Please be patient as the page loads
|
NUFP2_MOUSE
|
||||||
| SwissProt Accessions | Q5F2E7, Q3TCE2, Q3V195, Q80TF1 | Gene names | Nufip2, Kiaa1321 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear fragile X mental retardation-interacting protein 2 (FMRP- interacting protein 2) (82 kDa FMRP-interacting protein) (82-FIP). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
NUFP2_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.986251 (rank : 2) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q7Z417, Q9P2M5 | Gene names | NUFIP2, KIAA1321, PIG1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear fragile X mental retardation-interacting protein 2 (FMRP- interacting protein 2) (82 kDa FMRP-interacting protein) (82-FIP) (Proliferation-inducing gene 1 protein). | |||||
|
NUFP2_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q5F2E7, Q3TCE2, Q3V195, Q80TF1 | Gene names | Nufip2, Kiaa1321 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear fragile X mental retardation-interacting protein 2 (FMRP- interacting protein 2) (82 kDa FMRP-interacting protein) (82-FIP). | |||||
|
DDX21_MOUSE
|
||||||
| θ value | 0.0113563 (rank : 3) | NC score | 0.034556 (rank : 16) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9JIK5, Q9WV45 | Gene names | Ddx21 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar RNA helicase 2 (EC 3.6.1.-) (Nucleolar RNA helicase II) (Nucleolar RNA helicase Gu) (RH II/Gu) (Gu-alpha) (DEAD box protein 21). | |||||
|
RRBP1_MOUSE
|
||||||
| θ value | 0.0252991 (rank : 4) | NC score | 0.044543 (rank : 13) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1650 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q99PL5, Q99PK5, Q99PK6, Q99PK7, Q99PK8, Q99PK9, Q99PL0, Q99PL1, Q99PL2, Q99PL3, Q99PL4, Q9CS20 | Gene names | Rrbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (mRRp). | |||||
|
CYLC1_HUMAN
|
||||||
| θ value | 0.279714 (rank : 5) | NC score | 0.086892 (rank : 3) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P35663, Q5JQQ9 | Gene names | CYLC1, CYL, CYL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-1 (Cylicin I) (Multiple-band polypeptide I). | |||||
|
RFIP1_HUMAN
|
||||||
| θ value | 0.279714 (rank : 6) | NC score | 0.046022 (rank : 12) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6WKZ4, Q6AZK4, Q6WKZ2, Q6WKZ6, Q86YV4, Q8TDL1, Q9H642 | Gene names | RAB11FIP1, RCP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rab11 family-interacting protein 1 (Rab11-FIP1) (Rab-coupling protein). | |||||
|
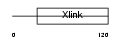
CD44_MOUSE
|
||||||
| θ value | 0.813845 (rank : 7) | NC score | 0.042498 (rank : 14) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P15379, Q05732, Q61395, Q62060, Q62061, Q62062, Q62063, Q62408, Q62409, Q64296, Q99J14, Q9QYX8 | Gene names | Cd44, Ly-24 | |||
|
Domain Architecture |
|
|||||
| Description | CD44 antigen precursor (Phagocytic glycoprotein 1) (PGP-1) (HUTCH-I) (Extracellular matrix receptor III) (ECMR-III) (GP90 lymphocyte homing/adhesion receptor) (Hermes antigen) (Hyaluronate receptor) (Lymphocyte antigen Ly-24). | |||||
|
CHIC2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 8) | NC score | 0.073667 (rank : 5) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UKJ5 | Gene names | CHIC2, BTL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cysteine-rich hydrophobic domain 2 protein (BrX-like translocated in leukemia). | |||||
|
CHIC2_MOUSE
|
||||||
| θ value | 0.813845 (rank : 9) | NC score | 0.073747 (rank : 4) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9D9G3, Q3T9U2, Q80ZT7 | Gene names | Chic2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cysteine-rich hydrophobic domain 2 protein. | |||||
|
CKAP2_HUMAN
|
||||||
| θ value | 0.813845 (rank : 10) | NC score | 0.065318 (rank : 7) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8WWK9, Q3KRA5, Q5VXB4, Q8IWV5, Q8IWV6, Q96FH9, Q9H012, Q9H0D0, Q9H988, Q9HC49, Q9NVG4 | Gene names | CKAP2, LB1, TMAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoskeleton-associated protein 2 (Tumor-associated microtubule- associated protein) (CTCL tumor antigen se20-10). | |||||
|
PLD1_HUMAN
|
||||||
| θ value | 0.813845 (rank : 11) | NC score | 0.050663 (rank : 8) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q13393 | Gene names | PLD1 | |||
|
Domain Architecture |

