Please be patient as the page loads
|
APEX1_MOUSE
|
||||||
| SwissProt Accessions | P28352 | Gene names | Apex1, Ape, Apex | |||
|
Domain Architecture |

|
|||||
| Description | DNA-(apurinic or apyrimidinic site) lyase (EC 4.2.99.18) (AP endonuclease 1) (APEX nuclease) (APEN). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
APEX1_HUMAN
|
||||||
| θ value | 1.08164e-170 (rank : 1) | NC score | 0.997663 (rank : 2) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P27695, Q969L5, Q99775 | Gene names | APEX1, APE, APEX, APX, HAP1, REF1 | |||
|
Domain Architecture |
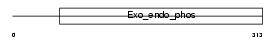
|
|||||
| Description | DNA-(apurinic or apyrimidinic site) lyase (EC 4.2.99.18) (AP endonuclease 1) (APEX nuclease) (APEN) (REF-1 protein). | |||||
|
APEX1_MOUSE
|
||||||
| θ value | 1.01238e-168 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P28352 | Gene names | Apex1, Ape, Apex | |||
|
Domain Architecture |

|
|||||
| Description | DNA-(apurinic or apyrimidinic site) lyase (EC 4.2.99.18) (AP endonuclease 1) (APEX nuclease) (APEN). | |||||
|
APEX2_MOUSE
|
||||||
| θ value | 8.10077e-17 (rank : 3) | NC score | 0.603800 (rank : 3) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q68G58, Q8BJP7, Q8BTR7, Q8BUZ2, Q8BYE9, Q8R018, Q8R328, Q9CS12 | Gene names | Apex2, Ape2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA-(apurinic or apyrimidinic site) lyase 2 (EC 4.2.99.18) (Apurinic- apyrimidinic endonuclease 2) (AP endonuclease 2) (APEX nuclease 2). | |||||
|
APEX2_HUMAN
|
||||||
| θ value | 1.99529e-15 (rank : 4) | NC score | 0.587422 (rank : 4) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UBZ4, Q9Y5X7 | Gene names | APEX2, APE2, APEXL2, XTH2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA-(apurinic or apyrimidinic site) lyase 2 (EC 4.2.99.18) (Apurinic- apyrimidinic endonuclease 2) (AP endonuclease 2) (APEX nuclease 2) (APEX nuclease-like 2) (AP endonuclease XTH2). | |||||
|
LIN1_HUMAN
|
||||||
| θ value | 1.17247e-07 (rank : 5) | NC score | 0.312475 (rank : 5) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P08547 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | LINE-1 reverse transcriptase homolog. | |||||
|
POL2_MOUSE
|
||||||
| θ value | 0.00390308 (rank : 6) | NC score | 0.237481 (rank : 6) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P11369, Q60713, Q61787 | Gene names | Pol | |||
|
Domain Architecture |

|
|||||
| Description | Retrovirus-related Pol polyprotein LINE-1 (Long interspersed element- 1) (L1) [Includes: Reverse transcriptase (EC 2.7.7.49); Endonuclease]. | |||||
|
FMN1A_MOUSE
|
||||||
| θ value | 0.125558 (rank : 7) | NC score | 0.042217 (rank : 10) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 423 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q05860 | Gene names | Fmn, Ld | |||
|
Domain Architecture |

|
|||||
| Description | Formin-1 isoforms I/II/III (Limb deformity protein). | |||||
|
ZCPW1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 8) | NC score | 0.039284 (rank : 11) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6IR42 | Gene names | Zcwpw1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CW-type PWWP domain protein 1 homolog. | |||||
|
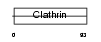
VPS41_HUMAN
|
||||||
| θ value | 2.36792 (rank : 9) | NC score | 0.062788 (rank : 8) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P49754, Q99851, Q99852 | Gene names | VPS41 | |||
|
Domain Architecture |
|
|||||
| Description | Vacuolar protein sorting-associated protein 41 homolog (S53). | |||||
|
VPS41_MOUSE
|
||||||
| θ value | 2.36792 (rank : 10) | NC score | 0.062874 (rank : 7) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5KU39, Q3TPS1, Q80V99, Q8BKK6, Q8CG01, Q8CJC3 | Gene names | Vps41 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Vacuolar protein sorting-associated protein 41 homolog (VAM2 homolog) (mVAM2). | |||||
|
TTRAP_MOUSE
|
||||||
| θ value | 4.03905 (rank : 11) | NC score | 0.029288 (rank : 12) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9JJX7 | Gene names | Ttrap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TRAF and TNF receptor-associated protein. | |||||
|
RNT2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 12) | NC score | 0.022518 (rank : 13) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O00584, Q5T8Q0, Q8TCU2, Q9BZ46, Q9BZ47 | Gene names | RNASET2, RNASE6PL | |||
|
Domain Architecture |

|
|||||
| Description | Ribonuclease T2 precursor (EC 3.1.27.-) (Ribonuclease 6). | |||||
|
TREX1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 13) | NC score | 0.015293 (rank : 14) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NSU2, Q8TEU2, Q9BPW1, Q9Y4X2 | Gene names | TREX1 | |||
|
Domain Architecture |
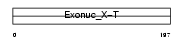
|
|||||
| Description | Three prime repair exonuclease 1 (EC 3.1.11.2) (3'-5' exonuclease TREX1) (DNase III). | |||||
|
NEIL3_MOUSE
|
||||||
| θ value | θ > 10 (rank : 14) | NC score | 0.055370 (rank : 9) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8K203, Q8CD85, Q8R3P4 | Gene names | Neil3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endonuclease VIII-like 3 (Nei-like 3) (DNA glycosylase FPG2). | |||||
|
APEX1_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 1.01238e-168 (rank : 2) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | P28352 | Gene names | Apex1, Ape, Apex | |||
|
Domain Architecture |

