Please be patient as the page loads
|
ACPH_HUMAN
|
||||||
| SwissProt Accessions | P13798, Q9BQ33, Q9P0Y2 | Gene names | APEH | |||
|
Domain Architecture |

|
|||||
| Description | Acylamino-acid-releasing enzyme (EC 3.4.19.1) (AARE) (Acyl-peptide hydrolase) (APH) (Acylaminoacyl-peptidase) (Oxidized protein hydrolase) (OPH) (DNF15S2 protein). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
ACPH_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P13798, Q9BQ33, Q9P0Y2 | Gene names | APEH | |||
|
Domain Architecture |

|
|||||
| Description | Acylamino-acid-releasing enzyme (EC 3.4.19.1) (AARE) (Acyl-peptide hydrolase) (APH) (Acylaminoacyl-peptidase) (Oxidized protein hydrolase) (OPH) (DNF15S2 protein). | |||||
|
APEH_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.993800 (rank : 2) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8R146, Q8K029, Q8R0M9 | Gene names | Apeh | |||
|
Domain Architecture |

|
|||||
| Description | Acylamino-acid-releasing enzyme (EC 3.4.19.1) (AARE) (Acyl-peptide hydrolase) (APH) (Acylaminoacyl-peptidase). | |||||
|
DPP9_MOUSE
|
||||||
| θ value | 5.43371e-13 (rank : 3) | NC score | 0.482621 (rank : 3) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8BVG4, Q6KAM9, Q8BWT9 | Gene names | Dpp9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dipeptidyl peptidase 9 (EC 3.4.14.5) (Dipeptidyl peptidase IX) (DP9) (Dipeptidyl peptidase-like protein 9) (DPLP9). | |||||
|
DPP9_HUMAN
|
||||||
| θ value | 1.58096e-12 (rank : 4) | NC score | 0.479510 (rank : 4) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q86TI2, Q6AI37, Q6UAL0, Q6ZMT2, Q6ZNJ5, Q8N2J7, Q8N3F5, Q8WXD8, Q96NT8, Q9BVR3 | Gene names | DPP9, DPRP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dipeptidyl peptidase 9 (EC 3.4.14.5) (Dipeptidyl peptidase IX) (DP9) (Dipeptidyl peptidase-like protein 9) (DPLP9) (Dipeptidyl peptidase IV-related protein 2) (DPRP-2). | |||||
|
SEPR_MOUSE
|
||||||
| θ value | 5.26297e-08 (rank : 5) | NC score | 0.389657 (rank : 7) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P97321 | Gene names | Fap | |||
|
Domain Architecture |

|
|||||
| Description | Seprase (EC 3.4.21.-) (Fibroblast activation protein alpha) (Integral membrane serine protease). | |||||
|
DPP8_HUMAN
|
||||||
| θ value | 8.97725e-08 (rank : 6) | NC score | 0.444808 (rank : 5) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q6V1X1, Q7Z4C8, Q7Z4D3, Q7Z4E1, Q8IWG7, Q8NEM5, Q96JX1, Q9HBM2, Q9HBM3, Q9HBM4, Q9HBM5, Q9NXF4 | Gene names | DPP8, DPRP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dipeptidyl peptidase 8 (EC 3.4.14.5) (Dipeptidyl peptidase VIII) (DP8) (Prolyl dipeptidase DPP8) (Dipeptidyl peptidase IV-related protein 1) (DPRP-1). | |||||
|
DPP8_MOUSE
|
||||||
| θ value | 1.99992e-07 (rank : 7) | NC score | 0.443969 (rank : 6) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q80YA7, Q9D4G6 | Gene names | Dpp8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dipeptidyl peptidase 8 (EC 3.4.14.5) (Dipeptidyl peptidase VIII) (DP8). | |||||
|
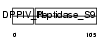
SEPR_HUMAN
|
||||||
| θ value | 9.92553e-07 (rank : 8) | NC score | 0.385857 (rank : 8) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q12884, O00199, Q86Z29, Q99998, Q9UID4 | Gene names | FAP | |||
|
Domain Architecture |
|
|||||
| Description | Seprase (EC 3.4.21.-) (Fibroblast activation protein alpha) (Integral membrane serine protease) (170 kDa melanoma membrane-bound gelatinase). | |||||
|
DPP6_HUMAN
|
||||||
| θ value | 2.21117e-06 (rank : 9) | NC score | 0.378532 (rank : 9) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P42658 | Gene names | DPP6 | |||
|
Domain Architecture |

