Please be patient as the page loads
|
LCAP_HUMAN
|
||||||
| SwissProt Accessions | Q9UIQ6, O00769, Q15145, Q9UIQ7 | Gene names | LNPEP, OTASE | |||
|
Domain Architecture |
|
|||||
| Description | Leucyl-cystinyl aminopeptidase (EC 3.4.11.3) (Cystinyl aminopeptidase) (Oxytocinase) (OTase) (Insulin-regulated membrane aminopeptidase) (Insulin-responsive aminopeptidase) (IRAP) (Placental leucine aminopeptidase) (P-LAP). | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
ARTS1_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.990507 (rank : 2) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9NZ08, O60278, Q6UWY6, Q8NEL4, Q8TAD0, Q9UHF8, Q9UKY2 | Gene names | ARTS1, APPILS, KIAA0525 | |||
|
Domain Architecture |

|
|||||
| Description | Adipocyte-derived leucine aminopeptidase precursor (EC 3.4.11.-) (A- LAP) (ARTS-1) (Aminopeptidase PILS) (Puromycin-insensitive leucyl- specific aminopeptidase) (PILS-AP) (Type 1 tumor necrosis factor receptor shedding aminopeptidase regulator). | |||||
|
ARTS1_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.989996 (rank : 3) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9EQH2, Q9ET63 | Gene names | Arts1, Appils | |||
|
Domain Architecture |

|
|||||
| Description | Adipocyte-derived leucine aminopeptidase precursor (EC 3.4.11.-) (A- LAP) (ARTS-1) (Aminopeptidase PILS) (Puromycin-insensitive leucyl- specific aminopeptidase) (PILS-AP) (VEGF-induced aminopeptidase). | |||||
|
LCAP_HUMAN
|
||||||
| θ value | 0 (rank : 3) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9UIQ6, O00769, Q15145, Q9UIQ7 | Gene names | LNPEP, OTASE | |||
|
Domain Architecture |

|
|||||
| Description | Leucyl-cystinyl aminopeptidase (EC 3.4.11.3) (Cystinyl aminopeptidase) (Oxytocinase) (OTase) (Insulin-regulated membrane aminopeptidase) (Insulin-responsive aminopeptidase) (IRAP) (Placental leucine aminopeptidase) (P-LAP). | |||||
|
AMPN_MOUSE
|
||||||
| θ value | 1.02182e-136 (rank : 4) | NC score | 0.975744 (rank : 4) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P97449, Q91YH8, Q99K63 | Gene names | Anpep, Lap-1, Lap1 | |||
|
Domain Architecture |
|
|||||
| Description | Aminopeptidase N (EC 3.4.11.2) (mAPN) (Alanyl aminopeptidase) (Microsomal aminopeptidase) (Aminopeptidase M) (Membrane protein p161) (CD13 antigen). | |||||
|
AMPE_HUMAN
|
||||||
| θ value | 1.80368e-133 (rank : 5) | NC score | 0.974423 (rank : 6) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q07075 | Gene names | ENPEP | |||
|
Domain Architecture |
|
|||||
| Description | Glutamyl aminopeptidase (EC 3.4.11.7) (EAP) (Aminopeptidase A) (APA) (Differentiation antigen gp160) (CD249 antigen). | |||||
|
AMPN_HUMAN
|
||||||
| θ value | 4.44261e-132 (rank : 6) | NC score | 0.975095 (rank : 5) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P15144, Q16728, Q8IUK3, Q8IVH3, Q9UCE0 | Gene names | ANPEP, APN, CD13, PEPN | |||
|
Domain Architecture |
|
|||||
| Description | Aminopeptidase N (EC 3.4.11.2) (hAPN) (Alanyl aminopeptidase) (Microsomal aminopeptidase) (Aminopeptidase M) (gp150) (Myeloid plasma membrane glycoprotein CD13) (CD13 antigen). | |||||
|
AMPE_MOUSE
|
||||||
| θ value | 9.26343e-130 (rank : 7) | NC score | 0.973706 (rank : 7) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P16406 | Gene names | Enpep | |||
|
Domain Architecture |
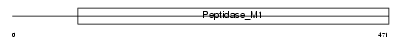
|
|||||
| Description | Glutamyl aminopeptidase (EC 3.4.11.7) (EAP) (Aminopeptidase A) (APA) (BP-1/6C3 antigen). | |||||
|
TRHDE_MOUSE
|
||||||
| θ value | 2.36115e-125 (rank : 8) | NC score | 0.967636 (rank : 10) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8K093 | Gene names | Trhde | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyrotropin-releasing hormone-degrading ectoenzyme (EC 3.4.19.6) (TRH- degrading ectoenzyme) (TRH-DE) (TRH-specific aminopeptidase) (Thyroliberinase) (Pyroglutamyl-peptidase II) (PAP-II). | |||||
|
TRHDE_HUMAN
|
||||||
| θ value | 2.61056e-124 (rank : 9) | NC score | 0.969045 (rank : 8) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9UKU6, Q6UWJ4 | Gene names | TRHDE | |||
|
Domain Architecture |