|
|||||
| Description | Phospholipase D1 (EC 3.1.4.4) (PLD 1) (Choline phosphatase 1) (Phosphatidylcholine-hydrolyzing phospholipase D1) (hPLD1). | |||||
|
SRCH_HUMAN
|
||||||
| θ value | 0.813845 (rank : 12) | NC score | 0.067112 (rank : 6) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P23327, Q504Y6 | Gene names | HRC, HCP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcoplasmic reticulum histidine-rich calcium-binding protein precursor. | |||||
|
ATX2_MOUSE
|
||||||
| θ value | 1.06291 (rank : 13) | NC score | 0.032285 (rank : 20) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O70305, P97421 | Gene names | Atxn2, Atx2, Sca2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-2 (Spinocerebellar ataxia type 2 protein homolog). | |||||
|
FILA_HUMAN
|
||||||
| θ value | 1.38821 (rank : 14) | NC score | 0.049202 (rank : 9) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 685 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P20930, Q01720, Q5T583 | Gene names | FLG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filaggrin. | |||||
|
EGR1_MOUSE
|
||||||
| θ value | 1.81305 (rank : 15) | NC score | 0.007866 (rank : 52) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 816 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P08046, Q61777 | Gene names | Egr1, Egr-1, Krox-24 | |||
|
Domain Architecture |

|
|||||
| Description | Early growth response protein 1 (EGR-1) (Krox-24 protein) (Transcription factor Zif268) (Nerve growth factor-induced protein A) (NGFI-A). | |||||
|
ZF106_MOUSE
|
||||||
| θ value | 1.81305 (rank : 16) | NC score | 0.021566 (rank : 33) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 472 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O88466, O55185, O88465, O88467, Q792M1, Q792P4, Q8CDZ8, Q8R3I4, Q9ESU3 | Gene names | Zfp106, H3a, Sh3bp3, Sirm, Znf474 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 106 (Zfp-106) (Zinc finger protein 474) (H3a minor histocompatibility antigen) (Son of insulin receptor mutant). | |||||
|
COAA1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 17) | NC score | 0.020745 (rank : 36) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 316 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q03692 | Gene names | COL10A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(X) chain precursor. | |||||
|
NUFP1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 18) | NC score | 0.032542 (rank : 18) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UHK0, Q8WVM5, Q96SG1 | Gene names | NUFIP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear fragile X mental retardation-interacting protein 1 (Nuclear FMRP-interacting protein 1). | |||||
|
RRBP1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 19) | NC score | 0.039840 (rank : 15) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1306 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9P2E9, O75300, O75301, Q5W165, Q96SB2, Q9BWP1, Q9H476 | Gene names | RRBP1, KIAA1398 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (180 kDa ribosome receptor homolog) (ES/130-related protein). | |||||
|
SFRS4_HUMAN
|
||||||
| θ value | 2.36792 (rank : 20) | NC score | 0.024557 (rank : 28) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q08170, Q5VXP1, Q9BUA4, Q9UEB5 | Gene names | SFRS4, SRP75 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 4 (Pre-mRNA-splicing factor SRP75) (SRP001LB). | |||||
|
CCD55_HUMAN
|
||||||
| θ value | 3.0926 (rank : 21) | NC score | 0.027772 (rank : 26) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9H0G5, Q6FI71 | Gene names | CCDC55 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 55. | |||||
|
GRIN1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 22) | NC score | 0.023074 (rank : 30) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q7Z2K8, Q8ND74, Q96PZ4 | Gene names | GPRIN1, KIAA1893 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein-regulated inducer of neurite outgrowth 1 (GRIN1). | |||||
|
NACAM_MOUSE
|
||||||
| θ value | 3.0926 (rank : 23) | NC score | 0.021508 (rank : 34) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
TDRD7_MOUSE
|
||||||
| θ value | 3.0926 (rank : 24) | NC score | 0.029704 (rank : 21) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8K1H1, Q8R181 | Gene names | Tdrd7, Pctaire2bp | |||
|
Domain Architecture |

|
|||||
| Description | Tudor domain-containing protein 7 (Tudor repeat associator with PCTAIRE 2) (Trap). | |||||
|
TNC6B_HUMAN
|
||||||
| θ value | 3.0926 (rank : 25) | NC score | 0.047518 (rank : 10) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 256 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UPQ9, Q5TH52, Q8TBX2 | Gene names | TNRC6B, KIAA1093 | |||
|
Domain Architecture |

|
|||||
| Description | Trinucleotide repeat-containing 6B protein. | |||||
|
ZN638_MOUSE
|
||||||
| θ value | 3.0926 (rank : 26) | NC score | 0.034278 (rank : 17) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q61464, Q6DFV9, Q8C941 | Gene names | Znf638, Np220, Zfml | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 638 (Nuclear protein 220) (Zinc-finger matrin-like protein). | |||||
|
CN155_HUMAN
|
||||||
| θ value | 4.03905 (rank : 27) | NC score | 0.022258 (rank : 32) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q5H9T9, Q5H9U7, Q86YI2, Q9H0J3 | Gene names | C14orf155 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf155. | |||||
|
CPNE7_HUMAN
|
||||||
| θ value | 4.03905 (rank : 28) | NC score | 0.015746 (rank : 44) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UBL6 | Gene names | CPNE7 | |||
|
Domain Architecture |

