19) | θ value | θ > 10 (rank : 164) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q969E4 | Gene names | TCEAL3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 3 (TCEA-like protein 3) (Transcription elongation factor S-II protein-like 3). | |||||
|
SH3BG_MOUSE
|
||||||
| NC score | 0.087104 (rank : 20) | θ value | 0.279714 (rank : 19) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9WUZ7 | Gene names | Sh3bgr | |||
|
Domain Architecture |

|
|||||
| Description | SH3 domain-binding glutamic acid-rich protein (SH3BGR protein). | |||||
|
DMP1_MOUSE
|
||||||
| NC score | 0.085439 (rank : 21) | θ value | 2.36792 (rank : 67) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 61 | |
| SwissProt Accessions | O55188 | Gene names | Dmp1, Dmp | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1) (AG1). | |||||
|
NASP_HUMAN
|
||||||
| NC score | 0.082982 (rank : 22) | θ value | θ > 10 (rank : 156) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 81 | |
| SwissProt Accessions | P49321, Q96A69, Q9BTW2 | Gene names | NASP | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear autoantigenic sperm protein (NASP). | |||||
|
CENPB_HUMAN
|
||||||
| NC score | 0.081104 (rank : 23) | θ value | 1.06291 (rank : 41) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 133 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P07199 | Gene names | CENPB | |||
|
Domain Architecture |

|
|||||
| Description | Major centromere autoantigen B (Centromere protein B) (CENP-B). | |||||
|
NOL3_MOUSE
|
||||||
| NC score | 0.080638 (rank : 24) | θ value | 0.62314 (rank : 32) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9D1X0 | Gene names | Nol3 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar protein 3. | |||||
|
TRI44_HUMAN
|
||||||
| NC score | 0.079147 (rank : 25) | θ value | 0.365318 (rank : 22) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 393 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q96DX7, Q96QY2, Q9UGK0 | Gene names | TRIM44, DIPB | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 44 (Protein DIPB). | |||||
|
RP1L1_HUMAN
|
||||||
| NC score | 0.077989 (rank : 26) | θ value | 0.125558 (rank : 11) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1072 | Shared Neighborhood Hits | 60 | |
| SwissProt Accessions | Q8IWN7, Q86SQ1, Q8IWN8, Q8IWN9, Q8IWP0, Q8IWP1, Q8IWP2 | Gene names | RP1L1 | |||
|
Domain Architecture |

|
|||||
| Description | Retinitis pigmentosa 1-like 1 protein. | |||||
|
TGON2_HUMAN
|
||||||
| NC score | 0.077400 (rank : 27) | θ value | 6.88961 (rank : 127) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1037 | Shared Neighborhood Hits | 80 | |
| SwissProt Accessions | O43493, O15282, O43492, O43499, O43500, O43501, Q92760 | Gene names | TGOLN2, TGN46, TGN51 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (Trans-Golgi network protein TGN51) (TGN46) (TGN48) (TGN38 homolog). | |||||
|
BASP_MOUSE
|
||||||
| NC score | 0.077272 (rank : 28) | θ value | 0.62314 (rank : 28) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q91XV3 | Gene names | Basp1, Nap22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
YTDC1_HUMAN
|
||||||
| NC score | 0.077160 (rank : 29) | θ value | 1.81305 (rank : 61) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 458 | Shared Neighborhood Hits | 44 | |
| SwissProt Accessions | Q96MU7, Q7Z622, Q8TF35 | Gene names | YTHDC1, KIAA1966, YT521 | |||
|
Domain Architecture |

|
|||||
| Description | YTH domain-containing protein 1 (Putative splicing factor YT521). | |||||
|
TCAL3_MOUSE
|
||||||
| NC score | 0.076799 (rank : 30) | θ value | 0.365318 (rank : 21) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 132 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q8R0A5, Q9CYK5, Q9D1S4 | Gene names | Tceal3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 3 (TCEA-like protein 3) (Transcription elongation factor S-II protein-like 3). | |||||
|
LIPS_HUMAN
|
||||||
| NC score | 0.075517 (rank : 31) | θ value | 0.0563607 (rank : 10) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 314 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q05469, Q3LRT2, Q6NSL7 | Gene names | LIPE | |||
|
Domain Architecture |

|
|||||
| Description | Hormone-sensitive lipase (EC 3.1.1.79) (HSL). | |||||
|
MYT1_MOUSE
|
||||||
| NC score | 0.073863 (rank : 32) | θ value | 0.47712 (rank : 26) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q8CFC2, O08995, Q8CFH1 | Gene names | Myt1, Kiaa0835, Nzf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myelin transcription factor 1 (MyT1) (Neural zinc finger factor 2) (NZF-2). | |||||
|
MATR3_HUMAN
|
||||||
| NC score | 0.072733 (rank : 33) | θ value | 1.06291 (rank : 44) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 325 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | P43243, Q9UHW0, Q9UQ27 | Gene names | MATR3, KIAA0723 | |||
|
Domain Architecture |

|
|||||
| Description | Matrin-3. | |||||
|
DMP1_HUMAN
|
||||||
| NC score | 0.072689 (rank : 34) | θ value | 8.99809 (rank : 136) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 491 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q13316, O43265 | Gene names | DMP1 | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1). | |||||
|
MDN1_HUMAN
|
||||||
| NC score | 0.072293 (rank : 35) | θ value | 6.88961 (rank : 117) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 662 | Shared Neighborhood Hits | 73 | |
| SwissProt Accessions | Q9NU22, O15019 | Gene names | MDN1, KIAA0301 | |||
|
Domain Architecture |

|
|||||
| Description | Midasin (MIDAS-containing protein). | |||||
|
TMAP1_HUMAN
|
||||||
| NC score | 0.072072 (rank : 36) | θ value | 1.38821 (rank : 51) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 140 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P84157 | Gene names | TMAP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane anchor protein 1. | |||||
|
DCDC2_MOUSE
|
||||||
| NC score | 0.069303 (rank : 37) | θ value | 6.88961 (rank : 114) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 63 | |
| SwissProt Accessions | Q5DU00, Q5SZU0, Q80Y99 | Gene names | Dcdc2, Kiaa1154 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Doublecortin domain-containing protein 2. | |||||
|
TXD13_HUMAN
|
||||||
| NC score | 0.069244 (rank : 38) | θ value | 0.62314 (rank : 36) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9H1E5, Q8N4P7, Q8NCC1, Q9UJA1, Q9ULQ8 | Gene names | TXNDC13, KIAA1162 | |||
|
Domain Architecture |

|
|||||
| Description | Thioredoxin domain-containing protein 13 precursor. | |||||
|
CASC3_HUMAN
|
||||||
| NC score | 0.068221 (rank : 39) | θ value | 0.813845 (rank : 37) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 204 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O15234 | Gene names | CASC3, MLN51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein CASC3 (Cancer susceptibility candidate gene 3 protein) (Metastatic lymph node protein 51) (Protein MLN 51) (Protein barentsz) (Btz). | |||||
|
SPRL1_MOUSE
|
||||||
| NC score | 0.067799 (rank : 40) | θ value | θ > 10 (rank : 162) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 403 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | P70663, P97810, Q99L82 | Gene names | Sparcl1, Ecm2, Sc1 | |||
|
Domain Architecture |