|
|||||
| Description | Thyrotropin-releasing hormone-degrading ectoenzyme (EC 3.4.19.6) (TRH- degrading ectoenzyme) (TRH-DE) (TRH-specific aminopeptidase) (Thyroliberinase) (Pyroglutamyl-peptidase II) (PAP-II). | |||||
|
PSA_HUMAN
|
||||||
| θ value | 1.34386e-112 (rank : 10) | NC score | 0.967735 (rank : 9) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P55786, Q6P145, Q9NP16, Q9UEM2 | Gene names | NPEPPS, PSA | |||
|
Domain Architecture |

|
|||||
| Description | Puromycin-sensitive aminopeptidase (EC 3.4.11.-) (PSA). | |||||
|
PSA_MOUSE
|
||||||
| θ value | 1.13764e-111 (rank : 11) | NC score | 0.967603 (rank : 11) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q11011, Q91VJ8 | Gene names | Npepps, Psa | |||
|
Domain Architecture |

|
|||||
| Description | Puromycin-sensitive aminopeptidase (EC 3.4.11.-) (PSA). | |||||
|
LAEVR_HUMAN
|
||||||
| θ value | 2.62879e-100 (rank : 12) | NC score | 0.961729 (rank : 12) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6Q4G3, Q32MR1, Q4G0I9, Q4G0V2, Q86XA3, Q8NBZ2 | Gene names | LVRN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Laeverin (EC 3.4.-.-) (CHL2 antigen). | |||||
|
LAEVR_MOUSE
|
||||||
| θ value | 5.53243e-66 (rank : 13) | NC score | 0.956069 (rank : 13) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q2KHK3, Q9D633 | Gene names | Lvrn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Laeverin (EC 3.4.-.-). | |||||
|
LKHA4_HUMAN
|
||||||
| θ value | 4.73814e-25 (rank : 14) | NC score | 0.731280 (rank : 14) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P09960 | Gene names | LTA4H, LTA4 | |||
|
Domain Architecture |

|
|||||
| Description | Leukotriene A-4 hydrolase (EC 3.3.2.6) (LTA-4 hydrolase) (Leukotriene A(4) hydrolase). | |||||
|
LKHA4_MOUSE
|
||||||
| θ value | 3.75424e-22 (rank : 15) | NC score | 0.724031 (rank : 15) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P24527, Q8VDR8 | Gene names | Lta4h | |||
|
Domain Architecture |

|
|||||
| Description | Leukotriene A-4 hydrolase (EC 3.3.2.6) (LTA-4 hydrolase) (Leukotriene A(4) hydrolase). | |||||
|
AMPB_MOUSE
|
||||||
| θ value | 8.65492e-19 (rank : 16) | NC score | 0.691912 (rank : 16) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8VCT3 | Gene names | Rnpep | |||
|
Domain Architecture |