|
|||||
| Description | Copine-7 (Copine VII). | |||||
|
HNF1B_HUMAN
|
||||||
| θ value | 4.03905 (rank : 29) | NC score | 0.028314 (rank : 24) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P35680 | Gene names | TCF2, HNF1B | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte nuclear factor 1-beta (HNF-1beta) (HNF-1B) (Variant hepatic nuclear factor 1) (VHNF1) (Homeoprotein LFB3) (Transcription factor 2) (TCF-2). | |||||
|
HNF1B_MOUSE
|
||||||
| θ value | 4.03905 (rank : 30) | NC score | 0.028516 (rank : 23) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P27889, Q8R162, Q9CS26, Q9R1W1, Q9R1W2, Q9WTL5, Q9WTL6 | Gene names | Tcf2, Hnf-1b, Hnf1b | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte nuclear factor 1-beta (HNF-1beta) (HNF-1B) (Variant hepatic nuclear factor 1) (VHNF1) (Homeoprotein LFB3). | |||||
|
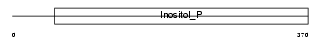
INPP_MOUSE
|
||||||
| θ value | 4.03905 (rank : 31) | NC score | 0.025881 (rank : 27) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P49442, Q8R3L1 | Gene names | Inpp1 | |||
|
Domain Architecture |
|
|||||
| Description | Inositol polyphosphate 1-phosphatase (EC 3.1.3.57) (IPPase) (IPP). | |||||
|
PEG3_MOUSE
|
||||||
| θ value | 4.03905 (rank : 32) | NC score | 0.008409 (rank : 51) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1101 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q3URU2, O54978, Q3TQ69, Q5EBP7, Q61138, Q6GQS0, Q80U47, Q8R5N0, Q9QX53 | Gene names | Peg3, Kiaa0287, Pw1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Paternally expressed gene 3 protein (ASF-1). | |||||
|
USP9X_HUMAN
|
||||||
| θ value | 4.03905 (rank : 33) | NC score | 0.020794 (rank : 35) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q93008, O75550, Q8WWT3, Q8WX12 | Gene names | USP9X, DFFRX, USP9 | |||
|
Domain Architecture |

|
|||||
| Description | Probable ubiquitin carboxyl-terminal hydrolase FAF-X (EC 3.1.2.15) (Ubiquitin thioesterase FAF-X) (Ubiquitin-specific-processing protease FAF-X) (Deubiquitinating enzyme FAF-X) (Fat facets protein-related, X- linked) (Ubiquitin-specific protease 9, X chromosome). | |||||
|
CYLC2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 34) | NC score | 0.046144 (rank : 11) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q14093 | Gene names | CYLC2, CYL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-2 (Cylicin II) (Multiple-band polypeptide II). | |||||
|
JARD2_HUMAN
|
||||||
| θ value | 5.27518 (rank : 35) | NC score | 0.018639 (rank : 39) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q92833, Q5U5L5 | Gene names | JARID2, JMJ | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji protein (Jumonji/ARID domain-containing protein 2). | |||||
|
KLHL1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 36) | NC score | 0.005146 (rank : 53) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 195 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NR64, Q5VZ64, Q9H4X4, Q9NR65, Q9P238 | Gene names | KLHL1, KIAA1490 | |||
|
Domain Architecture |

|
|||||
| Description | Kelch-like protein 1. | |||||
|
RPTN_HUMAN
|
||||||
| θ value | 5.27518 (rank : 37) | NC score | 0.028056 (rank : 25) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6XPR3 | Gene names | RPTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Repetin. | |||||
|
RYBP_HUMAN
|
||||||
| θ value | 5.27518 (rank : 38) | NC score | 0.022988 (rank : 31) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8N488, Q9P2W5, Q9UMW4 | Gene names | RYBP, DEDAF, YEAF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING1 and YY1-binding protein (Death effector domain-associated factor) (DED-associated factor) (YY1 and E4TF1-associated factor 1) (Apoptin-associating protein 1) (APAP-1). | |||||
|
BPA1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 39) | NC score | 0.010020 (rank : 50) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1477 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q91ZU6, Q91ZU7 | Gene names | Dst, Bpag1 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoforms 1/2/3/4 (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
BPAEA_MOUSE
|
||||||
| θ value | 6.88961 (rank : 40) | NC score | 0.010025 (rank : 49) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1480 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q91ZU8, Q8K5D4 | Gene names | Dst, Bpag1 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoform 5 (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
CENPF_HUMAN
|
||||||
| θ value | 6.88961 (rank : 41) | NC score | 0.012033 (rank : 47) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1932 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P49454, Q13171, Q13246 | Gene names | CENPF | |||
|
Domain Architecture |

|
|||||
| Description | Centromere protein F (Kinetochore protein CENP-F) (Mitosin) (AH antigen). | |||||
|
CXA5_MOUSE
|
||||||
| θ value | 6.88961 (rank : 42) | NC score | 0.013013 (rank : 45) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q01231 | Gene names | Gja5, Cxn-40 | |||
|
Domain Architecture |

