Please be patient as the page loads
|
UGGG2_HUMAN
|
||||||
| SwissProt Accessions | Q9NYU1, Q9UFC4 | Gene names | UGCGL2, UGT2 | |||
|
Domain Architecture |

|
|||||
| Description | UDP-glucose:glycoprotein glucosyltransferase 2 precursor (EC 2.4.1.-) (UDP-glucose ceramide glucosyltransferase-like 1) (UDP-- Glc:glycoprotein glucosyltransferase 2) (HUGT2). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
UGGG1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.987845 (rank : 2) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NYU2, Q8IW30, Q9H8I4 | Gene names | UGCGL1, GT, UGGT, UGT1, UGTR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UDP-glucose:glycoprotein glucosyltransferase 1 precursor (EC 2.4.1.-) (UDP-glucose ceramide glucosyltransferase-like 1) (UDP-- Glc:glycoprotein glucosyltransferase) (HUGT1). | |||||
|
UGGG1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.987036 (rank : 3) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6P5E4, Q6NV70 | Gene names | Ugcgl1, Gt, Uggt, Ugt1, Ugtr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UDP-glucose:glycoprotein glucosyltransferase 1 precursor (EC 2.4.1.-) (UDP-glucose ceramide glucosyltransferase-like 1) (UDP-- Glc:glycoprotein glucosyltransferase). | |||||
|
UGGG2_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9NYU1, Q9UFC4 | Gene names | UGCGL2, UGT2 | |||
|
Domain Architecture |

|
|||||
| Description | UDP-glucose:glycoprotein glucosyltransferase 2 precursor (EC 2.4.1.-) (UDP-glucose ceramide glucosyltransferase-like 1) (UDP-- Glc:glycoprotein glucosyltransferase 2) (HUGT2). | |||||
|
STAT4_MOUSE
|
||||||
| θ value | 0.163984 (rank : 4) | NC score | 0.034903 (rank : 4) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P42228 | Gene names | Stat4 | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and activator of transcription 4. | |||||
|
STAT4_HUMAN
|
||||||
| θ value | 0.279714 (rank : 5) | NC score | 0.033200 (rank : 5) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14765, Q96NZ6 | Gene names | STAT4 | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and activator of transcription 4. | |||||
|
KBTB3_HUMAN
|
||||||
| θ value | 0.365318 (rank : 6) | NC score | 0.014969 (rank : 14) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8NAB2, Q86X38, Q96NK5 | Gene names | KBTBD3, BKLHD3 | |||
|
Domain Architecture |

|
|||||
| Description | Kelch repeat and BTB domain-containing protein 3 (BTB and kelch domain-containing protein 3). | |||||
|
PDCD6_MOUSE
|
||||||
| θ value | 0.813845 (rank : 7) | NC score | 0.027158 (rank : 6) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P12815, Q545I0, Q61145 | Gene names | Pdcd6, Alg2 | |||
|
Domain Architecture |
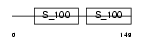
|
|||||
| Description | Programmed cell death protein 6 (Probable calcium-binding protein ALG- 2) (PMP41) (ALG-257). | |||||
|
OR5BH_HUMAN
|
||||||
| θ value | 1.81305 (rank : 8) | NC score | 0.003300 (rank : 26) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 714 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8NGF7, Q6IEX1 | Gene names | OR5B17 | |||
|
Domain Architecture |
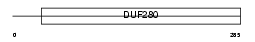
|
|||||
| Description | Olfactory receptor 5B17 (Olfactory receptor OR11-237). | |||||
|
PYGB_HUMAN
|
||||||
| θ value | 1.81305 (rank : 9) | NC score | 0.015735 (rank : 12) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P11216, Q96AK1, Q9NPX8 | Gene names | PYGB | |||
|
Domain Architecture |