|
|||||
| Description | SPARC-like protein 1 precursor (Matrix glycoprotein Sc1) (Extracellular matrix protein 2). | |||||
|
MYT1_HUMAN
|
||||||
| NC score | 0.067301 (rank : 41) | θ value | 3.0926 (rank : 81) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 475 | Shared Neighborhood Hits | 37 | |
| SwissProt Accessions | Q01538, O94922, Q9UPV2 | Gene names | MYT1, KIAA0835, KIAA1050, MTF1, MYTI, PLPB1 | |||
|
Domain Architecture |

|
|||||
| Description | Myelin transcription factor 1 (MyT1) (MyTI) (Proteolipid protein- binding protein) (PLPB1). | |||||
|
NSBP1_HUMAN
|
||||||
| NC score | 0.066785 (rank : 42) | θ value | θ > 10 (rank : 158) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 251 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P82970 | Gene names | NSBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1. | |||||
|
RPGR1_MOUSE
|
||||||
| NC score | 0.065630 (rank : 43) | θ value | 0.62314 (rank : 33) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 713 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9EPQ2, Q8CAC2, Q8CDJ9, Q91WE0, Q9CUK6, Q9D5Q1 | Gene names | Rpgrip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | X-linked retinitis pigmentosa GTPase regulator-interacting protein 1 (RPGR-interacting protein 1). | |||||
|
RGP1_MOUSE
|
||||||
| NC score | 0.065302 (rank : 44) | θ value | 1.06291 (rank : 45) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 105 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P46061, Q60801 | Gene names | Rangap1, Fug1 | |||
|
Domain Architecture |

|
|||||
| Description | Ran GTPase-activating protein 1. | |||||
|
TRI44_MOUSE
|
||||||
| NC score | 0.065146 (rank : 45) | θ value | 1.81305 (rank : 60) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 327 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9QXA7, Q80SV9, Q9JHR8 | Gene names | Trim44, Dipb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 44 (Protein DIPB) (Protein Mc7). | |||||
|
TXND2_MOUSE
|
||||||
| NC score | 0.063781 (rank : 46) | θ value | θ > 10 (rank : 166) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 828 | Shared Neighborhood Hits | 66 | |
| SwissProt Accessions | Q6P902, Q8CJD1 | Gene names | Txndc2, Sptrx, Sptrx1, Trx4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1) (Thioredoxin-4). | |||||
|
TXND2_HUMAN
|
||||||
| NC score | 0.062925 (rank : 47) | θ value | 4.03905 (rank : 100) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 793 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q86VQ3, Q8N7U4, Q96RX3, Q9H0L8 | Gene names | TXNDC2, SPTRX, SPTRX1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thioredoxin domain-containing protein 2 (Spermatid-specific thioredoxin-1) (Sptrx-1). | |||||
|
AN32E_HUMAN
|
||||||
| NC score | 0.062908 (rank : 48) | θ value | 2.36792 (rank : 64) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 291 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9BTT0, Q8N1S4, Q8WWW9 | Gene names | ANP32E | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acidic leucine-rich nuclear phosphoprotein 32 family member E (LANP- like protein) (LANP-L). | |||||
|
UBP35_HUMAN
|
||||||
| NC score | 0.062495 (rank : 49) | θ value | 0.21417 (rank : 14) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9P2H5 | Gene names | USP35, KIAA1372, USP34 | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 35 (EC 3.1.2.15) (Ubiquitin thioesterase 35) (Ubiquitin-specific-processing protease 35) (Deubiquitinating enzyme 35). | |||||
|
AN32B_MOUSE
|
||||||
| NC score | 0.061271 (rank : 50) | θ value | 2.36792 (rank : 63) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9EST5 | Gene names | Anp32b, Pal31 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acidic leucine-rich nuclear phosphoprotein 32 family member B (Proliferation-related acidic leucine-rich protein PAL31). | |||||
|
MATR3_MOUSE
|
||||||
| NC score | 0.061209 (rank : 51) | θ value | θ > 10 (rank : 155) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | Q8K310 | Gene names | Matr3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Matrin-3. | |||||
|
AN32C_HUMAN
|
||||||
| NC score | 0.060886 (rank : 52) | θ value | 1.81305 (rank : 53) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O43423 | Gene names | ANP32C, PP32R1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acidic leucine-rich nuclear phosphoprotein 32 family member C (Phosphoprotein 32-related protein 1) (Tumorigenic protein pp32r1). | |||||
|
CENPB_MOUSE
|
||||||
| NC score | 0.060237 (rank : 53) | θ value | 6.88961 (rank : 112) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 163 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P27790 | Gene names | Cenpb, Cenp-b | |||
|
Domain Architecture |

|
|||||
| Description | Major centromere autoantigen B (Centromere protein B) (CENP-B). | |||||
|
SMRC2_HUMAN
|
||||||
| NC score | 0.059955 (rank : 54) | θ value | 4.03905 (rank : 97) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 499 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q8TAQ2, Q92923, Q96E12, Q96GY4 | Gene names | SMARCC2, BAF170 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 2 (SWI/SNF complex 170 kDa subunit) (BRG1-associated factor 170). | |||||
|
NOLC1_HUMAN
|
||||||
| NC score | 0.059669 (rank : 55) | θ value | 1.38821 (rank : 49) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q14978, Q15030 | Gene names | NOLC1, KIAA0035 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar phosphoprotein p130 (Nucleolar 130 kDa protein) (140 kDa nucleolar phosphoprotein) (Nopp140) (Nucleolar and coiled-body phosphoprotein 1). | |||||
|
AN32A_HUMAN
|
||||||
| NC score | 0.058789 (rank : 56) | θ value | 2.36792 (rank : 62) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 234 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P39687 | Gene names | ANP32A, C15orf1, LANP, MAPM, PHAP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acidic leucine-rich nuclear phosphoprotein 32 family member A (Potent heat-stable protein phosphatase 2A inhibitor I1PP2A) (Acidic nuclear phosphoprotein pp32) (Leucine-rich acidic nuclear protein) (Lanp) (Putative HLA-DR-associated protein I) (PHAPI) (Mapmodulin). | |||||
|
SRCH_HUMAN
|
||||||
| NC score | 0.058599 (rank : 57) | θ value | θ > 10 (rank : 163) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P23327, Q504Y6 | Gene names | HRC, HCP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcoplasmic reticulum histidine-rich calcium-binding protein precursor. | |||||
|
OGFR_MOUSE
|
||||||
| NC score | 0.058077 (rank : 58) | θ value | θ > 10 (rank : 159) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 584 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q99PG2, Q99L33 | Gene names | Ogfr | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor). | |||||
|
LSD1_MOUSE
|
||||||
| NC score | 0.057537 (rank : 59) | θ value | 0.62314 (rank : 31) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 224 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q6ZQ88, Q6PB53, Q8VEA1 | Gene names | Aof2, Kiaa0601, Lsd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lysine-specific histone demethylase 1 (EC 1.-.-.-) (Flavin-containing amine oxidase domain-containing protein 2) (BRAF35-HDAC complex protein BHC110). | |||||
|
NIPBL_HUMAN
|
||||||
| NC score | 0.057179 (rank : 60) | θ value | θ > 10 (rank : 157) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1311 | Shared Neighborhood Hits | 86 | |
| SwissProt Accessions | Q6KC79, Q6KCD6, Q6N080, Q6ZT92, Q7Z2E6, Q8N4M5, Q9Y6Y3, Q9Y6Y4 | Gene names | NIPBL, IDN3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nipped-B-like protein (Delangin) (SCC2 homolog). | |||||
|
CCD96_MOUSE
|
||||||
| NC score | 0.057107 (rank : 61) | θ value | 0.0431538 (rank : 7) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 704 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9CR92 | Gene names | Ccdc96 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 96. | |||||
|
SEC63_MOUSE
|
||||||
| NC score | 0.056338 (rank : 62) | θ value | θ > 10 (rank : 161) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 601 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q8VHE0, Q8VEB9 | Gene names | Sec63, Sec63l | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
LSD1_HUMAN
|
||||||
| NC score | 0.056143 (rank : 63) | θ value | 0.62314 (rank : 30) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O60341, Q5TH94, Q5TH95, Q86VT7, Q8IXK4, Q8NDP6, Q8TAZ3, Q96AW4 | Gene names | AOF2, KIAA0601, LSD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lysine-specific histone demethylase 1 (EC 1.-.-.-) (Flavin-containing amine oxidase domain-containing protein 2) (BRAF35-HDAC complex protein BHC110). | |||||
|
SEC63_HUMAN
|
||||||
| NC score | 0.055975 (rank : 64) | θ value | θ > 10 (rank : 160) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 586 | Shared Neighborhood Hits | 67 | |
| SwissProt Accessions | Q9UGP8, O95380, Q86VS9, Q8IWL0, Q9NTE0 | Gene names | SEC63, SEC63L | |||
|
Domain Architecture |

