Please be patient as the page loads
|
SDHL_MOUSE
|
||||||
| SwissProt Accessions | Q8VBT2 | Gene names | Sds | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | L-serine dehydratase (EC 4.3.1.17) (L-serine deaminase). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
SDHL_MOUSE
|
||||||
| θ value | 6.35588e-163 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8VBT2 | Gene names | Sds | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | L-serine dehydratase (EC 4.3.1.17) (L-serine deaminase). | |||||
|
SDHL_HUMAN
|
||||||
| θ value | 1.99421e-132 (rank : 2) | NC score | 0.995041 (rank : 2) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P20132 | Gene names | SDS, SDH | |||
|
Domain Architecture |
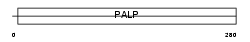
|
|||||
| Description | L-serine dehydratase (EC 4.3.1.17) (L-serine deaminase). | |||||
|
SRR_HUMAN
|
||||||
| θ value | 1.86321e-21 (rank : 3) | NC score | 0.681713 (rank : 3) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9GZT4 | Gene names | SRR | |||
|
Domain Architecture |

|
|||||
| Description | Serine racemase (EC 5.1.1.-). | |||||
|
SRR_MOUSE
|
||||||
| θ value | 3.40345e-15 (rank : 4) | NC score | 0.634584 (rank : 4) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9QZX7 | Gene names | Srr | |||
|
Domain Architecture |
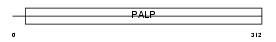
|
|||||
| Description | Serine racemase (EC 5.1.1.-). | |||||
|
CBS_HUMAN
|
||||||
| θ value | 0.813845 (rank : 5) | NC score | 0.181362 (rank : 5) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P35520, Q99425, Q9BWC5 | Gene names | CBS | |||
|
Domain Architecture |
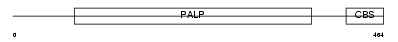
|
|||||
| Description | Cystathionine beta-synthase (EC 4.2.1.22) (Serine sulfhydrase) (Beta- thionase). | |||||
|
NPY2R_HUMAN
|
||||||
| θ value | 0.813845 (rank : 6) | NC score | 0.018329 (rank : 11) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 794 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P49146, Q13281, Q13457, Q6AZZ6, Q9UE67 | Gene names | NPY2R | |||
|
Domain Architecture |
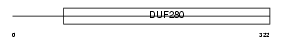
|
|||||
| Description | Neuropeptide Y receptor type 2 (NPY2-R) (NPY-Y2 receptor). | |||||
|
NPY2R_MOUSE
|
||||||
| θ value | 0.813845 (rank : 7) | NC score | 0.016694 (rank : 12) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 820 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P97295 | Gene names | Npy2r | |||
|
Domain Architecture |
|
|||||
| Description | Neuropeptide Y receptor type 2 (NPY2-R) (NPY-Y2 receptor). | |||||
|
E2F2_MOUSE
|
||||||
| θ value | 3.0926 (rank : 8) | NC score | 0.019538 (rank : 10) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P56931, Q8BID0 | Gene names | E2f2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor E2F2 (E2F-2). | |||||
|
GGTA1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 9) | NC score | 0.029648 (rank : 8) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P23336, Q91V22 | Gene names | Ggta1, Ggta-1 | |||
|
Domain Architecture |

|
|||||
| Description | N-acetyllactosaminide alpha-1,3-galactosyltransferase (EC 2.4.1.87) (Galactosyltransferase) (UDP-galactose:beta-D-galactosyl-1,4-N-acetyl- D-glucosaminide alpha-1,3-galactosyltransferase). | |||||
|
ETFD_MOUSE
|
||||||
| θ value | 4.03905 (rank : 10) | NC score | 0.035543 (rank : 6) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q921G7, Q3U7K2, Q8BK82, Q9DCT9 | Gene names | Etfdh | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Electron transfer flavoprotein-ubiquinone oxidoreductase, mitochondrial precursor (EC 1.5.5.1) (ETF-QO) (ETF-ubiquinone oxidoreductase) (ETF dehydrogenase) (Electron-transferring- flavoprotein dehydrogenase). | |||||
|
CHX10_HUMAN
|
||||||
| θ value | 8.99809 (rank : 11) | NC score | 0.004788 (rank : 13) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 340 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P58304 | Gene names | CHX10, VSX2 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein CHX10 (Ceh-10 homeodomain-containing homolog). | |||||
|
ETFD_HUMAN
|
||||||
| θ value | 8.99809 (rank : 12) | NC score | 0.030668 (rank : 7) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q16134, Q7Z347 | Gene names | ETFDH | |||
|
Domain Architecture |

|
|||||
| Description | Electron transfer flavoprotein-ubiquinone oxidoreductase, mitochondrial precursor (EC 1.5.5.1) (ETF-QO) (ETF-ubiquinone oxidoreductase) (ETF dehydrogenase) (Electron-transferring- flavoprotein dehydrogenase). | |||||
|
RPC5_HUMAN
|
||||||
| θ value | 8.99809 (rank : 13) | NC score | 0.022051 (rank : 9) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NVU0, Q9BWF7, Q9H8W8, Q9H907, Q9P276 | Gene names | POLR3E, KIAA1452 | |||
|
Domain Architecture |
|
|||||
| Description | DNA-directed RNA polymerases III 80 kDa polypeptide (EC 2.7.7.6) (RNA polymerase III subunit 5) (RPC5). | |||||
|
SDHL_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 6.35588e-163 (rank : 1) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8VBT2 | Gene names | Sds | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | L-serine dehydratase (EC 4.3.1.17) (L-serine deaminase). | |||||
|
SDHL_HUMAN
|
||||||
| NC score | 0.995041 (rank : 2) | θ value | 1.99421e-132 (rank : 2) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P20132 | Gene names | SDS, SDH | |||
|
Domain Architecture |