|
|||||
| Description | Glycogen phosphorylase, brain form (EC 2.4.1.1). | |||||
|
UBP53_MOUSE
|
||||||
| θ value | 1.81305 (rank : 10) | NC score | 0.020537 (rank : 9) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P15975, Q8CB37, Q8R251 | Gene names | Usp53, Phxr3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inactive ubiquitin carboxyl-terminal hydrolase 53 (Inactive ubiquitin- specific peptidase 53) (Per-hexamer repeat protein 3). | |||||
|
FAKD2_HUMAN
|
||||||
| θ value | 2.36792 (rank : 11) | NC score | 0.026125 (rank : 7) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NYY8, Q9NVX6, Q9Y2H7 | Gene names | FASTKD2, KIAA0971 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FAST kinase domains-containing protein 2. | |||||
|
PDCD6_HUMAN
|
||||||
| θ value | 2.36792 (rank : 12) | NC score | 0.023136 (rank : 8) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O75340 | Gene names | PDCD6, ALG2 | |||
|
Domain Architecture |
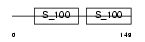
|
|||||
| Description | Programmed cell death protein 6 (Probable calcium-binding protein ALG- 2). | |||||
|
DYN1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 13) | NC score | 0.013973 (rank : 15) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P39053, Q61358, Q61359, Q61360, Q8JZZ4, Q9CSY7, Q9QXX1 | Gene names | Dnm1, Dnm | |||
|
Domain Architecture |

|
|||||
| Description | Dynamin-1 (EC 3.6.5.5). | |||||
|
HECD3_HUMAN
|
||||||
| θ value | 4.03905 (rank : 14) | NC score | 0.012107 (rank : 16) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q5T447, Q5T448, Q9H783 | Gene names | HECTD3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable E3 ubiquitin-protein ligase HECTD3 (HECT domain-containing protein 3). | |||||
|
SYNE1_HUMAN
|
||||||
| θ value | 4.03905 (rank : 15) | NC score | 0.011426 (rank : 17) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 1484 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8NF91, O94890, Q5JV19, Q5JV22, Q8N9P7, Q8TCP1, Q8WWW6, Q8WWW7, Q8WXF6, Q96N17, Q9C0A7, Q9H525, Q9H526, Q9NS36, Q9NU50, Q9UJ06, Q9UJ07, Q9ULF8 | Gene names | SYNE1, KIAA0796, KIAA1262, KIAA1756, MYNE1 | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-1 (Nuclear envelope spectrin repeat protein 1) (Synaptic nuclear envelope protein 1) (Syne-1) (Myocyte nuclear envelope protein 1) (Myne-1) (Enaptin). | |||||
|
TBCD8_MOUSE
|
||||||
| θ value | 4.03905 (rank : 16) | NC score | 0.018944 (rank : 10) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Z1A9 | Gene names | Tbc1d8, Hblp1, Vrp | |||
|
Domain Architecture |
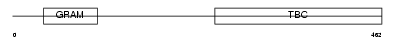
|
|||||
| Description | TBC1 domain family member 8 (Vascular Rab-GAP/TBC-containing protein) (BUB2-like protein 1). | |||||
|
CDC16_MOUSE
|
||||||
| θ value | 5.27518 (rank : 17) | NC score | 0.015796 (rank : 11) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8R349, Q9CYX9 | Gene names | Cdc16 | |||
|
Domain Architecture |
|
|||||
| Description | Cell division cycle protein 16 homolog (Anaphase-promoting complex subunit 6) (APC6) (Cyclosome subunit 6). | |||||
|
OR8J3_HUMAN
|
||||||
| θ value | 5.27518 (rank : 18) | NC score | 0.002774 (rank : 28) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 628 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8NGG0, Q96RC2 | Gene names | OR8J3 | |||
|
Domain Architecture |
|
|||||
| Description | Olfactory receptor 8J3. | |||||
|
II2A_MOUSE
|
||||||
| θ value | 6.88961 (rank : 19) | NC score | 0.010627 (rank : 20) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P15091 | Gene names | Ifi202a, Ifi202 | |||
|
Domain Architecture |
|
|||||
| Description | Interferon-activable protein 202a (Ifi-202a) (Interferon-inducible protein p202a). | |||||
|
O1038_MOUSE
|
||||||
| θ value | 6.88961 (rank : 20) | NC score | 0.002351 (rank : 30) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 748 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8VGS1 | Gene names | Olfr1038, Mor185-3 | |||
|
Domain Architecture |
|
|||||
| Description | Olfactory receptor 1038 (Olfactory receptor 185-3). | |||||
|
O1086_MOUSE
|
||||||
| θ value | 6.88961 (rank : 21) | NC score | 0.002323 (rank : 31) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8VFL9 | Gene names | Olfr1086, Mor179-2 | |||
|
Domain Architecture |
|
|||||
| Description | Olfactory receptor 1086 (Olfactory receptor 179-2). | |||||
|
OR5R1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 22) | NC score | 0.002465 (rank : 29) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 672 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8NH85 | Gene names | OR5R1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Olfactory receptor 5R1 (Olfactory receptor OR11-185). | |||||
|
AN13A_HUMAN
|
||||||
| θ value | 8.99809 (rank : 23) | NC score | 0.005745 (rank : 25) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8IZ07, O60736 | Gene names | ANKRD13A, ANKRD13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 13A (Protein KE03). | |||||
|
DESP_HUMAN
|
||||||
| θ value | 8.99809 (rank : 24) | NC score | 0.002006 (rank : 32) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P15924, O75993, Q14189, Q9UHN4 | Gene names | DSP | |||
|
Domain Architecture |

