Please be patient as the page loads
|
ATF5_MOUSE
|
||||||
| SwissProt Accessions | O70191, Q58E51, Q99NH6 | Gene names | Atf5, Atfx, Nap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-5 (Activating transcription factor 5-alpha/beta) (Transcription factor ATFx) (Transcription factor-like protein ODA-10) (BZIP protein ATF7) (NRIF3- associated protein) (NAP1). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
ATF5_HUMAN
|
||||||
| θ value | 1.15091e-71 (rank : 1) | NC score | 0.908225 (rank : 2) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9Y2D1, Q9BSA1, Q9UNQ3 | Gene names | ATF5, ATFX | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-5 (Activating transcription factor 5) (Transcription factor ATFx). | |||||
|
ATF5_MOUSE
|
||||||
| θ value | 8.55514e-59 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | O70191, Q58E51, Q99NH6 | Gene names | Atf5, Atfx, Nap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-5 (Activating transcription factor 5-alpha/beta) (Transcription factor ATFx) (Transcription factor-like protein ODA-10) (BZIP protein ATF7) (NRIF3- associated protein) (NAP1). | |||||
|
ATF4_MOUSE
|
||||||
| θ value | 1.58096e-12 (rank : 3) | NC score | 0.614334 (rank : 3) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q06507, Q61906 | Gene names | Atf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-4 (C/EBP-related ATF) (C/ATF) (TAXREB67 homolog). | |||||
|
ATF4_HUMAN
|
||||||
| θ value | 4.59992e-12 (rank : 4) | NC score | 0.605299 (rank : 4) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P18848, Q9UH31 | Gene names | ATF4 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-4 (Activating transcription factor 4) (DNA-binding protein TAXREB67) (Cyclic AMP response element-binding protein 2) (CREB2). | |||||
|
DBP_HUMAN
|
||||||
| θ value | 0.00035302 (rank : 5) | NC score | 0.170091 (rank : 8) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q10586 | Gene names | DBP | |||
|
Domain Architecture |

|
|||||
| Description | D site-binding protein (Albumin D box-binding protein) (Albumin D- element-binding protein) (TAXREB302). | |||||
|
DBP_MOUSE
|
||||||
| θ value | 0.00035302 (rank : 6) | NC score | 0.170838 (rank : 7) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q60925, Q8VCX3 | Gene names | Dbp | |||
|
Domain Architecture |

|
|||||
| Description | D site-binding protein (Albumin D box-binding protein) (Albumin D- element-binding protein). | |||||
|
SON_HUMAN
|
||||||
| θ value | 0.00509761 (rank : 7) | NC score | 0.080253 (rank : 28) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
ATF7_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 8) | NC score | 0.173960 (rank : 6) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P17544, Q13814 | Gene names | ATF7, ATFA | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-7 (Activating transcription factor 7) (Transcription factor ATF-A). | |||||
|
ATF7_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 9) | NC score | 0.175514 (rank : 5) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8R0S1 | Gene names | Atf7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-7 (Activating transcription factor 7) (Transcription factor ATF-A). | |||||
|
JUN_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 10) | NC score | 0.161265 (rank : 10) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P05412, Q96G93 | Gene names | JUN | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor AP-1 (Activator protein 1) (AP1) (Proto-oncogene c-jun) (V-jun avian sarcoma virus 17 oncogene homolog) (p39). | |||||
|
JUN_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 11) | NC score | 0.161359 (rank : 9) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P05627 | Gene names | Jun | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor AP-1 (Activator protein 1) (AP1) (Proto-oncogene c-jun) (V-jun avian sarcoma virus 17 oncogene homolog) (Jun A) (AH119). | |||||
|
MYO15_HUMAN
|
||||||
| θ value | 0.0736092 (rank : 12) | NC score | 0.031072 (rank : 56) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9UKN7 | Gene names | MYO15A, MYO15 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-15 (Myosin XV) (Unconventional myosin-15). | |||||
|
A4_MOUSE
|
||||||
| θ value | 0.0961366 (rank : 13) | NC score | 0.083084 (rank : 27) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 275 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P12023, P97487, P97942, Q99K32 | Gene names | App | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 protein precursor (APP) (ABPP) (Alzheimer disease amyloid protein homolog) (Amyloidogenic glycoprotein) (AG) [Contains: Soluble APP-alpha (S-APP-alpha); Soluble APP-beta (S-APP-beta); C99 (APP-C99); Beta-amyloid protein 42 (Beta-APP42); Beta-amyloid protein 40 (Beta-APP40); C83; P3(42); P3(40); Gamma-CTF(59) (Gamma-secretase C-terminal fragment 59) (Amyloid intracellular domain 59) (AID(59)) (APP-C59); Gamma-CTF(57) (Gamma-secretase C-terminal fragment 57) (Amyloid intracellular domain 57) (AID(57)) (APP-C57); Gamma-CTF(50) (Gamma-secretase C-terminal fragment 50) (Amyloid intracellular domain 50) (AID(50)); C31]. | |||||
|
ATF6B_HUMAN
|
||||||
| θ value | 0.125558 (rank : 14) | NC score | 0.079382 (rank : 29) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q99941, Q13269, Q14343, Q14345, Q99635, Q99637, Q9H3V9, Q9H3W1, Q9NPL0 | Gene names | CREBL1, G13 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-6 beta (Activating transcription factor 6 beta) (ATF6-beta) (cAMP-responsive element- binding protein-like 1) (cAMP response element-binding protein-related protein) (Creb-rp) (Protein G13). | |||||
|
YYAP1_HUMAN
|
||||||
| θ value | 0.21417 (rank : 15) | NC score | 0.066637 (rank : 33) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9H869, Q5VYZ4, Q5VYZ7, Q7L4C3, Q7L5E2, Q8IXA6, Q8TEW5, Q8TF04, Q96HB6, Q9BQ64, Q9NV84 | Gene names | YY1AP1, HCCA2, YY1AP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YY1-associated protein 1 (Hepatocellular carcinoma susceptibility protein) (Hepatocellular carcinoma-associated protein 2). | |||||
|
A4_HUMAN
|
||||||
| θ value | 0.279714 (rank : 16) | NC score | 0.069725 (rank : 32) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P05067, P09000, P78438, Q13764, Q13778, Q13793, Q16011, Q16014, Q16019, Q16020, Q9BT38, Q9UCA9, Q9UCB6, Q9UCC8, Q9UCD1, Q9UQ58 | Gene names | APP, A4, AD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 protein precursor (APP) (ABPP) (Alzheimer disease amyloid protein) (Cerebral vascular amyloid peptide) (CVAP) (Protease nexin-II) (PN-II) (APPI) (PreA4) [Contains: Soluble APP-alpha (S-APP- alpha); Soluble APP-beta (S-APP-beta); C99; Beta-amyloid protein 42 (Beta-APP42); Beta-amyloid protein 40 (Beta-APP40); C83; P3(42); P3(40); Gamma-CTF(59) (Gamma-secretase C-terminal fragment 59) (Amyloid intracellular domain 59) (AID(59)); Gamma-CTF(57) (Gamma- secretase C-terminal fragment 57) (Amyloid intracellular domain 57) (AID(57)); Gamma-CTF(50) (Gamma-secretase C-terminal fragment 50) (Amyloid intracellular domain 50) (AID(50)); C31]. | |||||
|
ZN687_HUMAN
|
||||||
| θ value | 0.365318 (rank : 17) | NC score | 0.017803 (rank : 69) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 1244 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8N1G0, Q68DQ8, Q9H937, Q9P2A7 | Gene names | ZNF687, KIAA1441 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 687. | |||||
|
LPP_HUMAN
|
||||||
| θ value | 0.47712 (rank : 18) | NC score | 0.029788 (rank : 58) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 775 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q93052, Q8NFX5 | Gene names | LPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lipoma-preferred partner (LIM domain-containing preferred translocation partner in lipoma). | |||||
|
RNF37_HUMAN
|
||||||
| θ value | 0.47712 (rank : 19) | NC score | 0.057863 (rank : 38) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O94941 | Gene names | UBOX5, KIAA0860, RNF37, UBCE7IP5 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 37 (Ubiquitin-conjugating enzyme 7-interacting protein 5) (U-box domain-containing protein 5). | |||||
|
LARP4_MOUSE
|
||||||
| θ value | 0.813845 (rank : 20) | NC score | 0.048875 (rank : 46) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8BWW4, Q3UR69 | Gene names | Larp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | La-related protein 4 (La ribonucleoprotein domain family member 4). | |||||
|
IRF3_HUMAN
|
||||||
| θ value | 1.06291 (rank : 21) | NC score | 0.039468 (rank : 50) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q14653 | Gene names | IRF3 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon regulatory factor 3 (IRF-3). | |||||
|
PLAL1_HUMAN
|
||||||
| θ value | 1.06291 (rank : 22) | NC score | 0.007948 (rank : 80) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 728 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UM63 | Gene names | PLAGL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein PLAGL1 (Pleiomorphic adenoma-like protein 1). | |||||
|
RGS3_HUMAN
|
||||||
| θ value | 1.06291 (rank : 23) | NC score | 0.036515 (rank : 51) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
EMIL2_MOUSE
|
||||||
| θ value | 1.38821 (rank : 24) | NC score | 0.029372 (rank : 61) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 485 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8K482 | Gene names | Emilin2 | |||
|
Domain Architecture |

