Please be patient as the page loads
|
LAP2B_HUMAN
|
||||||
| SwissProt Accessions | P42167, Q14861 | Gene names | TMPO, LAP2 | |||
|
Domain Architecture |

|
|||||
| Description | Lamina-associated polypeptide 2, isoforms beta/gamma (Thymopoietin, isoforms beta/gamma) (TP beta/gamma) (Thymopoietin-related peptide isoforms beta/gamma) (TPRP isoforms beta/gamma) [Contains: Thymopoietin (TP) (Splenin); Thymopentin (TP5)]. | |||||
| Rank Plots |
Jump to hits sorted by NC score
|
|||||
|
LAP2B_HUMAN
|
||||||
| θ value | 0 (rank : 1) | NC score | 0.999962 (rank : 1) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | P42167, Q14861 | Gene names | TMPO, LAP2 | |||
|
Domain Architecture |

|
|||||
| Description | Lamina-associated polypeptide 2, isoforms beta/gamma (Thymopoietin, isoforms beta/gamma) (TP beta/gamma) (Thymopoietin-related peptide isoforms beta/gamma) (TPRP isoforms beta/gamma) [Contains: Thymopoietin (TP) (Splenin); Thymopentin (TP5)]. | |||||
|
LAP2B_MOUSE
|
||||||
| θ value | 0 (rank : 2) | NC score | 0.988552 (rank : 2) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q61029, Q61030, Q61031, Q61032 | Gene names | Tmpo, Lap2 | |||
|
Domain Architecture |

|
|||||
| Description | Lamina-associated polypeptide 2 isoforms beta/delta/epsilon/gamma (Thymopoietin isoforms beta/delta/epsilon/gamma) (TP beta/delta/epsilon/gamma). | |||||
|
LAP2A_HUMAN
|
||||||
| θ value | 1.71185e-83 (rank : 3) | NC score | 0.913657 (rank : 3) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P42166, P08918, P08919, Q14860, Q16295 | Gene names | TMPO, LAP2 | |||
|
Domain Architecture |

|
|||||
| Description | Lamina-associated polypeptide 2 isoform alpha (Thymopoietin isoform alpha) (TP alpha) (Thymopoietin-related peptide isoform alpha) (TPRP isoform alpha) [Contains: Thymopoietin (TP) (Splenin); Thymopentin (TP5)]. | |||||
|
LAP2A_MOUSE
|
||||||
| θ value | 8.22888e-78 (rank : 4) | NC score | 0.911662 (rank : 4) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q61033, Q61028 | Gene names | Tmpo, Lap2 | |||
|
Domain Architecture |

|
|||||
| Description | Lamina-associated polypeptide 2 isoforms alpha/zeta (Thymopoietin isoforms alpha/zeta) (TP alpha/zeta). | |||||
|
MAP1B_MOUSE
|
||||||
| θ value | 0.00228821 (rank : 5) | NC score | 0.096881 (rank : 10) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 977 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P14873 | Gene names | Map1b, Mtap1b, Mtap5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) (MAP1.2) (MAP1(X)) [Contains: MAP1 light chain LC1]. | |||||
|
MAP1B_HUMAN
|
||||||
| θ value | 0.00390308 (rank : 6) | NC score | 0.098489 (rank : 9) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 903 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P46821 | Gene names | MAP1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) [Contains: MAP1 light chain LC1]. | |||||
|
EMD_MOUSE
|
||||||
| θ value | 0.00665767 (rank : 7) | NC score | 0.286373 (rank : 5) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O08579 | Gene names | Emd, Sta | |||
|
Domain Architecture |

|
|||||
| Description | Emerin. | |||||
|
EMD_HUMAN
|
||||||
| θ value | 0.0148317 (rank : 8) | NC score | 0.278834 (rank : 6) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P50402 | Gene names | EMD, EDMD, STA | |||
|
Domain Architecture |

|
|||||
| Description | Emerin. | |||||
|
CCNI_HUMAN
|
||||||
| θ value | 0.0330416 (rank : 9) | NC score | 0.125820 (rank : 8) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q14094 | Gene names | CCNI | |||
|
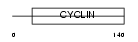
Domain Architecture |
|
|||||
| Description | Cyclin-I. | |||||
|
AP3B2_HUMAN
|
||||||
| θ value | 0.0563607 (rank : 10) | NC score | 0.063676 (rank : 14) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q13367, O14808 | Gene names | AP3B2 | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit beta-2 (Adapter-related protein complex 3 beta-2 subunit) (Beta3B-adaptin) (Adaptor protein complex AP-3 beta-2 subunit) (AP-3 complex beta-2 subunit) (Clathrin assembly protein complex 3 beta-2 large chain) (Neuron-specific vesicle coat protein beta-NAP). | |||||
|
CCD55_MOUSE
|
||||||
| θ value | 0.0736092 (rank : 11) | NC score | 0.059657 (rank : 15) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 519 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q5NCR9 | Gene names | Ccdc55 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 55. | |||||
|
MAN1_HUMAN
|
||||||
| θ value | 0.125558 (rank : 12) | NC score | 0.169262 (rank : 7) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y2U8, Q9NT47, Q9NYA5 | Gene names | LEMD3, MAN1 | |||
|
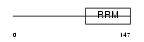
Domain Architecture |
|
|||||
| Description | Inner nuclear membrane protein Man1 (LEM domain-containing protein 3). | |||||
|
WAPL_MOUSE
|
||||||
| θ value | 0.163984 (rank : 13) | NC score | 0.051868 (rank : 16) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q65Z40, Q6ZQE9 | Gene names | Wapal, Kiaa0261, Wapl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wings apart-like protein homolog (Dioxin-inducible factor 2) (DIF-2). | |||||
|
FXR2_HUMAN
|
||||||
| θ value | 0.21417 (rank : 14) | NC score | 0.048554 (rank : 19) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P51116, Q8WUM2 | Gene names | FXR2 | |||
|
Domain Architecture |