|
|||||
| Description | Desmoplakin (DP) (250/210 kDa paraneoplastic pemphigus antigen). | |||||
|
DYN1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 25) | NC score | 0.010880 (rank : 18) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q05193 | Gene names | DNM1, DNM | |||
|
Domain Architecture |

|
|||||
| Description | Dynamin-1 (EC 3.6.5.5). | |||||
|
HIP1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 26) | NC score | 0.003185 (rank : 27) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 1016 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O00291, O00328, Q8TDL4 | Gene names | HIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin-interacting protein 1 (HIP-I). | |||||
|
LARG2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 27) | NC score | 0.008247 (rank : 22) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8N3Y3, Q8N8Y6, Q8NAK3, Q8WY62 | Gene names | GYLTL1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glycosyltransferase-like protein LARGE2 (EC 2.4.-.-) (Glycosyltransferase-like 1B). | |||||
|
LCB1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 28) | NC score | 0.009223 (rank : 21) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O15269 | Gene names | SPTLC1, LCB1 | |||
|
Domain Architecture |
|
|||||
| Description | Serine palmitoyltransferase 1 (EC 2.3.1.50) (Long chain base biosynthesis protein 1) (LCB 1) (Serine-palmitoyl-CoA transferase 1) (SPT 1) (SPT1). | |||||
|
NOC3L_HUMAN
|
||||||
| θ value | 8.99809 (rank : 29) | NC score | 0.010696 (rank : 19) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8WTT2, Q9H5M6, Q9H9D8 | Gene names | NOC3L, AD24, C10orf117, FAD24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleolar complex protein 3 homolog (NOC3 protein homolog) (NOC3-like protein) (Nucleolar complex-associated protein 3-like protein) (Factor for adipocyte differentiation 24). | |||||
|
TBCD8_HUMAN
|
||||||
| θ value | 8.99809 (rank : 30) | NC score | 0.015599 (rank : 13) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O95759, Q9UQ32 | Gene names | TBC1D8, VRP | |||
|
Domain Architecture |
|
|||||
| Description | TBC1 domain family member 8 (Vascular Rab-GAP/TBC-containing protein) (AD 3). | |||||
|
U3IP2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 31) | NC score | 0.006511 (rank : 23) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O43818, Q8IZ30 | Gene names | RNU3IP2, U355K | |||
|
Domain Architecture |

|
|||||
| Description | U3 small nucleolar RNA-interacting protein 2 (U3 small nucleolar ribonucleoprotein-associated 55 kDa protein) (U3 snoRNP-associated 55 kDa protein) (U3-55K). | |||||
|
U3IP2_MOUSE
|
||||||
| θ value | 8.99809 (rank : 32) | NC score | 0.006493 (rank : 24) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q91WM3, Q8CFB7 | Gene names | Rnu3ip2, U3-55k | |||
|
Domain Architecture |

|
|||||
| Description | U3 small nucleolar RNA-interacting protein 2 (U3 small nucleolar ribonucleoprotein-associated 55 kDa protein) (U3 snoRNP-associated 55 kDa protein) (U3-55K). | |||||
|
UGGG2_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 32 | |
| SwissProt Accessions | Q9NYU1, Q9UFC4 | Gene names | UGCGL2, UGT2 | |||
|
Domain Architecture |