|
|||||
| Description | Gap junction alpha-5 protein (Connexin-40) (Cx40). | |||||
|
MYCB2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 43) | NC score | 0.018192 (rank : 42) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q7TPH6, Q6PCM8 | Gene names | Mycbp2, Pam, Phr1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ubiquitin ligase protein MYCBP2 (EC 6.3.2.-) (Myc-binding protein 2) (Protein associated with Myc) (Pam/highwire/rpm-1 protein). | |||||
|
TEX2_MOUSE
|
||||||
| θ value | 6.88961 (rank : 44) | NC score | 0.018472 (rank : 40) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6ZPJ0 | Gene names | Tex2, Kiaa1738 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Testis-expressed sequence 2 protein. | |||||
|
USP9X_MOUSE
|
||||||
| θ value | 6.88961 (rank : 45) | NC score | 0.016256 (rank : 43) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P70398, Q62497 | Gene names | Usp9x, Fafl, Fam | |||
|
Domain Architecture |

|
|||||
| Description | Probable ubiquitin carboxyl-terminal hydrolase FAF-X (EC 3.1.2.15) (Ubiquitin thioesterase FAF-X) (Ubiquitin-specific-processing protease FAF-X) (Deubiquitinating enzyme FAF-X) (Fat facets protein-related, X- linked) (Ubiquitin-specific protease 9, X chromosome) (Ubiquitin carboxyl-terminal hydrolase FAM) (Fat facets homolog). | |||||
|
AFF4_MOUSE
|
||||||
| θ value | 8.99809 (rank : 46) | NC score | 0.032527 (rank : 19) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9ESC8, Q8C6K4, Q8C6W3, Q8CCH3 | Gene names | Aff4, Alf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4. | |||||
|
CIR_MOUSE
|
||||||
| θ value | 8.99809 (rank : 47) | NC score | 0.020040 (rank : 37) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9DA19, Q3V2N9, Q4KL44, Q52KL9, Q5FW66 | Gene names | Cir | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CBF1-interacting corepressor. | |||||
|
LUZP1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 48) | NC score | 0.011426 (rank : 48) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 739 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q86V48, Q5TH93, Q8N4X3, Q8TEH1 | Gene names | LUZP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine zipper protein 1. | |||||
|
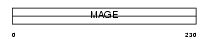
MAGB2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 49) | NC score | 0.012800 (rank : 46) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O15479, O75860 | Gene names | MAGEB2 | |||
|
Domain Architecture |
|
|||||
| Description | Melanoma-associated antigen B2 (MAGE-B2 antigen) (DSS-AHC critical interval MAGE superfamily 6) (DAM6) (MAGE XP-2). | |||||
|
MTSS1_HUMAN
|
||||||
| θ value | 8.99809 (rank : 50) | NC score | 0.019433 (rank : 38) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O43312, Q8TCA2, Q96RX2 | Gene names | MTSS1, KIAA0429, MIM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metastasis suppressor protein 1 (Missing in metastasis protein) (Metastasis suppressor YGL-1). | |||||
|
MTSS1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 51) | NC score | 0.018467 (rank : 41) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 592 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8R1S4, Q8BMM3, Q99LB3 | Gene names | Mtss1, Mim | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metastasis suppressor protein 1 (Missing in metastasis protein). | |||||
|
NSBP1_MOUSE
|
||||||
| θ value | 8.99809 (rank : 52) | NC score | 0.024341 (rank : 29) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9JL35, O88832, Q8VC71, Q9CUW1 | Gene names | Nsbp1, Garp45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1 (Nucleosome-binding protein 45) (NBP-45) (GARP45 protein). | |||||
|
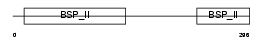
SIAL_HUMAN
|
||||||
| θ value | 8.99809 (rank : 53) | NC score | 0.028973 (rank : 22) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P21815 | Gene names | IBSP, BNSP | |||
|
Domain Architecture |
|
|||||
| Description | Bone sialoprotein-2 precursor (Bone sialoprotein II) (BSP II) (Cell- binding sialoprotein) (Integrin-binding sialoprotein). | |||||
|
NUFP2_MOUSE
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | Q5F2E7, Q3TCE2, Q3V195, Q80TF1 | Gene names | Nufip2, Kiaa1321 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear fragile X mental retardation-interacting protein 2 (FMRP- interacting protein 2) (82 kDa FMRP-interacting protein) (82-FIP). | |||||
|
NUFP2_HUMAN
|
||||||
| NC score | 0.986251 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 62 | Shared Neighborhood Hits | 26 | |
| SwissProt Accessions | Q7Z417, Q9P2M5 | Gene names | NUFIP2, KIAA1321, PIG1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear fragile X mental retardation-interacting protein 2 (FMRP- interacting protein 2) (82 kDa FMRP-interacting protein) (82-FIP) (Proliferation-inducing gene 1 protein). | |||||
|
CYLC1_HUMAN
|
||||||
| NC score | 0.086892 (rank : 3) | θ value | 0.279714 (rank : 5) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 408 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P35663, Q5JQQ9 | Gene names | CYLC1, CYL, CYL1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-1 (Cylicin I) (Multiple-band polypeptide I). | |||||
|
CHIC2_MOUSE
|
||||||
| NC score | 0.073747 (rank : 4) | θ value | 0.813845 (rank : 9) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9D9G3, Q3T9U2, Q80ZT7 | Gene names | Chic2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cysteine-rich hydrophobic domain 2 protein. | |||||
|
CHIC2_HUMAN
|
||||||
| NC score | 0.073667 (rank : 5) | θ value | 0.813845 (rank : 8) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 10 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q9UKJ5 | Gene names | CHIC2, BTL | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cysteine-rich hydrophobic domain 2 protein (BrX-like translocated in leukemia). | |||||
|
SRCH_HUMAN
|
||||||
| NC score | 0.067112 (rank : 6) | θ value | 0.813845 (rank : 12) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 225 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | P23327, Q504Y6 | Gene names | HRC, HCP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Sarcoplasmic reticulum histidine-rich calcium-binding protein precursor. | |||||
|
CKAP2_HUMAN
|
||||||
| NC score | 0.065318 (rank : 7) | θ value | 0.813845 (rank : 10) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 116 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q8WWK9, Q3KRA5, Q5VXB4, Q8IWV5, Q8IWV6, Q96FH9, Q9H012, Q9H0D0, Q9H988, Q9HC49, Q9NVG4 | Gene names | CKAP2, LB1, TMAP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cytoskeleton-associated protein 2 (Tumor-associated microtubule- associated protein) (CTCL tumor antigen se20-10). | |||||
|
PLD1_HUMAN
|
||||||
| NC score | 0.050663 (rank : 8) | θ value | 0.813845 (rank : 11) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 18 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q13393 | Gene names | PLD1 | |||
|
Domain Architecture |

