Please be patient as the page loads
|
ACAD8_MOUSE
|
||||||
| SwissProt Accessions | Q9D7B6, Q8BK36, Q8BKQ8, Q8CFY9, Q9D2L8 | Gene names | Acad8 | |||
|
Domain Architecture |
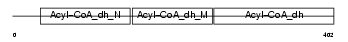
|
|||||
| Description | Acyl-CoA dehydrogenase family member 8, mitochondrial precursor (EC 1.3.99.-) (ACAD-8) (Isobutyryl-CoA dehydrogenase). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
ACAD8_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999244 (rank : 2) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9UKU7, Q9BUS8 | Gene names | ACAD8, ARC42 | |||
|
Domain Architecture |
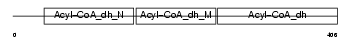
|
|||||
| Description | Acyl-CoA dehydrogenase family member 8, mitochondrial precursor (EC 1.3.99.-) (ACAD-8) (Isobutyryl-CoA dehydrogenase) (Activator- recruited cofactor 42 kDa component) (ARC42). | |||||
|
ACAD8_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9D7B6, Q8BK36, Q8BKQ8, Q8CFY9, Q9D2L8 | Gene names | Acad8 | |||
|
Domain Architecture |
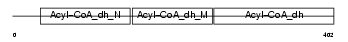
|
|||||
| Description | Acyl-CoA dehydrogenase family member 8, mitochondrial precursor (EC 1.3.99.-) (ACAD-8) (Isobutyryl-CoA dehydrogenase). | |||||
|
ACADS_MOUSE
|
||||||
| θ value | 3.4653e-68 (rank : 3) | NC score | 0.961961 (rank : 6) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q07417 | Gene names | Acads | |||
|
Domain Architecture |
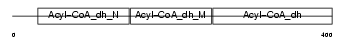
|
|||||
| Description | Short-chain specific acyl-CoA dehydrogenase, mitochondrial precursor (EC 1.3.99.2) (SCAD) (Butyryl-CoA dehydrogenase). | |||||
|
ACADS_HUMAN
|
||||||
| θ value | 5.00389e-67 (rank : 4) | NC score | 0.963679 (rank : 4) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P16219, P78331 | Gene names | ACADS | |||
|
Domain Architecture |
|
|||||
| Description | Short-chain specific acyl-CoA dehydrogenase, mitochondrial precursor (EC 1.3.99.2) (SCAD) (Butyryl-CoA dehydrogenase). | |||||
|
ACADM_MOUSE
|
||||||
| θ value | 3.71096e-62 (rank : 5) | NC score | 0.963926 (rank : 3) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P45952, Q64235 | Gene names | Acadm | |||
|
Domain Architecture |
|
|||||
| Description | Medium-chain specific acyl-CoA dehydrogenase, mitochondrial precursor (EC 1.3.99.3) (MCAD). | |||||
|
ACADM_HUMAN
|
||||||
| θ value | 4.10295e-61 (rank : 6) | NC score | 0.962542 (rank : 5) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P11310, Q9NYF1 | Gene names | ACADM | |||
|
Domain Architecture |
|
|||||
| Description | Medium-chain specific acyl-CoA dehydrogenase, mitochondrial precursor (EC 1.3.99.3) (MCAD). | |||||
|
ACDSB_HUMAN
|
||||||
| θ value | 3.59435e-57 (rank : 7) | NC score | 0.958973 (rank : 8) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P45954, Q96CX7 | Gene names | ACADSB | |||
|
Domain Architecture |
|
|||||
| Description | Short/branched chain specific acyl-CoA dehydrogenase, mitochondrial precursor (EC 1.3.99.-) (SBCAD) (2-methyl branched chain acyl-CoA dehydrogenase) (2-MEBCAD) (2-methylbutyryl-coenzyme A dehydrogenase) (2-methylbutyryl-CoA dehydrogenase). | |||||
|
IVD_MOUSE
|
||||||
| θ value | 1.15626e-55 (rank : 8) | NC score | 0.959131 (rank : 7) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9JHI5, Q9CYI3, Q9DBD7 | Gene names | Ivd | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Isovaleryl-CoA dehydrogenase, mitochondrial precursor (EC 1.3.99.10) (IVD). | |||||
|
IVD_HUMAN
|
||||||
| θ value | 1.66964e-54 (rank : 9) | NC score | 0.957460 (rank : 9) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P26440, Q96AF6 | Gene names | IVD | |||