|
|||||
| Description | Translocation protein SEC63 homolog. | |||||
|
NCKX1_HUMAN
|
||||||
| NC score | 0.055856 (rank : 65) | θ value | 3.0926 (rank : 82) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 29 | |
| SwissProt Accessions | O60721, O43485, O75184 | Gene names | SLC24A1, KIAA0702, NCKX1 | |||
|
Domain Architecture |

|
|||||
| Description | Sodium/potassium/calcium exchanger 1 (Na(+)/K(+)/Ca(2+)-exchange protein 1) (Retinal rod Na-Ca+K exchanger). | |||||
|
VPS72_MOUSE
|
||||||
| NC score | 0.055746 (rank : 66) | θ value | θ > 10 (rank : 168) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q62481, Q3U2N4, Q810A9, Q99K81 | Gene names | Vps72, Tcfl1, Yl1 | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar protein sorting-associated protein 72 homolog (Transcription factor-like 1) (Protein YL-1). | |||||
|
UBF1_HUMAN
|
||||||
| NC score | 0.055665 (rank : 67) | θ value | 1.38821 (rank : 52) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 415 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | P17480 | Gene names | UBTF, UBF, UBF1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar transcription factor 1 (Upstream-binding factor 1) (UBF-1) (Autoantigen NOR-90). | |||||
|
MARCS_MOUSE
|
||||||
| NC score | 0.055495 (rank : 68) | θ value | 2.36792 (rank : 70) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P26645 | Gene names | Marcks, Macs | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myristoylated alanine-rich C-kinase substrate (MARCKS). | |||||
|
LRRF1_HUMAN
|
||||||
| NC score | 0.054146 (rank : 69) | θ value | 4.03905 (rank : 90) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 625 | Shared Neighborhood Hits | 54 | |
| SwissProt Accessions | Q32MZ4, O75766, O75799, Q32MZ5, Q53T49, Q6PKG2 | Gene names | LRRFIP1, GCF2, TRIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine-rich repeat flightless-interacting protein 1 (LRR FLII- interacting protein 1) (TAR RNA-interacting protein) (GC-binding factor 2). | |||||
|
HSP3_MOUSE
|
||||||
| NC score | 0.053399 (rank : 70) | θ value | θ > 10 (rank : 152) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q62100 | Gene names | Prm3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sperm protamine P3. | |||||
|
MYST4_HUMAN
|
||||||
| NC score | 0.053064 (rank : 71) | θ value | 0.47712 (rank : 25) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 52 | |
| SwissProt Accessions | Q8WYB5, O15087, Q86Y05, Q8WU81, Q9UKW2, Q9UKW3, Q9UKX0 | Gene names | MYST4, KIAA0383, MORF, MOZ2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone acetyltransferase MYST4 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 4) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 4) (Histone acetyltransferase MOZ2) (Monocytic leukemia zinc finger protein- related factor) (Histone acetyltransferase MORF). | |||||
|
HTSF1_HUMAN
|
||||||
| NC score | 0.052678 (rank : 72) | θ value | 2.36792 (rank : 69) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | O43719, Q59G06, Q99730 | Gene names | HTATSF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | HIV Tat-specific factor 1 (Tat-SF1). | |||||
|
UBF1_MOUSE
|
||||||
| NC score | 0.052571 (rank : 73) | θ value | 3.0926 (rank : 86) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 419 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P25976 | Gene names | Ubtf, Tcfubf, Ubf-1, Ubf1 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar transcription factor 1 (Upstream-binding factor 1) (UBF-1). | |||||
|
TCAL6_HUMAN
|
||||||
| NC score | 0.052511 (rank : 74) | θ value | θ > 10 (rank : 165) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q6IPX3, Q5H9J8 | Gene names | TCEAL6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor A protein-like 6 (TCEA-like protein 6) (Transcription elongation factor S-II protein-like 6). | |||||
|
VPS72_HUMAN
|
||||||
| NC score | 0.052281 (rank : 75) | θ value | θ > 10 (rank : 167) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q15906, Q53GJ2, Q5U0R4 | Gene names | VPS72, TCFL1, YL1 | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar protein sorting-associated protein 72 homolog (Transcription factor-like 1) (Protein YL-1). | |||||
|
BASP_HUMAN
|
||||||
| NC score | 0.051767 (rank : 76) | θ value | θ > 10 (rank : 149) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 321 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | P80723, O43596, Q5U0S0 | Gene names | BASP1, NAP22 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Brain acid soluble protein 1 (BASP1 protein) (Neuronal axonal membrane protein NAP-22) (22 kDa neuronal tissue-enriched acidic protein). | |||||
|
CSTN1_HUMAN
|
||||||
| NC score | 0.051336 (rank : 77) | θ value | 0.47712 (rank : 24) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 183 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O94985 | Gene names | CLSTN1, CS1, KIAA0911 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Calsyntenin-1 precursor. | |||||
|
CC47_MOUSE
|
||||||
| NC score | 0.051336 (rank : 78) | θ value | θ > 10 (rank : 150) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 245 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q9D024, Q3TSB5, Q3V1S7, Q8C5D9, Q920S6 | Gene names | Ccdc47, Asp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 47 precursor (Adipocyte-specific protein 4). | |||||
|
NCOR1_MOUSE
|
||||||
| NC score | 0.050276 (rank : 79) | θ value | 1.81305 (rank : 55) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 521 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q60974, Q60812 | Gene names | Ncor1, Rxrip13 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear receptor corepressor 1 (N-CoR1) (N-CoR) (Retinoid X receptor- interacting protein 13) (RIP13). | |||||
|
SRCA_HUMAN
|
||||||
| NC score | 0.050208 (rank : 80) | θ value | 2.36792 (rank : 74) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 380 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | Q86TD4 | Gene names | SRL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcalumenin precursor. | |||||
|
CIR_HUMAN
|
||||||
| NC score | 0.049765 (rank : 81) | θ value | 1.06291 (rank : 42) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 249 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q86X95, O95367, Q12804, Q4G1B9, Q6PJI4, Q8IWI2 | Gene names | CIR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CBF1-interacting corepressor (Recepin). | |||||
|
MYT1L_HUMAN
|
||||||
| NC score | 0.048960 (rank : 82) | θ value | 4.03905 (rank : 92) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 471 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | Q9UL68, Q6IQ17, Q9UPP6 | Gene names | MYT1L, KIAA1106 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Myelin transcription factor 1-like protein (MyT1L protein) (MyT1-L). | |||||
|
RGP1_HUMAN
|
||||||
| NC score | 0.048197 (rank : 83) | θ value | 5.27518 (rank : 107) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 124 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P46060 | Gene names | RANGAP1 | |||
|
Domain Architecture |