|
|||||
| Description | EMILIN-2 precursor (Elastin microfibril interface-located protein 2) (Elastin microfibril interfacer 2) (Basilin). | |||||
|
K1802_MOUSE
|
||||||
| θ value | 1.38821 (rank : 25) | NC score | 0.034253 (rank : 54) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 1596 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8K327, Q3UZ85, Q6ZPI1 | Gene names | Kiaa1802, D8Ertd457e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
MKL1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 26) | NC score | 0.043679 (rank : 47) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 667 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8K4J6 | Gene names | Mkl1, Bsac | |||
|
Domain Architecture |

|
|||||
| Description | MKL/myocardin-like protein 1 (Myocardin-related transcription factor A) (MRTF-A) (Megakaryoblastic leukemia 1 protein homolog) (Basic SAP coiled-coil transcription activator). | |||||
|
MLL3_HUMAN
|
||||||
| θ value | 1.38821 (rank : 27) | NC score | 0.034194 (rank : 55) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
JUND_HUMAN
|
||||||
| θ value | 1.81305 (rank : 28) | NC score | 0.144417 (rank : 14) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P17535 | Gene names | JUND | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor jun-D. | |||||
|
JUND_MOUSE
|
||||||
| θ value | 2.36792 (rank : 29) | NC score | 0.144543 (rank : 13) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P15066 | Gene names | Jund, Jun-d, Jund1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor jun-D. | |||||
|
NDE1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 30) | NC score | 0.023426 (rank : 65) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 823 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NXR1, Q49AQ2 | Gene names | NDE1, NUDE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear distribution protein nudE homolog 1 (NudE). | |||||
|
AMPH_HUMAN
|
||||||
| θ value | 3.0926 (rank : 31) | NC score | 0.029820 (rank : 57) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P49418, O43538, Q75MJ8, Q75MK5, Q75MM3, Q8N4G0 | Gene names | AMPH, AMPH1 | |||
|
Domain Architecture |

|
|||||
| Description | Amphiphysin. | |||||
|
GCM2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 32) | NC score | 0.029626 (rank : 60) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O75603 | Gene names | GCM2, GCMB | |||
|
Domain Architecture |
|
|||||
| Description | Chorion-specific transcription factor GCMb (Glial cells missing homolog 2) (GCM motif protein 2) (hGCMb). | |||||
|
SH22A_MOUSE
|
||||||
| θ value | 3.0926 (rank : 33) | NC score | 0.029306 (rank : 62) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 179 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9QXK9, Q9JHP6 | Gene names | Sh2d2a, Lad, Ribp | |||
|
Domain Architecture |
|
|||||
| Description | SH2 domain protein 2A (Lck-associated adapter protein) (Lad) (Rlk/Itk- binding protein) (Ribp). | |||||
|
ATF3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 34) | NC score | 0.159336 (rank : 11) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P18847 | Gene names | ATF3 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-3 (Activating transcription factor 3). | |||||
|
ATF3_MOUSE
|
||||||
| θ value | 4.03905 (rank : 35) | NC score | 0.155817 (rank : 12) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q60765 | Gene names | Atf3 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-3 (Activating transcription factor 3) (Transcription factor LRG-21). | |||||
|
FETUB_MOUSE
|
||||||
| θ value | 4.03905 (rank : 36) | NC score | 0.029649 (rank : 59) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9QXC1 | Gene names | Fetub | |||
|
Domain Architecture |
|
|||||
| Description | Fetuin-B precursor (IRL685). | |||||
|
GR65_MOUSE
|
||||||
| θ value | 4.03905 (rank : 37) | NC score | 0.025271 (rank : 63) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q91X51, Q9D3L9 | Gene names | Gorasp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi reassembly-stacking protein 1 (Golgi reassembly-stacking protein of 65 kDa) (GRASP65) (Golgi peripheral membrane protein p65). | |||||
|
CREB3_MOUSE
|
||||||
| θ value | 5.27518 (rank : 38) | NC score | 0.091237 (rank : 24) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q61817, Q99M21, Q9CVK9 | Gene names | Creb3, Lzip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclic AMP-responsive element-binding protein 3 (Transcription factor LZIP). | |||||
|
HSF4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 39) | NC score | 0.036432 (rank : 52) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9ULV5, Q99472, Q9ULV6 | Gene names | HSF4 | |||
|
Domain Architecture |
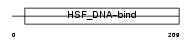
|
|||||
| Description | Heat shock factor protein 4 (HSF 4) (Heat shock transcription factor 4) (HSTF 4) (hHSF4). | |||||
|
TLE4_HUMAN
|
||||||
| θ value | 5.27518 (rank : 40) | NC score | 0.015859 (rank : 70) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 477 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q04727, Q5T1Y2, Q9BZ07, Q9BZ08, Q9BZ09, Q9NSL3, Q9ULF9 | Gene names | TLE4, KIAA1261 | |||
|
Domain Architecture |

|
|||||
| Description | Transducin-like enhancer protein 4. | |||||
|
TLE4_MOUSE
|
||||||
| θ value | 5.27518 (rank : 41) | NC score | 0.015747 (rank : 71) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 482 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q62441, Q9JKQ9 | Gene names | Tle4, Grg4 | |||
|
Domain Architecture |

|
|||||
| Description | Transducin-like enhancer protein 4 (Groucho-related protein 4) (Grg- 4). | |||||
|
CE290_HUMAN
|
||||||
| θ value | 6.88961 (rank : 42) | NC score | 0.009373 (rank : 78) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 1553 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O15078, Q1PSK5, Q66GS8, Q9H2G6, Q9H6Q7, Q9H8I0 | Gene names | CEP290, KIAA0373, NPHP6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein Cep290 (Nephrocystin-6) (Tumor antigen se2-2). | |||||
|
CRSP7_HUMAN
|
||||||
| θ value | 6.88961 (rank : 43) | NC score | 0.041553 (rank : 49) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O95402 | Gene names | CRSP7, ARC70 | |||
|
Domain Architecture |

