Please be patient as the page loads
|
OXSM_HUMAN
|
||||||
| SwissProt Accessions | Q9NWU1 | Gene names | OXSM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 3-oxoacyl-[acyl-carrier-protein] synthase, mitochondrial precursor (EC 2.3.1.41) (Beta-ketoacyl synthase). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
OXSM_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9NWU1 | Gene names | OXSM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 3-oxoacyl-[acyl-carrier-protein] synthase, mitochondrial precursor (EC 2.3.1.41) (Beta-ketoacyl synthase). | |||||
|
OXSM_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.998180 (rank : 2) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9D404 | Gene names | Oxsm | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 3-oxoacyl-[acyl-carrier-protein] synthase, mitochondrial precursor (EC 2.3.1.41) (Beta-ketoacyl synthase). | |||||
|
FAS_HUMAN
|
||||||
| θ value | 5.79196e-23 (rank : 3) | NC score | 0.531709 (rank : 3) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P49327, Q13479, Q16702, Q6P4U5, Q6SS02, Q969R1, Q96C68, Q96IT0 | Gene names | FASN, FAS | |||
|
Domain Architecture |

|
|||||
| Description | Fatty acid synthase (EC 2.3.1.85) [Includes: [Acyl-carrier-protein] S- acetyltransferase (EC 2.3.1.38); [Acyl-carrier-protein] S- malonyltransferase (EC 2.3.1.39); 3-oxoacyl-[acyl-carrier-protein] synthase (EC 2.3.1.41); 3-oxoacyl-[acyl-carrier-protein] reductase (EC 1.1.1.100); 3-hydroxypalmitoyl-[acyl-carrier-protein] dehydratase (EC 4.2.1.61); Enoyl-[acyl-carrier-protein] reductase (EC 1.3.1.10); Oleoyl-[acyl-carrier-protein] hydrolase (EC 3.1.2.14)]. | |||||
|
FAS_MOUSE
|
||||||
| θ value | 1.86321e-21 (rank : 4) | NC score | 0.523513 (rank : 4) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P19096, Q6PB72, Q8C4Z0, Q9EQR0 | Gene names | Fasn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fatty acid synthase (EC 2.3.1.85) [Includes: [Acyl-carrier-protein] S- acetyltransferase (EC 2.3.1.38); [Acyl-carrier-protein] S- malonyltransferase (EC 2.3.1.39); 3-oxoacyl-[acyl-carrier-protein] synthase (EC 2.3.1.41); 3-oxoacyl-[acyl-carrier-protein] reductase (EC 1.1.1.100); 3-hydroxypalmitoyl-[acyl-carrier-protein] dehydratase (EC 4.2.1.61); Enoyl-[acyl-carrier-protein] reductase (EC 1.3.1.10); Oleoyl-[acyl-carrier-protein] hydrolase (EC 3.1.2.14)]. | |||||
|
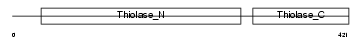
THIL_HUMAN
|
||||||
| θ value | 0.21417 (rank : 5) | NC score | 0.077089 (rank : 10) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P24752 | Gene names | ACAT1, ACAT, MAT | |||
|
Domain Architecture |
|
|||||
| Description | Acetyl-CoA acetyltransferase, mitochondrial precursor (EC 2.3.1.9) (Acetoacetyl-CoA thiolase) (T2). | |||||
|
ECHB_MOUSE
|
||||||
| θ value | 0.365318 (rank : 6) | NC score | 0.100519 (rank : 6) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q99JY0, Q3TEH9, Q8BJI5, Q8BJM0, Q8BK52 | Gene names | Hadhb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trifunctional enzyme subunit beta, mitochondrial precursor (TP-beta) [Includes: 3-ketoacyl-CoA thiolase (EC 2.3.1.16) (Acetyl-CoA acyltransferase) (Beta-ketothiolase)]. | |||||
|
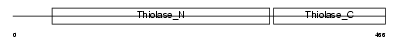
ECHB_HUMAN
|
||||||
| θ value | 0.47712 (rank : 7) | NC score | 0.091870 (rank : 7) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P55084, O14969, Q53TA6, Q96C77, Q9H3F5 | Gene names | HADHB | |||
|
Domain Architecture |
|
|||||
| Description | Trifunctional enzyme subunit beta, mitochondrial precursor (TP-beta) [Includes: 3-ketoacyl-CoA thiolase (EC 2.3.1.16) (Acetyl-CoA acyltransferase) (Beta-ketothiolase)]. | |||||
|
USH2A_HUMAN
|
||||||
| θ value | 3.0926 (rank : 8) | NC score | 0.008743 (rank : 25) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 600 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O75445, Q5VVM9, Q6S362, Q9NS27 | Gene names | USH2A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Usherin precursor (Usher syndrome type-2A protein) (Usher syndrome type IIa protein). | |||||
|
MLL2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 9) | NC score | 0.007744 (rank : 28) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||
|
THIL_MOUSE
|
||||||
| θ value | 5.27518 (rank : 10) | NC score | 0.062844 (rank : 18) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8QZT1, Q3TE92 | Gene names | Acat1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acetyl-CoA acetyltransferase, mitochondrial precursor (EC 2.3.1.9) (Acetoacetyl-CoA thiolase). | |||||
|
THIM_HUMAN
|
||||||
| θ value | 5.27518 (rank : 11) | NC score | 0.048168 (rank : 24) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P42765, Q9BUT6 | Gene names | ACAA2 | |||
|
Domain Architecture |