|
Domain Architecture |
|
|||||
| Description | Isovaleryl-CoA dehydrogenase, mitochondrial precursor (EC 1.3.99.10) (IVD). | |||||
|
ACADL_HUMAN
|
||||||
| θ value | 5.93839e-52 (rank : 10) | NC score | 0.951042 (rank : 11) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P28330 | Gene names | ACADL | |||
|
Domain Architecture |
|
|||||
| Description | Long-chain specific acyl-CoA dehydrogenase, mitochondrial precursor (EC 1.3.99.13) (LCAD). | |||||
|
ACAD9_MOUSE
|
||||||
| θ value | 7.75577e-52 (rank : 11) | NC score | 0.929634 (rank : 14) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8JZN5, Q8BK76, Q8C0B5 | Gene names | Acad9 | |||
|
Domain Architecture |
|
|||||
| Description | Acyl-CoA dehydrogenase family member 9, mitochondrial precursor (EC 1.3.99.-) (ACAD-9). | |||||
|
ACDSB_MOUSE
|
||||||
| θ value | 7.75577e-52 (rank : 12) | NC score | 0.955587 (rank : 10) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9DBL1 | Gene names | Acadsb | |||
|
Domain Architecture |
|
|||||
| Description | Short/branched chain specific acyl-CoA dehydrogenase, mitochondrial precursor (EC 1.3.99.-) (SBCAD) (2-methyl branched chain acyl-CoA dehydrogenase) (2-MEBCAD) (2-methylbutyryl-coenzyme A dehydrogenase) (2-methylbutyryl-CoA dehydrogenase). | |||||
|
ACADL_MOUSE
|
||||||
| θ value | 2.49495e-50 (rank : 13) | NC score | 0.949514 (rank : 12) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P51174, O35302, Q8QZR6, Q9CU29, Q9DB83 | Gene names | Acadl | |||
|
Domain Architecture |
|
|||||
| Description | Long-chain specific acyl-CoA dehydrogenase, mitochondrial precursor (EC 1.3.99.13) (LCAD). | |||||
|
ACAD9_HUMAN
|
||||||
| θ value | 5.55816e-50 (rank : 14) | NC score | 0.929634 (rank : 13) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9H845, Q8WXX3 | Gene names | ACAD9 | |||
|
Domain Architecture |
|
|||||
| Description | Acyl-CoA dehydrogenase family member 9, mitochondrial precursor (EC 1.3.99.-) (ACAD-9). | |||||
|
ACADV_HUMAN
|
||||||
| θ value | 2.3352e-48 (rank : 15) | NC score | 0.927474 (rank : 15) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P49748, O76056, Q8WUL0 | Gene names | ACADVL, VLCAD | |||
|
Domain Architecture |
|
|||||
| Description | Very-long-chain specific acyl-CoA dehydrogenase, mitochondrial precursor (EC 1.3.99.-) (VLCAD). | |||||
|
ACADV_MOUSE
|
||||||
| θ value | 2.18568e-46 (rank : 16) | NC score | 0.922861 (rank : 16) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P50544, O35289, O55133 | Gene names | Acadvl, Vlcad | |||
|
Domain Architecture |
|
|||||
| Description | Very-long-chain specific acyl-CoA dehydrogenase, mitochondrial precursor (EC 1.3.99.-) (VLCAD) (MVLCAD). | |||||
|
GCDH_HUMAN
|
||||||
| θ value | 2.96777e-27 (rank : 17) | NC score | 0.892623 (rank : 17) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q92947, O14719 | Gene names | GCDH | |||
|
Domain Architecture |
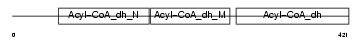
|
|||||
| Description | Glutaryl-CoA dehydrogenase, mitochondrial precursor (EC 1.3.99.7) (GCD). | |||||
|
GCDH_MOUSE
|
||||||
| θ value | 1.16704e-23 (rank : 18) | NC score | 0.882219 (rank : 18) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q60759 | Gene names | Gcdh | |||
|
Domain Architecture |
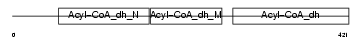
|
|||||
| Description | Glutaryl-CoA dehydrogenase, mitochondrial precursor (EC 1.3.99.7) (GCD). | |||||
|
ACOX3_HUMAN
|
||||||
| θ value | 0.000786445 (rank : 19) | NC score | 0.351919 (rank : 20) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O15254 | Gene names | ACOX3, BRCOX, PRCOX | |||
|
Domain Architecture |