|
|||||
| Description | Fragile X mental retardation syndrome-related protein 2. | |||||
|
FXR2_MOUSE
|
||||||
| θ value | 0.279714 (rank : 15) | NC score | 0.049033 (rank : 18) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9WVR4, Q9WVR5 | Gene names | Fxr2, Fxr2h | |||
|
Domain Architecture |

|
|||||
| Description | Fragile X mental retardation syndrome-related protein 2. | |||||
|
NOLC1_HUMAN
|
||||||
| θ value | 0.279714 (rank : 16) | NC score | 0.066320 (rank : 13) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q14978, Q15030 | Gene names | NOLC1, KIAA0035 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar phosphoprotein p130 (Nucleolar 130 kDa protein) (140 kDa nucleolar phosphoprotein) (Nopp140) (Nucleolar and coiled-body phosphoprotein 1). | |||||
|
SRRM2_HUMAN
|
||||||
| θ value | 0.365318 (rank : 17) | NC score | 0.042417 (rank : 20) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
ZFP37_HUMAN
|
||||||
| θ value | 0.365318 (rank : 18) | NC score | 0.005659 (rank : 50) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 797 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Y6Q3 | Gene names | ZFP37 | |||
|
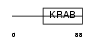
Domain Architecture |
|
|||||
| Description | Zinc finger protein 37 homolog (Zfp-37). | |||||
|
AP3B2_MOUSE
|
||||||
| θ value | 0.62314 (rank : 19) | NC score | 0.049290 (rank : 17) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9JME5, Q6QR53, Q8R1E5 | Gene names | Ap3b2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AP-3 complex subunit beta-2 (Adapter-related protein complex 3 beta-2 subunit) (Beta3B-adaptin) (Adaptor protein complex AP-3 beta-2 subunit) (Clathrin assembly protein complex 3 beta-2 large chain). | |||||
|
PTN13_MOUSE
|
||||||
| θ value | 0.62314 (rank : 20) | NC score | 0.017796 (rank : 36) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 426 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q64512, Q61494, Q62135, Q64499 | Gene names | Ptpn13, Ptp14 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 13 (EC 3.1.3.48) (Protein tyrosine phosphatase PTP-BL) (Protein-tyrosine phosphatase RIP) (protein tyrosine phosphatase DPZPTP) (PTP36). | |||||
|
CCNI_MOUSE
|
||||||
| θ value | 0.813845 (rank : 21) | NC score | 0.079573 (rank : 12) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Z2V9 | Gene names | Ccni | |||
|
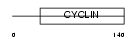
Domain Architecture |
|
|||||
| Description | Cyclin-I. | |||||
|
ARHGC_HUMAN
|
||||||
| θ value | 1.06291 (rank : 22) | NC score | 0.021481 (rank : 33) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NZN5, O15086 | Gene names | ARHGEF12, KIAA0382, LARG | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 12 (Leukemia-associated RhoGEF). | |||||
|
IWS1_MOUSE
|
||||||
| θ value | 1.06291 (rank : 23) | NC score | 0.038799 (rank : 22) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
AT2C2_HUMAN
|
||||||
| θ value | 1.81305 (rank : 24) | NC score | 0.024594 (rank : 31) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O75185 | Gene names | KIAA0703 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable calcium-transporting ATPase KIAA0703 (EC 3.6.3.8). | |||||
|
DEK_MOUSE
|
||||||
| θ value | 1.81305 (rank : 25) | NC score | 0.032197 (rank : 24) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q7TNV0, Q80VC5, Q8BZV6 | Gene names | Dek | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein DEK. | |||||
|
HEPH_HUMAN
|
||||||
| θ value | 1.81305 (rank : 26) | NC score | 0.028776 (rank : 28) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9BQS7, O75180, Q6UW45, Q9C058 | Gene names | HEPH, KIAA0698 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hephaestin precursor (EC 1.-.-.-). | |||||
|
HEPH_MOUSE
|
||||||
| θ value | 1.81305 (rank : 27) | NC score | 0.028562 (rank : 29) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Z0Z4, Q6ZQ65, Q80Y80, Q8C4S2 | Gene names | Heph, Kiaa0698 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hephaestin precursor (EC 1.-.-.-). | |||||
|
LMBL3_MOUSE
|
||||||
| θ value | 1.81305 (rank : 28) | NC score | 0.019846 (rank : 34) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8BLB7, Q641L7, Q6ZPI2, Q8C0G4 | Gene names | L3mbtl3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lethal(3)malignant brain tumor-like 3 protein (L(3)mbt-like 3 protein). | |||||
|
AHTF1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 29) | NC score | 0.031302 (rank : 26) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8WYP5, Q7Z4E3, Q8IZA4, Q96EH9, Q9Y4Q6 | Gene names | AHCTF1, ELYS, TMBS62 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-hook-containing transcription factor 1 (Embryonic large molecule derived from yolk sac). | |||||
|
MDC1_HUMAN
|
||||||
| θ value | 2.36792 (rank : 30) | NC score | 0.025722 (rank : 30) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
ZN592_HUMAN
|
||||||