|
|||||
| Description | Aminopeptidase B (EC 3.4.11.6) (Ap-B) (Arginyl aminopeptidase) (Arginine aminopeptidase) (Cytosol aminopeptidase IV). | |||||
|
AMPB_HUMAN
|
||||||
| θ value | 1.47631e-18 (rank : 17) | NC score | 0.679070 (rank : 17) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9H4A4, Q9BVM9, Q9H1D4, Q9NPT7 | Gene names | RNPEP, APB | |||
|
Domain Architecture |

|
|||||
| Description | Aminopeptidase B (EC 3.4.11.6) (Ap-B) (Arginyl aminopeptidase) (Arginine aminopeptidase). | |||||
|
RNPL1_HUMAN
|
||||||
| θ value | 2.28291e-11 (rank : 18) | NC score | 0.650127 (rank : 18) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9HAU8, Q96AC9, Q9H033, Q9NVD0 | Gene names | RNPEPL1 | |||
|
Domain Architecture |

|
|||||
| Description | Arginyl aminopeptidase-like 1 (EC 3.4.11.-) (RNPEP-like protein). | |||||
|
AMPO_MOUSE
|
||||||
| θ value | 1.80886e-08 (rank : 19) | NC score | 0.474836 (rank : 20) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8BXQ6, Q6P0W6, Q6P394, Q8BHX5 | Gene names | Onpep | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Aminopeptidase O (EC 3.4.11.-) (AP-O). | |||||
|
AMPO_HUMAN
|
||||||
| θ value | 5.26297e-08 (rank : 20) | NC score | 0.486437 (rank : 19) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8N6M6, Q5T9B1, Q5T9B3, Q5T9B4, Q8WUL6, Q96M23, Q96SS1 | Gene names | ONPEP, C9orf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Aminopeptidase O (EC 3.4.11.-) (AP-O). | |||||
|
TAF2_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 21) | NC score | 0.229379 (rank : 21) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q6P1X5, O43487, O43604, O60668, Q86WW7, Q8IWK4 | Gene names | TAF2, CIF150, TAF2B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID subunit 2 (Transcription initiation factor TFIID 150 kDa subunit) (TBP-associated factor 150 kDa) (TAFII-150) (TAFII150) (150 kDa cofactor of initiator function). | |||||
|
TAF2_MOUSE
|
||||||
| θ value | 0.00509761 (rank : 22) | NC score | 0.221307 (rank : 22) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8C176, Q8BKQ7 | Gene names | Taf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID subunit 2 (Transcription initiation factor TFIID 150 kDa subunit) (TBP-associated factor 150 kDa). | |||||
|
RPA2_HUMAN
|
||||||
| θ value | 0.125558 (rank : 23) | NC score | 0.030705 (rank : 23) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H9Y6, Q585T5, Q6ZRR2, Q9H9D3 | Gene names | POLR1B | |||
|
Domain Architecture |

|
|||||
| Description | DNA-directed RNA polymerase I 135 kDa polypeptide (EC 2.7.7.6) (RNA polymerase I subunit 2) (RPA135). | |||||
|
RPA2_MOUSE
|
||||||
| θ value | 1.06291 (rank : 24) | NC score | 0.026308 (rank : 24) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P70700 | Gene names | Polr1b, Rpa2, Rpo1-2 | |||
|
Domain Architecture |
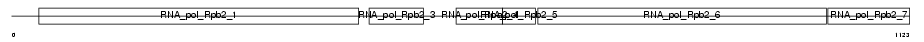
|
|||||
| Description | DNA-directed RNA polymerase I 135 kDa polypeptide (EC 2.7.7.6) (RNA polymerase I subunit 2) (RPA135). | |||||
|
PKHA6_HUMAN
|
||||||
| θ value | 5.27518 (rank : 25) | NC score | 0.004666 (rank : 30) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 556 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y2H5 | Gene names | PLEKHA6, KIAA0969, PEPP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family A member 6 (Phosphoinositol 3-phosphate-binding protein 3) (PEPP-3). | |||||
|
RA6I1_MOUSE
|
||||||
| θ value | 5.27518 (rank : 26) | NC score | 0.007689 (rank : 26) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6PAL8, Q62146, Q8C829, Q8VDF6, Q9QYZ2 | Gene names | Rab6ip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rab6-interacting protein 1 (Rab6IP1). | |||||
|
PIGQ_MOUSE
|
||||||
| θ value | 6.88961 (rank : 27) | NC score | 0.009817 (rank : 25) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 5 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9QYT7, O35120, O35456, Q99L11 | Gene names | Pigq, Gpi1h, Mgpi1 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol N-acetylglucosaminyltransferase subunit Q (EC 2.4.1.198) (Phosphatidylinositol-glycan biosynthesis class Q protein) (PIG-Q) (N-acetylglucosamyl transferase component GPI1) (MGpi1p). | |||||
|
RTN4_MOUSE
|
||||||
| θ value | 6.88961 (rank : 28) | NC score | 0.005313 (rank : 28) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99P72, Q5DTK9, Q7TNB7, Q80W95, Q8BGK7, Q8BGM9, Q8K290, Q8K3G8, Q9CTE3 | Gene names | Rtn4, Kiaa0886, Nogo | |||
|
Domain Architecture |