|
|||||
| Description | UDP-glucose:glycoprotein glucosyltransferase 2 precursor (EC 2.4.1.-) (UDP-glucose ceramide glucosyltransferase-like 1) (UDP-- Glc:glycoprotein glucosyltransferase 2) (HUGT2). | |||||
|
UGGG1_HUMAN
|
||||||
| NC score | 0.987845 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NYU2, Q8IW30, Q9H8I4 | Gene names | UGCGL1, GT, UGGT, UGT1, UGTR | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UDP-glucose:glycoprotein glucosyltransferase 1 precursor (EC 2.4.1.-) (UDP-glucose ceramide glucosyltransferase-like 1) (UDP-- Glc:glycoprotein glucosyltransferase) (HUGT1). | |||||
|
UGGG1_MOUSE
|
||||||
| NC score | 0.987036 (rank : 3) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q6P5E4, Q6NV70 | Gene names | Ugcgl1, Gt, Uggt, Ugt1, Ugtr | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | UDP-glucose:glycoprotein glucosyltransferase 1 precursor (EC 2.4.1.-) (UDP-glucose ceramide glucosyltransferase-like 1) (UDP-- Glc:glycoprotein glucosyltransferase). | |||||
|
STAT4_MOUSE
|
||||||
| NC score | 0.034903 (rank : 4) | θ value | 0.163984 (rank : 4) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P42228 | Gene names | Stat4 | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and activator of transcription 4. | |||||
|
STAT4_HUMAN
|
||||||
| NC score | 0.033200 (rank : 5) | θ value | 0.279714 (rank : 5) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 91 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q14765, Q96NZ6 | Gene names | STAT4 | |||
|
Domain Architecture |

|
|||||
| Description | Signal transducer and activator of transcription 4. | |||||
|
PDCD6_MOUSE
|
||||||
| NC score | 0.027158 (rank : 6) | θ value | 0.813845 (rank : 7) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 142 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P12815, Q545I0, Q61145 | Gene names | Pdcd6, Alg2 | |||
|
Domain Architecture |

|
|||||
| Description | Programmed cell death protein 6 (Probable calcium-binding protein ALG- 2) (PMP41) (ALG-257). | |||||
|
FAKD2_HUMAN
|
||||||
| NC score | 0.026125 (rank : 7) | θ value | 2.36792 (rank : 11) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NYY8, Q9NVX6, Q9Y2H7 | Gene names | FASTKD2, KIAA0971 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | FAST kinase domains-containing protein 2. | |||||
|
PDCD6_HUMAN
|
||||||
| NC score | 0.023136 (rank : 8) | θ value | 2.36792 (rank : 12) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 141 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O75340 | Gene names | PDCD6, ALG2 | |||
|
Domain Architecture |

|
|||||
| Description | Programmed cell death protein 6 (Probable calcium-binding protein ALG- 2). | |||||
|
UBP53_MOUSE
|
||||||
| NC score | 0.020537 (rank : 9) | θ value | 1.81305 (rank : 10) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 188 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P15975, Q8CB37, Q8R251 | Gene names | Usp53, Phxr3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Inactive ubiquitin carboxyl-terminal hydrolase 53 (Inactive ubiquitin- specific peptidase 53) (Per-hexamer repeat protein 3). | |||||
|
TBCD8_MOUSE
|
||||||
| NC score | 0.018944 (rank : 10) | θ value | 4.03905 (rank : 16) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Z1A9 | Gene names | Tbc1d8, Hblp1, Vrp | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 8 (Vascular Rab-GAP/TBC-containing protein) (BUB2-like protein 1). | |||||
|
CDC16_MOUSE
|
||||||
| NC score | 0.015796 (rank : 11) | θ value | 5.27518 (rank : 17) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8R349, Q9CYX9 | Gene names | Cdc16 | |||
|
Domain Architecture |

|
|||||
| Description | Cell division cycle protein 16 homolog (Anaphase-promoting complex subunit 6) (APC6) (Cyclosome subunit 6). | |||||
|
PYGB_HUMAN
|
||||||
| NC score | 0.015735 (rank : 12) | θ value | 1.81305 (rank : 9) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P11216, Q96AK1, Q9NPX8 | Gene names | PYGB | |||
|
Domain Architecture |