|
|||||
| Description | 3-ketoacyl-CoA thiolase, mitochondrial (EC 2.3.1.16) (Beta- ketothiolase) (Acetyl-CoA acyltransferase) (Mitochondrial 3-oxoacyl- CoA thiolase) (T1). | |||||
|
CO4B_MOUSE
|
||||||
| θ value | 8.99809 (rank : 12) | NC score | 0.007942 (rank : 26) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P01029, Q31201, Q61372, Q61859, Q62353 | Gene names | C4b, C4 | |||
|
Domain Architecture |

|
|||||
| Description | Complement C4-B precursor [Contains: Complement C4 beta chain; Complement C4 alpha chain; C4a anaphylatoxin; Complement C4 gamma chain]. | |||||
|
RGAP1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 13) | NC score | 0.007883 (rank : 27) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 185 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9WVM1, Q3THR5, Q3TI41, Q3TM81 | Gene names | Racgap1, Mgcracgap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rac GTPase-activating protein 1 (MgcRacGAP). | |||||
|
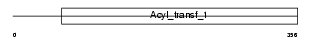
FABD_HUMAN
|
||||||
| θ value | θ > 10 (rank : 14) | NC score | 0.061945 (rank : 19) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 4 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8IVS2, O95510, O95511 | Gene names | MCAT, MT | |||
|
Domain Architecture |
|
|||||
| Description | Malonyl CoA-acyl carrier protein transacylase, mitochondrial precursor (EC 2.3.1.39) (MCT) (Mitochondrial malonyltransferase). | |||||
|
K1576_HUMAN
|
||||||
| θ value | θ > 10 (rank : 15) | NC score | 0.067582 (rank : 13) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9HCJ6, Q8IYW8 | Gene names | KIAA1576 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable oxidoreductase KIAA1576 (EC 1.-.-.-). | |||||
|
K1576_MOUSE
|
||||||
| θ value | θ > 10 (rank : 16) | NC score | 0.067107 (rank : 14) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q80TB8, Q6PGM9 | Gene names | Kiaa1576 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable oxidoreductase KIAA1576 (EC 1.-.-.-). | |||||
|
LTB4D_HUMAN
|
||||||
| θ value | θ > 10 (rank : 17) | NC score | 0.054214 (rank : 21) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q14914, Q8IYQ0, Q9H1X6 | Gene names | LTB4DH | |||
|
Domain Architecture |
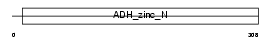
|
|||||
| Description | NADP-dependent leukotriene B4 12-hydroxydehydrogenase (EC 1.3.1.74) (15-oxoprostaglandin 13-reductase) (EC 1.3.1.48). | |||||
|
LTB4D_MOUSE
|
||||||
| θ value | θ > 10 (rank : 18) | NC score | 0.053359 (rank : 23) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q91YR9, Q3TKT6, Q9CPS1 | Gene names | Ltb4dh | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NADP-dependent leukotriene B4 12-hydroxydehydrogenase (EC 1.3.1.74) (15-oxoprostaglandin 13-reductase) (EC 1.3.1.48). | |||||
|
MECR_HUMAN
|
||||||
| θ value | θ > 10 (rank : 19) | NC score | 0.053761 (rank : 22) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9BV79, Q5SYU0, Q5SYU1, Q5SYU2, Q6IBU9, Q9Y373 | Gene names | MECR, NBRF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trans-2-enoyl-CoA reductase, mitochondrial precursor (EC 1.3.1.38) (HsNrbf-1) (NRBF-1). | |||||
|
QORL_HUMAN
|
||||||
| θ value | θ > 10 (rank : 20) | NC score | 0.061433 (rank : 20) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O95825, Q96DY0, Q9NVY7 | Gene names | CRYZL1 | |||
|
Domain Architecture |
|
|||||
| Description | Quinone oxidoreductase-like 1 (EC 1.-.-.-) (QOH-1) (Zeta-crystallin homolog) (4P11). | |||||