| θ value | 2.36792 (rank : 31) | NC score | 0.006722 (rank : 48) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 762 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q92610 | Gene names | ZNF592, KIAA0211 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 592. | |||||
|
CHD9_HUMAN
|
||||||
| θ value | 3.0926 (rank : 32) | NC score | 0.015595 (rank : 41) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q3L8U1, O15025, Q1WF12, Q6DTK9, Q9H9V7 | Gene names | CHD9, KIAA0308, KISH2, PRIC320 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain-helicase-DNA-binding protein 9 (EC 3.6.1.-) (ATP- dependent helicase CHD9) (CHD-9) (Chromatin-related mesenchymal modulator) (CReMM) (Chromatin remodeling factor CHROM1) (Peroxisomal proliferator-activated receptor A-interacting complex 320 kDa protein) (PPAR-alpha-interacting complex protein 320 kDa) (Kismet homolog 2). | |||||
|
DMP1_MOUSE
|
||||||
| θ value | 3.0926 (rank : 33) | NC score | 0.040745 (rank : 21) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O55188 | Gene names | Dmp1, Dmp | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1) (AG1). | |||||
|
LARP1_HUMAN
|
||||||
| θ value | 3.0926 (rank : 34) | NC score | 0.079794 (rank : 11) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q6PKG0, O94836, Q8N4M2, Q8NB73, Q9UFD7 | Gene names | LARP1, KIAA0731, LARP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | La-related protein 1 (La ribonucleoprotein domain family member 1). | |||||
|
ANK2_HUMAN
|
||||||
| θ value | 4.03905 (rank : 35) | NC score | 0.016032 (rank : 39) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
ARHG1_MOUSE
|
||||||
| θ value | 4.03905 (rank : 36) | NC score | 0.018564 (rank : 35) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q61210, O89074, Q80YE8, Q80YE9, Q91VL3 | Gene names | Arhgef1, Lbcl2, Lsc | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 1 (Lymphoid blast crisis-like 2) (Lbc's second cousin). | |||||
|
JADE2_MOUSE
|
||||||
| θ value | 4.03905 (rank : 37) | NC score | 0.011910 (rank : 44) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 560 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6ZQF7, Q3UHD5, Q6IE83 | Gene names | Phf15, Jade2, Kiaa0239 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Jade-2 (PHD finger protein 15). | |||||
|
AFF1_HUMAN
|
||||||
| θ value | 5.27518 (rank : 38) | NC score | 0.031338 (rank : 25) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P51825 | Gene names | AFF1, AF4, FEL, MLLT2 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 1 (Protein AF-4) (Proto-oncogene AF4) (Protein FEL). | |||||
|
NFM_MOUSE
|
||||||
| θ value | 5.27518 (rank : 39) | NC score | 0.006024 (rank : 49) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1159 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P08553, Q61961 | Gene names | Nef3, Nefm, Nfm | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet M protein (160 kDa neurofilament protein) (Neurofilament medium polypeptide) (NF-M). | |||||
|
RBM28_HUMAN
|
||||||
| θ value | 5.27518 (rank : 40) | NC score | 0.010533 (rank : 45) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9NW13, Q96CV3 | Gene names | RBM28 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein 28 (RNA-binding motif protein 28). | |||||
|
SRRM2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 41) | NC score | 0.034026 (rank : 23) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
TAOK2_MOUSE
|
||||||
| θ value | 5.27518 (rank : 42) | NC score | 0.000434 (rank : 52) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1373 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6ZQ29, Q7TSS8 | Gene names | Taok2, Kiaa0881 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase TAO2 (EC 2.7.11.1) (Thousand and one amino acid protein 2). | |||||
|
AFF1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 43) | NC score | 0.023884 (rank : 32) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O88573 | Gene names | Aff1, Mllt2, Mllt2h | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 1 (Protein AF-4) (Proto-oncogene AF4). | |||||
|
CHD1_HUMAN
|
||||||
| θ value | 6.88961 (rank : 44) | NC score | 0.014882 (rank : 42) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O14646 | Gene names | CHD1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
CHD1_MOUSE
|
||||||
| θ value | 6.88961 (rank : 45) | NC score | 0.016372 (rank : 37) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P40201 | Gene names | Chd1, Chd-1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
DEN2A_MOUSE
|
||||||
| θ value | 6.88961 (rank : 46) | NC score | 0.008654 (rank : 47) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8C4S8, Q3TJU2, Q3TUG3, Q3UXA3 | Gene names | Dennd2a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DENN domain-containing protein 2A. | |||||
|
MYO3A_HUMAN
|
||||||
| θ value | 6.88961 (rank : 47) | NC score | 0.000200 (rank : 53) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1033 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8NEV4, Q8WX17, Q9NYS8 | Gene names | MYO3A | |||
|
Domain Architecture |