|
|||||
| Description | CRSP complex subunit 7 (Cofactor required for Sp1 transcriptional activation subunit 7) (Transcriptional coactivator CRSP70) (Activator- recruited cofactor 70 kDa component) (ARC70). | |||||
|
DCC_MOUSE
|
||||||
| θ value | 6.88961 (rank : 44) | NC score | 0.008466 (rank : 79) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P70211 | Gene names | Dcc | |||
|
Domain Architecture |

|
|||||
| Description | Netrin receptor DCC precursor (Tumor suppressor protein DCC). | |||||
|
EFS_MOUSE
|
||||||
| θ value | 6.88961 (rank : 45) | NC score | 0.018641 (rank : 68) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 389 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q64355 | Gene names | Efs, Sin | |||
|
Domain Architecture |
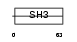
|
|||||
| Description | Embryonal Fyn-associated substrate (SRC-interacting protein) (Signal- integrating protein). | |||||
|
FOXH1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 46) | NC score | 0.010034 (rank : 76) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O75593 | Gene names | FOXH1, FAST1, FAST2 | |||
|
Domain Architecture |
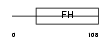
|
|||||
| Description | Forkhead box protein H1 (Forkhead activin signal transducer 1) (Fast- 1) (hFAST-1) (Forkhead activin signal transducer 2) (Fast-2). | |||||
|
IRF2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 47) | NC score | 0.025106 (rank : 64) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P23906 | Gene names | Irf2 | |||
|
Domain Architecture |
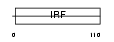
|
|||||
| Description | Interferon regulatory factor 2 (IRF-2). | |||||
|
NOTC2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 48) | NC score | 0.002142 (rank : 83) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 1113 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q04721, Q99734, Q9H240 | Gene names | NOTCH2 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 2 precursor (Notch 2) (hN2) [Contains: Notch 2 extracellular truncation; Notch 2 intracellular domain]. | |||||
|
PTN18_MOUSE
|
||||||
| θ value | 6.88961 (rank : 49) | NC score | 0.009663 (rank : 77) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q61152, Q4JFH4, Q62404, Q922E3 | Gene names | Ptpn18, Flp1, Ptpk1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 18 (EC 3.1.3.48) (Fetal liver phosphatase 1) (FLP-1) (PTP-K1). | |||||
|
ULK1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 50) | NC score | 0.004315 (rank : 82) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 1214 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O70405, Q6PGB2 | Gene names | Ulk1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase ULK1 (EC 2.7.11.1) (Unc-51-like kinase 1) (Serine/threonine-protein kinase Unc51.1). | |||||
|
AMOL2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 51) | NC score | 0.010648 (rank : 74) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 591 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8K371, Q3TPM1, Q7TPE4, Q8BP84, Q8BS08, Q9QUS0 | Gene names | Amotl2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin-like protein 2. | |||||
|
CCD45_HUMAN
|
||||||
| θ value | 8.99809 (rank : 52) | NC score | 0.010381 (rank : 75) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 525 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96GE4, Q96M81 | Gene names | CCDC45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 45. | |||||
|
CEL_HUMAN
|
||||||
| θ value | 8.99809 (rank : 53) | NC score | 0.022610 (rank : 66) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P19835, Q16398 | Gene names | CEL, BAL | |||
|
Domain Architecture |

|
|||||
| Description | Bile salt-activated lipase precursor (EC 3.1.1.3) (EC 3.1.1.13) (BAL) (Bile salt-stimulated lipase) (BSSL) (Carboxyl ester lipase) (Sterol esterase) (Cholesterol esterase) (Pancreatic lysophospholipase). | |||||
|
CSPG5_MOUSE
|
||||||
| θ value | 8.99809 (rank : 54) | NC score | 0.019288 (rank : 67) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q71M36, Q71M37, Q7TNT8, Q8BPJ5, Q9QY32 | Gene names | Cspg5, Caleb, Ngc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate proteoglycan 5 precursor (Neuroglycan C) (Acidic leucine-rich EGF-like domain-containing brain protein). | |||||
|
DOK3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 55) | NC score | 0.013193 (rank : 72) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7L591, Q8N864, Q9BQB3, Q9H666 | Gene names | DOK3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Docking protein 3 (Downstream of tyrosine kinase 3). | |||||
|
GON4L_HUMAN
|
||||||
| θ value | 8.99809 (rank : 56) | NC score | 0.042278 (rank : 48) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 555 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q3T8J9, Q3T8J8, Q5VYZ5, Q5W0D5, Q6AWA6, Q6P1Q6, Q7Z3L3, Q8IY79, Q9BQI1, Q9HCG6 | Gene names | GON4L, GON4, KIAA1606 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GON-4-like protein (GON-4 homolog). | |||||
|
JUNB_HUMAN
|
||||||
| θ value | 8.99809 (rank : 57) | NC score | 0.116080 (rank : 21) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P17275, Q96GH3 | Gene names | JUNB | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor jun-B. | |||||
|
JUNB_MOUSE
|
||||||
| θ value | 8.99809 (rank : 58) | NC score | 0.114781 (rank : 22) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P09450, Q8C2G9 | Gene names | Junb, Jun-b | |||
|
Domain Architecture |
|
|||||
| Description | Transcription factor jun-B. | |||||
|
MD1L1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 59) | NC score | 0.007312 (rank : 81) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 893 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Y6D9, Q13312, Q75MI0, Q86UM4, Q9UNH0 | Gene names | MAD1L1, MAD1, TXBP181 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitotic spindle assembly checkpoint protein MAD1 (Mitotic arrest deficient-like protein 1) (MAD1-like 1) (Mitotic checkpoint MAD1 protein-homolog) (HsMAD1) (hMAD1) (Tax-binding protein 181). | |||||
|
SON_MOUSE
|
||||||
| θ value | 8.99809 (rank : 60) | NC score | 0.060812 (rank : 36) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 957 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9QX47, Q9CQ12, Q9CQK6, Q9QXP5 | Gene names | Son | |||
|
Domain Architecture |

|
|||||
| Description | SON protein. | |||||
|
TARSH_HUMAN
|
||||||
| θ value | 8.99809 (rank : 61) | NC score | 0.034269 (rank : 53) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 768 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q7Z7G0, Q6ZW20, Q6ZW22, Q9C082, Q9UFI6 | Gene names | ABI3BP, NESHBP, TARSH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Target of Nesh-SH3 precursor (Tarsh) (Nesh-binding protein) (NeshBP) (ABI gene family member 3-binding protein). | |||||
|
YTHD2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 62) | NC score | 0.011599 (rank : 73) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y5A9, Q5VSZ9, Q8TDH0, Q9BUJ5 | Gene names | YTHDF2, HGRG8 | |||
|
Domain Architecture |