|
|||||
| Description | Acyl-coenzyme A oxidase 3, peroxisomal (EC 1.3.3.6) (Pristanoyl-CoA oxidase) (Branched-chain acyl-CoA oxidase) (BRCACox). | |||||
|
ACOX3_MOUSE
|
||||||
| θ value | 0.00869519 (rank : 20) | NC score | 0.361971 (rank : 19) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9EPL9 | Gene names | Acox3 | |||
|
Domain Architecture |

|
|||||
| Description | Acyl-coenzyme A oxidase 3, peroxisomal (EC 1.3.3.6) (Pristanoyl-CoA oxidase) (Branched-chain acyl-CoA oxidase) (BRCACox). | |||||
|
P3H1_MOUSE
|
||||||
| θ value | 1.38821 (rank : 21) | NC score | 0.012410 (rank : 26) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q3V1T4, Q3TWX8, Q8BSV2, Q8CFL3, Q9CWK5, Q9QZT6, Q9QZT7 | Gene names | Lepre1, Gros1, P3h1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prolyl 3-hydroxylase 1 precursor (EC 1.14.11.7) (Leucine- and proline- enriched proteoglycan 1) (Leprecan-1) (Growth suppressor 1). | |||||
|
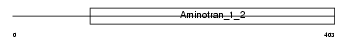
KBL_MOUSE
|
||||||
| θ value | 2.36792 (rank : 22) | NC score | 0.014548 (rank : 25) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O88986 | Gene names | Gcat, Kbl | |||
|
Domain Architecture |
|
|||||
| Description | 2-amino-3-ketobutyrate coenzyme A ligase, mitochondrial precursor (EC 2.3.1.29) (AKB ligase) (Glycine acetyltransferase). | |||||
|
ACOX2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 23) | NC score | 0.144440 (rank : 23) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q99424 | Gene names | ACOX2 | |||
|
Domain Architecture |

|
|||||
| Description | Acyl-coenzyme A oxidase 2, peroxisomal (EC 1.17.99.3) (3-alpha,7- alpha,12-alpha-trihydroxy-5-beta-cholestanoyl-CoA 24-hydroxylase) (3- alpha,7-alpha,12-alpha-trihydroxy-5-beta-cholestanoyl-CoA oxidase) (Trihydroxycoprostanoyl-CoA oxidase) (THCCox) (THCA-CoA oxidase). | |||||
|
TBAP_MOUSE
|
||||||
| θ value | 6.88961 (rank : 24) | NC score | 0.011314 (rank : 28) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q91WV0, Q3UT14 | Gene names | Dr1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TATA-binding protein-associated phosphoprotein (Down-regulator of transcription 1) (Negative cofactor 2 beta) (NC2 beta). | |||||
|
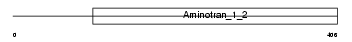
KBL_HUMAN
|
||||||
| θ value | 8.99809 (rank : 25) | NC score | 0.011490 (rank : 27) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O75600, Q96CA9 | Gene names | GCAT, KBL | |||
|
Domain Architecture |
|
|||||
| Description | 2-amino-3-ketobutyrate coenzyme A ligase, mitochondrial precursor (EC 2.3.1.29) (AKB ligase) (Glycine acetyltransferase). | |||||
|
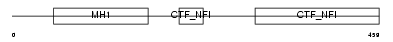
NFIA_MOUSE
|
||||||
| θ value | 8.99809 (rank : 26) | NC score | 0.004210 (rank : 29) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q02780, P70250, P70251, Q3UUZ2, Q61960, Q8VBT5, Q9R1G5 | Gene names | Nfia | |||
|
Domain Architecture |
|
|||||
| Description | Nuclear factor 1 A-type (Nuclear factor 1/A) (NF1-A) (NFI-A) (NF-I/A) (CCAAT-box-binding transcription factor) (CTF) (TGGCA-binding protein). | |||||
|
ACOX1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 27) | NC score | 0.193819 (rank : 22) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q15067, Q12863, Q15068, Q15101, Q16131, Q9UD31 | Gene names | ACOX1, ACOX | |||
|
Domain Architecture |