|
|||||
| Description | Myosin IIIA (EC 2.7.11.1). | |||||
|
TGON2_HUMAN
|
||||||
| θ value | 6.88961 (rank : 48) | NC score | 0.030387 (rank : 27) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1037 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O43493, O15282, O43492, O43499, O43500, O43501, Q92760 | Gene names | TGOLN2, TGN46, TGN51 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (Trans-Golgi network protein TGN51) (TGN46) (TGN48) (TGN38 homolog). | |||||
|
ARS2_HUMAN
|
||||||
| θ value | 8.99809 (rank : 49) | NC score | 0.016223 (rank : 38) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 186 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9BXP5, O95808, Q9BWP6, Q9BXP4 | Gene names | ARS2, ASR2 | |||
|
Domain Architecture |
|
|||||
| Description | Arsenite-resistance protein 2. | |||||
|
PPIG_HUMAN
|
||||||
| θ value | 8.99809 (rank : 50) | NC score | 0.014548 (rank : 43) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q13427, O00706, Q96DG9 | Gene names | PPIG | |||
|
Domain Architecture |

|
|||||
| Description | Peptidyl-prolyl cis-trans isomerase G (EC 5.2.1.8) (Peptidyl-prolyl isomerase G) (PPIase G) (Rotamase G) (Cyclophilin G) (Clk-associating RS-cyclophilin) (CARS-cyclophilin) (CARS-Cyp) (SR-cyclophilin) (SRcyp) (SR-cyp) (CASP10). | |||||
|
RERE_HUMAN
|
||||||
| θ value | 8.99809 (rank : 51) | NC score | 0.016028 (rank : 40) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 779 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9P2R6, O43393, O75046, O75359, Q5VXL9, Q6P6B9, Q9Y2W4 | Gene names | RERE, ARG, ARP, ATN1L, KIAA0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-1-like protein) (Atrophin-1-related protein). | |||||
|
SYTL5_MOUSE
|
||||||
| θ value | 8.99809 (rank : 52) | NC score | 0.005086 (rank : 51) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q80T23, Q812E5 | Gene names | Sytl5, Slp5 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-like protein 5. | |||||
|
TXLNB_HUMAN
|
||||||
| θ value | 8.99809 (rank : 53) | NC score | 0.009408 (rank : 46) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 912 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8N3L3, Q76L25, Q86T52, Q8N3S2 | Gene names | TXLNB, C6orf198, MDP77 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-taxilin (Muscle-derived protein 77) (hMDP77). | |||||
|
LAP2B_HUMAN
|
||||||
| NC score | 0.999962 (rank : 1) | θ value | 0 (rank : 1) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 53 | |
| SwissProt Accessions | P42167, Q14861 | Gene names | TMPO, LAP2 | |||
|
Domain Architecture |

|
|||||
| Description | Lamina-associated polypeptide 2, isoforms beta/gamma (Thymopoietin, isoforms beta/gamma) (TP beta/gamma) (Thymopoietin-related peptide isoforms beta/gamma) (TPRP isoforms beta/gamma) [Contains: Thymopoietin (TP) (Splenin); Thymopentin (TP5)]. | |||||
|
LAP2B_MOUSE
|
||||||
| NC score | 0.988552 (rank : 2) | θ value | 0 (rank : 2) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 28 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q61029, Q61030, Q61031, Q61032 | Gene names | Tmpo, Lap2 | |||
|
Domain Architecture |

|
|||||
| Description | Lamina-associated polypeptide 2 isoforms beta/delta/epsilon/gamma (Thymopoietin isoforms beta/delta/epsilon/gamma) (TP beta/delta/epsilon/gamma). | |||||
|
LAP2A_HUMAN
|
||||||
| NC score | 0.913657 (rank : 3) | θ value | 1.71185e-83 (rank : 3) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 29 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | P42166, P08918, P08919, Q14860, Q16295 | Gene names | TMPO, LAP2 | |||
|
Domain Architecture |

|
|||||
| Description | Lamina-associated polypeptide 2 isoform alpha (Thymopoietin isoform alpha) (TP alpha) (Thymopoietin-related peptide isoform alpha) (TPRP isoform alpha) [Contains: Thymopoietin (TP) (Splenin); Thymopentin (TP5)]. | |||||
|
LAP2A_MOUSE
|
||||||
| NC score | 0.911662 (rank : 4) | θ value | 8.22888e-78 (rank : 4) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 38 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q61033, Q61028 | Gene names | Tmpo, Lap2 | |||
|
Domain Architecture |

|
|||||
| Description | Lamina-associated polypeptide 2 isoforms alpha/zeta (Thymopoietin isoforms alpha/zeta) (TP alpha/zeta). | |||||
|
EMD_MOUSE
|
||||||
| NC score | 0.286373 (rank : 5) | θ value | 0.00665767 (rank : 7) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 9 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | O08579 | Gene names | Emd, Sta | |||
|
Domain Architecture |

