Please be patient as the page loads
|
PI3R5_MOUSE
|
||||||
| SwissProt Accessions | Q5SW28, Q3TU01, Q3UDZ2, Q8C215, Q8CGQ7 | Gene names | Pik3r5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphoinositide 3-kinase regulatory subunit 5 (PI3-kinase regulatory subunit 5) (PI3-kinase p101 subunit) (PtdIns-3-kinase p101) (p101- PI3K) (Phosphatidylinositol-4,5-bisphosphate 3-kinase regulatory subunit) (PtdIns-3-kinase regulatory subunit). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
PI3R5_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.994787 (rank : 2) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8WYR1, Q5G936, Q5G938, Q5G939, Q8IZ23, Q9Y2Y2 | Gene names | PIK3R5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphoinositide 3-kinase regulatory subunit 5 (PI3-kinase regulatory subunit 5) (PI3-kinase p101 subunit) (PtdIns-3-kinase p101) (p101- PI3K) (Phosphatidylinositol-4,5-bisphosphate 3-kinase regulatory subunit) (PtdIns-3-kinase regulatory subunit) (Protein FOAP-2). | |||||
|
PI3R5_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q5SW28, Q3TU01, Q3UDZ2, Q8C215, Q8CGQ7 | Gene names | Pik3r5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphoinositide 3-kinase regulatory subunit 5 (PI3-kinase regulatory subunit 5) (PI3-kinase p101 subunit) (PtdIns-3-kinase p101) (p101- PI3K) (Phosphatidylinositol-4,5-bisphosphate 3-kinase regulatory subunit) (PtdIns-3-kinase regulatory subunit). | |||||
|
PI3R6_MOUSE
|
||||||
| θ value | 7.80994e-28 (rank : 3) | NC score | 0.648200 (rank : 4) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q3U6Q4, Q3TBT5, Q3UAY4, Q8K323 | Gene names | Pik3r6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphoinositide 3-kinase regulatory subunit 6 (Phosphoinositide 3- kinase gamma adapter protein of 87 kDa) (p87 PI3K adapter protein) (p87PIKAP). | |||||
|
PI3R6_HUMAN
|
||||||
| θ value | 1.33217e-27 (rank : 4) | NC score | 0.652667 (rank : 3) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5UE93, Q658R3 | Gene names | PIK3R6, C17orf38 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphoinositide 3-kinase regulatory subunit 6 (Phosphoinositide 3- kinase gamma adapter protein of 87 kDa) (p87 PI3K adapter protein) (p87PIKAP). | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 5) | NC score | 0.037681 (rank : 8) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
NU160_HUMAN
|
||||||
| θ value | 0.21417 (rank : 6) | NC score | 0.097547 (rank : 5) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q12769, Q96GB3, Q9H660 | Gene names | NUP160, KIAA0197, NUP120 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear pore complex protein Nup160 (Nucleoporin Nup160) (160 kDa nucleoporin). | |||||
|
NU160_MOUSE
|
||||||
| θ value | 0.813845 (rank : 7) | NC score | 0.076304 (rank : 6) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Z0W3, Q3TBI7, Q3TP11, Q3U250, Q6A0A7, Q7TME1, Q9CZD9 | Gene names | Nup160, Gtl1-13, Kiaa0197 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear pore complex protein Nup160 (Nucleoporin Nup160) (160 kDa nucleoporin) (Gene trap locus 1-13) (GTL-13). | |||||
|
LAP2A_HUMAN
|
||||||
| θ value | 2.36792 (rank : 8) | NC score | 0.024676 (rank : 9) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P42166, P08918, P08919, Q14860, Q16295 | Gene names | TMPO, LAP2 | |||
|
Domain Architecture |