|
|||||
| Description | Acyl-coenzyme A oxidase 1, peroxisomal (EC 1.3.3.6) (Palmitoyl-CoA oxidase) (AOX) (Straight-chain acyl-CoA oxidase) (SCOX). | |||||
|
ACOX1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 28) | NC score | 0.195100 (rank : 21) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9R0H0, O35616, Q3TDG0 | Gene names | Acox1, Acox, Paox | |||
|
Domain Architecture |

|
|||||
| Description | Acyl-coenzyme A oxidase 1, peroxisomal (EC 1.3.3.6) (Palmitoyl-CoA oxidase) (AOX). | |||||
|
ACOX2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 29) | NC score | 0.104800 (rank : 24) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9QXD1 | Gene names | Acox2 | |||
|
Domain Architecture |

|
|||||
| Description | Acyl-coenzyme A oxidase 2, peroxisomal (EC 1.17.99.3) (3-alpha,7- alpha,12-alpha-trihydroxy-5-beta-cholestanoyl-CoA 24-hydroxylase) (3- alpha,7-alpha,12-alpha-trihydroxy-5-beta-cholestanoyl-CoA oxidase) (Trihydroxycoprostanoyl-CoA oxidase) (THCCox) (THCA-CoA oxidase). | |||||
|
ACAD8_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q9D7B6, Q8BK36, Q8BKQ8, Q8CFY9, Q9D2L8 | Gene names | Acad8 | |||
|
Domain Architecture |
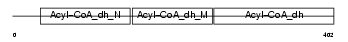
|
|||||
| Description | Acyl-CoA dehydrogenase family member 8, mitochondrial precursor (EC 1.3.99.-) (ACAD-8) (Isobutyryl-CoA dehydrogenase). | |||||
|
ACAD8_HUMAN
|
||||||
| NC score | 0.999244 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9UKU7, Q9BUS8 | Gene names | ACAD8, ARC42 | |||
|
Domain Architecture |
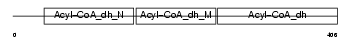
|
|||||
| Description | Acyl-CoA dehydrogenase family member 8, mitochondrial precursor (EC 1.3.99.-) (ACAD-8) (Isobutyryl-CoA dehydrogenase) (Activator- recruited cofactor 42 kDa component) (ARC42). | |||||
|
ACADM_MOUSE
|
||||||
| NC score | 0.963926 (rank : 3) | θ value | 3.71096e-62 (rank : 5) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P45952, Q64235 | Gene names | Acadm | |||
|
Domain Architecture |
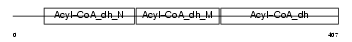
|
|||||
| Description | Medium-chain specific acyl-CoA dehydrogenase, mitochondrial precursor (EC 1.3.99.3) (MCAD). | |||||
|
ACADS_HUMAN
|
||||||
| NC score | 0.963679 (rank : 4) | θ value | 5.00389e-67 (rank : 4) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P16219, P78331 | Gene names | ACADS | |||
|
Domain Architecture |
|
|||||
| Description | Short-chain specific acyl-CoA dehydrogenase, mitochondrial precursor (EC 1.3.99.2) (SCAD) (Butyryl-CoA dehydrogenase). | |||||
|
ACADM_HUMAN
|
||||||
| NC score | 0.962542 (rank : 5) | θ value | 4.10295e-61 (rank : 6) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P11310, Q9NYF1 | Gene names | ACADM | |||
|
Domain Architecture |
|
|||||
| Description | Medium-chain specific acyl-CoA dehydrogenase, mitochondrial precursor (EC 1.3.99.3) (MCAD). | |||||
|
ACADS_MOUSE
|
||||||
| NC score | 0.961961 (rank : 6) | θ value | 3.4653e-68 (rank : 3) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q07417 | Gene names | Acads | |||
|
Domain Architecture |
|
|||||
| Description | Short-chain specific acyl-CoA dehydrogenase, mitochondrial precursor (EC 1.3.99.2) (SCAD) (Butyryl-CoA dehydrogenase). | |||||