|
|||||
| Description | Reticulon-4 (Neurite outgrowth inhibitor) (Nogo protein). | |||||
|
A1BG_HUMAN
|
||||||
| θ value | 8.99809 (rank : 29) | NC score | 0.005311 (rank : 29) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P04217, Q8IYJ6, Q96P39 | Gene names | A1BG | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-1B-glycoprotein precursor (Alpha-1-B glycoprotein). | |||||
|
NU107_MOUSE
|
||||||
| θ value | 8.99809 (rank : 30) | NC score | 0.007014 (rank : 27) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BH74, Q99KH5 | Gene names | Nup107 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup107 (Nucleoporin Nup107) (107 kDa nucleoporin). | |||||
|
LCAP_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 3) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q9UIQ6, O00769, Q15145, Q9UIQ7 | Gene names | LNPEP, OTASE | |||
|
Domain Architecture |

|
|||||
| Description | Leucyl-cystinyl aminopeptidase (EC 3.4.11.3) (Cystinyl aminopeptidase) (Oxytocinase) (OTase) (Insulin-regulated membrane aminopeptidase) (Insulin-responsive aminopeptidase) (IRAP) (Placental leucine aminopeptidase) (P-LAP). | |||||
|
ARTS1_HUMAN
|
||||||
| NC score | 0.990507 (rank : 2) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 27 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9NZ08, O60278, Q6UWY6, Q8NEL4, Q8TAD0, Q9UHF8, Q9UKY2 | Gene names | ARTS1, APPILS, KIAA0525 | |||
|
Domain Architecture |

|
|||||
| Description | Adipocyte-derived leucine aminopeptidase precursor (EC 3.4.11.-) (A- LAP) (ARTS-1) (Aminopeptidase PILS) (Puromycin-insensitive leucyl- specific aminopeptidase) (PILS-AP) (Type 1 tumor necrosis factor receptor shedding aminopeptidase regulator). | |||||
|
ARTS1_MOUSE
|
||||||
| NC score | 0.989996 (rank : 3) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9EQH2, Q9ET63 | Gene names | Arts1, Appils | |||
|
Domain Architecture |

|
|||||
| Description | Adipocyte-derived leucine aminopeptidase precursor (EC 3.4.11.-) (A- LAP) (ARTS-1) (Aminopeptidase PILS) (Puromycin-insensitive leucyl- specific aminopeptidase) (PILS-AP) (VEGF-induced aminopeptidase). | |||||
|
AMPN_MOUSE
|
||||||
| NC score | 0.975744 (rank : 4) | θ value | 1.02182e-136 (rank : 4) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 34 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P97449, Q91YH8, Q99K63 | Gene names | Anpep, Lap-1, Lap1 | |||
|
Domain Architecture |