|
QORL_MOUSE
|
||||||
| θ value | θ > 10 (rank : 21) | NC score | 0.065458 (rank : 16) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q921W4, Q3UKX2, Q8VCS1, Q9CWR9 | Gene names | Cryzl1 | |||
|
Domain Architecture |
|
|||||
| Description | Quinone oxidoreductase-like 1 (EC 1.-.-.-) (QOH-1) (Zeta-crystallin homolog). | |||||
|
QORX_HUMAN
|
||||||
| θ value | θ > 10 (rank : 22) | NC score | 0.113801 (rank : 5) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q53FA7, O14679, O14685, Q38G78, Q6JLE7, Q9BWB8 | Gene names | TP53I3, PIG3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative quinone oxidoreductase (EC 1.-.-.-) (Tumor protein p53- inducible protein 3) (p53-induced protein 3). | |||||
|
QOR_HUMAN
|
||||||
| θ value | θ > 10 (rank : 23) | NC score | 0.082685 (rank : 9) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q08257, Q53FT0, Q59EU7, Q6NSK9 | Gene names | CRYZ | |||
|
Domain Architecture |
|
|||||
| Description | Quinone oxidoreductase (EC 1.6.5.5) (NADPH:quinone reductase) (Zeta- crystallin). | |||||
|
QOR_MOUSE
|
||||||
| θ value | θ > 10 (rank : 24) | NC score | 0.084460 (rank : 8) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P47199, Q62508, Q99L63 | Gene names | Cryz | |||
|
Domain Architecture |
|
|||||
| Description | Quinone oxidoreductase (EC 1.6.5.5) (NADPH:quinone reductase) (Zeta- crystallin). | |||||
|
VAT1_HUMAN
|
||||||
| θ value | θ > 10 (rank : 25) | NC score | 0.064760 (rank : 17) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99536, Q13035, Q96A39, Q9BUT8 | Gene names | VAT1 | |||
|
Domain Architecture |
|
|||||
| Description | Synaptic vesicle membrane protein VAT-1 homolog (EC 1.-.-.-). | |||||
|
VAT1_MOUSE
|
||||||
| θ value | θ > 10 (rank : 26) | NC score | 0.066149 (rank : 15) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q62465, Q8R3G0, Q8R3T8 | Gene names | Vat1, Vat-1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptic vesicle membrane protein VAT-1 homolog (EC 1.-.-.-). | |||||
|
ZADH2_HUMAN
|
||||||
| θ value | θ > 10 (rank : 27) | NC score | 0.071016 (rank : 12) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8N4Q0 | Gene names | ZADH2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc-binding alcohol dehydrogenase domain-containing protein 2 (EC 1.-.-.-). | |||||
|
ZADH2_MOUSE
|
||||||
| θ value | θ > 10 (rank : 28) | NC score | 0.074071 (rank : 11) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BGC4 | Gene names | Zadh2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc-binding alcohol dehydrogenase domain-containing protein 2 (EC 1.-.-.-). | |||||
|
OXSM_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 13 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q9NWU1 | Gene names | OXSM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 3-oxoacyl-[acyl-carrier-protein] synthase, mitochondrial precursor (EC 2.3.1.41) (Beta-ketoacyl synthase). | |||||
|
OXSM_MOUSE
|
||||||
| NC score | 0.998180 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9D404 | Gene names | Oxsm | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | 3-oxoacyl-[acyl-carrier-protein] synthase, mitochondrial precursor (EC 2.3.1.41) (Beta-ketoacyl synthase). | |||||
|
FAS_HUMAN
|
||||||
| NC score | 0.531709 (rank : 3) | θ value | 5.79196e-23 (rank : 3) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P49327, Q13479, Q16702, Q6P4U5, Q6SS02, Q969R1, Q96C68, Q96IT0 | Gene names | FASN, FAS | |||
|
Domain Architecture |

