Please be patient as the page loads
|
CBS_HUMAN
|
||||||
| SwissProt Accessions | P35520, Q99425, Q9BWC5 | Gene names | CBS | |||
|
Domain Architecture |
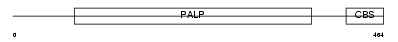
|
|||||
| Description | Cystathionine beta-synthase (EC 4.2.1.22) (Serine sulfhydrase) (Beta- thionase). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
CBS_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P35520, Q99425, Q9BWC5 | Gene names | CBS | |||
|
Domain Architecture |

|
|||||
| Description | Cystathionine beta-synthase (EC 4.2.1.22) (Serine sulfhydrase) (Beta- thionase). | |||||
|
SRR_HUMAN
|
||||||
| θ value | 2.98157e-11 (rank : 2) | NC score | 0.471098 (rank : 3) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9GZT4 | Gene names | SRR | |||
|
Domain Architecture |

|
|||||
| Description | Serine racemase (EC 5.1.1.-). | |||||
|
SRR_MOUSE
|
||||||
| θ value | 6.64225e-11 (rank : 3) | NC score | 0.478050 (rank : 2) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9QZX7 | Gene names | Srr | |||
|
Domain Architecture |
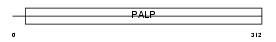
|
|||||
| Description | Serine racemase (EC 5.1.1.-). | |||||
|
PER3_HUMAN
|
||||||
| θ value | 0.21417 (rank : 4) | NC score | 0.048698 (rank : 8) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P56645, Q5H8X4, Q969K6, Q96S77, Q96S78, Q9C0J3, Q9NSP9, Q9UGU8 | Gene names | PER3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (hPER3). | |||||
|
CLCN6_MOUSE
|
||||||
| θ value | 0.47712 (rank : 5) | NC score | 0.060377 (rank : 6) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O35454 | Gene names | Clcn6, Clc6 | |||
|
Domain Architecture |

|
|||||
| Description | Chloride channel protein 6 (ClC-6). | |||||
|
SDHL_MOUSE
|
||||||
| θ value | 0.813845 (rank : 6) | NC score | 0.181362 (rank : 4) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8VBT2 | Gene names | Sds | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | L-serine dehydratase (EC 4.3.1.17) (L-serine deaminase). | |||||
|
CLCN6_HUMAN
|
||||||
| θ value | 1.06291 (rank : 7) | NC score | 0.055973 (rank : 7) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P51797, O60818, O60819, O60820, O60821, P78520, P78521, Q5SNW2, Q5SNW3, Q5SNX1, Q5SNX2, Q5SNX3, Q99427, Q99428, Q99429 | Gene names | CLCN6, KIAA0046 | |||
|
Domain Architecture |

|
|||||
| Description | Chloride channel protein 6 (ClC-6). | |||||
|
TITIN_HUMAN
|
||||||
| θ value | 1.38821 (rank : 8) | NC score | 0.009499 (rank : 20) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 2923 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8WZ42, Q10465, Q10466, Q15598, Q2XUS3, Q32Q60, Q4U1Z6, Q6NSG0, Q6PDB1, Q6PJP0, Q7KYM2, Q7KYN4, Q7KYN5, Q7LDM3, Q7Z2X3, Q8TCG8, Q8WZ51, Q8WZ52, Q8WZ53, Q8WZB3, Q92761, Q92762, Q9UD97, Q9UP84, Q9Y6L9 | Gene names | TTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Titin (EC 2.7.11.1) (Connectin) (Rhabdomyosarcoma antigen MU-RMS- 40.14). | |||||
|
GSCR1_HUMAN
|
||||||
| θ value | 1.81305 (rank : 9) | NC score | 0.036294 (rank : 9) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 674 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NZM4 | Gene names | GLTSCR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glioma tumor suppressor candidate region gene 1 protein. | |||||
|
EMID2_HUMAN
|
||||||
| θ value | 3.0926 (rank : 10) | NC score | 0.020550 (rank : 16) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 226 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96A83 | Gene names | EMID2, COL26A1, EMU2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XXVI) chain precursor (EMI domain-containing protein 2) (Protein Emu2) (Emilin and multimerin domain-containing protein 2). | |||||
|
PTN12_MOUSE
|
||||||
| θ value | 3.0926 (rank : 11) | NC score | 0.013153 (rank : 18) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P35831 | Gene names | Ptpn12 | |||
|
Domain Architecture |
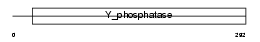
|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 12 (EC 3.1.3.48) (Protein-tyrosine phosphatase P19) (P19-PTP) (MPTP-PEST). | |||||
|
STAC3_MOUSE
|
||||||
| θ value | 3.0926 (rank : 12) | NC score | 0.023212 (rank : 13) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BZ71 | Gene names | Stac3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH3 and cysteine-rich domain-containing protein 3. | |||||
|
MSH5_HUMAN
|
||||||
| θ value | 4.03905 (rank : 13) | NC score | 0.036123 (rank : 10) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O43196, O60586, Q5BLU9, Q5SSR1, Q8IW44, Q9BQC7 | Gene names | MSH5 | |||
|
Domain Architecture |