|
|||||
| Description | YTH domain family protein 2 (High-glucose-regulated protein 8) (NY- REN-2 antigen) (CLL-associated antigen KW-14). | |||||
|
ATF2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 63) | NC score | 0.120422 (rank : 18) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 402 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P15336, Q13000 | Gene names | ATF2, CREB2, CREBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-2 (Activating transcription factor 2) (cAMP response element-binding protein CRE- BP1) (HB16). | |||||
|
ATF2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 64) | NC score | 0.120040 (rank : 19) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P16951, Q64089, Q64090, Q64091 | Gene names | Atf2 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-2 (Activating transcription factor 2) (cAMP response element-binding protein CRE- BP1) (MXBP protein). | |||||
|
ATF6A_HUMAN
|
||||||
| θ value | θ > 10 (rank : 65) | NC score | 0.056399 (rank : 39) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P18850, O15139, Q5VW62, Q6IPB5, Q9UEC9 | Gene names | ATF6 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-6 alpha (Activating transcription factor 6 alpha) (ATF6-alpha). | |||||
|
ATF6B_MOUSE
|
||||||
| θ value | θ > 10 (rank : 66) | NC score | 0.050645 (rank : 44) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O35451 | Gene names | Crebl1 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-6 beta (Activating transcription factor 6 beta) (ATF6-beta) (cAMP-responsive element- binding protein-like 1) (cAMP response element-binding protein-related protein) (Creb-rp). | |||||
|
BATF_HUMAN
|
||||||
| θ value | θ > 10 (rank : 67) | NC score | 0.119387 (rank : 20) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q16520 | Gene names | BATF | |||
|
Domain Architecture |
|
|||||
| Description | ATF-like basic leucine zipper transcriptional factor B-ATF (SF-HT- activated gene 2) (SFA-2). | |||||
|
BATF_MOUSE
|
||||||
| θ value | θ > 10 (rank : 68) | NC score | 0.124312 (rank : 15) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O35284 | Gene names | Batf | |||
|
Domain Architecture |
|
|||||
| Description | ATF-like basic leucine zipper transcription factor B-ATF. | |||||
|
CEBPB_HUMAN
|
||||||
| θ value | θ > 10 (rank : 69) | NC score | 0.053498 (rank : 40) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P17676, Q96IH2, Q9H4Z5 | Gene names | CEBPB, TCF5 | |||
|
Domain Architecture |

|
|||||
| Description | CCAAT/enhancer-binding protein beta (C/EBP beta) (Nuclear factor NF- IL6) (Transcription factor 5). | |||||
|
CEBPG_HUMAN
|
||||||
| θ value | θ > 10 (rank : 70) | NC score | 0.051148 (rank : 43) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P53567, Q5U052 | Gene names | CEBPG | |||
|
Domain Architecture |
|
|||||
| Description | CCAAT/enhancer-binding protein gamma (C/EBP gamma). | |||||
|
CEBPG_MOUSE
|
||||||
| θ value | θ > 10 (rank : 71) | NC score | 0.053484 (rank : 41) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P53568 | Gene names | Cebpg | |||
|
Domain Architecture |

|
|||||
| Description | CCAAT/enhancer-binding protein gamma (C/EBP gamma) (Immunoglobulin enhancer-binding protein 1) (IG/EBP-1) (Granulocyte colony-stimulating factor promoter element 1-binding protein) (GPE1-binding protein) (GPE1-BP). | |||||
|
CREB3_HUMAN
|
||||||
| θ value | θ > 10 (rank : 72) | NC score | 0.066283 (rank : 34) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O43889, O14671, O14919, Q96GK8, Q9H2W3, Q9UE77 | Gene names | CREB3, LZIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclic AMP-responsive element-binding protein 3 (Luman protein) (Transcription factor LZIP-alpha). | |||||
|
CREB5_HUMAN
|
||||||
| θ value | θ > 10 (rank : 73) | NC score | 0.114200 (rank : 23) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q02930, Q05886, Q06246 | Gene names | CREB5, CREBPA | |||
|
Domain Architecture |

|
|||||
| Description | cAMP response element-binding protein 5 (CRE-BPa). | |||||
|
FOSB_HUMAN
|
||||||
| θ value | θ > 10 (rank : 74) | NC score | 0.072314 (rank : 30) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P53539 | Gene names | FOSB, G0S3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein fosB (G0/G1 switch regulatory protein 3). | |||||
|
FOSB_MOUSE
|
||||||
| θ value | θ > 10 (rank : 75) | NC score | 0.071643 (rank : 31) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P13346 | Gene names | Fosb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein fosB. | |||||
|
FOSL2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 76) | NC score | 0.062641 (rank : 35) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P15408 | Gene names | FOSL2, FRA2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fos-related antigen 2. | |||||
|
FOSL2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 77) | NC score | 0.058327 (rank : 37) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P47930 | Gene names | Fosl2, Fra-2, Fra2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fos-related antigen 2. | |||||
|
HLF_HUMAN
|
||||||
| θ value | θ > 10 (rank : 78) | NC score | 0.088249 (rank : 25) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q16534 | Gene names | HLF | |||
|
Domain Architecture |

|
|||||
| Description | Hepatic leukemia factor. | |||||
|
HLF_MOUSE
|
||||||
| θ value | θ > 10 (rank : 79) | NC score | 0.087483 (rank : 26) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BW74, Q6PF83 | Gene names | Hlf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatic leukemia factor. | |||||
|
RU1C_HUMAN
|
||||||
| θ value | θ > 10 (rank : 80) | NC score | 0.052044 (rank : 42) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P09234 | Gene names | SNRPC | |||
|
Domain Architecture |

|
|||||
| Description | U1 small nuclear ribonucleoprotein C (U1 snRNP protein C) (U1C protein) (U1-C). | |||||
|
RU1C_MOUSE
|
||||||
| θ value | θ > 10 (rank : 81) | NC score | 0.050111 (rank : 45) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q62241 | Gene names | Snrp1c, Snrpc | |||
|
Domain Architecture |

|
|||||
| Description | U1 small nuclear ribonucleoprotein C (U1 snRNP protein C) (U1C protein) (U1-C). | |||||
|
TEF_HUMAN
|
||||||
| θ value | θ > 10 (rank : 82) | NC score | 0.122777 (rank : 16) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q10587, Q15729, Q7Z3J7, Q8IU94, Q96TG4 | Gene names | TEF | |||
|
Domain Architecture |

|
|||||
| Description | Thyrotroph embryonic factor. | |||||
|
TEF_MOUSE
|
||||||
| θ value | θ > 10 (rank : 83) | NC score | 0.122271 (rank : 17) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9JLC6, Q3U426, Q3UGF4, Q3UM49, Q3URC8, Q6QHT6, Q8C6I0, Q8VD02 | Gene names | Tef | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyrotroph embryonic factor. | |||||
|
ATF5_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 8.55514e-59 (rank : 2) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 62 | |
| SwissProt Accessions | O70191, Q58E51, Q99NH6 | Gene names | Atf5, Atfx, Nap1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-5 (Activating transcription factor 5-alpha/beta) (Transcription factor ATFx) (Transcription factor-like protein ODA-10) (BZIP protein ATF7) (NRIF3- associated protein) (NAP1). | |||||
|
ATF5_HUMAN
|
||||||
| NC score | 0.908225 (rank : 2) | θ value | 1.15091e-71 (rank : 1) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 99 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | Q9Y2D1, Q9BSA1, Q9UNQ3 | Gene names | ATF5, ATFX | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-5 (Activating transcription factor 5) (Transcription factor ATFx). | |||||
|
ATF4_MOUSE
|
||||||
| NC score | 0.614334 (rank : 3) | θ value | 1.58096e-12 (rank : 3) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 102 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q06507, Q61906 | Gene names | Atf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-4 (C/EBP-related ATF) (C/ATF) (TAXREB67 homolog). | |||||
|
ATF4_HUMAN
|
||||||
| NC score | 0.605299 (rank : 4) | θ value | 4.59992e-12 (rank : 4) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 110 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P18848, Q9UH31 | Gene names | ATF4 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-4 (Activating transcription factor 4) (DNA-binding protein TAXREB67) (Cyclic AMP response element-binding protein 2) (CREB2). | |||||
|
ATF7_MOUSE
|
||||||
| NC score | 0.175514 (rank : 5) | θ value | 0.0736092 (rank : 9) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q8R0S1 | Gene names | Atf7 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-7 (Activating transcription factor 7) (Transcription factor ATF-A). | |||||
|
ATF7_HUMAN
|
||||||
| NC score | 0.173960 (rank : 6) | θ value | 0.0736092 (rank : 8) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 333 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P17544, Q13814 | Gene names | ATF7, ATFA | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-7 (Activating transcription factor 7) (Transcription factor ATF-A). | |||||
|
DBP_MOUSE
|
||||||
| NC score | 0.170838 (rank : 7) | θ value | 0.00035302 (rank : 6) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q60925, Q8VCX3 | Gene names | Dbp | |||
|
Domain Architecture |