|
|||||
| Description | Aminopeptidase N (EC 3.4.11.2) (mAPN) (Alanyl aminopeptidase) (Microsomal aminopeptidase) (Aminopeptidase M) (Membrane protein p161) (CD13 antigen). | |||||
|
AMPN_HUMAN
|
||||||
| NC score | 0.975095 (rank : 5) | θ value | 4.44261e-132 (rank : 6) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P15144, Q16728, Q8IUK3, Q8IVH3, Q9UCE0 | Gene names | ANPEP, APN, CD13, PEPN | |||
|
Domain Architecture |

|
|||||
| Description | Aminopeptidase N (EC 3.4.11.2) (hAPN) (Alanyl aminopeptidase) (Microsomal aminopeptidase) (Aminopeptidase M) (gp150) (Myeloid plasma membrane glycoprotein CD13) (CD13 antigen). | |||||
|
AMPE_HUMAN
|
||||||
| NC score | 0.974423 (rank : 6) | θ value | 1.80368e-133 (rank : 5) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 46 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q07075 | Gene names | ENPEP | |||
|
Domain Architecture |

|
|||||
| Description | Glutamyl aminopeptidase (EC 3.4.11.7) (EAP) (Aminopeptidase A) (APA) (Differentiation antigen gp160) (CD249 antigen). | |||||
|
AMPE_MOUSE
|
||||||
| NC score | 0.973706 (rank : 7) | θ value | 9.26343e-130 (rank : 7) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P16406 | Gene names | Enpep | |||
|
Domain Architecture |

|
|||||
| Description | Glutamyl aminopeptidase (EC 3.4.11.7) (EAP) (Aminopeptidase A) (APA) (BP-1/6C3 antigen). | |||||
|
TRHDE_HUMAN
|
||||||
| NC score | 0.969045 (rank : 8) | θ value | 2.61056e-124 (rank : 9) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 49 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q9UKU6, Q6UWJ4 | Gene names | TRHDE | |||
|
Domain Architecture |

|
|||||
| Description | Thyrotropin-releasing hormone-degrading ectoenzyme (EC 3.4.19.6) (TRH- degrading ectoenzyme) (TRH-DE) (TRH-specific aminopeptidase) (Thyroliberinase) (Pyroglutamyl-peptidase II) (PAP-II). | |||||
|
PSA_HUMAN
|
||||||
| NC score | 0.967735 (rank : 9) | θ value | 1.34386e-112 (rank : 10) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P55786, Q6P145, Q9NP16, Q9UEM2 | Gene names | NPEPPS, PSA | |||
|
Domain Architecture |

|
|||||
| Description | Puromycin-sensitive aminopeptidase (EC 3.4.11.-) (PSA). | |||||
|
TRHDE_MOUSE
|
||||||
| NC score | 0.967636 (rank : 10) | θ value | 2.36115e-125 (rank : 8) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 58 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8K093 | Gene names | Trhde | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Thyrotropin-releasing hormone-degrading ectoenzyme (EC 3.4.19.6) (TRH- degrading ectoenzyme) (TRH-DE) (TRH-specific aminopeptidase) (Thyroliberinase) (Pyroglutamyl-peptidase II) (PAP-II). | |||||
|
PSA_MOUSE
|
||||||
| NC score | 0.967603 (rank : 11) | θ value | 1.13764e-111 (rank : 11) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q11011, Q91VJ8 | Gene names | Npepps, Psa | |||
|
Domain Architecture |

|
|||||
| Description | Puromycin-sensitive aminopeptidase (EC 3.4.11.-) (PSA). | |||||
|
LAEVR_HUMAN
|
||||||
| NC score | 0.961729 (rank : 12) | θ value | 2.62879e-100 (rank : 12) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 73 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q6Q4G3, Q32MR1, Q4G0I9, Q4G0V2, Q86XA3, Q8NBZ2 | Gene names | LVRN | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Laeverin (EC 3.4.-.-) (CHL2 antigen). | |||||
|
LAEVR_MOUSE
|
||||||
| NC score | 0.956069 (rank : 13) | θ value | 5.53243e-66 (rank : 13) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q2KHK3, Q9D633 | Gene names | Lvrn | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Laeverin (EC 3.4.-.-). | |||||
|
LKHA4_HUMAN
|
||||||
| NC score | 0.731280 (rank : 14) | θ value | 4.73814e-25 (rank : 14) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 24 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | P09960 | Gene names | LTA4H, LTA4 | |||
|
Domain Architecture |