|
|||||
| Description | MutS protein homolog 5. | |||||
|
TCOF_HUMAN
|
||||||
| θ value | 4.03905 (rank : 14) | NC score | 0.028955 (rank : 12) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q13428, Q99408, Q99860 | Gene names | TCOF1 | |||
|
Domain Architecture |

|
|||||
| Description | Treacle protein (Treacher Collins syndrome protein). | |||||
|
MSH5_MOUSE
|
||||||
| θ value | 5.27518 (rank : 15) | NC score | 0.034486 (rank : 11) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9QUM7, Q9Z1Q6 | Gene names | Msh5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MutS protein homolog 5. | |||||
|
ZN696_HUMAN
|
||||||
| θ value | 5.27518 (rank : 16) | NC score | 0.001172 (rank : 22) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 743 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9H7X3 | Gene names | ZNF696 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 696. | |||||
|
BHLH3_HUMAN
|
||||||
| θ value | 6.88961 (rank : 17) | NC score | 0.021491 (rank : 15) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9C0J9 | Gene names | BHLHB3, DEC2, SHARP1 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 3 (bHLHB3) (Differentially expressed in chondrocytes protein 2) (hDEC2) (Enhancer-of-split and hairy-related protein 1) (SHARP-1). | |||||
|
CO5A1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 18) | NC score | 0.009206 (rank : 21) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 434 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P20908, Q15094, Q5SUX4 | Gene names | COL5A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(V) chain precursor. | |||||
|
NKTR_HUMAN
|
||||||
| θ value | 6.88961 (rank : 19) | NC score | 0.011046 (rank : 19) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 658 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P30414 | Gene names | NKTR | |||
|
Domain Architecture |

|
|||||
| Description | NK-tumor recognition protein (Natural-killer cells cyclophilin-related protein) (NK-TR protein). | |||||
|
RIN2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 20) | NC score | 0.018075 (rank : 17) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8WYP3, Q00425, Q5TFT8, Q9BQL3, Q9H071 | Gene names | RIN2, RASSF4 | |||
|
Domain Architecture |

|
|||||
| Description | Ras and Rab interactor 2 (Ras interaction/interference protein 2) (Ras inhibitor JC265) (Ras association domain family 4). | |||||
|
LAG3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 21) | NC score | 0.022514 (rank : 14) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P18627 | Gene names | LAG3, FDC | |||
|
Domain Architecture |
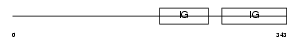
|
|||||
| Description | Lymphocyte activation gene 3 protein precursor (LAG-3) (FDC protein) (CD223 antigen). | |||||
|
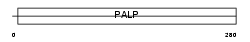
SDHL_HUMAN
|
||||||
| θ value | θ > 10 (rank : 22) | NC score | 0.135779 (rank : 5) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P20132 | Gene names | SDS, SDH | |||
|
Domain Architecture |
|
|||||
| Description | L-serine dehydratase (EC 4.3.1.17) (L-serine deaminase). | |||||
|
CBS_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P35520, Q99425, Q9BWC5 | Gene names | CBS | |||
|
Domain Architecture |