|
IVD_MOUSE
|
||||||
| NC score | 0.959131 (rank : 7) | θ value | 1.15626e-55 (rank : 8) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9JHI5, Q9CYI3, Q9DBD7 | Gene names | Ivd | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Isovaleryl-CoA dehydrogenase, mitochondrial precursor (EC 1.3.99.10) (IVD). | |||||
|
ACDSB_HUMAN
|
||||||
| NC score | 0.958973 (rank : 8) | θ value | 3.59435e-57 (rank : 7) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P45954, Q96CX7 | Gene names | ACADSB | |||
|
Domain Architecture |
|
|||||
| Description | Short/branched chain specific acyl-CoA dehydrogenase, mitochondrial precursor (EC 1.3.99.-) (SBCAD) (2-methyl branched chain acyl-CoA dehydrogenase) (2-MEBCAD) (2-methylbutyryl-coenzyme A dehydrogenase) (2-methylbutyryl-CoA dehydrogenase). | |||||
|
IVD_HUMAN
|
||||||
| NC score | 0.957460 (rank : 9) | θ value | 1.66964e-54 (rank : 9) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P26440, Q96AF6 | Gene names | IVD | |||
|
Domain Architecture |
|
|||||
| Description | Isovaleryl-CoA dehydrogenase, mitochondrial precursor (EC 1.3.99.10) (IVD). | |||||
|
ACDSB_MOUSE
|
||||||
| NC score | 0.955587 (rank : 10) | θ value | 7.75577e-52 (rank : 12) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9DBL1 | Gene names | Acadsb | |||
|
Domain Architecture |
|
|||||
| Description | Short/branched chain specific acyl-CoA dehydrogenase, mitochondrial precursor (EC 1.3.99.-) (SBCAD) (2-methyl branched chain acyl-CoA dehydrogenase) (2-MEBCAD) (2-methylbutyryl-coenzyme A dehydrogenase) (2-methylbutyryl-CoA dehydrogenase). | |||||
|
ACADL_HUMAN
|
||||||
| NC score | 0.951042 (rank : 11) | θ value | 5.93839e-52 (rank : 10) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P28330 | Gene names | ACADL | |||
|
Domain Architecture |
|
|||||
| Description | Long-chain specific acyl-CoA dehydrogenase, mitochondrial precursor (EC 1.3.99.13) (LCAD). | |||||
|
ACADL_MOUSE
|
||||||
| NC score | 0.949514 (rank : 12) | θ value | 2.49495e-50 (rank : 13) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P51174, O35302, Q8QZR6, Q9CU29, Q9DB83 | Gene names | Acadl | |||
|
Domain Architecture |
|
|||||
| Description | Long-chain specific acyl-CoA dehydrogenase, mitochondrial precursor (EC 1.3.99.13) (LCAD). | |||||
|
ACAD9_HUMAN
|
||||||
| NC score | 0.929634 (rank : 13) | θ value | 5.55816e-50 (rank : 14) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9H845, Q8WXX3 | Gene names | ACAD9 | |||
|
Domain Architecture |
|
|||||
| Description | Acyl-CoA dehydrogenase family member 9, mitochondrial precursor (EC 1.3.99.-) (ACAD-9). | |||||
|
ACAD9_MOUSE
|
||||||
| NC score | 0.929634 (rank : 14) | θ value | 7.75577e-52 (rank : 11) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q8JZN5, Q8BK76, Q8C0B5 | Gene names | Acad9 | |||
|
Domain Architecture |
|
|||||
| Description | Acyl-CoA dehydrogenase family member 9, mitochondrial precursor (EC 1.3.99.-) (ACAD-9). | |||||
|
ACADV_HUMAN
|
||||||
| NC score | 0.927474 (rank : 15) | θ value | 2.3352e-48 (rank : 15) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P49748, O76056, Q8WUL0 | Gene names | ACADVL, VLCAD | |||
|
Domain Architecture |