|
|||||
| Description | Emerin. | |||||
|
EMD_HUMAN
|
||||||
| NC score | 0.278834 (rank : 6) | θ value | 0.0148317 (rank : 8) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 11 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | P50402 | Gene names | EMD, EDMD, STA | |||
|
Domain Architecture |

|
|||||
| Description | Emerin. | |||||
|
MAN1_HUMAN
|
||||||
| NC score | 0.169262 (rank : 7) | θ value | 0.125558 (rank : 12) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 37 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9Y2U8, Q9NT47, Q9NYA5 | Gene names | LEMD3, MAN1 | |||
|
Domain Architecture |

|
|||||
| Description | Inner nuclear membrane protein Man1 (LEM domain-containing protein 3). | |||||
|
CCNI_HUMAN
|
||||||
| NC score | 0.125820 (rank : 8) | θ value | 0.0330416 (rank : 9) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 39 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q14094 | Gene names | CCNI | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin-I. | |||||
|
MAP1B_HUMAN
|
||||||
| NC score | 0.098489 (rank : 9) | θ value | 0.00390308 (rank : 6) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 903 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | P46821 | Gene names | MAP1B | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) [Contains: MAP1 light chain LC1]. | |||||
|
MAP1B_MOUSE
|
||||||
| NC score | 0.096881 (rank : 10) | θ value | 0.00228821 (rank : 5) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 977 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | P14873 | Gene names | Map1b, Mtap1b, Mtap5 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Microtubule-associated protein 1B (MAP 1B) (MAP1.2) (MAP1(X)) [Contains: MAP1 light chain LC1]. | |||||
|
LARP1_HUMAN
|
||||||
| NC score | 0.079794 (rank : 11) | θ value | 3.0926 (rank : 34) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 242 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q6PKG0, O94836, Q8N4M2, Q8NB73, Q9UFD7 | Gene names | LARP1, KIAA0731, LARP | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | La-related protein 1 (La ribonucleoprotein domain family member 1). | |||||
|
CCNI_MOUSE
|
||||||
| NC score | 0.079573 (rank : 12) | θ value | 0.813845 (rank : 21) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 35 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Z2V9 | Gene names | Ccni | |||
|
Domain Architecture |

|
|||||
| Description | Cyclin-I. | |||||
|
NOLC1_HUMAN
|
||||||
| NC score | 0.066320 (rank : 13) | θ value | 0.279714 (rank : 16) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 452 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | Q14978, Q15030 | Gene names | NOLC1, KIAA0035 | |||
|
Domain Architecture |

|
|||||
| Description | Nucleolar phosphoprotein p130 (Nucleolar 130 kDa protein) (140 kDa nucleolar phosphoprotein) (Nopp140) (Nucleolar and coiled-body phosphoprotein 1). | |||||
|
AP3B2_HUMAN
|
||||||
| NC score | 0.063676 (rank : 14) | θ value | 0.0563607 (rank : 10) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 15 | |
| SwissProt Accessions | Q13367, O14808 | Gene names | AP3B2 | |||
|
Domain Architecture |

|
|||||
| Description | AP-3 complex subunit beta-2 (Adapter-related protein complex 3 beta-2 subunit) (Beta3B-adaptin) (Adaptor protein complex AP-3 beta-2 subunit) (AP-3 complex beta-2 subunit) (Clathrin assembly protein complex 3 beta-2 large chain) (Neuron-specific vesicle coat protein beta-NAP). | |||||
|
CCD55_MOUSE
|
||||||
| NC score | 0.059657 (rank : 15) | θ value | 0.0736092 (rank : 11) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 519 | Shared Neighborhood Hits | 19 | |
| SwissProt Accessions | Q5NCR9 | Gene names | Ccdc55 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Coiled-coil domain-containing protein 55. | |||||
|
WAPL_MOUSE
|
||||||
| NC score | 0.051868 (rank : 16) | θ value | 0.163984 (rank : 13) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 158 | Shared Neighborhood Hits | 10 | |
| SwissProt Accessions | Q65Z40, Q6ZQE9 | Gene names | Wapal, Kiaa0261, Wapl | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Wings apart-like protein homolog (Dioxin-inducible factor 2) (DIF-2). | |||||
|
AP3B2_MOUSE
|
||||||
| NC score | 0.049290 (rank : 17) | θ value | 0.62314 (rank : 19) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 246 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9JME5, Q6QR53, Q8R1E5 | Gene names | Ap3b2 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AP-3 complex subunit beta-2 (Adapter-related protein complex 3 beta-2 subunit) (Beta3B-adaptin) (Adaptor protein complex AP-3 beta-2 subunit) (Clathrin assembly protein complex 3 beta-2 large chain). | |||||
|
FXR2_MOUSE
|
||||||
| NC score | 0.049033 (rank : 18) | θ value | 0.279714 (rank : 15) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 66 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9WVR4, Q9WVR5 | Gene names | Fxr2, Fxr2h | |||
|
Domain Architecture |