|
|||||
| Description | Leukotriene A-4 hydrolase (EC 3.3.2.6) (LTA-4 hydrolase) (Leukotriene A(4) hydrolase). | |||||
|
LKHA4_MOUSE
|
||||||
| NC score | 0.724031 (rank : 15) | θ value | 3.75424e-22 (rank : 15) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 23 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | P24527, Q8VDR8 | Gene names | Lta4h | |||
|
Domain Architecture |

|
|||||
| Description | Leukotriene A-4 hydrolase (EC 3.3.2.6) (LTA-4 hydrolase) (Leukotriene A(4) hydrolase). | |||||
|
AMPB_MOUSE
|
||||||
| NC score | 0.691912 (rank : 16) | θ value | 8.65492e-19 (rank : 16) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 30 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8VCT3 | Gene names | Rnpep | |||
|
Domain Architecture |

|
|||||
| Description | Aminopeptidase B (EC 3.4.11.6) (Ap-B) (Arginyl aminopeptidase) (Arginine aminopeptidase) (Cytosol aminopeptidase IV). | |||||
|
AMPB_HUMAN
|
||||||
| NC score | 0.679070 (rank : 17) | θ value | 1.47631e-18 (rank : 17) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9H4A4, Q9BVM9, Q9H1D4, Q9NPT7 | Gene names | RNPEP, APB | |||
|
Domain Architecture |

|
|||||
| Description | Aminopeptidase B (EC 3.4.11.6) (Ap-B) (Arginyl aminopeptidase) (Arginine aminopeptidase). | |||||
|
RNPL1_HUMAN
|
||||||
| NC score | 0.650127 (rank : 18) | θ value | 2.28291e-11 (rank : 18) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 26 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q9HAU8, Q96AC9, Q9H033, Q9NVD0 | Gene names | RNPEPL1 | |||
|
Domain Architecture |

|
|||||
| Description | Arginyl aminopeptidase-like 1 (EC 3.4.11.-) (RNPEP-like protein). | |||||
|
AMPO_HUMAN
|
||||||
| NC score | 0.486437 (rank : 19) | θ value | 5.26297e-08 (rank : 20) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8N6M6, Q5T9B1, Q5T9B3, Q5T9B4, Q8WUL6, Q96M23, Q96SS1 | Gene names | ONPEP, C9orf3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Aminopeptidase O (EC 3.4.11.-) (AP-O). | |||||
|
AMPO_MOUSE
|
||||||
| NC score | 0.474836 (rank : 20) | θ value | 1.80886e-08 (rank : 19) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 25 | Shared Neighborhood Hits | 22 | |
| SwissProt Accessions | Q8BXQ6, Q6P0W6, Q6P394, Q8BHX5 | Gene names | Onpep | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Aminopeptidase O (EC 3.4.11.-) (AP-O). | |||||
|
TAF2_HUMAN
|
||||||
| NC score | 0.229379 (rank : 21) | θ value | 0.00390308 (rank : 21) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 36 | Shared Neighborhood Hits | 18 | |
| SwissProt Accessions | Q6P1X5, O43487, O43604, O60668, Q86WW7, Q8IWK4 | Gene names | TAF2, CIF150, TAF2B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID subunit 2 (Transcription initiation factor TFIID 150 kDa subunit) (TBP-associated factor 150 kDa) (TAFII-150) (TAFII150) (150 kDa cofactor of initiator function). | |||||
|
TAF2_MOUSE
|
||||||
| NC score | 0.221307 (rank : 22) | θ value | 0.00509761 (rank : 22) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 31 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | Q8C176, Q8BKQ7 | Gene names | Taf2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Transcription initiation factor TFIID subunit 2 (Transcription initiation factor TFIID 150 kDa subunit) (TBP-associated factor 150 kDa). | |||||
|
RPA2_HUMAN
|
||||||
| NC score | 0.030705 (rank : 23) | θ value | 0.125558 (rank : 23) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 8 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9H9Y6, Q585T5, Q6ZRR2, Q9H9D3 | Gene names | POLR1B | |||
|
Domain Architecture |