|
|||||
| Description | Ran GTPase-activating protein 1. | |||||
|
AP3B2_HUMAN
|
||||||
| NC score | 0.047461 (rank : 84) | θ value | 1.06291 (rank : 40) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 28 | |
| SwissProt Accessions | Q13367, O14808 | Gene names | AP3B2 | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit beta-2 (Adapter-related protein complex 3 beta-2 subunit) (Beta3B-adaptin) (Adaptor protein complex AP-3 beta-2 subunit) (AP-3 complex beta-2 subunit) (Clathrin assembly protein complex 3 beta-2 large chain) (Neuron-specific vesicle coat protein beta-NAP). | |||||
|
ARI3A_HUMAN
|
||||||
| NC score | 0.046614 (rank : 85) | θ value | 5.27518 (rank : 104) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q99856, Q8IZA7 | Gene names | ARID3A, DRIL1, DRX, E2FBP1 | |||
|
Domain Architecture |

|
|||||
| Description | AT-rich interactive domain-containing protein 3A (ARID domain- containing protein 3A) (B-cell regulator of IgH transcription) (Bright) (E2F-binding protein 1). | |||||
|
ARS2_MOUSE
|
||||||
| NC score | 0.046519 (rank : 86) | θ value | 8.99809 (rank : 132) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q99MR6, Q99MR4, Q99MR5, Q99MR7 | Gene names | Ars2, Asr2 | |||
|
Domain Architecture |

|
|||||
| Description | Arsenite-resistance protein 2. | |||||
|
FOG1_MOUSE
|
||||||
| NC score | 0.045122 (rank : 87) | θ value | 0.0330416 (rank : 6) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1237 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | O35615 | Gene names | Zfpm1, Fog, Fog1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein ZFPM1 (Zinc finger protein multitype 1) (Friend of GATA protein 1) (Friend of GATA-1) (FOG-1). | |||||
|
PELP1_MOUSE
|
||||||
| NC score | 0.043901 (rank : 88) | θ value | 4.03905 (rank : 95) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 709 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9DBD5, Q5F2E2, Q6PEM0, Q91YM9 | Gene names | Pelp1, Mnar | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Proline-, glutamic acid- and leucine-rich protein 1 (Modulator of nongenomic activity of estrogen receptor). | |||||
|
ABTAP_MOUSE
|
||||||
| NC score | 0.043362 (rank : 89) | θ value | 4.03905 (rank : 88) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 381 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q3V1V3 | Gene names | Abtap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ABT1-associated protein. | |||||
|
SMRC1_MOUSE
|
||||||
| NC score | 0.043080 (rank : 90) | θ value | 6.88961 (rank : 124) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 459 | Shared Neighborhood Hits | 55 | |
| SwissProt Accessions | P97496, Q7TS80, Q7TT29 | Gene names | Smarcc1, Baf155, Srg3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily C member 1 (SWI/SNF complex 155 kDa subunit) (BRG1-associated factor 155) (SWI3-related protein). | |||||
|
OGFR_HUMAN
|
||||||
| NC score | 0.042332 (rank : 91) | θ value | 1.81305 (rank : 56) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1582 | Shared Neighborhood Hits | 56 | |
| SwissProt Accessions | Q9NZT2, O96029, Q4VXW5, Q96CM2, Q9BQW1, Q9H4H0, Q9H7J5, Q9NZT3, Q9NZT4 | Gene names | OGFR | |||
|
Domain Architecture |

|
|||||
| Description | Opioid growth factor receptor (OGFr) (Zeta-type opioid receptor) (7-60 protein). | |||||
|
SYTL1_MOUSE
|
||||||
| NC score | 0.041620 (rank : 92) | θ value | 6.88961 (rank : 126) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 314 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | Q99N80, Q99J26 | Gene names | Sytl1, Slp1 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-like protein 1 (Exophilin-7). | |||||
|
AP3B2_MOUSE
|
||||||
| NC score | 0.041610 (rank : 93) | θ value | 6.88961 (rank : 109) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9JME5, Q6QR53, Q8R1E5 | Gene names | Ap3b2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AP-3 complex subunit beta-2 (Adapter-related protein complex 3 beta-2 subunit) (Beta3B-adaptin) (Adaptor protein complex AP-3 beta-2 subunit) (Clathrin assembly protein complex 3 beta-2 large chain). | |||||
|
AP3B1_HUMAN
|
||||||
| NC score | 0.040931 (rank : 94) | θ value | 5.27518 (rank : 103) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 283 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | O00203, O00580, Q7Z393, Q9HD66 | Gene names | AP3B1, ADTB3A | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit beta-1 (Adapter-related protein complex 3 beta-1 subunit) (Beta3A-adaptin) (Adaptor protein complex AP-3 beta-1 subunit) (Clathrin assembly protein complex 3 beta-1 large chain). | |||||
|
RGS3_HUMAN
|
||||||
| NC score | 0.040742 (rank : 95) | θ value | 0.279714 (rank : 18) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
MYST3_HUMAN
|
||||||
| NC score | 0.039338 (rank : 96) | θ value | 5.27518 (rank : 106) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 772 | Shared Neighborhood Hits | 47 | |
| SwissProt Accessions | Q92794 | Gene names | MYST3, MOZ, RUNXBP2, ZNF220 | |||
|
Domain Architecture |

|
|||||
| Description | Histone acetyltransferase MYST3 (EC 2.3.1.48) (EC 2.3.1.-) (MYST protein 3) (MOZ, YBF2/SAS3, SAS2 and TIP60 protein 3) (Runt-related transcription factor-binding protein 2) (Monocytic leukemia zinc finger protein) (Zinc finger protein 220). | |||||
|
FBX34_MOUSE
|
||||||
| NC score | 0.037257 (rank : 97) | θ value | 2.36792 (rank : 68) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q80XI1, Q9D290 | Gene names | Fbxo34 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | F-box only protein 34. | |||||
|
DAXX_MOUSE
|
||||||
| NC score | 0.036023 (rank : 98) | θ value | 8.99809 (rank : 134) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 173 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O35613, Q9QWT8, Q9QWV3 | Gene names | Daxx | |||
|
Domain Architecture |

|
|||||
| Description | Death domain-associated protein 6 (Daxx). | |||||
|
CSTN1_MOUSE
|
||||||
| NC score | 0.035873 (rank : 99) | θ value | 4.03905 (rank : 89) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 175 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9EPL2 | Gene names | Clstn1, Cs1, Cstn1 | |||
|
Domain Architecture |