|
|||||
| Description | Dipeptidyl aminopeptidase-like protein 6 (Dipeptidylpeptidase VI) (Dipeptidylpeptidase 6) (Dipeptidyl peptidase IV-like protein) (Dipeptidyl aminopeptidase-related protein) (DPPX). | |||||
|
DPP6_MOUSE
|
||||||
| θ value | 6.43352e-06 (rank : 10) | NC score | 0.377465 (rank : 10) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9Z218, Q9QWW2, Q9Z219 | Gene names | Dpp6, Dpp-6 | |||
|
Domain Architecture |

|
|||||
| Description | Dipeptidyl aminopeptidase-like protein 6 (Dipeptidylpeptidase VI) (Dipeptidylpeptidase 6) (Dipeptidyl peptidase IV-like protein) (Dipeptidyl aminopeptidase-related protein) (DPPX). | |||||
|
DPP4_MOUSE
|
||||||
| θ value | 0.000786445 (rank : 11) | NC score | 0.357143 (rank : 11) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P28843, Q3U514 | Gene names | Dpp4, Cd26 | |||
|
Domain Architecture |

|
|||||
| Description | Dipeptidyl peptidase 4 (EC 3.4.14.5) (Dipeptidyl peptidase IV) (DPP IV) (T-cell activation antigen CD26) (Thymocyte-activating molecule) (THAM) [Contains: Dipeptidyl peptidase 4 membrane form (Dipeptidyl peptidase IV membrane form); Dipeptidyl peptidase 4 soluble form (Dipeptidyl peptidase IV soluble form)]. | |||||
|
DPP10_MOUSE
|
||||||
| θ value | 0.0431538 (rank : 12) | NC score | 0.332011 (rank : 13) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q6NXK7, Q6P554, Q8K1H0 | Gene names | Dpp10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inactive dipeptidyl peptidase 10 (Dipeptidyl peptidase X). | |||||
|
DPP4_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 13) | NC score | 0.338581 (rank : 12) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P27487 | Gene names | DPP4, ADCP2, CD26 | |||
|
Domain Architecture |

|
|||||
| Description | Dipeptidyl peptidase 4 (EC 3.4.14.5) (Dipeptidyl peptidase IV) (DPP IV) (T-cell activation antigen CD26) (TP103) (Adenosine deaminase complexing protein 2) (ADABP) [Contains: Dipeptidyl peptidase 4 membrane form (Dipeptidyl peptidase IV membrane form); Dipeptidyl peptidase 4 soluble form (Dipeptidyl peptidase IV soluble form)]. | |||||
|
PPCE_MOUSE
|
||||||
| θ value | 0.125558 (rank : 14) | NC score | 0.151900 (rank : 15) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9QUR6, Q80YS1 | Gene names | Prep, Pep | |||
|
Domain Architecture |

|
|||||
| Description | Prolyl endopeptidase (EC 3.4.21.26) (Post-proline cleaving enzyme) (PE). | |||||
|
CT135_HUMAN
|
||||||
| θ value | 0.21417 (rank : 15) | NC score | 0.058197 (rank : 17) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9H3Z7 | Gene names | C20orf135 | |||
|
Domain Architecture |

|
|||||
| Description | Uncharacterized protein C20orf135. | |||||
|
DPP10_HUMAN
|
||||||
| θ value | 0.279714 (rank : 16) | NC score | 0.326050 (rank : 14) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8N608, Q6TTV4, Q86YR9, Q9P236 | Gene names | DPP10, DPRP3, KIAA1492 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inactive dipeptidyl peptidase 10 (Dipeptidyl peptidase X) (Dipeptidyl peptidase-like protein 2) (DPL2) (Dipeptidyl peptidase IV-related protein 3) (DPRP-3). | |||||
|
PPCE_HUMAN
|
||||||
| θ value | 0.279714 (rank : 17) | NC score | 0.145774 (rank : 16) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P48147 | Gene names | PREP, PEP | |||
|
Domain Architecture |