|
|||||
| Description | Very-long-chain specific acyl-CoA dehydrogenase, mitochondrial precursor (EC 1.3.99.-) (VLCAD). | |||||
|
ACADV_MOUSE
|
||||||
| NC score | 0.922861 (rank : 16) | θ value | 2.18568e-46 (rank : 16) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P50544, O35289, O55133 | Gene names | Acadvl, Vlcad | |||
|
Domain Architecture |
|
|||||
| Description | Very-long-chain specific acyl-CoA dehydrogenase, mitochondrial precursor (EC 1.3.99.-) (VLCAD) (MVLCAD). | |||||
|
GCDH_HUMAN
|
||||||
| NC score | 0.892623 (rank : 17) | θ value | 2.96777e-27 (rank : 17) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q92947, O14719 | Gene names | GCDH | |||
|
Domain Architecture |
|
|||||
| Description | Glutaryl-CoA dehydrogenase, mitochondrial precursor (EC 1.3.99.7) (GCD). | |||||
|
GCDH_MOUSE
|
||||||
| NC score | 0.882219 (rank : 18) | θ value | 1.16704e-23 (rank : 18) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q60759 | Gene names | Gcdh | |||
|
Domain Architecture |
|
|||||
| Description | Glutaryl-CoA dehydrogenase, mitochondrial precursor (EC 1.3.99.7) (GCD). | |||||
|
ACOX3_MOUSE
|
||||||
| NC score | 0.361971 (rank : 19) | θ value | 0.00869519 (rank : 20) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q9EPL9 | Gene names | Acox3 | |||
|
Domain Architecture |

|
|||||
| Description | Acyl-coenzyme A oxidase 3, peroxisomal (EC 1.3.3.6) (Pristanoyl-CoA oxidase) (Branched-chain acyl-CoA oxidase) (BRCACox). | |||||
|
ACOX3_HUMAN
|
||||||
| NC score | 0.351919 (rank : 20) | θ value | 0.000786445 (rank : 19) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O15254 | Gene names | ACOX3, BRCOX, PRCOX | |||
|
Domain Architecture |

|
|||||
| Description | Acyl-coenzyme A oxidase 3, peroxisomal (EC 1.3.3.6) (Pristanoyl-CoA oxidase) (Branched-chain acyl-CoA oxidase) (BRCACox). | |||||
|
ACOX1_MOUSE
|
||||||
| NC score | 0.195100 (rank : 21) | θ value | θ > 10 (rank : 28) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q9R0H0, O35616, Q3TDG0 | Gene names | Acox1, Acox, Paox | |||
|
Domain Architecture |

|
|||||
| Description | Acyl-coenzyme A oxidase 1, peroxisomal (EC 1.3.3.6) (Palmitoyl-CoA oxidase) (AOX). | |||||
|
ACOX1_HUMAN
|
||||||
| NC score | 0.193819 (rank : 22) | θ value | θ > 10 (rank : 27) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q15067, Q12863, Q15068, Q15101, Q16131, Q9UD31 | Gene names | ACOX1, ACOX | |||
|
Domain Architecture |