|
|||||
| Description | Calsyntenin-1 precursor. | |||||
|
FGD5_HUMAN
|
||||||
| NC score | 0.035825 (rank : 100) | θ value | 0.163984 (rank : 12) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 235 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6ZNL6, Q6MZY1, Q7Z303, Q8IYP3, Q8N861, Q8N8G4 | Gene names | FGD5, ZFYVE23 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FYVE, RhoGEF and PH domain-containing protein 5 (Zinc finger FYVE domain-containing protein 23). | |||||
|
FOG1_HUMAN
|
||||||
| NC score | 0.034965 (rank : 101) | θ value | 0.62314 (rank : 29) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 981 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8IX07 | Gene names | ZFPM1, FOG1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein ZFPM1 (Zinc finger protein multitype 1) (Friend of GATA protein 1) (Friend of GATA-1) (FOG-1). | |||||
|
CAH9_HUMAN
|
||||||
| NC score | 0.033835 (rank : 102) | θ value | 0.279714 (rank : 15) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q16790 | Gene names | CA9, G250, MN | |||
|
Domain Architecture |

|
|||||
| Description | Carbonic anhydrase 9 precursor (EC 4.2.1.1) (Carbonic anhydrase IX) (Carbonate dehydratase IX) (CA-IX) (CAIX) (Membrane antigen MN) (P54/58N) (Renal cell carcinoma-associated antigen G250) (RCC- associated antigen G250) (pMW1). | |||||
|
TM16H_HUMAN
|
||||||
| NC score | 0.033726 (rank : 103) | θ value | 4.03905 (rank : 99) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 139 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9HCE9 | Gene names | TMEM16H, KIAA1623 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transmembrane protein 16H. | |||||
|
SYTL1_HUMAN
|
||||||
| NC score | 0.033626 (rank : 104) | θ value | 1.06291 (rank : 46) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 227 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8IYJ3, Q96BB6, Q96GU6, Q96S89, Q96SI0 | Gene names | SYTL1, SLP1 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-like protein 1 (Exophilin-7) (Protein JFC1) (SB146). | |||||
|
FOG2_MOUSE
|
||||||
| NC score | 0.033479 (rank : 105) | θ value | 3.0926 (rank : 78) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 714 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q8CCH7, Q9Z0F2 | Gene names | Zfpm2, Fog2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein ZFPM2 (Zinc finger protein multitype 2) (Friend of GATA protein 2) (Friend of GATA-2) (FOG-2) (mFOG-2). | |||||
|
MINT_MOUSE
|
||||||
| NC score | 0.033082 (rank : 106) | θ value | 6.88961 (rank : 118) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1514 | Shared Neighborhood Hits | 57 | |
| SwissProt Accessions | Q62504, Q80TN9, Q99PS4, Q9QZW2 | Gene names | Spen, Kiaa0929, Mint, Sharp | |||
|
Domain Architecture |

|
|||||
| Description | Msx2-interacting protein (SPEN homolog) (SMART/HDAC1-associated repressor protein). | |||||
|
JHD3B_HUMAN
|
||||||
| NC score | 0.031474 (rank : 107) | θ value | 0.813845 (rank : 39) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 128 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | O94953, O75274, Q6P3R5, Q9P1V1, Q9UF40 | Gene names | JMJD2B, JHDM3B, KIAA0876 | |||
|
Domain Architecture |

|
|||||
| Description | JmjC domain-containing histone demethylation protein 3B (EC 1.14.11.-) (Jumonji domain-containing protein 2B). | |||||
|
JHD2C_HUMAN
|
||||||
| NC score | 0.030811 (rank : 108) | θ value | 1.06291 (rank : 43) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 279 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q15652, Q5SQZ8, Q5SQZ9, Q5SR00, Q7Z3E7, Q8N3U0, Q96KB9, Q9P2G7 | Gene names | JMJD1C, JHDM2C, KIAA1380, TRIP8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable JmjC domain-containing histone demethylation protein 2C (EC 1.14.11.-) (Jumonji domain-containing protein 1C) (Thyroid receptor-interacting protein 8) (TRIP-8). | |||||
|
CLIC6_HUMAN
|
||||||
| NC score | 0.029674 (rank : 109) | θ value | 6.88961 (rank : 113) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 289 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | Q96NY7, Q8IX31 | Gene names | CLIC6, CLIC1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chloride intracellular channel 6. | |||||
|
NFH_HUMAN
|
||||||
| NC score | 0.029432 (rank : 110) | θ value | 8.99809 (rank : 142) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1242 | Shared Neighborhood Hits | 48 | |
| SwissProt Accessions | P12036, Q9UJS7, Q9UQ14 | Gene names | NEFH, KIAA0845, NFH | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet H protein (200 kDa neurofilament protein) (Neurofilament heavy polypeptide) (NF-H). | |||||
|
MLL2_HUMAN
|
||||||
| NC score | 0.029203 (rank : 111) | θ value | 4.03905 (rank : 91) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
DPOD3_HUMAN
|
||||||
| NC score | 0.027078 (rank : 112) | θ value | 6.88961 (rank : 115) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 109 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q15054 | Gene names | POLD3, KIAA0039 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA polymerase subunit delta 3 (DNA polymerase subunit delta p66). | |||||
|
TCF8_HUMAN
|
||||||
| NC score | 0.026977 (rank : 113) | θ value | 0.62314 (rank : 35) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1057 | Shared Neighborhood Hits | 51 | |
| SwissProt Accessions | P37275, Q12924, Q13800 | Gene names | TCF8, AREB6 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 8 (NIL-2-A zinc finger protein) (Negative regulator of IL2). | |||||
|
NPAS3_MOUSE
|
||||||
| NC score | 0.026834 (rank : 114) | θ value | 2.36792 (rank : 71) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9QZQ0, Q9EQP4 | Gene names | Npas3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 3 (Neuronal PAS3) (Member of PAS protein 6) (MOP6). | |||||
|
MADCA_HUMAN
|
||||||
| NC score | 0.026804 (rank : 115) | θ value | 8.99809 (rank : 140) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 221 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q13477, O60222, O75867 | Gene names | MADCAM1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mucosal addressin cell adhesion molecule 1 precursor (MAdCAM-1) (hMAdCAM-1). | |||||
|
MAP2_HUMAN
|
||||||
| NC score | 0.026055 (rank : 116) | θ value | 8.99809 (rank : 141) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 348 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | P11137, Q99975, Q99976 | Gene names | MAP2 | |||
|
Domain Architecture |