|
|||||
| Description | Prolyl endopeptidase (EC 3.4.21.26) (Post-proline cleaving enzyme) (PE). | |||||
|
CAC1A_HUMAN
|
||||||
| θ value | 0.365318 (rank : 18) | NC score | 0.017385 (rank : 26) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O00555, P78510, P78511, Q16290, Q92690, Q99790, Q99791, Q99792, Q99793 | Gene names | CACNA1A, CACH4, CACN3, CACNL1A4 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent P/Q-type calcium channel subunit alpha-1A (Voltage- gated calcium channel subunit alpha Cav2.1) (Calcium channel, L type, alpha-1 polypeptide isoform 4) (Brain calcium channel I) (BI). | |||||
|
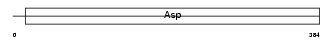
PEPA_HUMAN
|
||||||
| θ value | 1.38821 (rank : 19) | NC score | 0.018839 (rank : 25) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P00790, Q8N1E3 | Gene names | PGA3 | |||
|
Domain Architecture |
|
|||||
| Description | Pepsin A precursor (EC 3.4.23.1). | |||||
|
ATX1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 20) | NC score | 0.021948 (rank : 22) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 264 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P54254 | Gene names | Atxn1, Sca1 | |||
|
Domain Architecture |

|
|||||
| Description | Ataxin-1 (Spinocerebellar ataxia type 1 protein homolog). | |||||
|
CT135_MOUSE
|
||||||
| θ value | 5.27518 (rank : 21) | NC score | 0.035645 (rank : 18) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80YU0 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C20orf135 homolog. | |||||
|
ETFD_MOUSE
|
||||||
| θ value | 5.27518 (rank : 22) | NC score | 0.019580 (rank : 24) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q921G7, Q3U7K2, Q8BK82, Q9DCT9 | Gene names | Etfdh | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Electron transfer flavoprotein-ubiquinone oxidoreductase, mitochondrial precursor (EC 1.5.5.1) (ETF-QO) (ETF-ubiquinone oxidoreductase) (ETF dehydrogenase) (Electron-transferring- flavoprotein dehydrogenase). | |||||
|
ITIH1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 23) | NC score | 0.011597 (rank : 28) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q61702 | Gene names | Itih1 | |||
|
Domain Architecture |

|
|||||
| Description | Inter-alpha-trypsin inhibitor heavy chain H1 precursor (ITI heavy chain H1) (Inter-alpha-inhibitor heavy chain 1). | |||||
|
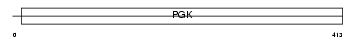
PGK2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 24) | NC score | 0.033742 (rank : 19) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P07205, Q9H107 | Gene names | PGK2, PGKB | |||
|
Domain Architecture |
|
|||||
| Description | Phosphoglycerate kinase, testis specific (EC 2.7.2.3). | |||||
|
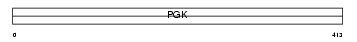
PGK2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 25) | NC score | 0.033686 (rank : 20) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P09041, Q6P8V2 | Gene names | Pgk2, Pgk-2 | |||
|
Domain Architecture |
|
|||||
| Description | Phosphoglycerate kinase, testis specific (EC 2.7.2.3). | |||||
|
PHLD2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 26) | NC score | 0.023568 (rank : 21) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q15127 | Gene names | GPLD2, PIGPLD2 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-glycan-specific phospholipase D 2 precursor (EC 3.1.4.50) (PI-G PLD) (Glycoprotein phospholipase D) (Glycosyl- phosphatidylinositol-specific phospholipase D). | |||||
|
CN029_HUMAN
|
||||||
| θ value | 8.99809 (rank : 27) | NC score | 0.017118 (rank : 27) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q7Z5M8, Q7Z5M6, Q7Z5M7, Q8N4D2 | Gene names | C14orf29 | |||
|
Domain Architecture |
|
|||||
| Description | Protein C14orf29. | |||||
|
HS12B_HUMAN
|
||||||
| θ value | 8.99809 (rank : 28) | NC score | 0.009819 (rank : 29) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96MM6, Q9BR52 | Gene names | HSPA12B, C20orf60 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heat shock 70 kDa protein 12B. | |||||
|
OR2M3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 29) | NC score | 0.000617 (rank : 30) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 780 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8NG83, Q6IEY0 | Gene names | OR2M3, OR2M6 | |||
|
Domain Architecture |
|
|||||
| Description | Olfactory receptor 2M3 (Olfactory receptor OR1-54). | |||||
|
PHLD1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 30) | NC score | 0.021468 (rank : 23) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P80108, Q15128 | Gene names | GPLD1, PIGPLD1 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-glycan-specific phospholipase D 1 precursor (EC 3.1.4.50) (PI-G PLD) (Glycoprotein phospholipase D) (Glycosyl- phosphatidylinositol-specific phospholipase D). | |||||
|
ACPH_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | P13798, Q9BQ33, Q9P0Y2 | Gene names | APEH | |||
|
Domain Architecture |