|
|||||
| Description | Fatty acid synthase (EC 2.3.1.85) [Includes: [Acyl-carrier-protein] S- acetyltransferase (EC 2.3.1.38); [Acyl-carrier-protein] S- malonyltransferase (EC 2.3.1.39); 3-oxoacyl-[acyl-carrier-protein] synthase (EC 2.3.1.41); 3-oxoacyl-[acyl-carrier-protein] reductase (EC 1.1.1.100); 3-hydroxypalmitoyl-[acyl-carrier-protein] dehydratase (EC 4.2.1.61); Enoyl-[acyl-carrier-protein] reductase (EC 1.3.1.10); Oleoyl-[acyl-carrier-protein] hydrolase (EC 3.1.2.14)]. | |||||
|
FAS_MOUSE
|
||||||
| NC score | 0.523513 (rank : 4) | θ value | 1.86321e-21 (rank : 4) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 76 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P19096, Q6PB72, Q8C4Z0, Q9EQR0 | Gene names | Fasn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Fatty acid synthase (EC 2.3.1.85) [Includes: [Acyl-carrier-protein] S- acetyltransferase (EC 2.3.1.38); [Acyl-carrier-protein] S- malonyltransferase (EC 2.3.1.39); 3-oxoacyl-[acyl-carrier-protein] synthase (EC 2.3.1.41); 3-oxoacyl-[acyl-carrier-protein] reductase (EC 1.1.1.100); 3-hydroxypalmitoyl-[acyl-carrier-protein] dehydratase (EC 4.2.1.61); Enoyl-[acyl-carrier-protein] reductase (EC 1.3.1.10); Oleoyl-[acyl-carrier-protein] hydrolase (EC 3.1.2.14)]. | |||||
|
QORX_HUMAN
|
||||||
| NC score | 0.113801 (rank : 5) | θ value | θ > 10 (rank : 22) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q53FA7, O14679, O14685, Q38G78, Q6JLE7, Q9BWB8 | Gene names | TP53I3, PIG3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Putative quinone oxidoreductase (EC 1.-.-.-) (Tumor protein p53- inducible protein 3) (p53-induced protein 3). | |||||
|
ECHB_MOUSE
|
||||||
| NC score | 0.100519 (rank : 6) | θ value | 0.365318 (rank : 6) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 20 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q99JY0, Q3TEH9, Q8BJI5, Q8BJM0, Q8BK52 | Gene names | Hadhb | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trifunctional enzyme subunit beta, mitochondrial precursor (TP-beta) [Includes: 3-ketoacyl-CoA thiolase (EC 2.3.1.16) (Acetyl-CoA acyltransferase) (Beta-ketothiolase)]. | |||||
|
ECHB_HUMAN
|
||||||
| NC score | 0.091870 (rank : 7) | θ value | 0.47712 (rank : 7) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P55084, O14969, Q53TA6, Q96C77, Q9H3F5 | Gene names | HADHB | |||
|
Domain Architecture |

|
|||||
| Description | Trifunctional enzyme subunit beta, mitochondrial precursor (TP-beta) [Includes: 3-ketoacyl-CoA thiolase (EC 2.3.1.16) (Acetyl-CoA acyltransferase) (Beta-ketothiolase)]. | |||||
|
QOR_MOUSE
|
||||||
| NC score | 0.084460 (rank : 8) | θ value | θ > 10 (rank : 24) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P47199, Q62508, Q99L63 | Gene names | Cryz | |||
|
Domain Architecture |