|
|||||
| Description | Lamina-associated polypeptide 2 isoform alpha (Thymopoietin isoform alpha) (TP alpha) (Thymopoietin-related peptide isoform alpha) (TPRP isoform alpha) [Contains: Thymopoietin (TP) (Splenin); Thymopentin (TP5)]. | |||||
|
MAML3_HUMAN
|
||||||
| θ value | 3.0926 (rank : 9) | NC score | 0.041847 (rank : 7) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96JK9 | Gene names | MAML3, KIAA1816 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mastermind-like protein 3 (Mam-3). | |||||
|
MBD1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 10) | NC score | 0.022130 (rank : 10) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Z2E2, Q792D6, Q8CCL9, Q9DC19 | Gene names | Mbd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 1 (Methyl-CpG-binding protein MBD1). | |||||
|
DHX29_HUMAN
|
||||||
| θ value | 4.03905 (rank : 11) | NC score | 0.013176 (rank : 17) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7Z478, O75549, Q63HN0, Q63HN3, Q8IWW2, Q8N3A1, Q9UMH2 | Gene names | DHX29 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative ATP-dependent RNA helicase DHX29 (EC 3.6.1.-) (DEAH box protein 29) (Nucleic acid helicase DDXx). | |||||
|
NIBL_HUMAN
|
||||||
| θ value | 5.27518 (rank : 12) | NC score | 0.017481 (rank : 12) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96TA1, Q9BUS2, Q9NT35 | Gene names | C9orf88 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Niban-like protein (Meg-3). | |||||
|
SELB_HUMAN
|
||||||
| θ value | 5.27518 (rank : 13) | NC score | 0.015473 (rank : 14) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P57772 | Gene names | EEFSEC, SELB | |||
|
Domain Architecture |
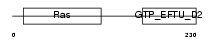
|
|||||
| Description | Selenocysteine-specific elongation factor (Elongation factor sec) (Eukaryotic elongation factor, selenocysteine-tRNA-specific). | |||||
|
TF3C1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 14) | NC score | 0.018204 (rank : 11) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q12789, Q12838, Q9Y4W9 | Gene names | GTF3C1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | General transcription factor 3C polypeptide 1 (Transcription factor IIIC-subunit alpha) (TF3C-alpha) (TFIIIC 220 kDa subunit) (TFIIIC220) (TFIIIC box B-binding subunit). | |||||
|
BRPF3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 15) | NC score | 0.006535 (rank : 20) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9ULD4, Q5R3K8 | Gene names | BRPF3, KIAA1286 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain and PHD finger-containing protein 3. | |||||
|
FAAH_HUMAN
|
||||||
| θ value | 8.99809 (rank : 16) | NC score | 0.014318 (rank : 15) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 4 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O00519, Q5TDF8 | Gene names | FAAH | |||
|
Domain Architecture |

|
|||||
| Description | Fatty-acid amide hydrolase (EC 3.1.-.-) (Oleamide hydrolase) (Anandamide amidohydrolase). | |||||
|
K0776_MOUSE
|
||||||
| θ value | 8.99809 (rank : 17) | NC score | 0.013318 (rank : 16) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CCJ3, Q3V145, Q6ZQ50, Q8BT70, Q8C484, Q9D8I8 | Gene names | Kiaa0776 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0776. | |||||
|
RHG07_HUMAN
|
||||||
| θ value | 8.99809 (rank : 18) | NC score | 0.009788 (rank : 18) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96QB1, O14868, O43199, Q9C0E0 | Gene names | DLC1, ARHGAP7, KIAA1723, STARD12 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 7 (Rho-type GTPase-activating protein 7) (Deleted in liver cancer 1 protein) (Dlc-1) (HP protein) (StAR-related lipid transfer protein 12) (StARD12) (START domain-containing protein 12). | |||||
|
RHG07_MOUSE
|
||||||
| θ value | 8.99809 (rank : 19) | NC score | 0.009692 (rank : 19) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9R0Z9, Q8R541 | Gene names | Dlc1, Arhgap7, Stard12 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 7 (Rho-type GTPase-activating protein 7) (Deleted in liver cancer 1 protein homolog) (Dlc-1) (StAR-related lipid transfer protein 12) (StARD12) (START domain-containing protein 12). | |||||
|
RRBP1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 20) | NC score | 0.001771 (rank : 21) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 1650 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99PL5, Q99PK5, Q99PK6, Q99PK7, Q99PK8, Q99PK9, Q99PL0, Q99PL1, Q99PL2, Q99PL3, Q99PL4, Q9CS20 | Gene names | Rrbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (mRRp). | |||||
|
STAG3_HUMAN
|
||||||
| θ value | 8.99809 (rank : 21) | NC score | 0.015554 (rank : 13) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UJ98, Q8NDP3 | Gene names | STAG3 | |||
|
Domain Architecture |