|
|||||
| Description | Cystathionine beta-synthase (EC 4.2.1.22) (Serine sulfhydrase) (Beta- thionase). | |||||
|
SRR_MOUSE
|
||||||
| NC score | 0.478050 (rank : 2) | θ value | 6.64225e-11 (rank : 3) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9QZX7 | Gene names | Srr | |||
|
Domain Architecture |

|
|||||
| Description | Serine racemase (EC 5.1.1.-). | |||||
|
SRR_HUMAN
|
||||||
| NC score | 0.471098 (rank : 3) | θ value | 2.98157e-11 (rank : 2) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 7 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9GZT4 | Gene names | SRR | |||
|
Domain Architecture |

|
|||||
| Description | Serine racemase (EC 5.1.1.-). | |||||
|
SDHL_MOUSE
|
||||||
| NC score | 0.181362 (rank : 4) | θ value | 0.813845 (rank : 6) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8VBT2 | Gene names | Sds | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | L-serine dehydratase (EC 4.3.1.17) (L-serine deaminase). | |||||
|
SDHL_HUMAN
|
||||||
| NC score | 0.135779 (rank : 5) | θ value | θ > 10 (rank : 22) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P20132 | Gene names | SDS, SDH | |||
|
Domain Architecture |

|
|||||
| Description | L-serine dehydratase (EC 4.3.1.17) (L-serine deaminase). | |||||
|
CLCN6_MOUSE
|
||||||
| NC score | 0.060377 (rank : 6) | θ value | 0.47712 (rank : 5) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O35454 | Gene names | Clcn6, Clc6 | |||
|
Domain Architecture |

|
|||||
| Description | Chloride channel protein 6 (ClC-6). | |||||
|
CLCN6_HUMAN
|
||||||
| NC score | 0.055973 (rank : 7) | θ value | 1.06291 (rank : 7) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P51797, O60818, O60819, O60820, O60821, P78520, P78521, Q5SNW2, Q5SNW3, Q5SNX1, Q5SNX2, Q5SNX3, Q99427, Q99428, Q99429 | Gene names | CLCN6, KIAA0046 | |||
|
Domain Architecture |

|
|||||
| Description | Chloride channel protein 6 (ClC-6). | |||||
|
PER3_HUMAN
|
||||||
| NC score | 0.048698 (rank : 8) | θ value | 0.21417 (rank : 4) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 479 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P56645, Q5H8X4, Q969K6, Q96S77, Q96S78, Q9C0J3, Q9NSP9, Q9UGU8 | Gene names | PER3 | |||
|
Domain Architecture |

|
|||||
| Description | Period circadian protein 3 (hPER3). | |||||
|
GSCR1_HUMAN
|
||||||
| NC score | 0.036294 (rank : 9) | θ value | 1.81305 (rank : 9) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 674 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9NZM4 | Gene names | GLTSCR1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Glioma tumor suppressor candidate region gene 1 protein. | |||||
|
MSH5_HUMAN
|
||||||
| NC score | 0.036123 (rank : 10) | θ value | 4.03905 (rank : 13) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 22 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O43196, O60586, Q5BLU9, Q5SSR1, Q8IW44, Q9BQC7 | Gene names | MSH5 | |||
|
Domain Architecture |

|
|||||
| Description | MutS protein homolog 5. | |||||
|
MSH5_MOUSE
|
||||||
| NC score | 0.034486 (rank : 11) | θ value | 5.27518 (rank : 15) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9QUM7, Q9Z1Q6 | Gene names | Msh5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | MutS protein homolog 5. | |||||
|
TCOF_HUMAN
|
||||||
| NC score | 0.028955 (rank : 12) | θ value | 4.03905 (rank : 14) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 543 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q13428, Q99408, Q99860 | Gene names | TCOF1 | |||
|
Domain Architecture |