|
|||||
| Description | Phospholipase D1 (EC 3.1.4.4) (PLD 1) (Choline phosphatase 1) (Phosphatidylcholine-hydrolyzing phospholipase D1) (hPLD1). | |||||
|
FILA_HUMAN
|
||||||
| NC score | 0.049202 (rank : 9) | θ value | 1.38821 (rank : 14) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 685 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P20930, Q01720, Q5T583 | Gene names | FLG | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Filaggrin. | |||||
|
TNC6B_HUMAN
|
||||||
| NC score | 0.047518 (rank : 10) | θ value | 3.0926 (rank : 25) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 256 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9UPQ9, Q5TH52, Q8TBX2 | Gene names | TNRC6B, KIAA1093 | |||
|
Domain Architecture |

|
|||||
| Description | Trinucleotide repeat-containing 6B protein. | |||||
|
CYLC2_HUMAN
|
||||||
| NC score | 0.046144 (rank : 11) | θ value | 5.27518 (rank : 34) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 223 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q14093 | Gene names | CYLC2, CYL2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Cylicin-2 (Cylicin II) (Multiple-band polypeptide II). | |||||
|
RFIP1_HUMAN
|
||||||
| NC score | 0.046022 (rank : 12) | θ value | 0.279714 (rank : 6) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 236 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6WKZ4, Q6AZK4, Q6WKZ2, Q6WKZ6, Q86YV4, Q8TDL1, Q9H642 | Gene names | RAB11FIP1, RCP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rab11 family-interacting protein 1 (Rab11-FIP1) (Rab-coupling protein). | |||||
|
RRBP1_MOUSE
|
||||||
| NC score | 0.044543 (rank : 13) | θ value | 0.0252991 (rank : 4) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1650 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q99PL5, Q99PK5, Q99PK6, Q99PK7, Q99PK8, Q99PK9, Q99PL0, Q99PL1, Q99PL2, Q99PL3, Q99PL4, Q9CS20 | Gene names | Rrbp1 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (mRRp). | |||||
|
CD44_MOUSE
|
||||||
| NC score | 0.042498 (rank : 14) | θ value | 0.813845 (rank : 7) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 64 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P15379, Q05732, Q61395, Q62060, Q62061, Q62062, Q62063, Q62408, Q62409, Q64296, Q99J14, Q9QYX8 | Gene names | Cd44, Ly-24 | |||
|
Domain Architecture |

|
|||||
| Description | CD44 antigen precursor (Phagocytic glycoprotein 1) (PGP-1) (HUTCH-I) (Extracellular matrix receptor III) (ECMR-III) (GP90 lymphocyte homing/adhesion receptor) (Hermes antigen) (Hyaluronate receptor) (Lymphocyte antigen Ly-24). | |||||
|
RRBP1_HUMAN
|
||||||
| NC score | 0.039840 (rank : 15) | θ value | 2.36792 (rank : 19) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1306 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9P2E9, O75300, O75301, Q5W165, Q96SB2, Q9BWP1, Q9H476 | Gene names | RRBP1, KIAA1398 | |||
|
Domain Architecture |