|
|||||
| Description | D site-binding protein (Albumin D box-binding protein) (Albumin D- element-binding protein). | |||||
|
DBP_HUMAN
|
||||||
| NC score | 0.170091 (rank : 8) | θ value | 0.00035302 (rank : 5) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 136 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q10586 | Gene names | DBP | |||
|
Domain Architecture |

|
|||||
| Description | D site-binding protein (Albumin D box-binding protein) (Albumin D- element-binding protein) (TAXREB302). | |||||
|
JUN_MOUSE
|
||||||
| NC score | 0.161359 (rank : 9) | θ value | 0.0736092 (rank : 11) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 145 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P05627 | Gene names | Jun | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor AP-1 (Activator protein 1) (AP1) (Proto-oncogene c-jun) (V-jun avian sarcoma virus 17 oncogene homolog) (Jun A) (AH119). | |||||
|
JUN_HUMAN
|
||||||
| NC score | 0.161265 (rank : 10) | θ value | 0.0736092 (rank : 10) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P05412, Q96G93 | Gene names | JUN | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor AP-1 (Activator protein 1) (AP1) (Proto-oncogene c-jun) (V-jun avian sarcoma virus 17 oncogene homolog) (p39). | |||||
|
ATF3_HUMAN
|
||||||
| NC score | 0.159336 (rank : 11) | θ value | 4.03905 (rank : 34) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P18847 | Gene names | ATF3 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-3 (Activating transcription factor 3). | |||||
|
ATF3_MOUSE
|
||||||
| NC score | 0.155817 (rank : 12) | θ value | 4.03905 (rank : 35) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 120 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q60765 | Gene names | Atf3 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-3 (Activating transcription factor 3) (Transcription factor LRG-21). | |||||
|
JUND_MOUSE
|
||||||
| NC score | 0.144543 (rank : 13) | θ value | 2.36792 (rank : 29) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P15066 | Gene names | Jund, Jun-d, Jund1 | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor jun-D. | |||||
|
JUND_HUMAN
|
||||||
| NC score | 0.144417 (rank : 14) | θ value | 1.81305 (rank : 28) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P17535 | Gene names | JUND | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor jun-D. | |||||
|
BATF_MOUSE
|
||||||
| NC score | 0.124312 (rank : 15) | θ value | θ > 10 (rank : 68) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 100 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | O35284 | Gene names | Batf | |||
|
Domain Architecture |

|
|||||
| Description | ATF-like basic leucine zipper transcription factor B-ATF. | |||||
|
TEF_HUMAN
|
||||||
| NC score | 0.122777 (rank : 16) | θ value | θ > 10 (rank : 82) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q10587, Q15729, Q7Z3J7, Q8IU94, Q96TG4 | Gene names | TEF | |||
|
Domain Architecture |

|
|||||
| Description | Thyrotroph embryonic factor. | |||||
|
TEF_MOUSE
|
||||||
| NC score | 0.122271 (rank : 17) | θ value | θ > 10 (rank : 83) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 68 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9JLC6, Q3U426, Q3UGF4, Q3UM49, Q3URC8, Q6QHT6, Q8C6I0, Q8VD02 | Gene names | Tef | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyrotroph embryonic factor. | |||||
|
ATF2_HUMAN
|
||||||
| NC score | 0.120422 (rank : 18) | θ value | θ > 10 (rank : 63) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 402 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P15336, Q13000 | Gene names | ATF2, CREB2, CREBP1 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-2 (Activating transcription factor 2) (cAMP response element-binding protein CRE- BP1) (HB16). | |||||
|
ATF2_MOUSE
|
||||||
| NC score | 0.120040 (rank : 19) | θ value | θ > 10 (rank : 64) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 397 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P16951, Q64089, Q64090, Q64091 | Gene names | Atf2 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-2 (Activating transcription factor 2) (cAMP response element-binding protein CRE- BP1) (MXBP protein). | |||||
|
BATF_HUMAN
|
||||||
| NC score | 0.119387 (rank : 20) | θ value | θ > 10 (rank : 67) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q16520 | Gene names | BATF | |||
|
Domain Architecture |

|
|||||
| Description | ATF-like basic leucine zipper transcriptional factor B-ATF (SF-HT- activated gene 2) (SFA-2). | |||||
|
JUNB_HUMAN
|
||||||
| NC score | 0.116080 (rank : 21) | θ value | 8.99809 (rank : 57) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P17275, Q96GH3 | Gene names | JUNB | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor jun-B. | |||||
|
JUNB_MOUSE
|
||||||
| NC score | 0.114781 (rank : 22) | θ value | 8.99809 (rank : 58) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P09450, Q8C2G9 | Gene names | Junb, Jun-b | |||
|
Domain Architecture |

|
|||||
| Description | Transcription factor jun-B. | |||||
|
CREB5_HUMAN
|
||||||
| NC score | 0.114200 (rank : 23) | θ value | θ > 10 (rank : 73) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 222 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q02930, Q05886, Q06246 | Gene names | CREB5, CREBPA | |||
|
Domain Architecture |

|
|||||
| Description | cAMP response element-binding protein 5 (CRE-BPa). | |||||
|
CREB3_MOUSE
|
||||||
| NC score | 0.091237 (rank : 24) | θ value | 5.27518 (rank : 38) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q61817, Q99M21, Q9CVK9 | Gene names | Creb3, Lzip | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclic AMP-responsive element-binding protein 3 (Transcription factor LZIP). | |||||
|
HLF_HUMAN
|
||||||
| NC score | 0.088249 (rank : 25) | θ value | θ > 10 (rank : 78) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 75 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q16534 | Gene names | HLF | |||
|
Domain Architecture |

|
|||||
| Description | Hepatic leukemia factor. | |||||
|
HLF_MOUSE
|
||||||
| NC score | 0.087483 (rank : 26) | θ value | θ > 10 (rank : 79) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 74 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8BW74, Q6PF83 | Gene names | Hlf | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hepatic leukemia factor. | |||||
|
A4_MOUSE
|
||||||
| NC score | 0.083084 (rank : 27) | θ value | 0.0961366 (rank : 13) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 275 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P12023, P97487, P97942, Q99K32 | Gene names | App | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 protein precursor (APP) (ABPP) (Alzheimer disease amyloid protein homolog) (Amyloidogenic glycoprotein) (AG) [Contains: Soluble APP-alpha (S-APP-alpha); Soluble APP-beta (S-APP-beta); C99 (APP-C99); Beta-amyloid protein 42 (Beta-APP42); Beta-amyloid protein 40 (Beta-APP40); C83; P3(42); P3(40); Gamma-CTF(59) (Gamma-secretase C-terminal fragment 59) (Amyloid intracellular domain 59) (AID(59)) (APP-C59); Gamma-CTF(57) (Gamma-secretase C-terminal fragment 57) (Amyloid intracellular domain 57) (AID(57)) (APP-C57); Gamma-CTF(50) (Gamma-secretase C-terminal fragment 50) (Amyloid intracellular domain 50) (AID(50)); C31]. | |||||
|
SON_HUMAN
|
||||||
| NC score | 0.080253 (rank : 28) | θ value | 0.00509761 (rank : 7) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 1568 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P18583, O14487, O95981, Q14120, Q9H7B1, Q9P070, Q9P072, Q9UKP9, Q9UPY0 | Gene names | SON, C21orf50, DBP5, KIAA1019, NREBP | |||
|
Domain Architecture |