|
|||||
| Description | Glycogen phosphorylase, brain form (EC 2.4.1.1). | |||||
|
TBCD8_HUMAN
|
||||||
| NC score | 0.015599 (rank : 13) | θ value | 8.99809 (rank : 30) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 65 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O95759, Q9UQ32 | Gene names | TBC1D8, VRP | |||
|
Domain Architecture |

|
|||||
| Description | TBC1 domain family member 8 (Vascular Rab-GAP/TBC-containing protein) (AD 3). | |||||
|
KBTB3_HUMAN
|
||||||
| NC score | 0.014969 (rank : 14) | θ value | 0.365318 (rank : 6) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 176 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8NAB2, Q86X38, Q96NK5 | Gene names | KBTBD3, BKLHD3 | |||
|
Domain Architecture |

|
|||||
| Description | Kelch repeat and BTB domain-containing protein 3 (BTB and kelch domain-containing protein 3). | |||||
|
DYN1_MOUSE
|
||||||
| NC score | 0.013973 (rank : 15) | θ value | 3.0926 (rank : 13) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 168 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P39053, Q61358, Q61359, Q61360, Q8JZZ4, Q9CSY7, Q9QXX1 | Gene names | Dnm1, Dnm | |||
|
Domain Architecture |

|
|||||
| Description | Dynamin-1 (EC 3.6.5.5). | |||||
|
HECD3_HUMAN
|
||||||
| NC score | 0.012107 (rank : 16) | θ value | 4.03905 (rank : 14) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q5T447, Q5T448, Q9H783 | Gene names | HECTD3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable E3 ubiquitin-protein ligase HECTD3 (HECT domain-containing protein 3). | |||||
|
SYNE1_HUMAN
|
||||||
| NC score | 0.011426 (rank : 17) | θ value | 4.03905 (rank : 15) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 1484 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8NF91, O94890, Q5JV19, Q5JV22, Q8N9P7, Q8TCP1, Q8WWW6, Q8WWW7, Q8WXF6, Q96N17, Q9C0A7, Q9H525, Q9H526, Q9NS36, Q9NU50, Q9UJ06, Q9UJ07, Q9ULF8 | Gene names | SYNE1, KIAA0796, KIAA1262, KIAA1756, MYNE1 | |||
|
Domain Architecture |

|
|||||
| Description | Nesprin-1 (Nuclear envelope spectrin repeat protein 1) (Synaptic nuclear envelope protein 1) (Syne-1) (Myocyte nuclear envelope protein 1) (Myne-1) (Enaptin). | |||||
|
DYN1_HUMAN
|
||||||
| NC score | 0.010880 (rank : 18) | θ value | 8.99809 (rank : 25) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 216 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q05193 | Gene names | DNM1, DNM | |||
|
Domain Architecture |

|
|||||
| Description | Dynamin-1 (EC 3.6.5.5). | |||||
|
NOC3L_HUMAN
|
||||||
| NC score | 0.010696 (rank : 19) | θ value | 8.99809 (rank : 29) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 135 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8WTT2, Q9H5M6, Q9H9D8 | Gene names | NOC3L, AD24, C10orf117, FAD24 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleolar complex protein 3 homolog (NOC3 protein homolog) (NOC3-like protein) (Nucleolar complex-associated protein 3-like protein) (Factor for adipocyte differentiation 24). | |||||
|
II2A_MOUSE
|
||||||
| NC score | 0.010627 (rank : 20) | θ value | 6.88961 (rank : 19) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P15091 | Gene names | Ifi202a, Ifi202 | |||
|
Domain Architecture |

|
|||||
| Description | Interferon-activable protein 202a (Ifi-202a) (Interferon-inducible protein p202a). | |||||
|
LCB1_HUMAN
|
||||||
| NC score | 0.009223 (rank : 21) | θ value | 8.99809 (rank : 28) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O15269 | Gene names | SPTLC1, LCB1 | |||
|
Domain Architecture |