|
|||||
| Description | Microtubule-associated protein 2 (MAP 2) (MAP-2). | |||||
|
NSD1_MOUSE
|
||||||
| NC score | 0.025669 (rank : 117) | θ value | 4.03905 (rank : 94) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 472 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | O88491, Q8C480, Q9CT70 | Gene names | Nsd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Histone-lysine N-methyltransferase, H3 lysine-36 and H4 lysine-20 specific (EC 2.1.1.43) (H3-K36-HMTase) (H4-K20-HMTase) (Nuclear receptor-binding SET domain-containing protein 1) (NR-binding SET domain-containing protein). | |||||
|
SPT5H_HUMAN
|
||||||
| NC score | 0.025459 (rank : 118) | θ value | 8.99809 (rank : 144) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 436 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | O00267, O43279, Q59G52, Q99639 | Gene names | SUPT5H, SPT5, SPT5H | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription elongation factor SPT5 (hSPT5) (DRB sensitivity-inducing factor large subunit) (DSIF large subunit) (DSIF p160) (Tat- cotransactivator 1 protein) (Tat-CT1 protein). | |||||
|
TXLNA_HUMAN
|
||||||
| NC score | 0.024841 (rank : 119) | θ value | 3.0926 (rank : 85) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 982 | Shared Neighborhood Hits | 43 | |
| SwissProt Accessions | P40222, Q66K62, Q86T54, Q86T85, Q86T86, Q86Y86, Q86YW3, Q8N2Y3 | Gene names | TXLNA, TXLN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Alpha-taxilin. | |||||
|
MYO9B_HUMAN
|
||||||
| NC score | 0.024767 (rank : 120) | θ value | 0.279714 (rank : 16) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 641 | Shared Neighborhood Hits | 33 | |
| SwissProt Accessions | Q13459, O75314, Q9NUJ2, Q9UHN0 | Gene names | MYO9B, MYR5 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-9B (Myosin IXb) (Unconventional myosin-9b). | |||||
|
CAC1C_MOUSE
|
||||||
| NC score | 0.024753 (rank : 121) | θ value | 1.81305 (rank : 54) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 130 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q01815, Q04476, Q61242, Q99242 | Gene names | Cacna1c, Cach2, Cacn2, Cacnl1a1, Cchl1a1 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1C (Voltage- gated calcium channel subunit alpha Cav1.2) (Calcium channel, L type, alpha-1 polypeptide, isoform 1, cardiac muscle) (Mouse brain class C) (MBC) (MELC-CC). | |||||
|
CAC1C_HUMAN
|
||||||
| NC score | 0.024496 (rank : 122) | θ value | 2.36792 (rank : 65) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q13936, Q13917, Q13918, Q13919, Q13920, Q13921, Q13922, Q13923, Q13924, Q13925, Q13926, Q13927, Q13928, Q13929, Q13930, Q13932, Q13933, Q14743, Q14744, Q15877, Q99025, Q99241, Q99875 | Gene names | CACNA1C, CACH2, CACN2, CACNL1A1, CCHL1A1 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent L-type calcium channel subunit alpha-1C (Voltage- gated calcium channel subunit alpha Cav1.2) (Calcium channel, L type, alpha-1 polypeptide, isoform 1, cardiac muscle). | |||||
|
TCF8_MOUSE
|
||||||
| NC score | 0.024453 (rank : 123) | θ value | 1.81305 (rank : 59) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1067 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | Q64318, Q62519 | Gene names | Tcf8, Zfhx1a, Zfx1a, Zfx1ha | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor 8 (Zinc finger homeobox protein 1a) (MEB1) (Delta EF1). | |||||
|
SCN5A_HUMAN
|
||||||
| NC score | 0.024247 (rank : 124) | θ value | 0.62314 (rank : 34) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q14524 | Gene names | SCN5A | |||
|
Domain Architecture |

|
|||||
| Description | Sodium channel protein type 5 subunit alpha (Sodium channel protein type V subunit alpha) (Voltage-gated sodium channel subunit alpha Nav1.5) (Sodium channel protein, cardiac muscle alpha-subunit) (HH1). | |||||
|
NPAS3_HUMAN
|
||||||
| NC score | 0.023881 (rank : 125) | θ value | 4.03905 (rank : 93) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8IXF0, Q86US6, Q86US7, Q8IXF2, Q9BY81, Q9H323, Q9Y4L8 | Gene names | NPAS3 | |||
|
Domain Architecture |

|
|||||
| Description | Neuronal PAS domain-containing protein 3 (Neuronal PAS3) (Member of PAS protein 6) (MOP6). | |||||
|
SSRP1_HUMAN
|
||||||
| NC score | 0.023556 (rank : 126) | θ value | 5.27518 (rank : 108) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q08945, Q5BJG8 | Gene names | SSRP1, FACT80 | |||
|
Domain Architecture |

|
|||||
| Description | FACT complex subunit SSRP1 (Facilitates chromatin transcription complex subunit SSRP1) (FACT 80 kDa subunit) (FACTp80) (Chromatin- specific transcription elongation factor 80 kDa subunit) (Structure- specific recognition protein 1) (hSSRP1) (Recombination signal sequence recognition protein 1) (T160). | |||||
|
NFM_MOUSE
|
||||||
| NC score | 0.022640 (rank : 127) | θ value | 1.38821 (rank : 47) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1159 | Shared Neighborhood Hits | 45 | |
| SwissProt Accessions | P08553, Q61961 | Gene names | Nef3, Nefm, Nfm | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet M protein (160 kDa neurofilament protein) (Neurofilament medium polypeptide) (NF-M). | |||||
|
SAS10_HUMAN
|
||||||
| NC score | 0.022618 (rank : 128) | θ value | 8.99809 (rank : 143) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NQZ2, Q6FI82 | Gene names | CRLZ1, SAS10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Something about silencing protein 10 (Disrupter of silencing SAS10) (Charged amino acid-rich leucine zipper 1). | |||||
|
DDX51_HUMAN
|
||||||
| NC score | 0.022544 (rank : 129) | θ value | 2.36792 (rank : 66) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 332 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q8N8A6, Q5CZ71, Q8IXK5, Q96ED1 | Gene names | DDX51 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | ATP-dependent RNA helicase DDX51 (EC 3.6.1.-) (DEAD box protein 51). | |||||
|
DDIT3_HUMAN
|
||||||
| NC score | 0.022395 (rank : 130) | θ value | 8.99809 (rank : 135) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 167 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P35638 | Gene names | DDIT3, CHOP, GADD153 | |||
|
Domain Architecture |

|
|||||
| Description | DNA damage-inducible transcript 3 (DDIT-3) (Growth arrest and DNA- damage-inducible protein GADD153) (C/EBP-homologous protein) (CHOP). | |||||
|
NGEF_MOUSE
|
||||||
| NC score | 0.021933 (rank : 131) | θ value | 1.38821 (rank : 48) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 228 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8CHT1, Q8R204, Q923H2, Q9JHT9 | Gene names | Ngef | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ephexin-1 (Eph-interacting exchange protein) (Neuronal guanine nucleotide exchange factor). | |||||
|
NFM_HUMAN
|
||||||
| NC score | 0.021628 (rank : 132) | θ value | 6.88961 (rank : 119) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1328 | Shared Neighborhood Hits | 50 | |
| SwissProt Accessions | P07197 | Gene names | NEF3, NEFM, NFM | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet M protein (160 kDa neurofilament protein) (Neurofilament medium polypeptide) (NF-M) (Neurofilament 3). | |||||
|
PEG3_MOUSE
|
||||||
| NC score | 0.021369 (rank : 133) | θ value | 6.88961 (rank : 120) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1101 | Shared Neighborhood Hits | 84 | |
| SwissProt Accessions | Q3URU2, O54978, Q3TQ69, Q5EBP7, Q61138, Q6GQS0, Q80U47, Q8R5N0, Q9QX53 | Gene names | Peg3, Kiaa0287, Pw1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Paternally expressed gene 3 protein (ASF-1). | |||||
|
B4GN4_MOUSE
|
||||||
| NC score | 0.020865 (rank : 134) | θ value | 3.0926 (rank : 77) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q766D5, Q8CHV2 | Gene names | B4galnt4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | N-acetyl-beta-glucosaminyl-glycoprotein 4-beta-N- acetylgalactosaminyltransferase 1 (EC 2.4.1.244) (NGalNAc-T1) (Beta- 1,4-N-acetylgalactosaminyltransferase IV) (Beta4GalNAc-T4) (Beta4GalNAcT4). | |||||
|
UBP8_MOUSE
|
||||||
| NC score | 0.019993 (rank : 135) | θ value | 8.99809 (rank : 148) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 362 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q80U87, Q80YP2, Q8R0D3, Q9EQU1, Q9WVP5 | Gene names | Usp8, Kiaa0055, Ubpy | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 8 (EC 3.1.2.15) (Ubiquitin thioesterase 8) (Ubiquitin-specific-processing protease 8) (Deubiquitinating enzyme 8) (mUBPy). | |||||
|
PCNT_MOUSE
|
||||||
| NC score | 0.019032 (rank : 136) | θ value | 2.36792 (rank : 72) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1207 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P48725 | Gene names | Pcnt, Pcnt2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pericentrin. | |||||
|
FIP1_HUMAN
|
||||||
| NC score | 0.017831 (rank : 137) | θ value | 8.99809 (rank : 137) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 118 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q6UN15, Q499Y4, Q49AU3, Q7Z608, Q8WVN3, Q96F80, Q9H077 | Gene names | FIP1L1, FIP1, RHE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pre-mRNA 3'-end-processing factor FIP1 (FIP1-like 1) (Factor interacting with PAP) (hFip1) (Rearranged in hypereosinophilia). | |||||
|
CD2L1_MOUSE
|
||||||
| NC score | 0.017105 (rank : 138) | θ value | 0.47712 (rank : 23) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1463 | Shared Neighborhood Hits | 40 | |
| SwissProt Accessions | P24788, Q3UI03, Q61399, Q7TST4, Q8BP53 | Gene names | Cdc2l1 | |||
|
Domain Architecture |