|
|||||
| Description | SON protein (SON3) (Negative regulatory element-binding protein) (NRE- binding protein) (DBP-5) (Bax antagonist selected in saccharomyces 1) (BASS1). | |||||
|
ATF6B_HUMAN
|
||||||
| NC score | 0.079382 (rank : 29) | θ value | 0.125558 (rank : 14) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 122 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q99941, Q13269, Q14343, Q14345, Q99635, Q99637, Q9H3V9, Q9H3W1, Q9NPL0 | Gene names | CREBL1, G13 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-6 beta (Activating transcription factor 6 beta) (ATF6-beta) (cAMP-responsive element- binding protein-like 1) (cAMP response element-binding protein-related protein) (Creb-rp) (Protein G13). | |||||
|
FOSB_HUMAN
|
||||||
| NC score | 0.072314 (rank : 30) | θ value | θ > 10 (rank : 74) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P53539 | Gene names | FOSB, G0S3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein fosB (G0/G1 switch regulatory protein 3). | |||||
|
FOSB_MOUSE
|
||||||
| NC score | 0.071643 (rank : 31) | θ value | θ > 10 (rank : 75) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 137 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P13346 | Gene names | Fosb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein fosB. | |||||
|
A4_HUMAN
|
||||||
| NC score | 0.069725 (rank : 32) | θ value | 0.279714 (rank : 16) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 274 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P05067, P09000, P78438, Q13764, Q13778, Q13793, Q16011, Q16014, Q16019, Q16020, Q9BT38, Q9UCA9, Q9UCB6, Q9UCC8, Q9UCD1, Q9UQ58 | Gene names | APP, A4, AD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Amyloid beta A4 protein precursor (APP) (ABPP) (Alzheimer disease amyloid protein) (Cerebral vascular amyloid peptide) (CVAP) (Protease nexin-II) (PN-II) (APPI) (PreA4) [Contains: Soluble APP-alpha (S-APP- alpha); Soluble APP-beta (S-APP-beta); C99; Beta-amyloid protein 42 (Beta-APP42); Beta-amyloid protein 40 (Beta-APP40); C83; P3(42); P3(40); Gamma-CTF(59) (Gamma-secretase C-terminal fragment 59) (Amyloid intracellular domain 59) (AID(59)); Gamma-CTF(57) (Gamma- secretase C-terminal fragment 57) (Amyloid intracellular domain 57) (AID(57)); Gamma-CTF(50) (Gamma-secretase C-terminal fragment 50) (Amyloid intracellular domain 50) (AID(50)); C31]. | |||||
|
YYAP1_HUMAN
|
||||||
| NC score | 0.066637 (rank : 33) | θ value | 0.21417 (rank : 15) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 70 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9H869, Q5VYZ4, Q5VYZ7, Q7L4C3, Q7L5E2, Q8IXA6, Q8TEW5, Q8TF04, Q96HB6, Q9BQ64, Q9NV84 | Gene names | YY1AP1, HCCA2, YY1AP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | YY1-associated protein 1 (Hepatocellular carcinoma susceptibility protein) (Hepatocellular carcinoma-associated protein 2). | |||||
|
CREB3_HUMAN
|
||||||
| NC score | 0.066283 (rank : 34) | θ value | θ > 10 (rank : 72) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 138 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | O43889, O14671, O14919, Q96GK8, Q9H2W3, Q9UE77 | Gene names | CREB3, LZIP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cyclic AMP-responsive element-binding protein 3 (Luman protein) (Transcription factor LZIP-alpha). | |||||
|
FOSL2_HUMAN
|
||||||
| NC score | 0.062641 (rank : 35) | θ value | θ > 10 (rank : 76) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P15408 | Gene names | FOSL2, FRA2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fos-related antigen 2. | |||||
|
SON_MOUSE
|
||||||
| NC score | 0.060812 (rank : 36) | θ value | 8.99809 (rank : 60) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 957 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9QX47, Q9CQ12, Q9CQK6, Q9QXP5 | Gene names | Son | |||
|
Domain Architecture |

|
|||||
| Description | SON protein. | |||||
|
FOSL2_MOUSE
|
||||||
| NC score | 0.058327 (rank : 37) | θ value | θ > 10 (rank : 77) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 151 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P47930 | Gene names | Fosl2, Fra-2, Fra2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fos-related antigen 2. | |||||
|
RNF37_HUMAN
|
||||||
| NC score | 0.057863 (rank : 38) | θ value | 0.47712 (rank : 19) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 82 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O94941 | Gene names | UBOX5, KIAA0860, RNF37, UBCE7IP5 | |||
|
Domain Architecture |

|
|||||
| Description | RING finger protein 37 (Ubiquitin-conjugating enzyme 7-interacting protein 5) (U-box domain-containing protein 5). | |||||
|
ATF6A_HUMAN
|
||||||
| NC score | 0.056399 (rank : 39) | θ value | θ > 10 (rank : 65) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 174 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P18850, O15139, Q5VW62, Q6IPB5, Q9UEC9 | Gene names | ATF6 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-6 alpha (Activating transcription factor 6 alpha) (ATF6-alpha). | |||||
|
CEBPB_HUMAN
|
||||||
| NC score | 0.053498 (rank : 40) | θ value | θ > 10 (rank : 69) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P17676, Q96IH2, Q9H4Z5 | Gene names | CEBPB, TCF5 | |||
|
Domain Architecture |

|
|||||
| Description | CCAAT/enhancer-binding protein beta (C/EBP beta) (Nuclear factor NF- IL6) (Transcription factor 5). | |||||
|
CEBPG_MOUSE
|
||||||
| NC score | 0.053484 (rank : 41) | θ value | θ > 10 (rank : 71) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P53568 | Gene names | Cebpg | |||
|
Domain Architecture |

|
|||||
| Description | CCAAT/enhancer-binding protein gamma (C/EBP gamma) (Immunoglobulin enhancer-binding protein 1) (IG/EBP-1) (Granulocyte colony-stimulating factor promoter element 1-binding protein) (GPE1-binding protein) (GPE1-BP). | |||||
|
RU1C_HUMAN
|
||||||
| NC score | 0.052044 (rank : 42) | θ value | θ > 10 (rank : 80) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 81 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P09234 | Gene names | SNRPC | |||
|
Domain Architecture |

|
|||||
| Description | U1 small nuclear ribonucleoprotein C (U1 snRNP protein C) (U1C protein) (U1-C). | |||||
|
CEBPG_HUMAN
|
||||||
| NC score | 0.051148 (rank : 43) | θ value | θ > 10 (rank : 70) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 101 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | P53567, Q5U052 | Gene names | CEBPG | |||
|
Domain Architecture |

|
|||||
| Description | CCAAT/enhancer-binding protein gamma (C/EBP gamma). | |||||
|
ATF6B_MOUSE
|
||||||
| NC score | 0.050645 (rank : 44) | θ value | θ > 10 (rank : 66) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 97 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O35451 | Gene names | Crebl1 | |||
|
Domain Architecture |