|
|||||
| Description | Serine palmitoyltransferase 1 (EC 2.3.1.50) (Long chain base biosynthesis protein 1) (LCB 1) (Serine-palmitoyl-CoA transferase 1) (SPT 1) (SPT1). | |||||
|
LARG2_HUMAN
|
||||||
| NC score | 0.008247 (rank : 22) | θ value | 8.99809 (rank : 27) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 14 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8N3Y3, Q8N8Y6, Q8NAK3, Q8WY62 | Gene names | GYLTL1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glycosyltransferase-like protein LARGE2 (EC 2.4.-.-) (Glycosyltransferase-like 1B). | |||||
|
U3IP2_HUMAN
|
||||||
| NC score | 0.006511 (rank : 23) | θ value | 8.99809 (rank : 31) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O43818, Q8IZ30 | Gene names | RNU3IP2, U355K | |||
|
Domain Architecture |

|
|||||
| Description | U3 small nucleolar RNA-interacting protein 2 (U3 small nucleolar ribonucleoprotein-associated 55 kDa protein) (U3 snoRNP-associated 55 kDa protein) (U3-55K). | |||||
|
U3IP2_MOUSE
|
||||||
| NC score | 0.006493 (rank : 24) | θ value | 8.99809 (rank : 32) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 312 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q91WM3, Q8CFB7 | Gene names | Rnu3ip2, U3-55k | |||
|
Domain Architecture |

|
|||||
| Description | U3 small nucleolar RNA-interacting protein 2 (U3 small nucleolar ribonucleoprotein-associated 55 kDa protein) (U3 snoRNP-associated 55 kDa protein) (U3-55K). | |||||
|
AN13A_HUMAN
|
||||||
| NC score | 0.005745 (rank : 25) | θ value | 8.99809 (rank : 23) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 243 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8IZ07, O60736 | Gene names | ANKRD13A, ANKRD13 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ankyrin repeat domain-containing protein 13A (Protein KE03). | |||||
|
OR5BH_HUMAN
|
||||||
| NC score | 0.003300 (rank : 26) | θ value | 1.81305 (rank : 8) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 714 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8NGF7, Q6IEX1 | Gene names | OR5B17 | |||
|
Domain Architecture |

|
|||||
| Description | Olfactory receptor 5B17 (Olfactory receptor OR11-237). | |||||
|
HIP1_HUMAN
|
||||||
| NC score | 0.003185 (rank : 27) | θ value | 8.99809 (rank : 26) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 1016 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O00291, O00328, Q8TDL4 | Gene names | HIP1 | |||
|
Domain Architecture |

|
|||||
| Description | Huntingtin-interacting protein 1 (HIP-I). | |||||
|
OR8J3_HUMAN
|
||||||
| NC score | 0.002774 (rank : 28) | θ value | 5.27518 (rank : 18) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 628 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8NGG0, Q96RC2 | Gene names | OR8J3 | |||
|
Domain Architecture |

|
|||||
| Description | Olfactory receptor 8J3. | |||||
|
OR5R1_HUMAN
|
||||||
| NC score | 0.002465 (rank : 29) | θ value | 6.88961 (rank : 22) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 672 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8NH85 | Gene names | OR5R1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Olfactory receptor 5R1 (Olfactory receptor OR11-185). | |||||
|
O1038_MOUSE
|
||||||
| NC score | 0.002351 (rank : 30) | θ value | 6.88961 (rank : 20) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 748 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8VGS1 | Gene names | Olfr1038, Mor185-3 | |||
|
Domain Architecture |

|
|||||
| Description | Olfactory receptor 1038 (Olfactory receptor 185-3). | |||||
|
O1086_MOUSE
|
||||||
| NC score | 0.002323 (rank : 31) | θ value | 6.88961 (rank : 21) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 650 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8VFL9 | Gene names | Olfr1086, Mor179-2 | |||
|
Domain Architecture |

|
|||||
| Description | Olfactory receptor 1086 (Olfactory receptor 179-2). | |||||
|
DESP_HUMAN
|
||||||
| NC score | 0.002006 (rank : 32) | θ value | 8.99809 (rank : 24) | |||
| Query Neighborhood Hits | 32 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | P15924, O75993, Q14189, Q9UHN4 | Gene names | DSP | |||
|
Domain Architecture |

|
|||||
| Description | Desmoplakin (DP) (250/210 kDa paraneoplastic pemphigus antigen). | |||||