|
|||||
| Description | Cohesin subunit SA-3 (Stromal antigen 3) (Stromalin 3) (SCC3 homolog 3). | |||||
|
PI3R5_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 21 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q5SW28, Q3TU01, Q3UDZ2, Q8C215, Q8CGQ7 | Gene names | Pik3r5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphoinositide 3-kinase regulatory subunit 5 (PI3-kinase regulatory subunit 5) (PI3-kinase p101 subunit) (PtdIns-3-kinase p101) (p101- PI3K) (Phosphatidylinositol-4,5-bisphosphate 3-kinase regulatory subunit) (PtdIns-3-kinase regulatory subunit). | |||||
|
PI3R5_HUMAN
|
||||||
| NC score | 0.994787 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8WYR1, Q5G936, Q5G938, Q5G939, Q8IZ23, Q9Y2Y2 | Gene names | PIK3R5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphoinositide 3-kinase regulatory subunit 5 (PI3-kinase regulatory subunit 5) (PI3-kinase p101 subunit) (PtdIns-3-kinase p101) (p101- PI3K) (Phosphatidylinositol-4,5-bisphosphate 3-kinase regulatory subunit) (PtdIns-3-kinase regulatory subunit) (Protein FOAP-2). | |||||
|
PI3R6_HUMAN
|
||||||
| NC score | 0.652667 (rank : 3) | θ value | 1.33217e-27 (rank : 4) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q5UE93, Q658R3 | Gene names | PIK3R6, C17orf38 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphoinositide 3-kinase regulatory subunit 6 (Phosphoinositide 3- kinase gamma adapter protein of 87 kDa) (p87 PI3K adapter protein) (p87PIKAP). | |||||
|
PI3R6_MOUSE
|
||||||
| NC score | 0.648200 (rank : 4) | θ value | 7.80994e-28 (rank : 3) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q3U6Q4, Q3TBT5, Q3UAY4, Q8K323 | Gene names | Pik3r6 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Phosphoinositide 3-kinase regulatory subunit 6 (Phosphoinositide 3- kinase gamma adapter protein of 87 kDa) (p87 PI3K adapter protein) (p87PIKAP). | |||||
|
NU160_HUMAN
|
||||||
| NC score | 0.097547 (rank : 5) | θ value | 0.21417 (rank : 6) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q12769, Q96GB3, Q9H660 | Gene names | NUP160, KIAA0197, NUP120 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear pore complex protein Nup160 (Nucleoporin Nup160) (160 kDa nucleoporin). | |||||
|
NU160_MOUSE
|
||||||
| NC score | 0.076304 (rank : 6) | θ value | 0.813845 (rank : 7) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9Z0W3, Q3TBI7, Q3TP11, Q3U250, Q6A0A7, Q7TME1, Q9CZD9 | Gene names | Nup160, Gtl1-13, Kiaa0197 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear pore complex protein Nup160 (Nucleoporin Nup160) (160 kDa nucleoporin) (Gene trap locus 1-13) (GTL-13). | |||||
|
MAML3_HUMAN
|
||||||
| NC score | 0.041847 (rank : 7) | θ value | 3.0926 (rank : 9) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 323 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q96JK9 | Gene names | MAML3, KIAA1816 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mastermind-like protein 3 (Mam-3). | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.037681 (rank : 8) | θ value | 0.0736092 (rank : 5) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
LAP2A_HUMAN
|
||||||
| NC score | 0.024676 (rank : 9) | θ value | 2.36792 (rank : 8) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P42166, P08918, P08919, Q14860, Q16295 | Gene names | TMPO, LAP2 | |||
|
Domain Architecture |

|
|||||
| Description | Lamina-associated polypeptide 2 isoform alpha (Thymopoietin isoform alpha) (TP alpha) (Thymopoietin-related peptide isoform alpha) (TPRP isoform alpha) [Contains: Thymopoietin (TP) (Splenin); Thymopentin (TP5)]. | |||||
|
MBD1_MOUSE
|
||||||
| NC score | 0.022130 (rank : 10) | θ value | 3.0926 (rank : 10) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 72 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Z2E2, Q792D6, Q8CCL9, Q9DC19 | Gene names | Mbd1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Methyl-CpG-binding domain protein 1 (Methyl-CpG-binding protein MBD1). | |||||
|
TF3C1_HUMAN
|
||||||
| NC score | 0.018204 (rank : 11) | θ value | 5.27518 (rank : 14) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 88 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q12789, Q12838, Q9Y4W9 | Gene names | GTF3C1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | General transcription factor 3C polypeptide 1 (Transcription factor IIIC-subunit alpha) (TF3C-alpha) (TFIIIC 220 kDa subunit) (TFIIIC220) (TFIIIC box B-binding subunit). | |||||
|
NIBL_HUMAN
|
||||||
| NC score | 0.017481 (rank : 12) | θ value | 5.27518 (rank : 12) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 61 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96TA1, Q9BUS2, Q9NT35 | Gene names | C9orf88 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Niban-like protein (Meg-3). | |||||
|
STAG3_HUMAN
|
||||||
| NC score | 0.015554 (rank : 13) | θ value | 8.99809 (rank : 21) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UJ98, Q8NDP3 | Gene names | STAG3 | |||
|
Domain Architecture |