|
|||||
| Description | Acylamino-acid-releasing enzyme (EC 3.4.19.1) (AARE) (Acyl-peptide hydrolase) (APH) (Acylaminoacyl-peptidase) (Oxidized protein hydrolase) (OPH) (DNF15S2 protein). | |||||
|
APEH_MOUSE
|
||||||
| NC score | 0.993800 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8R146, Q8K029, Q8R0M9 | Gene names | Apeh | |||
|
Domain Architecture |

|
|||||
| Description | Acylamino-acid-releasing enzyme (EC 3.4.19.1) (AARE) (Acyl-peptide hydrolase) (APH) (Acylaminoacyl-peptidase). | |||||
|
DPP9_MOUSE
|
||||||
| NC score | 0.482621 (rank : 3) | θ value | 5.43371e-13 (rank : 3) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8BVG4, Q6KAM9, Q8BWT9 | Gene names | Dpp9 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dipeptidyl peptidase 9 (EC 3.4.14.5) (Dipeptidyl peptidase IX) (DP9) (Dipeptidyl peptidase-like protein 9) (DPLP9). | |||||
|
DPP9_HUMAN
|
||||||
| NC score | 0.479510 (rank : 4) | θ value | 1.58096e-12 (rank : 4) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q86TI2, Q6AI37, Q6UAL0, Q6ZMT2, Q6ZNJ5, Q8N2J7, Q8N3F5, Q8WXD8, Q96NT8, Q9BVR3 | Gene names | DPP9, DPRP2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dipeptidyl peptidase 9 (EC 3.4.14.5) (Dipeptidyl peptidase IX) (DP9) (Dipeptidyl peptidase-like protein 9) (DPLP9) (Dipeptidyl peptidase IV-related protein 2) (DPRP-2). | |||||
|
DPP8_HUMAN
|
||||||
| NC score | 0.444808 (rank : 5) | θ value | 8.97725e-08 (rank : 6) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q6V1X1, Q7Z4C8, Q7Z4D3, Q7Z4E1, Q8IWG7, Q8NEM5, Q96JX1, Q9HBM2, Q9HBM3, Q9HBM4, Q9HBM5, Q9NXF4 | Gene names | DPP8, DPRP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dipeptidyl peptidase 8 (EC 3.4.14.5) (Dipeptidyl peptidase VIII) (DP8) (Prolyl dipeptidase DPP8) (Dipeptidyl peptidase IV-related protein 1) (DPRP-1). | |||||
|
DPP8_MOUSE
|
||||||
| NC score | 0.443969 (rank : 6) | θ value | 1.99992e-07 (rank : 7) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q80YA7, Q9D4G6 | Gene names | Dpp8 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Dipeptidyl peptidase 8 (EC 3.4.14.5) (Dipeptidyl peptidase VIII) (DP8). | |||||
|
SEPR_MOUSE
|
||||||
| NC score | 0.389657 (rank : 7) | θ value | 5.26297e-08 (rank : 5) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P97321 | Gene names | Fap | |||
|
Domain Architecture |

|
|||||
| Description | Seprase (EC 3.4.21.-) (Fibroblast activation protein alpha) (Integral membrane serine protease). | |||||
|
SEPR_HUMAN
|
||||||
| NC score | 0.385857 (rank : 8) | θ value | 9.92553e-07 (rank : 8) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q12884, O00199, Q86Z29, Q99998, Q9UID4 | Gene names | FAP | |||
|
Domain Architecture |

|
|||||
| Description | Seprase (EC 3.4.21.-) (Fibroblast activation protein alpha) (Integral membrane serine protease) (170 kDa melanoma membrane-bound gelatinase). | |||||
|
DPP6_HUMAN
|
||||||
| NC score | 0.378532 (rank : 9) | θ value | 2.21117e-06 (rank : 9) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P42658 | Gene names | DPP6 | |||
|
Domain Architecture |

|
|||||
| Description | Dipeptidyl aminopeptidase-like protein 6 (Dipeptidylpeptidase VI) (Dipeptidylpeptidase 6) (Dipeptidyl peptidase IV-like protein) (Dipeptidyl aminopeptidase-related protein) (DPPX). | |||||
|
DPP6_MOUSE
|
||||||
| NC score | 0.377465 (rank : 10) | θ value | 6.43352e-06 (rank : 10) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9Z218, Q9QWW2, Q9Z219 | Gene names | Dpp6, Dpp-6 | |||
|
Domain Architecture |