|
|||||
| Description | DNA-directed RNA polymerase I 135 kDa polypeptide (EC 2.7.7.6) (RNA polymerase I subunit 2) (RPA135). | |||||
|
RPA2_MOUSE
|
||||||
| NC score | 0.026308 (rank : 24) | θ value | 1.06291 (rank : 24) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | P70700 | Gene names | Polr1b, Rpa2, Rpo1-2 | |||
|
Domain Architecture |
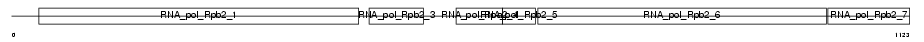
|
|||||
| Description | DNA-directed RNA polymerase I 135 kDa polypeptide (EC 2.7.7.6) (RNA polymerase I subunit 2) (RPA135). | |||||
|
PIGQ_MOUSE
|
||||||
| NC score | 0.009817 (rank : 25) | θ value | 6.88961 (rank : 27) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 5 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9QYT7, O35120, O35456, Q99L11 | Gene names | Pigq, Gpi1h, Mgpi1 | |||
|
Domain Architecture |

|
|||||
| Description | Phosphatidylinositol N-acetylglucosaminyltransferase subunit Q (EC 2.4.1.198) (Phosphatidylinositol-glycan biosynthesis class Q protein) (PIG-Q) (N-acetylglucosamyl transferase component GPI1) (MGpi1p). | |||||
|
RA6I1_MOUSE
|
||||||
| NC score | 0.007689 (rank : 26) | θ value | 5.27518 (rank : 26) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 57 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q6PAL8, Q62146, Q8C829, Q8VDF6, Q9QYZ2 | Gene names | Rab6ip1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Rab6-interacting protein 1 (Rab6IP1). | |||||
|
NU107_MOUSE
|
||||||
| NC score | 0.007014 (rank : 27) | θ value | 8.99809 (rank : 30) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 32 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q8BH74, Q99KH5 | Gene names | Nup107 | |||
|
Domain Architecture |

|
|||||
| Description | Nuclear pore complex protein Nup107 (Nucleoporin Nup107) (107 kDa nucleoporin). | |||||
|
RTN4_MOUSE
|
||||||
| NC score | 0.005313 (rank : 28) | θ value | 6.88961 (rank : 28) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 341 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q99P72, Q5DTK9, Q7TNB7, Q80W95, Q8BGK7, Q8BGM9, Q8K290, Q8K3G8, Q9CTE3 | Gene names | Rtn4, Kiaa0886, Nogo | |||
|
Domain Architecture |

|
|||||
| Description | Reticulon-4 (Neurite outgrowth inhibitor) (Nogo protein). | |||||
|
A1BG_HUMAN
|
||||||
| NC score | 0.005311 (rank : 29) | θ value | 8.99809 (rank : 29) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 94 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | P04217, Q8IYJ6, Q96P39 | Gene names | A1BG | |||
|
Domain Architecture |

|
|||||
| Description | Alpha-1B-glycoprotein precursor (Alpha-1-B glycoprotein). | |||||
|
PKHA6_HUMAN
|
||||||
| NC score | 0.004666 (rank : 30) | θ value | 5.27518 (rank : 25) | |||
| Query Neighborhood Hits | 30 | Target Neighborhood Hits | 556 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q9Y2H5 | Gene names | PLEKHA6, KIAA0969, PEPP3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Pleckstrin homology domain-containing family A member 6 (Phosphoinositol 3-phosphate-binding protein 3) (PEPP-3). | |||||