|
|||||
| Description | Cohesin subunit SA-3 (Stromal antigen 3) (Stromalin 3) (SCC3 homolog 3). | |||||
|
SELB_HUMAN
|
||||||
| NC score | 0.015473 (rank : 14) | θ value | 5.27518 (rank : 13) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P57772 | Gene names | EEFSEC, SELB | |||
|
Domain Architecture |

|
|||||
| Description | Selenocysteine-specific elongation factor (Elongation factor sec) (Eukaryotic elongation factor, selenocysteine-tRNA-specific). | |||||
|
FAAH_HUMAN
|
||||||
| NC score | 0.014318 (rank : 15) | θ value | 8.99809 (rank : 16) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 4 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O00519, Q5TDF8 | Gene names | FAAH | |||
|
Domain Architecture |

|
|||||
| Description | Fatty-acid amide hydrolase (EC 3.1.-.-) (Oleamide hydrolase) (Anandamide amidohydrolase). | |||||
|
K0776_MOUSE
|
||||||
| NC score | 0.013318 (rank : 16) | θ value | 8.99809 (rank : 17) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8CCJ3, Q3V145, Q6ZQ50, Q8BT70, Q8C484, Q9D8I8 | Gene names | Kiaa0776 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein KIAA0776. | |||||
|
DHX29_HUMAN
|
||||||
| NC score | 0.013176 (rank : 17) | θ value | 4.03905 (rank : 11) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 144 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q7Z478, O75549, Q63HN0, Q63HN3, Q8IWW2, Q8N3A1, Q9UMH2 | Gene names | DHX29 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative ATP-dependent RNA helicase DHX29 (EC 3.6.1.-) (DEAH box protein 29) (Nucleic acid helicase DDXx). | |||||
|
RHG07_HUMAN
|
||||||
| NC score | 0.009788 (rank : 18) | θ value | 8.99809 (rank : 18) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 108 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q96QB1, O14868, O43199, Q9C0E0 | Gene names | DLC1, ARHGAP7, KIAA1723, STARD12 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 7 (Rho-type GTPase-activating protein 7) (Deleted in liver cancer 1 protein) (Dlc-1) (HP protein) (StAR-related lipid transfer protein 12) (StARD12) (START domain-containing protein 12). | |||||
|
RHG07_MOUSE
|
||||||
| NC score | 0.009692 (rank : 19) | θ value | 8.99809 (rank : 19) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 103 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9R0Z9, Q8R541 | Gene names | Dlc1, Arhgap7, Stard12 | |||
|
Domain Architecture |

|
|||||
| Description | Rho-GTPase-activating protein 7 (Rho-type GTPase-activating protein 7) (Deleted in liver cancer 1 protein homolog) (Dlc-1) (StAR-related lipid transfer protein 12) (StARD12) (START domain-containing protein 12). | |||||
|
BRPF3_HUMAN
|
||||||
| NC score | 0.006535 (rank : 20) | θ value | 8.99809 (rank : 15) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 217 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9ULD4, Q5R3K8 | Gene names | BRPF3, KIAA1286 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Bromodomain and PHD finger-containing protein 3. | |||||
|
RRBP1_MOUSE
|
||||||
| NC score | 0.001771 (rank : 21) | θ value | 8.99809 (rank : 20) | |||
| Query Neighborhood Hits | 21 | Target Neighborhood Hits | 1650 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99PL5, Q99PK5, Q99PK6, Q99PK7, Q99PK8, Q99PK9, Q99PL0, Q99PL1, Q99PL2, Q99PL3, Q99PL4, Q9CS20 | Gene names | Rrbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (mRRp). | |||||