|
|||||
| Description | Cyclic AMP-dependent transcription factor ATF-6 beta (Activating transcription factor 6 beta) (ATF6-beta) (cAMP-responsive element- binding protein-like 1) (cAMP response element-binding protein-related protein) (Creb-rp). | |||||
|
RU1C_MOUSE
|
||||||
| NC score | 0.050111 (rank : 45) | θ value | θ > 10 (rank : 81) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 83 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q62241 | Gene names | Snrp1c, Snrpc | |||
|
Domain Architecture |

|
|||||
| Description | U1 small nuclear ribonucleoprotein C (U1 snRNP protein C) (U1C protein) (U1-C). | |||||
|
LARP4_MOUSE
|
||||||
| NC score | 0.048875 (rank : 46) | θ value | 0.813845 (rank : 20) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 260 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8BWW4, Q3UR69 | Gene names | Larp4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | La-related protein 4 (La ribonucleoprotein domain family member 4). | |||||
|
MKL1_MOUSE
|
||||||
| NC score | 0.043679 (rank : 47) | θ value | 1.38821 (rank : 26) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 667 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8K4J6 | Gene names | Mkl1, Bsac | |||
|
Domain Architecture |

|
|||||
| Description | MKL/myocardin-like protein 1 (Myocardin-related transcription factor A) (MRTF-A) (Megakaryoblastic leukemia 1 protein homolog) (Basic SAP coiled-coil transcription activator). | |||||
|
GON4L_HUMAN
|
||||||
| NC score | 0.042278 (rank : 48) | θ value | 8.99809 (rank : 56) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 555 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q3T8J9, Q3T8J8, Q5VYZ5, Q5W0D5, Q6AWA6, Q6P1Q6, Q7Z3L3, Q8IY79, Q9BQI1, Q9HCG6 | Gene names | GON4L, GON4, KIAA1606 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | GON-4-like protein (GON-4 homolog). | |||||
|
CRSP7_HUMAN
|
||||||
| NC score | 0.041553 (rank : 49) | θ value | 6.88961 (rank : 43) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 247 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O95402 | Gene names | CRSP7, ARC70 | |||
|
Domain Architecture |

|
|||||
| Description | CRSP complex subunit 7 (Cofactor required for Sp1 transcriptional activation subunit 7) (Transcriptional coactivator CRSP70) (Activator- recruited cofactor 70 kDa component) (ARC70). | |||||
|
IRF3_HUMAN
|
||||||
| NC score | 0.039468 (rank : 50) | θ value | 1.06291 (rank : 21) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q14653 | Gene names | IRF3 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon regulatory factor 3 (IRF-3). | |||||
|
RGS3_HUMAN
|
||||||
| NC score | 0.036515 (rank : 51) | θ value | 1.06291 (rank : 23) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 1375 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P49796, Q5VXC1, Q5VZ05, Q6ZRM5, Q8IUQ1, Q8NC47, Q8NFN4, Q8NFN5, Q8TD59, Q8TD68, Q8WXA0 | Gene names | RGS3 | |||
|
Domain Architecture |

|
|||||
| Description | Regulator of G-protein signaling 3 (RGS3) (RGP3). | |||||
|
HSF4_HUMAN
|
||||||
| NC score | 0.036432 (rank : 52) | θ value | 5.27518 (rank : 39) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9ULV5, Q99472, Q9ULV6 | Gene names | HSF4 | |||
|
Domain Architecture |

|
|||||
| Description | Heat shock factor protein 4 (HSF 4) (Heat shock transcription factor 4) (HSTF 4) (hHSF4). | |||||
|
TARSH_HUMAN
|
||||||
| NC score | 0.034269 (rank : 53) | θ value | 8.99809 (rank : 61) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 768 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q7Z7G0, Q6ZW20, Q6ZW22, Q9C082, Q9UFI6 | Gene names | ABI3BP, NESHBP, TARSH | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Target of Nesh-SH3 precursor (Tarsh) (Nesh-binding protein) (NeshBP) (ABI gene family member 3-binding protein). | |||||
|
K1802_MOUSE
|
||||||
| NC score | 0.034253 (rank : 54) | θ value | 1.38821 (rank : 25) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 1596 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q8K327, Q3UZ85, Q6ZPI1 | Gene names | Kiaa1802, D8Ertd457e | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein KIAA1802. | |||||
|
MLL3_HUMAN
|
||||||
| NC score | 0.034194 (rank : 55) | θ value | 1.38821 (rank : 27) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 1644 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8NEZ4, Q8NC02, Q8NDF6, Q9H9P4, Q9NR13, Q9P222, Q9UDR7 | Gene names | MLL3, HALR, KIAA1506 | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 3 homolog (EC 2.1.1.43) (Histone-lysine N-methyltransferase, H3 lysine-4 specific MLL3) (Homologous to ALR protein). | |||||
|
MYO15_HUMAN
|
||||||
| NC score | 0.031072 (rank : 56) | θ value | 0.0736092 (rank : 12) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 671 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9UKN7 | Gene names | MYO15A, MYO15 | |||
|
Domain Architecture |

|
|||||
| Description | Myosin-15 (Myosin XV) (Unconventional myosin-15). | |||||
|
AMPH_HUMAN
|
||||||
| NC score | 0.029820 (rank : 57) | θ value | 3.0926 (rank : 31) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 455 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P49418, O43538, Q75MJ8, Q75MK5, Q75MM3, Q8N4G0 | Gene names | AMPH, AMPH1 | |||
|
Domain Architecture |

|
|||||
| Description | Amphiphysin. | |||||
|
LPP_HUMAN
|
||||||
| NC score | 0.029788 (rank : 58) | θ value | 0.47712 (rank : 18) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 775 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q93052, Q8NFX5 | Gene names | LPP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lipoma-preferred partner (LIM domain-containing preferred translocation partner in lipoma). | |||||
|
FETUB_MOUSE
|
||||||
| NC score | 0.029649 (rank : 59) | θ value | 4.03905 (rank : 36) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 47 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9QXC1 | Gene names | Fetub | |||
|
Domain Architecture |

|
|||||
| Description | Fetuin-B precursor (IRL685). | |||||
|
GCM2_HUMAN
|
||||||
| NC score | 0.029626 (rank : 60) | θ value | 3.0926 (rank : 32) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O75603 | Gene names | GCM2, GCMB | |||
|
Domain Architecture |

|
|||||
| Description | Chorion-specific transcription factor GCMb (Glial cells missing homolog 2) (GCM motif protein 2) (hGCMb). | |||||
|
EMIL2_MOUSE
|
||||||
| NC score | 0.029372 (rank : 61) | θ value | 1.38821 (rank : 24) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 485 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q8K482 | Gene names | Emilin2 | |||
|
Domain Architecture |

|
|||||
| Description | EMILIN-2 precursor (Elastin microfibril interface-located protein 2) (Elastin microfibril interfacer 2) (Basilin). | |||||
|
SH22A_MOUSE
|
||||||
| NC score | 0.029306 (rank : 62) | θ value | 3.0926 (rank : 33) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 179 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9QXK9, Q9JHP6 | Gene names | Sh2d2a, Lad, Ribp | |||
|
Domain Architecture |