|
|||||
| Description | Quinone oxidoreductase (EC 1.6.5.5) (NADPH:quinone reductase) (Zeta- crystallin). | |||||
|
QOR_HUMAN
|
||||||
| NC score | 0.082685 (rank : 9) | θ value | θ > 10 (rank : 23) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q08257, Q53FT0, Q59EU7, Q6NSK9 | Gene names | CRYZ | |||
|
Domain Architecture |
|
|||||
| Description | Quinone oxidoreductase (EC 1.6.5.5) (NADPH:quinone reductase) (Zeta- crystallin). | |||||
|
THIL_HUMAN
|
||||||
| NC score | 0.077089 (rank : 10) | θ value | 0.21417 (rank : 5) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P24752 | Gene names | ACAT1, ACAT, MAT | |||
|
Domain Architecture |

|
|||||
| Description | Acetyl-CoA acetyltransferase, mitochondrial precursor (EC 2.3.1.9) (Acetoacetyl-CoA thiolase) (T2). | |||||
|
ZADH2_MOUSE
|
||||||
| NC score | 0.074071 (rank : 11) | θ value | θ > 10 (rank : 28) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BGC4 | Gene names | Zadh2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc-binding alcohol dehydrogenase domain-containing protein 2 (EC 1.-.-.-). | |||||
|
ZADH2_HUMAN
|
||||||
| NC score | 0.071016 (rank : 12) | θ value | θ > 10 (rank : 27) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8N4Q0 | Gene names | ZADH2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc-binding alcohol dehydrogenase domain-containing protein 2 (EC 1.-.-.-). | |||||
|
K1576_HUMAN
|
||||||
| NC score | 0.067582 (rank : 13) | θ value | θ > 10 (rank : 15) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9HCJ6, Q8IYW8 | Gene names | KIAA1576 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable oxidoreductase KIAA1576 (EC 1.-.-.-). | |||||
|
K1576_MOUSE
|
||||||
| NC score | 0.067107 (rank : 14) | θ value | θ > 10 (rank : 16) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q80TB8, Q6PGM9 | Gene names | Kiaa1576 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable oxidoreductase KIAA1576 (EC 1.-.-.-). | |||||
|
VAT1_MOUSE
|
||||||
| NC score | 0.066149 (rank : 15) | θ value | θ > 10 (rank : 26) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q62465, Q8R3G0, Q8R3T8 | Gene names | Vat1, Vat-1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Synaptic vesicle membrane protein VAT-1 homolog (EC 1.-.-.-). | |||||
|
QORL_MOUSE
|
||||||
| NC score | 0.065458 (rank : 16) | θ value | θ > 10 (rank : 21) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q921W4, Q3UKX2, Q8VCS1, Q9CWR9 | Gene names | Cryzl1 | |||
|
Domain Architecture |

|
|||||
| Description | Quinone oxidoreductase-like 1 (EC 1.-.-.-) (QOH-1) (Zeta-crystallin homolog). | |||||
|
VAT1_HUMAN
|
||||||
| NC score | 0.064760 (rank : 17) | θ value | θ > 10 (rank : 25) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 33 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q99536, Q13035, Q96A39, Q9BUT8 | Gene names | VAT1 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptic vesicle membrane protein VAT-1 homolog (EC 1.-.-.-). | |||||
|
THIL_MOUSE
|
||||||
| NC score | 0.062844 (rank : 18) | θ value | 5.27518 (rank : 10) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q8QZT1, Q3TE92 | Gene names | Acat1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Acetyl-CoA acetyltransferase, mitochondrial precursor (EC 2.3.1.9) (Acetoacetyl-CoA thiolase). | |||||
|
FABD_HUMAN
|
||||||
| NC score | 0.061945 (rank : 19) | θ value | θ > 10 (rank : 14) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 4 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8IVS2, O95510, O95511 | Gene names | MCAT, MT | |||
|
Domain Architecture |