|
|||||
| Description | Dipeptidyl aminopeptidase-like protein 6 (Dipeptidylpeptidase VI) (Dipeptidylpeptidase 6) (Dipeptidyl peptidase IV-like protein) (Dipeptidyl aminopeptidase-related protein) (DPPX). | |||||
|
DPP4_MOUSE
|
||||||
| NC score | 0.357143 (rank : 11) | θ value | 0.000786445 (rank : 11) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P28843, Q3U514 | Gene names | Dpp4, Cd26 | |||
|
Domain Architecture |

|
|||||
| Description | Dipeptidyl peptidase 4 (EC 3.4.14.5) (Dipeptidyl peptidase IV) (DPP IV) (T-cell activation antigen CD26) (Thymocyte-activating molecule) (THAM) [Contains: Dipeptidyl peptidase 4 membrane form (Dipeptidyl peptidase IV membrane form); Dipeptidyl peptidase 4 soluble form (Dipeptidyl peptidase IV soluble form)]. | |||||
|
DPP4_HUMAN
|
||||||
| NC score | 0.338581 (rank : 12) | θ value | 0.0563607 (rank : 13) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P27487 | Gene names | DPP4, ADCP2, CD26 | |||
|
Domain Architecture |

|
|||||
| Description | Dipeptidyl peptidase 4 (EC 3.4.14.5) (Dipeptidyl peptidase IV) (DPP IV) (T-cell activation antigen CD26) (TP103) (Adenosine deaminase complexing protein 2) (ADABP) [Contains: Dipeptidyl peptidase 4 membrane form (Dipeptidyl peptidase IV membrane form); Dipeptidyl peptidase 4 soluble form (Dipeptidyl peptidase IV soluble form)]. | |||||
|
DPP10_MOUSE
|
||||||
| NC score | 0.332011 (rank : 13) | θ value | 0.0431538 (rank : 12) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q6NXK7, Q6P554, Q8K1H0 | Gene names | Dpp10 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inactive dipeptidyl peptidase 10 (Dipeptidyl peptidase X). | |||||
|
DPP10_HUMAN
|
||||||
| NC score | 0.326050 (rank : 14) | θ value | 0.279714 (rank : 16) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q8N608, Q6TTV4, Q86YR9, Q9P236 | Gene names | DPP10, DPRP3, KIAA1492 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inactive dipeptidyl peptidase 10 (Dipeptidyl peptidase X) (Dipeptidyl peptidase-like protein 2) (DPL2) (Dipeptidyl peptidase IV-related protein 3) (DPRP-3). | |||||
|
PPCE_MOUSE
|
||||||
| NC score | 0.151900 (rank : 15) | θ value | 0.125558 (rank : 14) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q9QUR6, Q80YS1 | Gene names | Prep, Pep | |||
|
Domain Architecture |

|
|||||
| Description | Prolyl endopeptidase (EC 3.4.21.26) (Post-proline cleaving enzyme) (PE). | |||||
|
PPCE_HUMAN
|
||||||
| NC score | 0.145774 (rank : 16) | θ value | 0.279714 (rank : 17) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | P48147 | Gene names | PREP, PEP | |||
|
Domain Architecture |

|
|||||
| Description | Prolyl endopeptidase (EC 3.4.21.26) (Post-proline cleaving enzyme) (PE). | |||||
|
CT135_HUMAN
|
||||||
| NC score | 0.058197 (rank : 17) | θ value | 0.21417 (rank : 15) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9H3Z7 | Gene names | C20orf135 | |||
|
Domain Architecture |

|
|||||
| Description | Uncharacterized protein C20orf135. | |||||
|
CT135_MOUSE
|
||||||
| NC score | 0.035645 (rank : 18) | θ value | 5.27518 (rank : 21) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q80YU0 | Gene names | ||||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Uncharacterized protein C20orf135 homolog. | |||||
|
PGK2_HUMAN
|
||||||
| NC score | 0.033742 (rank : 19) | θ value | 5.27518 (rank : 24) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P07205, Q9H107 | Gene names | PGK2, PGKB | |||
|
Domain Architecture |