|
|||||
| Description | Acyl-coenzyme A oxidase 1, peroxisomal (EC 1.3.3.6) (Palmitoyl-CoA oxidase) (AOX) (Straight-chain acyl-CoA oxidase) (SCOX). | |||||
|
ACOX2_HUMAN
|
||||||
| NC score | 0.144440 (rank : 23) | θ value | 4.03905 (rank : 23) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q99424 | Gene names | ACOX2 | |||
|
Domain Architecture |

|
|||||
| Description | Acyl-coenzyme A oxidase 2, peroxisomal (EC 1.17.99.3) (3-alpha,7- alpha,12-alpha-trihydroxy-5-beta-cholestanoyl-CoA 24-hydroxylase) (3- alpha,7-alpha,12-alpha-trihydroxy-5-beta-cholestanoyl-CoA oxidase) (Trihydroxycoprostanoyl-CoA oxidase) (THCCox) (THCA-CoA oxidase). | |||||
|
ACOX2_MOUSE
|
||||||
| NC score | 0.104800 (rank : 24) | θ value | θ > 10 (rank : 29) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9QXD1 | Gene names | Acox2 | |||
|
Domain Architecture |

|
|||||
| Description | Acyl-coenzyme A oxidase 2, peroxisomal (EC 1.17.99.3) (3-alpha,7- alpha,12-alpha-trihydroxy-5-beta-cholestanoyl-CoA 24-hydroxylase) (3- alpha,7-alpha,12-alpha-trihydroxy-5-beta-cholestanoyl-CoA oxidase) (Trihydroxycoprostanoyl-CoA oxidase) (THCCox) (THCA-CoA oxidase). | |||||
|
KBL_MOUSE
|
||||||
| NC score | 0.014548 (rank : 25) | θ value | 2.36792 (rank : 22) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O88986 | Gene names | Gcat, Kbl | |||
|
Domain Architecture |

|
|||||
| Description | 2-amino-3-ketobutyrate coenzyme A ligase, mitochondrial precursor (EC 2.3.1.29) (AKB ligase) (Glycine acetyltransferase). | |||||
|
P3H1_MOUSE
|
||||||
| NC score | 0.012410 (rank : 26) | θ value | 1.38821 (rank : 21) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q3V1T4, Q3TWX8, Q8BSV2, Q8CFL3, Q9CWK5, Q9QZT6, Q9QZT7 | Gene names | Lepre1, Gros1, P3h1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Prolyl 3-hydroxylase 1 precursor (EC 1.14.11.7) (Leucine- and proline- enriched proteoglycan 1) (Leprecan-1) (Growth suppressor 1). | |||||
|
KBL_HUMAN
|
||||||
| NC score | 0.011490 (rank : 27) | θ value | 8.99809 (rank : 25) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O75600, Q96CA9 | Gene names | GCAT, KBL | |||
|
Domain Architecture |

|
|||||
| Description | 2-amino-3-ketobutyrate coenzyme A ligase, mitochondrial precursor (EC 2.3.1.29) (AKB ligase) (Glycine acetyltransferase). | |||||
|
TBAP_MOUSE
|
||||||
| NC score | 0.011314 (rank : 28) | θ value | 6.88961 (rank : 24) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q91WV0, Q3UT14 | Gene names | Dr1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | TATA-binding protein-associated phosphoprotein (Down-regulator of transcription 1) (Negative cofactor 2 beta) (NC2 beta). | |||||
|
NFIA_MOUSE
|
||||||
| NC score | 0.004210 (rank : 29) | θ value | 8.99809 (rank : 26) | |||
| Query Neighborhood Hits | 26 | Target Neighborhood Hits | 69 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q02780, P70250, P70251, Q3UUZ2, Q61960, Q8VBT5, Q9R1G5 | Gene names | Nfia | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear factor 1 A-type (Nuclear factor 1/A) (NF1-A) (NFI-A) (NF-I/A) (CCAAT-box-binding transcription factor) (CTF) (TGGCA-binding protein). | |||||