|
|||||
| Description | Ribosome-binding protein 1 (Ribosome receptor protein) (180 kDa ribosome receptor homolog) (ES/130-related protein). | |||||
|
DDX21_MOUSE
|
||||||
| NC score | 0.034556 (rank : 16) | θ value | 0.0113563 (rank : 3) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 350 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9JIK5, Q9WV45 | Gene names | Ddx21 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar RNA helicase 2 (EC 3.6.1.-) (Nucleolar RNA helicase II) (Nucleolar RNA helicase Gu) (RH II/Gu) (Gu-alpha) (DEAD box protein 21). | |||||
|
ZN638_MOUSE
|
||||||
| NC score | 0.034278 (rank : 17) | θ value | 3.0926 (rank : 26) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 318 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q61464, Q6DFV9, Q8C941 | Gene names | Znf638, Np220, Zfml | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 638 (Nuclear protein 220) (Zinc-finger matrin-like protein). | |||||
|
NUFP1_HUMAN
|
||||||
| NC score | 0.032542 (rank : 18) | θ value | 2.36792 (rank : 18) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 193 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9UHK0, Q8WVM5, Q96SG1 | Gene names | NUFIP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nuclear fragile X mental retardation-interacting protein 1 (Nuclear FMRP-interacting protein 1). | |||||
|
AFF4_MOUSE
|
||||||
| NC score | 0.032527 (rank : 19) | θ value | 8.99809 (rank : 46) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 581 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q9ESC8, Q8C6K4, Q8C6W3, Q8CCH3 | Gene names | Aff4, Alf4 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AF4/FMR2 family member 4. | |||||
|
ATX2_MOUSE
|
||||||
| NC score | 0.032285 (rank : 20) | θ value | 1.06291 (rank : 13) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 445 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | O70305, P97421 | Gene names | Atxn2, Atx2, Sca2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Ataxin-2 (Spinocerebellar ataxia type 2 protein homolog). | |||||
|
TDRD7_MOUSE
|
||||||
| NC score | 0.029704 (rank : 21) | θ value | 3.0926 (rank : 24) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8K1H1, Q8R181 | Gene names | Tdrd7, Pctaire2bp | |||
|
Domain Architecture |

|
|||||
| Description | Tudor domain-containing protein 7 (Tudor repeat associator with PCTAIRE 2) (Trap). | |||||
|
SIAL_HUMAN
|
||||||
| NC score | 0.028973 (rank : 22) | θ value | 8.99809 (rank : 53) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P21815 | Gene names | IBSP, BNSP | |||
|
Domain Architecture |

|
|||||
| Description | Bone sialoprotein-2 precursor (Bone sialoprotein II) (BSP II) (Cell- binding sialoprotein) (Integrin-binding sialoprotein). | |||||
|
HNF1B_MOUSE
|
||||||
| NC score | 0.028516 (rank : 23) | θ value | 4.03905 (rank : 30) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 78 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P27889, Q8R162, Q9CS26, Q9R1W1, Q9R1W2, Q9WTL5, Q9WTL6 | Gene names | Tcf2, Hnf-1b, Hnf1b | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte nuclear factor 1-beta (HNF-1beta) (HNF-1B) (Variant hepatic nuclear factor 1) (VHNF1) (Homeoprotein LFB3). | |||||
|
HNF1B_HUMAN
|
||||||
| NC score | 0.028314 (rank : 24) | θ value | 4.03905 (rank : 29) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 80 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P35680 | Gene names | TCF2, HNF1B | |||
|
Domain Architecture |

|
|||||
| Description | Hepatocyte nuclear factor 1-beta (HNF-1beta) (HNF-1B) (Variant hepatic nuclear factor 1) (VHNF1) (Homeoprotein LFB3) (Transcription factor 2) (TCF-2). | |||||
|
RPTN_HUMAN
|
||||||
| NC score | 0.028056 (rank : 25) | θ value | 5.27518 (rank : 37) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 406 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q6XPR3 | Gene names | RPTN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Repetin. | |||||
|
CCD55_HUMAN
|
||||||
| NC score | 0.027772 (rank : 26) | θ value | 3.0926 (rank : 21) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 513 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q9H0G5, Q6FI71 | Gene names | CCDC55 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 55. | |||||
|
INPP_MOUSE
|
||||||
| NC score | 0.025881 (rank : 27) | θ value | 4.03905 (rank : 31) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 16 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P49442, Q8R3L1 | Gene names | Inpp1 | |||
|
Domain Architecture |

|
|||||
| Description | Inositol polyphosphate 1-phosphatase (EC 3.1.3.57) (IPPase) (IPP). | |||||
|
SFRS4_HUMAN
|
||||||
| NC score | 0.024557 (rank : 28) | θ value | 2.36792 (rank : 20) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 384 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q08170, Q5VXP1, Q9BUA4, Q9UEB5 | Gene names | SFRS4, SRP75 | |||
|
Domain Architecture |