|
|||||
| Description | Fragile X mental retardation syndrome-related protein 2. | |||||
|
FXR2_HUMAN
|
||||||
| NC score | 0.048554 (rank : 19) | θ value | 0.21417 (rank : 14) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 77 | Shared Neighborhood Hits | 4 | |
| SwissProt Accessions | P51116, Q8WUM2 | Gene names | FXR2 | |||
|
Domain Architecture |

|
|||||
| Description | Fragile X mental retardation syndrome-related protein 2. | |||||
|
SRRM2_HUMAN
|
||||||
| NC score | 0.042417 (rank : 20) | θ value | 0.365318 (rank : 17) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1296 | Shared Neighborhood Hits | 24 | |
| SwissProt Accessions | Q9UQ35, O15038, O94803, Q6NSL3, Q6PIM3, Q6PK40, Q8IW17, Q96GY7, Q9P0G1, Q9UHA8, Q9UQ36, Q9UQ37, Q9UQ38, Q9UQ40 | Gene names | SRRM2, KIAA0324, SRL300, SRM300 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2 (Serine/arginine-rich splicing factor-related nuclear matrix protein of 300 kDa) (Ser/Arg- related nuclear matrix protein) (SR-related nuclear matrix protein of 300 kDa) (Splicing coactivator subunit SRm300) (300 kDa nuclear matrix antigen). | |||||
|
DMP1_MOUSE
|
||||||
| NC score | 0.040745 (rank : 21) | θ value | 3.0926 (rank : 33) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 498 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | O55188 | Gene names | Dmp1, Dmp | |||
|
Domain Architecture |

|
|||||
| Description | Dentin matrix acidic phosphoprotein 1 precursor (Dentin matrix protein 1) (DMP-1) (AG1). | |||||
|
IWS1_MOUSE
|
||||||
| NC score | 0.038799 (rank : 22) | θ value | 1.06291 (rank : 23) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1258 | Shared Neighborhood Hits | 30 | |
| SwissProt Accessions | Q8C1D8, Q3TQ25, Q3UWB0, Q6NVD1, Q8BY73, Q9D9H4 | Gene names | Iws1, Iws1l | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | IWS1 homolog (IWS1-like protein). | |||||
|
SRRM2_MOUSE
|
||||||
| NC score | 0.034026 (rank : 23) | θ value | 5.27518 (rank : 41) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1470 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q8BTI8, Q3TBY5, Q569P9, Q80U37, Q8K383 | Gene names | Srrm2, Kiaa0324 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/arginine repetitive matrix protein 2. | |||||
|
DEK_MOUSE
|
||||||
| NC score | 0.032197 (rank : 24) | θ value | 1.81305 (rank : 25) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 280 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q7TNV0, Q80VC5, Q8BZV6 | Gene names | Dek | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein DEK. | |||||
|
AFF1_HUMAN
|
||||||
| NC score | 0.031338 (rank : 25) | θ value | 5.27518 (rank : 38) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 506 | Shared Neighborhood Hits | 17 | |
| SwissProt Accessions | P51825 | Gene names | AFF1, AF4, FEL, MLLT2 | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 1 (Protein AF-4) (Proto-oncogene AF4) (Protein FEL). | |||||
|
AHTF1_HUMAN
|
||||||
| NC score | 0.031302 (rank : 26) | θ value | 2.36792 (rank : 29) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 286 | Shared Neighborhood Hits | 13 | |
| SwissProt Accessions | Q8WYP5, Q7Z4E3, Q8IZA4, Q96EH9, Q9Y4Q6 | Gene names | AHCTF1, ELYS, TMBS62 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | AT-hook-containing transcription factor 1 (Embryonic large molecule derived from yolk sac). | |||||
|
TGON2_HUMAN
|
||||||
| NC score | 0.030387 (rank : 27) | θ value | 6.88961 (rank : 48) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1037 | Shared Neighborhood Hits | 31 | |
| SwissProt Accessions | O43493, O15282, O43492, O43499, O43500, O43501, Q92760 | Gene names | TGOLN2, TGN46, TGN51 | |||
|
Domain Architecture |

|
|||||
| Description | Trans-Golgi network integral membrane protein 2 precursor (Trans-Golgi network protein TGN51) (TGN46) (TGN48) (TGN38 homolog). | |||||
|
HEPH_HUMAN
|
||||||
| NC score | 0.028776 (rank : 28) | θ value | 1.81305 (rank : 26) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 19 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9BQS7, O75180, Q6UW45, Q9C058 | Gene names | HEPH, KIAA0698 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hephaestin precursor (EC 1.-.-.-). | |||||
|
HEPH_MOUSE
|
||||||
| NC score | 0.028562 (rank : 29) | θ value | 1.81305 (rank : 27) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 17 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q9Z0Z4, Q6ZQ65, Q80Y80, Q8C4S2 | Gene names | Heph, Kiaa0698 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Hephaestin precursor (EC 1.-.-.-). | |||||
|
MDC1_HUMAN
|
||||||
| NC score | 0.025722 (rank : 30) | θ value | 2.36792 (rank : 30) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1446 | Shared Neighborhood Hits | 21 | |
| SwissProt Accessions | Q14676, Q5JP55, Q5JP56, Q5ST83, Q68CQ3, Q86Z06, Q96QC2 | Gene names | MDC1, KIAA0170, NFBD1 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Mediator of DNA damage checkpoint protein 1 (Nuclear factor with BRCT domains 1). | |||||
|
AT2C2_HUMAN
|
||||||
| NC score | 0.024594 (rank : 31) | θ value | 1.81305 (rank : 24) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 67 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | O75185 | Gene names | KIAA0703 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Probable calcium-transporting ATPase KIAA0703 (EC 3.6.3.8). | |||||
|
AFF1_MOUSE
|
||||||
| NC score | 0.023884 (rank : 32) | θ value | 6.88961 (rank : 43) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | O88573 | Gene names | Aff1, Mllt2, Mllt2h | |||
|
Domain Architecture |