|
|||||
| Description | Malonyl CoA-acyl carrier protein transacylase, mitochondrial precursor (EC 2.3.1.39) (MCT) (Mitochondrial malonyltransferase). | |||||
|
QORL_HUMAN
|
||||||
| NC score | 0.061433 (rank : 20) | θ value | θ > 10 (rank : 20) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | O95825, Q96DY0, Q9NVY7 | Gene names | CRYZL1 | |||
|
Domain Architecture |

|
|||||
| Description | Quinone oxidoreductase-like 1 (EC 1.-.-.-) (QOH-1) (Zeta-crystallin homolog) (4P11). | |||||
|
LTB4D_HUMAN
|
||||||
| NC score | 0.054214 (rank : 21) | θ value | θ > 10 (rank : 17) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q14914, Q8IYQ0, Q9H1X6 | Gene names | LTB4DH | |||
|
Domain Architecture |
|
|||||
| Description | NADP-dependent leukotriene B4 12-hydroxydehydrogenase (EC 1.3.1.74) (15-oxoprostaglandin 13-reductase) (EC 1.3.1.48). | |||||
|
MECR_HUMAN
|
||||||
| NC score | 0.053761 (rank : 22) | θ value | θ > 10 (rank : 19) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9BV79, Q5SYU0, Q5SYU1, Q5SYU2, Q6IBU9, Q9Y373 | Gene names | MECR, NBRF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Trans-2-enoyl-CoA reductase, mitochondrial precursor (EC 1.3.1.38) (HsNrbf-1) (NRBF-1). | |||||
|
LTB4D_MOUSE
|
||||||
| NC score | 0.053359 (rank : 23) | θ value | θ > 10 (rank : 18) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q91YR9, Q3TKT6, Q9CPS1 | Gene names | Ltb4dh | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | NADP-dependent leukotriene B4 12-hydroxydehydrogenase (EC 1.3.1.74) (15-oxoprostaglandin 13-reductase) (EC 1.3.1.48). | |||||
|
THIM_HUMAN
|
||||||
| NC score | 0.048168 (rank : 24) | θ value | 5.27518 (rank : 11) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 15 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P42765, Q9BUT6 | Gene names | ACAA2 | |||
|
Domain Architecture |

|
|||||
| Description | 3-ketoacyl-CoA thiolase, mitochondrial (EC 2.3.1.16) (Beta- ketothiolase) (Acetyl-CoA acyltransferase) (Mitochondrial 3-oxoacyl- CoA thiolase) (T1). | |||||
|
USH2A_HUMAN
|
||||||
| NC score | 0.008743 (rank : 25) | θ value | 3.0926 (rank : 8) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 600 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O75445, Q5VVM9, Q6S362, Q9NS27 | Gene names | USH2A | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Usherin precursor (Usher syndrome type-2A protein) (Usher syndrome type IIa protein). | |||||
|
CO4B_MOUSE
|
||||||
| NC score | 0.007942 (rank : 26) | θ value | 8.99809 (rank : 12) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P01029, Q31201, Q61372, Q61859, Q62353 | Gene names | C4b, C4 | |||
|
Domain Architecture |

|
|||||
| Description | Complement C4-B precursor [Contains: Complement C4 beta chain; Complement C4 alpha chain; C4a anaphylatoxin; Complement C4 gamma chain]. | |||||
|
RGAP1_MOUSE
|
||||||
| NC score | 0.007883 (rank : 27) | θ value | 8.99809 (rank : 13) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 185 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9WVM1, Q3THR5, Q3TI41, Q3TM81 | Gene names | Racgap1, Mgcracgap | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rac GTPase-activating protein 1 (MgcRacGAP). | |||||
|
MLL2_HUMAN
|
||||||
| NC score | 0.007744 (rank : 28) | θ value | 5.27518 (rank : 9) | |||
| Query Neighborhood Hits | 13 | Target Neighborhood Hits | 1788 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | O14686, O14687 | Gene names | MLL2, ALR | |||
|
Domain Architecture |

|
|||||
| Description | Myeloid/lymphoid or mixed-lineage leukemia protein 2 (ALL1-related protein). | |||||