|
|||||
| Description | L-serine dehydratase (EC 4.3.1.17) (L-serine deaminase). | |||||
|
SRR_HUMAN
|
||||||
| NC score | 0.681713 (rank : 3) | θ value | 1.86321e-21 (rank : 3) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9GZT4 | Gene names | SRR | |||
|
Domain Architecture |

|
|||||
| Description | Serine racemase (EC 5.1.1.-). | |||||
|
SRR_MOUSE
|
||||||
| NC score | 0.634584 (rank : 4) | θ value | 3.40345e-15 (rank : 4) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9QZX7 | Gene names | Srr | |||
|
Domain Architecture |

|
|||||
| Description | Serine racemase (EC 5.1.1.-). | |||||
|
CBS_HUMAN
|
||||||
| NC score | 0.181362 (rank : 5) | θ value | 0.813845 (rank : 5) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P35520, Q99425, Q9BWC5 | Gene names | CBS | |||
|
Domain Architecture |

|
|||||
| Description | Cystathionine beta-synthase (EC 4.2.1.22) (Serine sulfhydrase) (Beta- thionase). | |||||
|
ETFD_MOUSE
|
||||||
| NC score | 0.035543 (rank : 6) | θ value | 4.03905 (rank : 10) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q921G7, Q3U7K2, Q8BK82, Q9DCT9 | Gene names | Etfdh | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Electron transfer flavoprotein-ubiquinone oxidoreductase, mitochondrial precursor (EC 1.5.5.1) (ETF-QO) (ETF-ubiquinone oxidoreductase) (ETF dehydrogenase) (Electron-transferring- flavoprotein dehydrogenase). | |||||
|
ETFD_HUMAN
|
||||||
| NC score | 0.030668 (rank : 7) | θ value | 8.99809 (rank : 12) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q16134, Q7Z347 | Gene names | ETFDH | |||
|
Domain Architecture |

|
|||||
| Description | Electron transfer flavoprotein-ubiquinone oxidoreductase, mitochondrial precursor (EC 1.5.5.1) (ETF-QO) (ETF-ubiquinone oxidoreductase) (ETF dehydrogenase) (Electron-transferring- flavoprotein dehydrogenase). | |||||
|
GGTA1_MOUSE
|
||||||
| NC score | 0.029648 (rank : 8) | θ value | 3.0926 (rank : 9) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 12 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P23336, Q91V22 | Gene names | Ggta1, Ggta-1 | |||
|
Domain Architecture |

|
|||||
| Description | N-acetyllactosaminide alpha-1,3-galactosyltransferase (EC 2.4.1.87) (Galactosyltransferase) (UDP-galactose:beta-D-galactosyl-1,4-N-acetyl- D-glucosaminide alpha-1,3-galactosyltransferase). | |||||
|
RPC5_HUMAN
|
||||||
| NC score | 0.022051 (rank : 9) | θ value | 8.99809 (rank : 13) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9NVU0, Q9BWF7, Q9H8W8, Q9H907, Q9P276 | Gene names | POLR3E, KIAA1452 | |||
|
Domain Architecture |

|
|||||
| Description | DNA-directed RNA polymerases III 80 kDa polypeptide (EC 2.7.7.6) (RNA polymerase III subunit 5) (RPC5). | |||||
|
E2F2_MOUSE
|
||||||
| NC score | 0.019538 (rank : 10) | θ value | 3.0926 (rank : 8) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 42 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P56931, Q8BID0 | Gene names | E2f2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription factor E2F2 (E2F-2). | |||||
|
NPY2R_HUMAN
|
||||||
| NC score | 0.018329 (rank : 11) | θ value | 0.813845 (rank : 6) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 794 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P49146, Q13281, Q13457, Q6AZZ6, Q9UE67 | Gene names | NPY2R | |||
|
Domain Architecture |

|
|||||
| Description | Neuropeptide Y receptor type 2 (NPY2-R) (NPY-Y2 receptor). | |||||
|
NPY2R_MOUSE
|
||||||
| NC score | 0.016694 (rank : 12) | θ value | 0.813845 (rank : 7) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 820 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P97295 | Gene names | Npy2r | |||
|
Domain Architecture |

|
|||||
| Description | Neuropeptide Y receptor type 2 (NPY2-R) (NPY-Y2 receptor). | |||||
|
CHX10_HUMAN
|
||||||
| NC score | 0.004788 (rank : 13) | θ value | 8.99809 (rank : 11) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 340 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P58304 | Gene names | CHX10, VSX2 | |||
|
Domain Architecture |

|
|||||
| Description | Homeobox protein CHX10 (Ceh-10 homeodomain-containing homolog). | |||||