|
|||||
| Description | AF4/FMR2 family member 1 (Protein AF-4) (Proto-oncogene AF4). | |||||
|
ARHGC_HUMAN
|
||||||
| NC score | 0.021481 (rank : 33) | θ value | 1.06291 (rank : 22) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 351 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q9NZN5, O15086 | Gene names | ARHGEF12, KIAA0382, LARG | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 12 (Leukemia-associated RhoGEF). | |||||
|
LMBL3_MOUSE
|
||||||
| NC score | 0.019846 (rank : 34) | θ value | 1.81305 (rank : 28) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 212 | Shared Neighborhood Hits | 8 | |
| SwissProt Accessions | Q8BLB7, Q641L7, Q6ZPI2, Q8C0G4 | Gene names | L3mbtl3 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Lethal(3)malignant brain tumor-like 3 protein (L(3)mbt-like 3 protein). | |||||
|
ARHG1_MOUSE
|
||||||
| NC score | 0.018564 (rank : 35) | θ value | 4.03905 (rank : 36) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q61210, O89074, Q80YE8, Q80YE9, Q91VL3 | Gene names | Arhgef1, Lbcl2, Lsc | |||
|
Domain Architecture |

|
|||||
| Description | Rho guanine nucleotide exchange factor 1 (Lymphoid blast crisis-like 2) (Lbc's second cousin). | |||||
|
PTN13_MOUSE
|
||||||
| NC score | 0.017796 (rank : 36) | θ value | 0.62314 (rank : 20) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 426 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | Q64512, Q61494, Q62135, Q64499 | Gene names | Ptpn13, Ptp14 | |||
|
Domain Architecture |

|
|||||
| Description | Tyrosine-protein phosphatase non-receptor type 13 (EC 3.1.3.48) (Protein tyrosine phosphatase PTP-BL) (Protein-tyrosine phosphatase RIP) (protein tyrosine phosphatase DPZPTP) (PTP36). | |||||
|
CHD1_MOUSE
|
||||||
| NC score | 0.016372 (rank : 37) | θ value | 6.88961 (rank : 45) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 266 | Shared Neighborhood Hits | 11 | |
| SwissProt Accessions | P40201 | Gene names | Chd1, Chd-1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
ARS2_HUMAN
|
||||||
| NC score | 0.016223 (rank : 38) | θ value | 8.99809 (rank : 49) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 186 | Shared Neighborhood Hits | 9 | |
| SwissProt Accessions | Q9BXP5, O95808, Q9BWP6, Q9BXP4 | Gene names | ARS2, ASR2 | |||
|
Domain Architecture |

|
|||||
| Description | Arsenite-resistance protein 2. | |||||
|
ANK2_HUMAN
|
||||||
| NC score | 0.016032 (rank : 39) | θ value | 4.03905 (rank : 35) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1615 | Shared Neighborhood Hits | 23 | |
| SwissProt Accessions | Q01484, Q01485 | Gene names | ANK2 | |||
|
Domain Architecture |

|
|||||
| Description | Ankyrin-2 (Brain ankyrin) (Ankyrin-B) (Ankyrin, nonerythroid). | |||||
|
RERE_HUMAN
|
||||||
| NC score | 0.016028 (rank : 40) | θ value | 8.99809 (rank : 51) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 779 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q9P2R6, O43393, O75046, O75359, Q5VXL9, Q6P6B9, Q9Y2W4 | Gene names | RERE, ARG, ARP, ATN1L, KIAA0458 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Arginine-glutamic acid dipeptide repeats protein (Atrophin-1-like protein) (Atrophin-1-related protein). | |||||
|
CHD9_HUMAN
|
||||||
| NC score | 0.015595 (rank : 41) | θ value | 3.0926 (rank : 32) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 343 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q3L8U1, O15025, Q1WF12, Q6DTK9, Q9H9V7 | Gene names | CHD9, KIAA0308, KISH2, PRIC320 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Chromodomain-helicase-DNA-binding protein 9 (EC 3.6.1.-) (ATP- dependent helicase CHD9) (CHD-9) (Chromatin-related mesenchymal modulator) (CReMM) (Chromatin remodeling factor CHROM1) (Peroxisomal proliferator-activated receptor A-interacting complex 320 kDa protein) (PPAR-alpha-interacting complex protein 320 kDa) (Kismet homolog 2). | |||||
|
CHD1_HUMAN
|
||||||
| NC score | 0.014882 (rank : 42) | θ value | 6.88961 (rank : 44) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 337 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | O14646 | Gene names | CHD1 | |||
|
Domain Architecture |