|
|||||
| Description | DNA-(apurinic or apyrimidinic site) lyase (EC 4.2.99.18) (AP endonuclease 1) (APEX nuclease) (APEN). | |||||
|
APEX1_HUMAN
|
||||||
| NC score | 0.997663 (rank : 2) | θ value | 1.08164e-170 (rank : 1) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P27695, Q969L5, Q99775 | Gene names | APEX1, APE, APEX, APX, HAP1, REF1 | |||
|
Domain Architecture |
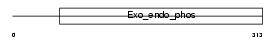
|
|||||
| Description | DNA-(apurinic or apyrimidinic site) lyase (EC 4.2.99.18) (AP endonuclease 1) (APEX nuclease) (APEN) (REF-1 protein). | |||||
|
APEX2_MOUSE
|
||||||
| NC score | 0.603800 (rank : 3) | θ value | 8.10077e-17 (rank : 3) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q68G58, Q8BJP7, Q8BTR7, Q8BUZ2, Q8BYE9, Q8R018, Q8R328, Q9CS12 | Gene names | Apex2, Ape2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA-(apurinic or apyrimidinic site) lyase 2 (EC 4.2.99.18) (Apurinic- apyrimidinic endonuclease 2) (AP endonuclease 2) (APEX nuclease 2). | |||||
|
APEX2_HUMAN
|
||||||
| NC score | 0.587422 (rank : 4) | θ value | 1.99529e-15 (rank : 4) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UBZ4, Q9Y5X7 | Gene names | APEX2, APE2, APEXL2, XTH2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DNA-(apurinic or apyrimidinic site) lyase 2 (EC 4.2.99.18) (Apurinic- apyrimidinic endonuclease 2) (AP endonuclease 2) (APEX nuclease 2) (APEX nuclease-like 2) (AP endonuclease XTH2). | |||||
|
LIN1_HUMAN
|
||||||
| NC score | 0.312475 (rank : 5) | θ value | 1.17247e-07 (rank : 5) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 187 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P08547 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | LINE-1 reverse transcriptase homolog. | |||||
|
POL2_MOUSE
|
||||||
| NC score | 0.237481 (rank : 6) | θ value | 0.00390308 (rank : 6) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P11369, Q60713, Q61787 | Gene names | Pol | |||
|
Domain Architecture |

|
|||||
| Description | Retrovirus-related Pol polyprotein LINE-1 (Long interspersed element- 1) (L1) [Includes: Reverse transcriptase (EC 2.7.7.49); Endonuclease]. | |||||
|
VPS41_MOUSE
|
||||||
| NC score | 0.062874 (rank : 7) | θ value | 2.36792 (rank : 10) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 40 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5KU39, Q3TPS1, Q80V99, Q8BKK6, Q8CG01, Q8CJC3 | Gene names | Vps41 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Vacuolar protein sorting-associated protein 41 homolog (VAM2 homolog) (mVAM2). | |||||
|
VPS41_HUMAN
|
||||||
| NC score | 0.062788 (rank : 8) | θ value | 2.36792 (rank : 9) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P49754, Q99851, Q99852 | Gene names | VPS41 | |||
|
Domain Architecture |

|
|||||
| Description | Vacuolar protein sorting-associated protein 41 homolog (S53). | |||||
|
NEIL3_MOUSE
|
||||||
| NC score | 0.055370 (rank : 9) | θ value | θ > 10 (rank : 14) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8K203, Q8CD85, Q8R3P4 | Gene names | Neil3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Endonuclease VIII-like 3 (Nei-like 3) (DNA glycosylase FPG2). | |||||
|
FMN1A_MOUSE
|
||||||
| NC score | 0.042217 (rank : 10) | θ value | 0.125558 (rank : 7) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 423 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q05860 | Gene names | Fmn, Ld | |||
|
Domain Architecture |

|
|||||
| Description | Formin-1 isoforms I/II/III (Limb deformity protein). | |||||
|
ZCPW1_MOUSE
|
||||||
| NC score | 0.039284 (rank : 11) | θ value | 1.81305 (rank : 8) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q6IR42 | Gene names | Zcwpw1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger CW-type PWWP domain protein 1 homolog. | |||||
|
TTRAP_MOUSE
|
||||||
| NC score | 0.029288 (rank : 12) | θ value | 4.03905 (rank : 11) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9JJX7 | Gene names | Ttrap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TRAF and TNF receptor-associated protein. | |||||
|
RNT2_HUMAN
|
||||||
| NC score | 0.022518 (rank : 13) | θ value | 6.88961 (rank : 12) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 6 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O00584, Q5T8Q0, Q8TCU2, Q9BZ46, Q9BZ47 | Gene names | RNASET2, RNASE6PL | |||
|
Domain Architecture |

|
|||||
| Description | Ribonuclease T2 precursor (EC 3.1.27.-) (Ribonuclease 6). | |||||
|
TREX1_HUMAN
|
||||||
| NC score | 0.015293 (rank : 14) | θ value | 8.99809 (rank : 13) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NSU2, Q8TEU2, Q9BPW1, Q9Y4X2 | Gene names | TREX1 | |||
|
Domain Architecture |
|
|||||
| Description | Three prime repair exonuclease 1 (EC 3.1.11.2) (3'-5' exonuclease TREX1) (DNase III). | |||||