|
|||||
| Description | Splicing factor, arginine/serine-rich 4 (Pre-mRNA-splicing factor SRP75) (SRP001LB). | |||||
|
NSBP1_MOUSE
|
||||||
| NC score | 0.024341 (rank : 29) | θ value | 8.99809 (rank : 52) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q9JL35, O88832, Q8VC71, Q9CUW1 | Gene names | Nsbp1, Garp45 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nucleosome-binding protein 1 (Nucleosome-binding protein 45) (NBP-45) (GARP45 protein). | |||||
|
GRIN1_HUMAN
|
||||||
| NC score | 0.023074 (rank : 30) | θ value | 3.0926 (rank : 22) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 681 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q7Z2K8, Q8ND74, Q96PZ4 | Gene names | GPRIN1, KIAA1893 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | G protein-regulated inducer of neurite outgrowth 1 (GRIN1). | |||||
|
RYBP_HUMAN
|
||||||
| NC score | 0.022988 (rank : 31) | θ value | 5.27518 (rank : 38) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 50 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q8N488, Q9P2W5, Q9UMW4 | Gene names | RYBP, DEDAF, YEAF1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RING1 and YY1-binding protein (Death effector domain-associated factor) (DED-associated factor) (YY1 and E4TF1-associated factor 1) (Apoptin-associating protein 1) (APAP-1). | |||||
|
CN155_HUMAN
|
||||||
| NC score | 0.022258 (rank : 32) | θ value | 4.03905 (rank : 27) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 385 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q5H9T9, Q5H9U7, Q86YI2, Q9H0J3 | Gene names | C14orf155 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein C14orf155. | |||||
|
ZF106_MOUSE
|
||||||
| NC score | 0.021566 (rank : 33) | θ value | 1.81305 (rank : 16) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 472 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O88466, O55185, O88465, O88467, Q792M1, Q792P4, Q8CDZ8, Q8R3I4, Q9ESU3 | Gene names | Zfp106, H3a, Sh3bp3, Sirm, Znf474 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 106 (Zfp-106) (Zinc finger protein 474) (H3a minor histocompatibility antigen) (Son of insulin receptor mutant). | |||||
|
NACAM_MOUSE
|
||||||
| NC score | 0.021508 (rank : 34) | θ value | 3.0926 (rank : 23) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1866 | Shared Neighborhood Hits | 25 | |
| SwissProt Accessions | P70670 | Gene names | Naca | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Nascent polypeptide-associated complex subunit alpha, muscle-specific form (Alpha-NAC, muscle-specific form). | |||||
|
USP9X_HUMAN
|
||||||
| NC score | 0.020794 (rank : 35) | θ value | 4.03905 (rank : 33) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 161 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | Q93008, O75550, Q8WWT3, Q8WX12 | Gene names | USP9X, DFFRX, USP9 | |||
|
Domain Architecture |

|
|||||
| Description | Probable ubiquitin carboxyl-terminal hydrolase FAF-X (EC 3.1.2.15) (Ubiquitin thioesterase FAF-X) (Ubiquitin-specific-processing protease FAF-X) (Deubiquitinating enzyme FAF-X) (Fat facets protein-related, X- linked) (Ubiquitin-specific protease 9, X chromosome). | |||||
|
COAA1_HUMAN
|
||||||
| NC score | 0.020745 (rank : 36) | θ value | 2.36792 (rank : 17) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 316 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q03692 | Gene names | COL10A1 | |||
|
Domain Architecture |

|
|||||
| Description | Collagen alpha-1(X) chain precursor. | |||||
|
CIR_MOUSE
|
||||||
| NC score | 0.020040 (rank : 37) | θ value | 8.99809 (rank : 47) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 255 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q9DA19, Q3V2N9, Q4KL44, Q52KL9, Q5FW66 | Gene names | Cir | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | CBF1-interacting corepressor. | |||||
|
MTSS1_HUMAN
|
||||||
| NC score | 0.019433 (rank : 38) | θ value | 8.99809 (rank : 50) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 504 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | O43312, Q8TCA2, Q96RX2 | Gene names | MTSS1, KIAA0429, MIM | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metastasis suppressor protein 1 (Missing in metastasis protein) (Metastasis suppressor YGL-1). | |||||
|
JARD2_HUMAN
|
||||||
| NC score | 0.018639 (rank : 39) | θ value | 5.27518 (rank : 35) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 119 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q92833, Q5U5L5 | Gene names | JARID2, JMJ | |||
|
Domain Architecture |