|
|||||
| Description | Chromodomain-helicase-DNA-binding protein 1 (EC 3.6.1.-) (ATP- dependent helicase CHD1) (CHD-1). | |||||
|
PPIG_HUMAN
|
||||||
| NC score | 0.014548 (rank : 43) | θ value | 8.99809 (rank : 50) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 670 | Shared Neighborhood Hits | 20 | |
| SwissProt Accessions | Q13427, O00706, Q96DG9 | Gene names | PPIG | |||
|
Domain Architecture |

|
|||||
| Description | Peptidyl-prolyl cis-trans isomerase G (EC 5.2.1.8) (Peptidyl-prolyl isomerase G) (PPIase G) (Rotamase G) (Cyclophilin G) (Clk-associating RS-cyclophilin) (CARS-cyclophilin) (CARS-Cyp) (SR-cyclophilin) (SRcyp) (SR-cyp) (CASP10). | |||||
|
JADE2_MOUSE
|
||||||
| NC score | 0.011910 (rank : 44) | θ value | 4.03905 (rank : 37) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 560 | Shared Neighborhood Hits | 12 | |
| SwissProt Accessions | Q6ZQF7, Q3UHD5, Q6IE83 | Gene names | Phf15, Jade2, Kiaa0239 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Protein Jade-2 (PHD finger protein 15). | |||||
|
RBM28_HUMAN
|
||||||
| NC score | 0.010533 (rank : 45) | θ value | 5.27518 (rank : 40) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 307 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q9NW13, Q96CV3 | Gene names | RBM28 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | RNA-binding protein 28 (RNA-binding motif protein 28). | |||||
|
TXLNB_HUMAN
|
||||||
| NC score | 0.009408 (rank : 46) | θ value | 8.99809 (rank : 53) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 912 | Shared Neighborhood Hits | 14 | |
| SwissProt Accessions | Q8N3L3, Q76L25, Q86T52, Q8N3S2 | Gene names | TXLNB, C6orf198, MDP77 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Beta-taxilin (Muscle-derived protein 77) (hMDP77). | |||||
|
DEN2A_MOUSE
|
||||||
| NC score | 0.008654 (rank : 47) | θ value | 6.88961 (rank : 46) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 53 | Shared Neighborhood Hits | 3 | |
| SwissProt Accessions | Q8C4S8, Q3TJU2, Q3TUG3, Q3UXA3 | Gene names | Dennd2a | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | DENN domain-containing protein 2A. | |||||
|
ZN592_HUMAN
|
||||||
| NC score | 0.006722 (rank : 48) | θ value | 2.36792 (rank : 31) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 762 | Shared Neighborhood Hits | 5 | |
| SwissProt Accessions | Q92610 | Gene names | ZNF592, KIAA0211 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Zinc finger protein 592. | |||||
|
NFM_MOUSE
|
||||||
| NC score | 0.006024 (rank : 49) | θ value | 5.27518 (rank : 39) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1159 | Shared Neighborhood Hits | 16 | |
| SwissProt Accessions | P08553, Q61961 | Gene names | Nef3, Nefm, Nfm | |||
|
Domain Architecture |

|
|||||
| Description | Neurofilament triplet M protein (160 kDa neurofilament protein) (Neurofilament medium polypeptide) (NF-M). | |||||
|
ZFP37_HUMAN
|
||||||
| NC score | 0.005659 (rank : 50) | θ value | 0.365318 (rank : 18) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 797 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q9Y6Q3 | Gene names | ZFP37 | |||
|
Domain Architecture |

|
|||||
| Description | Zinc finger protein 37 homolog (Zfp-37). | |||||
|
SYTL5_MOUSE
|
||||||
| NC score | 0.005086 (rank : 51) | θ value | 8.99809 (rank : 52) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 154 | Shared Neighborhood Hits | 2 | |
| SwissProt Accessions | Q80T23, Q812E5 | Gene names | Sytl5, Slp5 | |||
|
Domain Architecture |

|
|||||
| Description | Synaptotagmin-like protein 5. | |||||
|
TAOK2_MOUSE
|
||||||
| NC score | 0.000434 (rank : 52) | θ value | 5.27518 (rank : 42) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1373 | Shared Neighborhood Hits | 7 | |
| SwissProt Accessions | Q6ZQ29, Q7TSS8 | Gene names | Taok2, Kiaa0881 | |||
|
Domain Architecture |
No domain graphic available |
|||||
| Description | Serine/threonine-protein kinase TAO2 (EC 2.7.11.1) (Thousand and one amino acid protein 2). | |||||
|
MYO3A_HUMAN
|
||||||
| NC score | 0.000200 (rank : 53) | θ value | 6.88961 (rank : 47) | |||
| Query Neighborhood Hits | 53 | Target Neighborhood Hits | 1033 | Shared Neighborhood Hits | 6 | |
| SwissProt Accessions | Q8NEV4, Q8WX17, Q9NYS8 | Gene names | MYO3A | |||
|
Domain Architecture |

|
|||||
| Description | Myosin IIIA (EC 2.7.11.1). | |||||