|
|||||
| Description | PITSLRE serine/threonine-protein kinase CDC2L1 (EC 2.7.11.22) (Galactosyltransferase-associated protein kinase p58/GTA) (Cell division cycle 2-like protein kinase 1). | |||||
|
K0256_HUMAN
|
||||||
| NC score | 0.016927 (rank : 139) | θ value | 6.88961 (rank : 116) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q93073, Q8N767 | Gene names | KIAA0256 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0256. | |||||
|
TRI56_HUMAN
|
||||||
| NC score | 0.016822 (rank : 140) | θ value | 6.88961 (rank : 128) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 339 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9BRZ2, Q86VT6, Q8N2H8, Q8NAC0, Q9H031 | Gene names | TRIM56, RNF109 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tripartite motif-containing protein 56 (RING finger protein 109). | |||||
|
CD2L1_HUMAN
|
||||||
| NC score | 0.016543 (rank : 141) | θ value | 0.813845 (rank : 38) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1467 | Shared Neighborhood Hits | 39 | |
| SwissProt Accessions | P21127, O95265, Q12817, Q12818, Q12819, Q12820, Q12822, Q8N530, Q9NZS5, Q9UBJ0, Q9UBQ1, Q9UBR0, Q9UNY2, Q9UP57, Q9UP58, Q9UP59 | Gene names | CDC2L1, PITSLREA, PK58 | |||
|
Domain Architecture |

|
|||||
| Description | PITSLRE serine/threonine-protein kinase CDC2L1 (EC 2.7.11.22) (Galactosyltransferase-associated protein kinase p58/GTA) (Cell division cycle 2-like protein kinase 1) (CLK-1) (CDK11) (p58 CLK-1). | |||||
|
TAF1L_HUMAN
|
||||||
| NC score | 0.015773 (rank : 142) | θ value | 8.99809 (rank : 145) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q8IZX4 | Gene names | TAF1L | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID 210 kDa subunit (TBP-associated factor 210 kDa) (TAF(II)210) (TBP-associated factor 1-like). | |||||
|
UBP4_HUMAN
|
||||||
| NC score | 0.015680 (rank : 143) | θ value | 4.03905 (rank : 101) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q13107, O43452, O43453 | Gene names | USP4, UNP, UNPH | |||
|
Domain Architecture |

|
|||||
| Description | Ubiquitin carboxyl-terminal hydrolase 4 (EC 3.1.2.15) (Ubiquitin thioesterase 4) (Ubiquitin-specific-processing protease 4) (Deubiquitinating enzyme 4) (Ubiquitous nuclear protein homolog). | |||||
|
ZFHX2_HUMAN
|
||||||
| NC score | 0.015279 (rank : 144) | θ value | 3.0926 (rank : 87) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1178 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q9C0A1 | Gene names | ZFHX2, KIAA1762 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger homeobox protein 2. | |||||
|
ZEP2_HUMAN
|
||||||
| NC score | 0.015132 (rank : 145) | θ value | 6.88961 (rank : 131) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 937 | Shared Neighborhood Hits | 27 | |
| SwissProt Accessions | P31629, Q02646, Q5THT5, Q9NS05 | Gene names | HIVEP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Human immunodeficiency virus type I enhancer-binding protein 2 (HIV- EP2) (MHC-binding protein 2) (MBP-2). | |||||
|
BFSP1_HUMAN
|
||||||
| NC score | 0.014786 (rank : 146) | θ value | 6.88961 (rank : 110) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 508 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q12934, O43595, O76034, Q9HBX4 | Gene names | BFSP1 | |||
|
Domain Architecture |

|
|||||
| Description | Filensin (Beaded filament structural protein 1) (Lens fiber cell beaded-filament structural protein CP 115) (CP115) (Lens intermediate filament-like heavy) (LIFL-H). | |||||
|
PTPRG_HUMAN
|
||||||
| NC score | 0.014411 (rank : 147) | θ value | 3.0926 (rank : 84) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 231 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P23470, Q15623 | Gene names | PTPRG | |||
|
Domain Architecture |

|
|||||
| Description | Receptor-type tyrosine-protein phosphatase gamma precursor (EC 3.1.3.48) (Protein-tyrosine phosphatase gamma) (R-PTP-gamma). | |||||
|
BRD2_HUMAN
|
||||||
| NC score | 0.014182 (rank : 148) | θ value | 8.99809 (rank : 133) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P25440, O00699, O00700, Q15310, Q5STC9, Q969U4 | Gene names | BRD2, KIAA9001, RING3 | |||
|
Domain Architecture |

|
|||||
| Description | Bromodomain-containing protein 2 (Protein RING3) (O27.1.1). | |||||
|
SALL2_MOUSE
|
||||||
| NC score | 0.014138 (rank : 149) | θ value | 4.03905 (rank : 96) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 836 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9QX96 | Gene names | Sall2, Sal2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sal-like protein 2 (Spalt-like protein 2) (MSal-2). | |||||
|
CD2L2_HUMAN
|
||||||
| NC score | 0.013898 (rank : 150) | θ value | 6.88961 (rank : 111) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1439 | Shared Neighborhood Hits | 35 | |
| SwissProt Accessions | Q9UQ88, O95227, O95228, O96012, Q12821, Q12853, Q12854, Q5QPR0, Q5QPR1, Q5QPR2, Q9UBC4, Q9UBI3, Q9UEI1, Q9UEI2, Q9UP53, Q9UP54, Q9UP55, Q9UP56, Q9UQ86, Q9UQ87, Q9UQ89 | Gene names | CDC2L2, PITSLREB | |||
|
Domain Architecture |