|
|||||
| Description | Treacle protein (Treacher Collins syndrome protein). | |||||
|
STAC3_MOUSE
|
||||||
| NC score | 0.023212 (rank : 13) | θ value | 3.0926 (rank : 12) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 232 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BZ71 | Gene names | Stac3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | SH3 and cysteine-rich domain-containing protein 3. | |||||
|
LAG3_HUMAN
|
||||||
| NC score | 0.022514 (rank : 14) | θ value | 8.99809 (rank : 21) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 129 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P18627 | Gene names | LAG3, FDC | |||
|
Domain Architecture |

|
|||||
| Description | Lymphocyte activation gene 3 protein precursor (LAG-3) (FDC protein) (CD223 antigen). | |||||
|
BHLH3_HUMAN
|
||||||
| NC score | 0.021491 (rank : 15) | θ value | 6.88961 (rank : 17) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 106 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9C0J9 | Gene names | BHLHB3, DEC2, SHARP1 | |||
|
Domain Architecture |

|
|||||
| Description | Class B basic helix-loop-helix protein 3 (bHLHB3) (Differentially expressed in chondrocytes protein 2) (hDEC2) (Enhancer-of-split and hairy-related protein 1) (SHARP-1). | |||||
|
EMID2_HUMAN
|
||||||
| NC score | 0.020550 (rank : 16) | θ value | 3.0926 (rank : 10) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 226 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96A83 | Gene names | EMID2, COL26A1, EMU2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Collagen alpha-1(XXVI) chain precursor (EMI domain-containing protein 2) (Protein Emu2) (Emilin and multimerin domain-containing protein 2). | |||||
|
RIN2_HUMAN
|
||||||
| NC score | 0.018075 (rank : 17) | θ value | 6.88961 (rank : 20) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8WYP3, Q00425, Q5TFT8, Q9BQL3, Q9H071 | Gene names | RIN2, RASSF4 | |||
|
Domain Architecture |

|
|||||
| Description | Ras and Rab interactor 2 (Ras interaction/interference protein 2) (Ras inhibitor JC265) (Ras association domain family 4). | |||||
|
PTN12_MOUSE
|
||||||
| NC score | 0.013153 (rank : 18) | θ value | 3.0926 (rank : 11) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 123 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P35831 | Gene names | Ptpn12 | |||
|
Domain Architecture |
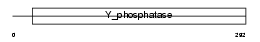
|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 12 (EC 3.1.3.48) (Protein-tyrosine phosphatase P19) (P19-PTP) (MPTP-PEST). | |||||
|
NKTR_HUMAN
|
||||||
| NC score | 0.011046 (rank : 19) | θ value | 6.88961 (rank : 19) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 658 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P30414 | Gene names | NKTR | |||
|
Domain Architecture |

|
|||||
| Description | NK-tumor recognition protein (Natural-killer cells cyclophilin-related protein) (NK-TR protein). | |||||
|
TITIN_HUMAN
|
||||||
| NC score | 0.009499 (rank : 20) | θ value | 1.38821 (rank : 8) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 2923 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8WZ42, Q10465, Q10466, Q15598, Q2XUS3, Q32Q60, Q4U1Z6, Q6NSG0, Q6PDB1, Q6PJP0, Q7KYM2, Q7KYN4, Q7KYN5, Q7LDM3, Q7Z2X3, Q8TCG8, Q8WZ51, Q8WZ52, Q8WZ53, Q8WZB3, Q92761, Q92762, Q9UD97, Q9UP84, Q9Y6L9 | Gene names | TTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Titin (EC 2.7.11.1) (Connectin) (Rhabdomyosarcoma antigen MU-RMS- 40.14). | |||||
|
CO5A1_HUMAN
|
||||||
| NC score | 0.009206 (rank : 21) | θ value | 6.88961 (rank : 18) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 434 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P20908, Q15094, Q5SUX4 | Gene names | COL5A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(V) chain precursor. | |||||
|
ZN696_HUMAN
|
||||||
| NC score | 0.001172 (rank : 22) | θ value | 5.27518 (rank : 16) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 743 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9H7X3 | Gene names | ZNF696 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 696. | |||||