|
|||||
| Description | Jumonji protein (Jumonji/ARID domain-containing protein 2). | |||||
|
TEX2_MOUSE
|
||||||
| NC score | 0.018472 (rank : 40) | θ value | 6.88961 (rank : 44) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 121 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q6ZPJ0 | Gene names | Tex2, Kiaa1738 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Testis-expressed sequence 2 protein. | |||||
|
MTSS1_MOUSE
|
||||||
| NC score | 0.018467 (rank : 41) | θ value | 8.99809 (rank : 51) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 592 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q8R1S4, Q8BMM3, Q99LB3 | Gene names | Mtss1, Mim | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Metastasis suppressor protein 1 (Missing in metastasis protein). | |||||
|
MYCB2_MOUSE
|
||||||
| NC score | 0.018192 (rank : 42) | θ value | 6.88961 (rank : 43) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q7TPH6, Q6PCM8 | Gene names | Mycbp2, Pam, Phr1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable ubiquitin ligase protein MYCBP2 (EC 6.3.2.-) (Myc-binding protein 2) (Protein associated with Myc) (Pam/highwire/rpm-1 protein). | |||||
|
USP9X_MOUSE
|
||||||
| NC score | 0.016256 (rank : 43) | θ value | 6.88961 (rank : 45) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 171 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P70398, Q62497 | Gene names | Usp9x, Fafl, Fam | |||
|
Domain Architecture |

|
|||||
| Description | Probable ubiquitin carboxyl-terminal hydrolase FAF-X (EC 3.1.2.15) (Ubiquitin thioesterase FAF-X) (Ubiquitin-specific-processing protease FAF-X) (Deubiquitinating enzyme FAF-X) (Fat facets protein-related, X- linked) (Ubiquitin-specific protease 9, X chromosome) (Ubiquitin carboxyl-terminal hydrolase FAM) (Fat facets homolog). | |||||
|
CPNE7_HUMAN
|
||||||
| NC score | 0.015746 (rank : 44) | θ value | 4.03905 (rank : 28) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 92 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9UBL6 | Gene names | CPNE7 | |||
|
Domain Architecture |

|
|||||
| Description | Copine-7 (Copine VII). | |||||
|
CXA5_MOUSE
|
||||||
| NC score | 0.013013 (rank : 45) | θ value | 6.88961 (rank : 42) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 41 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q01231 | Gene names | Gja5, Cxn-40 | |||
|
Domain Architecture |

|
|||||
| Description | Gap junction alpha-5 protein (Connexin-40) (Cx40). | |||||
|
MAGB2_HUMAN
|
||||||
| NC score | 0.012800 (rank : 46) | θ value | 8.99809 (rank : 49) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 55 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | O15479, O75860 | Gene names | MAGEB2 | |||
|
Domain Architecture |

|
|||||
| Description | Melanoma-associated antigen B2 (MAGE-B2 antigen) (DSS-AHC critical interval MAGE superfamily 6) (DAM6) (MAGE XP-2). | |||||
|
CENPF_HUMAN
|
||||||
| NC score | 0.012033 (rank : 47) | θ value | 6.88961 (rank : 41) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1932 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P49454, Q13171, Q13246 | Gene names | CENPF | |||
|
Domain Architecture |

|
|||||
| Description | Centromere protein F (Kinetochore protein CENP-F) (Mitosin) (AH antigen). | |||||
|
LUZP1_HUMAN
|
||||||
| NC score | 0.011426 (rank : 48) | θ value | 8.99809 (rank : 48) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 739 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q86V48, Q5TH93, Q8N4X3, Q8TEH1 | Gene names | LUZP1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Leucine zipper protein 1. | |||||
|
BPAEA_MOUSE
|
||||||
| NC score | 0.010025 (rank : 49) | θ value | 6.88961 (rank : 40) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1480 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q91ZU8, Q8K5D4 | Gene names | Dst, Bpag1 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoform 5 (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
BPA1_MOUSE
|
||||||
| NC score | 0.010020 (rank : 50) | θ value | 6.88961 (rank : 39) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1477 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q91ZU6, Q91ZU7 | Gene names | Dst, Bpag1 | |||
|
Domain Architecture |

|
|||||
| Description | Bullous pemphigoid antigen 1, isoforms 1/2/3/4 (BPA) (Hemidesmosomal plaque protein) (Dystonia musculorum protein) (Dystonin). | |||||
|
PEG3_MOUSE
|
||||||
| NC score | 0.008409 (rank : 51) | θ value | 4.03905 (rank : 32) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1101 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q3URU2, O54978, Q3TQ69, Q5EBP7, Q61138, Q6GQS0, Q80U47, Q8R5N0, Q9QX53 | Gene names | Peg3, Kiaa0287, Pw1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Paternally expressed gene 3 protein (ASF-1). | |||||
|
EGR1_MOUSE
|
||||||
| NC score | 0.007866 (rank : 52) | θ value | 1.81305 (rank : 15) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 816 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | P08046, Q61777 | Gene names | Egr1, Egr-1, Krox-24 | |||
|
Domain Architecture |

|
|||||
| Description | Early growth response protein 1 (EGR-1) (Krox-24 protein) (Transcription factor Zif268) (Nerve growth factor-induced protein A) (NGFI-A). | |||||
|
KLHL1_HUMAN
|
||||||
| NC score | 0.005146 (rank : 53) | θ value | 5.27518 (rank : 36) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 195 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9NR64, Q5VZ64, Q9H4X4, Q9NR65, Q9P238 | Gene names | KLHL1, KIAA1490 | |||
|
Domain Architecture |

|
|||||
| Description | Kelch-like protein 1. | |||||