|
|||||
| Description | PITSLRE serine/threonine-protein kinase CDC2L2 (EC 2.7.11.22) (Galactosyltransferase-associated protein kinase p58/GTA) (Cell division cycle 2-like protein kinase 2) (CDK11). | |||||
|
HLX1_MOUSE
|
||||||
| NC score | 0.013531 (rank : 151) | θ value | 8.99809 (rank : 139) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 362 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q61670 | Gene names | Hlx1, Hlx | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein HLX1. | |||||
|
SMCA2_HUMAN
|
||||||
| NC score | 0.013396 (rank : 152) | θ value | 6.88961 (rank : 123) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 503 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | P51531 | Gene names | SMARCA2, BRM, SNF2A, SNF2L2 | |||
|
Domain Architecture |

|
|||||
| Description | Probable global transcription activator SNF2L2 (EC 3.6.1.-) (ATP- dependent helicase SMARCA2) (SNF2-alpha) (SWI/SNF-related matrix- associated actin-dependent regulator of chromatin subfamily A member 2) (hBRM). | |||||
|
KIRR3_HUMAN
|
||||||
| NC score | 0.012799 (rank : 153) | θ value | 3.0926 (rank : 80) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 302 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8IZU9, Q6UWJ9, Q6UWL5, Q96JG0 | Gene names | KIRREL3, KIAA1867, NEPH2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Kin of IRRE-like protein 3 precursor (Kin of irregular chiasm-like protein 3) (Nephrin-like 2). | |||||
|
ZBTB5_HUMAN
|
||||||
| NC score | 0.012048 (rank : 154) | θ value | 6.88961 (rank : 129) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 811 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | O15062 | Gene names | ZBTB5, KIAA0354 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and BTB domain-containing protein 5. | |||||
|
G3BP_HUMAN
|
||||||
| NC score | 0.011818 (rank : 155) | θ value | 8.99809 (rank : 138) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 219 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q13283 | Gene names | G3BP, G3BP1 | |||
|
Domain Architecture |

|
|||||
| Description | Ras-GTPase-activating protein-binding protein 1 (EC 3.6.1.-) (ATP- dependent DNA helicase VIII) (GAP SH3-domain-binding protein 1) (G3BP- 1) (HDH-VIII). | |||||
|
TBC16_HUMAN
|
||||||
| NC score | 0.011743 (rank : 156) | θ value | 8.99809 (rank : 146) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8TBP0 | Gene names | TBC1D16 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TBC1 domain family member 16. | |||||
|
SPTN2_HUMAN
|
||||||
| NC score | 0.011741 (rank : 157) | θ value | 1.81305 (rank : 58) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 535 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O15020, O14872, O14873 | Gene names | SPTBN2, KIAA0302 | |||
|
Domain Architecture |

|
|||||
| Description | Spectrin beta chain, brain 2 (Spectrin, non-erythroid beta chain 2) (Beta-III spectrin). | |||||
|
SPEG_HUMAN
|
||||||
| NC score | 0.011582 (rank : 158) | θ value | 1.38821 (rank : 50) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1762 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q15772, Q27J74, Q695L1, Q6FGA6, Q6ZQW1, Q6ZTL8, Q9P2P9 | Gene names | SPEG, APEG1, KIAA1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle preferentially expressed protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
ZN295_HUMAN
|
||||||
| NC score | 0.011168 (rank : 159) | θ value | 2.36792 (rank : 76) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 847 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q9ULJ3, Q6P4R0 | Gene names | ZNF295, KIAA1227, ZBTB21 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 295 (Zinc finger and BTB domain-containing protein 21). | |||||
|
GP124_MOUSE
|
||||||
| NC score | 0.011010 (rank : 160) | θ value | 3.0926 (rank : 79) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 423 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q91ZV8 | Gene names | Gpr124, Tem5 | |||
|
Domain Architecture |

|
|||||
| Description | Probable G-protein coupled receptor 124 precursor (Tumor endothelial marker 5). | |||||
|
SPEG_MOUSE
|
||||||
| NC score | 0.010816 (rank : 161) | θ value | 4.03905 (rank : 98) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 1959 | Shared Neighborhood Hits | 34 | |
| SwissProt Accessions | Q62407, Q3TPH8, Q6P5V1, Q80TF7, Q80ZN0, Q8BZF4, Q9EQJ5 | Gene names | Speg, Apeg1, Kiaa1297 | |||
|
Domain Architecture |

|
|||||
| Description | Striated muscle-specific serine/threonine protein kinase (EC 2.7.11.1) (Aortic preferentially expressed protein 1) (APEG-1). | |||||
|
ZBTB5_MOUSE
|
||||||
| NC score | 0.010797 (rank : 162) | θ value | 6.88961 (rank : 130) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 801 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q7TQG0, Q3TVR7, Q6GV09, Q8BIB5 | Gene names | Zbtb5, Kiaa0354 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger and BTB domain-containing protein 5 (Transcription factor ZNF-POZ). | |||||
|
ZKSC1_HUMAN
|
||||||
| NC score | 0.009606 (rank : 163) | θ value | 2.36792 (rank : 75) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 949 | Shared Neighborhood Hits | 59 | |
| SwissProt Accessions | P17029, P52745, Q8TBW5, Q8TEK7 | Gene names | ZKSCAN1, KOX18, ZNF139, ZNF36 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger with KRAB and SCAN domain-containing protein 1 (Zinc finger protein 36) (Zinc finger protein KOX18). | |||||
|
SMRCD_MOUSE
|
||||||
| NC score | 0.009202 (rank : 164) | θ value | 6.88961 (rank : 125) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 96 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q04692 | Gene names | Smarcad1, Etl1, Kiaa1122 | |||
|
Domain Architecture |

|
|||||
| Description | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A containing DEAD/H box 1 (EC 3.6.1.-) (ATP- dependent helicase SMARCAD1) (Enhancer trap locus homolog 1) (Etl-1). | |||||
|
UBIP1_HUMAN
|
||||||
| NC score | 0.008900 (rank : 165) | θ value | 8.99809 (rank : 147) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NZI7, Q68CT0, Q86Y57, Q9H8V0, Q9UD76, Q9UD78 | Gene names | UBP1, LBP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Upstream-binding protein 1 (LBP-1). | |||||
|
PTN13_MOUSE
|
||||||
| NC score | 0.007957 (rank : 166) | θ value | 6.88961 (rank : 121) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 426 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q64512, Q61494, Q62135, Q64499 | Gene names | Ptpn13, Ptp14 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 13 (EC 3.1.3.48) (Protein tyrosine phosphatase PTP-BL) (Protein-tyrosine phosphatase RIP) (protein tyrosine phosphatase DPZPTP) (PTP36). | |||||
|
ZN174_HUMAN
|
||||||
| NC score | 0.006711 (rank : 167) | θ value | 4.03905 (rank : 102) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 744 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q15697, Q9BQ34 | Gene names | ZNF174, ZSCAN8 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 174 (AW-1) (Zinc finger and SCAN domain-containing protein 8). | |||||
|
REM1_HUMAN
|
||||||
| NC score | 0.003339 (rank : 168) | θ value | 6.88961 (rank : 122) | |||
| Query Neighborhood Hits | 148 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O75628, Q9NP57 | Gene names | REM1, GES, REM | |||
|
Domain Architecture |

|
|||||
| Description | GTP-binding protein REM 1 (Rad and Gem-like GTP-binding protein 1) (GTPase-regulating endothelial cell sprouting). | |||||