|
|||||
| Description | SH2 domain protein 2A (Lck-associated adapter protein) (Lad) (Rlk/Itk- binding protein) (Ribp). | |||||
|
GR65_MOUSE
|
||||||
| NC score | 0.025271 (rank : 63) | θ value | 4.03905 (rank : 37) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 54 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q91X51, Q9D3L9 | Gene names | Gorasp1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Golgi reassembly-stacking protein 1 (Golgi reassembly-stacking protein of 65 kDa) (GRASP65) (Golgi peripheral membrane protein p65). | |||||
|
IRF2_MOUSE
|
||||||
| NC score | 0.025106 (rank : 64) | θ value | 6.88961 (rank : 47) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P23906 | Gene names | Irf2 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon regulatory factor 2 (IRF-2). | |||||
|
NDE1_HUMAN
|
||||||
| NC score | 0.023426 (rank : 65) | θ value | 2.36792 (rank : 30) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 823 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9NXR1, Q49AQ2 | Gene names | NDE1, NUDE | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear distribution protein nudE homolog 1 (NudE). | |||||
|
CEL_HUMAN
|
||||||
| NC score | 0.022610 (rank : 66) | θ value | 8.99809 (rank : 53) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | P19835, Q16398 | Gene names | CEL, BAL | |||
|
Domain Architecture |

|
|||||
| Description | Bile salt-activated lipase precursor (EC 3.1.1.3) (EC 3.1.1.13) (BAL) (Bile salt-stimulated lipase) (BSSL) (Carboxyl ester lipase) (Sterol esterase) (Cholesterol esterase) (Pancreatic lysophospholipase). | |||||
|
CSPG5_MOUSE
|
||||||
| NC score | 0.019288 (rank : 67) | θ value | 8.99809 (rank : 54) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 211 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q71M36, Q71M37, Q7TNT8, Q8BPJ5, Q9QY32 | Gene names | Cspg5, Caleb, Ngc | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chondroitin sulfate proteoglycan 5 precursor (Neuroglycan C) (Acidic leucine-rich EGF-like domain-containing brain protein). | |||||
|
EFS_MOUSE
|
||||||
| NC score | 0.018641 (rank : 68) | θ value | 6.88961 (rank : 45) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 389 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q64355 | Gene names | Efs, Sin | |||
|
Domain Architecture |

|
|||||
| Description | Embryonal Fyn-associated substrate (SRC-interacting protein) (Signal- integrating protein). | |||||
|
ZN687_HUMAN
|
||||||
| NC score | 0.017803 (rank : 69) | θ value | 0.365318 (rank : 17) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 1244 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8N1G0, Q68DQ8, Q9H937, Q9P2A7 | Gene names | ZNF687, KIAA1441 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 687. | |||||
|
TLE4_HUMAN
|
||||||
| NC score | 0.015859 (rank : 70) | θ value | 5.27518 (rank : 40) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 477 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q04727, Q5T1Y2, Q9BZ07, Q9BZ08, Q9BZ09, Q9NSL3, Q9ULF9 | Gene names | TLE4, KIAA1261 | |||
|
Domain Architecture |

|
|||||
| Description | Transducin-like enhancer protein 4. | |||||
|
TLE4_MOUSE
|
||||||
| NC score | 0.015747 (rank : 71) | θ value | 5.27518 (rank : 41) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 482 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q62441, Q9JKQ9 | Gene names | Tle4, Grg4 | |||
|
Domain Architecture |

|
|||||
| Description | Transducin-like enhancer protein 4 (Groucho-related protein 4) (Grg- 4). | |||||
|
DOK3_HUMAN
|
||||||
| NC score | 0.013193 (rank : 72) | θ value | 8.99809 (rank : 55) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 60 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q7L591, Q8N864, Q9BQB3, Q9H666 | Gene names | DOK3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Docking protein 3 (Downstream of tyrosine kinase 3). | |||||
|
YTHD2_HUMAN
|
||||||
| NC score | 0.011599 (rank : 73) | θ value | 8.99809 (rank : 62) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 89 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Y5A9, Q5VSZ9, Q8TDH0, Q9BUJ5 | Gene names | YTHDF2, HGRG8 | |||
|
Domain Architecture |

|
|||||
| Description | YTH domain family protein 2 (High-glucose-regulated protein 8) (NY- REN-2 antigen) (CLL-associated antigen KW-14). | |||||
|
AMOL2_MOUSE
|
||||||
| NC score | 0.010648 (rank : 74) | θ value | 8.99809 (rank : 51) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 591 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8K371, Q3TPM1, Q7TPE4, Q8BP84, Q8BS08, Q9QUS0 | Gene names | Amotl2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Angiomotin-like protein 2. | |||||
|
CCD45_HUMAN
|
||||||
| NC score | 0.010381 (rank : 75) | θ value | 8.99809 (rank : 52) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 525 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96GE4, Q96M81 | Gene names | CCDC45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 45. | |||||
|
FOXH1_HUMAN
|
||||||
| NC score | 0.010034 (rank : 76) | θ value | 6.88961 (rank : 46) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 258 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | O75593 | Gene names | FOXH1, FAST1, FAST2 | |||
|
Domain Architecture |

|
|||||
| Description | Forkhead box protein H1 (Forkhead activin signal transducer 1) (Fast- 1) (hFAST-1) (Forkhead activin signal transducer 2) (Fast-2). | |||||
|
PTN18_MOUSE
|
||||||
| NC score | 0.009663 (rank : 77) | θ value | 6.88961 (rank : 49) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 134 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q61152, Q4JFH4, Q62404, Q922E3 | Gene names | Ptpn18, Flp1, Ptpk1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 18 (EC 3.1.3.48) (Fetal liver phosphatase 1) (FLP-1) (PTP-K1). | |||||
|
CE290_HUMAN
|
||||||
| NC score | 0.009373 (rank : 78) | θ value | 6.88961 (rank : 42) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 1553 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O15078, Q1PSK5, Q66GS8, Q9H2G6, Q9H6Q7, Q9H8I0 | Gene names | CEP290, KIAA0373, NPHP6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Centrosomal protein Cep290 (Nephrocystin-6) (Tumor antigen se2-2). | |||||
|
DCC_MOUSE
|
||||||
| NC score | 0.008466 (rank : 79) | θ value | 6.88961 (rank : 44) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 896 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P70211 | Gene names | Dcc | |||
|
Domain Architecture |

|
|||||
| Description | Netrin receptor DCC precursor (Tumor suppressor protein DCC). | |||||
|
PLAL1_HUMAN
|
||||||
| NC score | 0.007948 (rank : 80) | θ value | 1.06291 (rank : 22) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 728 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9UM63 | Gene names | PLAGL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein PLAGL1 (Pleiomorphic adenoma-like protein 1). | |||||
|
MD1L1_HUMAN
|
||||||
| NC score | 0.007312 (rank : 81) | θ value | 8.99809 (rank : 59) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 893 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9Y6D9, Q13312, Q75MI0, Q86UM4, Q9UNH0 | Gene names | MAD1L1, MAD1, TXBP181 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mitotic spindle assembly checkpoint protein MAD1 (Mitotic arrest deficient-like protein 1) (MAD1-like 1) (Mitotic checkpoint MAD1 protein-homolog) (HsMAD1) (hMAD1) (Tax-binding protein 181). | |||||
|
ULK1_MOUSE
|
||||||
| NC score | 0.004315 (rank : 82) | θ value | 6.88961 (rank : 50) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 1214 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | O70405, Q6PGB2 | Gene names | Ulk1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase ULK1 (EC 2.7.11.1) (Unc-51-like kinase 1) (Serine/threonine-protein kinase Unc51.1). | |||||
|
NOTC2_HUMAN
|
||||||
| NC score | 0.002142 (rank : 83) | θ value | 6.88961 (rank : 48) | |||
| Query Neighborhood Hits | 62 | Target Neighborhood Hits | 1113 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q04721, Q99734, Q9H240 | Gene names | NOTCH2 | |||
|
Domain Architecture |

|
|||||
| Description | Neurogenic locus notch homolog protein 2 precursor (Notch 2) (hN2) [Contains: Notch 2 extracellular truncation; Notch 2 intracellular domain]. | |||||