|
|||||
| Description | Phosphoglycerate kinase, testis specific (EC 2.7.2.3). | |||||
|
PGK2_MOUSE
|
||||||
| NC score | 0.033686 (rank : 20) | θ value | 6.88961 (rank : 25) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P09041, Q6P8V2 | Gene names | Pgk2, Pgk-2 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphoglycerate kinase, testis specific (EC 2.7.2.3). | |||||
|
PHLD2_HUMAN
|
||||||
| NC score | 0.023568 (rank : 21) | θ value | 6.88961 (rank : 26) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 44 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q15127 | Gene names | GPLD2, PIGPLD2 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-glycan-specific phospholipase D 2 precursor (EC 3.1.4.50) (PI-G PLD) (Glycoprotein phospholipase D) (Glycosyl- phosphatidylinositol-specific phospholipase D). | |||||
|
ATX1_MOUSE
|
||||||
| NC score | 0.021948 (rank : 22) | θ value | 3.0926 (rank : 20) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 264 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P54254 | Gene names | Atxn1, Sca1 | |||
|
Domain Architecture |

|
|||||
| Description | Ataxin-1 (Spinocerebellar ataxia type 1 protein homolog). | |||||
|
PHLD1_HUMAN
|
||||||
| NC score | 0.021468 (rank : 23) | θ value | 8.99809 (rank : 30) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 48 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P80108, Q15128 | Gene names | GPLD1, PIGPLD1 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol-glycan-specific phospholipase D 1 precursor (EC 3.1.4.50) (PI-G PLD) (Glycoprotein phospholipase D) (Glycosyl- phosphatidylinositol-specific phospholipase D). | |||||
|
ETFD_MOUSE
|
||||||
| NC score | 0.019580 (rank : 24) | θ value | 5.27518 (rank : 22) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q921G7, Q3U7K2, Q8BK82, Q9DCT9 | Gene names | Etfdh | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Electron transfer flavoprotein-ubiquinone oxidoreductase, mitochondrial precursor (EC 1.5.5.1) (ETF-QO) (ETF-ubiquinone oxidoreductase) (ETF dehydrogenase) (Electron-transferring- flavoprotein dehydrogenase). | |||||
|
PEPA_HUMAN
|
||||||
| NC score | 0.018839 (rank : 25) | θ value | 1.38821 (rank : 19) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P00790, Q8N1E3 | Gene names | PGA3 | |||
|
Domain Architecture |

|
|||||
| Description | Pepsin A precursor (EC 3.4.23.1). | |||||
|
CAC1A_HUMAN
|
||||||
| NC score | 0.017385 (rank : 26) | θ value | 0.365318 (rank : 18) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 387 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O00555, P78510, P78511, Q16290, Q92690, Q99790, Q99791, Q99792, Q99793 | Gene names | CACNA1A, CACH4, CACN3, CACNL1A4 | |||
|
Domain Architecture |

|
|||||
| Description | Voltage-dependent P/Q-type calcium channel subunit alpha-1A (Voltage- gated calcium channel subunit alpha Cav2.1) (Calcium channel, L type, alpha-1 polypeptide isoform 4) (Brain calcium channel I) (BI). | |||||
|
CN029_HUMAN
|
||||||
| NC score | 0.017118 (rank : 27) | θ value | 8.99809 (rank : 27) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q7Z5M8, Q7Z5M6, Q7Z5M7, Q8N4D2 | Gene names | C14orf29 | |||
|
Domain Architecture |

|
|||||
| Description | Protein C14orf29. | |||||
|
ITIH1_MOUSE
|
||||||
| NC score | 0.011597 (rank : 28) | θ value | 5.27518 (rank : 23) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q61702 | Gene names | Itih1 | |||
|
Domain Architecture |

|
|||||
| Description | Inter-alpha-trypsin inhibitor heavy chain H1 precursor (ITI heavy chain H1) (Inter-alpha-inhibitor heavy chain 1). | |||||
|
HS12B_HUMAN
|
||||||
| NC score | 0.009819 (rank : 29) | θ value | 8.99809 (rank : 28) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q96MM6, Q9BR52 | Gene names | HSPA12B, C20orf60 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Heat shock 70 kDa protein 12B. | |||||
|
OR2M3_HUMAN
|
||||||
| NC score | 0.000617 (rank : 30) | θ value | 8.99809 (rank : 29) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 780 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8NG83, Q6IEY0 | Gene names | OR2M3, OR2M6 | |||
|
Domain Architecture |

|
|||||
| Description | Olfactory receptor 2M3 (Olfactory receptor OR1-54). | |||||